

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147406.5 - phase: 0 /pseudo
(99 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
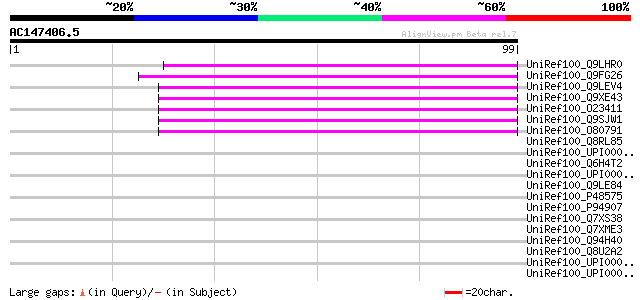
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9LHR0 Non-LTR retrolelement reverse transcriptase-lik... 59 4e-08
UniRef100_Q9FG26 Non-LTR retroelement reverse transcriptase-like... 59 4e-08
UniRef100_Q9LEV4 Hypothetical protein T30N20_120 [Arabidopsis th... 58 6e-08
UniRef100_Q9XE43 Putative non-LTR retrolelement reverse transcri... 57 1e-07
UniRef100_O23411 Hypothetical protein AT4g15600 [Arabidopsis tha... 56 2e-07
UniRef100_Q9SJW1 Hypothetical protein At2g01050 [Arabidopsis tha... 52 3e-06
UniRef100_O80791 Putative Ta11-like non-LTR retroelement protein... 52 3e-06
UniRef100_Q8RL85 2-isopropylmalate synthase [Bacillus stearother... 37 0.087
UniRef100_UPI00002A2BCD UPI00002A2BCD UniRef100 entry 37 0.15
UniRef100_Q6H4T2 Hypothetical protein B1008E06.28 [Oryza sativa] 35 0.33
UniRef100_UPI00002F416A UPI00002F416A UniRef100 entry 35 0.43
UniRef100_Q9LE84 Hypothetical protein AT4g08820 [Arabidopsis tha... 35 0.43
UniRef100_P48575 2-isopropylmalate synthase [Anabaena sp.] 35 0.43
UniRef100_P94907 2-isopropylmalate synthase [Microcystis aerugin... 33 1.3
UniRef100_Q7XS38 OSJNBa0081G05.2 protein [Oryza sativa] 33 1.7
UniRef100_Q7XME3 OSJNBa0061G20.16 protein [Oryza sativa] 33 1.7
UniRef100_Q94H40 Putative reverse transcriptase [Oryza sativa] 33 2.2
UniRef100_Q8U2A2 2-isopropylmalate synthase [Pyrococcus furiosus] 32 2.8
UniRef100_UPI000045353B UPI000045353B UniRef100 entry 32 3.7
UniRef100_UPI00002786D0 UPI00002786D0 UniRef100 entry 32 3.7
>UniRef100_Q9LHR0 Non-LTR retrolelement reverse transcriptase-like protein
[Arabidopsis thaliana]
Length = 591
Score = 58.5 bits (140), Expect = 4e-08
Identities = 24/69 (34%), Positives = 40/69 (57%)
Query: 31 VKLLGKNLGYHAMRERLQKTWRTQGGFEIMDNDNGFYMVKFDEAADKEKVITGGPWLIFD 90
VK+LG+ + A+ RL++ W+ G ++D F+MV+F++ + +TGGPW F
Sbjct: 83 VKVLGRKVSLAAVSHRLREMWKPSGAMYVLDLPRHFFMVRFEKEEEFLTALTGGPWRAFG 142
Query: 91 HCLAVTHWT 99
CL V W+
Sbjct: 143 SCLLVKAWS 151
>UniRef100_Q9FG26 Non-LTR retroelement reverse transcriptase-like [Arabidopsis
thaliana]
Length = 676
Score = 58.5 bits (140), Expect = 4e-08
Identities = 24/74 (32%), Positives = 44/74 (59%)
Query: 26 RMQFVVKLLGKNLGYHAMRERLQKTWRTQGGFEIMDNDNGFYMVKFDEAADKEKVITGGP 85
++ VVK+LG+++ A+ +L++ W+ +GG ++D F++V+FD D +TGGP
Sbjct: 92 KLCMVVKVLGRHVSIAALSRKLRELWKPRGGMYVLDLPRQFFVVRFDVEEDYLAAVTGGP 151
Query: 86 WLIFDHCLAVTHWT 99
W +F L W+
