

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147434.11 - phase: 0
(150 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
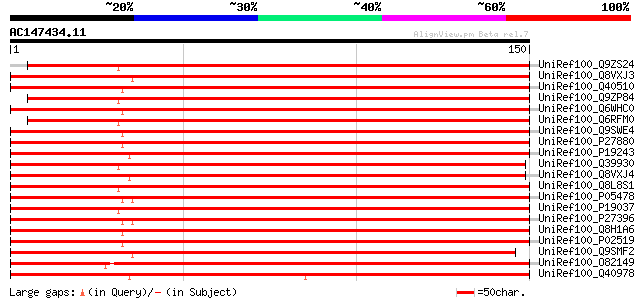
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9ZS24 Cytosolic class I small heat-shock protein HSP1... 243 5e-64
UniRef100_Q8VXJ3 17.6 kDa heat-shock protein [Helianthus annuus] 243 6e-64
UniRef100_Q40510 Nthsp18p [Nicotiana tabacum] 243 8e-64
UniRef100_Q9ZP84 Heat shock protein 17.4 [Quercus suber] 242 1e-63
UniRef100_Q6WHC0 Chloroplast small heat shock protein class I [C... 241 3e-63
UniRef100_Q6RFM0 Small heat shock protein [Prunus persica] 240 5e-63
UniRef100_Q9SWE4 Low molecular weight heat-shock protein [Nicoti... 239 1e-62
UniRef100_P27880 18.2 kDa class I heat shock protein [Medicago s... 239 1e-62
UniRef100_P19243 18.1 kDa class I heat shock protein [Pisum sati... 239 1e-62
UniRef100_Q39930 17.7 kDa heat shock protein [Helianthus annuus] 237 3e-62
UniRef100_Q8VXJ4 20.5 kDa heat-shock protein [Helianthus annuus] 237 4e-62
UniRef100_Q8L8S1 Heat shock protein 18 [Arabidopsis thaliana] 237 4e-62
UniRef100_P05478 18.5 kDa class I heat shock protein [Glycine max] 236 6e-62
UniRef100_P19037 18.2 kDa class I heat shock protein [Arabidopsi... 236 6e-62
UniRef100_P27396 17.8 kDa class I heat shock protein [Daucus car... 236 6e-62
UniRef100_Q8H1A6 Heat shock protein [Pisum sativum] 235 1e-61
UniRef100_P02519 17.3 kDa class I heat shock protein [Glycine max] 233 6e-61
UniRef100_Q9SMF2 17.9 kDa heat-shock protein [Helianthus annuus] 233 8e-61
UniRef100_O82149 Low-molecular-weight heat shock protein [Cuscut... 233 8e-61
UniRef100_Q40978 Low molecular weight heat-shock protein [Papave... 232 1e-60
>UniRef100_Q9ZS24 Cytosolic class I small heat-shock protein HSP17.5 [Castanea
sativa]
Length = 154
Score = 243 bits (621), Expect = 5e-64
Identities = 117/151 (77%), Positives = 131/151 (86%), Gaps = 6/151 (3%)
Query: 6 SFFGGRQNNVFDPFSMDIWDPLQGF------PSSARETTALANTRVDWKETQEAHVFSVD 59
S FGGR++NVFDPFS+DIWDP +GF P SARETTA A R+DWKET EAH+F D
Sbjct: 4 SLFGGRRSNVFDPFSLDIWDPFEGFSAVANVPPSARETTAFATARIDWKETPEAHIFKAD 63
Query: 60 LPGLKKEEVKVEIEDGNVLQISGERNKEQEEKDDKWHRVERSSGKFMRRFRLPENVKMDQ 119
