

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147006.6 + phase: 0 /pseudo
(797 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
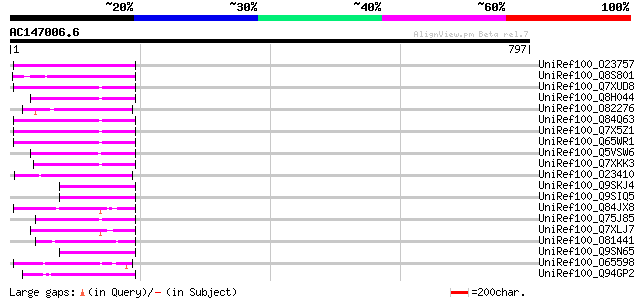
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_O23757 Reverse transcriptase [Beta vulgaris subsp. vul... 116 3e-24
UniRef100_Q8S801 Putative retroelement [Oryza sativa] 72 5e-11
UniRef100_Q7XUD8 OSJNBa0088A01.7 protein [Oryza sativa] 70 2e-10
UniRef100_Q8H044 Putative reverse transcriptase [Oryza sativa] 70 2e-10
UniRef100_O82276 Putative non-LTR retroelement reverse transcrip... 70 2e-10
UniRef100_Q84Q63 Putative reverse transcriptase [Oryza sativa] 70 3e-10
UniRef100_Q7X5Z1 OSJNBa0006A01.14 protein [Oryza sativa] 68 9e-10
UniRef100_Q65WR1 Putative polyprotein [Oryza sativa] 67 2e-09
UniRef100_Q5VSW6 OSJNBa0009P12.10 protein [Oryza sativa] 67 2e-09
UniRef100_Q7XKK3 OSJNBa0038O10.4 protein [Oryza sativa] 67 3e-09
UniRef100_O23410 STRONG homology to reverse transcriptase [Arabi... 66 4e-09
UniRef100_Q9SKJ4 Putative non-LTR retroelement reverse transcrip... 66 5e-09
UniRef100_Q9SIQ5 Putative non-LTR retroelement reverse transcrip... 66 5e-09
UniRef100_Q84JX8 Putative reverse transcriptase [Oryza sativa] 65 6e-09
UniRef100_Q75J85 Putative reverse transcriptase [Oryza sativa] 64 1e-08
UniRef100_Q7XLJ7 OSJNBa0009K15.13 protein [Oryza sativa] 64 2e-08
UniRef100_O81441 T24H24.17 protein [Arabidopsis thaliana] 64 2e-08
UniRef100_Q9SN65 Hypothetical protein F25I24.40 [Arabidopsis tha... 62 5e-08
UniRef100_O65598 Hypothetical protein M3E9.210 [Arabidopsis thal... 62 5e-08
UniRef100_Q94GP2 Putative reverse transcriptase [Oryza sativa] 62 5e-08
>UniRef100_O23757 Reverse transcriptase [Beta vulgaris subsp. vulgaris]
Length = 507
Score = 116 bits (290), Expect = 3e-24
Identities = 63/186 (33%), Positives = 103/186 (54%)
Query: 7 KLKGLKVILKE*IKKEYGGMEDRVELLVEDIKMLDEKGEEGTLTE*EVGLHKGKLVELWR 66
KL+ LK LK+ +KE+G ++D ++ E I LD L E+ + LW
Sbjct: 140 KLRTLKEALKQWNQKEFGIIDDNIKHCEEKIHFLDNIANSRLLCTNELQERDDAQLNLWT 199
Query: 67 LLKAKDTLITQRSRSRWLKKGDANSKFFHKCIKLRKSINSIKALEENEGWVVSPFEVCRK 126
LK ++ SR+RW+K+GD+N+++FH +R+ N ++ ++ + P E+ +
Sbjct: 200 WLKRSESYWALNSRARWIKEGDSNTRYFHTLATIRRHKNHLQYIKTEGKNISKPGEIKEE 259
