

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147430.8 + phase: 1 /pseudo
(437 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
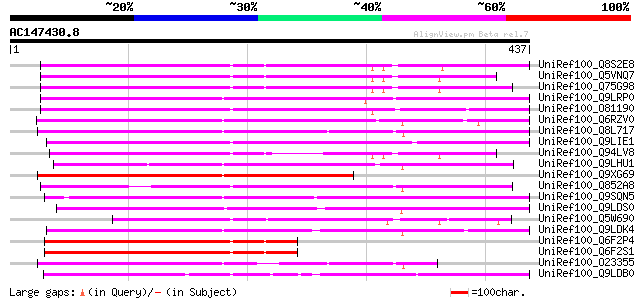
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q8S2E8 P0022F10.3 protein [Oryza sativa] 290 4e-77
UniRef100_Q5VNQ7 HAT dimerisation domain-containing protein-like... 283 9e-75
UniRef100_Q75G98 Hypothetical protein OSJNBa0042F15.17 [Oryza sa... 282 1e-74
UniRef100_Q9LRP0 Emb|CAB39405.1 [Arabidopsis thaliana] 278 2e-73
UniRef100_O81190 Putative transposase [Arabidopsis thaliana] 275 1e-72
UniRef100_Q6RZV0 Transposase-like protein [Musa acuminata] 249 8e-65
UniRef100_Q8L717 Hypothetical protein At4g15020 [Arabidopsis tha... 247 5e-64
UniRef100_Q9LIE1 Transposase-like protein [Arabidopsis thaliana] 245 2e-63
UniRef100_Q94LV8 Hypothetical protein OSJNBa0082M15.8 [Oryza sat... 229 1e-58
UniRef100_Q9LHU1 Similar to Arabidopsis thaliana chromosome 1 [O... 223 6e-57
UniRef100_Q9XG69 Hypothetical protein [Nicotiana tabacum] 212 2e-53
UniRef100_Q852A8 Hypothetical protein OSJNBb0081B07.17 [Oryza sa... 207 5e-52
UniRef100_Q9SQN5 Hypothetical protein F19K16.28 [Arabidopsis tha... 200 8e-50
UniRef100_Q9LDS0 Transposase-like protein [Arabidopsis thaliana] 199 1e-49
UniRef100_Q5W690 Hypothetical protein OSJNBa0076A09.5 [Oryza sat... 196 1e-48
UniRef100_Q9LDK4 Transposase-like protein [Arabidopsis thaliana] 196 1e-48
UniRef100_Q6F2P4 Hypothetical protein OSJNBa0075J06.6 [Oryza sat... 195 2e-48
UniRef100_Q6F2S1 Hypothetical protein OSJNBa0060G17.6 [Oryza sat... 191 5e-47
UniRef100_O23355 Hypothetical protein AT4g15020 [Arabidopsis tha... 167 4e-40
UniRef100_Q9LDB0 Transposase-like protein [Arabidopsis thaliana] 164 5e-39
>UniRef100_Q8S2E8 P0022F10.3 protein [Oryza sativa]
Length = 965
Score = 290 bits (743), Expect = 4e-77
Identities = 166/421 (39%), Positives = 241/421 (56%), Gaps = 16/421 (3%)
Query: 27 FNIMCDMISRHGNGFKPPSYYDIRVKLLNQEVKSTNDALEEHKREWKKTSCTIMTDEWTD 86
F M + I + G K PS Y++ L + + D +EHK W+ T C+IMTD WTD
Sbjct: 382 FAHMLEAIGQFGRSLKGPSPYEMSGSFLQKRKEKVMDGFKEHKESWELTGCSIMTDAWTD 441
Query: 87 RRRRTILNFLVNSPRGTVFLKSVDASNICKTSEKIFEMIDAVVEEVGEDNVVQVVTDNAA 146
R+ R ++N +V+S G +FL SV+ S K + IF+++D +EE+GE +VVQVVTDNA+
Sbjct: 442 RKGRGVMNLVVHSAHGVLFLDSVECSGDRKDGKYIFKLVDRYIEEIGEQHVVQVVTDNAS 501
Query: 147 NYKAAGEMLMRKRKKLYWTPCAAHCLDLMLEDLEKKIPIHGETIPKGKKITTFIYARTSL 206
A +L KR ++W CAAHCLDLMLED+ K P+ ETI +++T F+YA T +
Sbjct: 502 VNTIAASLLTAKRPSIFWNGCAAHCLDLMLEDIGKLGPVE-ETIANARQVTVFLYAHTRV 560
Query: 207 ISILHSHTKNRDLVRPGATRFATSYLTLGCLYENKEALIRMFTSKEWKSGKCAKLRDGKA 266
+ ++ NRDLVR G TRFAT+YL L L +NK+ L+R+F S E + K GK
Sbjct: 561 LDLMRKFL-NRDLVRSGVTRFATAYLNLKSLLDNKKELVRLFKSDEMEQLGYLKQTKGKK 619
Query: 267 LEDIILDKKFWKDIVICLKGATPLMKVLRLADLDKKTS---HGIHL*SNG--SSKGANSK 321
+I + FWK++ I + PL VLR D D + HG+ L + S + N K
Sbjct: 620 ASKVIRSETFWKNVDIAVNYFEPLANVLRRMDSDVPSMGFFHGLMLEAKKEISQRFDNDK 679
Query: 322 EF*WCQKEPVWNIIDDRWDRMLHRPLHAAAYYLNPQLHYRPG---FKADIEVRKGLMDCI 378
+ VW+IID RWD L PLH A YYLNP +Y P ++D R G++ CI
Sbjct: 680 S----RFIEVWDIIDKRWDNKLKTPLHLAGYYLNPYYYYYPNKQEIESDGSFRAGVISCI 735
Query: 379 TRMVEDPEEQAKIEVQMHDFKKQIGYFGTKIA-KLTLDKS-TPADWWESVVFEYLELQRF 436
++V+D + Q KI +++ ++ Q G FG +IA + +K+ PA WW + L L++
Sbjct: 736 DKLVDDEDIQDKIIEELNLYQDQHGSFGHEIAVRQRKNKNFNPAKWWLNHGTSTLNLRKL 795
Query: 437 A 437
A
Sbjct: 796 A 796
>UniRef100_Q5VNQ7 HAT dimerisation domain-containing protein-like [Oryza sativa]
Length = 622
Score = 283 bits (723), Expect = 9e-75
Identities = 157/391 (40%), Positives = 227/391 (57%), Gaps = 13/391 (3%)
Query: 27 FNIMCDMISRHGNGFKPPSYYDIRVKLLNQEVKSTNDALEEHKREWKKTSCTIMTDEWTD 86
F M + I + G K PS Y++ L + + D +EHK W+ T C+IMTD WTD
Sbjct: 221 FAHMLEAIGQFGRSLKGPSPYEMSGSFLQKRKEKVMDGFKEHKESWELTGCSIMTDAWTD 280
Query: 87 RRRRTILNFLVNSPRGTVFLKSVDASNICKTSEKIFEMIDAVVEEVGEDNVVQVVTDNAA 146
R+ R ++N +V+S G +FL SV+ S K + IFE++D +EE+GE +VVQVVTDNA+
Sbjct: 281 RKGRGVMNLVVHSAHGVLFLDSVECSGDRKDGKYIFELVDRYIEEIGEQHVVQVVTDNAS 340
