

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147010.4 - phase: 0 /pseudo
(618 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
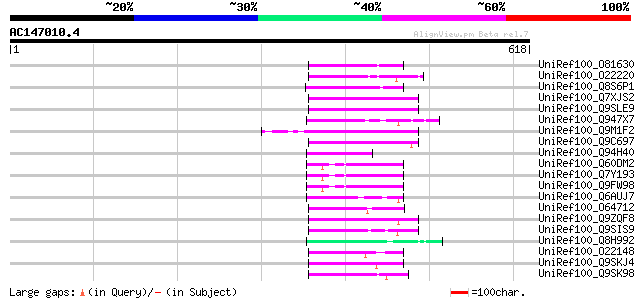
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_O81630 F8M12.22 protein [Arabidopsis thaliana] 78 7e-13
UniRef100_O22220 Putative non-LTR retroelement reverse transcrip... 76 3e-12
UniRef100_Q8S6P1 Putative reverse transcriptase [Oryza sativa] 74 1e-11
UniRef100_Q7XJS2 Putative non-LTR retroelement reverse transcrip... 73 2e-11
UniRef100_Q9SLE9 Putative non-LTR retroelement reverse transcrip... 72 5e-11
UniRef100_Q947X7 Hypothetical protein OSJNBa0067N01.20 [Oryza sa... 72 5e-11
UniRef100_Q9M1F2 Hypothetical protein F9K21.130 [Arabidopsis tha... 70 2e-10
UniRef100_Q9C697 Reverse transcriptase, putative; 16838-20266 [A... 68 7e-10
UniRef100_Q94H40 Putative reverse transcriptase [Oryza sativa] 65 6e-09
UniRef100_Q60DM2 F-box domain containing protein [Oryza sativa] 65 8e-09
UniRef100_Q7Y193 Putative reverse transcriptase [Oryza sativa] 65 8e-09
UniRef100_Q9FW98 Putative non-LTR retroelement reverse transcrip... 62 4e-08
UniRef100_Q6AUJ7 Hypothetical protein OSJNBa0052E20.4 [Oryza sat... 62 4e-08
UniRef100_O64712 Putative reverse transcriptase [Arabidopsis tha... 62 4e-08
UniRef100_Q9ZQF8 Putative non-LTR retroelement reverse transcrip... 60 1e-07
UniRef100_Q9SIS9 Putative non-LTR retroelement reverse transcrip... 60 2e-07
UniRef100_Q8H992 Oryza sativa (japonica cultivar-group) retrotra... 60 2e-07
UniRef100_O22148 Putative non-LTR retroelement reverse transcrip... 60 2e-07
UniRef100_Q9SKJ4 Putative non-LTR retroelement reverse transcrip... 59 3e-07
UniRef100_Q9SK98 Very similar to retrotransposon reverse transcr... 59 3e-07
>UniRef100_O81630 F8M12.22 protein [Arabidopsis thaliana]
Length = 1662
Score = 78.2 bits (191), Expect = 7e-13
Identities = 41/117 (35%), Positives = 61/117 (52%), Gaps = 5/117 (4%)
Query: 357 IWKLKVPPKFKNLVWCMCRGFQPTRVRLLDKGIQCPTNCVSCPSNHEDMSHIFFECPFAI 416
+WKL + PK K ++W PTR RLL +G+ +C CP+ E ++H+ F CP+A
Sbjct: 1362 LWKLPIIPKIKYMLWRTISKALPTRSRLLTRGMDIDPHCPRCPTEEETINHVLFTCPYAA 1421
Query: 417 HVW*MTSLWGTIQHALSSTTSAINVIFMLLETFSAELN--QRLTS--LFWSLWKHRN 469
