

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147178.10 + phase: 0
(161 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
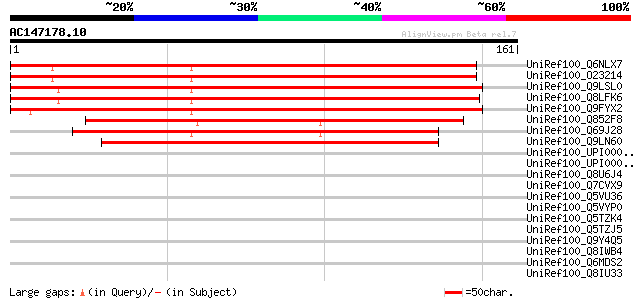
Sequences producing significant alignments: (bits) Value
UniRef100_Q6NLX7 At4g36660 [Arabidopsis thaliana] 226 2e-58
UniRef100_O23214 Hypothetical protein AT4g36660 [Arabidopsis tha... 226 2e-58
UniRef100_Q9LSL0 Emb|CAB16809.1 [Arabidopsis thaliana] 223 2e-57
UniRef100_Q8LFK6 Hypothetical protein At5g65650 [Arabidopsis tha... 221 5e-57
UniRef100_Q9FYX2 BAC19.4 [Lycopersicon esculentum] 205 3e-52
UniRef100_Q852F8 Hypothetical protein OJ1212_C05.9 [Oryza sativa] 147 9e-35
UniRef100_Q69J28 Hypothetical protein OSJNBa0072I06.26 [Oryza sa... 145 3e-34
UniRef100_Q9LN60 F18O14.10 [Arabidopsis thaliana] 120 1e-26
UniRef100_UPI0000457745 UPI0000457745 UniRef100 entry 34 1.1
UniRef100_UPI0000432C8B UPI0000432C8B UniRef100 entry 34 1.1
UniRef100_Q8U6J4 Hypothetical protein bme3 [Agrobacterium tumefa... 34 1.1
UniRef100_Q7CVX9 AGR_L_143p [Agrobacterium tumefaciens] 34 1.1
UniRef100_Q5VU36 Novel protein [Homo sapiens] 34 1.1
UniRef100_Q5VYP0 Novel protein [Homo sapiens] 34 1.1
UniRef100_Q5TZK4 Novel protein [Homo sapiens] 34 1.1
UniRef100_Q5TZJ5 OTTHUMP00000046132 [Homo sapiens] 34 1.1
UniRef100_Q9Y4Q5 Hypothetical protein DKFZp434B204 [Homo sapiens] 34 1.1
UniRef100_Q8IWB4 Chromosome 9 open reading frame 36 [Homo sapiens] 34 1.1
UniRef100_Q6MDS2 Hypothetical protein [Parachlamydia sp.] 34 1.4
UniRef100_Q8IU33 HrTRPV [Halocynthia roretzi] 34 1.4
>UniRef100_Q6NLX7 At4g36660 [Arabidopsis thaliana]
Length = 179
Score = 226 bits (576), Expect = 2e-58
Identities = 117/158 (74%), Positives = 129/158 (81%), Gaps = 10/158 (6%)
Query: 1 MKDDDGLPTTTA---------AINKKENMDSGLFGKGRYKFWALAAILLLAFWSMFTGTV 51
MK+DD LPTTT A +KKE+ DS LFG+GRYKFWA AAILLLAFWSMFTGTV
Sbjct: 1 MKEDDALPTTTTTGTATGTAMANSKKESSDSVLFGRGRYKFWAFAAILLLAFWSMFTGTV 60
Query: 52 SLRWS-GNLNTLSSDLDTPIHDDLDVLEMEEREKVVRHMWDVYTNSRRIRLPRFWQEAFE 110
+LR S GNLN LS DL P +D+LDVLEMEEREKVV+HMWDVYTNSRRI+LPRFWQEAF
Sbjct: 61 TLRLSTGNLNRLSEDLGIPNYDNLDVLEMEEREKVVKHMWDVYTNSRRIKLPRFWQEAFV 120
Query: 111 AAYEELTSDVPGVRDDAITEIAKMSVRSVNYDPPPIQS 148