Query: 127 VVNYFTNHFAEDRWDRPRLDRVDFESLTEVENGLLVAPFSLLEIEEVVRDSDGGKSPGPN 186
V YF + FAE+ RP + ++F L+E E L+APF+ EI+E V D K+PGP+
Sbjct: 260 AVAYFQHIFAEEFSTRPVFNGLNFNKLSESECSSLIAPFTHEEIDEAVDSCDAQKAPGPD 319
Query: 187 GYNFSF 192
G+NF F
Sbjct: 320 GFNFKF 325
>UniRef100_Q8S801 Putative retroelement [Oryza sativa]
Length = 288
Score = 72.4 bits (176), Expect = 5e-11
Identities = 56/192 (29%), Positives = 97/192 (50%), Gaps = 12/192 (6%)
Query: 5 KKKLKGLKVILKE*IKKEYGGMEDRVELLVEDIKMLDEKGEEGTLTE*EVGLHKGKLVEL 64
++K++G + K+ I KE E L+ I+ +DE+ E+ LT E G H+ +L
Sbjct: 96 RRKMRGWAMNKKKEINKEK-------EELISQIQKIDEEVEKKELTCEEWG-HRYELEGA 147
Query: 65 WRLLKAKDTLITQ-RSRSRWLKKGDANSKFFHKCIKLRKSINSIKALEENEGWVVSPFEV 123
+ + + Q RS +W+ +GDAN+ FFH RK +I++LE EG + + E+
Sbjct: 148 LNQIFISEEIYWQGRSGEKWMLEGDANAAFFHGVANGRKRKTTIRSLEGEEGPIENADEL 207
Query: 124 CRKVVNYFTNHFAEDRWDRPRLDRVDFES---LTEVENGLLVAPFSLLEIEEVVRDSDGG 180
+ ++ F + + L +E L+ +N L PFS+ EIE +R+
Sbjct: 208 KEHIYGFYKKLFGMELPPKFCLSHNMWEGRGRLSSEDNIELTKPFSMEEIEGALREMKTN 267
Query: 181 KSPGPNGYNFSF 192
+PGP+G++ SF
Sbjct: 268 TAPGPDGFSVSF 279
>UniRef100_Q7XUD8 OSJNBa0088A01.7 protein [Oryza sativa]
Length = 1324
Score = 70.5 bits (171), Expect = 2e-10
Identities = 54/188 (28%), Positives = 84/188 (43%), Gaps = 5/188 (2%)
Query: 7 KLKGLKVILKE*IKKEYGGMEDRVELLVEDIKMLDEKGEEGTLTE*EVGLHKGKLVELWR 66
KL + LK+ + ++ + L ++ I+ LD E LT+ E L G L
Sbjct: 734 KLSHTALALKKWARGICSEVKLKFHLALDVIQRLDVAQENRDLTQAEFRLRLGLKRRLLG 793
Query: 67 LLKAKDTLITQRSRSRWLKKGDANSKFFHKCIKLRKSINSIKALEENEGWVVSPFEVCRK 126
+ Q SR +K+GDAN+K+FH+ + RK N I L GW V+ E
Sbjct: 794 YAAIERARKKQASRITNIKEGDANTKYFHRKVNARKRRNHIFRLRRQNGWAVTHAEKEGV 853
Query: 127 VVNYFTNHFAEDRWDRPRLDRVDFESLTEVENGL--LVAPFSLLEIEEVVRDSDGGKSPG 184
+ ++F PR +++ SL L L PFS EI + + + G K+PG
Sbjct: 854 IADHFRQVMGR---PEPRTSDLNWPSLDLAPTDLTRLEEPFSEAEIRKAINEMPGDKAPG 910
Query: 185 PNGYNFSF 192
P+G+ F
Sbjct: 911 PDGFTGKF 918
>UniRef100_Q8H044 Putative reverse transcriptase [Oryza sativa]
Length = 1183
Score = 70.1 bits (170), Expect = 2e-10
Identities = 46/163 (28%), Positives = 83/163 (50%), Gaps = 5/163 (3%)
Query: 32 LLVEDIKMLDEKGEEGTLTE*EVGLHKGKLVELWRLLKAKDTLITQRSRSRWLKKGDANS 91