Sbjct: 152 WRLFGSVLMGQAWS 165
>UniRef100_Q9LEV4 Hypothetical protein T30N20_120 [Arabidopsis thaliana]
Length = 181
Score = 57.8 bits (138), Expect = 6e-08
Identities = 22/70 (31%), Positives = 41/70 (58%)
Query: 30 VVKLLGKNLGYHAMRERLQKTWRTQGGFEIMDNDNGFYMVKFDEAADKEKVITGGPWLIF 89
+VK+ G+N+ A+ +L++ W+ +G ++D F+M++F+ + +TGGPW F
Sbjct: 98 IVKVPGRNMTISALSRKLREMWKPRGAMSVVDLPRKFFMIRFEAEEEYMAALTGGPWRAF 157
Query: 90 DHCLAVTHWT 99
D L V W+
Sbjct: 158 DRYLMVQAWS 167
>UniRef100_Q9XE43 Putative non-LTR retrolelement reverse transcriptase [Arabidopsis
thaliana]
Length = 855
Score = 56.6 bits (135), Expect = 1e-07
Identities = 24/70 (34%), Positives = 40/70 (56%)
Query: 30 VVKLLGKNLGYHAMRERLQKTWRTQGGFEIMDNDNGFYMVKFDEAADKEKVITGGPWLIF 89
VVK+LG+N+ + RL++ W+ QG ++D F+M++F+ + +TGGPW F
Sbjct: 57 VVKVLGRNISIANLNRRLREMWKPQGAMFVVDLPCQFFMIRFEREDEYLSALTGGPWRAF 116
Query: 90 DHCLAVTHWT 99
L V W+
Sbjct: 117 GSYLLVQAWS 126
>UniRef100_O23411 Hypothetical protein AT4g15600 [Arabidopsis thaliana]
Length = 655
Score = 56.2 bits (134), Expect = 2e-07
Identities = 23/70 (32%), Positives = 38/70 (53%)
Query: 30 VVKLLGKNLGYHAMRERLQKTWRTQGGFEIMDNDNGFYMVKFDEAADKEKVITGGPWLIF 89
+VK+LG++ + +L+ WR GG ++D F+MV+F+ D +TGGPW +
Sbjct: 97 LVKILGRHTTIEVLSRKLRDLWRPTGGMSVLDLPRQFFMVRFEVEEDYMMALTGGPWRVL 156
Query: 90 DHCLAVTHWT 99
L V W+
Sbjct: 157 GSILMVQAWS 166
>UniRef100_Q9SJW1 Hypothetical protein At2g01050 [Arabidopsis thaliana]
Length = 515
Score = 52.4 bits (124), Expect = 3e-06
Identities = 20/70 (28%), Positives = 38/70 (53%)
Query: 30 VVKLLGKNLGYHAMRERLQKTWRTQGGFEIMDNDNGFYMVKFDEAADKEKVITGGPWLIF 89
+VK+LG + + +L++ W+ G +MD F+M++F+ + +TGGPW +
Sbjct: 82 IVKVLGSQIPISVLNRKLRELWKPSGVMTVMDLPRQFFMIRFELEEEYMAALTGGPWRVL 141
Query: 90 DHCLAVTHWT 99
+ L V W+
Sbjct: 142 GNYLLVQDWS 151
>UniRef100_O80791 Putative Ta11-like non-LTR retroelement protein [Arabidopsis
thaliana]
Length = 409
Score = 52.4 bits (124), Expect = 3e-06
Identities = 22/70 (31%), Positives = 37/70 (52%)
Query: 30 VVKLLGKNLGYHAMRERLQKTWRTQGGFEIMDNDNGFYMVKFDEAADKEKVITGGPWLIF 89
VVK+LG+++ + RL++ W+ GG +++ F+MVKFD D G PW +
Sbjct: 34 VVKVLGRHITIADLNRRLRELWKPNGGMSVLNLPRSFFMVKFDLEEDYLAAAIGDPWRVL 93
Query: 90 DHCLAVTHWT 99
L + W+
Sbjct: 94 GSILMIRAWS 103
>UniRef100_Q8RL85 2-isopropylmalate synthase [Bacillus stearothermophilus]
Length = 515
Score = 37.4 bits (85), Expect = 0.087
Identities = 20/50 (40%), Positives = 30/50 (60%), Gaps = 5/50 (10%)
Query: 33 LLGKNLGYHAMRERLQKTWRTQGGFEIMDNDNGFYMVKFDEAADKEKVIT 82
+LGKN G HA+R R+++ G+ + D + V+F E ADK+K IT