Query: 147 NYKAAGEMLMRKRKKLYWTPCAAHCLDLMLEDLEKKIPIHGETIPKGKKITTFIYARTSL 206
A +L KR ++W CAAHCLDLMLED+ K P+ ETI +++T F+YA T +
Sbjct: 341 VNTTAASLLTAKRPSIFWNGCAAHCLDLMLEDIGKLGPVE-ETIANARQVTVFLYAHTRV 399
Query: 207 ISILHSHTKNRDLVRPGATRFATSYLTLGCLYENKEALIRMFTSKEWKSGKCAKLRDGKA 266
+ ++ NRDLVR G TRFAT+YL L L +NK+ L+R+F S E + K GK
Sbjct: 400 LDLMRKFL-NRDLVRSGVTRFATAYLNLKSLLDNKKELVRLFKSNEMEQLGYLKQAKGKK 458
Query: 267 LEDIILDKKFWKDIVICLKGATPLMKVLRLADLDKKTS---HGIHL*SNG--SSKGANSK 321
+I+ + FWK++ I + PL VLR D D + HG+ L + S + N K
Sbjct: 459 ASKVIISETFWKNVDIAVNYFEPLANVLRRMDSDVPSMGFFHGLMLEAKKEISQRFDNDK 518
Query: 322 EF*WCQKEPVWNIIDDRWDRMLHRPLHAAAYYLNPQLHY--RPGFKADIEVRKGLMDCIT 379
+ VW+IID RWD L LH A YYLNP +Y + ++D R G++ CI
Sbjct: 519 S----RFIEVWDIIDKRWDNKLKTLLHLAGYYLNPYYYYPNKQEIESDGSFRAGVISCID 574
Query: 380 RMVEDPEEQAKIEVQMHDFKKQIGYFGTKIA 410
++V+D + Q KI +++ ++ Q G FG +IA
Sbjct: 575 KLVDDEDIQDKIIEELNLYQDQHGSFGHEIA 605
>UniRef100_Q75G98 Hypothetical protein OSJNBa0042F15.17 [Oryza sativa]
Length = 1005
Score = 282 bits (722), Expect = 1e-74
Identities = 159/406 (39%), Positives = 234/406 (57%), Gaps = 15/406 (3%)
Query: 27 FNIMCDMISRHGNGFKPPSYYDIRVKLLNQEVKSTNDALEEHKREWKKTSCTIMTDEWTD 86
F M + I + G K PS Y++ L + + D +EHK W+ T C+IMTD WTD
Sbjct: 427 FAHMLEAIGQFGRSLKGPSPYEMSGSFLQKRKEKVMDGFKEHKESWELTGCSIMTDAWTD 486
Query: 87 RRRRTILNFLVNSPRGTVFLKSVDASNICKTSEKIFEMIDAVVEEVGEDNVVQVVTDNAA 146
R+ R ++N +V+S G +FL SV+ S K + IFE++D +EE+GE +VVQVVTDNA+
Sbjct: 487 RKGRGVMNLVVHSAHGVLFLDSVECSGDRKDGKYIFELVDRYIEEIGEQHVVQVVTDNAS 546
Query: 147 NYKAAGEMLMRKRKKLYWTPCAAHCLDLMLEDLEKKIPIHGETIPKGKKITTFIYARTSL 206
A +L KR ++W AAHCLDLMLED+ K P+ ETI +++T F+YA T +
Sbjct: 547 VNTTAASLLTAKRPSIFWNGYAAHCLDLMLEDIGKLGPVE-ETIANARQVTVFLYAHTRV 605
Query: 207 ISILHSHTKNRDLVRPGATRFATSYLTLGCLYENKEALIRMFTSKEWKSGKCAKLRDGKA 266
+ ++ NRDLVR G TRFAT+YL L L +NK+ L+R+F S E + K GK
Sbjct: 606 LDLMRKFL-NRDLVRSGVTRFATAYLNLKSLLDNKKELVRLFKSDEMEQLGYLKQAKGKK 664
Query: 267 LEDIILDKKFWKDIVICLKGATPLMKVLRLADLDKKTS---HGIHL*SNG--SSKGANSK 321
+I + FWK++ I + PL VLR D D + HG+ L + S + N K
Sbjct: 665 ASKVIRSETFWKNVDIAVNYFEPLANVLRRMDSDVPSMGFFHGLMLEAKKEISQRFDNDK 724
Query: 322 EF*WCQKEPVWNIIDDRWDRMLHRPLHAAAYYLNPQLHY--RPGFKADIEVRKGLMDCIT 379
+ VW+IID RWD + PLH A YYLNP ++ + ++D R G++ CI
Sbjct: 725 S----RFIEVWDIIDKRWDNKIKAPLHLAGYYLNPYYYFPNKQEIESDGSFRAGVISCID 780
Query: 380 RMVEDPEEQAKIEVQMHDFKKQIGYFGTKIA-KLTLDKS-TPADWW 423
++++D + Q KI +++ ++ Q G FG +IA + +K+ PA WW
Sbjct: 781 KLIDDEDIQDKIIEELNLYQDQHGSFGHEIAVRQRKNKNFNPAKWW 826
>UniRef100_Q9LRP0 Emb|CAB39405.1 [Arabidopsis thaliana]
Length = 883
Score = 278 bits (712), Expect = 2e-73
Identities = 149/418 (35%), Positives = 239/418 (56%), Gaps = 10/418 (2%)
Query: 27 FNIMCDMISRHGNGFKPPSYYDIRVKLLNQEVKSTNDALEEHKREWKKTSCTIMTDEWTD 86
F M ++I +G GF PS +LL +E+ + L E++ W T C+IM D WT+
Sbjct: 325 FQKMIELIGMYGEGFVVPSSQLFSGRLLQEEMSTIKSYLREYRSSWVVTGCSIMADTWTN 384
Query: 87 RRRRTILNFLVNSPRGTVFLKSVDASNICKTSEKIFEMIDAVVEEVGEDNVVQVVTDNAA 146
+ +++FLV+ PRG F S+DA++I + + +F+ +D +V+++GE+NVVQV+T N A
Sbjct: 385 TEGKKMISFLVSCPRGVYFHSSIDATDIVEDALSLFKCLDKLVDDIGEENVVQVITQNTA 444
Query: 147 NYKAAGEMLMRKRKKLYWTPCAAHCLDLMLEDLEKKIPIHGETIPKGKKITTFIYARTSL 206
+++AG++L KRK LYWTPCA HC +L+LED K+ E + K ++IT FIY +T L
Sbjct: 445 IFRSAGKLLEEKRKNLYWTPCAIHCTELVLEDF-SKLEFVSECLEKAQRITRFIYNQTWL 503
Query: 207 ISIL-HSHTKNRDLVRPGATRFATSYLTLGCLYENKEALIRMFTSKEW-KSGKCAKLRDG 264
++++ + T+ DL+RP R A+ + TL L ++K +L +F S W S AK +G
Sbjct: 504 LNLMKNEFTQGLDLLRPAVMRHASGFTTLQSLMDHKASLRGLFQSDGWILSQTAAKSEEG 563
Query: 265 KALEDIILDKKFWKDIVICLKGATPLMKVLRLAD-----LDKKTSHGIHL*SNGSSKGAN 319
+ +E ++L FWK + LK P+M+V+ + + L ++G + + K +
Sbjct: 564 REVEKMVLSAVFWKKVQYVLKSVDPVMQVIHMINDGGDRLSMPYAYGYMCCAKMAIKSIH 623
Query: 320 SKEF*WCQKEPVWNIIDDRWDRMLHRPLHAAAYYLNPQLHYRPGFKADIEVRKGLMDCIT 379
S + + P W +I+ RW+ + H PL+ AAY+ NP YRP F A EV +G+ +CI
Sbjct: 624 SDDA--RKYGPFWRVIEYRWNPLFHHPLYVAAYFFNPAYKYRPDFMAQSEVVRGVNECIV 681