+ ++ I+ LD E L+ E+ L ++ L + Q SR +++GDAN+
Sbjct: 32 MALDVIQRLDVAQEHRDLSPQEMWLRAALKRKILGLASIERARKKQASRVTHIREGDANT 91
Query: 92 KFFHKCIKLRKSINSIKALEENEGWVVSPFEVCRKVVNYFTNHFAEDRWDRPRLDRVDFE 151
KFFH I +RK N+I+ L ++ GW V+ E + ++F + PR + +++E
Sbjct: 92 KFFHLKINVRKRKNTIQRLRKDNGWAVTHGEKESTIHDHFASFMGR---SGPRPNELNWE 148
Query: 152 SL--TEVENGLLVAPFSLLEIEEVVRDSDGGKSPGPNGYNFSF 192
+L V+ L PFS E+ +++ K+PGP+G+ +F
Sbjct: 149 ALDIMPVDLATLGKPFSEEEVHRAIKEMPADKAPGPDGFTGAF 191
>UniRef100_O82276 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1231
Score = 70.1 bits (170), Expect = 2e-10
Identities = 53/176 (30%), Positives = 92/176 (52%), Gaps = 12/176 (6%)
Query: 20 KKEYGGMEDRVELLVEDIK----MLDEKGEEGTLTE*EVGLHKGKLVELWRLLKAKDTLI 75
++ +G + R E LV DIK +L + L + EV L + LV L+ ++TL
Sbjct: 145 REVFGDIHVRKEKLVADIKEVQDLLGVVLSDDLLAKEEVLLKEMDLV-----LEQEETLW 199
Query: 76 TQRSRSRWLKKGDANSKFFHKCIKLRKSINSIKALE-ENEGWVVSPFEVCRKVVNYFTNH 134
Q+SR ++++ GD N+ FFH +R+ N I++L+ +++ WV E+ + Y+
Sbjct: 200 FQKSREKYIELGDRNTTFFHTSTIIRRRRNRIESLKGDDDRWVTDKVELEAMALTYYKRL 259
Query: 135 FA-EDRWD-RPRLDRVDFESLTEVENGLLVAPFSLLEIEEVVRDSDGGKSPGPNGY 188
++ ED + R L F S++E E L+ F+ E+ V+ K+PGP+GY
Sbjct: 260 YSLEDVSEVRNMLPTGGFASISEAEKAALLQAFTKAEVVSAVKSMGRFKAPGPDGY 315
>UniRef100_Q84Q63 Putative reverse transcriptase [Oryza sativa]
Length = 870
Score = 69.7 bits (169), Expect = 3e-10
Identities = 49/189 (25%), Positives = 94/189 (48%), Gaps = 5/189 (2%)
Query: 6 KKLKGLKVILKE*IKKEYGGMEDRVELLVEDIKMLDEKGEEGTLTE*EVGLHKGKLVELW 65
+KL+ V L++ K + + + + I+ LD E L+ E L ++
Sbjct: 281 QKLQRTAVGLRDWTKGICSEAKLQFHMALNVIQRLDVAQEHRELSPPETRLRAALKRKVL 340
Query: 66 RLLKAKDTLITQRSRSRWLKKGDANSKFFHKCIKLRKSINSIKALEENEGWVVSPFEVCR 125
RL + Q SR +++GDAN+KFFH + R+ N+I+ L+++ GW V+ E
Sbjct: 341 RLASIERARKKQASRVTHIREGDANTKFFHLKVNARRRRNTIQRLKKDSGWAVTHGEKES 400
Query: 126 KVVNYFTNHFAEDRWDRPRLDRVDFESL--TEVENGLLVAPFSLLEIEEVVRDSDGGKSP 183
+ ++F + PR + +++E+L T ++ L PF+ E+ +++ K+P
Sbjct: 401 TIHDHFASFMGR---PGPRPNELNWETLDITLIDLATLGEPFTEEEVHRAIKEMPADKAP 457
Query: 184 GPNGYNFSF 192
GP+G+ +F
Sbjct: 458 GPDGFTGAF 466
>UniRef100_Q7X5Z1 OSJNBa0006A01.14 protein [Oryza sativa]
Length = 1189
Score = 68.2 bits (165), Expect = 9e-10
