

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
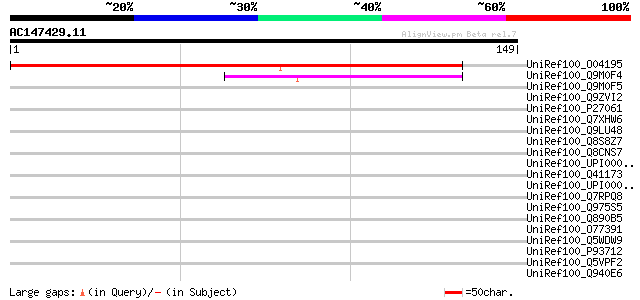
Query= AC147429.11 - phase: 0
(149 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_O04195 Expressed protein [Arabidopsis thaliana] 130 5e-30
UniRef100_Q9M0F4 Acid phosphatase-like protein [Arabidopsis thal... 49 2e-05
UniRef100_Q9M0F5 Putative acid phosphatase [Arabidopsis thaliana] 41 0.005
UniRef100_Q9ZVI2 Putative acid phosphatase [Arabidopsis thaliana] 40 0.011
UniRef100_P27061 Acid phosphatase 1 precursor [Lycopersicon escu... 40 0.014
UniRef100_Q7XHW6 Putative syringolide-induced protein [Oryza sat... 38 0.040
UniRef100_Q9LU48 Acid phosphatase [Arabidopsis thaliana] 38 0.040
UniRef100_Q8S8Z7 Syringolide-induced protein B15-3-5 [Glycine max] 37 0.068
UniRef100_Q8CNS7 Replication-associated protein [Staphylococcus ... 36 0.20
UniRef100_UPI00002D6890 UPI00002D6890 UniRef100 entry 35 0.26
UniRef100_Q41173 Acid phosphatase-1 [Lycopersicon esculentum] 35 0.34
UniRef100_UPI000027D636 UPI000027D636 UniRef100 entry 34 0.76
UniRef100_Q7RPQ8 Hypothetical protein [Plasmodium yoelii yoelii] 34 0.76
UniRef100_Q975S5 Probable spermidine synthase [Sulfolobus tokodaii] 34 0.76
UniRef100_Q890B5 Amino acid transport protein [Lactobacillus pla... 33 0.99
UniRef100_O77391 Hypothetical protein MAL3P6.4 [Plasmodium falci... 33 0.99
UniRef100_Q5WDW9 D-alanine--D-alanine ligase A [Bacillus clausii] 33 1.3
UniRef100_P93712 Pod storage protein [Phaseolus vulgaris] 33 1.3
UniRef100_Q5VPF2 Putative acid phosphatase [Oryza sativa] 33 1.3
UniRef100_Q940E6 Putative defense associated acid phosphatase [P... 33 1.3
>UniRef100_O04195 Expressed protein [Arabidopsis thaliana]
Length = 283
Score = 130 bits (328), Expect = 5e-30
Identities = 63/134 (47%), Positives = 94/134 (70%), Gaps = 1/134 (0%)
Query: 1 MGSSFVLESGFYITSFSATIFIAGFAALGLLLVTLLVSMAMMLQSCQNSNGGIIELRNIN 60
+GS + +ESG Y+TS +A+IFIA G+L++TLL++++ MLQSC+N N I+E + ++
Sbjct: 23 LGSRYSIESGCYMTSLAASIFIASLVTFGVLMITLLIALSTMLQSCENRNIAIVEAQRLD 82
Query: 61 NDYSYCKIHSLHAKLNNL-EEHNVPNICKDLALQYIKGGQYARDLDSTKSVIEDYFNGVK 119
+ YCKI SLH++LN+L EE +P +C+D+AL IK G Y R+L+ T + YF +K
Sbjct: 83 ESFGYCKILSLHSQLNSLDEESELPLLCRDVALHRIKQGIYLRELNFTIQMALTYFQTIK 142
Query: 120 PSEDGFDVVLIDID 133
P D DVV+IDID
Sbjct: 143 PMNDNCDVVVIDID 156
>UniRef100_Q9M0F4 Acid phosphatase-like protein [Arabidopsis thaliana]
Length = 256