AAYEELTSDVPGVR+ AI EIAKMS RS+ DPPP +S
Sbjct: 121 AAYEELTSDVPGVREAAIGEIAKMSARSITLDPPPSRS 158
>UniRef100_O23214 Hypothetical protein AT4g36660 [Arabidopsis thaliana]
Length = 159
Score = 226 bits (576), Expect = 2e-58
Identities = 117/158 (74%), Positives = 129/158 (81%), Gaps = 10/158 (6%)
Query: 1 MKDDDGLPTTTA---------AINKKENMDSGLFGKGRYKFWALAAILLLAFWSMFTGTV 51
MK+DD LPTTT A +KKE+ DS LFG+GRYKFWA AAILLLAFWSMFTGTV
Sbjct: 1 MKEDDALPTTTTTGTATGTAMANSKKESSDSVLFGRGRYKFWAFAAILLLAFWSMFTGTV 60
Query: 52 SLRWS-GNLNTLSSDLDTPIHDDLDVLEMEEREKVVRHMWDVYTNSRRIRLPRFWQEAFE 110
+LR S GNLN LS DL P +D+LDVLEMEEREKVV+HMWDVYTNSRRI+LPRFWQEAF
Sbjct: 61 TLRLSTGNLNRLSEDLGIPNYDNLDVLEMEEREKVVKHMWDVYTNSRRIKLPRFWQEAFV 120
Query: 111 AAYEELTSDVPGVRDDAITEIAKMSVRSVNYDPPPIQS 148
AAYEELTSDVPGVR+ AI EIAKMS RS+ DPPP +S
Sbjct: 121 AAYEELTSDVPGVREAAIGEIAKMSARSITLDPPPSRS 158
>UniRef100_Q9LSL0 Emb|CAB16809.1 [Arabidopsis thaliana]
Length = 159
Score = 223 bits (567), Expect = 2e-57
Identities = 112/157 (71%), Positives = 127/157 (80%), Gaps = 7/157 (4%)
Query: 1 MKDDDGLPTTTAAI------NKKENMDSGLFGKGRYKFWALAAILLLAFWSMFTGTVSLR 54
MKD D LP +T+++ KKE S LF KGRYKFWALAAILLLAFWSM TGTV+LR
Sbjct: 1 MKDGDSLPISTSSVAATTVTGKKETGYSALFSKGRYKFWALAAILLLAFWSMLTGTVNLR 60
Query: 55 WS-GNLNTLSSDLDTPIHDDLDVLEMEEREKVVRHMWDVYTNSRRIRLPRFWQEAFEAAY 113
WS GN+N + DL PIH+DLDVLEMEEREKVV+HMWDVY N RRIRLPRFWQEAFEAAY
Sbjct: 61 WSAGNINHFTDDLVFPIHEDLDVLEMEEREKVVKHMWDVYNNGRRIRLPRFWQEAFEAAY 120
Query: 114 EELTSDVPGVRDDAITEIAKMSVRSVNYDPPPIQSTV 150
EELTSDVP V + AI+EIA+MS+RS+ DPPP+ STV
Sbjct: 121 EELTSDVPDVVEAAISEIARMSIRSIVIDPPPLHSTV 157
>UniRef100_Q8LFK6 Hypothetical protein At5g65650 [Arabidopsis thaliana]
Length = 185
Score = 221 bits (563), Expect = 5e-57
Identities = 111/156 (71%), Positives = 126/156 (80%), Gaps = 7/156 (4%)
Query: 1 MKDDDGLPTTTAAI------NKKENMDSGLFGKGRYKFWALAAILLLAFWSMFTGTVSLR 54
MKD D LP +T+++ KKE S LF KGRYKFWALAAILLLAFWSM TGTV+LR
Sbjct: 1 MKDGDSLPISTSSVAATTVTGKKETGYSALFSKGRYKFWALAAILLLAFWSMLTGTVNLR 60
Query: 55 WS-GNLNTLSSDLDTPIHDDLDVLEMEEREKVVRHMWDVYTNSRRIRLPRFWQEAFEAAY 113
WS GN+N + DL PIH+DLDVLEMEEREKVV+HMWDVY N RRIRLPRFWQEAFEAAY