LPGLKKEEVKVE+EDGNVLQISGER+KE EEK+DKWHRVERS GKF+RRFRLPEN K++Q
Sbjct: 64 LPGLKKEEVKVEVEDGNVLQISGERSKEHEEKNDKWHRVERSCGKFLRRFRLPENAKVEQ 123
Query: 120 VKAGMENGVLTVTVPKEEEKKSEVKSIEISG 150
VKA MENGVLTV VPKEE+KK+EVKSIEISG
Sbjct: 124 VKANMENGVLTVIVPKEEQKKTEVKSIEISG 154
>UniRef100_Q8VXJ3 17.6 kDa heat-shock protein [Helianthus annuus]
Length = 155
Score = 243 bits (620), Expect = 6e-64
Identities = 116/155 (74%), Positives = 134/155 (85%), Gaps = 5/155 (3%)
Query: 1 MSLIPSFFGGRQNNVFDPFSMDIWDPLQGFPSSA-----RETTALANTRVDWKETQEAHV 55
MS+IPS F GR+++VFDPFS+D+WDP + FP S+ RET+AL N RVDWKET EAHV
Sbjct: 1 MSIIPSLFAGRRSSVFDPFSLDVWDPFRDFPISSSSDVSRETSALVNARVDWKETPEAHV 60
Query: 56 FSVDLPGLKKEEVKVEIEDGNVLQISGERNKEQEEKDDKWHRVERSSGKFMRRFRLPENV 115
F DLPG+KKEEVKVE+EDGN+LQI+GERN E+E+K+DKWHRVERSSGKF RRFRLPEN
Sbjct: 61 FKADLPGIKKEEVKVEVEDGNILQITGERNVEKEDKNDKWHRVERSSGKFTRRFRLPENA 120
Query: 116 KMDQVKAGMENGVLTVTVPKEEEKKSEVKSIEISG 150
KMDQVKA MENGVLT+TVPKEE KK +VKSIEISG
Sbjct: 121 KMDQVKAAMENGVLTITVPKEEVKKPDVKSIEISG 155
>UniRef100_Q40510 Nthsp18p [Nicotiana tabacum]
Length = 159
Score = 243 bits (619), Expect = 8e-64
Identities = 113/159 (71%), Positives = 140/159 (87%), Gaps = 9/159 (5%)
Query: 1 MSLIPSFFGGRQNNVFDPFSMDIWDPLQGFP---------SSARETTALANTRVDWKETQ 51
M++IPSFFGGR++N+FDPFS+DI+DP +GFP SSARET+A AN R+DWKET
Sbjct: 1 MAMIPSFFGGRRSNIFDPFSLDIFDPFEGFPFSGTVANVPSSARETSAFANARIDWKETP 60
Query: 52 EAHVFSVDLPGLKKEEVKVEIEDGNVLQISGERNKEQEEKDDKWHRVERSSGKFMRRFRL 111
++H+F +D+PG+KKEEVKVE+E+G VLQISGER++EQEEK+D WHR+ERSSGKFMRRFRL
Sbjct: 61 DSHIFKMDVPGIKKEEVKVEVEEGRVLQISGERSREQEEKNDTWHRMERSSGKFMRRFRL 120
Query: 112 PENVKMDQVKAGMENGVLTVTVPKEEEKKSEVKSIEISG 150
P N KM+++KA MENGVLTVTVPKEEEKKSEVK+I+ISG
Sbjct: 121 PGNAKMEEIKAAMENGVLTVTVPKEEEKKSEVKAIDISG 159
>UniRef100_Q9ZP84 Heat shock protein 17.4 [Quercus suber]
Length = 154
Score = 242 bits (617), Expect = 1e-63
Identities = 117/151 (77%), Positives = 129/151 (84%), Gaps = 6/151 (3%)
Query: 6 SFFGGRQNNVFDPFSMDIWDPLQGF------PSSARETTALANTRVDWKETQEAHVFSVD 59
S FGGR++NVFDPFS+DIWDP +GF P SARETTA A R+DWKET EAH+F D
Sbjct: 4 SLFGGRRSNVFDPFSLDIWDPFEGFSAVASVPPSARETTAFATARIDWKETPEAHIFKAD 63
Query: 60 LPGLKKEEVKVEIEDGNVLQISGERNKEQEEKDDKWHRVERSSGKFMRRFRLPENVKMDQ 119