Query: 380 RMVEDPEEQAKIEVQMHDFKKQIGYFGTKIAKLTLDKSTPADWWESVVFEYLELQRFA 437
R+ D + +Q+ D+ FGT IA T + P+ WW+ LELQR A
Sbjct: 682 RLEPDNTRRITALMQIPDYTCAKADFGTDIAIGTRTELDPSAWWQQHGISCLELQRVA 739
>UniRef100_O81190 Putative transposase [Arabidopsis thaliana]
Length = 729
Score = 275 bits (704), Expect = 1e-72
Identities = 150/417 (35%), Positives = 237/417 (55%), Gaps = 12/417 (2%)
Query: 27 FNIMCDMISRHGNGFKPPSYYDIRVKLLNQEVKSTNDALEEHKREWKKTSCTIMTDEWTD 86
F M M++ G+G+ PSY+ +RV LL + +++ K W T CT+M D W D
Sbjct: 188 FQPMMSMVASMGHGYVGPSYHALRVGLLRDAKLQVSLIIDKFKSSWASTGCTLMADGWKD 247
Query: 87 RRRRTILNFLVNSPRGTVFLKSVDASNICKTSEKIFEMIDAVVEEVGEDNVVQVVTDNAA 146
R+R ++NFLV P+G FLKSVDAS+I ++E + + +V +G +N+V VTD+A
Sbjct: 248 TRQRPLINFLVYCPKGITFLKSVDASDIYASAENLCNLFAELVGIIGSENIVHFVTDSAP 307
Query: 147 NYKAAGEMLMRKRKKLYWTPCAAHCLDLMLEDLEKKIPIHGETIPKGKKITTFIYARTSL 206
NYKAAG++L+ K + W+PC+AHC++L+LED+ K +H + + K+T F+Y
Sbjct: 308 NYKAAGKLLVEKFPTIAWSPCSAHCINLILEDVAKLPHVH-HIVRRMSKVTIFVYNHKPA 366
Query: 207 ISILHSHTKNRDLVRPGATRFATSYLTLGCLYENKEALIRMFTSKEWKSGKCAKLRDGKA 266
++ + + R+++RPG TRFAT+++ L LY++KE L + TS + + + K K
Sbjct: 367 LNWVRKRSGWREIIRPGETRFATTFIALQSLYQHKEDLQALVTSADPELKQLFKTSKAKV 426
Query: 267 LEDIILDKKFWKDIVICLKGATPLMKVLRLADLDKKTS-----HGIHL*SNGSSKGANSK 321
+ +ILD++ W D +I +K TP++++LR+ D D+K S G++ G K
Sbjct: 427 AKSVILDERMWNDCLIIVKVMTPIIRLLRICDADEKPSLPYVYEGMYRARLGIKNIFQEK 486
Query: 322 EF*WCQKEPVWNIIDDRWDRMLHRPLHAAAYYLNPQLHY-RPGFKADIEVRKGLMDCITR 380
E + +P NIID RWDRML LHAAAYYLNP Y +P F EV GLM+ +
Sbjct: 487 ETLY---KPYTNIIDRRWDRMLRHDLHAAAYYLNPAFMYDQPTFCEKPEVMSGLMNLFEK 543
Query: 381 MVEDPEEQAKIEVQMHDFKKQIGYFGTKIAKLTLDKSTPADWWESVVFEYLELQRFA 437
D + K+ ++ ++++ G F +A S P +WW + LQ+ A
Sbjct: 544 QKND--SKTKLFQELRVYREREGSFSLDMALTCSKTSQPDEWWRYFGHDAPNLQKMA 598
>UniRef100_Q6RZV0 Transposase-like protein [Musa acuminata]
Length = 670
Score = 249 bits (637), Expect = 8e-65
Identities = 146/419 (34%), Positives = 233/419 (54%), Gaps = 8/419 (1%)
Query: 23 RSKEFNIMCDMISRHGNGFKPPSYYDIRVKLLNQEVKSTNDALEEHKREWKKTSCTIMTD 82
+S + M D I+ G G+KPP+Y +R LL + + N+ + K EWK T CTI++D
Sbjct: 160 KSIYYQAMIDAIADCGAGYKPPTYEGLRSTLLEKVKEEINENHRKLKDEWKDTGCTILSD 219
Query: 83 EWTDRRRRTILNFLVNSPRGTVFLKSVDASNICKTSEKIFEMIDAVVEEVGEDNVVQVVT 142
W+D R +++L V SP+GT FLK VD S+ + +FE++D+V+ EVG +NVVQV+T
Sbjct: 220 NWSDGRSKSLLVLSVASPKGTQFLKLVDISSRADDAYYLFELLDSVIMEVGAENVVQVIT 279
Query: 143 DNAANYKAAGEMLMRKRKKLYWTPCAAHCLDLMLEDLEKKIPIHGETIPKGKKITTFIYA 202
D+A +Y A +L++K L+W PCA++ ++ MLED+ K+ T+ + + I FI +
Sbjct: 280 DSATSYTYAAGLLLKKYPSLFWFPCASYSIEKMLEDI-SKLEWVSTTLEETRTIARFICS 338
Query: 203 RTSLISILHSHTKNRDLVRPGATRFATSYLTLGCLYENKEALIRMFTSKEWKSGKCAKLR 262
++S++ T R+LVRP RF T +LTL + ++ L F+ +W S ++
Sbjct: 339 DGWILSLMKKLTGGRELVRPKVARFMTHFLTLRSIVNQEDDLKHFFSHADWLSSVHSRRP 398
Query: 263 DGKALEDIILDKKFWKDIVICLKGATPLMKVLRLADLDKKTSHGIHL*SNGSSKGANSKE 322
D A++ ++ ++FWK + + PL+K+LRL D D I+ +K A
Sbjct: 399 DALAIKSLLYLERFWKSAHEIIGMSEPLLKLLRLVDGDMPAMGYIYE-GIERAKMAIKAF 457
Query: 323 F*WCQKE--PVWNIIDDRWDRMLHRPLHAAAYYLNPQLHYRPGFKADIEVRKGLMDCITR 380
+ C+++ V II+ RW H LHAAA +LNP + Y P FK D+ +R G + +
Sbjct: 458 YKGCEEKYMSVLEIIERRWSMHCHSHLHAAAAFLNPSIFYDPSFKFDVNMRNGFHAAMWK 517
Query: 381 MVEDPEEQAKIEV--QMHDFKKQIGYFGTKIAKLTLDKSTPADWWESVVFEYLELQRFA 437
M PEE +IE+ + K G G+K A + ++P DWW + +E LQR A
Sbjct: 518 MF--PEENDRIELIKDQPVYIKAQGALGSKFAIMGRTLNSPGDWWATYGYEIPVLQRAA 574
>UniRef100_Q8L717 Hypothetical protein At4g15020 [Arabidopsis thaliana]
Length = 768
Score = 247 bits (630), Expect = 5e-64
Identities = 145/417 (34%), Positives = 222/417 (52%), Gaps = 7/417 (1%)
Query: 24 SKEFNIMCDMISRHGNGFKPPSYYDIRVKLLNQEVKSTNDALEEHKREWKKTSCTIMTDE 83
S F M D I+ G G P++ D+R +L V+ ++E K WK+T C+I+ +E
Sbjct: 214 SVNFQPMIDAIASGGFGVSAPTHDDLRGWILKNCVEEMAKEIDECKAMWKRTGCSILVEE 273
Query: 84 WTDRRRRTILNFLVNSPRGTVFLKSVDASNICKTSEKIFEMIDAVVEEVGEDNVVQVVTD 143
+ +LNFLV P VFLKSVDAS + +++K+FE++ +VEEVG NVVQV+T
Sbjct: 274 LNSDKGFKVLNFLVYCPEKVVFLKSVDASEVLSSADKLFELLSELVEEVGSTNVVQVITK 333