Sbjct: 61 WSAGNINHFTDDLVFPIHEDLDVLEMEEREKVVKHMWDVYNNGRRIRLPRFWQEAFEAAY 120
Query: 114 EELTSDVPGVRDDAITEIAKMSVRSVNYDPPPIQST 149
EELTSDVP V + AI+EIA+MS+RS+ DPPP+ ST
Sbjct: 121 EELTSDVPDVVEAAISEIARMSIRSIVIDPPPLHST 156
>UniRef100_Q9FYX2 BAC19.4 [Lycopersicon esculentum]
Length = 182
Score = 205 bits (522), Expect = 3e-52
Identities = 101/157 (64%), Positives = 124/157 (78%), Gaps = 7/157 (4%)
Query: 1 MKDDD------GLPTTTAAINKKENMDSGLFGKGRYKFWALAAILLLAFWSMFTGTVSLR 54
M+DDD P+++ +KKE +FG+GRYKFWA AAI LLA WSMFTGTV+LR
Sbjct: 1 MRDDDLPISTPTAPSSSFTTSKKETSYFAIFGRGRYKFWAFAAISLLALWSMFTGTVTLR 60
Query: 55 WS-GNLNTLSSDLDTPIHDDLDVLEMEEREKVVRHMWDVYTNSRRIRLPRFWQEAFEAAY 113
WS GNLN +S D+D P+ +DLDVLEMEEREK+V+HMWDVYTNSR I+L +FWQEAFEAAY
Sbjct: 61 WSAGNLNAISDDIDIPLPEDLDVLEMEEREKLVKHMWDVYTNSRGIKLLKFWQEAFEAAY 120
Query: 114 EELTSDVPGVRDDAITEIAKMSVRSVNYDPPPIQSTV 150
EEL SDVPGV +DA++EIAKMSVR + + PP+ S+V
Sbjct: 121 EELASDVPGVPEDALSEIAKMSVRYIPIESPPLHSSV 157
>UniRef100_Q852F8 Hypothetical protein OJ1212_C05.9 [Oryza sativa]
Length = 178
Score = 147 bits (371), Expect = 9e-35
Identities = 77/125 (61%), Positives = 87/125 (69%), Gaps = 5/125 (4%)
Query: 25 FGKGRYKFWALAAILLLAFWSMFTGTVSLRWSGN----LNTLSSDLDTPIHDDLDVLEME 80
FGKGRYK WALAAI LLA WSM + SLRWS T S DLD P+ DDLD LEME
Sbjct: 35 FGKGRYKVWALAAIALLALWSMSAASASLRWSSGRFLLAATASEDLDAPLLDDLDSLEME 94
Query: 81 EREKVVRHMWDVYTNSR-RIRLPRFWQEAFEAAYEELTSDVPGVRDDAITEIAKMSVRSV 139
EREK+V MWD+YT + +RLPRFWQEAFEAAYEEL D VRD AI+EIA+MS +
Sbjct: 95 EREKLVGRMWDMYTRTGDEVRLPRFWQEAFEAAYEELAGDDMQVRDAAISEIARMSAHRL 154
Query: 140 NYDPP 144
+ P
Sbjct: 155 ELEQP 159
>UniRef100_Q69J28 Hypothetical protein OSJNBa0072I06.26 [Oryza sativa]
Length = 164
Score = 145 bits (367), Expect = 3e-34
Identities = 74/119 (62%), Positives = 89/119 (74%), Gaps = 3/119 (2%)
Query: 21 DSGLFGKGRYKFWALAAILLLAFWSMFTGTVSLRWS-GNLNTLSSDLDTPIHDDLDVLEM 79
++ L GKGRYK WALAAI LLA WSMF +V++RWS G+L DL P+ DDLD LEM
Sbjct: 34 EAALLGKGRYKAWALAAIALLALWSMFAASVTIRWSSGDLAAEFGDLPDPLIDDLDPLEM 93
Query: 80 EEREKVVRHMWDVYTNSR--RIRLPRFWQEAFEAAYEELTSDVPGVRDDAITEIAKMSV 136
E+REK+VR MWDVYT + R+RLPRFWQEAFEAAYEEL D + A++EIA+MSV
Sbjct: 94 EDREKLVRRMWDVYTRTGVDRVRLPRFWQEAFEAAYEELAGDDTQASETAVSEIARMSV 152