LPGLKKEEVKVE+EDGNVLQISGER+KE EEK+DKWHRVERS GKFMRRFRLPEN K+DQ
Sbjct: 64 LPGLKKEEVKVEVEDGNVLQISGERSKEHEEKNDKWHRVERSCGKFMRRFRLPENAKVDQ 123
Query: 120 VKAGMENGVLTVTVPKEEEKKSEVKSIEISG 150
VKA MENGVLTV VPKEE+KK VK+IEISG
Sbjct: 124 VKANMENGVLTVMVPKEEQKKPAVKAIEISG 154
>UniRef100_Q6WHC0 Chloroplast small heat shock protein class I [Capsicum frutescens]
Length = 159
Score = 241 bits (614), Expect = 3e-63
Identities = 109/159 (68%), Positives = 139/159 (86%), Gaps = 9/159 (5%)
Query: 1 MSLIPSFFGGRQNNVFDPFSMDIWDPLQGFP---------SSARETTALANTRVDWKETQ 51
MS+IPSFFGGR++N+FDP S+D+WDP +GFP SSARET+A N R+DWKET
Sbjct: 1 MSMIPSFFGGRRSNIFDPVSLDLWDPFEGFPISSTIANTPSSARETSAFPNARIDWKETP 60
Query: 52 EAHVFSVDLPGLKKEEVKVEIEDGNVLQISGERNKEQEEKDDKWHRVERSSGKFMRRFRL 111
+AH+F VD+PG+K+EEVKV++E+G +LQI+GER++EQEEK+D+WHR+ERSSGKF+RRFRL
Sbjct: 61 QAHIFKVDVPGIKREEVKVQVEEGRILQITGERSREQEEKNDQWHRMERSSGKFLRRFRL 120
Query: 112 PENVKMDQVKAGMENGVLTVTVPKEEEKKSEVKSIEISG 150
PEN KM ++KA MENGVLTVTVPKEEEK+SEVK+I+ISG
Sbjct: 121 PENTKMGEIKAAMENGVLTVTVPKEEEKRSEVKAIDISG 159
>UniRef100_Q6RFM0 Small heat shock protein [Prunus persica]
Length = 154
Score = 240 bits (612), Expect = 5e-63
Identities = 113/151 (74%), Positives = 133/151 (87%), Gaps = 6/151 (3%)
Query: 6 SFFGGRQNNVFDPFSMDIWDPLQGF------PSSARETTALANTRVDWKETQEAHVFSVD 59
S FGGR++NVFDPFS+DIWDPL+G P SARETTA+ANTR+DWKET EAH+F D
Sbjct: 4 SLFGGRRSNVFDPFSLDIWDPLEGLGTLANIPPSARETTAIANTRIDWKETPEAHIFIAD 63
Query: 60 LPGLKKEEVKVEIEDGNVLQISGERNKEQEEKDDKWHRVERSSGKFMRRFRLPENVKMDQ 119
LPGLKKEEVKVE++DG VL ISGER++EQEEK+DKWHR+ERS+GKF RRFRLP+N K+DQ
Sbjct: 64 LPGLKKEEVKVEVDDGKVLHISGERSREQEEKNDKWHRIERSTGKFSRRFRLPDNAKIDQ 123
Query: 120 VKAGMENGVLTVTVPKEEEKKSEVKSIEISG 150
VKA MENGVLTVTVPKEEEK+ +VK+I+ISG
Sbjct: 124 VKASMENGVLTVTVPKEEEKRPQVKAIDISG 154
>UniRef100_Q9SWE4 Low molecular weight heat-shock protein [Nicotiana tabacum]
Length = 159
Score = 239 bits (609), Expect = 1e-62
Identities = 112/159 (70%), Positives = 137/159 (85%), Gaps = 9/159 (5%)
Query: 1 MSLIPSFFGGRQNNVFDPFSMDIWDPLQGFP---------SSARETTALANTRVDWKETQ 51
MSLIPSFF GR++N+FDPFS++IWDP +GFP +S RET A ++ R+DWKET
Sbjct: 1 MSLIPSFFDGRRSNIFDPFSLNIWDPFEGFPFSGTVANIPTSTRETAAFSSARIDWKETP 60