Query: 144 NAANYKAAGEMLMRKRKKLYWTPCAAHCLDLMLEDLEKKIPIHGETIPKGKKITTFIYAR 203
Y AG+ LM LYW PCAAHC+D MLE+ K+ ETI + + IT F+Y
Sbjct: 334 CDDYYVDAGKRLMLVYPSLYWVPCAAHCIDQMLEEF-GKLGWISETIEQAQAITRFVYNH 392
Query: 204 TSLISILHSHTKNRDLVRPGATRFATSYLTLGCLYENKEALIRMFTSKEWKSGKCAKLRD 263
+ +++++ T D++ P + AT++ TLG + E K L M TS EW ++
Sbjct: 393 SGVLNLMWKFTSGNDILLPAFSSSATNFATLGRIAELKSNLQAMVTSAEWNECSYSEEPS 452
Query: 264 GKALEDIILDKKFWKDIVICLKGATPLMKVLRLADLDKKTSHGIHL*SNGSSKGANSKEF 323
G + + + D+ FWK + + +PL++ LR+ +K+ + G + +K A
Sbjct: 453 GLVM-NALTDEAFWKAVALVNHLTSPLLRALRIVCSEKRPAMGYVYAALYRAKDAIKTHL 511
Query: 324 *WCQKEP---VWNIIDDRWDRMLHRPLHAAAYYLNPQLHYRPGFKADIEVRKGLMDCITR 380
+E W IID W++ H PL AA ++LNP+L Y + E+ ++DCI R
Sbjct: 512 --VNREDYIIYWKIIDRWWEQQQHIPLLAAGFFLNPKLFYNTNEEMRSELILSVLDCIER 569
Query: 381 MVEDPEEQAKIEVQMHDFKKQIGYFGTKIAKLTLDKSTPADWWESVVFEYLELQRFA 437
+V D + Q KI ++ +K G FG +A D PA+WW + L L RFA
Sbjct: 570 LVPDDKIQDKIIKELTSYKTAGGVFGRNLAIRARDTMLPAEWWSTYGESCLNLSRFA 626
>UniRef100_Q9LIE1 Transposase-like protein [Arabidopsis thaliana]
Length = 761
Score = 245 bits (625), Expect = 2e-63
Identities = 141/407 (34%), Positives = 216/407 (52%), Gaps = 5/407 (1%)
Query: 32 DMISRHGNGFKPPSYYDIRVKLLNQEVKSTNDALEEHKREWKKTSCTIMTDEWTDRRRRT 91
D I G G P++ D+R +L V+ ++E K WK+T C+++ E
Sbjct: 218 DAIVSGGFGVSIPTHEDLRGWILKSCVEEVKKEIDECKTLWKRTGCSVLVQELNSNEGPL 277
Query: 92 ILNFLVNSPRGTVFLKSVDASNICKTSEKIFEMIDAVVEEVGEDNVVQVVTDNAANYKAA 151
IL FLV P VFLKSVDAS I + +K++E++ VVEE+G+ NVVQV+T +Y AA
Sbjct: 278 ILKFLVYCPEKVVFLKSVDASEILDSEDKLYELLKEVVEEIGDTNVVQVITKCEDHYAAA 337
Query: 152 GEMLMRKRKKLYWTPCAAHCLDLMLEDLEKKIPIHGETIPKGKKITTFIYARTSLISILH 211
G+ LM LYW PCAAHC+D MLE+ K I E I + + +T IY + +++++
Sbjct: 338 GKKLMDVYPSLYWVPCAAHCIDKMLEEFGKMDWIR-EIIEQARTVTRIIYNHSGVLNLMR 396
Query: 212 SHTKNRDLVRPGATRFATSYLTLGCLYENKEALIRMFTSKEWKSGKCAKLRDGKALEDII 271
T D+V+P T AT++ T+G + + K L M TS EW +K G A+ + I
Sbjct: 397 KFTFGNDIVQPVCTSSATNFTTMGRIADLKPYLQAMVTSSEWNDCSYSKEAGGLAMTETI 456
Query: 272 LDKKFWKDIVICLKGATPLMKVLRLADLDKKTSHGIHL*SNGSSKGANSKEF*WCQKEPV 331
D+ FWK + + P+++VLR+ ++K + G + +K A ++ V
Sbjct: 457 NDEDFWKALTLANHITAPILRVLRIVCSERKPAMGYVYAAMYRAKEAIKTNLAHREEYIV 516
Query: 332 -WNIIDDRWDRMLHRPLHAAAYYLNPQLHYRPGFKADIEVRKGLMDCITRMVEDPEEQAK 390
W IID W L +PL+AA +YLNP+ Y + E+ ++DCI ++V D Q
Sbjct: 517 YWKIIDRWW---LQQPLYAAGFYLNPKFFYSIDEEMRSEIHLAVVDCIEKLVPDVNIQDI 573
Query: 391 IEVQMHDFKKQIGYFGTKIAKLTLDKSTPADWWESVVFEYLELQRFA 437
+ ++ +K +G FG +A D PA+WW + L L RFA
Sbjct: 574 VIKDINSYKNAVGIFGRNLAIRARDTMLPAEWWSTYGESCLNLSRFA 620
>UniRef100_Q94LV8 Hypothetical protein OSJNBa0082M15.8 [Oryza sativa]
Length = 645
Score = 229 bits (584), Expect = 1e-58
Identities = 136/384 (35%), Positives = 199/384 (51%), Gaps = 55/384 (14%)
Query: 34 ISRHGNGFKPPSYYDIRVKLLNQEVKSTNDALEEHKREWKKTSCTIMTDEWTDRRRRTIL 93
I + G K PS Y++ L + + D +EHK W+ T C+IMTD WTDR+ R ++
Sbjct: 172 IGQFGRSLKGPSPYEMSGSFLQKRKEKVMDGFKEHKESWELTGCSIMTDAWTDRKGRGVM 231
Query: 94 NFLVNSPRGTVFLKSVDASNICKTSEKIFEMIDAVVEEVGEDNVVQVVTDNAANYKAAGE 153
N +V+S G +FL SV+ S K + IFE++D +EE+GE +VVQVVTDNA+ A
Sbjct: 232 NLVVHSAHGVLFLDSVECSGDRKDGKYIFELVDRYIEEIGEQHVVQVVTDNASVNTTAAS 291
Query: 154 MLMRKRKKLYWTPCAAHCLDLMLEDLEKKIPIHGETIPKGKKITTFIYARTSLISILHSH 213
+L KR ++W CAAHCLDLMLED+ K P+ ETI +++T F+YA T ++ ++
Sbjct: 292 LLTAKRPSIFWNGCAAHCLDLMLEDIGKLGPVE-ETIANARQVTVFLYAHTRVLDLMRKF 350
Query: 214 TKNRDLVRPGATRFATSYLTLGCLYENKEALIRMFTSKEWKSGKCAKLRDGKALEDIILD 273
NRDL + GK +I
Sbjct: 351 L-NRDLAK------------------------------------------GKKASKVIRS 367
Query: 274 KKFWKDIVICLKGATPLMKVLRLADLDKKTS---HGIHL*SNG--SSKGANSKEF*WCQK 328
+ FWK++ I + PL VLR D D + HG+ L + S + N K +
Sbjct: 368 ETFWKNVDIAVNYFEPLANVLRRMDSDVPSMGFFHGLMLEAKKEISQRFDNDKS----RF 423
Query: 329 EPVWNIIDDRWDRMLHRPLHAAAYYLNPQLHY--RPGFKADIEVRKGLMDCITRMVEDPE 386
VW+IID RWD L PLH A YYLNP +Y + ++D R G++ CI ++V+D +
Sbjct: 424 IEVWDIIDKRWDNKLKTPLHLAGYYLNPYYYYPNKQEIESDGSFRAGVISCIDKLVDDED 483
