

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147364.8 + phase: 0
(954 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
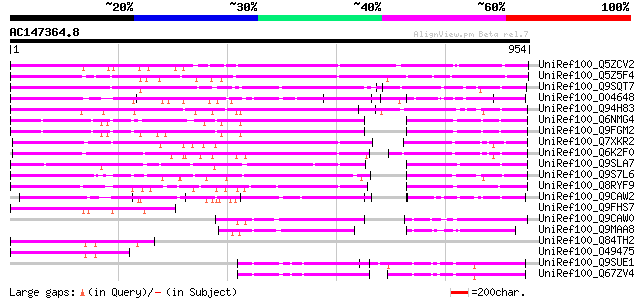
UniRef100_Q5ZCV2 DNAJ heat shock N-terminal domain-containing pr... 600 e-170
UniRef100_Q5Z5F4 Putative DNAJ heat shock N-terminal domain-cont... 583 e-165
UniRef100_Q9SQT7 Putative DnaJ protein [Arabidopsis thaliana] 477 e-133
UniRef100_O04648 A_TM021B04.9 protein [Arabidopsis thaliana] 386 e-105
UniRef100_Q94H83 Putative heat shock protein [Oryza sativa] 349 2e-94
UniRef100_Q6NMG4 At5g53150 [Arabidopsis thaliana] 323 2e-86
UniRef100_Q9FGM2 DnaJ protein-like [Arabidopsis thaliana] 320 1e-85
UniRef100_Q7XKR2 OSJNBa0053B21.9 protein [Oryza sativa] 319 3e-85
UniRef100_Q6K2F0 Heat shock protein-like [Oryza sativa] 308 5e-82
UniRef100_Q9SLA7 At2g25560/F13B15.22 [Arabidopsis thaliana] 304 1e-80
UniRef100_Q9S7L6 Expressed protein [Arabidopsis thaliana] 292 3e-77
UniRef100_Q8RYF9 Heat shock protein-like [Oryza sativa] 291 6e-77
UniRef100_Q9CAW2 Hypothetical protein T9J14.7 [Arabidopsis thali... 202 3e-50
UniRef100_Q9FHS7 Similarity to DnaJ protein [Arabidopsis thaliana] 182 4e-44
UniRef100_Q9CAW0 Hypothetical protein T9J14.9 [Arabidopsis thali... 177 2e-42
UniRef100_Q9MAA8 T12H1.7 protein [Arabidopsis thaliana] 176 2e-42
UniRef100_Q84TH2 Hypothetical protein At4g19570 [Arabidopsis tha... 157 1e-36
UniRef100_O49475 Hypothetical protein F24J7.130 [Arabidopsis tha... 153 3e-35
UniRef100_Q9SUE1 Hypothetical protein T13J8.90 [Arabidopsis thal... 152 4e-35
UniRef100_Q67ZV4 Hypothetical protein At4g27980 [Arabidopsis tha... 147 1e-33
>UniRef100_Q5ZCV2 DNAJ heat shock N-terminal domain-containing protein-like [Oryza
sativa]
Length = 1008
Score = 600 bits (1547), Expect = e-170
Identities = 377/1033 (36%), Positives = 537/1033 (51%), Gaps = 110/1033 (10%)
Query: 1 MDCNKEEALRAKEIAEKKMENRDFAGARKFALKAQRLYPVLENIAQMLVVCDVHCSAEQK 60
M+CNKEEAL+A++IA KKME++DF GA++ ALKAQR++P LENI+QML VC+VHC+AE K
Sbjct: 2 MECNKEEALKARDIAAKKMESKDFVGAKRIALKAQRIFPELENISQMLTVCEVHCAAEAK 61
Query: 61 VFGDEINWYGILQLERTAGDAMIKKQFRKFALQLHPDKNKFAGAEAAFKLIGEAQRVLSD 120
+ G +++YG+LQ++ A +A IKKQFRK A LHPDKN FAGAEAAFKL+ EAQ LSD
Sbjct: 62 MNG-LLDFYGVLQVDVMADEATIKKQFRKLAFSLHPDKNGFAGAEAAFKLVAEAQSTLSD 120
Query: 121 REKRTRYDMKLNV--------NKTAMPPRSNQPKVPTNFNSATKNNVRTNFTNSNTQQPP 172
R KR YD+K + + A P ++ QPK T ATK N T++ Q
Sbjct: 121 RTKRRAYDIKWRIASKQATQPKQGAQPAQAAQPKQCTQPPLATKRNQSAQPTHNTQQSAQ 180
Query: 173 QQQNKQP-----------PQQQNGVRRT-------------------FWTACPFCSVKYE 202
+Q+ QP P ++ R FWT C C KY+
Sbjct: 181 PKQSTQPMQATQPKHATEPMEKTDANRASNAKEGYGSSVRPPSAGEAFWTMCVNCKTKYQ 240
Query: 203 YYREILNKSLRCQQCHRLFVAYILDMQGTSPTTN-------------PSQQASKANVGSQ 249
YY +LN LRCQ C + F A +L+ Q + P Q+ S
Sbjct: 241 YYSNVLNHKLRCQNCKKDFRAVMLNEQDVPSVFSSSAAKSAGQHCDVPKQEDCSTKFSSA 300
Query: 250 GNSHA-----------EKSNTKPFKKKGPVGVSRKPDVKRKRNQVEEFSQSSDSTSSSDS 298
N A + N+ + G V+ +++ + SS + S +
Sbjct: 301 ANRDAKPMVNGGQHDEQMKNSASVRAGGEGTVNHTESIRKGGLEFSTLHVSSAANVGSKA 360
Query: 299 EDET-------VAGKNGFPGVGNHSTE-------QPRRSVRQKHNVSYSDNMNGTDNDLL 344
+ VAG+ N S E PRRS R+K N S +
Sbjct: 361 GGKMTSCPTPDVAGRQNPGNRVNTSAETGVMNIPNPRRSARRKENADASIIQD------- 413
Query: 345 RPSKRGQENGSHCGDGRSYRETAKTNDQNGLAADPKNEHEKVKQKQEEKIRAGGKEAAEG 404
PSK+ + + S R+ K D N + AD + V + + G + EG
Sbjct: 414 TPSKKRRTILDWFSNPDSSRK--KVADDNVVRADGQACEPHVSSEAHNHQK--GTTSNEG 469
Query: 405 SKQMDKTFEHSSPGSTSKTSNCPNAYVYPDAEFSDFDKDRKKECFAPGQIWAIYDSIDGM 464
+++ K H + + K S P + YPD EF DFD+ R FA QIWA+YD DGM
Sbjct: 470 NQEKRKDVAHDT--NAQKKSGIPGNFSYPDPEFFDFDRCRDVSMFAVDQIWALYDDRDGM 527
Query: 465 PRFYALIRKVLSPGFQLQATWLEPRPDDNDEIKWVDEELPVACGKFKLCNTEIIEDHLTF 524
PR+YA IR++ + F++Q TWLE + +E KW DEELPVACG F L T + +D L F
Sbjct: 528 PRYYARIRRIDTTNFRVQFTWLEHDAKNEEEDKWTDEELPVACGNFFLGKTVVSQDALMF 587
Query: 525 SHLVMF-KRNGRNTFQVYPRKGETWALFKNWDITWYKDEESHRQYEYEFVEILSDYVEGE 583
SH+V + K R+++++YPRKGE WAL+K W + W D + HR YEYE VEILS++
Sbjct: 588 SHIVSWVKGRKRSSYEIYPRKGEVWALYKGWSMQWSSDADKHRAYEYEAVEILSNFTVEA 647
Query: 584 GVHVAYLGKLKGFVSIFIQIMKEDNQPFQIPSAELFRFSHRIPSFKMTGQEGVDVHLGYL 643
G V L K+KGFVS+F ++ ++ F IP +E+ RFSH IP F+ G E V V G+L
Sbjct: 648 GAAVGPLVKIKGFVSLFAKV--KEKPSFVIPPSEMLRFSHSIPFFRTKGDEKVGVAGGFL 705
Query: 644 EFDPASLPMNLEEIAVTQNLDMRTGHSSCGSENARTSKRSKPSMSPEDIVSTPKVKVDTS 703
E D ASLP NL+ + LD + + E++++ K K +
Sbjct: 706 ELDTASLPSNLDVAFPSVTLDSCMPVCKTMNSGFNDFTGYEQGALKENLMNEGKRKDHSL 765
Query: 704 NLTDVKDSLDDMDDCHASASTPEAFEIPDAQFFNFETGRSLDKFQVGQIWAFYSDEDGMP 763
T V A+ S+P F+ P+++F NFE RS KF+ GQIWA YSD D P
Sbjct: 766 ERTPVHQQ-------SAAYSSPSTFDYPNSEFHNFEEYRSYSKFERGQIWALYSDLDQFP 818
Query: 764 KYYGQIKKVVTSPTIELHVYWLACCWLPENTTKWEDDGMLTSCGRFKVIKTKDFLSIYSN 823
KYYG + KV T P +H+ WL C E W + + SCG FK+ + + + +N
Sbjct: 819 KYYGWVTKVDTDP-FRVHLTWLEVCPQLEQENMWLEQNIPVSCGTFKIRNWR--IKLDTN 875
Query: 824 LSCISHQVQADPIG--KNYTIYPRKGEVWALYRKWSNKIKCSDLKNWDYDIVEVLEVADL 881
SH V+ +G + + I+P+ GE+WA+Y W+ S ++Y I E+ + +
Sbjct: 876 -DAFSHLVETSQVGWKRYFEIHPQVGEIWAIYNNWAPGWVPSSKDTFEYTIGEITDCTEA 934
Query: 882 FIETSILEHVTGFSSVFRGKSIEGSSGNLRIPKKELLRFSHQIPAFKLTEEH-GDLRGFW 940
+ +L V G+ +VF+ S+ G+ L IP E +RFSH IP+F+LT+E+ G L GF+
Sbjct: 935 STKVLLLTRVDGYRAVFKPDSVRGT---LEIPTNENIRFSHLIPSFRLTKENGGKLCGFY 991
Query: 941 ELDPGALPPSFLW 953
ELDP ++P +FL+
Sbjct: 992 ELDPASVPDTFLF 1004
>UniRef100_Q5Z5F4 Putative DNAJ heat shock N-terminal domain-containing protein [Oryza
sativa]
Length = 1018
Score = 583 bits (1503), Expect = e-165
Identities = 378/1036 (36%), Positives = 527/1036 (50%), Gaps = 105/1036 (10%)
Query: 1 MDCNKEEALRAKEIAEKKMENRDFAGARKFALKAQRLYPVLE-NIAQMLVVCDVHCSAEQ 59
MDCNK+EA++AK +AEKKM +DFAGA++ KAQ L ++ NI+QML VCD+HC++
Sbjct: 1 MDCNKDEAVKAKALAEKKMREKDFAGAKRMINKAQNLSKDVDSNISQMLTVCDIHCASAT 60
Query: 60 KVFGDEINWYGILQLERTAGDAMIKKQFRKFALQLHPDKNKFAGAEAAFKLIGEAQRVLS 119
KV G EI+WYGILQ+ TA D +IKKQ+RK AL LHPDKN FAGAEAAFKL+GEA L+
Sbjct: 61 KVNG-EIDWYGILQVPVTADDTLIKKQYRKLALLLHPDKNNFAGAEAAFKLVGEANMTLT 119
Query: 120 DREKRTRYDMKLNVNKTAMPPRSNQPKVPTNFNSATKNNVRTNFTNSNTQQPPQQQNKQP 179
DR KR+ YDMK N + R +VP + T VR T N Q Q +P
Sbjct: 120 DRSKRSVYDMKRNAS-----VRIGSARVPYQQSRRTA-PVRPTTTPVNLHNVHQSQQHKP 173
Query: 180 PQQQNGVRRTFWTACPFCSVKYEYYREILNKSLRCQQCHRLFVAYILDMQGTSPTTNPSQ 239
+ +TFWT CP C ++Y+YY IL K+LRCQ C + FVA L+ Q N
Sbjct: 174 SNPSDS--QTFWTICPTCGMRYQYYLSILKKALRCQNCLKPFVALDLNEQAVPSGANQRS 231
Query: 240 -------------QASKANVGSQG--------------NSHAEKSNTKPFKKKGPVGVSR 272
S+ANVG Q N+H E + G + R
Sbjct: 232 AGVWKSSGAPQNFPGSQANVGQQAQNSANPVHANFGSHNAHVETKRGADGNEAGGLKNKR 291
Query: 273 K--------------PDVKRKRNQVEEFSQSSDSTSSSDSEDETVAGKNGFPGVGNHSTE 318
K K++R + E S+SS S +S+DSE+E + VG +
Sbjct: 292 KFAKATGNSSKASSVAGSKKRRKAMFESSESSASDTSTDSEEEIIEDGPAASNVG--PDQ 349
Query: 319 QPRRSVRQKHNVSYSDNMNGTDND---------LLRPSKRGQENGS--HCGDGRSYRETA 367
PRRS RQK V Y+++ +G D D + PS + G H G+ + A
Sbjct: 350 HPRRSSRQKQEVKYNEDSDGDDTDCHGNGDDGFVSSPSLKRLRKGGLFHGGENNETKLNA 409
Query: 368 KT----NDQNGLAADPKNEHEKVK------QKQEEKIRAGGKEAAEGSKQMDKTFEHSSP 417
T +D + N E ++ ++ + + +GG +AE K S+
Sbjct: 410 DTTGPGHDGPTNGVNNYNNTEDIERGSACAEQIKRETMSGGGNSAEKEKLSHSV---SNN 466
Query: 418 GSTSKTSNCPNAYVYPDAEFSDFDKDRKKECFAPGQIWAIYDSIDGMPRFYALIRKVLS- 476
G S + + PN + D+EF DF++ R F QIWA YDS MPR+YA I KV
Sbjct: 467 GLESNSDDAPNEVICADSEFFDFNQLRHINQFKANQIWACYDSQSCMPRYYARITKVKHV 526
Query: 477 PGFQLQATWLEPRPDDNDEIKWVDEELPVACGKFKLCNTEIIEDHLTFSHLVMFKRN-GR 535
P F L WLE P + E W +LPV+CG+FK ++ ++ FSH + +++N R
Sbjct: 527 PKFMLNFIWLEFDPKNKAEAAWSSGDLPVSCGRFKHGVSDTAKESSMFSHAIFYEKNKTR 586
Query: 536 NTFQVYPRKGETWALFKNWDITWYKDEESHRQYEYEFVEILSDYVEGEGVHVAYLGKLKG 595
N++++YPRKGE WALFK WDI W D + H+ YEYE V++LSD + V L K+KG
Sbjct: 587 NSYEIYPRKGEVWALFKGWDIDWSADADKHKNYEYEVVQVLSDLTSSTSIIVMPLVKIKG 646
Query: 596 FVSIFIQIMKEDNQPFQIPSAELFRFSHRIPSFKMTGQEGVDVHLGYLEFDPASLPMN-- 653
FVS+FIQ ++ P+ IP + RFSH +P M G E + G +E DPA+LP+N
Sbjct: 647 FVSLFIQ--SKEASPYVIPQDDTLRFSHCVPRHTMIGTEKEGIPEGAIELDPAALPLNFG 704
Query: 654 LEEIAVTQNLDMRTGHSSCGSENARTSKRSKPSMSPEDI--------------VSTPKVK 699
+ +V G+E+ +S + D+ T K +
Sbjct: 705 VAFASVVPESCCSVKVQGSGAEHIGSSSGNNCHKGSVDVGESQHATCANTGFATRTTKAE 764
Query: 700 VDTSNLTDVKDSLDDMDDCHASASTPEAFEIPDAQFFNFETGRSLDKFQVGQIWAFYSDE 759
++ N + DD ++ A + P+++FF F RS+ KFQ GQIWA YSD
Sbjct: 765 INEHNARSAVEGTDDDEEPDDFAQAEVLY--PESEFFEFSEIRSIHKFQPGQIWALYSDV 822
