

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147404.4 - phase: 0
(583 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
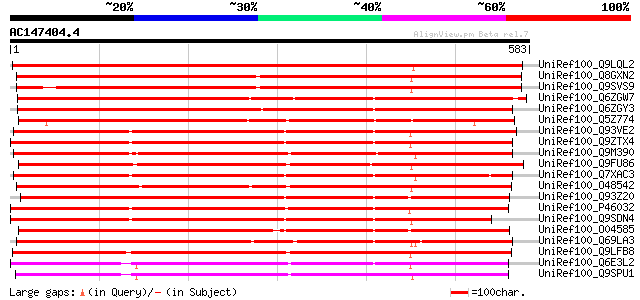
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9LQL2 F5D14.23 protein [Arabidopsis thaliana] 880 0.0
UniRef100_Q8GXN2 Putative peptide transporter [Arabidopsis thali... 780 0.0
UniRef100_Q9SVS9 Peptide transporter-like protein [Arabidopsis t... 749 0.0
UniRef100_Q6ZGW7 Peptide transporter-like [Oryza sativa] 712 0.0
UniRef100_Q6ZGY3 Putative peptide transporter [Oryza sativa] 680 0.0
UniRef100_Q5Z774 Putative peptide transporter [Oryza sativa] 539 e-151
UniRef100_Q93VE2 Putative peptide transporter 1 [Oryza sativa] 475 e-132
UniRef100_Q9ZTX4 LeOPT1 [Lycopersicon esculentum] 473 e-132
UniRef100_Q9M390 Putative peptide transport protein [Arabidopsis... 468 e-130
UniRef100_Q9FU86 Putative peptide transport protein [Oryza sativa] 464 e-129
UniRef100_Q7XAC3 Peptide transporter 1 [Vicia faba] 463 e-129
UniRef100_O48542 Peptide transporter [Hordeum vulgare] 461 e-128
UniRef100_Q93Z20 At1g62200/F19K23_13 [Arabidopsis thaliana] 460 e-128
UniRef100_P46032 Peptide transporter PTR2-B [Arabidopsis thaliana] 459 e-127
UniRef100_Q9SDN4 Amino acid/peptide transporter [Prunus dulcis] 441 e-122
UniRef100_O04585 F19K23.13 protein [Arabidopsis thaliana] 441 e-122
UniRef100_Q69LA3 Putative peptide transport protein [Oryza sativa] 438 e-121
UniRef100_Q9LFB8 Oligopeptide transporter-like protein [Arabidop... 421 e-116
UniRef100_Q6E3L2 Low affinity nitrate transporter NRT1.1 [Tritic... 404 e-111
UniRef100_Q9SPU1 Nitrate transporter [Oryza sativa] 402 e-110
>UniRef100_Q9LQL2 F5D14.23 protein [Arabidopsis thaliana]
Length = 606
Score = 880 bits (2273), Expect = 0.0
Identities = 433/581 (74%), Positives = 500/581 (85%), Gaps = 8/581 (1%)
Query: 4 KEETEELTLDGSVDWHGRPSIRATSGRWFAGTIILLNQGLATLAFFGVGVNLVLFLTRVL 63
K+E EE T DG+VD++GRPSIR+ SG+W AG +ILLNQGLATLAFFGVGVNLVLFLTRVL
Sbjct: 6 KKEGEEETRDGTVDYYGRPSIRSNSGQWVAGIVILLNQGLATLAFFGVGVNLVLFLTRVL 65
Query: 64 GQDNAAAANNVSKWTGTVYIFSLVGAFLSDSYWGRYKTCAIFQGIFVTGLVSLSVTTYLA 123
Q+NA AANNVSKWTGTVYIFSLVGAFLSDSYWGRYKTCAIFQ IFV GL SLS+++Y+
Sbjct: 66 QQNNADAANNVSKWTGTVYIFSLVGAFLSDSYWGRYKTCAIFQVIFVIGLSSLSLSSYMF 125
Query: 124 LLRPKGCGNGKLECGEHSSLEMGMFYLSIYLIALGNGGYQPNIATFGADQFDEDHSKESY 183
L+RP+GCG+ CG HS +E+ MFY SIYLIALG GGYQPNIAT GADQFDE+H KE Y
Sbjct: 126 LIRPRGCGDEVTPCGSHSMMEITMFYFSIYLIALGYGGYQPNIATLGADQFDEEHPKEGY 185
Query: 184 SKVAFFSYFYLALNLGSLFSNTILGYFEDEGLWALGFWASAGSAFLALVLFLVGTPKYRH 243
SK+AFFSYFYLALNLGSLFSNTILGYFEDEG+WALGFWAS GSA + L+LFLVGTP+YR+
Sbjct: 186 SKIAFFSYFYLALNLGSLFSNTILGYFEDEGMWALGFWASTGSAIIGLILFLVGTPRYRY 245
Query: 244 FKPCGNPLSRFCQVFFAASRKLGVQMTSNG-DDLYVIDE--KESSNNSNRKILHTHGFKF 300
FKP GNPLSRFCQV AA++K V+ G +++Y D K +S N+ R+I+HT FKF
Sbjct: 246 FKPTGNPLSRFCQVLVAATKKSSVEAPLRGREEMYDGDSEGKNASVNTGRRIVHTDEFKF 305
Query: 301 LDRAAYITSRDLEVQKGGQHNPWYLCPITQVEEVKCILRLLPIWLCTIIYSVVFTQMASL 360
LD+AAYIT+RDL+ +K NPW LCP+TQVEEVKCILRL+PIWLCTIIYSVVFTQMASL
Sbjct: 306 LDKAAYITARDLDDKKQDSVNPWRLCPVTQVEEVKCILRLMPIWLCTIIYSVVFTQMASL 365
Query: 361 FVEQGAAMKTTISSFKIPPASMSSFDILSVAIFIFFYRRVLDPLVGKLKKSSSKGLTELQ 420
FVEQGAAM T++S FKIPPASMSSFDILSVA+FIF YRRVL+P+ + KK+ SKG+TEL
Sbjct: 366 FVEQGAAMNTSVSDFKIPPASMSSFDILSVALFIFLYRRVLEPVANRFKKNGSKGITELH 425
Query: 421 RMGIGLVIAIIAMVTAGIVECYRLKYAKQGDT-----SSLSIFWQIPQYALIGASEVFMY 475
RMGIGLVIA+IAM+ AGIVECYRLKYA + T SSLSIFWQ PQY+LIGASEVFMY
Sbjct: 426 RMGIGLVIAVIAMIAAGIVECYRLKYADKSCTHCDGSSSLSIFWQAPQYSLIGASEVFMY 485
Query: 476 VGQLEFFNAQTPDGLKSFGSALCMTSISLGNYVSSLIVSIVMKISTEDHMPGWIPGNLNR 535
VGQLEFFNAQTPDGLKSFGSALCM S+S+GN+VSSL+V++V+KISTEDHMPGWIP NLN+
Sbjct: 486 VGQLEFFNAQTPDGLKSFGSALCMMSMSMGNFVSSLLVTMVVKISTEDHMPGWIPRNLNK 545
Query: 536 GHLDRFFFLLAVLTSLDLIAYIACAKWFQNIQMACKYDNND 576
GHLDRF+FLLA LTS+DL+ YIACAKW++ IQ+ K + D
Sbjct: 546 GHLDRFYFLLAALTSIDLVVYIACAKWYKPIQLEGKDEMQD 586
>UniRef100_Q8GXN2 Putative peptide transporter [Arabidopsis thaliana]
Length = 589
Score = 780 bits (2015), Expect = 0.0
Identities = 392/577 (67%), Positives = 460/577 (78%), Gaps = 12/577 (2%)
Query: 8 EELTLDGSVDWHGRPSIRATSGRWFAGTIILLNQGLATLAFFGVGVNLVLFLTRVLGQDN 67
E T DGSVD HG P+IRA +G+W +IL+NQGLATLAFFGVGVNLVLFLTRV+GQDN
Sbjct: 9 EVCTQDGSVDRHGNPAIRANTGKWLTAILILVNQGLATLAFFGVGVNLVLFLTRVMGQDN 68
Query: 68 AAAANNVSKWTGTVYIFSLVGAFLSDSYWGRYKTCAIFQGIFVTGLVSLSVTTYLALLRP 127
A AANNVSKWTGTVYIFSL+GAFLSDSYWGRYKTCAIFQ FV GL+ LS++T LL P
Sbjct: 69 AEAANNVSKWTGTVYIFSLLGAFLSDSYWGRYKTCAIFQASFVAGLMMLSLSTGALLLEP 128
Query: 128 KGCGNGKLECGEHSSLEMGMFYLSIYLIALGNGGYQPNIATFGADQFDEDHSKESYSKVA 187
GCG C HS+ + +FYLS+YLIALG GGYQPNIATFGADQFD + S E +SK+A
Sbjct: 129 SGCGVEDSPCKPHSTFKTVLFYLSVYLIALGYGGYQPNIATFGADQFDAEDSVEGHSKIA 188
Query: 188 FFSYFYLALNLGSLFSNTILGYFEDEGLWALGFWASAGSAFLALVLFLVGTPKYRHFKPC 247
FFSYFYLALNLGSLFSNT+LGYFED+G W LGFWASAGSAF LVLFL+GTPKYRHF P
Sbjct: 189 FFSYFYLALNLGSLFSNTVLGYFEDQGEWPLGFWASAGSAFAGLVLFLIGTPKYRHFTPR 248
Query: 248 GNPLSRFCQVFFAASRKLGVQMTSNGDDLYVIDEKESSNNSNRKILHTHGFKFLDRAAYI 307
+P SRFCQV AA+RK + + +LY + E+ ++KILHT GF+FLDRAA +
Sbjct: 249 ESPWSRFCQVLVAATRKAKIDVHHEELNLY---DSETQYTGDKKILHTKGFRFLDRAAIV 305
Query: 308 TSRD--LEVQKGGQHNPWYLCPITQVEEVKCILRLLPIWLCTIIYSVVFTQMASLFVEQG 365
T D +V+ G +++PW LC +TQVEEVKC+LRLLPIWLCTI+YSVVFTQMASLFV QG
Sbjct: 306 TPDDEAEKVESGSKYDPWRLCSVTQVEEVKCVLRLLPIWLCTILYSVVFTQMASLFVVQG 365
Query: 366 AAMKTTISSFKIPPASMSSFDILSVAIFIFFYRRVLDPLVGKLKKSS-SKGLTELQRMGI 424
AAMKT I +F+IP +SMSSFDIL+VA FIF YRR LDPL +L K+ +KGLTELQRMGI
Sbjct: 366 AAMKTNIKNFRIPASSMSSFDILNVAFFIFAYRRFLDPLFARLNKTERNKGLTELQRMGI 425
Query: 425 GLVIAIIAMVTAGIVECYRLKYAKQ------GDTSSLSIFWQIPQYALIGASEVFMYVGQ 478
GLVIAI+AM++AGIVE +RLK + +S+LSIFWQ+PQY LIGASEVFMYVGQ
