

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147000.4 + phase: 1 /pseudo
(187 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
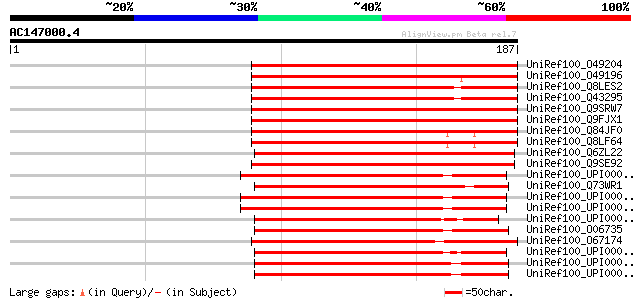
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_O49204 Adenylyl-sulfate kinase, chloroplast precursor ... 156 3e-37
UniRef100_O49196 Adenylyl-sulfate kinase 2, chloroplast precurso... 156 3e-37
UniRef100_Q8LES2 Putative adenosine phosphosulfate kinase [Arabi... 152 5e-36
UniRef100_Q43295 Adenylyl-sulfate kinase 1, chloroplast precurso... 152 5e-36
UniRef100_Q9SRW7 Putative adenylylsulfate kinase [Arabidopsis th... 142 6e-33
UniRef100_Q9FJX1 Adenylylsulfate kinase-like protein [Arabidopsi... 140 1e-32
UniRef100_Q84JF0 Putative adenylylsulfate kinase [Arabidopsis th... 138 6e-32
UniRef100_Q8LF64 Adenylylsulfate kinase-like protein [Arabidopsi... 138 6e-32
UniRef100_Q6ZL22 Putative adenosine-5'-phosphosulfate kinase [Or... 135 7e-31
UniRef100_Q9SE92 Adenosine-5'-phosphosulfate kinase [Zea mays] 130 2e-29
UniRef100_UPI00002E654A UPI00002E654A UniRef100 entry 114 1e-24
UniRef100_Q73WR1 Hypothetical protein [Mycobacterium paratubercu... 112 4e-24
UniRef100_UPI0000278622 UPI0000278622 UniRef100 entry 109 4e-23
UniRef100_UPI000026030C UPI000026030C UniRef100 entry 108 5e-23
UniRef100_UPI000031C323 UPI000031C323 UniRef100 entry 106 3e-22
UniRef100_O06735 Probable adenylyl-sulfate kinase [Bacillus subt... 103 2e-21
UniRef100_O67174 Probable bifunctional SAT/APS kinase [Includes:... 99 5e-20
UniRef100_UPI00002F3BB3 UPI00002F3BB3 UniRef100 entry 99 7e-20
UniRef100_UPI0000340E6D UPI0000340E6D UniRef100 entry 98 9e-20
UniRef100_UPI00002F681C UPI00002F681C UniRef100 entry 98 9e-20
>UniRef100_O49204 Adenylyl-sulfate kinase, chloroplast precursor [Catharanthus
roseus]
Length = 312
Score = 156 bits (394), Expect = 3e-37
Identities = 75/98 (76%), Positives = 85/98 (86%)
Query: 90 LISPYQKDRDACRALLPEGDFIEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGIDDPY 149
LISPY+K DACR+LLPEGDFIEVF+DVPL VCEARDPKGLYKLARAGKIKGFTGIDDPY
Sbjct: 214 LISPYRKPPDACRSLLPEGDFIEVFMDVPLKVCEARDPKGLYKLARAGKIKGFTGIDDPY 273
Query: 150 EPPCCCEIILQQKGSDCKSPKDMAETVISYLEKSGHLQ 187
EPP EI+L QK C SP D+A+ VISYLE++G+L+
Sbjct: 274 EPPLKSEIVLHQKLGMCDSPCDLADIVISYLEENGYLK 311
>UniRef100_O49196 Adenylyl-sulfate kinase 2, chloroplast precursor [Arabidopsis
thaliana]
Length = 293
Score = 156 bits (394), Expect = 3e-37
Identities = 71/99 (71%), Positives = 86/99 (86%), Gaps = 1/99 (1%)
Query: 90 LISPYQKDRDACRALLPEGDFIEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGIDDPY 149
LISPY++DRDACR+LLP+GDF+EVF+DVPLHVCE+RDPKGLYKLARAGKIKGFTGIDDPY
Sbjct: 194 LISPYRRDRDACRSLLPDGDFVEVFMDVPLHVCESRDPKGLYKLARAGKIKGFTGIDDPY 253
Query: 150 EPPCCCEIILQQKGSD-CKSPKDMAETVISYLEKSGHLQ 187
E P CE++L+ G D SP+ MAE +ISYL+ G+L+
Sbjct: 254 EAPVNCEVVLKHTGDDESCSPRQMAENIISYLQNKGYLE 292
>UniRef100_Q8LES2 Putative adenosine phosphosulfate kinase [Arabidopsis thaliana]
