

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147363.1 - phase: 0 /pseudo
(427 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
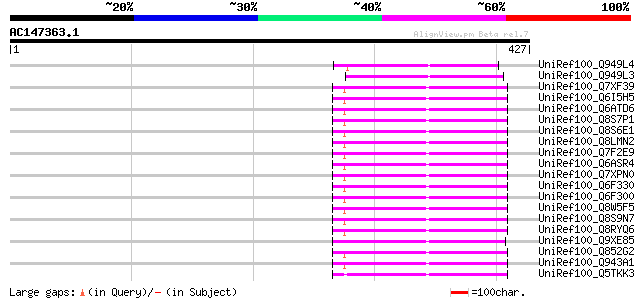
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q949L4 Putative polyprotein [Cicer arietinum] 89 3e-16
UniRef100_Q949L3 Putative polyprotein [Cicer arietinum] 87 9e-16
UniRef100_Q7XF39 Similar to Sorghum bicolor 22 kDakafirincluster... 70 1e-10
UniRef100_Q6I5H5 Putative polyprotein [Oryza sativa] 69 2e-10
UniRef100_Q6ATD6 Putative polyprotein [Oryza sativa] 69 2e-10
UniRef100_Q8S7P1 Putative polyprotein [Oryza sativa] 69 3e-10
UniRef100_Q8S6E1 Putative polyprotein [Oryza sativa] 69 3e-10
UniRef100_Q8LMN2 Putative polyprotein [Oryza sativa] 69 3e-10
UniRef100_Q7F2E9 Putative polyprotein [Oryza sativa] 69 3e-10
UniRef100_Q6ASR4 Putative polyprotein [Oryza sativa] 69 3e-10
UniRef100_Q7XPN0 OSJNBa0060D06.14 protein [Oryza sativa] 69 3e-10
UniRef100_Q6F330 Putative polyprotein [Oryza sativa] 69 3e-10
UniRef100_Q6F300 Putative polyprotein [Oryza sativa] 69 3e-10
UniRef100_Q8W5F5 Putative polyprotein [Oryza sativa] 69 3e-10
UniRef100_Q8S9N7 Similar to polyprotein [Oryza sativa] 69 3e-10
UniRef100_Q8RYQ6 Putative Sorghum bicolor 22 kDa kafirin cluster... 69 3e-10
UniRef100_Q9XE85 Polyprotein [Sorghum bicolor] 68 4e-10
UniRef100_Q852G2 Putative polyprotein [Oryza sativa] 68 4e-10
UniRef100_Q943A1 Putative polyprotein [Oryza sativa] 67 7e-10
UniRef100_Q5TKK3 Hypothetical protein OJ1532_D06.4 [Oryza sativa] 67 7e-10
>UniRef100_Q949L4 Putative polyprotein [Cicer arietinum]
Length = 318
Score = 88.6 bits (218), Expect = 3e-16
Identities = 50/147 (34%), Positives = 76/147 (51%), Gaps = 12/147 (8%)
Query: 267 CKQPKKAPTI-----------GRVFALTGTQTENEDRLIIGTCYINNTPLVAIIDIGATH 315
C PK P + GR++ + + LI C I L A+ D GATH
Sbjct: 108 CTAPKVEPVVNVARATRPTARGRIYCMNAEEGNPSGNLIQRDCEIAGNTLTALFDSGATH 167
Query: 316 CFIAFDCASTLGLDMSNMNGEMVVETPAKGSVTTSLVCLKSSLSMFGRDFEMDLVCLPLS 375
FIA DC + L L +S + ++VV TP K ++T + CL +++ RD ++L+CLPL
Sbjct: 168 SFIAMDCVNRLKLSVSALPFDLVVSTPDK-TLTVNSACLHCRMTIQNRDLLVNLICLPLQ 226
Query: 376 GMYVILGMNWFE*NHVHINYFSKSVYF 402
+ VILGM+W ++V ++ K V F
Sbjct: 227 SLEVILGMDWMSYHYVILDCARKLVIF 253
>UniRef100_Q949L3 Putative polyprotein [Cicer arietinum]
Length = 318
Score = 87.0 bits (214), Expect = 9e-16