Query: 387 EQAKIEVQMHDFKKQIGYFGTKIA 410
Q KI +++ ++ Q G FG +IA
Sbjct: 484 IQDKIIEELNLYQDQHGSFGHEIA 507
>UniRef100_Q9LHU1 Similar to Arabidopsis thaliana chromosome 1 [Oryza sativa]
Length = 521
Score = 223 bits (569), Expect = 6e-57
Identities = 122/391 (31%), Positives = 212/391 (54%), Gaps = 9/391 (2%)
Query: 38 GNGFKPPSYYDIRVKLLNQEVKSTNDALEEHKREWKKTSCTIMTDEWTDRRRRTILNFLV 97
G GF+ PS ++ K L++ +E +++W T CTI+ D WTD + + ++NF V
Sbjct: 7 GQGFRGPSAEVLKTKWLHKLKSEVLQKTKEIEKDWATTGCTILADSWTDNKSKALINFSV 66
Query: 98 NSPRGTVFLKSVDASNICKTSEKIFEMIDAVVEEVGEDNVVQVVTDNAANYKAAGEMLMR 157
+SP GT FLK+VDAS K S +++E+ D V+ EVG DNVVQ++TD NY + +++M+
Sbjct: 67 SSPLGTFFLKTVDASPHIK-SHQLYELFDDVIREVGPDNVVQIITDRNINYGSVDKLIMQ 125
Query: 158 KRKKLYWTPCAAHCLDLMLEDLEKKIPIHGETIPKGKKITTFIYARTSLISILHSHTKNR 217
++W+PCA+ C++ ML+D KI I + + IT F+Y ++ ++ +
Sbjct: 126 NYNTIFWSPCASSCVNSMLDDF-SKIDWVNRCICQAQTITRFVYNNKWVLDLMRKCIAGQ 184
Query: 218 DLVRPGATRFATSYLTLGCLYENKEALIRMFTSKEWKSGKCAKLRDGKALEDIILDKKFW 277
+LV G T+ + +LTL L ++ L +MF S ++ S A + +I+ D +FW
Sbjct: 185 ELVCSGITKCVSDFLTLQSLLRHRPKLKQMFHSSDYASSSYANRSLSSSCVEILDDDEFW 244
Query: 278 KDIVICLKGATPLMKVLRLADLDKKTSHGIHL*SNGSSKGANSKEF*WCQKE----PVWN 333
+ + + PL++V+R K I+ +K +S + E +
Sbjct: 245 RAVEEIAAVSEPLLRVMRDVLGGKAAIGYIY---ESMTKVMDSIRTYYIMDEGKCKSFLD 301
Query: 334 IIDDRWDRMLHRPLHAAAYYLNPQLHYRPGFKADIEVRKGLMDCITRMVEDPEEQAKIEV 393
I++ +W LH PLH+AA +LNP + Y P K +++ + +++ P+++ I V
Sbjct: 302 IVEQKWQVELHSPLHSAAAFLNPSIQYNPEVKFFTSIKEEFYHVLDKVLTVPDQRQGITV 361
Query: 394 QMHDFKKQIGYFGTKIAKLTLDKSTPADWWE 424
++H F+K G FG+ IAK + ++P WWE
Sbjct: 362 ELHAFRKAQGMFGSNIAKEARNNTSPGMWWE 392
>UniRef100_Q9XG69 Hypothetical protein [Nicotiana tabacum]
Length = 593
Score = 212 bits (539), Expect = 2e-53
Identities = 106/267 (39%), Positives = 167/267 (61%), Gaps = 2/267 (0%)
Query: 24 SKEFNIMCDMISRHGNGFKPPSYYDIRVKLLNQEVKSTNDALEEHKREWKKTSCTIMTDE 83
S F M +++ ++G G PS + +LL E+ S + L E+K W T +I+ D
Sbjct: 327 SPYFQKMLELVGQYGEGLVGPSSRVLSGRLLQDEIVSIRNYLSEYKASWAVTGYSILADS 386
Query: 84 WTDRRRRTILNFLVNSPRGTVFLKSVDASNICKTSEKIFEMIDAVVEEVGEDNVVQVVTD 143
W D + RT++N LV+ P G F+ SVDA+ + + + IF+++D VVE++GE+NVVQV+T
Sbjct: 387 WQDTQGRTLINVLVSCPHGMYFVCSVDATGVVEDATYIFKLLDRVVEDMGEENVVQVITQ 446
Query: 144 NAANYKAAGEMLMRKRKKLYWTPCAAHCLDLMLEDLEKKIPIHGETIPKGKKITTFIYAR 203
N NY+AAG+ML KR+ L+WTPCAA+C+D +LED KI E + K +KIT FIY
Sbjct: 447 NTPNYQAAGKMLEEKRRNLFWTPCAAYCIDRILEDF-VKIKWVRECMEKAQKITKFIYNS 505
Query: 204 TSLISILHSH-TKNRDLVRPGATRFATSYLTLGCLYENKEALIRMFTSKEWKSGKCAKLR 262
L+S++ T ++L++P TR+++++ T+ L +++ L RMF S +W S + +KL
Sbjct: 506 FWLLSLMKKEFTAGQELLKPSFTRYSSTFATVQSLLDHRNGLKRMFQSNKWLSSRYSKLE 565
Query: 263 DGKALEDIILDKKFWKDIVICLKGATP 289
DGK +E I+L+ FW+ + K P
Sbjct: 566 DGKEVEKIVLNATFWRKMQYVRKSVDP 592
>UniRef100_Q852A8 Hypothetical protein OSJNBb0081B07.17 [Oryza sativa]
Length = 779
Score = 207 bits (527), Expect = 5e-52
Identities = 127/401 (31%), Positives = 203/401 (49%), Gaps = 24/401 (5%)
Query: 27 FNIMCDMISRHGNGFKPPSYYDIRVKLLNQEVKSTNDALEEHKREWKKTSCTIMTDEWTD 86
F M + ++ G + SY+D R +L + + LE +K W +T CT++ DEWT
Sbjct: 250 FQPMLEAVASAGGKPEAFSYHDFRGSILKKSLDEVTAQLEFYKGSWTRTGCTLLADEWTT 309
Query: 87 RRRRTILNFLVNSPRGTVFLKSVDASNICKTSEKIFEMIDAVVEEVGEDNVVQVVTDNAA 146
R RT++NF V P + ++E++ VVEEVGE NVVQV+T+N+
Sbjct: 310 DRGRTLINFSVYCPE-----------------DPLYELLKNVVEEVGEKNVVQVITNNSE 352
Query: 147 NYKAAGEMLMRKRKKLYWTPCAAHCLDLMLEDLEKKIPIHGETIPKGKKITTFIYARTSL 206
+ AG+ L L+W+ C+ C+D MLED K I+ E I K IT FIY
Sbjct: 353 IHAVAGKRLCETFPTLFWSQCSFQCIDGMLEDFSKVGAIN-EIICNAKVITGFIYNSAFA 411
Query: 207 ISILHSHTKNRDLVRPGATRFATSYLTLGCLYENKEALIRMFTSKEWKSGKCAKLRDGKA 266
+++ H +DL+ P TR A +++TL +Y K++L M +S EW K G
Sbjct: 412 FNLMKRHLHGKDLLVPAETRAAMNFVTLKNMYNLKDSLEAMISSDEWIHYLLPKKPGGVE 471
Query: 267 LEDIILDKKFWKDIVICLKGATPLMKVLRLADLDKKTSHGIHL*SNGSSKGANSKEF*WC 326
+ ++I + +FW ++ PL+ +L+L +K+ S G +K A KE
Sbjct: 472 VTNLIGNLQFWSSCAAVVRITEPLVHLLKLVGSNKRPSMGYVYAGLYQAKAAIKKEL--V 529
Query: 327 QKE---PVWNIIDDRWDRMLHRPLHAAAYYLNPQLHYRPGFKADIEVRKGLMDCITRMVE 383