Query: 52 EAHVFSVDLPGLKKEEVKVEIEDGNVLQISGERNKEQEEKDDKWHRVERSSGKFMRRFRL 111
E+HVF VDLPG+KKEEVKVE+E+G VLQISGER++EQEEK+DKWH +ERSSGKF+RRFRL
Sbjct: 61 ESHVFKVDLPGIKKEEVKVEVEEGRVLQISGERSREQEEKNDKWHSMERSSGKFLRRFRL 120
Query: 112 PENVKMDQVKAGMENGVLTVTVPKEEEKKSEVKSIEISG 150
PEN+KM+++KA MENGVLTVTVPK EEKK EVK+I+ISG
Sbjct: 121 PENIKMEEIKATMENGVLTVTVPKMEEKKPEVKAIDISG 159
>UniRef100_P27880 18.2 kDa class I heat shock protein [Medicago sativa]
Length = 158
Score = 239 bits (609), Expect = 1e-62
Identities = 117/158 (74%), Positives = 133/158 (84%), Gaps = 8/158 (5%)
Query: 1 MSLIPSFFGGRQNNVFDPFSMDIWDPLQGFP--------SSARETTALANTRVDWKETQE 52
MSLIPSFFGGR++NVFDPFS+D+WDP + FP S RE +A +TRVDWKET E
Sbjct: 1 MSLIPSFFGGRRSNVFDPFSLDVWDPFKDFPFNNSALSASFPRENSAFVSTRVDWKETPE 60
Query: 53 AHVFSVDLPGLKKEEVKVEIEDGNVLQISGERNKEQEEKDDKWHRVERSSGKFMRRFRLP 112
AHVF DLPG+KKEEVKVEIED VLQISGER+ E+E+K+D+WHR+ERSSGKFMRRFRLP
Sbjct: 61 AHVFKADLPGMKKEEVKVEIEDDRVLQISGERSVEKEDKNDQWHRLERSSGKFMRRFRLP 120
Query: 113 ENVKMDQVKAGMENGVLTVTVPKEEEKKSEVKSIEISG 150
EN KMDQVKA MENGVLTVTVPKEE KK EVK+I+ISG
Sbjct: 121 ENAKMDQVKAAMENGVLTVTVPKEEVKKPEVKTIDISG 158
>UniRef100_P19243 18.1 kDa class I heat shock protein [Pisum sativum]
Length = 158
Score = 239 bits (609), Expect = 1e-62
Identities = 118/158 (74%), Positives = 133/158 (83%), Gaps = 8/158 (5%)
Query: 1 MSLIPSFFGGRQNNVFDPFSMDIWDPLQGFPSS--------ARETTALANTRVDWKETQE 52
MSLIPSFF GR++NVFDPFS+D+WDPL+ FP S RE A +TRVDWKET E
Sbjct: 1 MSLIPSFFSGRRSNVFDPFSLDVWDPLKDFPFSNSSPSASFPRENPAFVSTRVDWKETPE 60
Query: 53 AHVFSVDLPGLKKEEVKVEIEDGNVLQISGERNKEQEEKDDKWHRVERSSGKFMRRFRLP 112
AHVF DLPGLKKEEVKVE+ED VLQISGER+ E+E+K+D+WHRVERSSGKF+RRFRLP
Sbjct: 61 AHVFKADLPGLKKEEVKVEVEDDRVLQISGERSVEKEDKNDEWHRVERSSGKFLRRFRLP 120
Query: 113 ENVKMDQVKAGMENGVLTVTVPKEEEKKSEVKSIEISG 150
EN KMD+VKA MENGVLTVTVPKEE KK+EVKSIEISG
Sbjct: 121 ENAKMDKVKASMENGVLTVTVPKEEIKKAEVKSIEISG 158
>UniRef100_Q39930 17.7 kDa heat shock protein [Helianthus annuus]
Length = 156
Score = 237 bits (605), Expect = 3e-62
Identities = 115/156 (73%), Positives = 131/156 (83%), Gaps = 7/156 (4%)
Query: 1 MSLIPSFFGGRQNNVFDPFSMDIWDPLQGF-------PSSARETTALANTRVDWKETQEA 53
MS+IPSFF G +N+FDPFS +IWDP QG P S+RETTA+ANTR+DWKET EA