Identities = 47/130 (36%), Positives = 72/130 (55%), Gaps = 1/130 (0%)
Query: 277 GRVFALTGTQTENEDRLIIGTCYINNTPLVAIIDIGATHCFIAFDCASTLGLDMSNMNGE 336
GR++ + + LI C I L A+ D GATH FIA C + L L +S + +
Sbjct: 129 GRIYCVNAEEGNPSGNLIQRDCEIAGNTLTALFDSGATHSFIAMGCVNRLKLSVSALPFD 188
Query: 337 MVVETPAKGSVTTSLVCLKSSLSMFGRDFEMDLVCLPLSGMYVILGMNWFE*NHVHINYF 396
+VV TPAK ++T + CL +++ RDF ++L+CLPL + VILGM+W ++V ++
Sbjct: 189 LVVSTPAK-TLTVNSACLHCRMTIQNRDFLVNLICLPLQSLEVILGMDWMSYHYVILDCA 247
Query: 397 SKSVYFSSVE 406
V F E
Sbjct: 248 RTLVLFPEPE 257
>UniRef100_Q7XF39 Similar to Sorghum bicolor 22 kDakafirinclusterpolyprotein [Oryza
sativa]
Length = 1484
Score = 69.7 bits (169), Expect = 1e-10
Identities = 43/147 (29%), Positives = 78/147 (52%), Gaps = 4/147 (2%)
Query: 266 QCKQPKKAP---TIGRVFALTGTQTENEDRLIIGTCYINNTPLVAIIDIGATHCFIAFDC 322
+C++P++A RV + + ++ +I+GT +N+TP V + D GATH FI+
Sbjct: 343 KCQKPRRAGPKFVQARVNHASAKEAQSAPEVILGTFPVNSTPAVILFDSGATHSFISKRF 402
Query: 323 ASTLGLDMSNMNGEMVVETPAKGSVTTSLVCLKSSLSMFGRDFEMDLVCLPLSGMYVILG 382
A GL + + M V TP G +TT+ C ++ + G F +L+ L + VILG
Sbjct: 403 AGAHGLSLVKLKIPMRVHTPG-GGMTTTHYCPSVTVEIQGLIFPANLILLESKDLDVILG 461
Query: 383 MNWFE*NHVHINYFSKSVYFSSVEEEI 409
M+W + I+ S+++ ++ + E+
Sbjct: 462 MDWLTRHRGVIDCASRTIELTNAKGEV 488
>UniRef100_Q6I5H5 Putative polyprotein [Oryza sativa]
Length = 1524
Score = 69.3 bits (168), Expect = 2e-10
Identities = 43/147 (29%), Positives = 77/147 (52%), Gaps = 4/147 (2%)
Query: 266 QCKQPKKAP---TIGRVFALTGTQTENEDRLIIGTCYINNTPLVAIIDIGATHCFIAFDC 322
+C +P++A RV + + ++ +I+GT +N+TP V + D GATH FI+
Sbjct: 370 KCPKPRRAGPKFVQARVNHASAEEAQSAPEVILGTFPVNSTPAVILFDSGATHSFISKRF 429
Query: 323 ASTLGLDMSNMNGEMVVETPAKGSVTTSLVCLKSSLSMFGRDFEMDLVCLPLSGMYVILG 382
A GL + + M V TP G +TT+ C ++ + G F +L+ L + VILG
Sbjct: 430 AGAHGLSLVKLKIPMRVHTPG-GGMTTTHYCPSVTVEIQGSIFPANLILLESKDLDVILG 488
Query: 383 MNWFE*NHVHINYFSKSVYFSSVEEEI 409
M+W + I+ S+++ ++ + E+
Sbjct: 489 MDWLTRHRGVIDCASRTIKLTNAKGEV 515
>UniRef100_Q6ATD6 Putative polyprotein [Oryza sativa]
Length = 1524
Score = 69.3 bits (168), Expect = 2e-10
Identities = 43/147 (29%), Positives = 77/147 (52%), Gaps = 4/147 (2%)
Query: 266 QCKQPKKAP---TIGRVFALTGTQTENEDRLIIGTCYINNTPLVAIIDIGATHCFIAFDC 322
+C +P++A RV + + ++ +I+GT +N+TP V + D GATH FI+
Sbjct: 370 KCPKPRRAGPKFVQARVNHASAEEAQSAPEVILGTFPVNSTPAVILFDSGATHSFISKRF 429