Identities = 48/189 (25%), Positives = 94/189 (49%), Gaps = 5/189 (2%)
Query: 6 KKLKGLKVILKE*IKKEYGGMEDRVELLVEDIKMLDEKGEEGTLTE*EVGLHKGKLVELW 65
+KL+ V L++ K + + + + I+ LD E LT E L ++
Sbjct: 281 QKLQHTAVGLRDWTKGICSEAKLQFHMALNVIQRLDVAQEHRELTPLETRLRAALKRKVL 340
Query: 66 RLLKAKDTLITQRSRSRWLKKGDANSKFFHKCIKLRKSINSIKALEENEGWVVSPFEVCR 125
L + Q SR +++GDAN+KFFH + R+ N+I+ L+++ GW V+ +
Sbjct: 341 GLASIERARKKQASRVTHIREGDANTKFFHLKVNARRRRNTIQRLKKDSGWAVTHGDKEL 400
Query: 126 KVVNYFTNHFAEDRWDRPRLDRVDFESL--TEVENGLLVAPFSLLEIEEVVRDSDGGKSP 183
+ ++F + PR + +++E+L T ++ L PF+ E+ + +++ K+P
Sbjct: 401 TIHDHFASFMGR---PGPRPNELNWETLDITPIDLATLGEPFTEEEVHKAIKEMPADKAP 457
Query: 184 GPNGYNFSF 192
GP+G+ +F
Sbjct: 458 GPDGFTGAF 466
>UniRef100_Q65WR1 Putative polyprotein [Oryza sativa]
Length = 1023
Score = 67.4 bits (163), Expect = 2e-09
Identities = 48/189 (25%), Positives = 93/189 (48%), Gaps = 5/189 (2%)
Query: 6 KKLKGLKVILKE*IKKEYGGMEDRVELLVEDIKMLDEKGEEGTLTE*EVGLHKGKLVELW 65
+KL+ V L++ K + + + + I+ LD E L+ E L ++
Sbjct: 115 QKLQRTAVGLRDWTKGICSEAKLQFHMALNVIQRLDVAQEHRELSPPETRLRAALKRKVL 174
Query: 66 RLLKAKDTLITQRSRSRWLKKGDANSKFFHKCIKLRKSINSIKALEENEGWVVSPFEVCR 125
L + Q SR +++GDAN+KFFH + R+ N+I+ L+++ GW V+ E
Sbjct: 175 GLASIERARKKQASRVTHIREGDANTKFFHLRVNARRRRNTIQRLKKDSGWAVTHGEKEL 234
Query: 126 KVVNYFTNHFAEDRWDRPRLDRVDFESL--TEVENGLLVAPFSLLEIEEVVRDSDGGKSP 183
+ ++F + PR + +++E+L T ++ L PF+ E+ +++ K+P
Sbjct: 235 TIHDHFASFMGR---PGPRPNELNWETLDITPIDLATLGEPFTEEEVHRAIKEMPADKAP 291
Query: 184 GPNGYNFSF 192
GP+G+ +F
Sbjct: 292 GPDGFTGAF 300
>UniRef100_Q5VSW6 OSJNBa0009P12.10 protein [Oryza sativa]
Length = 1230
Score = 67.0 bits (162), Expect = 2e-09
Identities = 48/163 (29%), Positives = 78/163 (47%), Gaps = 5/163 (3%)
Query: 32 LLVEDIKMLDEKGEEGTLTE*EVGLHKGKLVELWRLLKAKDTLITQRSRSRWLKKGDANS 91
+ ++ I LD E LT+ E L G L + T Q SR +K+ DAN+
Sbjct: 564 MALDVIHRLDVAQEHRDLTQAEYRLRLGLKRILLGFAVIERTRKKQASRITNIKEADANT 623
Query: 92 KFFHKCIKLRKSINSIKALEENEGWVVSPFEVCRKVVNYFTNHFAEDRWDRPRLDRVDFE 151
KFFH+ I R+ N I L+++ GW V+ + + +F + W PR +++
Sbjct: 624 KFFHRKINARRRKNHIHRLKKHNGWAVTHEDKESTIAEHFRHVMG---WPEPRSCDLNWP 680
Query: 152 SL--TEVENGLLVAPFSLLEIEEVVRDSDGGKSPGPNGYNFSF 192
L T ++ L PF+ EI E +++ K+PGP+G+ F