Query: 760 DGMPKYYGQIKKVVTSPTIELHVYWLACCWLPENTTKWEDDGMLTSCGRFKVIKTKDFLS 819
D P YY IK V EL V WL C E + + + +CG FK I + +
Sbjct: 823 DKFPNYYACIKTVDVKNN-ELQVRWLDACPQSEEERRLVREDLTVACGTFK-ISSFHGIQ 880
Query: 820 IYSNLSCISHQVQADPIGKN-YTIYPRKGEVWALYRKWSNKIKCSDLKNWDYDIVEVLEV 878
Y+ +SH VQA P +N Y I P +G++WA+++ W D K DY++VE+
Sbjct: 881 TYNGTEYLSHPVQAKPGRRNEYEIVPCQGDIWAVFKNWRTGWTAKDYKKCDYELVEIFGH 940
Query: 879 ADLFIETSILEHVTGFSSVFRGKSIEGSSGNLRIPKKELLRFSHQIPAFKLTEEH-GDLR 937
D I+ +L V G+ +VF EG+ +R K E +FSHQIP F LT E G LR
Sbjct: 941 TDSSIQVQLLRKVDGYRAVFMPDRREGAVKTIR--KDEYPKFSHQIPCFHLTNERGGKLR 998
Query: 938 GFWELDPGALPPSFLW 953
GF ELDP ++P FL+
Sbjct: 999 GFLELDPLSVPEMFLF 1014
>UniRef100_Q9SQT7 Putative DnaJ protein [Arabidopsis thaliana]
Length = 673
Score = 477 bits (1228), Expect = e-133
Identities = 296/702 (42%), Positives = 391/702 (55%), Gaps = 52/702 (7%)
Query: 1 MDCNKEEALRAKEIAEKKMENRDFAGARKFALKAQRLYPVLENIAQMLVVCDVHCSAEQK 60
M N++EALRAK++AE M+ DF ARK A+KAQ++ LENI++M++VCDVHC+A +K
Sbjct: 1 MSINRDEALRAKDLAEGLMKKTDFTAARKLAMKAQKMDSSLENISRMIMVCDVHCAATEK 60
Query: 61 VFGDEINWYGILQLERTAGDAMIKKQFRKFALQLHPDKNKFAGAEAAFKLIGEAQRVLSD 120
+FG E++WYGILQ+E+ A D +IKKQ+++ AL LHPDKNK GAE+AFKLIGEAQR+L D
Sbjct: 61 LFGTEMDWYGILQVEQIANDVIIKKQYKRLALLLHPDKNKLPGAESAFKLIGEAQRILLD 120
Query: 121 REKRTRYDMKLNV-NKTAMPPRSNQ--PKVPTNFNSATKNNVRTNFT--NSNTQQPPQQQ 175
REKRT +D K K A PP Q P T + N R FT + P Q+
Sbjct: 121 REKRTLHDNKRKTWRKPAAPPYKAQQMPNYHTQPHFRASVNTRNIFTELRPEIRHPFQKA 180
Query: 176 NKQPPQQQNGVRRTFWTACPFCSVKYEYYREILNKSLRCQQCHRLFVAYILDMQGTSPTT 235
QP + +TF T+C FC V+YEY R +NK + C+ C + F A+ +Q
Sbjct: 181 QAQPAAFTH--LKTFGTSCVFCRVRYEYDRAHVNKEVTCETCKKRFTAFEEPLQSAPQAK 238
Query: 236 NPSQ------QASK---ANVGSQGNSHAEKSNTKPFKKKG-PV-GVSRKPDVKRKRNQVE 284
PSQ Q SK S+ + E T K P+ G + K + KRKR V
Sbjct: 239 GPSQTTYCFPQQSKFPDQRACSEPHKRPENPPTVSSSKASFPMPGSTAKHNGKRKRKNVA 298
Query: 285 EFSQSSDSTSSSDSEDETVAGKNGFPGVGNHSTEQPRRSVRQKHNVSYSDNMNGTDNDLL 344
E S+SSDS SSS+SED+ G++ EQPRRSVR K VSY++N++ D DL+
Sbjct: 299 ECSESSDSESSSESEDDVNNDTTAAQDSGSNGGEQPRRSVRSKQKVSYNENLSDDDVDLV 358
Query: 345 RPSKRGQENGSHCGDGRSYRETAKTNDQNGLAADPKNEHEKVKQKQEEKIRAGGKEAAEG 404
+ G+ +G + D +ET + N +N KI E G
Sbjct: 359 --NDNGEGSGKNI-DTEREKETEEEKQTN------ENHSSTESIDMNGKIEVDQVETPSG 409
Query: 405 SKQMDKTFEHSSPGSTSKTSNCPNAYVYPDAEFSDFDKDRKKECFAPGQIWAIYDSIDGM 464
+ + E S GS K PN Y D +F+DFDK R+K CF GQIWA+YD +GM
Sbjct: 410 ASDSE---EDLSSGSAEK----PNLINYDDPDFNDFDKLREKSCFQAGQIWAVYDEEEGM 462
Query: 465 PRFYALIRKVLSPGFQLQATWLEPRPDDNDEIKWVDEELPVACGKFKLCNTEIIEDHLTF 524
PRFYALI+KV +P F L+ W E D +E LPV+ GKF + N E F
Sbjct: 463 PRFYALIKKVTTPDFMLRYVWFEVDQDQENE----TPNLPVSVGKFVVGNIEETNLCSIF 518
Query: 525 SHLVMFKRNGR-NTFQVYPRKGETWALFKNWDITWYKDEESHRQYEYEFVEILSDYVEGE 583
SH V R F V+P+KGE WALFKNWDI D S +YEYEFVEILSD+ EG
Sbjct: 519 SHFVYSTTKIRTRKFTVFPKKGEIWALFKNWDINCSADSVSPMKYEYEFVEILSDHAEGA 578
Query: 584 GVHVAYLGKLKGFVSIFIQIMKEDNQPFQIPSAELFRFSHRIPSFKMTGQEGVDVHLGYL 643
V V +L K++GF +F + K+++ +IP E RFSH IPSF++TG EG + G+
Sbjct: 579 TVSVGFLSKVQGFNCVFCPMPKDESNTCEIPPHEFCRFSHSIPSFRLTGTEGRGITKGWY 638
Query: 644 EFDPASLPMNLEEIAVTQNLDMRTGHSSCGSENARTSKRSKP 685
E DPA+LP +V+QNL G E A+ R P
Sbjct: 639 ELDPAALP-----ASVSQNLS--------GEEAAQDRDRQSP 667
Score = 136 bits (343), Expect = 3e-30
Identities = 95/287 (33%), Positives = 140/287 (48%), Gaps = 25/287 (8%)
Query: 675 ENARTSKRSKPSMSPEDIVSTPKVKVD-TSNLTDVKDSLDDMDDCHASASTPEAFEIPDA 733
E + ++ S E I K++VD + DS +D+ SA P D
Sbjct: 376 ETEEEKQTNENHSSTESIDMNGKIEVDQVETPSGASDSEEDLSS--GSAEKPNLINYDDP 433
Query: 734 QFFNFETGRSLDKFQVGQIWAFYSDEDGMPKYYGQIKKVVTSPTIELHVYWLACCWLPEN 793
F +F+ R FQ GQIWA Y +E+GMP++Y IKK VT+P L W EN
Sbjct: 434 DFNDFDKLREKSCFQAGQIWAVYDEEEGMPRFYALIKK-VTTPDFMLRYVWFEVDQDQEN 492
Query: 794 TTKWEDDGMLTSCGRFKV--IKTKDFLSIYSNLSCISHQVQADPIGKNYTIYPRKGEVWA 851
E + S G+F V I+ + SI+S+ + +++ + +T++P+KGE+WA
Sbjct: 493 ----ETPNLPVSVGKFVVGNIEETNLCSIFSHFVYSTTKIRT----RKFTVFPKKGEIWA 544
Query: 852 LYRKWSNKIKCS----DLKNWDYDIVEVL--EVADLFIETSILEHVTGFSSVFRGKSIEG 905
L++ W I CS ++Y+ VE+L + L V GF+ VF +
Sbjct: 545 LFKNWD--INCSADSVSPMKYEYEFVEILSDHAEGATVSVGFLSKVQGFNCVFCPMP-KD 601
Query: 906 SSGNLRIPKKELLRFSHQIPAFKL--TEEHGDLRGFWELDPGALPPS 950
S IP E RFSH IP+F+L TE G +G++ELDP ALP S
Sbjct: 602 ESNTCEIPPHEFCRFSHSIPSFRLTGTEGRGITKGWYELDPAALPAS 648
>UniRef100_O04648 A_TM021B04.9 protein [Arabidopsis thaliana]
Length = 1609
Score = 386 bits (992), Expect = e-105
Identities = 273/770 (35%), Positives = 377/770 (48%), Gaps = 140/770 (18%)
Query: 1 MDCNKEEALRAKEIAEKKMENRDFAGARKFALKAQRLYPVLENIAQMLVVCDVHCSAEQK 60
MD NKEEA RAK +AE KM+ DF GA+K LKAQ L+ LE++ QML VCDVH SAE+K
Sbjct: 1 MDWNKEEACRAKTLAEDKMKEGDFVGAQKLLLKAQSLFSGLESLPQMLAVCDVHNSAEKK 60
Query: 61 VFGDEINWYGILQLERTAGDAMIKKQFRKFALQLHPDKNKFAGAEAAFKLIGEAQRVLSD 120
+ E NWYGILQ+ A DA IKKQ RK AL LHPDKN+F GAEAAFKL+ +A R L+D
Sbjct: 61 INCLE-NWYGILQVMHFADDATIKKQVRKLALLLHPDKNQFPGAEAAFKLVWDASRFLAD 119
Query: 121 REKRTRYDMKLNVNKTAMPPRSNQPKVPTNFNSATKNNVRTNFTNSNTQQPPQQQNKQPP 180
++KR++YD++ + ++ TN +A + ++ TNS T
Sbjct: 120 KDKRSQYDIRRRIYL----------RLATNQLNAN-SGLQCAATNSATD----------- 157
Query: 181 QQQNGVRRTFWTACPFCSVKYEYYREILNKSLRCQQCHRLFVAYIL-------------- 226
TFWT C C +Y+Y R+ +N L C C R ++AY
Sbjct: 158 --------TFWTCCEHCGYRYKYLRKYVNILLNCNICQRSYMAYDTGFNEAPSKSNTGQK 209
Query: 227 DMQGTSPTTNPSQQASKANVGSQGNSHA---EKSNT--KPFKKKGPVGVSRKPDVKRK-- 279
++Q P N S + ++G+Q S A +K T K F K+ G S+K +VK
Sbjct: 210 EVQNQGPC-NTSLNTNGESIGAQPGSVAAEVDKKGTFNKKFNKRNGGGESKKTEVKNSKI 268
Query: 280 -----RNQVEEFSQSS-----------------------DSTSSSDSEDETVAGKNGFPG 311
RN EE +S+ +S S SD T KN
Sbjct: 269 ETEFVRNDDEEKMKSAAKLHKPHPEVTEPEIGASKSVPDESVSRSDEAPSTSKDKNKMKK 328
Query: 312 VGNHSTE----------------QPRRSVRQKHNVSYSDNMNGTDNDLLRPSKRGQENGS 355
S R+S R+ SY++ +DN L P KR + N
Sbjct: 329 GVEESINVDGSDFIKDKAYSKDNNKRKSPRRSQQSSYAEEEKISDNSLGPPKKRLRSNVG 388
Query: 356 HCGDGRSYRE-----TAKTNDQNGLAADPK---------------NEHEKVKQKQEEKIR 395
+ + + ++K D G ++ P E VK K+ E +
Sbjct: 389 LKSEQTTRKGLVGVGSSKRFDSGGGSSAPSCAFKRKAKKFVDSGYQEISSVKDKEREVRK 448
Query: 396 AGGKEAAEGSKQMDKTFEHSSPGSTSKTSNCPNAYVYPDAEFSDFDKDRKKECFAPGQIW 455
A G EG K + +P T + P D EFS+F+ CF Q+W
Sbjct: 449 ASG----EGVVMEAKIDNNHNPNENLITEDLP------DPEFSNFELTTS--CFGVNQVW 496
Query: 456 AIYDSIDGMPRFYALIRKVLSPGFQLQATWLEPRPDDNDEIKWVDEELPVACGKFKLCNT 515
++YD IDGMPR YA I KVL P F+L TW++P D+ D +P+ACG F+ +
Sbjct: 497 SMYDPIDGMPRLYARIDKVLVPEFKLWITWIDPLQDNKDN------SIPIACGIFQGGGS 550
Query: 516 EIIEDHLTFSHLVMFKRNGRNTFQVYPRKGETWALFKNWDITWYKDEESHRQ-YEYEFVE 574
E DHL FS MF N+ +YPRKGE WA+F+ WDI+W E+H+ YEY+FVE
Sbjct: 551 EEENDHLKFS-CQMFHLTRNNSVVIYPRKGEIWAIFRGWDISWSASSENHKHPYEYDFVE 609
Query: 575 ILSDYVEGEGVHVAYLGKLKGFVSIFIQIMKEDNQPFQIPSAELFRFSHRIPSFKMTGQE 634
+LS++ + G+ V +LGK++GFVS+F Q ++ QIP +++ RFSH++PSFKMTG+E
Sbjct: 610 VLSNFNDENGLGVGFLGKVEGFVSLFRQDAQDGVLQLQIPPSQMLRFSHKVPSFKMTGKE 669
Query: 635 GVDVHLGYLEFDPASLPMNLEEI---AVTQNLDMRTGHSSCGSENARTSK 681
V G E DPA+LP L E+ V LD + G SK
Sbjct: 670 REGVPPGCFELDPAALPKELFEVYDSKVDVGLDRELPNGKTGGPFPEASK 719
Score = 113 bits (283), Expect = 2e-23
Identities = 79/227 (34%), Positives = 123/227 (53%), Gaps = 21/227 (9%)
Query: 729 EIPDAQFFNFETGRSLDKFQVGQIWAFYSDEDGMPKYYGQIKKVVTSPTIELHVYWLACC 788
++PD +F NFE S F V Q+W+ Y DGMP+ Y +I KV+ P +L + W+
Sbjct: 474 DLPDPEFSNFELTTSC--FGVNQVWSMYDPIDGMPRLYARIDKVLV-PEFKLWITWIDP- 529
Query: 789 WLPENTTKWEDDGMLTSCGRFKVIKTKDFLSIYSNLSCISHQVQADPIGKNYTIYPRKGE 848
L +N +D+ + +CG F+ +++ + + SC + + + IYPRKGE
Sbjct: 530 -LQDN----KDNSIPIACGIFQGGGSEE-ENDHLKFSCQMFHLTRN---NSVVIYPRKGE 580
Query: 849 VWALYRKWSNKIKCSDLKN---WDYDIVEVLE--VADLFIETSILEHVTGFSSVFRGKSI 903
+WA++R W S + ++YD VEVL + + L V GF S+FR +
Sbjct: 581 IWAIFRGWDISWSASSENHKHPYEYDFVEVLSNFNDENGLGVGFLGKVEGFVSLFRQDAQ 640
Query: 904 EGSSGNLRIPKKELLRFSHQIPAFKLT--EEHGDLRGFWELDPGALP 948
+G L+IP ++LRFSH++P+FK+T E G G +ELDP ALP
Sbjct: 641 DGVL-QLQIPPSQMLRFSHKVPSFKMTGKEREGVPPGCFELDPAALP 686
Score = 98.2 bits (243), Expect = 1e-18
Identities = 75/234 (32%), Positives = 108/234 (46%), Gaps = 23/234 (9%)
Query: 666 RTGHSSCGSENARTSKRSKPSMSPEDIVS-TPKVKVDTSNLTDVKDSLDD---------M 715
+ G S E ++ +P+DI T VK N + S D M
Sbjct: 814 KPGGLSLSGEKMSEKTITRMRKTPQDIHKPTVNVKKHGRNDESLSQSRGDGLLTQLNGNM 873
Query: 716 DDCHASASTPEAFEIPDAQFFNFETGRSLDKFQVGQIWAFYSDEDGMPKYYGQIKKVVTS 775
P + + FNFE RS DKFQ+ QIWA YS++ G P+ Y QIKK+ TS
Sbjct: 874 KSSEPETRVPSSCKTVKENTFNFENQRSWDKFQIDQIWAIYSNDKGSPRKYAQIKKIDTS 933