Query: 323 ASTLGLDMSNMNGEMVVETPAKGSVTTSLVCLKSSLSMFGRDFEMDLVCLPLSGMYVILG 382
A GL + + M V TP G +TT+ C ++ + G F +L+ L + VILG
Sbjct: 430 AGAHGLSLVKLKIPMRVHTPG-GGMTTTHYCPSVTVEIQGSIFPANLILLESKDLDVILG 488
Query: 383 MNWFE*NHVHINYFSKSVYFSSVEEEI 409
M+W + I+ S+++ ++ + E+
Sbjct: 489 MDWLTRHRGVIDCASRTIKLTNAKGEV 515
>UniRef100_Q8S7P1 Putative polyprotein [Oryza sativa]
Length = 1524
Score = 68.9 bits (167), Expect = 3e-10
Identities = 43/147 (29%), Positives = 77/147 (52%), Gaps = 4/147 (2%)
Query: 266 QCKQPKKAP---TIGRVFALTGTQTENEDRLIIGTCYINNTPLVAIIDIGATHCFIAFDC 322
+C +P++A RV + + ++ +I+GT +N+TP V + D GATH FI+
Sbjct: 370 KCPKPRRAGPKFVQARVNHASAEEAQSAPEVILGTFPVNSTPAVILFDSGATHSFISKRF 429
Query: 323 ASTLGLDMSNMNGEMVVETPAKGSVTTSLVCLKSSLSMFGRDFEMDLVCLPLSGMYVILG 382
A GL + + M V TP G +TT+ C ++ + G F +L+ L + VILG
Sbjct: 430 AGAHGLSLVKLKIPMRVHTPG-GGMTTTHYCPSVTVEIQGLIFPANLILLESKDLDVILG 488
Query: 383 MNWFE*NHVHINYFSKSVYFSSVEEEI 409
M+W + I+ S+++ ++ + E+
Sbjct: 489 MDWLTRHRGVIDCASRTIKLTNAKGEV 515
>UniRef100_Q8S6E1 Putative polyprotein [Oryza sativa]
Length = 1229
Score = 68.9 bits (167), Expect = 3e-10
Identities = 43/147 (29%), Positives = 77/147 (52%), Gaps = 4/147 (2%)
Query: 266 QCKQPKKAP---TIGRVFALTGTQTENEDRLIIGTCYINNTPLVAIIDIGATHCFIAFDC 322
+C +P++A RV + + ++ +I+GT +N+TP V + D GATH FI+
Sbjct: 75 KCPKPRRAGPKFVQARVNHASAEEAQSAPEVILGTFPVNSTPAVILFDSGATHSFISKRF 134
Query: 323 ASTLGLDMSNMNGEMVVETPAKGSVTTSLVCLKSSLSMFGRDFEMDLVCLPLSGMYVILG 382
A GL + + M V TP G +TT+ C ++ + G F +L+ L + VILG
Sbjct: 135 AGAHGLSLVKLKIPMRVHTPG-GGMTTTHYCPSVTVEIQGLIFPANLILLESKDLDVILG 193
Query: 383 MNWFE*NHVHINYFSKSVYFSSVEEEI 409
M+W + I+ S+++ ++ + E+
Sbjct: 194 MDWLARHRGVIDCASRTIKLTNAKGEV 220
>UniRef100_Q8LMN2 Putative polyprotein [Oryza sativa]
Length = 1413
Score = 68.9 bits (167), Expect = 3e-10
Identities = 43/147 (29%), Positives = 77/147 (52%), Gaps = 4/147 (2%)
Query: 266 QCKQPKKAP---TIGRVFALTGTQTENEDRLIIGTCYINNTPLVAIIDIGATHCFIAFDC 322
+C +P++A RV + + ++ +I+GT +N+TP V + D GATH FI+
Sbjct: 342 KCPKPRRAGPKFVQARVNHASAEEAQSAPEVILGTFPVNSTPAVILFDSGATHSFISKRF 401
Query: 323 ASTLGLDMSNMNGEMVVETPAKGSVTTSLVCLKSSLSMFGRDFEMDLVCLPLSGMYVILG 382
A GL + + M V TP G +TT+ C ++ + G F +L+ L + VILG
Sbjct: 402 AGAHGLSLVKLKIPMRVHTPG-GGMTTTHYCPSVTVEIQGLIFPANLILLESKDLDVILG 460