Sbjct: 426 GLVIAIMAMISAGIVEIHRLKNKEPESATSISSSSTLSIFWQVPQYMLIGASEVFMYVGQ 485
Query: 479 LEFFNAQTPDGLKSFGSALCMTSISLGNYVSSLIVSIVMKISTEDHMPGWIPGNLNRGHL 538
LEFFN+Q P GLKSF SALCM SISLGNYVSSL+VSIVMKIST D + GWIP NLN+GHL
Sbjct: 486 LEFFNSQAPTGLKSFASALCMASISLGNYVSSLLVSIVMKISTTDDVHGWIPENLNKGHL 545
Query: 539 DRFFFLLAVLTSLDLIAYIACAKWFQNIQMACKYDNN 575
+RF+FLLA LT+ D + Y+ CAKW++ I+ + +
Sbjct: 546 ERFYFLLAGLTAADFVVYLICAKWYKYIKSEASFSES 582
>UniRef100_Q9SVS9 Peptide transporter-like protein [Arabidopsis thaliana]
Length = 576
Score = 749 bits (1933), Expect = 0.0
Identities = 381/577 (66%), Positives = 447/577 (77%), Gaps = 25/577 (4%)
Query: 8 EELTLDGSVDWHGRPSIRATSGRWFAGTIILLNQGLATLAFFGVGVNLVLFLTRVLGQDN 67
E T DGSVD HG P+IRA +G+W +IL GVNLVLFLTRV+GQDN
Sbjct: 9 EVCTQDGSVDRHGNPAIRANTGKWLTAILIL-------------GVNLVLFLTRVMGQDN 55
Query: 68 AAAANNVSKWTGTVYIFSLVGAFLSDSYWGRYKTCAIFQGIFVTGLVSLSVTTYLALLRP 127
A AANNVSKWTGTVYIFSL+GAFLSDSYWGRYKTCAIFQ FV GL+ LS++T LL P
Sbjct: 56 AEAANNVSKWTGTVYIFSLLGAFLSDSYWGRYKTCAIFQASFVAGLMMLSLSTGALLLEP 115
Query: 128 KGCGNGKLECGEHSSLEMGMFYLSIYLIALGNGGYQPNIATFGADQFDEDHSKESYSKVA 187
GCG C HS+ + +FYLS+YLIALG GGYQPNIATFGADQFD + S E +SK+A
Sbjct: 116 SGCGVEDSPCKPHSTFKTVLFYLSVYLIALGYGGYQPNIATFGADQFDAEDSVEGHSKIA 175
Query: 188 FFSYFYLALNLGSLFSNTILGYFEDEGLWALGFWASAGSAFLALVLFLVGTPKYRHFKPC 247
FFSYFYLALNLGSLFSNT+LGYFED+G W LGFWASAGSAF LVLFL+GTPKYRHF P
Sbjct: 176 FFSYFYLALNLGSLFSNTVLGYFEDQGEWPLGFWASAGSAFAGLVLFLIGTPKYRHFTPR 235
Query: 248 GNPLSRFCQVFFAASRKLGVQMTSNGDDLYVIDEKESSNNSNRKILHTHGFKFLDRAAYI 307
+P SRFCQV AA+RK + + +LY + E+ ++KILHT GF+FLDRAA +
Sbjct: 236 ESPWSRFCQVLVAATRKAKIDVHHEELNLY---DSETQYTGDKKILHTKGFRFLDRAAIV 292
Query: 308 TSRD--LEVQKGGQHNPWYLCPITQVEEVKCILRLLPIWLCTIIYSVVFTQMASLFVEQG 365
T D +V+ G +++PW LC +TQVEEVKC+LRLLPIWLCTI+YSVVFTQMASLFV QG
Sbjct: 293 TPDDEAEKVESGSKYDPWRLCSVTQVEEVKCVLRLLPIWLCTILYSVVFTQMASLFVVQG 352
Query: 366 AAMKTTISSFKIPPASMSSFDILSVAIFIFFYRRVLDPLVGKLKKSS-SKGLTELQRMGI 424
AAMKT I +F+IP +SMSSFDILSVA FIF YRR LDPL +L K+ +KGLTELQRMGI
Sbjct: 353 AAMKTNIKNFRIPASSMSSFDILSVAFFIFAYRRFLDPLFARLNKTERNKGLTELQRMGI 412
Query: 425 GLVIAIIAMVTAGIVECYRLKYAKQ------GDTSSLSIFWQIPQYALIGASEVFMYVGQ 478
GLVIAI+AM++AGIVE +RLK + +S+LSIFWQ+PQY LIGASEVFMYVGQ
Sbjct: 413 GLVIAIMAMISAGIVEIHRLKNKEPESATSISSSSTLSIFWQVPQYMLIGASEVFMYVGQ 472
Query: 479 LEFFNAQTPDGLKSFGSALCMTSISLGNYVSSLIVSIVMKISTEDHMPGWIPGNLNRGHL 538
LEFFN+Q P GLKSF SALCM SISLGNYVSSL+VSIVMKIST D + GWIP NLN+GHL
Sbjct: 473 LEFFNSQAPTGLKSFASALCMASISLGNYVSSLLVSIVMKISTTDDVHGWIPENLNKGHL 532
Query: 539 DRFFFLLAVLTSLDLIAYIACAKWFQNIQMACKYDNN 575
+RF+FLLA LT+ D + Y+ CAKW++ I+ + +
Sbjct: 533 ERFYFLLAGLTAADFVVYLICAKWYKYIKSEASFSES 569
>UniRef100_Q6ZGW7 Peptide transporter-like [Oryza sativa]
Length = 584
Score = 712 bits (1839), Expect = 0.0
Identities = 356/573 (62%), Positives = 437/573 (76%), Gaps = 8/573 (1%)
Query: 9 ELTLDGSVDWHGRPSIRATSGRWFAGTIILLNQGLATLAFFGVGVNLVLFLTRVLGQDNA 68
++T DGS+D G P+++A +G W + ++L+N GL T AFFGVGVNLV+FL RVL QDNA
Sbjct: 13 DITEDGSMDRRGNPAVKAKTGNWRSSILLLVNYGLVTCAFFGVGVNLVVFLRRVLHQDNA 72
Query: 69 AAANNVSKWTGTVYIFSLVGAFLSDSYWGRYKTCAIFQGIFVTGLVSLSVTTYLALLRPK 128
AAN++SKWTGTVYIFSL+GAF+SDSYWGRY TCAIFQ I+VTGLV LS+ ++ L++P
Sbjct: 73 EAANSISKWTGTVYIFSLIGAFMSDSYWGRYITCAIFQMIYVTGLVILSLASWFLLVKPT 132
Query: 129 GCGNGKLECGEHSSLEMGMFYLSIYLIALGNGGYQPNIATFGADQFDEDHSKESYSKVAF 188
GCG C SS + +FYLS Y+IA GNGGYQP+IATFG+DQFDE +E+ SKVAF
Sbjct: 133 GCGAAGEHCDAPSSAGVALFYLSTYMIAFGNGGYQPSIATFGSDQFDETDPREARSKVAF 192
Query: 189 FSYFYLALNLGSLFSNTILGYFEDEGLWALGFWASAGSAFLALVLFLVGTPKYRHFKPCG 248
FSYFYLALN+GSLFSNT+L Y+EDEG W +GFW SA +A +ALVLFL+GTP YRHFKP G
Sbjct: 193 FSYFYLALNVGSLFSNTVLVYYEDEGRWVMGFWVSAAAAAMALVLFLLGTPNYRHFKPTG 252
Query: 249 NPLSRFCQVFFAASRKLGVQMTSNGDDLYVIDEKESSNNSNRKILHTHGFKFLDRAAYIT 308
NPL+R QVF AA RK ++ + L+ +D ES RKILH+ +FLD+AA +T
Sbjct: 253 NPLTRIAQVFVAAFRKWRAEV-PRSELLHEVDGDESQIAGIRKILHSDQIRFLDKAATVT 311
Query: 309 SRDLEVQKGGQHNPWYLCPITQVEEVKCILRLLPIWLCTIIYSVVFTQMASLFVEQGAAM 368
D + Q +PW LC +TQVEEVKCIL++LPIWLCTI+YSVVFTQMASLFVEQG M
Sbjct: 312 EEDYCTPENMQ-DPWRLCTVTQVEEVKCILKMLPIWLCTIVYSVVFTQMASLFVEQGTTM 370
Query: 369 KTTISSFKIPPASMSSFDILSVAIFIFFYRRVLDPLVGKLKKSSSKGLTELQRMGIGLVI 428
T I SF +P ASMS FDILSV FI YRRVL P++ +L + +GLTELQRMG+GLV+
Sbjct: 371 NTNIGSFHVPAASMSVFDILSVLAFIAIYRRVLVPVMSRL-SGNPQGLTELQRMGVGLVV 429
Query: 429 AIIAMVTAGIVECYRLKYAKQGD-TSSLSIFWQIPQYALIGASEVFMYVGQLEFFNAQTP 487
+ AMV AG+VE RLK D SSLS+ WQ+PQYALIGASEVFMYVGQLEFFN Q P
Sbjct: 430 GMAAMVVAGVVEVERLKRVGAPDQPSSLSVLWQVPQYALIGASEVFMYVGQLEFFNGQAP 489
Query: 488 DGLKSFGSALCMTSISLGNYVSSLIVSIVMKISTEDHMPGWIPGNLNRGHLDRFFFLLAV 547
DG+KSFGS+LCM SISLGNYVS ++VS+V ++ D PGWIPGNLN GHLDRF+FLLA
Sbjct: 490 DGVKSFGSSLCMASISLGNYVSIMLVSVVTSLTAGDRRPGWIPGNLNSGHLDRFYFLLAA 549
Query: 548 LTSLDLIAYIACAKWFQNIQMACKYDNNDEPSS 580
L+ +DL Y+ACA W++ I K D+N+E ++
Sbjct: 550 LSLVDLAVYVACAVWYKGI----KLDSNEEKAN 578
>UniRef100_Q6ZGY3 Putative peptide transporter [Oryza sativa]
Length = 610
Score = 680 bits (1754), Expect = 0.0
Identities = 348/559 (62%), Positives = 410/559 (73%), Gaps = 5/559 (0%)
Query: 9 ELTLDGSVDWHGRPSIRATSGRWFAGTIILLNQGLATLAFFGVGVNLVLFLTRVLGQDNA 68
E TLDGSVD G P+++ SG W AG +ILLNQGLATLAFFGV VNLVLFLTRVL Q N
Sbjct: 30 EYTLDGSVDIKGSPAVKGKSGGWLAGGLILLNQGLATLAFFGVNVNLVLFLTRVLQQSNG 89
Query: 69 AAANNVSKWTGTVYIFSLVGAFLSDSYWGRYKTCAIFQGIFVTGLVSLSVTTYLALLRPK 128
AANNVSKWTGTVY+FSL+GAFLSDSYWGRYKTCAIFQ IFV GL LS+++ L L+RP
Sbjct: 90 DAANNVSKWTGTVYMFSLIGAFLSDSYWGRYKTCAIFQAIFVLGLALLSLSSRLYLIRPV 149
Query: 129 GCGNGKLECGEHSSLEMGMFYLSIYLIALGNGGYQPNIATFGADQFDEDHSKESYSKVAF 188
GCG + C HS E+G+FY+++Y+IA GNGGYQPN+ATFGADQFD + ES+SKV+F
Sbjct: 150 GCGTEHVPCEPHSGAELGIFYIALYMIAFGNGGYQPNVATFGADQFDGEDPAESHSKVSF 209
Query: 189 FSYFYLALNLGSLFSNTILGYFEDEGLWALGFWASAGSAFLALVLFLVGTPKYRHFKPCG 248