>UniRef100_Q9LN60 F18O14.10 [Arabidopsis thaliana]
Length = 147
Score = 120 bits (301), Expect = 1e-26
Identities = 61/107 (57%), Positives = 77/107 (71%)
Query: 30 YKFWALAAILLLAFWSMFTGTVSLRWSGNLNTLSSDLDTPIHDDLDVLEMEEREKVVRHM 89
YK W L A+LLLAF SM TG+VSL+ G ++ DDLDVLE+EEREKVVR M
Sbjct: 25 YKLWVLIAVLLLAFGSMLTGSVSLKGIGLFHSADGVNAFSFGDDLDVLEIEEREKVVRQM 84
Query: 90 WDVYTNSRRIRLPRFWQEAFEAAYEELTSDVPGVRDDAITEIAKMSV 136
WDVY S +++PRFW+EAFEAAYE L SD VR+ A+++IAK+S+
Sbjct: 85 WDVYGRSGGVKVPRFWREAFEAAYEFLISDSAAVRNAAVSDIAKLSL 131
>UniRef100_UPI0000457745 UPI0000457745 UniRef100 entry
Length = 474
Score = 34.3 bits (77), Expect = 1.1
Identities = 19/75 (25%), Positives = 36/75 (47%), Gaps = 6/75 (8%)
Query: 77 LEMEEREKVVRHMWDVYTNSR------RIRLPRFWQEAFEAAYEELTSDVPGVRDDAITE 130
L+ E+ + W++ NS+ ++ PR WQE+F Y +L +P + +++
Sbjct: 376 LDAEQDTTNPKPFWNMGENSKQLPGPQKLSDPRLWQESFWKNYSQLFWGLPSLHSESLVA 435
Query: 131 IAKMSVRSVNYDPPP 145
A ++ RS PP
Sbjct: 436 NAWVTDRSYTLQSPP 450
>UniRef100_UPI0000432C8B UPI0000432C8B UniRef100 entry
Length = 316
Score = 34.3 bits (77), Expect = 1.1
Identities = 19/69 (27%), Positives = 41/69 (58%), Gaps = 8/69 (11%)
Query: 83 EKVVRHMWDVYTNSRRI--RLPRFWQEAFEAAYEELTSDVPGVRDDAITEIAKMSVRSVN 140
E +V+ +WD Y N+ R+ R+ R W+E F+ ++L +++ +RD+ + + + ++N
Sbjct: 35 ETMVKTIWDTYMNNTRLADRIIR-WRELFQNLLDKLVAEIKLLRDEKVD--TEKEIETLN 91
Query: 141 YDPPPIQST 149
+ P+Q T
Sbjct: 92 F---PLQLT 97
>UniRef100_Q8U6J4 Hypothetical protein bme3 [Agrobacterium tumefaciens]
Length = 405
Score = 34.3 bits (77), Expect = 1.1
Identities = 20/68 (29%), Positives = 36/68 (52%), Gaps = 3/68 (4%)
Query: 30 YKFWALAAILLLAFWSMFTGTVSLRWSGNLNTLSSDLDTPIHDDLDVLE-MEEREKVVRH 88
YK WA+ I ++ ++F GT+S+ G+ ++ D P+ DD+ ++ + EKV R
Sbjct: 179 YKGWAIPRIFVII--ALFLGTLSMNSMGDYRAITKANDAPVWDDIKRIDFVGNLEKVFRD 236
Query: 89 MWDVYTNS 96
D N+
Sbjct: 237 GGDEMRNA 244
>UniRef100_Q7CVX9 AGR_L_143p [Agrobacterium tumefaciens]
Length = 425
Score = 34.3 bits (77), Expect = 1.1
Identities = 20/68 (29%), Positives = 36/68 (52%), Gaps = 3/68 (4%)
Query: 30 YKFWALAAILLLAFWSMFTGTVSLRWSGNLNTLSSDLDTPIHDDLDVLE-MEEREKVVRH 88
YK WA+ I ++ ++F GT+S+ G+ ++ D P+ DD+ ++ + EKV R