Query: 383 MNWFE*NHVHINYFSKSVYFSSVEEEI 409
M+W + I+ S+++ ++ + E+
Sbjct: 461 MDWLARHRGVIDCASRTIKLTNAKGEV 487
>UniRef100_Q7F2E9 Putative polyprotein [Oryza sativa]
Length = 1524
Score = 68.9 bits (167), Expect = 3e-10
Identities = 43/147 (29%), Positives = 77/147 (52%), Gaps = 4/147 (2%)
Query: 266 QCKQPKKAP---TIGRVFALTGTQTENEDRLIIGTCYINNTPLVAIIDIGATHCFIAFDC 322
+C +P++A RV + + ++ +I+GT +N+TP V + D GATH FI+
Sbjct: 370 KCPKPRRAGPKFVQARVNHASAEEAQSAPEVILGTFPVNSTPAVILFDSGATHSFISKRF 429
Query: 323 ASTLGLDMSNMNGEMVVETPAKGSVTTSLVCLKSSLSMFGRDFEMDLVCLPLSGMYVILG 382
A GL + + M V TP G +TT+ C ++ + G F +L+ L + VILG
Sbjct: 430 AGAHGLSLVKLKIPMRVHTPG-GGMTTTHYCPSVTVEIQGLIFPANLILLESKDLDVILG 488
Query: 383 MNWFE*NHVHINYFSKSVYFSSVEEEI 409
M+W + I+ S+++ ++ + E+
Sbjct: 489 MDWLTRHRGVIDCASRTIKLTNAKGEV 515
>UniRef100_Q6ASR4 Putative polyprotein [Oryza sativa]
Length = 1500
Score = 68.9 bits (167), Expect = 3e-10
Identities = 43/147 (29%), Positives = 77/147 (52%), Gaps = 4/147 (2%)
Query: 266 QCKQPKKAP---TIGRVFALTGTQTENEDRLIIGTCYINNTPLVAIIDIGATHCFIAFDC 322
+C +P++A RV + + ++ +I+GT +N+TP V + D GATH FI+
Sbjct: 346 KCPKPRRAGPKFVQARVNHASAEEAQSAPEVILGTFPVNSTPAVILFDSGATHSFISKRF 405
Query: 323 ASTLGLDMSNMNGEMVVETPAKGSVTTSLVCLKSSLSMFGRDFEMDLVCLPLSGMYVILG 382
A GL + + M V TP G +TT+ C ++ + G F +L+ L + VILG
Sbjct: 406 AGAHGLSLVKLKIPMRVHTPG-GGMTTTHYCPSVTVEIQGLIFPANLILLESKDLDVILG 464
Query: 383 MNWFE*NHVHINYFSKSVYFSSVEEEI 409
M+W + I+ S+++ ++ + E+
Sbjct: 465 MDWLTRHRGVIDCASRTIKLTNAKGEV 491
>UniRef100_Q7XPN0 OSJNBa0060D06.14 protein [Oryza sativa]
Length = 1458
Score = 68.9 bits (167), Expect = 3e-10
Identities = 43/147 (29%), Positives = 77/147 (52%), Gaps = 4/147 (2%)
Query: 266 QCKQPKKAP---TIGRVFALTGTQTENEDRLIIGTCYINNTPLVAIIDIGATHCFIAFDC 322
+C +P++A RV + + ++ +I+GT +N+TP V + D GATH FI+
Sbjct: 370 KCPKPRRAGPKFVQARVNHASAEEAQSAPEVILGTFPVNSTPAVILFDSGATHSFISKRF 429
Query: 323 ASTLGLDMSNMNGEMVVETPAKGSVTTSLVCLKSSLSMFGRDFEMDLVCLPLSGMYVILG 382
A GL + + M V TP G +TT+ C ++ + G F +L+ L + VILG
Sbjct: 430 AGAHGLSLVKLKIPMRVHTPG-GGMTTTHYCPSVTVEIQGLIFPANLILLESKDLDVILG 488
Query: 383 MNWFE*NHVHINYFSKSVYFSSVEEEI 409
M+W + I+ S+++ ++ + E+
Sbjct: 489 MDWLARHRGVIDCASRTIKLTNAKGEV 515
>UniRef100_Q6F330 Putative polyprotein [Oryza sativa]
Length = 1458
Score = 68.9 bits (167), Expect = 3e-10