FSYFYLALNLGSLFSNT L + EDEG WALGFW S +A AL+LFL GT +YR+ +P G
Sbjct: 210 FSYFYLALNLGSLFSNTFLSFLEDEGNWALGFWVSTAAAATALLLFLGGTLRYRYIRPSG 269
Query: 249 NPLSRFCQVFFAASRKLGVQMTSNGDDLYVIDEKESSNNSNRKILHTHGFKFLDRAAYIT 308
NP+ R QV FAA R + LY DEK +++ RK+LHT GF+FLDRAA +
Sbjct: 270 NPVGRIFQVAFAACRNWKAGESPGAVTLYESDEK--ADSGGRKLLHTEGFRFLDRAAVVG 327
Query: 309 SRDLEVQKGGQHNPWYLCPITQVEEVKCILRLLPIWLCTIIYSVVFTQMASLFVEQGAAM 368
+ +PW LC +TQVEEVK ILRLLPIWLCTI+YSVVFTQMASLFV QGAAM
Sbjct: 328 ANPKLGTCTQPRDPWKLCTVTQVEEVKSILRLLPIWLCTILYSVVFTQMASLFVVQGAAM 387
Query: 369 K--TTISSFKIPPASMSSFDILSVAIFIFFYRRVLDPLVGKLKKSSSKGLTELQRMGIGL 426
+ T F +PP+SMS+FDIL+VA IF YRR + PLV +L G TELQRMG+GL
Sbjct: 388 RRTTRFPGFSVPPSSMSAFDILTVATTIFLYRRAVCPLVSRL-TGRHTGPTELQRMGLGL 446
Query: 427 VIAIIAMVTAGIVECYRLKYAKQGDTSSLSIFWQIPQYALIGASEVFMYVGQLEFFNAQT 486
V+ +AM TAG VE +R A +S L I WQ+PQYALIG SEV MYVGQLEFFN +
Sbjct: 447 VLGAMAMATAGTVEHFRKAGATTAMSSDLHIMWQVPQYALIGVSEVMMYVGQLEFFNGEM 506
Query: 487 PDGLKSFGSALCMTSISLGNYVSSLIVSIVMKISTEDHMPGWIPGNLNRGHLDRFFFLLA 546
PD LKSFGSALCM S+SLGNY S +IVS V K + PGWIP +LN GHLD+FFFLLA
Sbjct: 507 PDALKSFGSALCMMSMSLGNYFSDVIVSAVTKATAVRGRPGWIPADLNEGHLDKFFFLLA 566
Query: 547 VLTSLDLIAYIACAKWFQN 565
VL D Y+ CA +++
Sbjct: 567 VLAVADFAVYLVCASRYRS 585
>UniRef100_Q5Z774 Putative peptide transporter [Oryza sativa]
Length = 606
Score = 539 bits (1388), Expect = e-151
Identities = 286/569 (50%), Positives = 380/569 (66%), Gaps = 17/569 (2%)
Query: 11 TLDGSVDWHGRPSIRATSGRWFAGTIILL-----NQGLATLAFFGVGVNLVLFLTRVLGQ 65
T D + + G P ++ + G+ ++ L + LA AFFGV V LV+FL +VL Q
Sbjct: 25 TEDQTQHFQGSPELKTSRGKMTMALLLALLDDAVSYVLANFAFFGVAVGLVVFLRQVLHQ 84
Query: 66 DNAAAANNVSKWTGTVYIFSLVGAFLSDSYWGRYKTCAIFQGIFVTGLVSLSVTTYLALL 125
+NA AAN+VS W GTVYIFSL AFLSDSY GRY TC +FQ IF+ GL+ LS+ ++ L+
Sbjct: 85 ENAEAANSVSMWMGTVYIFSLFCAFLSDSYMGRYITCIMFQFIFIVGLMLLSLLSWFLLV 144
Query: 126 RPKGCGNGK--LECGEHSSLEMGMFYLSIYLIALGNGGYQPNIATFGADQFDEDHSKESY 183
P GCG+G +C S + +FYLSIY+ A GNGGYQP++ATFGADQFD+ E
Sbjct: 145 EPPGCGDGGGLRQCAAPSRRGVAVFYLSIYMAAFGNGGYQPSVATFGADQFDDADPGERR 204
Query: 184 SKVAFFSYFYLALNLGSLFSNTILGYFEDEGLWALGFWASAGSAFLALVLFLVGTPKYRH 243
K AFF FYL+LN+GSLF N++L +FED G W GFW S +A LAL LFL+GTP+YR
Sbjct: 205 RKQAFFCLFYLSLNVGSLFYNSVLVFFEDRGRWVAGFWVSTAAAALALALFLLGTPRYRR 264
Query: 244 FKPCGNPLSRFCQVFFAASRKLGVQMTSNGDDLYVIDEKESSNNSNRKILHTHGFKFLDR 303
+P GNPL+R QVF AA RK + + GD L+ +D + S+ K+ H+ +FLD+
Sbjct: 265 VRPAGNPLTRIAQVFVAAYRKRHI-VPPPGDHLHEVDGEGSAIRGVGKLAHSDQLRFLDK 323
Query: 304 AAYITSRDLEVQKGGQHNPWYLCPITQVEEVKCILRLLPIWLCTIIYSVVFTQMASLFVE 363
AA T D G NPW LC +TQVEE KC++ ++PIW+C+I+YSV FTQM+SLFVE
Sbjct: 324 AATATEED--YHDGNAKNPWRLCTVTQVEEAKCVVSMVPIWICSIVYSVEFTQMSSLFVE 381
Query: 364 QGAAMKTTI-SSFKIPPASMSSFDILSVAIFIFFYRRVLDPLVGKLKKSSSKGLTELQRM 422
QGAAM T I F P ASMS FD+ V + F VL P +L K + +G+ EL+RM
Sbjct: 382 QGAAMDTDILGLFNAPAASMSVFDVAGVLATLAFSHYVLVPAAARLTK-NPRGVGELKRM 440
Query: 423 GIGLVIAIIAMVTAGIVECYRLKYAKQGDTSSLSIFWQIPQYALIGASEVFMYVGQLEFF 482
G GLVIA++ MV A +VE +R + + G ++S+ WQ PQYA++GASEVF+YVGQLEFF
Sbjct: 441 GAGLVIALLGMVAAAVVEVHRRRRSGAGG-RAMSVLWQAPQYAVMGASEVFVYVGQLEFF 499
Query: 483 NAQTPDGLKSFGSALCMTSISLGNYVSSLIVSIVMKIS----TEDHMPGWIPGNLNRGHL 538
N Q+P+G+KS GS+LCM SISLGNY S ++VS + ++ T GWI L+RGHL
Sbjct: 500 NVQSPEGVKSLGSSLCMASISLGNYASMVMVSAISGVASRRRTGGGTAGWILAELDRGHL 559
Query: 539 DRFFFLLAVLTSLDLIAYIACAKWFQNIQ 567
DR F LAVL+++DL+ +I A+ F+ I+
Sbjct: 560 DRSFITLAVLSAVDLVVFIVFARLFKGIE 588
>UniRef100_Q93VE2 Putative peptide transporter 1 [Oryza sativa]
Length = 593
Score = 475 bits (1222), Expect = e-132
Identities = 245/573 (42%), Positives = 367/573 (63%), Gaps = 13/573 (2%)
Query: 5 EETEELTL--DGSVDWHGRPSIRATSGRWFAGTIILLNQGLATLAFFGVGVNLVLFLTRV 62
EE+ +LT DGSVD+ G P ++ +GRW A IL N+ LA++G+ NLV +LT+
Sbjct: 26 EESNQLTYTGDGSVDFSGNPVVKERTGRWRACPFILGNECCERLAYYGISTNLVTYLTKK 85
Query: 63 LGQDNAAAANNVSKWTGTVYIFSLVGAFLSDSYWGRYKTCAIFQGIFVTGLVSLSVTTYL 122
L NA+AA+NV+ W GT Y+ L+GA L+D+YWGRY T A F I+ G+ L+++ +
Sbjct: 86 LHDGNASAASNVTAWQGTCYLTPLIGAILADAYWGRYWTIATFSTIYFIGMAVLTLSASV 145
Query: 123 ALLRPKGCGNGKLECGEHSSLEMGMFYLSIYLIALGNGGYQPNIATFGADQFDEDHSKES 182
P C C + L+ +F+L +YLIALG GG +P +++FGADQFD+ E
Sbjct: 146 PTFMPPPCEGSF--CPPANPLQYTVFFLGLYLIALGTGGIKPCVSSFGADQFDDTDPVER 203
Query: 183 YSKVAFFSYFYLALNLGSLFSNTILGYFEDEGLWALGFWASAGSAFLALVLFLVGTPKYR 242
K +FF++FY ++N+G+L S++ L + +D W +GF LA++ F GT YR
Sbjct: 204 IQKGSFFNWFYFSINIGALISSSFLVWVQDNIGWGIGFGIPTIFMGLAIISFFSGTSLYR 263
Query: 243 HFKPCGNPLSRFCQVFFAASRKLGVQMTSNGDDLYVIDEKESSNNSNRKILHTHGFKFLD 302
KP G+P++R CQV A+ RK V + + LY + + S+ +R++ HT + LD
Sbjct: 264 FQKPGGSPITRVCQVVVASFRKWNVHVPEDSSRLYELPDGASAIEGSRQLEHTDELRCLD 323
Query: 303 RAAYITSRDLEVQKGGQHNPWYLCPITQVEEVKCILRLLPIWLCTIIYSVVFTQMASLFV 362
+AA IT DL+V+ NPW +C +TQVEE+K ++R+ P+W TI++S V+ QM+++FV
Sbjct: 324 KAATIT--DLDVKADSFTNPWRICTVTQVEELKILVRMFPVWATTIVFSAVYAQMSTMFV 381
Query: 363 EQGAAMKTTISSFKIPPASMSSFDILSVAIFIFFYRRVLDPLVGKLKKSSSKGLTELQRM 422
EQG + T++ FKIPPAS+S+FD++SV I++ Y +L P+ + + +G TELQRM
Sbjct: 382 EQGMMLDTSVGPFKIPPASLSTFDVVSVIIWVPLYDSILVPIARRF-TGNPRGFTELQRM 440
Query: 423 GIGLVIAIIAMVTAGIVECYRLKYAK------QGDTSSLSIFWQIPQYALIGASEVFMYV 476
GIGLVI+I +M A ++E RL A+ Q L+I WQIPQY L+GASEVF +V
Sbjct: 441 GIGLVISIFSMAAAAVLEIKRLDIARAEHLVDQNVPVPLNICWQIPQYFLVGASEVFTFV 500
Query: 477 GQLEFFNAQTPDGLKSFGSALCMTSISLGNYVSSLIVSIVMKISTEDHMPGWIPGNLNRG 536
G LEFF Q+PD ++S SAL + + +LGNY+S+ I+++V +T PGWIP NLN+G
Sbjct: 501 GSLEFFYDQSPDAMRSLCSALQLVTTALGNYLSAFILTLVAYFTTRGGNPGWIPDNLNQG 560
Query: 537 HLDRFFFLLAVLTSLDLIAYIACAKWFQNIQMA 569
HLD FF+LLA L+ L+ + Y+ CA +++ + A
Sbjct: 561 HLDYFFWLLAGLSFLNFVIYVICANKYKSKKAA 593
>UniRef100_Q9ZTX4 LeOPT1 [Lycopersicon esculentum]
Length = 580
Score = 473 bits (1217), Expect = e-132