Sbjct: 1 MSIIPSFFTGNGSNIFDPFSSEIWDPFQGLSSVINNLPESSRETTAIANTRIDWKETPEA 60
Query: 54 HVFSVDLPGLKKEEVKVEIEDGNVLQISGERNKEQEEKDDKWHRVERSSGKFMRRFRLPE 113
HVF DLPGLKKEEVKVE+E+G VLQISGER++E EK+DKWHR+ERSSGKF+RRFRLPE
Sbjct: 61 HVFKADLPGLKKEEVKVEVEEGRVLQISGERSRENVEKNDKWHRMERSSGKFLRRFRLPE 120
Query: 114 NVKMDQVKAGMENGVLTVTVPKEEEKKSEVKSIEIS 149
N KMDQVKA MENGVLTVTVPK E KK EVK+I+IS
Sbjct: 121 NAKMDQVKAAMENGVLTVTVPKAEVKKPEVKAIDIS 156
>UniRef100_Q8VXJ4 20.5 kDa heat-shock protein [Helianthus annuus]
Length = 177
Score = 237 bits (604), Expect = 4e-62
Identities = 112/154 (72%), Positives = 132/154 (84%), Gaps = 5/154 (3%)
Query: 1 MSLIPSFFGGRQNNVFDPFSMDIWDPLQGFPSS-----ARETTALANTRVDWKETQEAHV 55
MS++PS FG R++++FDPFS+ +WDP + FP S +RET+AL N RVDWKET EAHV
Sbjct: 1 MSIVPSLFGSRRSSIFDPFSLYVWDPFRDFPISTSSEVSRETSALVNARVDWKETPEAHV 60
Query: 56 FSVDLPGLKKEEVKVEIEDGNVLQISGERNKEQEEKDDKWHRVERSSGKFMRRFRLPENV 115
F DLPG+KKEEVKVE+EDGN+LQI+GERN E+E+K+DKWHRVERSSGKF RRFRLPEN
Sbjct: 61 FKADLPGIKKEEVKVEVEDGNILQITGERNVEKEDKNDKWHRVERSSGKFTRRFRLPENA 120
Query: 116 KMDQVKAGMENGVLTVTVPKEEEKKSEVKSIEIS 149
KMDQVKA MENGVLT+TVPKEE KK +VKSIEIS
Sbjct: 121 KMDQVKAAMENGVLTITVPKEEAKKPDVKSIEIS 154
>UniRef100_Q8L8S1 Heat shock protein 18 [Arabidopsis thaliana]
Length = 161
Score = 237 bits (604), Expect = 4e-62
Identities = 116/159 (72%), Positives = 130/159 (80%), Gaps = 9/159 (5%)
Query: 1 MSLIPSFFGGRQNNVFDPFSMDIWDPLQGF---------PSSARETTALANTRVDWKETQ 51
MSLIPS FGGR++NVFDPFS D+WDP +GF S+AR+ A N RVDWKET
Sbjct: 1 MSLIPSIFGGRRSNVFDPFSQDVWDPFEGFFTPSSALANASTARDVAAFTNARVDWKETP 60
Query: 52 EAHVFSVDLPGLKKEEVKVEIEDGNVLQISGERNKEQEEKDDKWHRVERSSGKFMRRFRL 111
EAHVF DLPGLKKEEVKVE+ED NVLQISGER+KE EEK+DKWHRVER+SGKFMRRFRL
Sbjct: 61 EAHVFKADLPGLKKEEVKVEVEDKNVLQISGERSKENEEKNDKWHRVERASGKFMRRFRL 120
Query: 112 PENVKMDQVKAGMENGVLTVTVPKEEEKKSEVKSIEISG 150
PEN KM++VKA MENGVLTV VPK EKK +VKSI+ISG
Sbjct: 121 PENAKMEEVKATMENGVLTVVVPKAPEKKPQVKSIDISG 159
>UniRef100_P05478 18.5 kDa class I heat shock protein [Glycine max]
Length = 161
Score = 236 bits (603), Expect = 6e-62
Identities = 118/161 (73%), Positives = 134/161 (82%), Gaps = 11/161 (6%)
Query: 1 MSLIPSFFGGRQNNVFDPFSMDIWDPLQGFP-----SSA------RETTALANTRVDWKE 49