Identities = 43/147 (29%), Positives = 77/147 (52%), Gaps = 4/147 (2%)
Query: 266 QCKQPKKAP---TIGRVFALTGTQTENEDRLIIGTCYINNTPLVAIIDIGATHCFIAFDC 322
+C +P++A RV + + ++ +I+GT +N+TP V + D GATH FI+
Sbjct: 370 KCPKPRRAGPKFVQARVNHASAEEAQSAPEVILGTFPVNSTPAVILFDSGATHSFISKRF 429
Query: 323 ASTLGLDMSNMNGEMVVETPAKGSVTTSLVCLKSSLSMFGRDFEMDLVCLPLSGMYVILG 382
A GL + + M V TP G +TT+ C ++ + G F +L+ L + VILG
Sbjct: 430 AGAHGLSLVKLKIPMRVHTPG-GGMTTTHYCPSVTVEIQGLIFPANLILLESKDLDVILG 488
Query: 383 MNWFE*NHVHINYFSKSVYFSSVEEEI 409
M+W + I+ S+++ ++ + E+
Sbjct: 489 MDWLTRHRGVIDCASRTIKLTNAKGEV 515
>UniRef100_Q6F300 Putative polyprotein [Oryza sativa]
Length = 1524
Score = 68.9 bits (167), Expect = 3e-10
Identities = 43/147 (29%), Positives = 77/147 (52%), Gaps = 4/147 (2%)
Query: 266 QCKQPKKAP---TIGRVFALTGTQTENEDRLIIGTCYINNTPLVAIIDIGATHCFIAFDC 322
+C +P++A RV + + ++ +I+GT +N+TP V + D GATH FI+
Sbjct: 370 KCPKPRRAGPKFVQARVNHASAEEAQSAPEVILGTFPVNSTPAVILFDSGATHSFISKRF 429
Query: 323 ASTLGLDMSNMNGEMVVETPAKGSVTTSLVCLKSSLSMFGRDFEMDLVCLPLSGMYVILG 382
A GL + + M V TP G +TT+ C ++ + G F +L+ L + VILG
Sbjct: 430 AGAHGLSLVKLKIPMRVHTPG-GGMTTTHYCPSVTVEIQGLIFPANLILLESKDLDVILG 488
Query: 383 MNWFE*NHVHINYFSKSVYFSSVEEEI 409
M+W + I+ S+++ ++ + E+
Sbjct: 489 MDWLARHRGVIDCASRTIKLTNAKGEV 515
>UniRef100_Q8W5F5 Putative polyprotein [Oryza sativa]
Length = 1524
Score = 68.9 bits (167), Expect = 3e-10
Identities = 43/147 (29%), Positives = 77/147 (52%), Gaps = 4/147 (2%)
Query: 266 QCKQPKKAP---TIGRVFALTGTQTENEDRLIIGTCYINNTPLVAIIDIGATHCFIAFDC 322
+C +P++A RV + + ++ +I+GT +N+TP V + D GATH FI+
Sbjct: 370 KCPKPRRAGPKFVQARVNHASAEEAQSAPEVILGTFPVNSTPAVILFDSGATHSFISKRF 429
Query: 323 ASTLGLDMSNMNGEMVVETPAKGSVTTSLVCLKSSLSMFGRDFEMDLVCLPLSGMYVILG 382
A GL + + M V TP G +TT+ C ++ + G F +L+ L + VILG
Sbjct: 430 AGAHGLSLVKLKIPMRVHTPG-GGMTTTHYCPSVTVEIQGLIFPANLILLESKDLDVILG 488
Query: 383 MNWFE*NHVHINYFSKSVYFSSVEEEI 409
M+W + I+ S+++ ++ + E+
Sbjct: 489 MDWLTRHRGVIDCASRTIKLTNAKGEV 515
>UniRef100_Q8S9N7 Similar to polyprotein [Oryza sativa]
Length = 1524
Score = 68.9 bits (167), Expect = 3e-10
Identities = 43/147 (29%), Positives = 77/147 (52%), Gaps = 4/147 (2%)
Query: 266 QCKQPKKAP---TIGRVFALTGTQTENEDRLIIGTCYINNTPLVAIIDIGATHCFIAFDC 322
+C +P++A RV + + ++ +I+GT +N+TP V + D GATH FI+
Sbjct: 370 KCPKPRRAGPKFVQARVNHASAEEAQSAPEVILGTFPVNSTPAVILFDSGATHSFISKRF 429