Identities = 239/572 (41%), Positives = 362/572 (62%), Gaps = 12/572 (2%)
Query: 1 GKFKEETEEL-TLDGSVDWHGRPSIRATSGRWFAGTIILLNQGLATLAFFGVGVNLVLFL 59
G ++E L T DGSVD G P +++ +G W A IL N+ LA++G+ NLV +L
Sbjct: 10 GLLEDENSGLYTRDGSVDIKGNPVLKSETGNWRACPFILGNECCERLAYYGIAANLVTYL 69
Query: 60 TRVLGQDNAAAANNVSKWTGTVYIFSLVGAFLSDSYWGRYKTCAIFQGIFVTGLVSLSVT 119
T+ L + N +AA NV+ W GT YI L+GA L+D+YWGRY T A F I+ G+ +L+++
Sbjct: 70 TKKLHEGNVSAARNVTTWQGTCYITPLIGAVLADAYWGRYWTIATFSTIYFIGMCTLTLS 129
Query: 120 TYLALLRPKGCGNGKLECGEHSSLEMGMFYLSIYLIALGNGGYQPNIATFGADQFDEDHS 179
+ +P C C S + +F+ +YLIALG GG +P +++FGADQFD+
Sbjct: 130 ASVPAFKPPQCVGSV--CPSASPAQYAIFFFGLYLIALGTGGIKPCVSSFGADQFDDTDP 187
Query: 180 KESYSKVAFFSYFYLALNLGSLFSNTILGYFEDEGLWALGFWASAGSAFLALVLFLVGTP 239
KE K +FF++FY ++N+G+L S++++ + ++ W LGF A +A+ F GTP
Sbjct: 188 KERVKKGSFFNWFYFSINIGALISSSLIVWIQENAGWGLGFGIPAVFMGIAIASFFFGTP 247
Query: 240 KYRHFKPCGNPLSRFCQVFFAASRKLGVQMTSNGDDLYVIDEKESSNNSNRKILHTHGFK 299
YR KP G+PL+R CQV A K + + + LY +K S+ +RK+LHT +
Sbjct: 248 LYRFQKPGGSPLTRMCQVLVAVFHKWNLSVPDDSTLLYETPDKSSAIEGSRKLLHTDELR 307
Query: 300 FLDRAAYITSRDLEVQKGGQHNPWYLCPITQVEEVKCILRLLPIWLCTIIYSVVFTQMAS 359
LD+AA ++ D E+ G N W LC +TQVEE+K ++R+ PIW I++S V+ QM++
Sbjct: 308 CLDKAAVVS--DNELTTGDYSNAWRLCTVTQVEELKILIRMFPIWATGIVFSAVYAQMST 365
Query: 360 LFVEQGAAMKTTISSFKIPPASMSSFDILSVAIFIFFYRRVLDPLVGKLKKSSSKGLTEL 419
+FVEQG M T + SFKIP AS+S+FD +SV +++ Y ++L P+ + +G +EL
Sbjct: 366 MFVEQGMVMDTAVGSFKIPAASLSTFDTISVIVWVPVYDKILVPIARRF-TGIERGFSEL 424
Query: 420 QRMGIGLVIAIIAMVTAGIVECYRLKYAK------QGDTSSLSIFWQIPQYALIGASEVF 473
QRMGIGL ++++ M A IVE RL+ A+ + + LSIFWQIPQY ++GA+E+F
Sbjct: 425 QRMGIGLFLSMLCMSAAAIVEIRRLQLARDLGLVDEAVSVPLSIFWQIPQYFILGAAEIF 484
Query: 474 MYVGQLEFFNAQTPDGLKSFGSALCMTSISLGNYVSSLIVSIVMKISTEDHMPGWIPGNL 533
++GQLEFF Q+PD ++S SAL + + +LGNY+SS I+++V I+T PGWIP NL
Sbjct: 485 TFIGQLEFFYDQSPDAMRSLCSALSLLTTALGNYLSSFILTVVTSITTRGGKPGWIPNNL 544
Query: 534 NRGHLDRFFFLLAVLTSLDLIAYIACAKWFQN 565
N GHLD FF+LLA L+ +L+ Y+ + +++
Sbjct: 545 NGGHLDYFFWLLAALSFFNLVIYVFLCQMYKS 576
>UniRef100_Q9M390 Putative peptide transport protein [Arabidopsis thaliana]
Length = 570
Score = 468 bits (1204), Expect = e-130
Identities = 244/566 (43%), Positives = 369/566 (65%), Gaps = 12/566 (2%)
Query: 5 EETEELTLDGSVDWHGRPSIRATSGRWFAGTIILLNQGLATLAFFGVGVNLVLFLTRVLG 64
EE + T DG+VD H P+ + +G W A IL N+ LA++G+G NLV +L L
Sbjct: 2 EEKDVYTQDGTVDIHKNPANKEKTGNWKACRFILGNECCERLAYYGMGTNLVNYLESRLN 61
Query: 65 QDNAAAANNVSKWTGTVYIFSLVGAFLSDSYWGRYKTCAIFQGIFVTGLVSLSVTTYLAL 124
Q NA AANNV+ W+GT YI L+GAF++D+Y GRY T A F I+V+G+ L+++ +
Sbjct: 62 QGNATAANNVTNWSGTCYITPLIGAFIADAYLGRYWTIATFVFIYVSGMTLLTLSASVPG 121
Query: 125 LRPKGCGNGKLECGEHSSLEMGMFYLSIYLIALGNGGYQPNIATFGADQFDEDHSKESYS 184
L+P C C +SS + +F++++Y+IALG GG +P +++FGADQFDE+ E
Sbjct: 122 LKPGNCNADT--CHPNSS-QTAVFFVALYMIALGTGGIKPCVSSFGADQFDENDENEKIK 178
Query: 185 KVAFFSYFYLALNLGSLFSNTILGYFEDEGLWALGFWASAGSAFLALVLFLVGTPKYRHF 244
K +FF++FY ++N+G+L + T+L + + W GF + +A+ F G+ YR
Sbjct: 179 KSSFFNWFYFSINVGALIAATVLVWIQMNVGWGWGFGVPTVAMVIAVCFFFFGSRFYRLQ 238
Query: 245 KPCGNPLSRFCQVFFAASRKLGVQMTSNGDDLYVIDEKESSNNSNRKILHTHGFKFLDRA 304
+P G+PL+R QV AA RK+ V++ + L+ + ES+ +RK++HT KF D+A
Sbjct: 239 RPGGSPLTRIFQVIVAAFRKISVKVPEDKSLLFETADDESNIKGSRKLVHTDNLKFFDKA 298
Query: 305 AYITSRDLEVQKGGQHNPWYLCPITQVEEVKCILRLLPIWLCTIIYSVVFTQMASLFVEQ 364
A + D K G+ NPW LC +TQVEE+K I+ LLP+W I+++ V++QM+++FV Q
Sbjct: 299 AVESQSDSI--KDGEVNPWRLCSVTQVEELKSIITLLPVWATGIVFATVYSQMSTMFVLQ 356
Query: 365 GAAMKTTIS-SFKIPPASMSSFDILSVAIFIFFYRRVLDPLVGKLKKSSSKGLTELQRMG 423
G M + +F+IP AS+S FD +SV + Y + + PL K ++ +G T+LQRMG
Sbjct: 357 GNTMDQHMGKNFEIPSASLSLFDTVSVLFWTPVYDQFIIPLARKFTRNE-RGFTQLQRMG 415
Query: 424 IGLVIAIIAMVTAGIVECYRLKYAKQGDTSS-----LSIFWQIPQYALIGASEVFMYVGQ 478
IGLV++I AM+TAG++E RL Y K + +SIFWQIPQY LIG +EVF ++GQ
Sbjct: 416 IGLVVSIFAMITAGVLEVVRLDYVKTHNAYDQKQIHMSIFWQIPQYLLIGCAEVFTFIGQ 475
Query: 479 LEFFNAQTPDGLKSFGSALCMTSISLGNYVSSLIVSIVMKISTEDHMPGWIPGNLNRGHL 538
LEFF Q PD ++S SAL +T+++LGNY+S+++V++VMKI+ ++ PGWIP NLNRGHL
Sbjct: 476 LEFFYDQAPDAMRSLCSALSLTTVALGNYLSTVLVTVVMKITKKNGKPGWIPDNLNRGHL 535
Query: 539 DRFFFLLAVLTSLDLIAYIACAKWFQ 564
D FF+LLA L+ L+ + Y+ +K ++
Sbjct: 536 DYFFYLLATLSFLNFLVYLWISKRYK 561
>UniRef100_Q9FU86 Putative peptide transport protein [Oryza sativa]
Length = 580
Score = 464 bits (1195), Expect = e-129
Identities = 248/574 (43%), Positives = 370/574 (64%), Gaps = 15/574 (2%)
Query: 11 TLDGSVDWHGRPSIRATSGRWFAGTIILLNQGLATLAFFGVGVNLVLFLTRVLGQDNAAA 70
T DG+VD G P+ + +G W A IL N+ LA++G+ NLV ++ LGQ++A A
Sbjct: 10 TQDGTVDVKGNPATKKNTGNWRACPYILANECCERLAYYGMSTNLVNYMKTRLGQESAIA 69
Query: 71 ANNVSKWTGTVYIFSLVGAFLSDSYWGRYKTCAIFQGIFVTGLVSLSVTTYLALLRPKGC 130
ANNV+ W+GT YI L+GAFL+D+Y GR+ T A F I++ GL L++ + + L P C
Sbjct: 70 ANNVTNWSGTCYITPLLGAFLADAYMGRFWTIASFMIIYILGLALLTMASSVKGLVP-AC 128
Query: 131 GNGKLECGEHSSLEMGMFYLSIYLIALGNGGYQPNIATFGADQFDEDHSKESYSKVAFFS 190
G E + G+ +L++YLIALG GG +P +++FGADQFDE+ E SK +FF+
Sbjct: 129 DGGACHPTE---AQTGVVFLALYLIALGTGGIKPCVSSFGADQFDENDEGEKRSKSSFFN 185
Query: 191 YFYLALNLGSLFSNTILGYFEDEGLWALGFWASAGSAFLALVLFLVGTPKYRHFKPCGNP 250
+FY ++N+G+L ++++L Y + W GF A +A+ F VGTP YRH +P G+P
Sbjct: 186 WFYFSINIGALVASSVLVYVQTHVGWGWGFGIPAVVMAVAVASFFVGTPLYRHQRPGGSP 245
Query: 251 LSRFCQVFFAASRKLGVQMTSNGDDLYVIDEKESSNNSNRKILHTHGFKFLDRAAYITSR 310
L+R QV A++RK GV++ ++G L+ ++ES +RK+ HT F LDRAA T
Sbjct: 246 LTRIAQVLVASARKWGVEVPADGSRLHETLDRESGIEGSRKLEHTGQFACLDRAAVETPE 305
Query: 311 DLEVQKGGQHNPWYLCPITQVEEVKCILRLLPIWLCTIIYSVVFTQMASLFVEQGAAMKT 370
D + + W LC +TQVEE+K ++RLLPIW I+++ V+ QM+++FV QG +
Sbjct: 306 D---RSAANASAWRLCTVTQVEELKSVVRLLPIWASGIVFATVYGQMSTMFVLQGNTLDA 362
Query: 371 TIS-SFKIPPASMSSFDILSVAIFIFFYRRVLDPLVGKLKKSSSKGLTELQRMGIGLVIA 429