Score = 48.9 bits (115), Expect = 2e-05
Identities = 28/71 (39%), Positives = 37/71 (51%), Gaps = 1/71 (1%)
Query: 64 SYCKIHSLHAKLNNLEEHNV-PNICKDLALQYIKGGQYARDLDSTKSVIEDYFNGVKPSE 122
SYC+ L A+ NN+ V P+ C++ YI GGQ+ +D D S DY VK
Sbjct: 42 SYCESWRLAAETNNVGPWKVIPSQCENYIKNYINGGQFDKDYDVVASYAIDYAKTVKVGG 101
Query: 123 DGFDVVLIDID 133
DG D + DID
Sbjct: 102 DGKDAWVFDID 112
>UniRef100_Q9M0F5 Putative acid phosphatase [Arabidopsis thaliana]
Length = 255
Score = 41.2 bits (95), Expect = 0.005
Identities = 29/80 (36%), Positives = 39/80 (48%), Gaps = 9/80 (11%)
Query: 65 YCKIHSLHAKLNNLEEHN-VPNICKDLALQYIKGGQYARDLDSTKSVIEDYF----NGVK 119
YC L A+ NN+ + +P+IC D +Y+ G Q+ D SVI DY V+
Sbjct: 42 YCDSWRLAAETNNVGTWDLIPSICVDSVAEYLNGDQFLSDY----SVIVDYALAFAKSVE 97
Query: 120 PSEDGFDVVLIDIDSLFQWN 139
S DG DV + DID N
Sbjct: 98 ISGDGKDVWIFDIDETLLTN 117
>UniRef100_Q9ZVI2 Putative acid phosphatase [Arabidopsis thaliana]
Length = 251
Score = 40.0 bits (92), Expect = 0.011
Identities = 25/75 (33%), Positives = 34/75 (45%), Gaps = 1/75 (1%)
Query: 60 NNDYSYCKIHSLHAKLNNLEEHN-VPNICKDLALQYIKGGQYARDLDSTKSVIEDYFNGV 118
N SYC L + NN+ VP C Y+ GQY RD+ T I+ Y N +
Sbjct: 32 NYGASYCLSWRLAVETNNVRAWRIVPLQCLRYVEVYMLAGQYDRDVQLTVDQIKVYLNEI 91
Query: 119 KPSEDGFDVVLIDID 133
DG D ++D+D
Sbjct: 92 ILPGDGMDAWILDVD 106
>UniRef100_P27061 Acid phosphatase 1 precursor [Lycopersicon esculentum]
Length = 255
Score = 39.7 bits (91), Expect = 0.014
Identities = 22/81 (27%), Positives = 34/81 (41%), Gaps = 1/81 (1%)
Query: 66 CKIHSLHAKLNNLEE-HNVPNICKDLALQYIKGGQYARDLDSTKSVIEDYFNGVKPSEDG 124
C + NNL +P C D +Y+ G Y ++D +Y V +DG
Sbjct: 43 CTTWRFVVETNNLSPWKTIPEECADYVKEYMVGPGYKMEIDRVSDEAGEYAKSVDLGDDG 102
Query: 125 FDVVLIDIDSLFQWNPPHSSN 145
DV + D+D N P+ S+
Sbjct: 103 RDVWIFDVDETLLSNLPYYSD 123
>UniRef100_Q7XHW6 Putative syringolide-induced protein [Oryza sativa]
Length = 244
Score = 38.1 bits (87), Expect = 0.040
Identities = 22/73 (30%), Positives = 32/73 (43%), Gaps = 1/73 (1%)
Query: 62 DYSYCKIHSLHAKLNNLEEH-NVPNICKDLALQYIKGGQYARDLDSTKSVIEDYFNGVKP 120
D YC + + NN + VP C +Y+ GQYARD+ I Y +
Sbjct: 29 DEPYCLSWRVMVEANNAKNWPTVPPPCVGYVWRYMAWGQYARDVAGVADQIAAYAAQLAA 88
Query: 121 SEDGFDVVLIDID 133
+DG D + D+D
Sbjct: 89 GDDGLDAWVFDVD 101
>UniRef100_Q9LU48 Acid phosphatase [Arabidopsis thaliana]
Length = 257
Score = 38.1 bits (87), Expect = 0.040
Identities = 22/79 (27%), Positives = 34/79 (42%), Gaps = 1/79 (1%)