Sbjct: 681 VLGYTPIDLSGLDDPFTKAEIHEAIKEMPIDKAPGPDGFTGKF 723
>UniRef100_Q7XKK3 OSJNBa0038O10.4 protein [Oryza sativa]
Length = 1045
Score = 66.6 bits (161), Expect = 3e-09
Identities = 46/158 (29%), Positives = 76/158 (47%), Gaps = 5/158 (3%)
Query: 37 IKMLDEKGEEGTLTE*EVGLHKGKLVELWRLLKAKDTLITQRSRSRWLKKGDANSKFFHK 96
+K LD E L+ E+ L + L + Q SR +++G+AN+KFFH
Sbjct: 262 VKRLDVAQEHRDLSGPELRLRAALKRKTLTLATIERERKRQASRLTHIREGEANTKFFHL 321
Query: 97 CIKLRKSINSIKALEENEGWVVSPFEVCRKVVNYFTNHFAEDRWDRPRLDRVDFESL--T 154
+ R+ NSI+ L + GW V+ E ++++F + PR + +++E+L
Sbjct: 322 RVNTRRRKNSIQRLRKENGWAVTHDEKEMTILDHFAHLMGR---PGPRTNELNWEALDIQ 378
Query: 155 EVENGLLVAPFSLLEIEEVVRDSDGGKSPGPNGYNFSF 192
+ L PFS EI +++ KSPGP+GY F
Sbjct: 379 PFDLSTLEEPFSEQEIHRAIKEMPVDKSPGPDGYTGIF 416
>UniRef100_O23410 STRONG homology to reverse transcriptase [Arabidopsis thaliana]
Length = 929
Score = 66.2 bits (160), Expect = 4e-09
Identities = 55/184 (29%), Positives = 89/184 (47%), Gaps = 4/184 (2%)
Query: 8 LKGLKVILKE*IKKEYGGMEDRVELLVEDIKMLDEKGEEGTLTE*EVGLHKGKLVELWRL 67
L L+ LK+ K+ +G + R E +V D+K + + E T+ + L E L
Sbjct: 85 LNRLRWQLKKWNKEVFGNIHVRKEKVVSDLKAVQDLLEV-VQTDDLLMKEDTLLKEFDVL 143
Query: 68 LKAKDTLITQRSRSRWLKKGDANSKFFHKCIKLRKSINSIKALEENEG-WVVSPFEVCRK 126
L ++TL Q+SR + L GD N+ FFH +R+ N I+ L+++E WV + +
Sbjct: 144 LHQEETLWFQKSREKLLALGDRNTTFFHTSTVIRRRRNRIEMLKDSEDRWVTEKEALEKL 203
Query: 127 VVNYFTNHFAEDRWD--RPRLDRVDFESLTEVENGLLVAPFSLLEIEEVVRDSDGGKSPG 184
++Y+ ++ + R L F LT E L PF+ E+ VR K+PG
Sbjct: 204 AMDYYRKLYSLEDVSVVRGTLPTEGFPRLTREEKNNLNRPFTRDEVVVAVRSMGRFKAPG 263
Query: 185 PNGY 188
P+GY
Sbjct: 264 PDGY 267
>UniRef100_Q9SKJ4 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1750
Score = 65.9 bits (159), Expect = 5e-09
Identities = 38/118 (32%), Positives = 62/118 (52%), Gaps = 2/118 (1%)
Query: 77 QRSRSRWLKKGDANSKFFHKCIKLRKSINSIKALEENEG-WVVSPFEVCRKVVNYFTNHF 135
Q+SR++W+K+GD N+ +FH C K R S N + + +++G E+ ++FTN F
Sbjct: 720 QKSRNQWMKEGDQNTGYFHACTKTRYSQNRVNTIMDDQGRMFTGDKEIGNHAQDFFTNIF 779
Query: 136 AEDRWDRPRLDRVDFES-LTEVENGLLVAPFSLLEIEEVVRDSDGGKSPGPNGYNFSF 192
+ + +D DF+S +T N L FS EI + + K+PGP+G F
Sbjct: 780 STNGIKVSPIDFADFKSTVTNTVNLDLTKEFSDTEIYDAICQIGDDKAPGPDGLTARF 837