++ F IP AS+S FD LSV +++ Y R++ P V + +G T+LQRMGIGLVI+
Sbjct: 363 SMGPHFSIPAASLSIFDTLSVIVWVPVYDRLIVPAV-RAVTGRPRGFTQLQRMGIGLVIS 421
Query: 430 IIAMVTAGIVECYRLK-YAKQG-----DTSSLSIFWQIPQYALIGASEVFMYVGQLEFFN 483
+ +M+ AG+++ RL+ A+ G D +SIFWQ+PQY +IGA+EVF +VGQLEFF
Sbjct: 422 VFSMLAAGVLDVVRLRAIARHGLYGDKDVVPISIFWQVPQYFIIGAAEVFTFVGQLEFFY 481
Query: 484 AQTPDGLKSFGSALCMTSISLGNYVSSLIVSIVMKISTEDHMPGWIPGNLNRGHLDRFFF 543
Q PD ++S SAL +T+++LGNY+S+L+V+IV ++T + GWIP NLNRGHLD FF+
Sbjct: 482 DQAPDAMRSMCSALSLTTVALGNYLSTLLVTIVTHVTTRNGAVGWIPDNLNRGHLDYFFW 541
Query: 544 LLAVLTSLDLIAYIACAKWFQNIQMACKYDNNDE 577
LLAVL+ ++ Y+ A W+ + A D+ E
Sbjct: 542 LLAVLSLINFGVYLVIASWYTYKKTADSPDDKAE 575
>UniRef100_Q7XAC3 Peptide transporter 1 [Vicia faba]
Length = 584
Score = 463 bits (1191), Expect = e-129
Identities = 246/568 (43%), Positives = 364/568 (63%), Gaps = 13/568 (2%)
Query: 5 EETEELTLDGSVDWHGRPSIRATSGRWFAGTIILLNQGLATLAFFGVGVNLVL-FLTRVL 63
EE++ T DGSVD+ GRP ++ +G W A IL N+ LA++G+ NLV L L
Sbjct: 19 EESKLYTGDGSVDFKGRPVLKKNTGNWKACPFILGNECCERLAYYGIATNLVKPILLAKL 78
Query: 64 GQDNAAAANNVSKWTGTVYIFSLVGAFLSDSYWGRYKTCAIFQGIFVTGLVSLSVTTYLA 123
+ N +AA NV+ W GT Y+ L+GA L+DSYWGRY T AIF I+ G+ +L+++ +
Sbjct: 79 HEGNVSAARNVTTWQGTCYLAPLIGAVLADSYWGRYWTIAIFSMIYFIGMGTLTLSASIP 138
Query: 124 LLRPKGCGNGKLECGEHSSLEMGMFYLSIYLIALGNGGYQPNIATFGADQFDEDHSKESY 183
L+P C C + + +F++ +YLIALG GG +P +++FGADQFD+ S+E
Sbjct: 139 ALKPAECLGAV--CPPATPAQYAVFFIGLYLIALGTGGIKPCVSSFGADQFDDTDSRERV 196
Query: 184 SKVAFFSYFYLALNLGSLFSNTILGYFEDEGLWALGFWASAGSAFLALVLFLVGTPKYRH 243
K +FF++FY ++N+G+L S++ + + ++ W LGF A LA+ F +GTP YR
Sbjct: 197 KKGSFFNWFYFSINIGALISSSFIVWIQENAGWGLGFGIPALFMGLAIGSFFLGTPLYRF 256
Query: 244 FKPCGNPLSRFCQVFFAASRKLGVQMTSNGDDLYVIDEKESSNNSNRKILHTHGFKFLDR 303
KP G+PL+R CQV A+ RK + + + LY +K S+ +RK+ H+ + LDR
Sbjct: 257 QKPGGSPLTRMCQVVAASFRKRNLTVPEDSSLLYETPDKSSAIEGSRKLQHSDELRCLDR 316
Query: 304 AAYITSRDLEVQKGGQHNPWYLCPITQVEEVKCILRLLPIWLCTIIYSVVFTQMASLFVE 363
AA I+ D E ++G N W LC +TQVEE+K ++R+ P+W I++S V+ QM+++FVE
Sbjct: 317 AAVIS--DDERKRGDYSNLWRLCTVTQVEELKILIRMFPVWATGIVFSAVYAQMSTMFVE 374
Query: 364 QGAAMKTTISSFKIPPASMSSFDILSVAIFIFFYRRVLDPLVGKLKKSSSKGLTELQRMG 423
QG M T++ SFKIP AS+S+FD++SV ++ Y R + P+ K +G +ELQRMG
Sbjct: 375 QGTMMDTSVGSFKIPAASLSTFDVISVIFWVPVYDRFIVPIARKF-TGKERGFSELQRMG 433
Query: 424 IGLVIAIIAMVTAGIVECYRLKYAKQGDTSS------LSIFWQIPQYALIGASEVFMYVG 477
IGL I+++ M A IVE RL+ AK+ D L+IF QIPQY L+GA+EVF +VG
Sbjct: 434 IGLFISVLCMSAAAIVEIKRLQLAKELDLVDKAVPVPLTIFLQIPQYFLLGAAEVFTFVG 493
Query: 478 QLEFFNAQTPDGLKSFGSALCMTSISLGNYVSSLIVSIVMKISTEDHMPGWIPGNLNRGH 537
QLEFF Q+PD ++S SAL + + SLGNY+SS I+++V+ +T PGWIP NLN+GH
Sbjct: 494 QLEFFYDQSPDAMRSLCSALSLLTTSLGNYLSSFILTVVLYFTTRGGNPGWIPDNLNKGH 553
Query: 538 LDRFFFLLAVLTSLDLIAYIACAKWFQN 565
LD +F LA L+ L++ YI AK +++
Sbjct: 554 LD-YFSGLAGLSFLNMFLYIVAAKRYKS 580
>UniRef100_O48542 Peptide transporter [Hordeum vulgare]
Length = 579
Score = 461 bits (1186), Expect = e-128
Identities = 248/563 (44%), Positives = 359/563 (63%), Gaps = 16/563 (2%)
Query: 8 EELTLDGSVDWHGRPSIRATSGRWFAGTIILLNQGLATLAFFGVGVNLVLFLTRVLGQDN 67
E T DG+VD G P+++ +G W A IL N+ LA++G+ NLV F+ +G N
Sbjct: 7 EMYTQDGTVDIKGNPALKKDTGNWRACPYILANECCERLAYYGMSTNLVNFMKDRMGMAN 66
Query: 68 AAAANNVSKWTGTVYIFSLVGAFLSDSYWGRYKTCAIFQGIFVTGLVSLSVTTYLALLRP 127
AAAANNV+ W GT YI L+GAFL+D+Y GR+ T A F I++ GL L++ T + L P
Sbjct: 67 AAAANNVTNWGGTCYITPLIGAFLADAYLGRFWTIASFMIIYIFGLGLLTMATSVHGLVP 126
Query: 128 KGCGNGKLECGEHSSLEMGMFYLSIYLIALGNGGYQPNIATFGADQFDEDHSKESYSKVA 187
G + S + ++++YLIALG GG +P +++FGADQFDE E SK +
Sbjct: 127 ACASKGVCDPTPGQSAAV---FIALYLIALGTGGIKPCVSSFGADQFDEHDDVERKSKSS 183
Query: 188 FFSYFYLALNLGSLFSNTILGYFEDEGLWALGFWASAGSAFLALVLFLVGTPKYRHFKPC 247
FF++FY ++N+G+L ++++L Y + W+ GF A +A+ F VGT YRH +P
Sbjct: 184 FFNWFYFSINIGALVASSVLVYVQTHVGWSWGFGIPAVVMAIAVGSFFVGTSLYRHQRPG 243
Query: 248 GNPLSRFCQVFFAASRKLGVQMTSNGDDLYVIDEKESSNNSNRKILHTHGFKFLDRAAYI 307
G+PL+R QV AA+RKLGV + +G LY +KES +RK+ HT F+FLD+AA
Sbjct: 244 GSPLTRIAQVLVAATRKLGVAV--DGSALYETADKESGIEGSRKLEHTRQFRFLDKAAVE 301
Query: 308 TSRDLEVQKGGQHNPWYLCPITQVEEVKCILRLLPIWLCTIIYSVVFTQMASLFVEQGAA 367
T D + +PW LC +TQVEE+K ++RLLPIW I+++ V+ QM+++FV QG
Sbjct: 302 THAD---RTAAAPSPWRLCTVTQVEELKSVVRLLPIWASGIVFATVYGQMSTMFVLQGNT 358
Query: 368 MKTTIS-SFKIPPASMSSFDILSVAIFIFFYRRVLDPLVGKLKKSSSKGLTELQRMGIGL 426
+ ++ FKIP AS+S FD LSV ++ Y R+L P V + +G T+LQRMGIGL
Sbjct: 359 LDASMGPKFKIPSASLSIFDTLSVIAWVPVYDRILVPAVRSVT-GRPRGFTQLQRMGIGL 417
Query: 427 VIAIIAMVTAGIVECYRLKYAKQG------DTSSLSIFWQIPQYALIGASEVFMYVGQLE 480
V+++ AM+ AG++E RL+ Q D +SIFWQ+PQY +IG +EVF +VGQLE
Sbjct: 418 VVSMFAMLAAGVLELVRLRTIAQHGLYGEKDVVPISIFWQVPQYFIIGCAEVFTFVGQLE 477
Query: 481 FFNAQTPDGLKSFGSALCMTSISLGNYVSSLIVSIVMKISTEDHMPGWIPGNLNRGHLDR 540
FF Q PD ++S SAL +T+++LGNY+S+L+V++V K++T GWIP NLN GHLD
Sbjct: 478 FFYDQAPDAMRSMCSALSLTTVALGNYLSTLLVTVVAKVTTRGGKQGWIPDNLNVGHLDY 537
Query: 541 FFFLLAVLTSLDLIAYIACAKWF 563
FF+LLA L+ ++ Y+ A W+
Sbjct: 538 FFWLLAALSLVNFAVYLLIASWY 560
>UniRef100_Q93Z20 At1g62200/F19K23_13 [Arabidopsis thaliana]
Length = 590
Score = 460 bits (1183), Expect = e-128
Identities = 236/551 (42%), Positives = 354/551 (63%), Gaps = 7/551 (1%)
Query: 13 DGSVDWHGRPSIRATSGRWFAGTIILLNQGLATLAFFGVGVNLVLFLTRVLGQDNAAAAN 72
DGS+D +G P + +G W A IL N+ LA++G+ NL+ + T L + N +AA+
Sbjct: 38 DGSIDIYGNPPSKKKTGNWKACPFILGNECCERLAYYGIAKNLITYYTSELHESNVSAAS 97
Query: 73 NVSKWTGTVYIFSLVGAFLSDSYWGRYKTCAIFQGIFVTGLVSLSVTTYLALLRPKGC-G 131
+V W GT YI L+GA ++DSYWGRY T A F I+ G+ L+++ L +L+P C G
Sbjct: 98 DVMIWQGTCYITPLIGAVIADSYWGRYWTIASFSAIYFIGMALLTLSASLPVLKPAACAG 157
Query: 132 NGKLECGEHSSLEMGMFYLSIYLIALGNGGYQPNIATFGADQFDEDHSKESYSKVAFFSY 191
C ++++ +F+ +YLIALG GG +P +++FGADQFD+ +E K +FF++