Sbjct: 326 VLGKNSGRHALRYRVEEL-----GYTLSDEEINQLFVRFKELADKKKDIT 370
>UniRef100_UPI00002A2BCD UPI00002A2BCD UniRef100 entry
Length = 287
Score = 36.6 bits (83), Expect = 0.15
Identities = 18/49 (36%), Positives = 30/49 (60%), Gaps = 5/49 (10%)
Query: 33 LLGKNLGYHAMRERLQKTWRTQGGFEIMDNDNGFYMVKFDEAADKEKVI 81
+LGK+ G HA R+RL T+ G+E+ +++ +KF E DK+K +
Sbjct: 106 VLGKHSGRHAFRDRL-----TEFGYELSEDNFEIAFIKFKELCDKKKYV 149
>UniRef100_Q6H4T2 Hypothetical protein B1008E06.28 [Oryza sativa]
Length = 417
Score = 35.4 bits (80), Expect = 0.33
Identities = 13/45 (28%), Positives = 24/45 (52%)
Query: 42 AMRERLQKTWRTQGGFEIMDNDNGFYMVKFDEAADKEKVITGGPW 86
++R +L + W G +G ++++F E AD +V+ G PW
Sbjct: 64 SLRTKLLRDWNPSGAVNFWRATDGAFLIQFYERADLARVLDGAPW 108
>UniRef100_UPI00002F416A UPI00002F416A UniRef100 entry
Length = 431
Score = 35.0 bits (79), Expect = 0.43
Identities = 18/49 (36%), Positives = 29/49 (58%), Gaps = 5/49 (10%)
Query: 33 LLGKNLGYHAMRERLQKTWRTQGGFEIMDNDNGFYMVKFDEAADKEKVI 81
+LGK+ G HA R+RL Q G+E+ +++ +KF E DK+K +
Sbjct: 311 VLGKHSGRHAFRDRL-----NQFGYELSEDNFEIAFLKFKELCDKKKYV 354
>UniRef100_Q9LE84 Hypothetical protein AT4g08820 [Arabidopsis thaliana]
Length = 404
Score = 35.0 bits (79), Expect = 0.43
Identities = 16/53 (30%), Positives = 28/53 (52%), Gaps = 9/53 (16%)
Query: 56 GFEIMDNDNG---------FYMVKFDEAADKEKVITGGPWLIFDHCLAVTHWT 99
G E+++ NG F+M++F+ + +TGGPW +F + L V W+
Sbjct: 46 GEEVLEAMNGLWKRCMIVKFFMIRFELEEEYLAALTGGPWRVFGNYLMVQDWS 98
>UniRef100_P48575 2-isopropylmalate synthase [Anabaena sp.]
Length = 531
Score = 35.0 bits (79), Expect = 0.43
Identities = 20/50 (40%), Positives = 30/50 (60%), Gaps = 5/50 (10%)
Query: 33 LLGKNLGYHAMRERLQKTWRTQGGFEIMDNDNGFYMVKFDEAADKEKVIT 82
+LGK+ G +A R RL++ GFE+ + + VKF E ADK+K I+
Sbjct: 344 VLGKHSGRNAFRTRLKEL-----GFELSETELNKAFVKFKEVADKKKEIS 388
>UniRef100_P94907 2-isopropylmalate synthase [Microcystis aeruginosa]
Length = 533
Score = 33.5 bits (75), Expect = 1.3
Identities = 19/50 (38%), Positives = 29/50 (58%), Gaps = 5/50 (10%)
Query: 33 LLGKNLGYHAMRERLQKTWRTQGGFEIMDNDNGFYMVKFDEAADKEKVIT 82
+LGK G +A R RLQ+ GFE+ + + ++F E AD++K IT
Sbjct: 345 VLGKLSGRNAFRSRLQEL-----GFELSETELNNAFIQFKEMADRKKEIT 389
>UniRef100_Q7XS38 OSJNBa0081G05.2 protein [Oryza sativa]
Length = 1711
Score = 33.1 bits (74), Expect = 1.7
Identities = 13/46 (28%), Positives = 23/46 (49%)
Query: 42 AMRERLQKTWRTQGGFEIMDNDNGFYMVKFDEAADKEKVITGGPWL 87
A+ E +++ WR + + ++V F D + V+ GGPWL
Sbjct: 184 ALFEDMRRAWRLRAEMSYKSLKDNLFIVSFSAEGDHKFVLQGGPWL 229
>UniRef100_Q7XME3 OSJNBa0061G20.16 protein [Oryza sativa]
Length = 1285