Query: 776 PTIELHVYWLACCWLPENTTKWEDDGMLTSCGRFKVIKTKDFLSIYSNLSCISHQVQADP 835
P +LHV L P + + CGRFK+ K + + S+ SHQV+A
Sbjct: 934 PEFKLHVAPLELYRPPIHMPR------PVCCGRFKLKTGKAEVYVPSS---FSHQVKAVK 984
Query: 836 IGKN-YTIYPRKGEVWALYRKWSNKIKCSDLKNWDYDIVEVLEVADLFIETSIL 888
KN + +YP KGE+WALY+ + + + + +IVEV+E + I+ L
Sbjct: 985 TKKNRFEVYPGKGEIWALYKNCNTR---DYTETEELEIVEVVETDEQRIQAMTL 1035
Score = 89.4 bits (220), Expect = 5e-16
Identities = 95/353 (26%), Positives = 146/353 (40%), Gaps = 49/353 (13%)
Query: 234 TTNPSQQASKANVGSQGNSHAEKSNTKPFKKKGPVGVSRKPDVKRKRNQVEEFSQSSDST 293
T P +ASK + Q N+ E S PFKKK P + + FS+ SD
Sbjct: 710 TGGPFPEASKVEM--QANTTRESS---PFKKKRP----------KVHDNHGSFSKESDGR 754
Query: 294 SSSDSEDETVAGKNGFPGVGNHSTEQPRRSVRQKHNVSYSDNMNGTDNDLLRPSKRGQEN 353
++ + + V + + R + + + S +++ +G K+ N
Sbjct: 755 RTNHERNSAKESRKSVKAVDALNLRKSPRLLSETN--SQANSRSGRQI----AEKKSANN 808
Query: 354 GSHCGDGRSYRETAKTNDQNGLAADPKNEHEKVKQKQEEKIRAGGKEAAEGSK------Q 407
G CG + + + + K + K K E+ S+ Q
Sbjct: 809 GESCGKPGGLSLSGEKMSEKTITRMRKTPQDIHKPTVNVKKHGRNDESLSQSRGDGLLTQ 868
Query: 408 MDKTFEHSSPGSTSKTSNCPNAYVYPDAEFSDFDKDRKKECFAPGQIWAIYDSIDGMPRF 467
++ + S P T S+C +F+ R + F QIWAIY + G PR
Sbjct: 869 LNGNMKSSEP-ETRVPSSCKTV----KENTFNFENQRSWDKFQIDQIWAIYSNDKGSPRK 923
Query: 468 YALIRKV-LSPGFQLQATWLEP-RPDDNDEIKWVDEELPVACGKFKLCNTEIIEDHL--T 523
YA I+K+ SP F+L LE RP + PV CG+FKL T E ++ +
Sbjct: 924 YAQIKKIDTSPEFKLHVAPLELYRPP-------IHMPRPVCCGRFKL-KTGKAEVYVPSS 975
Query: 524 FSHLVMFKRNGRNTFQVYPRKGETWALFKNWDITWYKDEESHRQYEYEFVEIL 576
FSH V + +N F+VYP KGE WAL+KN + Y + E E E VE++
Sbjct: 976 FSHQVKAVKTKKNRFEVYPGKGEIWALYKNCNTRDYTETE-----ELEIVEVV 1023
>UniRef100_Q94H83 Putative heat shock protein [Oryza sativa]
Length = 748
Score = 349 bits (896), Expect = 2e-94
Identities = 233/754 (30%), Positives = 362/754 (47%), Gaps = 88/754 (11%)
Query: 1 MDCNKEEALRAKEIAEKKMENRDFAGARKFALKAQRLYPVLENIAQMLVVCDVHCSAEQK 60
M+CNK+EALRAKEIAE+K E++D GA+KFALKAQ L+P LE I QM+ D++ ++E
Sbjct: 1 MECNKDEALRAKEIAERKFESKDLQGAKKFALKAQALFPGLEGIVQMITTLDLYLASEVL 60
Query: 61 VFGDEINWYGILQLERTAGDAMIKKQFRKFALQLHPDKNKFAGAEAAFKLIGEAQRVLSD 120
+ G++ +WY IL +E +A D +KKQ+RK LQLHPDKNK GAE AFK++ EA VLSD
Sbjct: 61 ISGEK-DWYSILSVESSADDETLKKQYRKLVLQLHPDKNKSVGAEGAFKMVQEAWTVLSD 119
Query: 121 REKRTRYDMK-----LNVNKTAMPPRSNQPKVPTNFNSATKNNVRTNFTNSNTQQ----- 170
+ KR YD K L N + S P F + N + T N Q+
Sbjct: 120 KTKRALYDQKRKLMVLKRNTSQTNKASAAPGASNGFYNFAANAAASKVTRGNKQKAGPAT 179
Query: 171 --------PPQQQNKQPPQQQNGVRRTFWTACPFCSVKYEYYREILNKSLRCQQCHRLFV 222
PP +Q P TFWT+C C + YEY + LN +L C C F+
Sbjct: 180 SSVRQRPPPPPPPPRQAPAPPPAKPPTFWTSCNKCKMNYEYLKVYLNHNLLCPTCREPFL 239
Query: 223 AYILDM--------------QGTSPTTNPSQQASKANVGSQGNSHAEKSNTKPFKKKGPV 268
A + M G + TN S+ + + +++ + V
Sbjct: 240 AQEVPMPPTESVHAVHDPNISGANQNTNGSRNFQWGPFSRTAGAASATASSAAAAQAANV 299
Query: 269 GVSRKPDVKRKRNQVEEFSQSSDSTSSSDSEDETVA--GKNGFPGVGNHSTEQPRRSVRQ 326
V+R+R + + ++ ++ + + A +N G G +S+++ R++
Sbjct: 300 VHHTYEKVRREREEAQAAARREEALRRKYNPPKRQANISENLNLGTGGNSSKK-MRTMGN 358
Query: 327 KHNVSYSDNMNGT--------DNDLLRPSKRGQENGSHCGDGRSYRETAKTN-------- 370
+ S ++G+ ++ + G + G S++ T
Sbjct: 359 DIGIGSSSILSGSGANYFGVPGGNISFSTNSGAHHFQGVNGGFSWKPRPPTRISLVKTFT 418
Query: 371 --DQNGLAADPKNEHEKVKQKQEEKIRAGGKEAAEGSKQMDKTFEHS--------SPGST 420
D G+ + K K K+ + R+ + AA G K F+ S S ST
Sbjct: 419 QFDVRGILMEKAKSDLKDKLKEMQTKRS--QVAANGKKNKKNMFKESGGDDESLASDDST 476
Query: 421 SK--------------------TSNCPNAYVYPDAEFSDFDKDRKKECFAPGQIWAIYDS 460
++ ++ P +Y PD +F DFDKDR +ECF QIWA YD
Sbjct: 477 ARQAAHVDPEDNASVNSTDADDENDDPLSYNVPDPDFHDFDKDRTEECFQSDQIWATYDD 536
Query: 461 IDGMPRFYALIRKVLS-PGFQLQATWLEPRPDDN-DEIKWVDEELPVACGKFKLCNTEII 518
DGMPR+YA I+KVLS FQL+ ++L R + + WV CG F++C E
Sbjct: 537 EDGMPRYYAFIQKVLSLEPFQLKISFLTSRTNSEFGSLNWVSSGFTKTCGDFRICRYETC 596
Query: 519 EDHLTFSHLVMFKRNGRNTFQVYPRKGETWALFKNWDITWYKDEESHRQYEYEFVEILSD 578
+ FSH + +++ R ++YP+KG WA+++NW W +D + Y+ VE+L +
Sbjct: 597 DILNMFSHQIKWEKGPRGVIKIYPQKGNIWAVYRNWSPDWDEDTPDKVLHAYDVVEVLDE 656
Query: 579 YVEGEGVHVAYLGKLKGFVSIFIQIMKEDNQPFQIPSAELFRFSHRIPSFKMTGQEGVDV 638
Y E G+ V L K+ GF ++F Q ++ N +IP E+FRFSH +P ++M+G+E +V
Sbjct: 657 YDEDLGISVIPLVKVAGFRTVF-QRNQDLNAIKKIPKEEMFRFSHEVPFYRMSGEEAPNV 715
Query: 639 HLGYLEFDPASLPMNLEEIAVTQNLDMRTGHSSC 672
E DPA++ L + +T+ ++ S C
Sbjct: 716 PKDSYELDPAAISKELLQ-EITETVESSKATSEC 748
Score = 135 bits (339), Expect = 8e-30
Identities = 101/340 (29%), Positives = 163/340 (47%), Gaps = 53/340 (15%)
Query: 642 YLEFDPASLPMNLEEIAVTQNL-DMRTGHSSCGSENARTSKR-------SKPSMSPEDIV 693
+ +FD + M + + L +M+T S + + K S++ +D
Sbjct: 417 FTQFDVRGILMEKAKSDLKDKLKEMQTKRSQVAANGKKNKKNMFKESGGDDESLASDDST 476
Query: 694 STPKVKVDTSNLTDVK--DSLDDMDDCHASASTPEAFEIPDAQFFNFETGRSLDKFQVGQ 751
+ VD + V D+ D+ DD P ++ +PD F +F+ R+ + FQ Q
Sbjct: 477 ARQAAHVDPEDNASVNSTDADDENDD-------PLSYNVPDPDFHDFDKDRTEECFQSDQ 529
Query: 752 IWAFYSDEDGMPKYYGQIKKVVTSPTIELHVYWLACCWLPE-NTTKWEDDGMLTSCGRFK 810
IWA Y DEDGMP+YY I+KV++ +L + +L E + W G +CG F+
Sbjct: 530 IWATYDDEDGMPRYYAFIQKVLSLEPFQLKISFLTSRTNSEFGSLNWVSSGFTKTCGDFR 589
Query: 811 VIK--TKDFLSIYSNLSCISHQV--QADPIGKNYTIYPRKGEVWALYRKWSNKIKCSDLK 866
+ + T D L+++ SHQ+ + P G IYP+KG +WA+YR WS
Sbjct: 590 ICRYETCDILNMF------SHQIKWEKGPRGV-IKIYPQKGNIWAVYRNWS--------P 634
Query: 867 NWD----------YDIVEVLEV--ADLFIETSILEHVTGFSSVFRGKSIEGSSGNLRIPK 914
+WD YD+VEVL+ DL I L V GF +VF+ + + +IPK
Sbjct: 635 DWDEDTPDKVLHAYDVVEVLDEYDEDLGISVIPLVKVAGFRTVFQRN--QDLNAIKKIPK 692
Query: 915 KELLRFSHQIPAFKLTEEHGD--LRGFWELDPGALPPSFL 952
+E+ RFSH++P ++++ E + +ELDP A+ L
Sbjct: 693 EEMFRFSHEVPFYRMSGEEAPNVPKDSYELDPAAISKELL 732
>UniRef100_Q6NMG4 At5g53150 [Arabidopsis thaliana]
Length = 726
Score = 323 bits (828), Expect = 2e-86
Identities = 229/700 (32%), Positives = 335/700 (47%), Gaps = 78/700 (11%)
Query: 1 MDCNKEEALRAKEIAEKKMENRDFAGARKFALKAQRLYPVLENIAQMLVVCDVHCSAEQK 60
M+CNK+EA RA +IAE+KM +D+ GA+KFA KAQ L+P L+ + Q+ V +V+ S E K
Sbjct: 1 MECNKDEAKRAMDIAERKMTEKDYTGAKKFANKAQNLFPELDGLKQLFVAINVYISGE-K 59
Query: 61 VFGDEINWYGILQLERTAGDAMIKKQFRKFALQLHPDKNKFAGAEAAFKLIGEAQRVLSD 120
F E +WYG+L ++ A D +KKQ+RK L LHPDKNK GAE AF L+ EA +LSD
Sbjct: 60 TFAGEADWYGVLGVDPFASDEALKKQYRKLVLMLHPDKNKCKGAEGAFNLVAEAWALLSD 119
Query: 121 REKRTRYDMKLNVNKTAMPPR--SNQPKVPTNFNSATKN---NVRTNFTNS-------NT 168
++KR Y++K + A R + Q ++P+ + T N NVR + +S T
Sbjct: 120 KDKRILYNVKRGKDVKAAQQRFPTTQREIPS--HQPTSNGIPNVREHVVSSARARYKPAT 177
Query: 169 QQPPQQQNKQ---------PPQQQNGVRRTFWTACPFCSVKYEYYREILNKSLRCQQCHR 219
++P + ++ P Q+ + TFWT C C +YEY R LN++L C CH
Sbjct: 178 RKPAARMDRSRTGSPAFVYPTQESS----TFWTMCNKCDTQYEYQRVYLNQTLLCPHCHH 233
Query: 220 LFVAYILDMQGTSPTTNPSQQASKANVGSQGNSHAEKS-NTKPFKKKGPVGVSRKPDVKR 278
FVA + T PT P K V N H S N K R+P
Sbjct: 234 GFVA----EEKTPPTNIP-----KPPVNISSNQHHRSSKNQASNKNSNGSSYRREPATSV 284
Query: 279 KRNQVEEFSQSSDSTSSSDSEDETVAGKNGFPGVGNHSTEQPRRSVRQKHNVSYSDNMNG 338
N + S+ S S + + + + G + E R + G
Sbjct: 285 NHNFQWDSSRMGGSYSRNATNETANVVQQGQDKLKRVFWETQEREAAR-----------G 333
Query: 339 TDNDLLRPSKRGQENGSH-----CGDGRSYRETAKTNDQNGLAADPKNEHEKVKQ----- 388
N L KR + + SH G Y + +D D + + E K+
Sbjct: 334 FTNSDLGNFKRQKTDDSHMRGPSAGSRHPYVQALLRSDIKKALMD-RGQSEIFKRLPMMI 392
Query: 389 -KQEEKIR--AGGKEAAEGSKQMDKTFEHSSPGSTSK-----------TSNCPNAYVYPD 434
K E K+ G K + + E S ST+ S+ V PD
Sbjct: 393 AKMEGKVNPTEGEKNSTKAMSSKASEVERSKMSSTANEVERSVEVIPHESDEVKEIVVPD 452
Query: 435 AEFSDFDKDRKKECFAPGQIWAIYDSIDGMPRFYALIRKVLSPG-FQLQATWLEPR-PDD 492
++F +FD DR + F QIWA YD DGMPRFYA I+KV+S F+L+ +WL + +
Sbjct: 453 SDFHNFDLDRSESAFKDDQIWAAYDDADGMPRFYARIQKVISVNPFKLKISWLNSKTTSE 512
Query: 493 NDEIKWVDEELPVACGKFKLCNTEIIEDHLTFSHLVMFKRNGRNTFQVYPRKGETWALFK 552
I W+ +CG F+ E + FSH V F + R + P+KG+ WAL++
Sbjct: 513 FGPIDWMGAGFAKSCGDFRCGRYESTDTLNAFSHSVDFTKGARGLLHILPKKGQVWALYR 572
Query: 553 NWDITWYKDEESHRQYEYEFVEILSDYVE-GEGVHVAYLGKLKGFVSIFIQIMKEDNQPF 611
NW W K+ +++YE VE+L DY E + + VA L K +GF +F + E
Sbjct: 573 NWSPEWDKNTPDEVKHKYEMVEVLDDYTEDDQSLTVALLLKAEGFRVVF-RRCTEKLGVR 631
Query: 612 QIPSAELFRFSHRIPSFKMTGQEGVDVHLGYLEFDPASLP 651
+I E+ RFSH++P + +TG+E + G+LE DPA+ P
Sbjct: 632 KIAKEEMLRFSHQVPHYILTGKEADNAPEGFLELDPAATP 671
Score = 127 bits (320), Expect = 1e-27
Identities = 85/232 (36%), Positives = 121/232 (51%), Gaps = 17/232 (7%)
Query: 730 IPDAQFFNFETGRSLDKFQVGQIWAFYSDEDGMPKYYGQIKKVVTSPTIELHVYWLACCW 789
+PD+ F NF+ RS F+ QIWA Y D DGMP++Y +I+KV++ +L + WL
Sbjct: 450 VPDSDFHNFDLDRSESAFKDDQIWAAYDDADGMPRFYARIQKVISVNPFKLKISWLNSKT 509