>UniRef100_Q9SIQ5 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1524
Score = 65.9 bits (159), Expect = 5e-09
Identities = 38/118 (32%), Positives = 62/118 (52%), Gaps = 2/118 (1%)
Query: 77 QRSRSRWLKKGDANSKFFHKCIKLRKSINSIKALEENEG-WVVSPFEVCRKVVNYFTNHF 135
Q+SR++W+K+GD N+ +FH C K R S N + + +++G E+ ++FTN F
Sbjct: 494 QKSRNQWMKEGDQNTGYFHACTKTRYSQNRVNTIMDDQGRMFTGDKEIGNHAQDFFTNIF 553
Query: 136 AEDRWDRPRLDRVDFES-LTEVENGLLVAPFSLLEIEEVVRDSDGGKSPGPNGYNFSF 192
+ + +D DF+S +T N L FS EI + + K+PGP+G F
Sbjct: 554 STNGIKVSPIDFADFKSTVTNTVNLDLTKEFSDTEIYDAICQIGDDKAPGPDGLTARF 611
>UniRef100_Q84JX8 Putative reverse transcriptase [Oryza sativa]
Length = 1349
Score = 65.5 bits (158), Expect = 6e-09
Identities = 63/196 (32%), Positives = 90/196 (45%), Gaps = 21/196 (10%)
Query: 7 KLKGLKVILKE*IKKEYGGMEDRVELLVEDIKMLDEKGEEGTLTE*EVGLH---KGKLVE 63
KLK L LK K G + + + E I LD EE LT+ E+ L K +++
Sbjct: 607 KLKRLARDLKRWSKFRIGDIRLQHAIANEVIFQLDVAQEERALTDEELLLKSTLKSRVLG 666
Query: 64 LWRLLKAKDTLITQRSRSRWLKKGDANSKFFHKCIKLRKSINSIKALEENEGWVVSPFEV 123
L L+K + I QR+R WLK GD NSKFFH RK N I AL G V+ +
Sbjct: 667 LATLVKIR---IRQRARLTWLKCGDVNSKFFHLKANSRKRKNFIPALITPSGIAVTTEQK 723
Query: 124 CRKVVNYFTNHFAED-------RWDRPRLDRVDFESLTEVENGLLVAPFSLLEIEEVVRD 176
++ YF+ + W L +D L+EVE F+ E++E + +
Sbjct: 724 EAELERYFSQRLGQYVPRSLTLNWSELSLPSLD---LSEVEE-----DFTEEELKEAISN 775
Query: 177 SDGGKSPGPNGYNFSF 192
K+PGP+ Y +F
Sbjct: 776 LPPQKAPGPDSYIGAF 791
>UniRef100_Q75J85 Putative reverse transcriptase [Oryza sativa]
Length = 817
Score = 64.3 bits (155), Expect = 1e-08
Identities = 46/155 (29%), Positives = 73/155 (46%), Gaps = 5/155 (3%)
Query: 40 LDEKGEEGTLTE*EVGLHKGKLVELWRLLKAKDTLITQRSRSRWLKKGDANSKFFHKCIK 99
LD E LT+ E L G L + Q +R +K+GDAN+KFFH+ I
Sbjct: 90 LDVAQEHRELTQAEYRLRMGLKRRLLGFAVIERARKKQAARITNIKEGDANTKFFHRKIN 149
Query: 100 LRKSINSIKALEENEGWVVSPFEVCRKVVNYFTNHFAEDRWDRPRLDRVDFESL--TEVE 157
RK N I L++ GW V+ E + +F + + PR +++ L T ++
Sbjct: 150 ARKRKNHIHRLKKQNGWAVTHAEKELVIAEHFQHVMGQ---PEPRSCDLNWPELGYTPLD 206
Query: 158 NGLLVAPFSLLEIEEVVRDSDGGKSPGPNGYNFSF 192
L PF+ EI + +++ K+PGP+G+ F
Sbjct: 207 LSGLDDPFTEAEIHKAIKEMPADKAPGPDGFTGKF 241
>UniRef100_Q7XLJ7 OSJNBa0009K15.13 protein [Oryza sativa]
Length = 779
Score = 63.9 bits (154), Expect = 2e-08