Score = 33.1 bits (74), Expect = 1.7
Identities = 13/46 (28%), Positives = 23/46 (49%)
Query: 42 AMRERLQKTWRTQGGFEIMDNDNGFYMVKFDEAADKEKVITGGPWL 87
A+ E +++ WR + + ++V F D + V+ GGPWL
Sbjct: 184 ALFEDMRRAWRLRAEMSYKSLKDNLFIVSFSAEGDHKFVLQGGPWL 229
>UniRef100_Q94H40 Putative reverse transcriptase [Oryza sativa]
Length = 1185
Score = 32.7 bits (73), Expect = 2.2
Identities = 13/52 (25%), Positives = 24/52 (46%)
Query: 36 KNLGYHAMRERLQKTWRTQGGFEIMDNDNGFYMVKFDEAADKEKVITGGPWL 87
K + A+ ++ W + D+ +MV+F D +V+ GGPW+
Sbjct: 74 KPFSHSALLNAMRNAWSSAQDISFKVMDSNLFMVQFQCLGDWNRVMEGGPWI 125
>UniRef100_Q8U2A2 2-isopropylmalate synthase [Pyrococcus furiosus]
Length = 499
Score = 32.3 bits (72), Expect = 2.8
Identities = 16/50 (32%), Positives = 29/50 (58%), Gaps = 5/50 (10%)
Query: 33 LLGKNLGYHAMRERLQKTWRTQGGFEIMDNDNGFYMVKFDEAADKEKVIT 82
+LGK+ G HA R++L++ G+ + + + KF E AD++K +T
Sbjct: 327 VLGKHSGRHAFRKKLEEM-----GYRLSEEEINRLFAKFKEIADRKKGLT 371
>UniRef100_UPI000045353B UPI000045353B UniRef100 entry
Length = 1507
Score = 32.0 bits (71), Expect = 3.7
Identities = 20/70 (28%), Positives = 33/70 (46%), Gaps = 10/70 (14%)
Query: 5 IVFYPRFISNNQSF--NNYVRHGRMQFVVKLLGKNLGYHAMRERLQKTWRTQGGFEIMDN 62
+ F RF S+ ++ NN+ G +F+++ +G +GYH L WR E DN
Sbjct: 1034 VKFSTRFESSTDTYSVNNF---GSKEFIIQFVGNAIGYH-----LDSQWRVLLALENKDN 1085
Query: 63 DNGFYMVKFD 72
N + + D
Sbjct: 1086 FNQVLVSQLD 1095
>UniRef100_UPI00002786D0 UPI00002786D0 UniRef100 entry
Length = 315
Score = 32.0 bits (71), Expect = 3.7
Identities = 17/49 (34%), Positives = 26/49 (52%), Gaps = 5/49 (10%)
Query: 33 LLGKNLGYHAMRERLQKTWRTQGGFEIMDNDNGFYMVKFDEAADKEKVI 81
+LGK+ G HA + +L + G+E +N KF E ADK+K +
Sbjct: 117 VLGKHSGRHAFKVKLNEL-----GYEFSENKINLLFKKFKELADKKKQV 160
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.327 0.141 0.460
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 183,395,297
Number of Sequences: 2790947
Number of extensions: 6921639
Number of successful extensions: 16795
Number of sequences better than 10.0: 38
Number of HSP's better than 10.0 without gapping: 20
Number of HSP's successfully gapped in prelim test: 18
Number of HSP's that attempted gapping in prelim test: 16769
Number of HSP's gapped (non-prelim): 38
length of query: 99
length of database: 848,049,833
effective HSP length: 75
effective length of query: 24
effective length of database: 638,728,808
effective search space: 15329491392
effective search space used: 15329491392
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 68 (30.8 bits)
Medicago: description of AC147406.5