Query: 790 LPE-NTTKWEDDGMLTSCGRFKVIKTKDFLSIYSNLSCISHQVQADPIGKNYT-IYPRKG 847
E W G SCG F+ + + L+ SH V + I P+KG
Sbjct: 510 TSEFGPIDWMGAGFAKSCGDFRCGRYES----TDTLNAFSHSVDFTKGARGLLHILPKKG 565
Query: 848 EVWALYRKWS---NKIKCSDLKNWDYDIVEVLE---VADLFIETSILEHVTGFSSVFRGK 901
+VWALYR WS +K ++K+ Y++VEVL+ D + ++L GF VFR
Sbjct: 566 QVWALYRNWSPEWDKNTPDEVKH-KYEMVEVLDDYTEDDQSLTVALLLKAEGFRVVFR-- 622
Query: 902 SIEGSSGNLRIPKKELLRFSHQIPAFKLTEEHGD--LRGFWELDPGALPPSF 951
G +I K+E+LRFSHQ+P + LT + D GF ELDP A P +F
Sbjct: 623 RCTEKLGVRKIAKEEMLRFSHQVPHYILTGKEADNAPEGFLELDPAATPCAF 674
>UniRef100_Q9FGM2 DnaJ protein-like [Arabidopsis thaliana]
Length = 755
Score = 320 bits (820), Expect = 1e-85
Identities = 225/713 (31%), Positives = 341/713 (47%), Gaps = 75/713 (10%)
Query: 1 MDCNKEEALRAKEIAEKKMENRDFAGARKFALKAQRLYPVLENIAQMLVVCDVHCSAEQK 60
M+CNK+EA RA +IAE+KM +D+ GA+KFA KAQ L+P L+ + Q+ V +V+ S E K
Sbjct: 1 MECNKDEAKRAMDIAERKMTEKDYTGAKKFANKAQNLFPELDGLKQLFVAINVYISGE-K 59
Query: 61 VFGDEINWYGILQLERTAGDAMIKKQFRKFALQLHPDKNKFAGAEAAFKLIGEAQRVLSD 120
F E +WYG+L ++ A D +KKQ+RK L LHPDKNK GAE AF L+ EA +LSD
Sbjct: 60 TFAGEADWYGVLGVDPFASDEALKKQYRKLVLMLHPDKNKCKGAEGAFNLVAEAWALLSD 119
Query: 121 REKRTRYDMKLNVNKTAMPPR--SNQPKVPTNFNSATKN---NVRTNFTNS-------NT 168
++KR Y++K + A R + Q ++P+ + T N NVR + +S T
Sbjct: 120 KDKRILYNVKRGKDVKAAQQRFPTTQREIPS--HQPTSNGIPNVREHVVSSARARYKPAT 177
Query: 169 QQPPQQQNKQ---------PPQQQNGVRRTFWTACPFCSVKYEYYREILNKSLRCQQCHR 219
++P + ++ P Q+ + TFWT C C +YEY R LN++L C CH
Sbjct: 178 RKPAARMDRSRTGSPAFVYPTQESS----TFWTMCNKCDTQYEYQRVYLNQTLLCPHCHH 233
Query: 220 LFVAYILDMQGTSPTTNPSQ----QASKANVGSQGNSHAEKSNTKPFKKKGPVGV----- 270
FVA + T PT P +++ + S+ + + SN ++++ V
Sbjct: 234 GFVA----EEKTPPTNIPKPPVNISSNQHHRSSKNQASNKNSNGSSYRREPATSVNHNFQ 289
Query: 271 --SRKPDVKRKRNQVEE----FSQSSDSTSSSDSEDETVAGKNGFPG--VGNHSTEQPRR 322
S + RN E Q D E + GF +GN ++
Sbjct: 290 WDSSRMGGSYSRNATNETANVVQQGQDKLKRVFWETQEREAARGFTNSDLGNFKRQKTDD 349
Query: 323 SVRQKHNVSYSDNMNGTDNDL--LRPSKRGQENGSHCGDGRSYRETAKTNDQNGLAADPK 380
S + + N G + E+ + G Y + +D D +
Sbjct: 350 SHMRGPSAGKLSNQIGASRSSTPFYAANENTESSNTIGSRHPYVQALLRSDIKKALMD-R 408
Query: 381 NEHEKVKQ------KQEEKIR--AGGKEAAEGSKQMDKTFEHSSPGSTSK---------- 422
+ E K+ K E K+ G K + + E S ST+
Sbjct: 409 GQSEIFKRLPMMIAKMEGKVNPTEGEKNSTKAMSSKASEVERSKMSSTANEVERSVEVIP 468
Query: 423 -TSNCPNAYVYPDAEFSDFDKDRKKECFAPGQIWAIYDSIDGMPRFYALIRKVLSPG-FQ 480
S+ V PD++F +FD DR + F QIWA YD DGMPRFYA I+KV+S F+
Sbjct: 469 HESDEVKEIVVPDSDFHNFDLDRSESAFKDDQIWAAYDDADGMPRFYARIQKVISVNPFK 528
Query: 481 LQATWLEPR-PDDNDEIKWVDEELPVACGKFKLCNTEIIEDHLTFSHLVMFKRNGRNTFQ 539
L+ +WL + + I W+ +CG F+ E + FSH V F + R
Sbjct: 529 LKISWLNSKTTSEFGPIDWMGAGFAKSCGDFRCGRYESTDTLNAFSHSVDFTKGARGLLH 588
Query: 540 VYPRKGETWALFKNWDITWYKDEESHRQYEYEFVEILSDYVE-GEGVHVAYLGKLKGFVS 598
+ P+KG+ WAL++NW W K+ +++YE VE+L DY E + + VA L K +GF
Sbjct: 589 ILPKKGQVWALYRNWSPEWDKNTPDEVKHKYEMVEVLDDYTEDDQSLTVALLLKAEGFRV 648
Query: 599 IFIQIMKEDNQPFQIPSAELFRFSHRIPSFKMTGQEGVDVHLGYLEFDPASLP 651
+F + E +I E+ RFSH++P + +TG+E + G+LE DPA+ P
Sbjct: 649 VF-RRCTEKLGVRKIAKEEMLRFSHQVPHYILTGKEADNAPEGFLELDPAATP 700
Score = 127 bits (320), Expect = 1e-27
Identities = 85/232 (36%), Positives = 121/232 (51%), Gaps = 17/232 (7%)
Query: 730 IPDAQFFNFETGRSLDKFQVGQIWAFYSDEDGMPKYYGQIKKVVTSPTIELHVYWLACCW 789
+PD+ F NF+ RS F+ QIWA Y D DGMP++Y +I+KV++ +L + WL
Sbjct: 479 VPDSDFHNFDLDRSESAFKDDQIWAAYDDADGMPRFYARIQKVISVNPFKLKISWLNSKT 538
Query: 790 LPE-NTTKWEDDGMLTSCGRFKVIKTKDFLSIYSNLSCISHQVQADPIGKNYT-IYPRKG 847
E W G SCG F+ + + L+ SH V + I P+KG
Sbjct: 539 TSEFGPIDWMGAGFAKSCGDFRCGRYES----TDTLNAFSHSVDFTKGARGLLHILPKKG 594
Query: 848 EVWALYRKWS---NKIKCSDLKNWDYDIVEVLE---VADLFIETSILEHVTGFSSVFRGK 901
+VWALYR WS +K ++K+ Y++VEVL+ D + ++L GF VFR
Sbjct: 595 QVWALYRNWSPEWDKNTPDEVKH-KYEMVEVLDDYTEDDQSLTVALLLKAEGFRVVFR-- 651
Query: 902 SIEGSSGNLRIPKKELLRFSHQIPAFKLTEEHGD--LRGFWELDPGALPPSF 951
G +I K+E+LRFSHQ+P + LT + D GF ELDP A P +F
Sbjct: 652 RCTEKLGVRKIAKEEMLRFSHQVPHYILTGKEADNAPEGFLELDPAATPCAF 703
>UniRef100_Q7XKR2 OSJNBa0053B21.9 protein [Oryza sativa]
Length = 729
Score = 319 bits (817), Expect = 3e-85
Identities = 230/732 (31%), Positives = 339/732 (45%), Gaps = 78/732 (10%)
Query: 6 EEALRAKEIAEKKMENRDFAGARKFALKAQRLYPVLENIAQMLVVCDVHCSAEQKVFGDE 65
+EA++A+ +AE + +RD GARK+A+KAQ L P LE I+QM+ +VH +AE K+ G E
Sbjct: 2 DEAIKARGVAESRFHSRDIRGARKYAIKAQNLCPSLEGISQMVSTLEVHLAAESKIDG-E 60
Query: 66 INWYGILQLERTAGDAMIKKQFRKFALQLHPDKNKFAGAEAAFKLIGEAQRVLSDREKRT 125
+WY IL L A + +KKQ+RK ALQLHPDKNK GAE AFKLI EA VLSD K+
Sbjct: 61 SDWYRILSLTAFADEEEVKKQYRKLALQLHPDKNKSVGAEEAFKLISEAWSVLSDNSKKV 120
Query: 126 RYDMKLNVNKTAMPPRSNQPKVPTNFNSATKNNVRTNFTNSNTQQPPQQQNKQPPQQQNG 185
YD K + N ++ R N + + + G
Sbjct: 121 LYDQKRKDHSVV-----NVTNGMYTYDKKANKRARKNAAAAAAAAAAAAAAAEATTRPAG 175
Query: 186 VRRTFWTACPFCSVKYEYYREILNKSLRCQQCHRLFVAYILDM--QGTSPTTNPSQQASK 243
V TFWT+C C ++YEY R LN +L C CH F+A G+S + + S +
Sbjct: 176 VD-TFWTSCNRCRMQYEYLRIYLNHNLLCPNCHHAFLAVETGFPCNGSSSSFSWSTKQQP 234
Query: 244 ANVGSQGNSHAEKSNTKPFKKKGPVGVSRKPDV--------------------------- 276
N S +S+ S T G G +
Sbjct: 235 QNNNSTKHSYGSTSRTSSIPGTGHGGYQQDGTYDSYNNQSFQWNQYSKTTPAAGTNAYGT 294
Query: 277 ----KRKRNQVEEFSQSSDSTSSSDSEDETVAGKNGFPGVGNHSTE---------QPRRS 323
K KR E +S + +T +S + T + + F HS + R +
Sbjct: 295 QALEKPKRKHEESYSYNYSATGNSYGHERTNSRRGRFSKRRRHSNDGYTTMDFGGDNRET 354
Query: 324 VRQK-HNVSYSD----NMNGTDNDLLRPSKRGQENG-----SHCGDGRSYRETAKTNDQN 373
V +++D +NGT + LR + G+ S E AK Q
Sbjct: 355 VAASTETTAFTDVAVAQVNGTSGEKLRSAVSGRRANVLREISQIDTRALLIEKAKAAIQE 414
Query: 374 GLA----------ADPKNEHEKVKQKQEEKIRAGG--KEAAEGSKQMD-KTFEHSSPGST 420
L A+ KV + GG + +G KQ ++ + +P
Sbjct: 415 KLQEWNITSSSRLAERGKSQGKVYPSDNNIKQNGGLSDKHVKGLKQCSSRSVDTQAPTVD 474
Query: 421 SKTSN---CPNAYVYPDAEFSDFDKDRKKECFAPGQIWAIYDSIDGMPRFYALIRKVLSP 477
K P + PD +F DFDKDR + F Q+WA YDS DGMPR YA+++KVLS
Sbjct: 475 EKNPEQRRVPVSIDVPDPDFHDFDKDRTERAFDSDQVWATYDSEDGMPRLYAMVQKVLSM 534
Query: 478 G-FQLQATWLEPRPDDN-DEIKWVDEELPVACGKFKLCNTEIIEDHLTFSHLVMFKRNGR 535
F+++ ++L + + I WV CG F++ +I E FSH V + + R
Sbjct: 535 RPFRIRMSFLNSKSNSELAPISWVASGFQKTCGDFRVGRYQISETVNIFSHKVSWTKGPR 594
Query: 536 NTFQVYPRKGETWALFKNWDITWYKDEESHRQYEYEFVEILSDYVEGEGVHVAYLGKLKG 595
++ P+KG+TWAL++NW W + Y+YE VEI+ D+ + +G+ V L K+ G
Sbjct: 595 GIIRIVPQKGDTWALYRNWSPDWNELTPDDVIYKYEIVEIIDDFTDEQGLTVIPLLKVAG 654
Query: 596 FVSIFIQIMKEDNQPFQIPSAELFRFSHRIPSFKMTGQEGVDVHLGYLEFDPASLPMNLE 655
F ++F + M + + +IP ELFRFSHR+PS +TG+EG + G E DPA+ P++L
Sbjct: 655 FKAVFHRHM-DPKEARRIPKEELFRFSHRVPSRLLTGEEGNNAPKGCHELDPAATPVDLL 713
Query: 656 EIAVTQNLDMRT 667
++ D T
Sbjct: 714 KVITEVTEDTAT 725
Score = 112 bits (281), Expect = 4e-23
Identities = 81/240 (33%), Positives = 123/240 (50%), Gaps = 22/240 (9%)
Query: 725 PEAFEIPDAQFFNFETGRSLDKFQVGQIWAFYSDEDGMPKYYGQIKKVVTSPTIELHVYW 784
P + ++PD F +F+ R+ F Q+WA Y EDGMP+ Y ++KV++ + + +
Sbjct: 484 PVSIDVPDPDFHDFDKDRTERAFDSDQVWATYDSEDGMPRLYAMVQKVLSMRPFRIRMSF 543
Query: 785 LACCWLPE-NTTKWEDDGMLTSCGRFKVIKTKDFLSIYSNLSCISHQVQ--ADPIGKNYT 841
L E W G +CG F+V + I ++ SH+V P G
Sbjct: 544 LNSKSNSELAPISWVASGFQKTCGDFRVGR----YQISETVNIFSHKVSWTKGPRG-IIR 598
Query: 842 IYPRKGEVWALYRKWS---NKIKCSDLKNWDYDIVEVLEVADLFIETSI----LEHVTGF 894
I P+KG+ WALYR WS N++ D+ + Y+IVE+++ D E + L V GF
Sbjct: 599 IVPQKGDTWALYRNWSPDWNELTPDDV-IYKYEIVEIID--DFTDEQGLTVIPLLKVAGF 655
Query: 895 SSVFRGKSIEGSSGNLRIPKKELLRFSHQIPAFKLTEEHGD--LRGFWELDPGALPPSFL 952
+VF + ++ RIPK+EL RFSH++P+ LT E G+ +G ELDP A P L
Sbjct: 656 KAVFH-RHMDPKEAR-RIPKEELFRFSHRVPSRLLTGEEGNNAPKGCHELDPAATPVDLL 713
>UniRef100_Q6K2F0 Heat shock protein-like [Oryza sativa]
Length = 734
Score = 308 bits (789), Expect = 5e-82
Identities = 227/731 (31%), Positives = 351/731 (47%), Gaps = 86/731 (11%)
Query: 6 EEALRAKEIAEKKMENRDFAGARKFALKAQRLYPVLENIAQMLVVCDVHCSAEQKVFGDE 65
+EAL+A++ AE+K RD GAR+ A+KAQ L P L+ I+QM+ +V ++E KV G+
Sbjct: 12 DEALKARDAAERKFHARDVKGARRSAIKAQNLCPSLDGISQMVSTLEVLLASESKVDGEN 71
Query: 66 INWYGILQLERTAGDAMIKKQFRKFALQLHPDKNKFAGAEAAFKLIGEAQRVLSDREKRT 125
+WY IL L +A + +KKQ+RK ALQLHPDKNK GAE AFKLI EA VLSD+ ++
Sbjct: 72 -DWYRILSLSASADEEEVKKQYRKLALQLHPDKNKSVGAEGAFKLISEAWAVLSDKSRKM 130
Query: 126 RYDMKLNVNKTAMPPRSNQPKVPTNFNSATKNNVRTNFTNSNTQQPPQQQNKQPPQQQNG 185
+YD K + P +N ++ R N S +
Sbjct: 131 QYDQKRKDH-----PVTNGANGLYTYDKKAHKRARKNAAASAAAAAAAAAAAAEATTRPV 185
Query: 186 VRRTFWTACPFCSVKYEYYREILNKSLRCQQCHRLFVAYILDM--QGTSPTTNPSQQASK 243