Identities = 45/171 (26%), Positives = 80/171 (46%), Gaps = 19/171 (11%)
Query: 33 LVEDIKMLDEKGEEGTLTE*EVGLHKGKLVELWRLLKAKDTLITQRSRSRWLKKGDANSK 92
L+++++ D+ + L++ E +L + + ++ +RS +WL +GDAN+
Sbjct: 19 LLKEMQKWDDAADSRNLSQGEWKQRYELEDQLNIIFQEEEIYWQERSGEKWLLEGDANTT 78
Query: 93 FFHKCIKLRKSINSIKALEENEGWVVSPFEVCRKVVNYFTNHFAED-----------RWD 141
FFH +K I +I++LEE+ + E+ + Y+ F + R D
Sbjct: 79 FFHGVANGKKRIITIRSLEEDGRVIEGNSELREHITRYYKGLFGSELPPKVFLSQDMRRD 138
Query: 142 RPRLDRVDFESLTEVENGLLVAPFSLLEIEEVVRDSDGGKSPGPNGYNFSF 192
R R+D+ D N LV PFS+ EIE + + +PGP+G F
Sbjct: 139 RGRVDQED--------NDFLVRPFSMEEIERALAEMKTNTAPGPDGLPVCF 181
>UniRef100_O81441 T24H24.17 protein [Arabidopsis thaliana]
Length = 1077
Score = 63.5 bits (153), Expect = 2e-08
Identities = 46/156 (29%), Positives = 77/156 (48%), Gaps = 7/156 (4%)
Query: 40 LDEKGEEGTLTE*EVGLHKGKLVELWRLLKAKDTLITQRSRSRWLKKGDANSKFFHKCIK 99
L+E + T+ V L L+ +R ++T QRSR+ WL GD NS FFH
Sbjct: 280 LEEALTDDAATQDSVDLLNQNLLLAYR---KEETFWKQRSRNLWLALGDKNSCFFHAATN 336
Query: 100 LRKSINSIKALEENEGW-VVSPFEVCRKVVNYFTNHFAEDRWDRPRL--DRVDFESLTEV 156
R++IN+I +E + G V E+ + +YF + F DR + + + T++
Sbjct: 337 SRRAINTISVIENSAGTSVYEDSEIIETISDYFQDIFTSQEGDRTSVVAEALSPCITTDM 396
Query: 157 ENGLLVAPFSLLEIEEVVRDSDGGKSPGPNGYNFSF 192
N L+ P ++ EI++ K+PG +G++ SF
Sbjct: 397 NNTLISLP-TVAEIKQACLSIHPDKAPGRDGFSASF 431
>UniRef100_Q9SN65 Hypothetical protein F25I24.40 [Arabidopsis thaliana]
Length = 1294
Score = 62.4 bits (150), Expect = 5e-08
Identities = 37/118 (31%), Positives = 60/118 (50%), Gaps = 2/118 (1%)
Query: 77 QRSRSRWLKKGDANSKFFHKCIKLRKSINSIKALEENEGWVV-SPFEVCRKVVNYFTNHF 135
Q+SR++W+K+GD N++FFH C K R S+N + +++ EG + E+ +FT +
Sbjct: 700 QKSRNQWMKEGDRNTEFFHACTKTRFSVNRLVTIKDEEGMIYRGDKEIGVHAQEFFTKVY 759
Query: 136 AEDRWDRPRLDRVDFESL-TEVENGLLVAPFSLLEIEEVVRDSDGGKSPGPNGYNFSF 192
+ +D F+ + TE N L S LEI + K+PGP+G F
Sbjct: 760 ESNGRPVSIIDFAGFKPIVTEQINDDLTKDLSDLEIYNAICHIGDDKAPGPDGLTARF 817
>UniRef100_O65598 Hypothetical protein M3E9.210 [Arabidopsis thaliana]
Length = 1141
Score = 62.4 bits (150), Expect = 5e-08
Identities = 52/195 (26%), Positives = 88/195 (44%), Gaps = 20/195 (10%)
Query: 6 KKLKGLKVILKE*IKKEYGGMEDRVELLVEDIKMLDEKGEEGTLTE*EVGLHKGKLVELW 65