Sbjct: 158 VAAALCSPATTVQYAVFFTGLYLIALGTGGIKPCVSSFGADQFDDTDPRERVRKASFFNW 217
Query: 192 FYLALNLGSLFSNTILGYFEDEGLWALGFWASAGSAFLALVLFLVGTPKYRHFKPCGNPL 251
FY ++N+GS S+T+L + ++ W LGF +++ F +GTP YR KP G+P+
Sbjct: 218 FYFSINIGSFISSTLLVWVQENVGWGLGFLIPTVFMGVSIASFFIGTPLYRFQKPGGSPI 277
Query: 252 SRFCQVFFAASRKLGVQMTSNGDDLYVIDEKESSNNSNRKILHTHGFKFLDRAAYITSRD 311
+R CQV AA RKL + + + LY EK S +RKI HT G+KFLD+AA I+ +
Sbjct: 278 TRVCQVLVAAYRKLKLNLPEDISFLYETREKNSMIAGSRKIQHTDGYKFLDKAAVIS--E 335
Query: 312 LEVQKGGQHNPWYLCPITQVEEVKCILRLLPIWLCTIIYSVVFTQMASLFVEQGAAMKTT 371
E + G NPW LC +TQVEEVK ++R+ PIW I+YSV+++Q+++LFV+QG +M
Sbjct: 336 YESKSGAFSNPWKLCTVTQVEEVKTLIRMFPIWASGIVYSVLYSQISTLFVQQGRSMNRI 395
Query: 372 ISSFKIPPASMSSFDILSVAIFIFFYRRVLDPLVGKLKKSSSKGLTELQRMGIGLVIAII 431
I SF+IPPAS FD L V I I Y R L P V + KGLT+LQRMGIGL ++++
Sbjct: 396 IRSFEIPPASFGVFDTLIVLISIPIYDRFLVPFVRRF-TGIPKGLTDLQRMGIGLFLSVL 454
Query: 432 AMVTAGIVECYRLKYAKQGDTSSLSIFWQIPQYALIGASEVFMYVGQLEFFNAQTPDGLK 491
++ A IVE RL+ A+ D ++SIFWQIPQY L+G +EVF ++G++EFF ++PD ++
Sbjct: 455 SIAAAAIVETVRLQLAQ--DFVAMSIFWQIPQYILMGIAEVFFFIGRVEFFYDESPDAMR 512
Query: 492 SFGSALCMTSISLGNYVSSLIVSIVMKISTEDHMPGWIPGNLNRGHLDRFFFLLAVLTSL 551
S SAL + + ++G+Y+SSLI+++V + GW+P +LN+GHLD FF+LL L +
Sbjct: 513 SVCSALALLNTAVGSYLSSLILTLVAYFTALGGKDGWVPDDLNKGHLDYFFWLLVSLGLV 572
Query: 552 DLIAY-IACAK 561
++ Y + C K
Sbjct: 573 NIPVYALICVK 583
>UniRef100_P46032 Peptide transporter PTR2-B [Arabidopsis thaliana]
Length = 585
Score = 459 bits (1180), Expect = e-127
Identities = 238/566 (42%), Positives = 353/566 (62%), Gaps = 11/566 (1%)
Query: 1 GKFKEETEELTLDGSVDWHGRPSIRATSGRWFAGTIILLNQGLATLAFFGVGVNLVLFLT 60
G +E + DGSVD++G P ++ +G W A IL N+ LA++G+ NL+ +LT
Sbjct: 15 GLILQEVKLYAEDGSVDFNGNPPLKEKTGNWKACPFILGNECCERLAYYGIAGNLITYLT 74
Query: 61 RVLGQDNAAAANNVSKWTGTVYIFSLVGAFLSDSYWGRYKTCAIFQGIFVTGLVSLSVTT 120
L Q N +AA NV+ W GT Y+ L+GA L+D+YWGRY T A F GI+ G+ +L+++
Sbjct: 75 TKLHQGNVSAATNVTTWQGTCYLTPLIGAVLADAYWGRYWTIACFSGIYFIGMSALTLSA 134
Query: 121 YLALLRPKGCGNGKLECGEHSSLEMGMFYLSIYLIALGNGGYQPNIATFGADQFDEDHSK 180
+ L+P C C + + MF+ +YLIALG GG +P +++FGADQFD+ S+
Sbjct: 135 SVPALKPAECIGDF--CPSATPAQYAMFFGGLYLIALGTGGIKPCVSSFGADQFDDTDSR 192
Query: 181 ESYSKVAFFSYFYLALNLGSLFSNTILGYFEDEGLWALGFWASAGSAFLALVLFLVGTPK 240
E K +FF++FY ++N+G+L S+++L + ++ W LGF LA+ F GTP
Sbjct: 193 ERVRKASFFNWFYFSINIGALVSSSLLVWIQENRGWGLGFGIPTVFMGLAIASFFFGTPL 252
Query: 241 YRHFKPCGNPLSRFCQVFFAASRKLGVQMTSNGDDLYVIDEKESSNNSNRKILHTHGFKF 300
YR KP G+P++R QV A+ RK V++ + LY +K S+ +RKI HT ++
Sbjct: 253 YRFQKPGGSPITRISQVVVASFRKSSVKVPEDATLLYETQDKNSAIAGSRKIEHTDDCQY 312
Query: 301 LDRAAYITSRDLEVQKGGQHNPWYLCPITQVEEVKCILRLLPIWLCTIIYSVVFTQMASL 360
LD+AA I+ E + G N W LC +TQVEE+K ++R+ PIW II+S V+ QM+++
Sbjct: 313 LDKAAVISEE--ESKSGDYSNSWRLCTVTQVEELKILIRMFPIWASGIIFSAVYAQMSTM 370
Query: 361 FVEQGAAMKTTISSFKIPPASMSSFDILSVAIFIFFYRRVLDPLVGKLKKSSSKGLTELQ 420
FV+QG AM I SF++PPA++ +FD SV I++ Y R + PL K KG TE+Q
Sbjct: 371 FVQQGRAMNCKIGSFQLPPAALGTFDTASVIIWVPLYDRFIVPLARKF-TGVDKGFTEIQ 429
Query: 421 RMGIGLVIAIIAMVTAGIVECYRLKYA------KQGDTSSLSIFWQIPQYALIGASEVFM 474
RMGIGL ++++ M A IVE RL A + G +S+ WQIPQY ++GA+EVF
Sbjct: 430 RMGIGLFVSVLCMAAAAIVEIIRLHMANDLGLVESGAPVPISVLWQIPQYFILGAAEVFY 489
Query: 475 YVGQLEFFNAQTPDGLKSFGSALCMTSISLGNYVSSLIVSIVMKISTEDHMPGWIPGNLN 534
++GQLEFF Q+PD ++S SAL + + +LGNY+SSLI+++V +T + GWI NLN
Sbjct: 490 FIGQLEFFYDQSPDAMRSLCSALALLTNALGNYLSSLILTLVTYFTTRNGQEGWISDNLN 549
Query: 535 RGHLDRFFFLLAVLTSLDLIAYIACA 560
GHLD FF+LLA L+ +++ Y A
Sbjct: 550 SGHLDYFFWLLAGLSLVNMAVYFFSA 575
>UniRef100_Q9SDN4 Amino acid/peptide transporter [Prunus dulcis]
Length = 559
Score = 441 bits (1135), Expect = e-122
Identities = 231/548 (42%), Positives = 339/548 (61%), Gaps = 12/548 (2%)
Query: 1 GKFKEETEEL-TLDGSVDWHGRPSIRATSGRWFAGTIILLNQGLATLAFFGVGVNLVLFL 59
G ++ET L T DGSVD G+P ++ ++G W A IL + LAF+G+ NLV +L
Sbjct: 14 GLIQDETNGLYTGDGSVDITGKPVLKQSTGNWXACPFILGTECCERLAFYGISTNLVTYL 73
Query: 60 TRVLGQDNAAAANNVSKWTGTVYIFSLVGAFLSDSYWGRYKTCAIFQGIFVTGLVSLSVT 119
T L + N +AA NV+ W+GT Y+ L+GA L+D+YWGRY T AIF I+ G+ +L+++
Sbjct: 74 THKLHEGNVSAARNVTTWSGTCYLTPLIGAVLADAYWGRYWTIAIFSTIYFIGMCTLTIS 133
Query: 120 TYLALLRPKGCGNGKLECGEHSSLEMGMFYLSIYLIALGNGGYQPNIATFGADQFDEDHS 179
+ L+P C + C S + G+F+ +YLIAL GG +P +++FGADQFD+ S
Sbjct: 134 ASVPALKPPQCVDSV--CPSASPAQYGVFFFGLYLIALRTGGIKPCVSSFGADQFDDTDS 191
Query: 180 KESYSKVAFFSYFYLALNLGSLFSNTILGYFEDEGLWALGFWASAGSAFLALVLFLVGTP 239
+E K +FF++FY ++N+G+L S+T++ + +D W LGF A +A+V F GTP
Sbjct: 192 RERVKKGSFFNWFYFSINIGALVSSTLIVWVQDNAGWGLGFGIPALFMGIAIVSFFSGTP 251
Query: 240 KYRHFKPCGNPLSRFCQVFFAASRKLGVQMTSNGDDLYVIDEKESSNNSNRKILHTHGFK 299
YR KP G+PL+R CQV A+ RK + + + LY +K S+ +RK+ H+
Sbjct: 252 LYRFQKPGGSPLTRMCQVLVASFRKWNLDVPRDSSLLYETQDKGSAIKGSRKLEHSDELN 311
Query: 300 FLDRAAYITSRDLEVQKGGQHNPWYLCPITQVEEVKCILRLLPIWLCTIIYSVVFTQMAS 359
LD+AA I+ + E + G NPW +C +TQVEE+K ++R+ PIW I++S V+ QMA+
Sbjct: 312 CLDKAAVIS--ETETKTGDFSNPWRICTVTQVEELKILIRMFPIWATGIVFSAVYAQMAT 369
Query: 360 LFVEQGAAMKTTISSFKIPPASMSSFDILSVAIFIFFYRRVLDPLVGKLKKSSSKGLTEL 419
+FVEQG M T++ SF IPPAS+SSFD++SV ++ Y R + P+ K +G +EL
Sbjct: 370 MFVEQGMMMDTSVGSFTIPPASLSSFDVISVIFWVPIYDRFIVPIARKF-TGKERGFSEL 428
Query: 420 QRMGIGLVIAIIAMVTAGIVECYRLKYAKQ------GDTSSLSIFWQIPQYALIGASEVF 473
QRMGIGL ++++ M A +VE RL+ A + LSIFWQIPQY L+GA+E+F
Sbjct: 429 QRMGIGLFLSVLCMSAAAVVEMKRLQLATELGLVDKEVAVPLSIFWQIPQYFLLGAAEIF 488
Query: 474 MYVGQLEFFNAQTPDGLKSFGSALCMTSISLGNYVSSLIVSIVMKISTEDHMPGWIPGNL 533
++GQLEFF Q+ D ++S SAL + +G IV +T+ GWIP NL
Sbjct: 489 TFIGQLEFFYDQSSDAMRSLCSALSASDDFIGKLSELFDSDIVTYFTTQGGKAGWIPDNL 548
Query: 534 NRGHLDRF 541
N GHLD F
Sbjct: 549 NDGHLDYF 556
>UniRef100_O04585 F19K23.13 protein [Arabidopsis thaliana]
Length = 568
Score = 441 bits (1135), Expect = e-122