Query: 65 YCKIHSLHAKLNNLEE-HNVPNICKDLALQYIKGGQYARDLDSTKSVIEDYFNGVKPSED 123
+C A++NNL +P C D Y+ G Y DL+ + ++ S D
Sbjct: 44 HCTTWRFAAEMNNLAPWKTIPVECADYVKDYVMGKGYLTDLERVSEEALIFARSIEFSGD 103
Query: 124 GFDVVLIDIDSLFQWNPPH 142
G D+ + DID N P+
Sbjct: 104 GKDIWIFDIDETLLSNLPY 122
>UniRef100_Q8S8Z7 Syringolide-induced protein B15-3-5 [Glycine max]
Length = 234
Score = 37.4 bits (85), Expect = 0.068
Identities = 22/71 (30%), Positives = 34/71 (46%), Gaps = 1/71 (1%)
Query: 64 SYCKIHSLHAKLNNLEEHN-VPNICKDLALQYIKGGQYARDLDSTKSVIEDYFNGVKPSE 122
SY + L + NN VP+ C + Y+ GGQY DL+ I Y + + +
Sbjct: 19 SYGRSWRLTVEANNARPWRIVPDNCYNHLQNYMSGGQYQLDLNLVVQHILSYAHEIPLAA 78
Query: 123 DGFDVVLIDID 133
DG D ++D+D
Sbjct: 79 DGMDAWILDVD 89
>UniRef100_Q8CNS7 Replication-associated protein [Staphylococcus epidermidis]
Length = 285
Score = 35.8 bits (81), Expect = 0.20
Identities = 33/137 (24%), Positives = 61/137 (44%), Gaps = 17/137 (12%)
Query: 8 ESGFYITSFSATIFIAGFAALGLLLVTLLVSMAMMLQSCQN-------SNGGIIELRNIN 60
++G Y+ + AT + L+ + V M L + SN I+E++N+
Sbjct: 76 DNGLYVLKYKATSTANKTVVVNSELLNVNVFMEKGLTKLNDVKVFYPLSNYFIVEIKNLP 135
Query: 61 NDYSYCKIHSLHAKLNNLEEHNVPNICKDLALQYIKGGQYARDLDSTKSVIEDYFNGVKP 120
DY KI + KL + E+ NV + K+++ QY + RD + +Y+
Sbjct: 136 KDYKALKI---NFKLLDKEDENVSSNSKEMS-QYTSAEKNNRDYNLQPKTKSEYY----- 186
Query: 121 SEDGFDVVLIDIDSLFQ 137
D F + L ++DS ++
Sbjct: 187 -VDAFQLRLKELDSKYK 202
>UniRef100_UPI00002D6890 UPI00002D6890 UniRef100 entry
Length = 427
Score = 35.4 bits (80), Expect = 0.26
Identities = 31/94 (32%), Positives = 51/94 (53%), Gaps = 11/94 (11%)
Query: 54 IELRNINNDYSYCKIHSLHAKLNNLEEHNVPNICKDLALQYIKGGQYARDLDSTKSVIED 113
++ +NINN KI+ + KL+N ++ + K L L+ KG QY D + + D
Sbjct: 188 LDEKNINN-----KIYDIENKLDNSKDQLIDAKNKILDLKN-KGNQYLNDPTQVGTDVAD 241
Query: 114 YFNGVKPSEDGFDVVLIDIDSLFQWNPPHSSNLL 147
Y +G K +EDG + IDI ++ + P +SNL+
Sbjct: 242 YLDG-KVTEDGI-LQNIDIGNI---SMPLNSNLI 270
>UniRef100_Q41173 Acid phosphatase-1 [Lycopersicon esculentum]
Length = 174
Score = 35.0 bits (79), Expect = 0.34
Identities = 20/71 (28%), Positives = 32/71 (44%), Gaps = 1/71 (1%)
Query: 76 NNLEE-HNVPNICKDLALQYIKGGQYARDLDSTKSVIEDYFNGVKPSEDGFDVVLIDIDS 134
NNL +P D +Y+ G Y ++D +Y V +DG DV + D+D
Sbjct: 6 NNLSPGKTIPEEGADYVKEYMVGPGYKMEIDRVSDEAGEYAKSVDLGDDGRDVWIFDVDE 65
Query: 135 LFQWNPPHSSN 145
+ N P+ S+
Sbjct: 66 TWLSNLPYYSD 76