TFWT+C C ++YEY R LN +L C CH F+A GTS + + S + +
Sbjct: 186 GLDTFWTSCNRCRMQYEYLRIYLNHNLLCPNCHHAFMAVETGYPCNGTSSSFSWSTKQQQ 245
Query: 244 ANVGSQGNSHAEKSNTKPFKKKGPVGVSRKPDVKRKRNQVEEFSQSSDSTSS------SD 297
N +S++ S T G ++ + NQ +++Q S + +S S
Sbjct: 246 QN---HKHSYSSASRTSGVPGTGHGVYQQENTYETYNNQSFQWNQYSKTNNSAGTNAYSS 302
Query: 298 SEDETVAGKNGFPGVGNHST-------EQPR----RSVRQKHNVSY---SDNMNGTDNDL 343
+ E K+ + N+S+ E+P R +++ N++ S + NG + +
Sbjct: 303 TASEKPKRKHEESYIYNYSSSGNEFGQERPTSGRGRFSKRRQNINNGYASVDCNGDNKET 362
Query: 344 LRPSKR-------GQENGSHCGDGRSYRETAKTN--------DQNGLAADPKNEHEKVKQ 388
+ + G+ NG+ RS + N D GL + + K
Sbjct: 363 VAATAGTTVLADVGRVNGTSVEKFRSAVSGRRANVMREIFQLDTRGLLIEKAKAAIREKL 422
Query: 389 KQEEKIRAGGKEAAEGSKQMDKTFEHS---------SPGSTSKTSNCPNAYV-------- 431
Q+ I A AA+G + +H +P K N A V
Sbjct: 423 -QDLNISATRHIAAKGKAERKNHVDHDVKGNGILPHNPSHKFKICNSKGADVENPATDEN 481
Query: 432 ------------YPDAEFSDFDKDRKKECFAPGQIWAIYDSIDGMPRFYALIRKVLS-PG 478
PD +F DFDKDR + F Q+WA YDS DGMPR YA+++KV+S
Sbjct: 482 NLEQKRVPVSIDVPDPDFYDFDKDRTERTFDNDQVWATYDSEDGMPRLYAMVQKVISRKP 541
Query: 479 FQLQATWLEPRPD-DNDEIKWVDEELPVACGKFKLCNTEIIEDHLTFSHLVMFKRNGRNT 537
F+++ ++L + + + I WV CG F++ +I E FSH V + + R
Sbjct: 542 FRIRMSFLNSKSNIELSPINWVASGFSKTCGDFRVGRYQIFETVNIFSHRVSWSKGPRGI 601
Query: 538 FQVYPRKGETWALFKNWDITWYKDEESHRQYEYEFVEILSDYVEGEGVHVAYLGKLKGFV 597
++ P+KG+TWAL++NW W + Y+YE VE++ D+ + +GV V L K+ GF
Sbjct: 602 IKIVPKKGDTWALYRNWSSDWNELTPDDVIYKYEIVEVIDDFTDEQGVTVIPLLKVAGFK 661
Query: 598 SIFIQIMKEDNQPFQIPSAELFRFSHRIPSFKMTGQEGVDVHLGYLEFDPASLPMNL--- 654
++F + + + +IP ELFRFSHR+PS +TG+EG + G E DPA+ P++L
Sbjct: 662 AVFHR-RTDSDVVRRIPKEELFRFSHRVPSRLLTGEEGNNAPKGCHELDPAATPVDLLKV 720
Query: 655 ----EEIAVTQ 661
+E+A T+
Sbjct: 721 ITEVKEVATTE 731
Score = 124 bits (311), Expect = 1e-26
Identities = 88/268 (32%), Positives = 138/268 (50%), Gaps = 23/268 (8%)
Query: 697 KVKVDTSNLTDVKDSLDDMDDCHASASTPEAFEIPDAQFFNFETGRSLDKFQVGQIWAFY 756
K K+ S DV++ D ++ P + ++PD F++F+ R+ F Q+WA Y
Sbjct: 462 KFKICNSKGADVENPATDENNLEQKR-VPVSIDVPDPDFYDFDKDRTERTFDNDQVWATY 520
Query: 757 SDEDGMPKYYGQIKKVVTSPTIELHVYWL-ACCWLPENTTKWEDDGMLTSCGRFKVIKTK 815
EDGMP+ Y ++KV++ + + +L + + + W G +CG F+V +
Sbjct: 521 DSEDGMPRLYAMVQKVISRKPFRIRMSFLNSKSNIELSPINWVASGFSKTCGDFRVGR-- 578
Query: 816 DFLSIYSNLSCISHQV--QADPIGKNYTIYPRKGEVWALYRKWS---NKIKCSDLKNWDY 870
I+ ++ SH+V P G I P+KG+ WALYR WS N++ D+ + Y
Sbjct: 579 --YQIFETVNIFSHRVSWSKGPRG-IIKIVPKKGDTWALYRNWSSDWNELTPDDV-IYKY 634
Query: 871 DIVEVLEVADLFIETSI----LEHVTGFSSVFRGKSIEGSSGNLRIPKKELLRFSHQIPA 926
+IVEV++ D E + L V GF +VF ++ S RIPK+EL RFSH++P+
Sbjct: 635 EIVEVID--DFTDEQGVTVIPLLKVAGFKAVFHRRT--DSDVVRRIPKEELFRFSHRVPS 690
Query: 927 FKLTEEHGD--LRGFWELDPGALPPSFL 952
LT E G+ +G ELDP A P L
Sbjct: 691 RLLTGEEGNNAPKGCHELDPAATPVDLL 718
>UniRef100_Q9SLA7 At2g25560/F13B15.22 [Arabidopsis thaliana]
Length = 656
Score = 304 bits (778), Expect = 1e-80
Identities = 214/671 (31%), Positives = 315/671 (46%), Gaps = 40/671 (5%)
Query: 1 MDCNKEEALRAKEIAEKKMENRDFAGARKFALKAQRLYPVLENIAQMLVVCDVHCSAEQK 60
M+ NKEEA RA+EIA++K DFAGARKFALKAQ LYP L+ IAQM+ DVH SA+
Sbjct: 1 MEFNKEEATRAREIAKRKFLANDFAGARKFALKAQFLYPELDGIAQMVATFDVHLSAQNI 60
Query: 61 VFGDEINWYGILQLERTAGDAMIKKQFRKFALQLHPDKNKFAGAEAAFKLIGEAQRVLSD 120
++GD ++ YG+L L A D +++K++RK A+ LHPD+NK GAE AFK + +A V SD
Sbjct: 61 IYGD-VDHYGVLGLNPEADDEIVRKRYRKLAVMLHPDRNKSVGAEEAFKFLSQAWGVFSD 119
Query: 121 REKRTRYDMKLNVNKTAMPPRSNQPKVPTNFNSATKNNVRTNFTNSN------TQQPPQQ 174
+ KR YD+K NV S+ F TK + T S+
Sbjct: 120 KAKRADYDLKRNVGLYKGGGASSSRPATNGFQKVTKASGNTTKVKSSKRGIKRASDASAA 179
Query: 175 QNKQPPQQQNGVRRTFWTACPFCSVKYEYYREILNKSLRCQQCHRLFVAYILDMQGTSPT 234
Q+ TFWT C C +YEY+ LN++L C C + F+A D G+
Sbjct: 180 ATTSTSAQKTTADGTFWTVCRTCRTQYEYHSVYLNQNLLCPNCRKPFIAVETDPPGSGSI 239
Query: 235 TNPSQQASKANVGSQGNSHAEKSNTKPFKKKGPVGVSRKPDVKRKRNQVEEFSQSSDSTS 294
+ ++ H K + GV + D + S+ +T
Sbjct: 240 RKTFHEHQFDSL-----RHTTDGRKKNVPGRDNNGVYGEYD-SFEWGVFTGTKTSAHATP 293
Query: 295 SSDSEDETVAGKNGFPGVGNHSTEQPRRSVRQKHNVSYSDNMNGTDNDLLRPSKRGQENG 354
+ +DE V + G ST P+R ++ V+ G L P G +
Sbjct: 294 TGSRKDEVVRREYTKRTAGPSSTIPPKRRKVMENAVA-----GGNIASCLAPKSTGVKEV 348
Query: 355 SHCGDGRSYRETAKTNDQNGL------AADPKNEHEKVKQKQEEKIRAG---GKEAAEGS 405
S ++ AK+ L A+ + ++ + AG K A E S
Sbjct: 349 SEDELKNLLKKKAKSVISRNLPELCTIVAETETNGRGMETEDLNGFNAGSSVNKNAIE-S 407
Query: 406 KQMDKTFEHSSPGSTSKTSNCPNAYVYPDAEFSDFDKDRKKECFAPGQIWAIYDSIDGMP 465
M+ E +S G+ + P +F DFDKDR ++ QIWA YDS +G+P
Sbjct: 408 CCMNTDKELNSLGALTLDVTAP--------DFCDFDKDRTEKSVKDNQIWAFYDSHEGLP 459
Query: 466 RFYALIRKVLS-PGFQLQATWLEPRPD-DNDEIKWVDEELPVACGKFKLCNTEIIEDHLT 523
R YALI V+S F+++ +WL P + + W+ +P +CG F++ T I +
Sbjct: 460 RSYALIHNVISVDPFKVRMSWLTPVTNGEPSSTNWLGFGIPKSCGGFRVRKTLIYRSPYS 519
Query: 524 FSHLVMFKRNGRNTFQVYPRKGETWALFKNWDITWYKDEESHRQYEYEFVEILSDYVEGE 583
FSH V + F +YPR G+ WAL++ W W EY+ VE++ Y E
Sbjct: 520 FSHKVNLVKGNHGEFLIYPRTGDVWALYRKWSPDW-NYLTGVETVEYDIVEVVEGYTEEY 578
Query: 584 GVHVAYLGKLKGFVSIFIQIMKEDNQPFQIPSAELFRFSHRIPSFKMTGQEGVDVHLGYL 643
GV V L K+ GF ++F + + + + E+ RFSH+IPS+ +TGQE G
Sbjct: 579 GVVVVPLVKVAGFKAVFHHHL-DSKETKRFLRDEISRFSHKIPSYLLTGQEAPGAPRGCR 637
Query: 644 EFDPASLPMNL 654
+ DPA+ P L
Sbjct: 638 QLDPAATPSQL 648
Score = 115 bits (289), Expect = 5e-24
Identities = 81/231 (35%), Positives = 117/231 (50%), Gaps = 13/231 (5%)
Query: 729 EIPDAQFFNFETGRSLDKFQVGQIWAFYSDEDGMPKYYGQIKKVVTSPTIELHVYWLACC 788
++ F +F+ R+ + QIWAFY +G+P+ Y I V++ ++ + WL
Sbjct: 425 DVTAPDFCDFDKDRTEKSVKDNQIWAFYDSHEGLPRSYALIHNVISVDPFKVRMSWLTPV 484
Query: 789 WLPE-NTTKWEDDGMLTSCGRFKVIKTKDFLSIYSNLSCISHQVQ-ADPIGKNYTIYPRK 846
E ++T W G+ SCG F+V KT + S YS SH+V + IYPR
Sbjct: 485 TNGEPSSTNWLGFGIPKSCGGFRVRKTLIYRSPYS----FSHKVNLVKGNHGEFLIYPRT 540
Query: 847 GEVWALYRKWSNKIK-CSDLKNWDYDIVEVLE--VADLFIETSILEHVTGFSSVFRGKSI 903
G+VWALYRKWS + ++ +YDIVEV+E + + L V GF +VF
Sbjct: 541 GDVWALYRKWSPDWNYLTGVETVEYDIVEVVEGYTEEYGVVVVPLVKVAGFKAVFHHHL- 599
Query: 904 EGSSGNLRIPKKELLRFSHQIPAFKLT--EEHGDLRGFWELDPGALPPSFL 952
S R + E+ RFSH+IP++ LT E G RG +LDP A P L
Sbjct: 600 -DSKETKRFLRDEISRFSHKIPSYLLTGQEAPGAPRGCRQLDPAATPSQLL 649
>UniRef100_Q9S7L6 Expressed protein [Arabidopsis thaliana]
Length = 706
Score = 292 bits (748), Expect = 3e-77
Identities = 222/742 (29%), Positives = 353/742 (46%), Gaps = 116/742 (15%)
Query: 1 MDCNKEEALRAKEIAEKKMENRDFAGARKFALKAQRLYPVLENIAQMLVVCDVHCSAEQK 60
M+ +EEALR K+IAE++ +DF AR +ALKA+ L+P LE ++QM+ +V+ +++ +
Sbjct: 1 MEAYREEALRVKQIAERRFAEKDFTSARSYALKAKSLFPDLEGLSQMVATFEVYLASQTR 60
Query: 61 VFGDEINWYGILQLERTAGDAMIKKQFRKFALQLHPDKNKFAGAEAAFKLIGEAQRVLSD 120
G +I++Y +L L+ +AG +KKQ++K A+ LHPDKNK GA+ AF LI EA LS+
Sbjct: 61 S-GGQIDYYAVLGLKPSAGKREVKKQYKKMAVLLHPDKNKCIGADGAFHLISEAWSFLSN 119
Query: 121 REKRTR--YDMKLNVNKTAMPPRSNQPKVPTNFNSATKNNVRTNFTNSNTQQPPQQQNKQ 178
++ Y K +++ T + S + T + T V F PP +
Sbjct: 120 EFNKSTFYYKRKKHIDSTEVQKHSTEYMPGTGTGTGTA--VFDRF-------PPSSERLD 170
Query: 179 PPQQQNGVRRTFWTACPFCSVKYEYYREILNKSLRCQQCHRLFVAYILDMQGTSPTTNPS 238
TFWT C C V+YEY R+ +NK L C+ C F+A G +P + P
Sbjct: 171 ----------TFWTVCTSCKVQYEYLRKYVNKRLSCKNCRGAFIAV---ETGPAPVSAPF 217
Query: 239 QQASKANVGSQGNSHAEKSNTKPFKKKGPVG---VSRKPDVKR---------------KR 280
++ SHA S+ P G G +SR P
Sbjct: 218 HYTPPSHAPP---SHAPPSHAPPSNGYGAHGYDAMSRMPTNSTYFLGHYPGQGHGYDYST 274
Query: 281 NQVEEFSQSSDSTSSSDSED--ETVAGKNGFP----------GV--------GNHSTEQP 320
N E+S S +T+S + D + NG+P G+ G S +
Sbjct: 275 NGSYEWSSYSGTTTSPGNLDLKRVSSVSNGYPYKHSNSVVSGGINKVKDGSNGTCSKKST 334
Query: 321 RRSVRQKHNVSYSDNMNGTDNDLLRPSKRGQENGSHCGDG-------------------- 360
+ + +S S + + N + RP K+ + +G
Sbjct: 335 PGGLIYPNQISMSAHASA--NKVGRPGKKSKVFMEAAANGFVENPLRSVSVSKTANTDAK 392
Query: 361 --RSYRETAKTNDQNGLAADPKNEHEKVKQKQEEKIRAG---------GKEAAEGSKQMD 409
+ Y+ +++ + AA + + + QK I+ AAE + +D
Sbjct: 393 MDQDYKLHIQSSTRRWSAASVLDTRKPLIQKARTDIKQRLEMMRLALEAAAAAEDATPLD 452
Query: 410 -KTFEHSSPGS-TSKTSNCPNAYVYPDAEFSDFDKDRKKECFAPGQIWAIYDSIDGMPRF 467
KT G T + +N P PD++F DFDK+R +E F P QIWAIYD DGMPR
Sbjct: 453 EKTVLSCKLGDVTGRKTNGP--ITVPDSDFHDFDKNRSEESFEPRQIWAIYDEDDGMPRL 510
Query: 468 YALIRKVLS-PGFQLQATWLEPRPD-DNDEIKWVDEELPVACGKFKLCNTEIIEDHLTFS 525
Y ++R+VLS F++ +L + D + +KWV +CG F++ N++I++ FS
Sbjct: 511 YCVVREVLSVQPFKIDIAYLSSKTDIEFGSMKWVQYGFTKSCGHFRIRNSDIVDHVNIFS 570
Query: 526 HLVMFKRNGR-NTFQVYPRKGETWALFKNWDITWYKDEESHRQYEYEFVEILSDYVEGEG 584
HL+ K+ GR +++P GE WA++KNW + W +++YE VEIL +Y E G