+W +++ H S T N+ F++ + LN QRL L W LWK RN
Sbjct: 1422 SIWGLSNFPWLPGHTFSQDTEE-NISFLINSFSNNTLNTEQRLAPFWLIWRLWKARN 1477
>UniRef100_O22220 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1094
Score = 76.3 bits (186), Expect = 3e-12
Identities = 44/140 (31%), Positives = 68/140 (48%), Gaps = 7/140 (5%)
Query: 357 IWKLKVPPKFKNLVWCMCRGFQPTRVRLLDKGIQCPTNCVSCPSNHEDMSHIFFECPFAI 416
IW L PK K +W + +G RL +G+ C CP +E ++HI F+CP A
Sbjct: 775 IWNLNSAPKIKVFLWKVLKGAVAVEDRLRTRGVLIEDGCSMCPEKNETLNHILFQCPLAR 834
Query: 417 HVW*MTSLWGTIQHALSSTTSAINVIFMLLETFSAELNQRLTS----LFWSLWKHRNLRA 472
VW +T + H + NV ++ + EL+ L + W LWK+RN R
Sbjct: 835 QVWALTPMQSP-NHGFGDSIFT-NVNHVIGNCHNTELSPHLRYVSPWIIWILWKNRNKRL 892
Query: 473 WEDVTEVSVVVVERA*LAEC 492
+E + VS+ +V +A L +C
Sbjct: 893 FEGIGSVSLSIVGKA-LEDC 911
>UniRef100_Q8S6P1 Putative reverse transcriptase [Oryza sativa]
Length = 1509
Score = 74.3 bits (181), Expect = 1e-11
Identities = 41/119 (34%), Positives = 57/119 (47%), Gaps = 6/119 (5%)
Query: 353 FWSGIWKLKVPPKFKNLVWCMCRGFQPTRVRLLDKGIQCPTNCVSCPSNHEDMSHIFFEC 412
FW +WKL VP K K+ +W MC R L +G+ T CV C +ED H+FF+C
Sbjct: 1246 FWKKLWKLGVPGKIKHFLWRMCHNTLALRANLHHRGMDVDTRCVMCGRYNEDAGHLFFKC 1305
Query: 413 PFAIHVW*MTSLWGTIQHALSSTTSAINVIFMLLETFSAELNQRLTSL--FWSLWKHRN 469
VW +L ++ L TS NV L + N+R +++ W WK RN
Sbjct: 1306 KPVKKVWQALNL-EELRSMLEQQTSGKNV---LQSIYCRPENERTSAIVCLWQWWKERN 1360
>UniRef100_Q7XJS2 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1344
Score = 73.2 bits (178), Expect = 2e-11
Identities = 39/133 (29%), Positives = 64/133 (47%), Gaps = 2/133 (1%)
Query: 357 IWKLKVPPKFKNLVWCMCRGFQPTRVRLLDKGIQCPTNCVSCPSNHEDMSHIFFECPFAI 416
IW L PK + +W G P RL +GI+ C+ C + +E ++HI FECP A
Sbjct: 1024 IWNLHTAPKIRIFLWKALHGAIPVEDRLRTRGIRSDDGCLMCDTENETINHILFECPLAR 1083
Query: 417 HVW*MTSLWGTIQHALSSTTSAINVIFMLLETFSAELNQRLTS--LFWSLWKHRNLRAWE 474
VW +T L +S + ++ + L + + R S + W LWK+RN +E
Sbjct: 1084 QVWAITHLSSAGSEFSNSVYTNMSRLIDLTQQNDLPHHLRFVSPWILWFLWKNRNALLFE 1143
Query: 475 DVTEVSVVVVERA 487
++ +V++A
Sbjct: 1144 GKGSITTTLVDKA 1156
>UniRef100_Q9SLE9 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1319
Score = 72.0 bits (175), Expect = 5e-11
Identities = 38/133 (28%), Positives = 62/133 (46%), Gaps = 2/133 (1%)