Query: 323 ASTLGLDMSNMNGEMVVETPAKGSVTTSLVCLKSSLSMFGRDFEMDLVCLPLSGMYVILG 382
A GL + + M V TP G +TT+ C ++ + G F +L+ L + VILG
Sbjct: 430 AGAHGLSLVKLKIPMRVHTPG-GGMTTTHYCPSVTVEIQGLIFPANLILLESKDLDVILG 488
Query: 383 MNWFE*NHVHINYFSKSVYFSSVEEEI 409
M+W + I+ S+++ ++ + E+
Sbjct: 489 MDWLARHRGVIDCASRTIKLTNAKGEV 515
>UniRef100_Q8RYQ6 Putative Sorghum bicolor 22 kDa kafirin cluster polyprotein [Oryza
sativa]
Length = 1524
Score = 68.9 bits (167), Expect = 3e-10
Identities = 43/147 (29%), Positives = 77/147 (52%), Gaps = 4/147 (2%)
Query: 266 QCKQPKKAP---TIGRVFALTGTQTENEDRLIIGTCYINNTPLVAIIDIGATHCFIAFDC 322
+C +P++A RV + + ++ +I+GT +N+TP V + D GATH FI+
Sbjct: 370 KCPKPRRAGPKFVQARVNHASAEEAQSAPEVILGTFPVNSTPAVILFDSGATHSFISKRF 429
Query: 323 ASTLGLDMSNMNGEMVVETPAKGSVTTSLVCLKSSLSMFGRDFEMDLVCLPLSGMYVILG 382
A GL + + M V TP G +TT+ C ++ + G F +L+ L + VILG
Sbjct: 430 AGAHGLSLVKLKIPMRVHTPG-GGMTTTHYCPSVTVEIQGLIFPANLILLESKDLDVILG 488
Query: 383 MNWFE*NHVHINYFSKSVYFSSVEEEI 409
M+W + I+ S+++ ++ + E+
Sbjct: 489 MDWLARHRGVIDCASRTIKLTNAKGEV 515
>UniRef100_Q9XE85 Polyprotein [Sorghum bicolor]
Length = 1484
Score = 68.2 bits (165), Expect = 4e-10
Identities = 41/143 (28%), Positives = 72/143 (49%), Gaps = 1/143 (0%)
Query: 266 QCKQPKKAPTIGRVFALTGTQTENEDRLIIGTCYINNTPLVAIIDIGATHCFIAFDCAST 325
Q + P +V + + +++GT IN+ P + D GA+H FI+
Sbjct: 357 QYNDRRSTPNSAKVNNVAAETVQEGPEIMMGTFSINSIPATVLFDSGASHTFISQAFVRV 416
Query: 326 LGLDMSNMNGEMVVETPAKGSVTTSLVCLKSSLSMFGRDFEMDLVCLPLSGMYVILGMNW 385
L + MN M+V +P G++ SL C +SLS+ G DF + + + SG+ VILG++W
Sbjct: 417 HSLPLVAMNTPMLVNSPG-GTIPVSLRCPSASLSLRGVDFPISPMVMRTSGIDVILGLDW 475
Query: 386 FE*NHVHINYFSKSVYFSSVEEE 408
+ + +IN + V ++ + E
Sbjct: 476 MKQHATNINCKERVVVLTTPKGE 498
>UniRef100_Q852G2 Putative polyprotein [Oryza sativa]
Length = 1079
Score = 68.2 bits (165), Expect = 4e-10
Identities = 43/147 (29%), Positives = 77/147 (52%), Gaps = 4/147 (2%)
Query: 266 QCKQPKKAP---TIGRVFALTGTQTENEDRLIIGTCYINNTPLVAIIDIGATHCFIAFDC 322
+C +P++A RV + + ++ +I+GT +N+TP V + D GATH FI+
Sbjct: 370 KCSKPRRAGPKFVQARVNHASAEEAQSAPEVILGTFPVNSTPAVILFDSGATHSFISKLF 429
Query: 323 ASTLGLDMSNMNGEMVVETPAKGSVTTSLVCLKSSLSMFGRDFEMDLVCLPLSGMYVILG 382
A GL + + M V TP G +TT+ C ++ + G F +L+ L + VILG
Sbjct: 430 AGAHGLSLVKLKIPMRVHTPG-GGMTTTHYCPAVTVEIQGLIFPANLILLESKDLDVILG 488
Query: 383 MNWFE*NHVHINYFSKSVYFSSVEEEI 409