KKLK LK +K+ + Y +E R E E + + E H+ + W
Sbjct: 276 KKLKALKKPIKDFSRLNYSNLEKRTEEAHETLLSFQNLTLDNPSLE--NAAHELEAQRKW 333
Query: 66 RLLK-AKDTLITQRSRSRWLKKGDANSKFFHKCIKLRKSINSIKALEENEGWVV-SPFEV 123
++L A+++ QRSR W +GD N+++FH+ RKS+N+I L ++ G + S +
Sbjct: 334 QILATAEESFFRQRSRVTWFAEGDGNTRYFHRMADSRKSVNTITTLVDDSGTQIDSQQGI 393
Query: 124 CRKVVNYFTNHFAEDRWDRPRLDRVDFESLTEVENGLLVAPFS-LLEIEEVVRDSD---- 178
YF N ++D D L++ D L P+S + ++E + D D
Sbjct: 394 ADHCALYFENLLSDDN-DPYSLEQDDMNLLLTYR-----CPYSQVADLEAMFSDEDIKAA 447
Query: 179 -----GGKSPGPNGY 188
K+ GP+G+
Sbjct: 448 FFGLPSNKACGPDGF 462
>UniRef100_Q94GP2 Putative reverse transcriptase [Oryza sativa]
Length = 1833
Score = 62.4 bits (150), Expect = 5e-08
Identities = 49/175 (28%), Positives = 83/175 (47%), Gaps = 7/175 (4%)
Query: 20 KKEYGGMEDRVELLVEDIKMLDEKGEEGTLTE*EVGLHKGKLVELWRLLKAKDTLITQRS 79
K + G + +++ L E + L G T E +H K EL +L ++ QRS
Sbjct: 271 KDKIGNVRKKIKDLREKLGELRNIGLLDTDNE----VHSVKK-ELEEMLHREEIWWKQRS 325
Query: 80 RSRWLKKGDANSKFFHKCIKLRKSINSIKALEENEGWVVSPFEVCRKVVNYFTN--HFAE 137
R WLK+GD N+++FH R N IK L++N+G + +++ F + +
Sbjct: 326 RITWLKEGDLNTRYFHLKASWRAKKNKIKKLKKNDGSTTMNKKEMKEINRSFFQQLYTKD 385
Query: 138 DRWDRPRLDRVDFESLTEVENGLLVAPFSLLEIEEVVRDSDGGKSPGPNGYNFSF 192
D + L + E ++E N L+ PF+ EI + + K+PGP+G+ F
Sbjct: 386 DNLNPVNLLNMFHEKISEQMNADLIKPFTNEEISDALFQIGPLKAPGPDGFPARF 440
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.356 0.160 0.625
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,231,872,242
Number of Sequences: 2790947
Number of extensions: 47173718
Number of successful extensions: 205095
Number of sequences better than 10.0: 225
Number of HSP's better than 10.0 without gapping: 122
Number of HSP's successfully gapped in prelim test: 103
Number of HSP's that attempted gapping in prelim test: 204654
Number of HSP's gapped (non-prelim): 416
length of query: 797
length of database: 848,049,833
effective HSP length: 136
effective length of query: 661
effective length of database: 468,481,041
effective search space: 309665968101
effective search space used: 309665968101
T: 11
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.7 bits)
S2: 79 (35.0 bits)
Medicago: description of AC147006.6