MSLIP+FFGGR+NNVFDPFS+D+WDP + FP SSA RE +A +TRVDWKE
Sbjct: 1 MSLIPNFFGGRRNNVFDPFSLDVWDPFKDFPFPNTLSSASFPEFSRENSAFVSTRVDWKE 60
Query: 50 TQEAHVFSVDLPGLKKEEVKVEIEDGNVLQISGERNKEQEEKDDKWHRVERSSGKFMRRF 109
T EAHVF D+PGLKKEEVKV+IED VLQISGERN E+E+K+D WHRVERSSGKFMRRF
Sbjct: 61 TPEAHVFKADIPGLKKEEVKVQIEDDKVLQISGERNVEKEDKNDTWHRVERSSGKFMRRF 120
Query: 110 RLPENVKMDQVKAGMENGVLTVTVPKEEEKKSEVKSIEISG 150
RLPEN K++QVKA MENGVLTVTVPKEE KK +VK+IEISG
Sbjct: 121 RLPENAKVEQVKASMENGVLTVTVPKEEVKKPDVKAIEISG 161
>UniRef100_P19037 18.2 kDa class I heat shock protein [Arabidopsis thaliana]
Length = 161
Score = 236 bits (603), Expect = 6e-62
Identities = 116/159 (72%), Positives = 130/159 (80%), Gaps = 9/159 (5%)
Query: 1 MSLIPSFFGGRQNNVFDPFSMDIWDPLQGF---------PSSARETTALANTRVDWKETQ 51
MSLIPS FGGR++NVFDPFS D+WDP +GF S+AR+ A N RVDWKET
Sbjct: 1 MSLIPSIFGGRRSNVFDPFSQDLWDPFEGFFTPSSALANASTARDVAAFTNARVDWKETP 60
Query: 52 EAHVFSVDLPGLKKEEVKVEIEDGNVLQISGERNKEQEEKDDKWHRVERSSGKFMRRFRL 111
EAHVF DLPGLKKEEVKVE+ED NVLQISGER+KE EEK+DKWHRVER+SGKFMRRFRL
Sbjct: 61 EAHVFKADLPGLKKEEVKVEVEDKNVLQISGERSKENEEKNDKWHRVERASGKFMRRFRL 120
Query: 112 PENVKMDQVKAGMENGVLTVTVPKEEEKKSEVKSIEISG 150
PEN KM++VKA MENGVLTV VPK EKK +VKSI+ISG
Sbjct: 121 PENAKMEEVKATMENGVLTVVVPKAPEKKPQVKSIDISG 159
>UniRef100_P27396 17.8 kDa class I heat shock protein [Daucus carota]
Length = 157
Score = 236 bits (603), Expect = 6e-62
Identities = 115/157 (73%), Positives = 134/157 (85%), Gaps = 7/157 (4%)
Query: 1 MSLIPSFFGGRQNNVFDPFSMDIWDPLQGFP---SSA----RETTALANTRVDWKETQEA 53
MS+IPSFFGGR++NVFDPFS+D+WDP + FP SSA +ET A NT +DWKET +A
Sbjct: 1 MSIIPSFFGGRRSNVFDPFSLDVWDPFKDFPLVTSSASEFGKETAAFVNTHIDWKETPQA 60
Query: 54 HVFSVDLPGLKKEEVKVEIEDGNVLQISGERNKEQEEKDDKWHRVERSSGKFMRRFRLPE 113
HVF DLPGLKKEEVKVE+E+G VLQISGERNKE+EEK+DKWHRVERSSGKF+RRFRLPE
Sbjct: 61 HVFKADLPGLKKEEVKVELEEGKVLQISGERNKEKEEKNDKWHRVERSSGKFLRRFRLPE 120
Query: 114 NVKMDQVKAGMENGVLTVTVPKEEEKKSEVKSIEISG 150
N K+D+VKA M NGV+TVTVPK E KK EVK+I+ISG
Sbjct: 121 NAKVDEVKAAMANGVVTVTVPKVEIKKPEVKAIDISG 157
>UniRef100_Q8H1A6 Heat shock protein [Pisum sativum]
Length = 158
Score = 235 bits (600), Expect = 1e-61
Identities = 116/158 (73%), Positives = 132/158 (83%), Gaps = 8/158 (5%)