Identities = 232/553 (41%), Positives = 348/553 (61%), Gaps = 14/553 (2%)
Query: 11 TLDGSVDWHGRPSIRATSGRWFAGTIILLNQGLATLAFFGVGVNLVLFLTRVLGQDNAAA 70
T DGS+D +G P + +G W A IL N+ LA++G+ NL+ + T L + N +A
Sbjct: 21 TEDGSIDIYGNPPSKKKTGNWKACPFILGNECCERLAYYGIAKNLITYYTSELHESNVSA 80
Query: 71 ANNVSKWTGTVYIFSLVGAFLSDSYWGRYKTCAIFQGIFVTGLVSLSVTTYLALLRPKGC 130
A++V W GT YI L+GA ++DSYWGRY T A F I+ G+ L+++ L +L+P C
Sbjct: 81 ASDVMIWQGTCYITPLIGAVIADSYWGRYWTIASFSAIYFIGMALLTLSASLPVLKPAAC 140
Query: 131 -GNGKLECGEHSSLEMGMFYLSIYLIALGNGGYQPNIATFGADQFDEDHSKESYSKVAFF 189
G C ++++ +F+ +YLIALG GG +P +++FGADQFD+ +E K +FF
Sbjct: 141 AGVAAALCSPATTVQYAVFFTGLYLIALGTGGIKPCVSSFGADQFDDTDPRERVRKASFF 200
Query: 190 SYFYLALNLGSLFSNTILGYFEDEGLWALGFWASAGSAFLALVLFLVGTPKYRHFKPCGN 249
++FY ++N+GS S+T+L + ++ W LGF +++ F +GTP YR KP G+
Sbjct: 201 NWFYFSINIGSFISSTLLVWVQENVGWGLGFLIPTVFMGVSIASFFIGTPLYRFQKPGGS 260
Query: 250 PLSRFCQVFFAASRKLGVQMTSNGDDLYVIDEKESSNNSNRKILHTHGFKFLDRAAYITS 309
P++R CQV AA RKL + + + LY EK S +RKI HT AA I+
Sbjct: 261 PITRVCQVLVAAYRKLKLNLPEDISFLYETREKNSMIAGSRKIQHTD-------AAVIS- 312
Query: 310 RDLEVQKGGQHNPWYLCPITQVEEVKCILRLLPIWLCTIIYSVVFTQMASLFVEQGAAMK 369
+ E + G NPW LC +TQVEEVK ++R+ PIW I+YSV+++Q+++LFV+QG +M
Sbjct: 313 -EYESKSGAFSNPWKLCTVTQVEEVKTLIRMFPIWASGIVYSVLYSQISTLFVQQGRSMN 371
Query: 370 TTISSFKIPPASMSSFDILSVAIFIFFYRRVLDPLVGKLKKSSSKGLTELQRMGIGLVIA 429
I SF+IPPAS FD L V I I Y R L P V + KGLT+LQRMGIGL ++
Sbjct: 372 RIIRSFEIPPASFGVFDTLIVLISIPIYDRFLVPFVRRF-TGIPKGLTDLQRMGIGLFLS 430
Query: 430 IIAMVTAGIVECYRLKYAKQGDTSSLSIFWQIPQYALIGASEVFMYVGQLEFFNAQTPDG 489
++++ A IVE RL+ A+ D ++SIFWQIPQY L+G +EVF ++G++EFF ++PD
Sbjct: 431 VLSIAAAAIVETVRLQLAQ--DFVAMSIFWQIPQYILMGIAEVFFFIGRVEFFYDESPDA 488
Query: 490 LKSFGSALCMTSISLGNYVSSLIVSIVMKISTEDHMPGWIPGNLNRGHLDRFFFLLAVLT 549
++S SAL + + ++G+Y+SSLI+++V + GW+P +LN+GHLD FF+LL L
Sbjct: 489 MRSVCSALALLNTAVGSYLSSLILTLVAYFTALGGKDGWVPDDLNKGHLDYFFWLLVSLG 548
Query: 550 SLDLIAY-IACAK 561
+++ Y + C K
Sbjct: 549 LVNIPVYALICVK 561
>UniRef100_Q69LA3 Putative peptide transport protein [Oryza sativa]
Length = 572
Score = 438 bits (1126), Expect = e-121
Identities = 234/565 (41%), Positives = 364/565 (64%), Gaps = 15/565 (2%)
Query: 8 EELTLDGSVDWHGRPSIRATSGRWFAGTIILLNQGLATLAFFGVGVNLVLFLTRVLGQDN 67
E T DG+ D G+P++R+ SG W A IL N+ LA++G+ NLV ++ L Q N
Sbjct: 2 EATTTDGTTDHAGKPAVRSKSGTWRACPFILGNECCERLAYYGMSANLVNYMVDRLRQGN 61
Query: 68 AAAANNVSKWTGTVYIFSLVGAFLSDSYWGRYKTCAIFQGIFVTGLVSLSVTTYLALLRP 127
A AA +V+ W+GT Y+ LVGAFL+D+Y GRY+T A F +++ GL L+++ + ++P
Sbjct: 62 AGAAASVNNWSGTCYVMPLVGAFLADAYLGRYRTIAAFMALYIVGLALLTMSASVPGMKP 121
Query: 128 KGCGNGKLECGEHSSLEMGMFYLSIYLIALGNGGYQPNIATFGADQFDEDHSKESYSKVA 187
C S + F++++YLIALG GG +P +++FGADQFD+ +E SK +
Sbjct: 122 PNCATISASSCGPSPGQSAAFFVALYLIALGTGGIKPCVSSFGADQFDDADPREHRSKAS 181
Query: 188 FFSYFYLALNLGSLFSNTILGYFEDEGLWALGFWASAGSAFLALVLFLVGTPKYRHFKPC 247
FF++FY+++N+G+L ++++L + + W GF A + +A+ FL+G+ YRH KP
Sbjct: 182 FFNWFYMSINVGALVASSVLVWVQMNVGWGWGFGIPAVAMAVAVASFLMGSSLYRHQKPG 241
Query: 248 GNPLSRFCQVFFAASRKLGVQMTSNGDDLYVIDEKESSNNSNRKILHTHGFKFLDRAAYI 307
G+PL+R QV AA+RK V + + D ++ E + R++ HT F++LDRAA +
Sbjct: 242 GSPLTRMLQVVVAAARKSRVALPA--DAAALLYEGDKLACGTRRLAHTEQFRWLDRAAVV 299
Query: 308 T-SRDLEVQKGGQHNPWYLCPITQVEEVKCILRLLPIWLCTIIYSVVFTQMASLFVEQGA 366
T + D + G + W LCP+TQVEE+K ++RLLP+W I+ S V+ QM+++FV QG
Sbjct: 300 TPTTDKDDDTGSR---WRLCPVTQVEELKAVVRLLPVWASGIVMSAVYGQMSTMFVLQGN 356
Query: 367 AMKTTI-SSFKIPPASMSSFDILSVAIFIFFYRRVLDPLVGKLKKSSSKGLTELQRMGIG 425
+ + ++FKIP AS+S FD L+V ++ Y R++ P + +G T+LQRMGIG
Sbjct: 357 TLDPRMGATFKIPSASLSIFDTLAVLAWVPVYDRLIVPAARRF-TGHPRGFTQLQRMGIG 415
Query: 426 LVIAIIAMVTAGIVECYRLKYAKQG---DTSS---LSIFWQIPQYALIGASEVFMYVGQL 479
L+I++ +MV AG++E RL+ A D++S +SIFWQ+ QY +IGA+EVF ++GQ+
Sbjct: 416 LLISVFSMVAAGVLEVVRLRVAAAHGMLDSTSYLPISIFWQV-QYFIIGAAEVFAFIGQI 474
Query: 480 EFFNAQTPDGLKSFGSALCMTSISLGNYVSSLIVSIVMKISTEDHMPGWIPGNLNRGHLD 539
+FF Q PD ++S +AL +TS +LGNY+S+L+V IV ST GWIP NLNRGHLD
Sbjct: 475 DFFYDQAPDDMRSTCTALSLTSSALGNYLSTLLVVIVTAASTRGGGLGWIPDNLNRGHLD 534
Query: 540 RFFFLLAVLTSLDLIAYIACAKWFQ 564
FF+LLA L++++ + Y+ A W++
Sbjct: 535 YFFWLLAALSAVNFLVYLWIANWYR 559
>UniRef100_Q9LFB8 Oligopeptide transporter-like protein [Arabidopsis thaliana]
Length = 570
Score = 421 bits (1083), Expect = e-116
Identities = 224/567 (39%), Positives = 353/567 (61%), Gaps = 15/567 (2%)
Query: 4 KEETEELTLDGSVDWHGRPSIRATSGRWFAGTIILLNQGLATLAFFGVGVNLVLFLTRVL 63
+++ + T DG++D H +P+ + +G W A IL + LA++G+ NL+ +L + +
Sbjct: 2 EDDKDIYTKDGTLDIHKKPANKNKTGTWKACRFILGTECCERLAYYGMSTNLINYLEKQM 61
Query: 64 GQDNAAAANNVSKWTGTVYIFSLVGAFLSDSYWGRYKTCAIFQGIFVTGLVSLSVTTYLA 123
+N +A+ +VS W+GT Y L+GAF++D+Y GRY T A F I++ G+ L+++ +
Sbjct: 62 NMENVSASKSVSNWSGTCYATPLIGAFIADAYLGRYWTIASFVVIYIAGMTLLTISASVP 121
Query: 124 LLRPKGCGNGKLECGEHSSLEMGMFYLSIYLIALGNGGYQPNIATFGADQFDEDHSKESY 183
L P G E ++ + + ++++YLIALG GG +P +++FGADQFD+ KE
Sbjct: 122 GLTPTCSG----ETCHATAGQTAITFIALYLIALGTGGIKPCVSSFGADQFDDTDEKEKE 177
Query: 184 SKVAFFSYFYLALNLGSLFSNTILGYFEDEGLWALGFWASAGSAFLALVLFLVGTPKYRH 243
SK +FF++FY +N+G++ ++++L + + W G + +A+V F G+ YR
Sbjct: 178 SKSSFFNWFYFVINVGAMIASSVLVWIQMNVGWGWGLGVPTVAMAIAVVFFFAGSNFYRL 237
Query: 244 FKPCGNPLSRFCQVFFAASRKLGVQMTSNGDDLYVIDEKESSNNSNRKILHTHGFKFLDR 303
KP G+PL+R QV A+ RK V++ + LY + ESS +RK+ HT F D+
Sbjct: 238 QKPGGSPLTRMLQVIVASCRKSKVKIPEDESLLYENQDAESSIIGSRKLEHTKILTFFDK 297
Query: 304 AAYITSRDLEVQKGG-QHNPWYLCPITQVEEVKCILRLLPIWLCTIIYSVVFTQMASLFV 362
AA T D KG + + W LC +TQVEE+K ++RLLPIW I+++ V++QM ++FV
Sbjct: 298 AAVETESD---NKGAAKSSSWKLCTVTQVEELKALIRLLPIWATGIVFASVYSQMGTVFV 354
Query: 363 EQGAAMKTTIS-SFKIPPASMSSFDILSVAIFIFFYRRVLDPLVGKLKKSSSKGLTELQR 421
QG + + +FKIP AS+S FD LSV + Y +++ P K +G T+LQR