Query: 357 IWKLKVPPKFKNLVWCMCRGFQPTRVRLLDKGIQCPTNCVSCPSNHEDMSHIFFECPFAI 416
+W L+ PK K +W +G RL +GI+ C+ C E ++H+ F+CPFA
Sbjct: 998 VWSLETVPKIKVFMWKALKGALAVEDRLRSRGIRTADGCLFCKEEIETINHLLFQCPFAR 1057
Query: 417 HVW*MTSLWGTIQHALSSTTSAINVIFMLLETFSAELNQRLTS--LFWSLWKHRNLRAWE 474
VW ++ + +S S IN + + F + R S L W +WK+RN ++
Sbjct: 1058 QVWALSLIQAPATGFGTSIFSNINHVIQNSQNFGIPRHMRTVSPWLLWEIWKNRNKTLFQ 1117
Query: 475 DVTEVSVVVVERA 487
S +V +A
Sbjct: 1118 GTGLTSSEIVAKA 1130
>UniRef100_Q947X7 Hypothetical protein OSJNBa0067N01.20 [Oryza sativa]
Length = 387
Score = 72.0 bits (175), Expect = 5e-11
Identities = 52/163 (31%), Positives = 70/163 (42%), Gaps = 17/163 (10%)
Query: 354 WSGIWKLKVPPKFKNLVWCMCRGFQPTRVRLLDKGIQCPTNCVSCPSNHEDMSHIFFECP 413
WS IW+ PP+ K W M + PTRV L K I C C +N ED HIF CP
Sbjct: 226 WSAIWRSAAPPRVKFFAWLMSKNRLPTRVNLHKKTILPTPTCELCNANLEDTYHIFLRCP 285
Query: 414 FAIHVW*MTSLWGTIQHALSSTTSAINVIFMLLETFSAELNQRLTSLF-----WSLWKHR 468
A W M I +SS + N+ S L L+S F W LW HR
Sbjct: 286 MAAAFWNMIQ----IIPEISSLSDLHNL------ELSGPLPTFLSSTFFLLCCWRLWNHR 335
Query: 469 NLRAWEDVTEVSVVVVERA*LAEC*LPWCS*FHPTSASATACW 511
N ++++ S+ + + L + L W F+P+ S W
Sbjct: 336 NEVVFQNLPP-SISRLINSCLQDAKL-WAHRFNPSQKSVLPLW 376
>UniRef100_Q9M1F2 Hypothetical protein F9K21.130 [Arabidopsis thaliana]
Length = 851
Score = 70.1 bits (170), Expect = 2e-10
Identities = 52/190 (27%), Positives = 84/190 (43%), Gaps = 21/190 (11%)
Query: 301 NSTGVQC*HC*ENSSNAASWSSTRGPSHLVLCS*CLLVMCHGTR*LFSPYVGFWSGIWKL 360
NSTG + + + W ST PS+ + + HG+ V + IW L
Sbjct: 629 NSTG-------DYTVRSGYWLSTHDPSNTIPT----MAKPHGS-------VDLKTKIWNL 670
Query: 361 KVPPKFKNLVWCMCRGFQPTRVRLLDKGIQCPTNCVSCPSNHEDMSHIFFECPFAIHVW* 420
+ PK K+ +W + PT RL +G++ C C +E ++H F CPFA W
Sbjct: 671 PIMPKLKHFLWRILSKALPTTDRLTTRGMRIDPGCPRCRRENESINHALFTCPFATMAWR 730
Query: 421 MTSLWGTIQHALSST-TSAINVIFMLLETFSAELNQRLTS--LFWSLWKHRNLRAWEDVT 477
++ LS+ I+ I +LL+ + +Q+L L W +WK RN + ++
Sbjct: 731 LSDTPLYRSSILSNNIEDNISNILLLLQNTTITDSQKLIPFWLLWRIWKARNNVVFNNLR 790
Query: 478 EVSVVVVERA 487
E + V RA
Sbjct: 791 ESPSITVVRA 800
>UniRef100_Q9C697 Reverse transcriptase, putative; 16838-20266 [Arabidopsis thaliana]
Length = 1142
Score = 68.2 bits (165), Expect = 7e-10
Identities = 37/135 (27%), Positives = 66/135 (48%), Gaps = 5/135 (3%)