Sbjct: 571 HLLKGKKTGRGGCVRIFPTAGEIWAVYKNWSLNWDGSTPDEVRHQYEMVEILDEYTEQYG 630
Query: 585 VHVAYLGKLKGFVSIFIQIMKEDNQPFQIPSAELFRFSHRIPSFKMTGQEGVDVHLGYLE 644
V V L KL+G+ +++ + +ED++ + IP E+ RFSH++PS+ + D G+ E
Sbjct: 631 VCVTPLVKLEGYKTVYHRSTREDSKKW-IPRCEMLRFSHQVPSWFLK-----DATSGFPE 684
Query: 645 ----FDPASLPMNLEEIAVTQN 662
DPA++P L I N
Sbjct: 685 NCWDLDPAAIPEELLHIGAGTN 706
Score = 115 bits (288), Expect = 6e-24
Identities = 70/239 (29%), Positives = 123/239 (51%), Gaps = 30/239 (12%)
Query: 730 IPDAQFFNFETGRSLDKFQVGQIWAFYSDEDGMPKYYGQIKKVVTSPTIELHVYWLAC-C 788
+PD+ F +F+ RS + F+ QIWA Y ++DGMP+ Y +++V++ ++ + +L+
Sbjct: 475 VPDSDFHDFDKNRSEESFEPRQIWAIYDEDDGMPRLYCVVREVLSVQPFKIDIAYLSSKT 534
Query: 789 WLPENTTKWEDDGMLTSCGRFKVIKTKDFLSIYSNLSCISHQVQADPIGKN--YTIYPRK 846
+ + KW G SCG F++ + I +++ SH ++ G+ I+P
Sbjct: 535 DIEFGSMKWVQYGFTKSCGHFRIRNS----DIVDHVNIFSHLLKGKKTGRGGCVRIFPTA 590
Query: 847 GEVWALYRKWSNKIKCSDLKNWD----------YDIVEVLE--VADLFIETSILEHVTGF 894
GE+WA+Y+ WS NWD Y++VE+L+ + + L + G+
Sbjct: 591 GEIWAVYKNWS--------LNWDGSTPDEVRHQYEMVEILDEYTEQYGVCVTPLVKLEGY 642
Query: 895 SSVFRGKSIEGSSGNLRIPKKELLRFSHQIPAFKLTE-EHGDLRGFWELDPGALPPSFL 952
+V+ + E S IP+ E+LRFSHQ+P++ L + G W+LDP A+P L
Sbjct: 643 KTVYHRSTREDS--KKWIPRCEMLRFSHQVPSWFLKDATSGFPENCWDLDPAAIPEELL 699
>UniRef100_Q8RYF9 Heat shock protein-like [Oryza sativa]
Length = 744
Score = 291 bits (745), Expect = 6e-77
Identities = 220/768 (28%), Positives = 349/768 (44%), Gaps = 142/768 (18%)
Query: 1 MDCNKEEALRAKEIAEKKMENRDFAGARKFALKAQRLYPVLENIAQMLVVCDVHCSAEQK 60
M+CN+++A+R+KEIAE+K D AGA++FALKA+ L+ LE I M+ D+H A+ K
Sbjct: 1 MECNRDDAIRSKEIAERKFNENDIAGAKRFALKAKTLFDSLEGIDNMISALDIHIRAQTK 60
Query: 61 VFGDEINWYGILQLERTAGDAMIKKQFRKFALQLHPDKNKFAGAEAAFKLIGEAQRVLSD 120
+ G+ + YGIL + + D IKKQ+RK ALQ HPDKNKF+GAE+AFKLI +A VLSD
Sbjct: 61 IEGEN-DLYGILDISASDDDEKIKKQYRKLALQTHPDKNKFSGAESAFKLIQDAWDVLSD 119
Query: 121 REKRTRYDMKLNVNKTAMPPRSNQPKVPTNFNS---ATKNNVRTNFTNSNTQQPPQQQNK 177
++K+ YD K + R Q N N+ +T +++ F ++ + P +
Sbjct: 120 KDKKRSYDQK----RFGGSSRVYQNGFAENANATPGSTMSSMNGFFWQNSGRHPSYATD- 174
Query: 178 QPPQQQNGVRRTFWTACPFCSVKYEYYREILNKSLRCQQCHRLFVAY------------I 225
TFWT C C + ++Y RE +N++L C C FVA +
Sbjct: 175 -----------TFWTYCDSCQMSFQYSREYVNRNLACSFCQTEFVAVETPPPTAPVYYNV 223
Query: 226 LDMQGTSPTTNPSQQAS-------------KANVGSQGNSHAEK----SNTKPFKKK--G 266
++ TS + Q + GS G +HA + KP +K+
Sbjct: 224 TNLMDTSSNMDDPQGTGVPYSSNKIFDPVLQPVFGSVGGAHASRYPVQQTCKPARKEEVA 283
Query: 267 PVGVSRKPDVKRKRNQVEEFSQSSDSTSSSDSEDETVAGKNGFPGVGNHSTEQPRRSVRQ 326
V V+R+ + +++++ Q+S S SS S + + + V + RR +
Sbjct: 284 EVNVARREEATKRKHE-----QASSSLGSSSSAAKVIHRRK---AVTKEMEAEKRRCINN 335
Query: 327 KHNVSYSDN-------------MNGTDNDLLRPSKRGQENGSHCGDGRSYRETAKTNDQN 373
K VS N +G + P+KR ++ S G + R ++ +
Sbjct: 336 KSKVSGQKNNTNKVVGKSTSSAADGDSGPQMHPAKR--KSASSIGTSGTKRRKMPSDHNS 393
Query: 374 GLAAD-------------PKNEHEKVKQKQEEKI--------RAGGKEAAEGSKQMDKTF 412
G A P + EK+K + +K+ K S + KT+
Sbjct: 394 GNARTSFGKVFLQLETEIPGLKMEKMKLQIRDKLEEFKSRRANVENKGNVHVSLEKKKTW 453
Query: 413 EHSSP-----------------------GSTSKTSNCPNAY----------------VYP 433
+ P G+ S + Y P
Sbjct: 454 KWKKPATLFVYTRRNRKEHRKEPGVDAIGAGSSHKHLDGKYSCLDQVPSSDEGSCVMPVP 513
Query: 434 DAEFSDFDKDRKKECFAPGQIWAIYDSIDGMPRFYALIRKVLS-PGFQLQATWLEPRPDD 492
+A+F F D + F GQIWA YD DGMPR+YALI+KVLS F+++ +L+ + D
Sbjct: 514 EADFYTFG-DHPETSFQNGQIWAAYDEEDGMPRYYALIQKVLSRHPFKVRLAFLKAK--D 570
Query: 493 NDEI---KWVDEELPVACGKFKLCNTEIIEDHLTFSHLVMFKRNGRNTFQVYPRKGETWA 549
E W+ CG F + + + TFSH+V +++ +++PRKG+ WA
Sbjct: 571 CSEFVTSNWISYGYSKTCGDFIVGTPKNTDQLNTFSHVVTWEKGPGGIIRIFPRKGDIWA 630
Query: 550 LFKNWDITWYKDEESHRQYEYEFVEILSDYVEGEGVHVAYLGKLKGFVSIFIQIMKEDNQ 609
L++NW W Y+Y+ V++L Y G+ V + K+ GFVS+F ++ + +
Sbjct: 631 LYQNWSPEWNTCTPDDTIYKYDLVQVLDSYNPSAGISVMPIVKVPGFVSVFTPLL-DPTK 689
Query: 610 PFQIPSAELFRFSHRIPSFKMTGQEGVDVHLGYLEFDPASLPMNLEEI 657
IP E+ RFSH++P +TG+E + G E DP S P L ++
Sbjct: 690 SRTIPKEEMLRFSHQVPFHVLTGEEAKNSPKGCYELDPGSTPKELLQV 737
Score = 117 bits (293), Expect = 2e-24
Identities = 84/231 (36%), Positives = 123/231 (52%), Gaps = 15/231 (6%)
Query: 730 IPDAQFFNFETGRSLDKFQVGQIWAFYSDEDGMPKYYGQIKKVVTSPTIELHVYWLACCW 789
+P+A F+ F FQ GQIWA Y +EDGMP+YY I+KV++ ++ + +L
Sbjct: 512 VPEADFYTFGDHPETS-FQNGQIWAAYDEEDGMPRYYALIQKVLSRHPFKVRLAFLKAKD 570
Query: 790 LPE-NTTKWEDDGMLTSCGRFKVIKTKDFLSIYSNLSCISHQVQADP-IGKNYTIYPRKG 847
E T+ W G +CG F V K+ L+ SH V + G I+PRKG
Sbjct: 571 CSEFVTSNWISYGYSKTCGDFIVGTPKN----TDQLNTFSHVVTWEKGPGGIIRIFPRKG 626
Query: 848 EVWALYRKWSNKIK-CS-DLKNWDYDIVEVLEVADLFIETSILE--HVTGFSSVFRGKSI 903
++WALY+ WS + C+ D + YD+V+VL+ + S++ V GF SVF +
Sbjct: 627 DIWALYQNWSPEWNTCTPDDTIYKYDLVQVLDSYNPSAGISVMPIVKVPGFVSVF--TPL 684
Query: 904 EGSSGNLRIPKKELLRFSHQIPAFKLT--EEHGDLRGFWELDPGALPPSFL 952
+ + IPK+E+LRFSHQ+P LT E +G +ELDPG+ P L
Sbjct: 685 LDPTKSRTIPKEEMLRFSHQVPFHVLTGEEAKNSPKGCYELDPGSTPKELL 735
>UniRef100_Q9CAW2 Hypothetical protein T9J14.7 [Arabidopsis thaliana]
Length = 1165
Score = 202 bits (515), Expect = 3e-50
Identities = 154/468 (32%), Positives = 225/468 (47%), Gaps = 58/468 (12%)
Query: 227 DMQGTSPTT-----NPSQQASKANVGSQGNSHAEKSNTKPFKKKGPVGVSRKPDVKRKRN 281
D +G P T N S + KA+V + E+ +T K + +K +K+
Sbjct: 263 DAEGRKPQTSETGTNISAEMPKADVLKPQHQVKEEPDTSAEKSIPDLSAPQKNRAPKKKR 322
Query: 282 QVEEFSQSSDSTSSSDSEDETVAGKNGFPGVGNHSTEQPRRSVRQKHNVSYSDNMNGTDN 341
+V E S S SSD T K + + R+S R+K VSY+ G+D
Sbjct: 323 KVVEESSKSFEVDSSD----TAGAKT------DTNEHNKRKSSRKKPQVSYAKE--GSDG 370
Query: 342 DLLRPSKRGQENGSHCGDGRSYRETAKTNDQNGLAAD--------------PKNEHE--- 384
D + P + ++G + ++TA+ N LA KN H
Sbjct: 371 DFVSPPNKKTKSGFEFESEPNKKQTAEDNKSPKLAVSGVSSASSHSYKGKAKKNAHSGNE 430
Query: 385 ---KVKQKQEEKIRAGGKEAAEGSK--QMDKTFEHSSPGSTSKTSNCPNAYVYPDAEFSD 439
K K E G++AA SK +++K + K + N PD EFS
Sbjct: 431 DNLSAKNKVSEGCDGNGEDAALLSKIGRVEKGY---------KANENSNPLDIPDLEFSV 481
Query: 440 FDKDRKKECFAPGQIWAIY-DSIDGMPRFYALIRKVLSPGFQLQATWLEPRPDDNDEIKW 498
F +RK E FA Q+W+ D DGMPR YA ++KVL+ F+L+ T+L+P D DE
Sbjct: 482 FKVERKTEDFAVNQVWSTTTDCRDGMPRKYARVKKVLNGEFKLRITYLDPVLDKTDE--- 538
Query: 499 VDEELPVACGKFKLCNTEIIEDHLTFSHLVMFKRNGRNTFQVYPRKGETWALFKNWDITW 558
+PVACGKFK T ++D FS + R N +YPRKGE WA+F+ W+ W
Sbjct: 539 ---SIPVACGKFKNGKTMEVKDSSIFSGQMHHLRCN-NIVSIYPRKGEIWAIFREWEEEW 594
Query: 559 YKDEESHR-QYEYEFVEILSDYVEGEGVHVAYLGKLKGFVSIFIQIMKEDNQPFQIPSAE 617
+ H+ Y+Y+FVEI+SD+ + GV VAYLGKLKG V +F + Q +
Sbjct: 595 NTSLKKHKFPYKYDFVEIVSDFHDLNGVGVAYLGKLKGSVQLFHWEPQHGICQIQCSPKD 654
Query: 618 LFRFSHRIPSFKMTGQEGVDVHLGYLEFDPASLPMNLEEI-AVTQNLD 664
+ RFSH++P+ KMTG+E V E DPA+LP ++ ++ AV +D
Sbjct: 655 MLRFSHKVPAVKMTGKEKESVPPNSYELDPAALPKDIFQVDAVDMEMD 702
Score = 192 bits (489), Expect = 3e-47
Identities = 144/426 (33%), Positives = 208/426 (48%), Gaps = 50/426 (11%)
Query: 19 MENRDFAGARKFALKAQRLYPVLENIAQMLVVCDVHCSAEQKVFGDEINWYGILQLERTA 78
ME DF GA KF KAQRL+P LENI QM+ +CDVH SA +K+ G + +WYG+LQ++ A
Sbjct: 1 MEAGDFVGAHKFVTKAQRLFPNLENIVQMMTICDVHSSAIKKIKGLD-DWYGVLQVQPYA 59
Query: 79 GDAMIKKQFRKFALQLHPDKNKFAGAEAAFKLIGEAQRVLSDREKRTRYDMKLNVNKTAM 138
IKKQ+RK AL LHPDKNKFAGAEAAFKL+GEA R+LSD+ KR++YD + +
Sbjct: 60 DADTIKKQYRKLALLLHPDKNKFAGAEAAFKLVGEANRLLSDQIKRSQYDNRYRSHSMFA 119
Query: 139 PPRSNQPKVPTNFNSATKNNVRTNFTNSNTQQPPQQQNKQPPQQQNGVRRTFWTACPFCS 198
N V + + A NN N GV TFWT C C
Sbjct: 120 NRHVN---VYSGRHCAATNNAAENIA--------------------GV-FTFWTRCRHCG 155
Query: 199 VKYEYYREILNKSLRCQQCHRLFVAYILDMQGTSPTTNPS-----QQASKANVGSQGNSH 253
Y+Y RE +N S+ C C + FVA + G P+++ + Q +N Q S
Sbjct: 156 QCYKYLREYMNTSMHCSSCQKSFVACKMRCDGVPPSSSTAGRKEFQDQVMSNTSRQNAST 215
Query: 254 AEKSNTK--PFKKKGPVG--VSRKPDVKRKRNQVEEFSQSSDSTSSSDSED-ETVAGKNG 308
A +S + K G VG V++K K+K V ++ + + +D+E + + G
Sbjct: 216 AAESGSSAADMGKNGKVGGKVNKKNQEKQKNGAVNRGTKKEEGCTENDAEGRKPQTSETG 275
Query: 309 FPGVGNHSTEQPRRSV-RQKHNVSYSDNMNGTDN--DLLRPSK-RGQENGSHCGDGRSYR 364
N S E P+ V + +H V + + + DL P K R + + S
Sbjct: 276 ----TNISAEMPKADVLKPQHQVKEEPDTSAEKSIPDLSAPQKNRAPKKKRKVVEESSKS 331
Query: 365 ETAKTNDQNGLAADPKNEHEKVKQKQEEKIRAGGKEAAEG------SKQMDKTFEHSSPG 418
++D G D NEH K K +++ + KE ++G +K+ FE S
Sbjct: 332 FEVDSSDTAGAKTD-TNEHNKRKSSRKKPQVSYAKEGSDGDFVSPPNKKTKSGFEFESEP 390
Query: 419 STSKTS 424
+ +T+
Sbjct: 391 NKKQTA 396
Score = 117 bits (293), Expect = 2e-24
Identities = 112/351 (31%), Positives = 152/351 (42%), Gaps = 45/351 (12%)
Query: 324 VRQKHNVSYSDNMNGTDNDLLRPSKRGQENGSHCGD-------GRSYRETAKTN-----D 371
V +K N TD LR S R Q S GD G KT+ D