M+W + I+ S+++ ++ + E+
Sbjct: 489 MDWLTRHRGVIDCASRTIKLTNAKGEV 515
>UniRef100_Q943A1 Putative polyprotein [Oryza sativa]
Length = 1524
Score = 67.4 bits (163), Expect = 7e-10
Identities = 42/147 (28%), Positives = 76/147 (51%), Gaps = 4/147 (2%)
Query: 266 QCKQPKKAP---TIGRVFALTGTQTENEDRLIIGTCYINNTPLVAIIDIGATHCFIAFDC 322
+C +P++A RV + + ++ +I+GT +N+TP + D GATH FI+
Sbjct: 370 KCSKPRRAGPKFVQARVNHASAEEAQSAPEVILGTFPVNSTPAKILFDSGATHSFISKRF 429
Query: 323 ASTLGLDMSNMNGEMVVETPAKGSVTTSLVCLKSSLSMFGRDFEMDLVCLPLSGMYVILG 382
A GL + + M V TP G +TT+ C ++ + G F +L+ L + VILG
Sbjct: 430 AGAHGLSLVKLKIPMRVHTPG-GGMTTTHYCPSVTVEIQGSIFPANLILLESKDLDVILG 488
Query: 383 MNWFE*NHVHINYFSKSVYFSSVEEEI 409
M+W + I+ S+++ ++ + E+
Sbjct: 489 MDWLTRHRGVIDCASRTIKLTNAKGEV 515
>UniRef100_Q5TKK3 Hypothetical protein OJ1532_D06.4 [Oryza sativa]
Length = 1238
Score = 67.4 bits (163), Expect = 7e-10
Identities = 43/144 (29%), Positives = 74/144 (50%), Gaps = 3/144 (2%)
Query: 266 QCKQPKKAPTIGRVFALTGTQTENEDRLIIGTCYINNTPLVAIIDIGATHCFIAFDCAST 325
+C PK RV + + ++ +I+GT +N+TP V + D GATH FI+ A
Sbjct: 89 RCAGPKFVQA--RVNHASAEEAQSAPEVILGTFPVNSTPAVILFDSGATHSFISKRFAGA 146
Query: 326 LGLDMSNMNGEMVVETPAKGSVTTSLVCLKSSLSMFGRDFEMDLVCLPLSGMYVILGMNW 385
GL + + M V TP G +TT+ C ++ + G F +L+ L + VILGM+W
Sbjct: 147 HGLSLVKLKIPMRVHTPG-GGMTTTHYCPSVTVEIQGLIFPANLILLESKDLDVILGMDW 205
Query: 386 FE*NHVHINYFSKSVYFSSVEEEI 409
+ I+ S+++ ++ + E+
Sbjct: 206 LTRHRGVIDCASRTIKLTNAKGEV 229
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.343 0.151 0.478
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 580,679,777
Number of Sequences: 2790947
Number of extensions: 21079990
Number of successful extensions: 70467
Number of sequences better than 10.0: 352
Number of HSP's better than 10.0 without gapping: 289
Number of HSP's successfully gapped in prelim test: 63
Number of HSP's that attempted gapping in prelim test: 69846
Number of HSP's gapped (non-prelim): 364
length of query: 427
length of database: 848,049,833
effective HSP length: 130
effective length of query: 297
effective length of database: 485,226,723
effective search space: 144112336731
effective search space used: 144112336731
T: 11
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.5 bits)
S2: 76 (33.9 bits)
Medicago: description of AC147363.1