Query: 357 IWKLKVPPKFKNLVWCMCRGFQPTRVRLLDKGIQCPTNCVSCPSNHEDMSHIFFECPFAI 416
IWK++ PPK ++ +W + G P L +GI C CVSC ++ E ++H F+C A
Sbjct: 828 IWKVQCPPKLRHFLWQILSGCVPVSENLRKRGILCDKGCVSCGASEESINHTLFQCHPAR 887
Query: 417 HVW*MTSLWGTIQHALSSTTSAINVIFMLLETFSAELNQRLTSLFWSLWKHRNLRAWEDV 476
+W ++ + T S + N+ + S + + W +WK RN + +E+V
Sbjct: 888 QIWALSQI-PTAPGIFPSNSIFTNLDHLFWRIPSGVDSAPYPWIIWYIWKARNEKVFENV 946
Query: 477 ----TEVSVVVVERA 487
E+ ++ V+ A
Sbjct: 947 DKDPMEILLLAVKEA 961
>UniRef100_Q94H40 Putative reverse transcriptase [Oryza sativa]
Length = 1185
Score = 65.1 bits (157), Expect = 6e-09
Identities = 30/79 (37%), Positives = 38/79 (47%)
Query: 354 WSGIWKLKVPPKFKNLVWCMCRGFQPTRVRLLDKGIQCPTNCVSCPSNHEDMSHIFFECP 413
W +WKL++PPK K W M G P + L DK I C C S ED+ H+ F C
Sbjct: 860 WDNLWKLEIPPKIKIFAWRMLHGVIPCKGILADKHIAVQGGCPVCSSGVEDIKHMVFTCR 919
Query: 414 FAIHVW*MTSLWGTIQHAL 432
A +W +W IQ L
Sbjct: 920 RAKLIWKQLGIWSRIQPIL 938
>UniRef100_Q60DM2 F-box domain containing protein [Oryza sativa]
Length = 855
Score = 64.7 bits (156), Expect = 8e-09
Identities = 35/121 (28%), Positives = 56/121 (45%), Gaps = 11/121 (9%)
Query: 354 WSGIWKLKVPPKFKNLVW-----CMCRGFQPTRVRLLDKGIQCPTNCVSCPSNHEDMSHI 408
W G+WK+ P K K ++W C+ GFQ R + I CV C + + + H+
Sbjct: 536 WKGLWKINAPGKMKIILWRAAHECLATGFQLPR-----RHIPSTDGCVFC-NRDDTVEHV 589
Query: 409 FFECPFAIHVW*MTSLWGTIQHALSSTTSAINVIFMLLETFSAELNQRLTSLFWSLWKHR 468
F CPFA +W ++ + ++ + IF L+ S+ N L FW +W+ R
Sbjct: 590 FLFCPFAAQIWEEIKRKCAVKLGRNGFSTMRHWIFDFLKRGSSHTNTLLAVTFWHIWEAR 649
Query: 469 N 469
N
Sbjct: 650 N 650
>UniRef100_Q7Y193 Putative reverse transcriptase [Oryza sativa]
Length = 404
Score = 64.7 bits (156), Expect = 8e-09
Identities = 35/121 (28%), Positives = 56/121 (45%), Gaps = 11/121 (9%)
Query: 354 WSGIWKLKVPPKFKNLVW-----CMCRGFQPTRVRLLDKGIQCPTNCVSCPSNHEDMSHI 408
W G+WK+ P K K ++W C+ GFQ R + I CV C + + + H+
Sbjct: 85 WKGLWKINAPGKMKIILWRAAHECLATGFQLPR-----RHIPSTDGCVFC-NRDDTVEHV 138
Query: 409 FFECPFAIHVW*MTSLWGTIQHALSSTTSAINVIFMLLETFSAELNQRLTSLFWSLWKHR 468
F CPFA +W ++ + ++ + IF L+ S+ N L FW +W+ R
Sbjct: 139 FLFCPFAAQIWEEIKRKCAVKLGRNGFSTMRHWIFDFLKRGSSHTNTLLAVTFWHIWEAR 198
Query: 469 N 469
N
Sbjct: 199 N 199
>UniRef100_Q9FW98 Putative non-LTR retroelement reverse transcriptase [Oryza sativa]