Sbjct: 830 VDEKKTSKSRKNGEATDVFKLRKSPRLQTIPSQQGDEMKSTKQGNKMNTPKKTDKGLETD 889
Query: 372 QNGLAADPKNEHEKVKQKQEEKIRAGG--KEAAEGSKQMDKTFE-----HSSPGSTSKTS 424
G+ P H+ + ++ E + G E SKQ D + + SP +T +
Sbjct: 890 SLGVRKSPNGIHQPAESQEGESSKKQGCNGEIPSLSKQNDLPTQLGGSTYKSPRTTHVSP 949
Query: 425 NCPNAYVYPDAEFSDFDKDRKKECFAPGQIWAIYDSIDGMPRFYALIRKV-LSPGFQLQA 483
+C P DF R ++ F QIWAIY + +GMP Y I+K+ P F L+
Sbjct: 950 HCKT----PRRNAFDFQNLRSEDKFEVNQIWAIYSNDNGMPLEYVKIKKIETKPKFVLRG 1005
Query: 484 TWLEPRPDDNDEIKWVDEELPVACGKFKLCNTEI-IEDHLTFSHLVM-FKRNGRNTFQVY 541
T E P + + V+CG+FKL I H +FSHLV F + R F+VY
Sbjct: 1006 TPTELYPPSTEPVTRT-----VSCGEFKLLKGRPKIIPHASFSHLVKPFDSSKRFRFKVY 1060
Query: 542 PRKGETWALFKNWDITWYKDEESHRQYEYEFVEILSDYVEGEGVHVAYLGKLKGFVSIFI 601
PRKGE WAL+KN D T E + VE++ D +GE V V L S F
Sbjct: 1061 PRKGEIWALYKNCDST----------EEPDIVEVVEDNCDGEIVKVV---ALTAMGSSFQ 1107
Query: 602 QIMKEDNQPFQIPSAELFRFSHRIPSFKMTGQEGVDVHLGYL-EFDPASLP 651
+ D I AE+ RFSH+IP+ + + V GY E DP ++P
Sbjct: 1108 RKQGSDVGLIDISKAEMSRFSHQIPAIRHPKKTTRLVKGGYYWELDPIAIP 1158
Score = 105 bits (261), Expect = 8e-21
Identities = 83/224 (37%), Positives = 120/224 (53%), Gaps = 25/224 (11%)
Query: 731 PDAQFFNFETGRSLDKFQVGQIWAFYSDEDGMPKYYGQIKKVVTSPTIELHVYWLACCWL 790
P F+F+ RS DKF+V QIWA YS+++GMP Y +IKK+ T P L
Sbjct: 954 PRRNAFDFQNLRSEDKFEVNQIWAIYSNDNGMPLEYVKIKKIETKPKFVLR--GTPTELY 1011
Query: 791 PENTTKWEDDGMLTSCGRFKVIKTKDFLSIYSNLSCISHQVQADPIGK--NYTIYPRKGE 848
P +T E SCG FK++K + + +++ SH V+ K + +YPRKGE
Sbjct: 1012 PPST---EPVTRTVSCGEFKLLKGRPKIIPHAS---FSHLVKPFDSSKRFRFKVYPRKGE 1065
Query: 849 VWALYRKWSNKIKCSDLKNWDYDIVEVLEVADLFIETSILEHVTGFSSVFRGKSIEGSS- 907
+WALY+ C + + DIVEV+E + E + +T S F+ K +GS
Sbjct: 1066 IWALYK------NCDSTE--EPDIVEVVE-DNCDGEIVKVVALTAMGSSFQRK--QGSDV 1114
Query: 908 GNLRIPKKELLRFSHQIPAFKLTEEHGDL-RG--FWELDPGALP 948
G + I K E+ RFSHQIPA + ++ L +G +WELDP A+P
Sbjct: 1115 GLIDISKAEMSRFSHQIPAIRHPKKTTRLVKGGYYWELDPIAIP 1158
Score = 102 bits (254), Expect = 5e-20
Identities = 72/228 (31%), Positives = 119/228 (51%), Gaps = 20/228 (8%)
Query: 729 EIPDAQFFNFETGRSLDKFQVGQIWAFYSD-EDGMPKYYGQIKKVVTSPTIELHVYWLAC 787
+IPD +F F+ R + F V Q+W+ +D DGMP+ Y ++KKV+ +L + +L
Sbjct: 473 DIPDLEFSVFKVERKTEDFAVNQVWSTTTDCRDGMPRKYARVKKVLNGE-FKLRITYL-- 529
Query: 788 CWLPENTTKWEDDGMLTSCGRFKVIKTKDFLSIYSNLSCISHQVQADPIGKNYTIYPRKG 847
+ D+ + +CG+FK KT + + S S Q+ +IYPRKG
Sbjct: 530 ----DPVLDKTDESIPVACGKFKNGKTMEV----KDSSIFSGQMHHLRCNNIVSIYPRKG 581
Query: 848 EVWALYRKWSNKIKCSDLKN---WDYDIVEVL-EVADL-FIETSILEHVTGFSSVFRGKS 902
E+WA++R+W + S K+ + YD VE++ + DL + + L + G +F +
Sbjct: 582 EIWAIFREWEEEWNTSLKKHKFPYKYDFVEIVSDFHDLNGVGVAYLGKLKGSVQLFHWEP 641
Query: 903 IEGSSGNLRIPKKELLRFSHQIPAFKLT--EEHGDLRGFWELDPGALP 948
G ++ K++LRFSH++PA K+T E+ +ELDP ALP
Sbjct: 642 QHGIC-QIQCSPKDMLRFSHKVPAVKMTGKEKESVPPNSYELDPAALP 688
>UniRef100_Q9FHS7 Similarity to DnaJ protein [Arabidopsis thaliana]
Length = 431
Score = 182 bits (462), Expect = 4e-44
Identities = 123/361 (34%), Positives = 175/361 (48%), Gaps = 58/361 (16%)
Query: 1 MDCNKEEALRAKEIAEKKMENRDFAGARKFALKAQRLYPVLENIAQMLVVCDVHCSAEQK 60
M+CNK+EA RAKEIAE K + +D AGA+KFALKAQ LYP +E ++QML DV+ +AE K
Sbjct: 1 MECNKDEATRAKEIAENKFKMKDIAGAKKFALKAQNLYPEIEGVSQMLATLDVYIAAENK 60
Query: 61 VFGDEINWYGILQLERTAGDAMIKKQFRKFALQLHPDKNKFAGAEAAFKLIGEAQRVLSD 120
V ++++WYGIL D +K+++RK AL LHPDKNK GAE AFK + EA + LSD
Sbjct: 61 V-NEDVDWYGILNASPRDDDETLKRKYRKLALMLHPDKNKSIGAEGAFKHVSEAWKFLSD 119
Query: 121 REKRTRYDMKLNVN----KTAMPPRSN--------------QPKVPTNFNSATKNNVRTN 162
+EKR YD + +++ K ++ +N + P N A K N
Sbjct: 120 KEKRAAYDRRKSLHSVYQKVSVSSSNNGFCNFAKTTFTTNARTTTPRNNPPAQKTNPPAQ 179
Query: 163 FTNSNTQQ--PPQQQNKQPPQQQNGVR--------------------------------R 188
TN Q+ PP Q+N P Q+ N +
Sbjct: 180 KTNPPAQKTNPPAQKNNPPTQKNNPQKPVGTTQKTGRTDNHTTTPNSFTASGSSDQSKSN 239
Query: 189 TFWTACPFCSVKYEYYREILNKSLRCQQCHRLFVAYILDMQG-TSPTTNPSQQASKAN-- 245
TFWT C C ++YEY R +N +LRC C + ++A + G +S ++ S+ S AN
Sbjct: 240 TFWTVCRRCMMQYEYLRVYVNCNLRCPNCLQSYLAVEVPKPGISSRWSSCSRLKSAANHN 299
Query: 246 --VGSQGNSHAEKSNTKPFKKKGPVGVSRKPDVKRKRNQVEEFSQSSDSTSSSDSEDETV 303
G NS S T V VK+ R Q + ++ + S ++
Sbjct: 300 TTSGLFNNSKWTFSRTSSAAHAASVVQHAYEKVKKDREQAKATARREKKNAKRKSTTDSS 359
Query: 304 A 304
A
Sbjct: 360 A 360
>UniRef100_Q9CAW0 Hypothetical protein T9J14.9 [Arabidopsis thaliana]
Length = 603
Score = 177 bits (448), Expect = 2e-42
Identities = 105/293 (35%), Positives = 157/293 (52%), Gaps = 31/293 (10%)
Query: 378 DPKNEHEKVKQKQEEK--IRAGGKEAAEGSKQMDKTFEHSS-------PGSTSKTSNCPN 428
+ + +H ++KQ+++EK IRA ++ + + + E PG +
Sbjct: 303 EEREQHLELKQRKKEKPAIRAETRKRSRLEYESPLSAEKGRYAETLIRPGKKQVQKREAH 362
Query: 429 AYVY---------PDAEFSDFDKDRKKECFAPGQIWAIYDSIDGMPRFYALIRKVLSPGF 479
+Y PD +F DF+ FA GQ+WA+YD +D MPR+YA IRKVL P
Sbjct: 363 EIIYIDEDEPFNCPDPDFHDFNNTMSS--FAVGQVWALYDPVDDMPRYYAEIRKVLQPQL 420
Query: 480 QLQATWLEPRPDDNDEIKWVDEELPVACGKFKLCNTEIIEDHLTFSHLVMFKRNGRNTFQ 539
L+ TWLE ++ +E +P ACG+F+ +E HL FSH + G+
Sbjct: 421 SLRVTWLE-------SLQTTEEPIP-ACGRFEHGKSET-SSHLMFSHEMYHTIRGQYV-T 470
Query: 540 VYPRKGETWALFKNWDITWYKDEESHRQ-YEYEFVEILSDYVEGEGVHVAYLGKLKGFVS 598
+ PRKGETWALF +W TW E + Y Y+FVE+++++ G+ VAYLG+++GF S
Sbjct: 471 INPRKGETWALFGDWTKTWKSHSEQQKTPYSYDFVEVVTEFDSDRGIGVAYLGRVEGFTS 530
Query: 599 IFIQIMKEDNQPFQIPSAELFRFSHRIPSFKMTGQEGVDVHLGYLEFDPASLP 651
++ + + I E+ RFSHR+PSFKMTG E V G E DPA++P
Sbjct: 531 VYERAAQNGLVEIMISCDEMLRFSHRVPSFKMTGDEKEGVPAGSFELDPAAVP 583
Score = 124 bits (311), Expect = 1e-26
Identities = 82/234 (35%), Positives = 122/234 (52%), Gaps = 24/234 (10%)
Query: 726 EAFEIPDAQFFNFETGRSLDKFQVGQIWAFYSDEDGMPKYYGQIKKVVTSPTIELHVYWL 785
E F PD F +F ++ F VGQ+WA Y D MP+YY +I+KV+ P + L V WL
Sbjct: 371 EPFNCPDPDFHDFNN--TMSSFAVGQVWALYDPVDDMPRYYAEIRKVL-QPQLSLRVTWL 427
Query: 786 ACCWLPENTTKWEDDGMLTSCGRFKVIKTKDFLSIYSNLSCISHQVQADPIGKNYTIYPR 845
E + +CGRF+ K++ S+ SH++ G+ TI PR
Sbjct: 428 ESLQTTEEP--------IPACGRFEHGKSET-----SSHLMFSHEMYHTIRGQYVTINPR 474
Query: 846 KGEVWALYRKWSNKIKCSDLKN---WDYDIVEVLEV--ADLFIETSILEHVTGFSSVFRG 900
KGE WAL+ W+ K + + YD VEV+ +D I + L V GF+SV+
Sbjct: 475 KGETWALFGDWTKTWKSHSEQQKTPYSYDFVEVVTEFDSDRGIGVAYLGRVEGFTSVYE- 533
Query: 901 KSIEGSSGNLRIPKKELLRFSHQIPAFKLT--EEHGDLRGFWELDPGALPPSFL 952
++ + + I E+LRFSH++P+FK+T E+ G G +ELDP A+P +L
Sbjct: 534 RAAQNGLVEIMISCDEMLRFSHRVPSFKMTGDEKEGVPAGSFELDPAAVPRVYL 587
>UniRef100_Q9MAA8 T12H1.7 protein [Arabidopsis thaliana]
Length = 372
Score = 176 bits (447), Expect = 2e-42
Identities = 104/262 (39%), Positives = 141/262 (53%), Gaps = 23/262 (8%)
Query: 385 KVKQKQEEKIRAGGKEAAEGSKQM------DKTFEHSSPGSTS-----KTSNCPNAYVYP 433
K +Q QE I+ G AE Q DK + +S S S +
Sbjct: 103 KQRQVQERSIQDGPSVDAEPLTQQRNHNDEDKEKDSASVLSASVQIIENDEDHEPVMCVV 162
Query: 434 DAEFSDFDKDRKKECFAPGQIWAIYDSIDGMPRFYALIRKVLSPGFQLQATWLEPRPDDN 493
D+EF+DF K F GQ+WA+YD ID MPR Y I+KV LQ TWLEP+
Sbjct: 163 DSEFNDFRKTMSS--FMAGQVWALYDGIDSMPRCYGRIKKVNKCQSSLQVTWLEPK---- 216
Query: 494 DEIKWVDEELPVACGKFKLCNTEIIEDHLTFSHLVMFKRNGRNTFQVYPRKGETWALFKN 553
+E + ACG+FK NT+ I+ HL FSH + G++ V P KGETWALF++
Sbjct: 217 -----AEESVLAACGRFKWENTDTIQSHLAFSHEIHPIIRGKHFIAVNPSKGETWALFRD 271
Query: 554 WDITWYKDEESHRQ-YEYEFVEILSDYVEGEGVHVAYLGKLKGFVSIFIQIMKEDNQPFQ 612
W +W D E H+ Y Y+FVE+L + + GV VAYLGK++GF S++ Q ++ F
Sbjct: 272 WSKSWNNDPEQHKTPYRYDFVEVLVSFDDSLGVGVAYLGKVQGFASVYKQAVQHGVISFM 331
Query: 613 IPSAELFRFSHRIPSFKMTGQE 634
I E+ RFSHR+PSF++ G E
Sbjct: 332 ITPEEMQRFSHRVPSFRLNGDE 353
Score = 100 bits (249), Expect = 2e-19
Identities = 71/206 (34%), Positives = 113/206 (54%), Gaps = 23/206 (11%)
Query: 730 IPDAQFFNFETGRSLDKFQVGQIWAFYSDEDGMPKYYGQIKKVVTSPTIELHVYWLACCW 789
+ D++F +F +++ F GQ+WA Y D MP+ YG+IKKV + L V WL
Sbjct: 161 VVDSEFNDFR--KTMSSFMAGQVWALYDGIDSMPRCYGRIKKVNKCQS-SLQVTWLE--- 214
Query: 790 LPENTTKWEDDGMLTSCGRFKVIKTKDFLSIYSNLSCISHQVQADPIGKNY-TIYPRKGE 848
P+ ++ +L +CGRFK T +I S+L+ SH++ GK++ + P KGE
Sbjct: 215 -PK-----AEESVLAACGRFKWENTD---TIQSHLA-FSHEIHPIIRGKHFIAVNPSKGE 264
Query: 849 VWALYRKWS---NKIKCSDLKNWDYDIVEVLEVAD--LFIETSILEHVTGFSSVFRGKSI 903
WAL+R WS N + YD VEVL D L + + L V GF+SV++ +++
Sbjct: 265 TWALFRDWSKSWNNDPEQHKTPYRYDFVEVLVSFDDSLGVGVAYLGKVQGFASVYK-QAV 323
Query: 904 EGSSGNLRIPKKELLRFSHQIPAFKL 929
+ + I +E+ RFSH++P+F+L
Sbjct: 324 QHGVISFMITPEEMQRFSHRVPSFRL 349
>UniRef100_Q84TH2 Hypothetical protein At4g19570 [Arabidopsis thaliana]
Length = 558
Score = 157 bits (398), Expect = 1e-36
Identities = 101/293 (34%), Positives = 152/293 (51%), Gaps = 29/293 (9%)