Query: 1 MSLIPSFFGGRQNNVFDPFSMDIWDPLQGFP--------SSARETTALANTRVDWKETQE 52
MSLIPSFFG R++NV+DPFS+D+WDPL+ FP S RE +A +TRVDWKET E
Sbjct: 1 MSLIPSFFGNRRSNVYDPFSLDVWDPLKDFPFPSSALSASFPRENSAFVSTRVDWKETPE 60
Query: 53 AHVFSVDLPGLKKEEVKVEIEDGNVLQISGERNKEQEEKDDKWHRVERSSGKFMRRFRLP 112
AHVF DLPGLKKEEVKVE+ED VLQISGER+ E+E+K+D+WHRVERSSGKF+RRFRLP
Sbjct: 61 AHVFKADLPGLKKEEVKVEVEDDRVLQISGERSVEKEDKNDEWHRVERSSGKFLRRFRLP 120
Query: 113 ENVKMDQVKAGMENGVLTVTVPKEEEKKSEVKSIEISG 150
EN KM QVKA MENGVLTVTVPKEE KK +VKSIEISG
Sbjct: 121 ENAKMGQVKASMENGVLTVTVPKEEIKKPDVKSIEISG 158
>UniRef100_P02519 17.3 kDa class I heat shock protein [Glycine max]
Length = 153
Score = 233 bits (594), Expect = 6e-61
Identities = 112/153 (73%), Positives = 132/153 (86%), Gaps = 3/153 (1%)
Query: 1 MSLIPSFFGGRQNNVFDPFSMDIWDPLQGFP---SSARETTALANTRVDWKETQEAHVFS 57
MSLIPSFFGGR+++VFDPFS+D+WDP + FP S + E +A +TRVDWKET EAHVF
Sbjct: 1 MSLIPSFFGGRRSSVFDPFSLDVWDPFKDFPFPSSLSAENSAFVSTRVDWKETPEAHVFK 60
Query: 58 VDLPGLKKEEVKVEIEDGNVLQISGERNKEQEEKDDKWHRVERSSGKFMRRFRLPENVKM 117
D+PGLKKEEVK+EI+DG VLQISGERN E+E+K+D WHRVERSSGK +RRFRLPEN K+
Sbjct: 61 ADIPGLKKEEVKLEIQDGRVLQISGERNVEKEDKNDTWHRVERSSGKLVRRFRLPENAKV 120
Query: 118 DQVKAGMENGVLTVTVPKEEEKKSEVKSIEISG 150
DQVKA MENGVLTVTVPKEE KK +VK+I+ISG
Sbjct: 121 DQVKASMENGVLTVTVPKEEIKKPDVKAIDISG 153
>UniRef100_Q9SMF2 17.9 kDa heat-shock protein [Helianthus annuus]
Length = 157
Score = 233 bits (593), Expect = 8e-61
Identities = 110/151 (72%), Positives = 130/151 (85%), Gaps = 5/151 (3%)
Query: 1 MSLIPSFFGGRQNNVFDPFSMDIWDPLQGFPSSA-----RETTALANTRVDWKETQEAHV 55
MS++ S FGGR+++VFDPFS+D+WDP + FP S+ RET+AL N RVDWKET EAHV
Sbjct: 1 MSIVSSLFGGRRSSVFDPFSLDVWDPFRDFPISSSSDVSRETSALVNARVDWKETPEAHV 60
Query: 56 FSVDLPGLKKEEVKVEIEDGNVLQISGERNKEQEEKDDKWHRVERSSGKFMRRFRLPENV 115
F DLPG+KKEEVKVE+EDGN+L+I+GERN E+E+K+DKWHRVERSSGKF RRFRLPEN
Sbjct: 61 FKADLPGIKKEEVKVEVEDGNILKITGERNIEKEDKNDKWHRVERSSGKFTRRFRLPENA 120
Query: 116 KMDQVKAGMENGVLTVTVPKEEEKKSEVKSI 146
KMDQVKA MENGVLT+TVPKEE KK +VKSI
Sbjct: 121 KMDQVKAAMENGVLTITVPKEEVKKPDVKSI 151
>UniRef100_O82149 Low-molecular-weight heat shock protein [Cuscuta japonica]
Length = 157
Score = 233 bits (593), Expect = 8e-61
Identities = 115/158 (72%), Positives = 133/158 (83%), Gaps = 9/158 (5%)