Length = 1382
Score = 62.4 bits (150), Expect = 4e-08
Identities = 35/121 (28%), Positives = 54/121 (43%), Gaps = 11/121 (9%)
Query: 354 WSGIWKLKVPPKFKNLVW-----CMCRGFQPTRVRLLDKGIQCPTNCVSCPSNHEDMSHI 408
W G+WK+ P K K +W C+ GFQ R + I CV C + + + H+
Sbjct: 1063 WKGLWKINAPGKMKITLWRAAHECLATGFQLRR-----RHIPSTDGCVFC-NRDDTVEHV 1116
Query: 409 FFECPFAIHVW*MTSLWGTIQHALSSTTSAINVIFMLLETFSAELNQRLTSLFWSLWKHR 468
F CPFA +W ++ + ++ IF L+ S+ N L FW +W+ R
Sbjct: 1117 FLFCPFAAQIWEEIKGKCAVKLGRNGFSTMRQWIFDFLKRGSSHANTLLAVTFWHIWEAR 1176
Query: 469 N 469
N
Sbjct: 1177 N 1177
>UniRef100_Q6AUJ7 Hypothetical protein OSJNBa0052E20.4 [Oryza sativa]
Length = 144
Score = 62.4 bits (150), Expect = 4e-08
Identities = 42/122 (34%), Positives = 53/122 (43%), Gaps = 17/122 (13%)
Query: 354 WSGIWKLKVPPKFKNLVWCMCRGFQPTRVRLLDKGIQCPTNCVSCPSNHEDMSHIFFECP 413
W IWK PP+ K W M PT V L K C C + ED HIF P
Sbjct: 10 WEFIWKSSAPPRVKFFGWLMTMNRLPTAVNLHKKSTIPSLTCQLCNTCPEDTDHIFLAYP 69
Query: 414 FAIHVW*MTSLWGTIQ-HALSSTTSAINVIFMLLETFSAELNQRLTSLF-----WSLWKH 467
A ++ WG IQ + ++ S I+ I +S +L +RL S F W LW H
Sbjct: 70 LA------SAFWGLIQVLPMLTSLSEIHTIH-----WSGDLPERLNSTFFLLCCWRLWNH 118
Query: 468 RN 469
RN
Sbjct: 119 RN 120
>UniRef100_O64712 Putative reverse transcriptase [Arabidopsis thaliana]
Length = 365
Score = 62.4 bits (150), Expect = 4e-08
Identities = 38/121 (31%), Positives = 51/121 (41%), Gaps = 11/121 (9%)
Query: 357 IWKLKVPPKFKNLVWCMCRGFQPTRVRLLDKGIQCPTNCVSCPSNHEDMSHIFFECPFAI 416
IWKL V PK K+ +W G T RL + I C C E + HI F CP+
Sbjct: 37 IWKLHVAPKIKHFLWRCVTGALATNTRLRSRNIDADPICQRCCIEEETIHHIMFNCPYTQ 96
Query: 417 HVW*MTSL-----WGTIQHALSSTTSAINVIFMLLETFSAELNQRLTS--LFWSLWKHRN 469
VW ++ WG SS +N + L +T + R + W LWK RN
Sbjct: 97 SVWRSANIIIGNQWG----PPSSFEDNLNRLIQLSKTQTTNSLDRFLPFWIMWRLWKSRN 152
Query: 470 L 470
+
Sbjct: 153 V 153
>UniRef100_Q9ZQF8 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1225
Score = 60.5 bits (145), Expect = 1e-07
Identities = 34/135 (25%), Positives = 63/135 (46%), Gaps = 5/135 (3%)
Query: 357 IWKLKVPPKFKNLVWCMCRGFQPTRVRLLDKGIQCPTNCVSCPSNHEDMSHIFFECPFAI 416
+WK+K KFK+ W G T RL + I C C + E ++H+ F CP +
Sbjct: 926 VWKIKTTRKFKHFEWQCLSGCLATNQRLFSRHIGTEKVCPRCGAEEESINHLLFLCPPSR 985