Sbjct: 355 LQGNTLDQHMGPNFKIPSASLSLFDTLSVLFWAPVYDKLIVPFARKYT-GHERGFTQLQR 413
Query: 422 MGIGLVIAIIAMVTAGIVECYRLKYAK-----QGDTSSLSIFWQIPQYALIGASEVFMYV 476
+GIGLVI+I +MV+AGI+E RL Y + +T ++IFWQ+PQY L+G +EVF ++
Sbjct: 414 IGIGLVISIFSMVSAGILEVARLNYVQTHNLYNEETIPMTIFWQVPQYFLVGCAEVFTFI 473
Query: 477 GQLEFFNAQTPDGLKSFGSALCMTSISLGNYVSSLIVSIVMKISTEDHMPGWIPGNLNRG 536
GQLEFF Q PD ++S SAL +T+I+ GNY+S+ +V++V K++ PGWI NLN G
Sbjct: 474 GQLEFFYDQAPDAMRSLCSALSLTAIAFGNYLSTFLVTLVTKVTRSGGRPGWIAKNLNNG 533
Query: 537 HLDRFFFLLAVLTSLDLIAYIACAKWF 563
HLD FF+LLA L+ L+ + Y+ AKW+
Sbjct: 534 HLDYFFWLLAGLSFLNFLVYLWIAKWY 560
>UniRef100_Q6E3L2 Low affinity nitrate transporter NRT1.1 [Triticum aestivum]
Length = 586
Score = 404 bits (1038), Expect = e-111
Identities = 220/574 (38%), Positives = 333/574 (57%), Gaps = 27/574 (4%)
Query: 1 GKFKEETEELTLDGSVDWHGRPSIRATSGRWFAGTIILLNQGLATLAFFGVGVNLVLFLT 60
G +E T E T DGSV G P+ R +G W A +I++ LA+ +G NLV +LT
Sbjct: 17 GSSQENTTEYTGDGSVCISGHPASRKHTGNWKASFLIIVCSFCCYLAYSSIGKNLVSYLT 76
Query: 61 RVLGQDNAAAANNVSKWTGTVYIFSLVGAFLSDSYWGRYKTCAIFQGIFVTGLVSLSVTT 120
+VL + N AA +V+ W GT Y+ LVGAF++DSY G+Y+T I IF+ G++ L ++
Sbjct: 77 KVLHETNLDAARHVATWQGTSYLAPLVGAFVADSYLGKYRTALIACKIFIIGMMMLLLSA 136
Query: 121 YLALLRPKGCGNGKLECGEHS--------SLEMGMFYLSIYLIALGNGGYQPNIATFGAD 172
L L+ G H+ S + +F + +Y++ LG G +P + +FGAD
Sbjct: 137 ALQLI----------SAGPHAWTVWVHLVSSQYTIFLIGLYMVGLGYGAQRPCVTSFGAD 186
Query: 173 QFDEDHSKESYSKVAFFSYFYLALNLGSLFSNTILGYFEDEGLWALGFWASAGSAFLALV 232
QFD+ E K +FF++ Y A+N GSL + T++ + ++ W GF S L +
Sbjct: 187 QFDDTDYVEKTRKSSFFNWHYFAINAGSLIAGTVIVWVQEHEGWLWGFTISTLFVTLGVC 246
Query: 233 LFLVGTPKYRHFKPCGNPLSRFCQVFFAASRKLGVQMTSNGDDLYVIDEKESSNNSNRKI 292
+F +G+ YR KP G+PL+R CQV AA+R + + LY + S+ RK+
Sbjct: 247 IFFLGSIVYRFQKPRGSPLTRLCQVVIAATRNFDKVLPCDSSALYEFMGQGSAIEGRRKL 306
Query: 293 LHTHGFKFLDRAAYITSRDLEVQKGGQHNPWYLCPITQVEEVKCILRLLPIWLCTIIYSV 352
HT G F D+AA +T D E GQHN W +C +TQVEE+K ++R+ PIW I+++
Sbjct: 307 EHTTGLGFFDKAAIVTLPDCE--SPGQHNKWKICTVTQVEELKILIRMFPIWSAMILFAA 364
Query: 353 VFTQMASLFVEQGAAMKTTISSFKIPPASMSSFDILSVAIFIFFYRRVLDPLVGKLKKSS 412
V QM+S FVEQG AM I SF+IP AS D ++V + + Y R++ P++ K
Sbjct: 365 VQEQMSSTFVEQGMAMDKHIGSFEIPSASFQCVDTITVIVLVPIYERLIVPVIRKF-TGR 423
Query: 413 SKGLTELQRMGIGLVIAIIAMVTAGIVECYRLKYAK------QGDTSSLSIFWQIPQYAL 466
+ G+T QR+GIGL ++ +MV+A +VE RL+ A+ + +SI WQ PQY L
Sbjct: 424 ANGITSPQRIGIGLCFSMFSMVSAALVEGNRLQIAQAEGLVHRKVAVPMSIMWQGPQYFL 483
Query: 467 IGASEVFMYVGQLEFFNAQTPDGLKSFGSALCMTSISLGNYVSSLIVSIVMKISTEDHMP 526
+G +EVF +G E F ++PDG++S A + ++S GNY+SSLI+S+V + P
Sbjct: 484 LGVAEVFSNIGLTEAFYDESPDGMRSLCMAFSLVNMSAGNYLSSLILSLVPVFTARGGSP 543
Query: 527 GWIPGNLNRGHLDRFFFLLAVLTSLDLIAYIACA 560
GWIP NLN GHLDRF+ ++A L+ +++ ++ CA
Sbjct: 544 GWIPDNLNEGHLDRFYLMMAGLSFFNIVVFVFCA 577
>UniRef100_Q9SPU1 Nitrate transporter [Oryza sativa]
Length = 584
Score = 402 bits (1034), Expect = e-110
Identities = 225/568 (39%), Positives = 327/568 (56%), Gaps = 27/568 (4%)
Query: 7 TEELTLDGSVDWHGRPSIRATSGRWFAGTIILLNQGLATLAFFGVGVNLVLFLTRVLGQD 66
T E T DGSV G P++R +G W ++ ++ + LAF + NLV +LT+VL +
Sbjct: 21 TLEYTGDGSVCIRGHPALRKHTGNWKGSSLAIVFSFCSYLAFTSIVKNLVSYLTKVLHET 80
Query: 67 NAAAANNVSKWTGTVYIFSLVGAFLSDSYWGRYKTCAIFQGIFVTGLVSLSVTTYLALLR 126
N AAA +V+ W+GT Y+ LVGAFL+DSY G+Y T IF IF+ GL+ L ++ + L+
Sbjct: 81 NVAAARDVATWSGTSYLAPLVGAFLADSYLGKYCTILIFCTIFIIGLMMLLLSAAVPLI- 139
Query: 127 PKGCGNGKLECGEHS--------SLEMGMFYLSIYLIALGNGGYQPNIATFGADQFDEDH 178
G HS S + +F++ +Y++ALG G P I++FGADQFD+
Sbjct: 140 ---------STGPHSWIIWTDPVSSQNIIFFVGLYMVALGYGAQCPCISSFGADQFDDTD 190
Query: 179 SKESYSKVAFFSYFYLALNLGSLFSNTILGYFEDEGLWALGFWASAGSAFLALVLFLVGT 238
E K +FF++ Y N GSL S T++ + +D W GF SA +L F+ G+
Sbjct: 191 ENERTKKSSFFNWTYFVANAGSLISGTVIVWVQDHKGWIWGFTISALFVYLGFGTFIFGS 250
Query: 239 PKYRHFKPCGNPLSRFCQVFFAASRKLGVQMTSNGDDLYVIDEKESSNNSNRKILHTHGF 298
Y PL+R CQV AA K + + LY + S+ +RK+ HT G
Sbjct: 251 SMYDFRNLEEAPLARICQVVVAAIHKRDKDLPCDSSVLYEFLGQSSAIEGSRKLEHTTGL 310
Query: 299 KFLDRAAYITSRDLEVQKGGQHNPWYLCPITQVEEVKCILRLLPIWLCTIIYSVVFTQMA 358
KF DRAA +T D E G N W +C +TQVEE+K ++R+ P+W I+++ V M
Sbjct: 311 KFFDRAAMVTPSDFE--SDGLLNTWKICTVTQVEELKILIRMFPVWATMILFAAVLDNMF 368
Query: 359 SLFVEQGAAMKTTISSFKIPPASMSSFDILSVAIFIFFYRRVLDPLVGKLKKSSSKGLTE 418
S F+EQG M+ I SF+IP AS S D+++V I + Y RVL P+ K + G+T
Sbjct: 369 STFIEQGMVMEKHIGSFEIPAASFQSIDVIAVLILVPVYERVLVPVFRKFT-GRANGITP 427
Query: 419 LQRMGIGLVIAIIAMVTAGIVECYRLKYAKQGD------TSSLSIFWQIPQYALIGASEV 472
LQRMGIGL ++++MV+A +VE RL+ A+ +SI WQ PQY LIG EV
Sbjct: 428 LQRMGIGLFFSMLSMVSAALVESNRLRIAQDEGLVHRKVAVPMSILWQGPQYFLIGVGEV 487
Query: 473 FMYVGQLEFFNAQTPDGLKSFGSALCMTSISLGNYVSSLIVSIVMKISTEDHMPGWIPGN 532
F +G EFF ++PD ++S A + ++S G+Y+SS IVS+V + + PGWIP N
Sbjct: 488 FSNIGLTEFFYQESPDAMRSLCLAFSLANVSAGSYLSSFIVSLVPVFTAREGSPGWIPDN 547
Query: 533 LNRGHLDRFFFLLAVLTSLDLIAYIACA 560
LN GHLDRFF+++A L L+++A++ CA
Sbjct: 548 LNEGHLDRFFWMMAGLCFLNMLAFVFCA 575
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.324 0.139 0.426
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 968,981,647
Number of Sequences: 2790947
Number of extensions: 40727173
Number of successful extensions: 117719
Number of sequences better than 10.0: 626
Number of HSP's better than 10.0 without gapping: 442
Number of HSP's successfully gapped in prelim test: 184
Number of HSP's that attempted gapping in prelim test: 115184
Number of HSP's gapped (non-prelim): 968
length of query: 583
length of database: 848,049,833
effective HSP length: 133
effective length of query: 450
effective length of database: 476,853,882
effective search space: 214584246900
effective search space used: 214584246900
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.5 bits)
S2: 78 (34.7 bits)
Medicago: description of AC147404.4