Sbjct: 199 YKGWAIPRIFVII--ALFLGTLSMNSMGDYRAITKANDAPVWDDIKRIDFVGNLEKVFRD 256
Query: 89 MWDVYTNS 96
D N+
Sbjct: 257 GGDEMRNA 264
>UniRef100_Q5VU36 Novel protein [Homo sapiens]
Length = 1347
Score = 34.3 bits (77), Expect = 1.1
Identities = 19/75 (25%), Positives = 36/75 (47%), Gaps = 6/75 (8%)
Query: 77 LEMEEREKVVRHMWDVYTNSR------RIRLPRFWQEAFEAAYEELTSDVPGVRDDAITE 130
L+ E+ + W++ NS+ ++ PR WQE+F Y +L +P + +++
Sbjct: 370 LDAEQDTTNPKPFWNMGENSKQLPGPQKLSDPRLWQESFWKNYSQLFWGLPSLHSESLVA 429
Query: 131 IAKMSVRSVNYDPPP 145
A ++ RS PP
Sbjct: 430 NAWVTDRSYTLQSPP 444
>UniRef100_Q5VYP0 Novel protein [Homo sapiens]
Length = 1347
Score = 34.3 bits (77), Expect = 1.1
Identities = 19/75 (25%), Positives = 36/75 (47%), Gaps = 6/75 (8%)
Query: 77 LEMEEREKVVRHMWDVYTNSR------RIRLPRFWQEAFEAAYEELTSDVPGVRDDAITE 130
L+ E+ + W++ NS+ ++ PR WQE+F Y +L +P + +++
Sbjct: 370 LDAEQDTTNPKPFWNMGENSKQLPGPQKLSDPRLWQESFWKNYSQLFWGLPSLHSESLVA 429
Query: 131 IAKMSVRSVNYDPPP 145
A ++ RS PP
Sbjct: 430 NAWVTDRSYTLQSPP 444
>UniRef100_Q5TZK4 Novel protein [Homo sapiens]
Length = 1347
Score = 34.3 bits (77), Expect = 1.1
Identities = 19/75 (25%), Positives = 36/75 (47%), Gaps = 6/75 (8%)
Query: 77 LEMEEREKVVRHMWDVYTNSR------RIRLPRFWQEAFEAAYEELTSDVPGVRDDAITE 130
L+ E+ + W++ NS+ ++ PR WQE+F Y +L +P + +++
Sbjct: 370 LDAEQDTTNPKPFWNMGENSKQLPGPQKLSDPRLWQESFWKNYSQLFWGLPSLHSESLVA 429
Query: 131 IAKMSVRSVNYDPPP 145
A ++ RS PP
Sbjct: 430 NAWVTDRSYTLQSPP 444
>UniRef100_Q5TZJ5 OTTHUMP00000046132 [Homo sapiens]
Length = 1347
Score = 34.3 bits (77), Expect = 1.1
Identities = 19/75 (25%), Positives = 36/75 (47%), Gaps = 6/75 (8%)
Query: 77 LEMEEREKVVRHMWDVYTNSR------RIRLPRFWQEAFEAAYEELTSDVPGVRDDAITE 130
L+ E+ + W++ NS+ ++ PR WQE+F Y +L +P + +++
Sbjct: 370 LDAEQDTTNPKPFWNMGENSKQLPGPQKLSDPRLWQESFWKNYSQLFWGLPSLHSESLVA 429
Query: 131 IAKMSVRSVNYDPPP 145
A ++ RS PP
Sbjct: 430 NAWVTDRSYTLQSPP 444
>UniRef100_Q9Y4Q5 Hypothetical protein DKFZp434B204 [Homo sapiens]
Length = 1008
Score = 34.3 bits (77), Expect = 1.1
Identities = 19/75 (25%), Positives = 36/75 (47%), Gaps = 6/75 (8%)
Query: 77 LEMEEREKVVRHMWDVYTNSR------RIRLPRFWQEAFEAAYEELTSDVPGVRDDAITE 130
L+ E+ + W++ NS+ ++ PR WQE+F Y +L +P + +++