Query: 417 HVW*MTSLWGTIQHALSSTTSAINVIFMLLETFSAELNQRLTSLF----WSLWKHRNLRA 472
+W ++ + + ++ + N F+L ++ + + +F W +WK RN
Sbjct: 986 QIWALSPI-PSSEYIFPRNSLFYNFDFLLSRGKEFDIAEDIMEIFPWILWYIWKSRNRFI 1044
Query: 473 WEDVTEVSVVVVERA 487
+E+V E V+++ A
Sbjct: 1045 FENVIESPQVILDFA 1059
>UniRef100_Q9SIS9 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1715
Score = 60.1 bits (144), Expect = 2e-07
Identities = 39/139 (28%), Positives = 63/139 (45%), Gaps = 13/139 (9%)
Query: 357 IWKLKVPPKFKNLVWCMCRGFQPTRVRLLDKGIQCPTNCVSCPSNHEDMSHIFFECPFAI 416
IW+LK+ PK K+ +W G T +L ++ I C C + E ++HI F C +A
Sbjct: 1388 IWRLKITPKIKHFIWRCLSGALSTTTQLRNRNIPADPTCQRCCNADETINHIIFTCSYAQ 1447
Query: 417 HVW*MTSLWGTIQHALSSTTSAINVIFMLLETFSAELNQRLTSL--------FWSLWKHR 468
VW + G+ + L T + I ++L+ + NQ L L W LWK R
Sbjct: 1448 VVWRSANFSGS--NRLCFTDNLEENIRLILQ---GKKNQNLPILNGLMPFWIMWRLWKSR 1502
Query: 469 NLRAWEDVTEVSVVVVERA 487
N ++ + V ++A
Sbjct: 1503 NEYLFQQLDRFPWKVAQKA 1521
>UniRef100_Q8H992 Oryza sativa (japonica cultivar-group) retrotransposon Karma DNA,
complete sequence [Oryza sativa]
Length = 1197
Score = 60.1 bits (144), Expect = 2e-07
Identities = 43/164 (26%), Positives = 62/164 (37%), Gaps = 11/164 (6%)
Query: 354 WSGIWKLKVPPKFKNLVWCMCRGFQPTRVRLLDKGIQCPTNCVSCPSNHEDMSHIFFECP 413
W IW ++P K K W + R T++ LL K I C C + E H+ F CP
Sbjct: 1033 WKFIWDCRMPLKIKLFAWLLVRDRLSTKLNLLKKKIVQTATCDICSTTDETADHLSFNCP 1092
Query: 414 FAIHVW*MTSLWGTIQHA--LSSTTSAINVIFMLLETFSAELNQRLTSLFWSLWKHRNLR 471
FAI W + I LS + + ++F FW LW HR+
Sbjct: 1093 FAISFWQALHIQPIINETKYLSQLKAPATIPTRHFQSF-------FMLCFWVLWNHRHDV 1145
Query: 472 AWEDVTEVSVVVVERA*LAEC*LPWCS*FHPTSASATACWRGEF 515
+ ++R ++E L W F P W+G F
Sbjct: 1146 VFRGRPPSIASCLQRG-ISESSL-WAETFTPDDRFVIDVWKGVF 1187
>UniRef100_O22148 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1374
Score = 59.7 bits (143), Expect = 2e-07
Identities = 35/125 (28%), Positives = 56/125 (44%), Gaps = 21/125 (16%)
Query: 357 IWKLKVPPKFKNLVWCMCRGFQPTRVRLLDKGIQCPTNCVSCPSNHEDMSHIFFECPFAI 416
IWKL VPPK + +W L + + +CV CPS+ E ++H+ F+CPFA
Sbjct: 1054 IWKLDVPPKIHHFLWRCVNNCLSVASNLAYRHLAREKSCVRCPSHGETVNHLLFKCPFAR 1113
Query: 417 HVW*MT------------SLWGTIQHALSSTTSAINVIFMLLETFSAELNQRLTSLFWSL 464
W ++ SL+ + H LS S + ++ + + + W L