Query: 1 MSLIPSFFGGRQNNVFDPFSMDIWDP--------LQGFPSSARETTALANTRVDWKETQE 52
MSLIPSFF GR++N FDPFS+++WDP L G SSARE +A AN R+DWKET E
Sbjct: 1 MSLIPSFFEGRRSNAFDPFSLELWDPFFSNTVANLSG-SSSAREASAFANARIDWKETPE 59
Query: 53 AHVFSVDLPGLKKEEVKVEIEDGNVLQISGERNKEQEEKDDKWHRVERSSGKFMRRFRLP 112
AH+F D+PGLKKEEVKVE+E+G VLQISGER+KE+EEK+D WHRVERSSGKF+R FRLP
Sbjct: 60 AHIFKADVPGLKKEEVKVEVEEGKVLQISGERSKEKEEKNDTWHRVERSSGKFLRSFRLP 119
Query: 113 ENVKMDQVKAGMENGVLTVTVPKEEEKKSEVKSIEISG 150
EN K+DQVKA MENGVLTVTVPK EEKK+EVKSI+ISG
Sbjct: 120 ENAKVDQVKAAMENGVLTVTVPKVEEKKAEVKSIQISG 157
>UniRef100_Q40978 Low molecular weight heat-shock protein [Papaver somniferum]
Length = 210
Score = 232 bits (592), Expect = 1e-60
Identities = 116/165 (70%), Positives = 134/165 (80%), Gaps = 15/165 (9%)
Query: 1 MSLIPSFFGGRQNNVFDPFSMDIWDPLQGFPSS--------------ARETTALANTRVD 46
MS+IPSFF +++NVFDPFS+DIWDP QGFP S ARET+ LANTR+D
Sbjct: 1 MSIIPSFFSNQRSNVFDPFSLDIWDPFQGFPFSTGALTANWQGGSDTARETSQLANTRID 60
Query: 47 WKETQEAHVFSVDLPGLKKEEVKVEIEDGNVLQISGER-NKEQEEKDDKWHRVERSSGKF 105
WKET EAHVF DLPG+ KEEVKVE+E+G VLQISGER ++E EEK+DKWHRVERSSGKF
Sbjct: 61 WKETPEAHVFRADLPGVTKEEVKVEVEEGRVLQISGERRSRESEEKNDKWHRVERSSGKF 120
Query: 106 MRRFRLPENVKMDQVKAGMENGVLTVTVPKEEEKKSEVKSIEISG 150
+RRFRLPEN KMD+VKA MENGVLTV VPK E+++ EVKSIEISG
Sbjct: 121 LRRFRLPENTKMDEVKATMENGVLTVCVPKVEQRRPEVKSIEISG 165
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.312 0.131 0.372
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 253,896,004
Number of Sequences: 2790947
Number of extensions: 10205231
Number of successful extensions: 33294
Number of sequences better than 10.0: 1036
Number of HSP's better than 10.0 without gapping: 559
Number of HSP's successfully gapped in prelim test: 479
Number of HSP's that attempted gapping in prelim test: 32032
Number of HSP's gapped (non-prelim): 1205
length of query: 150
length of database: 848,049,833
effective HSP length: 126
effective length of query: 24
effective length of database: 496,390,511
effective search space: 11913372264
effective search space used: 11913372264
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 67 (30.4 bits)
Medicago: description of AC147434.11