+K W+IID RW++ RPLH A ++LNP E+ G++DC+ R+V
Sbjct: 530 RKNDYMAYWDIIDWRWNKDAPRPLHLAGFFLNPLFFDGVRGGTSSEIFSGMLDCVERLVS 589
Query: 384 DPEEQAKIEVQMHDFKKQ-IGYFGTKIAKLTLDKSTPADWW 423
D + Q KI+ +++ ++ + G F ++A PA+WW
Sbjct: 590 DVKIQDKIQKELNVYRSEAAGDFRRQMAIRARHTLPPAEWW 630
>UniRef100_Q9SQN5 Hypothetical protein F19K16.28 [Arabidopsis thaliana]
Length = 518
Score = 200 bits (508), Expect = 8e-50
Identities = 114/409 (27%), Positives = 212/409 (50%), Gaps = 7/409 (1%)
Query: 30 MCDMISRHGNGFKPPSYYDIRVKLLNQEVKSTNDALEEHKREWKKTSCTIMTDEWTDRRR 89
M D +++ G GF PS + + L++ + L++ ++EW T CTI+ + WTD +
Sbjct: 1 MLDAVAKCGPGFVAPSP---KTEWLDRVKSDISLQLKDTEKEWVTTGCTIIAEAWTDNKS 57
Query: 90 RTILNFLVNSPRGTVFLKSVDASNICKTSEKIFEMIDAVVEEVGEDNVVQVVTDNAANYK 149
R ++NF V+SP F KSVDAS+ K S+ + ++ D+V++++G++++VQ++ DN+ Y
Sbjct: 58 RALINFSVSSPSRIFFHKSVDASSYFKNSKCLADLFDSVIQDIGQEHIVQIIMDNSFCYT 117
Query: 150 AAGEMLMRKRKKLYWTPCAAHCLDLMLEDLEKKIPIHGETIPKGKKITTFIYARTSLISI 209
L++ ++ +PCA+ CL+++LE+ K+ + I + + I+ F+Y + ++ +
Sbjct: 118 GISNHLLQNYATIFVSPCASQCLNIILEEF-SKVDWVNQCISQAQVISKFVYNNSPVLDL 176
Query: 210 LHSHTKNRDLVRPGATRFATSYLTLGCLYENKEALIRMFTSKEWKSGKCAKLRDGKALED 269
L T +D++R G TR +++L+L + + K L MF E+ + + +
Sbjct: 177 LRKLTGGQDIIRSGVTRSVSNFLSLQSMMKQKARLKHMFNCPEYTTN--TNKPQSISCVN 234
Query: 270 IILDKKFWKDIVICLKGATPLMKVLRLADLDKKTSHGIHL*SNGSSKGANSKEF*WCQKE 329
I+ D FW+ + + + P++KVLR K I+ + + + + K
Sbjct: 235 ILEDNDFWRAVEESVAISEPILKVLREVSTGKPAVGSIYELMSKAKESIRTYYIMDENKH 294
Query: 330 PVW-NIIDDRWDRMLHRPLHAAAYYLNPQLHYRPGFKADIEVRKGLMDCITRMVEDPEEQ 388
V+ +I+D W LH PLHAAA +LNP + Y P K +++ + +++ + +
Sbjct: 295 KVFSDIVDTNWCEHLHSPLHAAAAFLNPSIQYNPEIKFLTSLKEDFFKVLEKLLPTSDLR 354
Query: 389 AKIEVQMHDFKKQIGYFGTKIAKLTLDKSTPADWWESVVFEYLELQRFA 437
I Q+ F + G FG +A D +P WWE LQR A
Sbjct: 355 RDITNQIFTFTRAKGMFGCNLAMEARDSVSPGLWWEQFGDSAPVLQRVA 403
>UniRef100_Q9LDS0 Transposase-like protein [Arabidopsis thaliana]
Length = 544
Score = 199 bits (507), Expect = 1e-49
Identities = 119/402 (29%), Positives = 195/402 (47%), Gaps = 11/402 (2%)
Query: 40 GFKPPSYYDIRVKLLNQEVKSTNDALEEHKREWKKTSCTIMTDEWTDRRRRTILNFLVNS 99
G + +D+ L ++ D +E+ K W T C+I+ D W D++ R ++ F+ +
Sbjct: 84 GLESSDCHDLNGWRLQDALEEVQDRVEKIKESWAITGCSILLDAWVDQKGRDLVTFVADC 143
Query: 100 PRGTVFLKSVDASNICKTSEKIFEMIDAVVEEVGEDNVVQVVTDNAANYKAA-GEMLMRK 158
P G V+L S D S+ + +++ +VEEVG NV Q++ + + + GE+
Sbjct: 144 PAGLVYLISFDVSDFKDDVTALLSLVNGLVEEVGVRNVTQIIACSTSGWVGELGELFAGH 203
Query: 159 RKKLYWTPCAAHCLDLMLEDLEKKIPIHGETIPKGKKITTFIYARTSLISILHSHTKNRD 218
++++W+ +HC +LML + K I G+ K I FI S+++I D
Sbjct: 204 DREVFWSVSVSHCFELMLVKISK-IRSFGDIFDKVNNIWLFINNNPSVLNIFRDQCHGID 262
Query: 219 L-VRPGATRFATSYLTLGCLYENKEALIRMFTSKEWKSGKCAKLRDGKALEDIILDKKFW 277
+ V F T YL L +++ K+ L MF S W + +C A+ +++ D FW
Sbjct: 263 ITVSSSEFEFVTPYLILESIFKAKKNLTAMFASSNWNNEQCI------AISNLVSDSSFW 316
Query: 278 KDIVICLKGATPLMKVLRLADLDKKTSHGIHL*SNGSSKGANSKEF*WCQK--EPVWNII 335
+ + LK +PL+ L L G + S K + ++EF + +P+W++I
Sbjct: 317 ETVESVLKCTSPLIHGLLLFSTANNQHLGYVYDTMDSIKESIAREFNHKPQFYKPLWDVI 376
Query: 336 DDRWDRMLHRPLHAAAYYLNPQLHYRPGFKADIEVRKGLMDCITRMVEDPEEQAKIEVQM 395
DD W++ LH PLHAA Y+LNP Y F DIEV GL+ + MVED Q KI Q+
Sbjct: 377 DDVWNKHLHNPLHAAGYFLNPTAFYSTNFHLDIEVVTGLISSLIHMVEDCHVQFKISTQI 436
Query: 396 HDFKKQIGYFGTKIAKLTLDKSTPADWWESVVFEYLELQRFA 437
++ F + +PA+WW +Y ELQ A
Sbjct: 437 DMYRLGKDCFNEASQADQITGISPAEWWAHKASQYPELQSLA 478
>UniRef100_Q5W690 Hypothetical protein OSJNBa0076A09.5 [Oryza sativa]
Length = 1030
Score = 196 bits (498), Expect = 1e-48
Identities = 122/352 (34%), Positives = 185/352 (51%), Gaps = 24/352 (6%)
Query: 87 RRRRTILNFLVNSPRGTVFLKSVDASNICKTSEKIFEMIDAVVEEVGEDNVVQVVTDNAA 146
RR R+++N + + RG F+ ++DAS + I+ ++ + ++E+G VVQVVTDNA+
Sbjct: 521 RRGRSLMNLVAHCARGMCFIDAIDASLEVHDGKYIYSLVSSCIDEIGPKKVVQVVTDNAS 580
Query: 147 NYKAAGEMLMRKRKKLYWTPCAAHCLDLMLEDLEKKIPIHGETIPKGKKITTFIYARTSL 206
N +A +ML K ++WT CAAH +DLMLED+ K +H I GK IT +YA+ L
Sbjct: 581 NNMSASKMLQVKHPHIFWTSCAAHYIDLMLEDIGKITMVH-NIIRDGKSITNLLYAQVRL 639