Sbjct: 1114 LTWAISPLPAPPGGEWAESLFRNMHHVLSVHKS---------QPEESDHHALIPWILWRL 1164
Query: 465 WKHRN 469
WK+RN
Sbjct: 1165 WKNRN 1169
>UniRef100_Q9SKJ4 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1750
Score = 59.3 bits (142), Expect = 3e-07
Identities = 33/117 (28%), Positives = 57/117 (48%), Gaps = 5/117 (4%)
Query: 357 IWKLKVPPKFKNLVWCMCRGFQPTRVRLLDKGIQCPTNCVSCPSNHEDMSHIFFECPFAI 416
IW L + PK K+ +W T RL +G++ +C C +E ++H F CPFA
Sbjct: 1428 IWNLPIMPKLKHFLWRALSQALATTERLTTRGMRIDPSCPRCHRENESINHALFTCPFAT 1487
Query: 417 HVW*MTSLWGTIQHALSST---TSAINVIFMLLETFSAELNQRL-TSLFWSLWKHRN 469
W ++ I++ L S + N++ + +T ++ ++ L L W +WK RN
Sbjct: 1488 MAWRLSDS-SLIRNQLMSNDFEENISNILNFVQDTTMSDFHKLLPVWLIWRIWKARN 1543
>UniRef100_Q9SK98 Very similar to retrotransposon reverse transcriptase [Arabidopsis
thaliana]
Length = 1231
Score = 59.3 bits (142), Expect = 3e-07
Identities = 33/122 (27%), Positives = 57/122 (46%), Gaps = 5/122 (4%)
Query: 357 IWKLKVPPKFKNLVWCMCRGFQPTRVRLLDKGIQCPTNCVSCPSNHEDMSHIFFECPFAI 416
+WKL+ PK K +W + G P L +G++ + C +C E + H+ F C F
Sbjct: 894 VWKLQTEPKIKVFLWKVLSGAIPVVDLLSYRGMKLDSRCQTCGCEGESIQHVLFSCSFPR 953
Query: 417 HVW*MTSLWGTIQHALSSTTSAINVIFMLLE----TFSAELNQRLTSLFWSLWKHRNLRA 472
VW M+++ + + A N+ L+ + EL + + W +WK+RNL
Sbjct: 954 QVWAMSNIHVPLLGFECGSVYA-NLYHFLINRDNLKWPVELRRSFPWIIWRIWKNRNLFF 1012
Query: 473 WE 474
+E
Sbjct: 1013 FE 1014
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.364 0.161 0.683
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 878,682,666
Number of Sequences: 2790947
Number of extensions: 31302327
Number of successful extensions: 139227
Number of sequences better than 10.0: 147
Number of HSP's better than 10.0 without gapping: 98
Number of HSP's successfully gapped in prelim test: 49
Number of HSP's that attempted gapping in prelim test: 138988
Number of HSP's gapped (non-prelim): 185
length of query: 618
length of database: 848,049,833
effective HSP length: 133
effective length of query: 485
effective length of database: 476,853,882
effective search space: 231274132770
effective search space used: 231274132770
T: 11
A: 40
X1: 14 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.5 bits)
S2: 78 (34.7 bits)
Medicago: description of AC147010.4