>UniRef100_UPI000027D636 UPI000027D636 UniRef100 entry
Length = 242
Score = 33.9 bits (76), Expect = 0.76
Identities = 22/58 (37%), Positives = 31/58 (52%), Gaps = 5/58 (8%)
Query: 4 SFVLESGFYITSFSATIFIAGFAALGLLLVTLLVSMAMMLQSCQNSNGGIIELRNINN 61
SF L SGF++ F T+F F AL L TLL+S+ ++ S +G + INN
Sbjct: 69 SFFLISGFFV--FFGTVF---FEALNLTTTTLLLSLVIIFYSLFALSGKVFSADKINN 121
>UniRef100_Q7RPQ8 Hypothetical protein [Plasmodium yoelii yoelii]
Length = 702
Score = 33.9 bits (76), Expect = 0.76
Identities = 23/84 (27%), Positives = 37/84 (43%), Gaps = 1/84 (1%)
Query: 43 LQSCQNSNGGIIELRNINNDYSYCKIHSLHAKLNNLEEHNVPNICKDLALQYIKGGQYAR 102
L+ C N + I N YCK H K +N+ +H P + K L+ G QY R
Sbjct: 149 LKLCNNLDMSFIYNTYDKNGLLYCKKIKFHVKGDNINDHITPLVVKINTLKNNMGKQYMR 208
Query: 103 DLDSTKSVIEDYFNGVKPSEDGFD 126
L+ + E +F + E+ ++
Sbjct: 209 -LELSDDKNESFFYYLDLFEENYE 231
>UniRef100_Q975S5 Probable spermidine synthase [Sulfolobus tokodaii]
Length = 300
Score = 33.9 bits (76), Expect = 0.76
Identities = 20/70 (28%), Positives = 38/70 (53%), Gaps = 1/70 (1%)
Query: 72 HAKLNNLEEHNVPNICKDLALQYIKGG-QYARDLDSTKSVIEDYFNGVKPSEDGFDVVLI 130
H + N+ ++ + D A +Y++ Q A D +K VIED + +K +++ FD V+I
Sbjct: 98 HKTIKNVTMVDIDPVVIDFAKKYLQEWHQGAFDNPKSKLVIEDGYKFIKETKEKFDAVVI 157
Query: 131 DIDSLFQWNP 140
D+ + +P
Sbjct: 158 DLTDPIKDSP 167
>UniRef100_Q890B5 Amino acid transport protein [Lactobacillus plantarum]
Length = 454
Score = 33.5 bits (75), Expect = 0.99
Identities = 21/86 (24%), Positives = 39/86 (44%), Gaps = 4/86 (4%)
Query: 1 MGSSFVLESGFYITSFS--ATIFIAGFAALGLLLVTLLVSMAMMLQSCQNSNGGIIELRN 58
+G+ FV+ S + S + F+ FA +G+ +++ ++ S +N GI
Sbjct: 242 VGAIFVIVSIYPWDKLSQIGSPFVQTFAKIGITAAASIINFVVITASLSGTNSGIYSASR 301
Query: 59 INNDYSYCKIHSLHAKLNNLEEHNVP 84
+ Y+ + H L K+ L H VP
Sbjct: 302 MT--YTLAESHQLPRKMTLLNRHGVP 325
>UniRef100_O77391 Hypothetical protein MAL3P6.4 [Plasmodium falciparum]
Length = 1008
Score = 33.5 bits (75), Expect = 0.99
Identities = 32/108 (29%), Positives = 49/108 (44%), Gaps = 27/108 (25%)
Query: 48 NSNGGIIELRN----INND--YSYCKIHSLHAKLNNLEEHN-----VPNICKDLAL---- 92
N GII+ INND Y++ IH+ NN++E+N +PN D L
Sbjct: 164 NKTEGIIDKNKSSNVINNDSEYNHPNIHNNEGTYNNIDENNPMYEGMPNHLSDSNLESAM 223
Query: 93 ----QYIKGGQ----YARDLDSTKSVIEDYFNGVKPSEDGFDVVLIDI 132
+YIKG + R+ + K ++E +K ED F L+D+
Sbjct: 224 KKAEEYIKGKEDDIIERREKEKKKKIVEK----MKDDEDVFTDKLLDL 267
>UniRef100_Q5WDW9 D-alanine--D-alanine ligase A [Bacillus clausii]