Query: 207 ISILHSHTKNRDLVRPGATRFATSYLTLGCLYENKEALIRMFTSKEWKSGKCAKLRDGKA 266
++I+ TK DLVR G TRFATSYL L LY+ + L ++F S++W AK G+
Sbjct: 640 LAIMRQFTKG-DLVRAGTTRFATSYLNLKSLYDKRNELKQLFASQDWAKSSWAKKIKGQN 698
Query: 267 LEDIILDKKFWKDIVICLKGATPLMKVLRLADLDKKTSHGIHL*SNGSSK------GANS 320
+++++ KFW ++ + PL VLR D D I+ + K N
Sbjct: 699 AHNLVMNNKFWSQMLEVINYFEPLAYVLRRVDGDVPAMGYIYGDLIKAKKDVAACLNGNE 758
Query: 321 KEF*WCQKEPVWNIIDDRWDRMLHRPLHAAAYYLNPQLHY--RPGFKADIEVRKGLMDCI 378
K++ +W IID RWD L LH A Y+LNP Y + K D + + +++C
Sbjct: 759 KKY-----SHIWKIIDARWDSKLKTTLHKAGYFLNPCFFYENKREIKEDF-LMQAVVECA 812
Query: 379 TRMVEDPEEQAKIEV-QMHDFKKQIGYFGTKIA-------KLTLDKSTPADW 422
T M D I V Q+ + + + FGT +A +T D P +W
Sbjct: 813 TCMYRDDITVQDICVAQLSLYTEAMDSFGTTMAIRQRNSPTITPDDDPPNEW 864
>UniRef100_Q9LDK4 Transposase-like protein [Arabidopsis thaliana]
Length = 605
Score = 196 bits (497), Expect = 1e-48
Identities = 122/411 (29%), Positives = 202/411 (48%), Gaps = 14/411 (3%)
Query: 32 DMISRHGNGFKPPSYYDIRVKLLNQEVKSTNDALEEHKREWKKTSCTIMTDEWTDRRRRT 91
+M+ G G K P +D+ +LL + +K D ++ K WK T C+I+ D W D +
Sbjct: 149 EMMMALGVGQKIPDSHDLNGRLLQEAMKEVQDYVKNIKDSWKITGCSILLDAWIDPKGHD 208
Query: 92 ILNFLVNSPRGTVFLKSVDASNICKTSEKIFEMIDAVVEEVGEDNVVQVVTDNAANYKA- 150
+++F+ + P G V+LKS+D S + + +++ +VEEVG NV Q++ + + +
Sbjct: 209 LVSFVADCPAGPVYLKSIDVSVVKNDVTALLSLVNGLVEEVGVHNVTQIIACSTSGWVGE 268
Query: 151 AGEMLMRKRKKLYWTPCAAHCLDLMLEDLEKKIPIHGETIPKGKKITTFIYARTSLISIL 210
G++ ++++W+ +HC +LML + K+ G+ + K I FI S + I
Sbjct: 269 LGKLFSGHDREVFWSVSLSHCFELMLVKI-GKMRSFGDILDKVNTIWEFINNNPSALKIY 327
Query: 211 HSHTKNRDL-VRPGATRFATSYLTLGCLYENKEALIRMFTSKEWKSGKCAKLRDGKALED 269
+ +D+ V F YL L +++ K+ L MF S WK +GK++ +
Sbjct: 328 RDQSHGKDITVSSSEFEFVKPYLILKSVFKAKKNLAAMFASSVWKK------EEGKSVSN 381
Query: 270 IILDKKFWKDIVICLKGATPLMKVLRL-ADLDKKTSHGIHL*SNGSSKGANSKEF*WCQK 328
++ D FW+ + LK +PL LRL ++ D G + K + KEF +K
Sbjct: 382 LVNDSSFWEAVEEILKCTSPLTDGLRLFSNADNNQHVGYIYDTLDGIKLSIKKEFNDEKK 441
Query: 329 E--PVWNIIDDRWDRMLHRPLHAAAYYLNPQLHYRPGFKADIEVRKGLMDCITRMVEDPE 386
+W++IDD W++ LH PLHAA YYLNP Y F D EV GL + + + E
Sbjct: 442 HYLTLWDVIDDVWNKHLHNPLHAAGYYLNPTSFYSTDFHLDPEVSSGLTHSLVHVAK--E 499
Query: 387 EQAKIEVQMHDFKKQIGYFGTKIAKLTLDKSTPADWWESVVFEYLELQRFA 437
Q KI Q+ ++ F + +P DWW ++ ELQ FA
Sbjct: 500 GQIKIASQLDRYRLGKDCFNEASQPDQISGISPIDWWTEKASQHPELQSFA 550
>UniRef100_Q6F2P4 Hypothetical protein OSJNBa0075J06.6 [Oryza sativa]
Length = 217
Score = 195 bits (495), Expect = 2e-48
Identities = 96/213 (45%), Positives = 135/213 (63%), Gaps = 2/213 (0%)
Query: 30 MCDMISRHGNGFKPPSYYDIRVKLLNQEVKSTNDALEEHKREWKKTSCTIMTDEWTDRRR 89
M + I + G K PS Y++ L + + D +EHK W+ T C+IMTD WTDR+
Sbjct: 1 MLEAIGQFGRSLKGPSPYEMSGSFLQKRKEKVMDGFKEHKESWELTGCSIMTDAWTDRKG 60
Query: 90 RTILNFLVNSPRGTVFLKSVDASNICKTSEKIFEMIDAVVEEVGEDNVVQVVTDNAANYK 149
R ++N +V+S G +FL SV+ S K + IFE++D +EE+GE +VVQVVTDNA+
Sbjct: 61 RGVMNLVVHSAHGVLFLDSVECSGDRKDDKYIFELVDRYIEEIGEQHVVQVVTDNASVNT 120
Query: 150 AAGEMLMRKRKKLYWTPCAAHCLDLMLEDLEKKIPIHGETIPKGKKITTFIYARTSLISI 209
A +L KR ++W CAAHCLDLMLED+ K P+ ETI +++T F+YA T ++ +
Sbjct: 121 TAASLLTAKRPSIFWNGCAAHCLDLMLEDIGKLGPVE-ETIANARQVTVFLYAHTRVLDL 179
Query: 210 LHSHTKNRDLVRPGATRFATSYLTLGCLYENKE 242
+ NRDLVR G TRFAT+YL L L +NK+
Sbjct: 180 MRKFL-NRDLVRSGVTRFATAYLNLKSLLDNKK 211
>UniRef100_Q6F2S1 Hypothetical protein OSJNBa0060G17.6 [Oryza sativa]
Length = 217
Score = 191 bits (484), Expect = 5e-47
Identities = 95/213 (44%), Positives = 134/213 (62%), Gaps = 2/213 (0%)
Query: 30 MCDMISRHGNGFKPPSYYDIRVKLLNQEVKSTNDALEEHKREWKKTSCTIMTDEWTDRRR 89
M + I + G K PS Y++ L + + D +EHK W+ T C+IMTD WTDR+
Sbjct: 1 MLEAIGQFGRSLKGPSPYEMSGSFLQKRKEKVMDGFKEHKESWELTGCSIMTDAWTDRKG 60
Query: 90 RTILNFLVNSPRGTVFLKSVDASNICKTSEKIFEMIDAVVEEVGEDNVVQVVTDNAANYK 149
R ++N +V+S G +FL SV+ S K + IFE++D +EE+GE +VVQVVTDNA+
Sbjct: 61 RGVMNLVVHSAHGVLFLDSVECSGDRKDGKYIFELVDRYIEEIGEQHVVQVVTDNASVNT 120
Query: 150 AAGEMLMRKRKKLYWTPCAAHCLDLMLEDLEKKIPIHGETIPKGKKITTFIYARTSLISI 209