Query: 1 MDCNKEEALRAKEIAEKKMENRDFAGARKFALKAQRLYPVLENIAQMLVVCDVHCSAEQK 60
M+ NKEEA RA +IAEKK+ D+ A+++A KA R+YP L + Q+L++ DV+ SA K
Sbjct: 1 MESNKEEAKRALDIAEKKLSKNDYNRAKRYAKKAHRMYPNLVGLEQVLIMIDVYISATNK 60
Query: 61 VFGDEINWYGILQLERTAGDAMIKKQFRKFALQLHPDKNKFAGAEAAFKLIGEAQRVLSD 120
+ G E +WY +L ++ A D +KK++RK AL LHPDKN+F GAE AFKLI EA +LSD
Sbjct: 61 ING-EADWYRVLGVDPLADDEAVKKRYRKLALLLHPDKNRFTGAEGAFKLILEAWDLLSD 119
Query: 121 REKRTRYDMKLNVNKTAM-------PPRSNQPKVPTNFNSAT------------KNNVRT 161
+ +R+ YD K N+ P RS+ PK + A+ K R
Sbjct: 120 KSQRSSYDQKRKSNQVKQRTSGMQKPKRSSTPKPTESDKPASSYGPTPPPEPRPKRRPRP 179
Query: 162 NFTNSNTQQPPQQQNK---QPPQQQNGVR-RTFWTACPFCSVKYEYYR-EILNKSLRCQQ 216
N + P ++K +P V+ TFWT C C E+ R LNK++ C
Sbjct: 180 NIPEPDIPMPMPTRHKPKSKPDISLTTVKVGTFWTVCNRCKTYCEFMRASCLNKTVPCPN 239
Query: 217 CHRLFVAYILDMQGTSP----TTNPSQQASKANVGSQGNSHAEKSNTKPFKKK 265
C + F+A ++ + + +P Q++ +S + + T P + K
Sbjct: 240 CGKYFIATVIPSELVNGRLVIRLSPCNQSTWKKASDDTSSTSAMAKTVPERVK 292
Score = 35.8 bits (81), Expect = 6.3
Identities = 31/130 (23%), Positives = 61/130 (46%), Gaps = 12/130 (9%)
Query: 148 PTNFNSATKNNVRTNFTNSNTQQPPQQQNK---QPPQQQNGVRR--TFWTACPFCSVKYE 202
P N ++ K + T+ T++ + P++ + P+ + ++ TFWT C C +
Sbjct: 264 PCNQSTWKKASDDTSSTSAMAKTVPERVKRWFDPMPEMDSSTQKIGTFWTVCSRCKTHCK 323
Query: 203 YYR-EILNKSLRCQQCHRLFVA--YILDMQGTSPT--TNPSQQAS--KANVGSQGNSHAE 255
R LNK+ C C + FVA I+++ P +PS Q++ + S S ++
Sbjct: 324 LVRANHLNKTFPCPNCSQEFVAAEMIIEVINGGPVIKLSPSVQSNSKRTRDASSTTSASD 383
Query: 256 KSNTKPFKKK 265
K+ ++K+
Sbjct: 384 KAQRTAYQKE 393
>UniRef100_O49475 Hypothetical protein F24J7.130 [Arabidopsis thaliana]
Length = 539
Score = 153 bits (386), Expect = 3e-35
Identities = 94/244 (38%), Positives = 133/244 (53%), Gaps = 25/244 (10%)
Query: 1 MDCNKEEALRAKEIAEKKMENRDFAGARKFALKAQRLYPVLENIAQMLVVCDVHCSAEQK 60
M+ NKEEA RA +IAEKK+ D+ A+++A KA R+YP L + Q+L++ DV+ SA K
Sbjct: 1 MESNKEEAKRALDIAEKKLSKNDYNRAKRYAKKAHRMYPNLVGLEQVLIMIDVYISATNK 60
Query: 61 VFGDEINWYGILQLERTAGDAMIKKQFRKFALQLHPDKNKFAGAEAAFKLIGEAQRVLSD 120
+ G E +WY +L ++ A D +KK++RK AL LHPDKN+F GAE AFKLI EA +LSD
Sbjct: 61 ING-EADWYRVLGVDPLADDEAVKKRYRKLALLLHPDKNRFTGAEGAFKLILEAWDLLSD 119
Query: 121 REKRTRYDMKLNVNKTAM-------PPRSNQPKVPTNFNSAT------------KNNVRT 161
+ +R+ YD K N+ P RS+ PK + A+ K R
Sbjct: 120 KSQRSSYDQKRKSNQVKQRTSGMQKPKRSSTPKPTESDKPASSYGPTPPPEPRPKRRPRP 179
Query: 162 NFTNSNTQQPPQQQNK---QPPQQQNGVR-RTFWTACPFCSVKYEYYR-EILNKSLRCQQ 216
N + P ++K +P V+ TFWT C C E+ R LNK++ C
Sbjct: 180 NIPEPDIPMPMPTRHKPKSKPDISLTTVKVGTFWTVCNRCKTYCEFMRASCLNKTVPCPN 239
Query: 217 CHRL 220
C +L
Sbjct: 240 CGKL 243
Score = 37.4 bits (85), Expect = 2.2
Identities = 35/145 (24%), Positives = 67/145 (46%), Gaps = 13/145 (8%)
Query: 133 VNKTAMPPRSNQPKVPTNFNSATKNNVRTNFTNSNTQQPPQQQNK---QPPQQQNGVRR- 188
+NKT P + P N ++ K + T+ T++ + P++ + P+ + ++
Sbjct: 231 LNKTVPCPNCGKLS-PCNQSTWKKASDDTSSTSAMAKTVPERVKRWFDPMPEMDSSTQKI 289
Query: 189 -TFWTACPFCSVKYEYYR-EILNKSLRCQQCHRLFVA--YILDMQGTSPT--TNPSQQAS 242
TFWT C C + R LNK+ C C + FVA I+++ P +PS Q++
Sbjct: 290 GTFWTVCSRCKTHCKLVRANHLNKTFPCPNCSQEFVAAEMIIEVINGGPVIKLSPSVQSN 349
Query: 243 --KANVGSQGNSHAEKSNTKPFKKK 265
+ S S ++K+ ++K+
Sbjct: 350 SKRTRDASSTTSASDKAQRTAYQKE 374
>UniRef100_Q9SUE1 Hypothetical protein T13J8.90 [Arabidopsis thaliana]
Length = 565
Score = 152 bits (385), Expect = 4e-35
Identities = 88/246 (35%), Positives = 136/246 (54%), Gaps = 13/246 (5%)
Query: 420 TSKTSNCPNAYVYPDAEFSDFDKDRKKECFAPGQIWAIYDSIDGMPRFYALIRKVLSPGF 479
T K+ PN+ PD + +DF K FA Q+WA+YD D MPR YA IR++
Sbjct: 306 TDKSYEDPNSLTCPDTKLNDFSKSMSS--FAVDQVWALYDPRDDMPRNYAQIREIFESQL 363
Query: 480 QLQATWLEPRPDDNDEIKWVDEELPVACGKFKLCNTEIIEDHLTFSHLVMFKRNGRNTFQ 539
LQ T LE DE + + CG+F+ +TE I+ HL F+H + ++
Sbjct: 364 SLQVTLLEHVKTTKDE-----QSILSGCGRFEYGDTE-IKSHLMFAHEMDHIKSAEEVI- 416
Query: 540 VYPRKGETWALFKNWDITWYKD-EESHRQYEYEFVEILSDYVEGEGVHVAYLGKLKGFVS 598
V PRKGETWALF +W+ +W E Y Y+FVE++S++ + G+ VAY+G+++G+ S
Sbjct: 417 VNPRKGETWALFSDWNASWNSHLELQELPYRYDFVEVISEFDDLIGIQVAYMGRVEGYES 476
Query: 599 IFIQIMKEDNQPFQIPSAELFRFSHRIPSFKMTGQEGVDVHLGYLEFDPASLPMN---LE 655
+F + IP AE+ RFSH++ S K++G+E + + +PA++P LE
Sbjct: 477 VFNHAEQYGCIKIVIPPAEMQRFSHKVESVKLSGKEEEGIPFRSFKLNPAAMPRYYHVLE 536
Query: 656 EIAVTQ 661
E+ T+
Sbjct: 537 EVVETE 542
Score = 100 bits (248), Expect = 3e-19
Identities = 92/322 (28%), Positives = 150/322 (46%), Gaps = 34/322 (10%)
Query: 643 LEFDPASLPMNLEEIAVTQNLDMRTGHSSCGSENARTSKRSKPSMSPEDIVS-------- 694
LE +L L+E+ + Q T +R KRS + P +V
Sbjct: 226 LEIKEKTLEKRLKELELKQMELEETSRPQLVEAESR--KRSNLEIEPPLLVKNDSDADSC 283
Query: 695 TPKVKVDTSNLTDVKDSLDDMDDCHASASTPEAFEIPDAQFFNFETGRSLDKFQVGQIWA 754
TP+ K S + D ++ + S P + PD + +F +S+ F V Q+WA
Sbjct: 284 TPQAKKQKSQEANDGD-IEGIVCTDKSYEDPNSLTCPDTKLNDFS--KSMSSFAVDQVWA 340
Query: 755 FYSDEDGMPKYYGQIKKVVTSPTIELHVYWLACCWLPENTTKWEDDGMLTSCGRFKVIKT 814
Y D MP+ Y QI+++ S + L V L TTK ++ +L+ CGRF+ T
Sbjct: 341 LYDPRDDMPRNYAQIREIFES-QLSLQVTLLE----HVKTTK-DEQSILSGCGRFEYGDT 394
Query: 815 KDFLSIYSNLSCISHQVQADPIGKNYTIYPRKGEVWALY----RKWSNKIKCSDLKNWDY 870
+ I S+L +H++ + + PRKGE WAL+ W++ ++ +L + Y
Sbjct: 395 E----IKSHL-MFAHEMDHIKSAEEVIVNPRKGETWALFSDWNASWNSHLELQELP-YRY 448
Query: 871 DIVEVLEVAD--LFIETSILEHVTGFSSVFRGKSIEGSSGNLRIPKKELLRFSHQIPAFK 928
D VEV+ D + I+ + + V G+ SVF G + IP E+ RFSH++ + K
Sbjct: 449 DFVEVISEFDDLIGIQVAYMGRVEGYESVFNHAEQYGCI-KIVIPPAEMQRFSHKVESVK 507
Query: 929 LT--EEHGDLRGFWELDPGALP 948
L+ EE G ++L+P A+P
Sbjct: 508 LSGKEEEGIPFRSFKLNPAAMP 529
>UniRef100_Q67ZV4 Hypothetical protein At4g27980 [Arabidopsis thaliana]
Length = 291
Score = 147 bits (371), Expect = 1e-33
Identities = 86/246 (34%), Positives = 134/246 (53%), Gaps = 13/246 (5%)
Query: 420 TSKTSNCPNAYVYPDAEFSDFDKDRKKECFAPGQIWAIYDSIDGMPRFYALIRKVLSPGF 479
T K+ PN+ P + +DF K FA Q+WA+YD D MPR YA IR++
Sbjct: 32 TDKSYEDPNSLTCPGTKLNDFSKSMSS--FAVDQVWALYDPRDDMPRNYAQIREIFESQL 89
Query: 480 QLQATWLEPRPDDNDEIKWVDEELPVACGKFKLCNTEIIEDHLTFSHLVMFKRNGRNTFQ 539
LQ T LE E + + CG+F+ +TE I+ HL F+H + ++
Sbjct: 90 SLQVTLLEHVKTTKGE-----QSILSGCGRFEYGDTE-IKSHLMFAHEMDHIKSAEEVI- 142
Query: 540 VYPRKGETWALFKNWDITWYKD-EESHRQYEYEFVEILSDYVEGEGVHVAYLGKLKGFVS 598
V PRKGETWALF +W+ +W E Y Y+FVE++S++ + G+ VAY+G+++G+ S
Sbjct: 143 VNPRKGETWALFSDWNASWNSHLELQELPYRYDFVEVISEFDDLIGIQVAYMGRVEGYES 202
Query: 599 IFIQIMKEDNQPFQIPSAELFRFSHRIPSFKMTGQEGVDVHLGYLEFDPASLPMN---LE 655
+F + IP AE+ RFSH++ S K++G+E + + +PA++P LE
Sbjct: 203 VFNHAEQYGCIKIVIPPAEMQRFSHKVESVKLSGKEEEGIPFRSFKLNPAAMPRYYHVLE 262
Query: 656 EIAVTQ 661
E+ T+
Sbjct: 263 EVVETE 268
Score = 94.4 bits (233), Expect = 1e-17
Identities = 79/262 (30%), Positives = 128/262 (48%), Gaps = 24/262 (9%)
Query: 695 TPKVKVDTSNLTDVKDSLDDMDDCHASASTPEAFEIPDAQFFNFETGRSLDKFQVGQIWA 754
TP+ K S + D ++ + S P + P + +F +S+ F V Q+WA
Sbjct: 10 TPQAKKQKSQEANDGD-IEGIVCTDKSYEDPNSLTCPGTKLNDFS--KSMSSFAVDQVWA 66
Query: 755 FYSDEDGMPKYYGQIKKVVTSPTIELHVYWLACCWLPENTTKWEDDGMLTSCGRFKVIKT 814
Y D MP+ Y QI+++ S + L V L TTK E +L+ CGRF+ T
Sbjct: 67 LYDPRDDMPRNYAQIREIFES-QLSLQVTLLE----HVKTTKGE-QSILSGCGRFEYGDT 120
Query: 815 KDFLSIYSNLSCISHQVQADPIGKNYTIYPRKGEVWALY----RKWSNKIKCSDLKNWDY 870
+ I S+L +H++ + + PRKGE WAL+ W++ ++ +L + Y
Sbjct: 121 E----IKSHL-MFAHEMDHIKSAEEVIVNPRKGETWALFSDWNASWNSHLELQELP-YRY 174
Query: 871 DIVEVLEVAD--LFIETSILEHVTGFSSVFRGKSIEGSSGNLRIPKKELLRFSHQIPAFK 928
D VEV+ D + I+ + + V G+ SVF G + IP E+ RFSH++ + K
Sbjct: 175 DFVEVISEFDDLIGIQVAYMGRVEGYESVFNHAEQYGCI-KIVIPPAEMQRFSHKVESVK 233
Query: 929 LT--EEHGDLRGFWELDPGALP 948
L+ EE G ++L+P A+P
Sbjct: 234 LSGKEEEGIPFRSFKLNPAAMP 255
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.316 0.132 0.400
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,727,756,156
Number of Sequences: 2790947
Number of extensions: 78136147
Number of successful extensions: 276442
Number of sequences better than 10.0: 3042
Number of HSP's better than 10.0 without gapping: 1234
Number of HSP's successfully gapped in prelim test: 1871
Number of HSP's that attempted gapping in prelim test: 267654
Number of HSP's gapped (non-prelim): 6642
length of query: 954
length of database: 848,049,833
effective HSP length: 137
effective length of query: 817
effective length of database: 465,690,094
effective search space: 380468806798
effective search space used: 380468806798
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 80 (35.4 bits)
Medicago: description of AC147364.8