Length = 368
Score = 33.1 bits (74), Expect = 1.3
Identities = 19/53 (35%), Positives = 27/53 (50%), Gaps = 6/53 (11%)
Query: 96 KGGQYARDLDSTKSVIEDYFNGVKPSEDGFDVVLIDIDSLFQWNPPHSSNLLL 148
K ++ L S K+++E +D FDV LI ID QW+ +SN LL
Sbjct: 13 KSAEHEVSLQSAKNIVEAI------DKDRFDVTLIGIDKQGQWHINEASNFLL 59
>UniRef100_P93712 Pod storage protein [Phaseolus vulgaris]
Length = 255
Score = 33.1 bits (74), Expect = 1.3
Identities = 21/62 (33%), Positives = 27/62 (42%), Gaps = 2/62 (3%)
Query: 83 VPNICKDLALQYIKGGQYARDLDSTKSVIEDYFNGVKPSEDGFDVVLIDIDSLFQWNPPH 142
+P C D YI+GGQY D + I YF DV+L +ID N P+
Sbjct: 62 IPQQCVDATANYIEGGQYRSDSKTVNQQI--YFFARDRHVHENDVILFNIDGTALSNIPY 119
Query: 143 SS 144
S
Sbjct: 120 YS 121
>UniRef100_Q5VPF2 Putative acid phosphatase [Oryza sativa]
Length = 293
Score = 33.1 bits (74), Expect = 1.3
Identities = 20/78 (25%), Positives = 31/78 (39%), Gaps = 1/78 (1%)
Query: 66 CKIHSLHAKLNNLEE-HNVPNICKDLALQYIKGGQYARDLDSTKSVIEDYFNGVKPSEDG 124
C L + NNL ++P C +Y+ G Y DL+ Y + +DG
Sbjct: 81 CASWRLAGEANNLAPWKSLPEECAAYVREYLTGVAYRSDLEVVAREASAYARTARVGDDG 140
Query: 125 FDVVLIDIDSLFQWNPPH 142
D + D+D N P+
Sbjct: 141 RDAWVFDVDETLLSNLPY 158
>UniRef100_Q940E6 Putative defense associated acid phosphatase [Phaseolus vulgaris]
Length = 264
Score = 33.1 bits (74), Expect = 1.3
Identities = 20/59 (33%), Positives = 27/59 (44%), Gaps = 1/59 (1%)
Query: 76 NNLEE-HNVPNICKDLALQYIKGGQYARDLDSTKSVIEDYFNGVKPSEDGFDVVLIDID 133
NN+ VP C Y+ GQY RDL I Y + + S DG D ++D+D
Sbjct: 61 NNVRRWRTVPPQCYHHLQNYMCAGQYERDLSLAVEHILLYASQIPLSPDGMDAWILDVD 119
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.321 0.138 0.404
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 252,279,995
Number of Sequences: 2790947
Number of extensions: 10071029
Number of successful extensions: 28523
Number of sequences better than 10.0: 62
Number of HSP's better than 10.0 without gapping: 22
Number of HSP's successfully gapped in prelim test: 40
Number of HSP's that attempted gapping in prelim test: 28497
Number of HSP's gapped (non-prelim): 66
length of query: 149
length of database: 848,049,833
effective HSP length: 125
effective length of query: 24
effective length of database: 499,181,458
effective search space: 11980354992
effective search space used: 11980354992
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 67 (30.4 bits)
Medicago: description of AC147429.11