Sbjct: 31 LDAEQDTTNPKPFWNMGENSKQLPGPQKLSDPRLWQESFWKNYSQLFWGLPSLHSESLVA 90
Query: 131 IAKMSVRSVNYDPPP 145
A ++ RS PP
Sbjct: 91 NAWVTDRSYTLQSPP 105
>UniRef100_Q8IWB4 Chromosome 9 open reading frame 36 [Homo sapiens]
Length = 1272
Score = 34.3 bits (77), Expect = 1.1
Identities = 19/75 (25%), Positives = 36/75 (47%), Gaps = 6/75 (8%)
Query: 77 LEMEEREKVVRHMWDVYTNSR------RIRLPRFWQEAFEAAYEELTSDVPGVRDDAITE 130
L+ E+ + W++ NS+ ++ PR WQE+F Y +L +P + +++
Sbjct: 295 LDAEQDTTNPKPFWNMGENSKQLPGPQKLSDPRLWQESFWKNYSQLFWGLPSLHSESLVA 354
Query: 131 IAKMSVRSVNYDPPP 145
A ++ RS PP
Sbjct: 355 NAWVTDRSYTLQSPP 369
>UniRef100_Q6MDS2 Hypothetical protein [Parachlamydia sp.]
Length = 245
Score = 33.9 bits (76), Expect = 1.4
Identities = 13/36 (36%), Positives = 20/36 (55%)
Query: 87 RHMWDVYTNSRRIRLPRFWQEAFEAAYEELTSDVPG 122
+ WD YT+S+ I LP W E F+ + ++PG
Sbjct: 83 QEFWDQYTHSKGITLPANWNELFDQVRFKSIKEIPG 118
>UniRef100_Q8IU33 HrTRPV [Halocynthia roretzi]
Length = 786
Score = 33.9 bits (76), Expect = 1.4
Identities = 21/91 (23%), Positives = 44/91 (48%), Gaps = 1/91 (1%)
Query: 8 PTTTAAINKKENMDSGLFGKGRYKFWALAAILLLAFWSMFTGTVSLRWSGNLNT-LSSDL 66
P AA K+ M G+ K KFW++ I+ + +++ + + N N+ L +
Sbjct: 331 PMVLAAFLGKQEMFEGIMKKTSKKFWSINNIVCSGYPLKDLDSINPQGNTNWNSALIQII 390
Query: 67 DTPIHDDLDVLEMEEREKVVRHMWDVYTNSR 97
++ LD+L+ + ++++ WDV+ R
Sbjct: 391 HNNSNEHLDMLQADVITRLLQEKWDVFGKRR 421
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.318 0.133 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 277,277,098
Number of Sequences: 2790947
Number of extensions: 10478412
Number of successful extensions: 29315
Number of sequences better than 10.0: 36
Number of HSP's better than 10.0 without gapping: 19
Number of HSP's successfully gapped in prelim test: 17
Number of HSP's that attempted gapping in prelim test: 29281
Number of HSP's gapped (non-prelim): 36
length of query: 161
length of database: 848,049,833
effective HSP length: 117
effective length of query: 44
effective length of database: 521,509,034
effective search space: 22946397496
effective search space used: 22946397496
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 69 (31.2 bits)
Medicago: description of AC147178.10