A +L KR ++W AAHCLDLMLED+ K P+ ETI +++T F+YA T ++ +
Sbjct: 121 TAASLLTAKRPSIFWNGYAAHCLDLMLEDIGKLGPVE-ETIANARQVTVFLYAHTRVLDL 179
Query: 210 LHSHTKNRDLVRPGATRFATSYLTLGCLYENKE 242
+ NRDLVR G TRFAT+YL L L +NK+
Sbjct: 180 MRKFL-NRDLVRSGVTRFATAYLNLKSLLDNKK 211
>UniRef100_O23355 Hypothetical protein AT4g15020 [Arabidopsis thaliana]
Length = 522
Score = 167 bits (424), Expect = 4e-40
Identities = 112/340 (32%), Positives = 168/340 (48%), Gaps = 25/340 (7%)
Query: 24 SKEFNIMCDMISRHGNGFKPPSYYDIRVKLLNQEVKSTNDALEEHKREWKKTSCTIMTDE 83
S F M D I+ G G P++ D+R +L V+ ++E K WK+T C+I+ +E
Sbjct: 198 SVNFQPMIDAIASGGFGVSAPTHDDLRGWILKNCVEEMAKEIDECKAMWKRTGCSILVEE 257
Query: 84 WTDRRRRTILNFLVNSPRGTVFLKSVDASNICKTSEKIFEMIDAVVEEVGEDNVVQVVTD 143
+ +LNFLV P VFLKSVDAS + +++K+FE++ +VEEVG NVVQV+T
Sbjct: 258 LNSDKGFKVLNFLVYCPEKVVFLKSVDASEVLSSADKLFELLSELVEEVGSTNVVQVITK 317
Query: 144 NAANYKAAGEMLMRKRKKLYWTPCAAHCLDLMLEDLEKKIPIHGETIPKGKKITTFIYAR 203
Y AG+ LM LYW PCAAHC+D MLE+ K+ ETI + + IT F+Y
Sbjct: 318 CDDYYVDAGKRLMLVYPSLYWVPCAAHCIDQMLEEF-GKLGWISETIEQAQAITRFVYNH 376
Query: 204 TSLISILHSHTKNRDLVRPGATRFATSYLTLGCLYENKEALIRMFTSKEWKSGKCAKLRD 263
++ S AT++ TLG + E K L M TS EW ++
Sbjct: 377 SAFSS------------------SATNFATLGRIAELKSNLQAMVTSAEWNECSYSEEPS 418
Query: 264 GKALEDIILDKKFWKDIVICLKGATPLMKVLRLADLDKKTSHGIHL*SNGSSKGANSKEF 323
G + + + D+ FWK + + +PL++ LR+ +K+ + G + +K A
Sbjct: 419 GLVM-NALTDEAFWKAVALVNHLTSPLLRALRIVCSEKRPAMGYVYAALYRAKDAIKTHL 477
Query: 324 *WCQKEP---VWNIIDDRWDRMLHRPLHAAAYYLNPQLHY 360
+E W IID ++LNP+L Y
Sbjct: 478 --VNREDYIIYWKIIDHGGSNNSIFLCLLLGFFLNPKLFY 515
>UniRef100_Q9LDB0 Transposase-like protein [Arabidopsis thaliana]
Length = 572
Score = 164 bits (415), Expect = 5e-39
Identities = 109/410 (26%), Positives = 186/410 (44%), Gaps = 15/410 (3%)
Query: 29 IMCDMISRHGNGFKPPSYYDIRVKLLNQEVKSTNDALEEHKREWKKTSCTIMTDEWTDRR 88
+M + G G P D+ + + +K D ++E K W+ T C+I+ D W +
Sbjct: 122 MMIKTVGGEGGGQMIPDSRDLNGWMFQEALKKVQDRVKEIKASWEITGCSILFDAWIGPK 181
Query: 89 RRTILNFLVNSPRGTVFLKSVDASNICKTSEKIFEMIDAVVEEVGEDNVVQVVTDNAANY 148
R ++ F+ + P G V+LKS D S+I + +++ +VEEVG NV Q++ + + +
Sbjct: 182 GRDLVTFVADCPAGAVYLKSADVSDIKTDVTALTSLVNGIVEEVGVRNVTQIIACSTSGW 241
Query: 149 KAAGEMLMRKRKKLYWTPCAAHCLDLMLEDLEKKIPIHGETIPKGKKITTFIYARTSLIS 208
G++ + +++W+ ++CL LML ++ K + K K + I S +
Sbjct: 242 --VGDLGKQLAGQVFWSVSLSYCLKLMLVEIGKMYSFE-DIFEKVKLLLDLINNNPSFLY 298
Query: 209 ILHSHTKNRDLVRPGATRFATSYLTLGCLYENKEALIRMFTSKEWKSGKCAKLRDGKALE 268
+ ++ D+ F YLTL +Y + A +F S EWK G A+
Sbjct: 299 VFRENSHKVDV--SSECEFVMPYLTLEHIYWVRRA--GLFASPEWKK------EQGIAIS 348
Query: 269 DIILDKKFWKDIVICLKGATPLMK-VLRLADLDKKTSHGIHL*SNGSSKGANSKEF*WCQ 327
+ D FW+ + + + L+ L + K ++ H + A + ++
Sbjct: 349 SFVNDSTFWESLDKIVGSTSSLVHGWLWFSRGSKHVAYAYHFIESIKKNVAWTFKYERQF 408
Query: 328 KEPVWNIIDDRWDRMLHRPLHAAAYYLNPQLHYRPGFKADIEVRKGLMDCITRMVEDPEE 387
EP WN+IDD W H PLHAA Y+LNP +Y F V GL + +V++P
Sbjct: 409 YEPTWNVIDDVWHNN-HNPLHAAGYFLNPMAYYSDDFHTYQHVYTGLAFSLVHLVKEPHL 467
Query: 388 QAKIEVQMHDFKKQIGYFGTKIAKLTLDKSTPADWWESVVFEYLELQRFA 437
Q KI Q+ ++ G F L+ +P +WW +Y ELQ A
Sbjct: 468 QVKIGTQLDVYRYGRGCFMKASQAGQLNGVSPVNWWTQKANQYPELQNLA 517
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.325 0.138 0.432
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 719,909,374
Number of Sequences: 2790947
Number of extensions: 29584867
Number of successful extensions: 90072
Number of sequences better than 10.0: 58
Number of HSP's better than 10.0 without gapping: 31
Number of HSP's successfully gapped in prelim test: 27
Number of HSP's that attempted gapping in prelim test: 89933
Number of HSP's gapped (non-prelim): 71
length of query: 437
length of database: 848,049,833
effective HSP length: 130
effective length of query: 307
effective length of database: 485,226,723
effective search space: 148964603961
effective search space used: 148964603961
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 76 (33.9 bits)
Medicago: description of AC147430.8