Length = 276
Score = 152 bits (383), Expect = 5e-36
Identities = 73/98 (74%), Positives = 83/98 (84%), Gaps = 2/98 (2%)
Query: 90 LISPYQKDRDACRALLPEGDFIEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGIDDPY 149
LISPY+ DRDACR+LLPEGDF+EVF+DVPL VCEARDPKGLYKLARAGKIKGFTGIDDPY
Sbjct: 180 LISPYRTDRDACRSLLPEGDFVEVFMDVPLSVCEARDPKGLYKLARAGKIKGFTGIDDPY 239
Query: 150 EPPCCCEIILQQKGSDCKSPKDMAETVISYLEKSGHLQ 187
EPP CEI L ++G SP +MAE V+ YL+ G+LQ
Sbjct: 240 EPPLNCEISLGREGG--TSPIEMAEKVVGYLDNKGYLQ 275
>UniRef100_Q43295 Adenylyl-sulfate kinase 1, chloroplast precursor [Arabidopsis
thaliana]
Length = 276
Score = 152 bits (383), Expect = 5e-36
Identities = 73/98 (74%), Positives = 83/98 (84%), Gaps = 2/98 (2%)
Query: 90 LISPYQKDRDACRALLPEGDFIEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGIDDPY 149
LISPY+ DRDACR+LLPEGDF+EVF+DVPL VCEARDPKGLYKLARAGKIKGFTGIDDPY
Sbjct: 180 LISPYRTDRDACRSLLPEGDFVEVFMDVPLSVCEARDPKGLYKLARAGKIKGFTGIDDPY 239
Query: 150 EPPCCCEIILQQKGSDCKSPKDMAETVISYLEKSGHLQ 187
EPP CEI L ++G SP +MAE V+ YL+ G+LQ
Sbjct: 240 EPPLNCEISLGREGG--TSPIEMAEKVVGYLDNKGYLQ 275
>UniRef100_Q9SRW7 Putative adenylylsulfate kinase [Arabidopsis thaliana]
Length = 208
Score = 142 bits (357), Expect = 6e-33
Identities = 68/98 (69%), Positives = 78/98 (79%)
Query: 90 LISPYQKDRDACRALLPEGDFIEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGIDDPY 149
LISPY+KDRDACR ++ FIEVF+++ L +CEARDPKGLYKLARAGKIKGFTGIDDPY
Sbjct: 109 LISPYRKDRDACREMIQNSSFIEVFMNMSLQLCEARDPKGLYKLARAGKIKGFTGIDDPY 168
Query: 150 EPPCCCEIILQQKGSDCKSPKDMAETVISYLEKSGHLQ 187
E P CEI L++K +C SP MAE VISYLE G LQ
Sbjct: 169 ESPLNCEIELKEKEGECPSPVAMAEEVISYLEDKGFLQ 206
>UniRef100_Q9FJX1 Adenylylsulfate kinase-like protein [Arabidopsis thaliana]
Length = 290
Score = 140 bits (354), Expect = 1e-32
Identities = 66/98 (67%), Positives = 81/98 (82%)
Query: 90 LISPYQKDRDACRALLPEGDFIEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGIDDPY 149
LISPY+ +R ACRALLP+GDFIEVF+DVPLHVCEARDPKGLYK ARAGKIKGFTG+DDPY
Sbjct: 183 LISPYRIERAACRALLPQGDFIEVFMDVPLHVCEARDPKGLYKRARAGKIKGFTGVDDPY 242
Query: 150 EPPCCCEIILQQKGSDCKSPKDMAETVISYLEKSGHLQ 187
E P CE+ + S S +MA+ V+SYL+++G+L+
Sbjct: 243 EAPLDCEVHIISNFSSSSSLCEMADIVVSYLDQNGYLK 280
>UniRef100_Q84JF0 Putative adenylylsulfate kinase [Arabidopsis thaliana]
Length = 310
Score = 138 bits (348), Expect = 6e-32
Identities = 69/113 (61%), Positives = 84/113 (74%), Gaps = 15/113 (13%)
Query: 90 LISPYQKDRDACRALLPEGDFIEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGIDDPY 149
LISPY+ +R ACRALLP+GDFIEVF+DVPLHVCEARDPKGLYK ARAGKIKGFTG+DDPY
Sbjct: 188 LISPYRIERAACRALLPQGDFIEVFMDVPLHVCEARDPKGLYKRARAGKIKGFTGVDDPY 247
Query: 150 EPPCCCEIILQ--------QKGSDCKSPK-------DMAETVISYLEKSGHLQ 187
E P CEI++Q S SP +MA+ V+SYL+++G+L+
Sbjct: 248 EAPLDCEIVIQNSRDKGLSSSSSSSSSPSSSSSSLCEMADIVVSYLDQNGYLK 300
>UniRef100_Q8LF64 Adenylylsulfate kinase-like protein [Arabidopsis thaliana]
Length = 305
Score = 138 bits (348), Expect = 6e-32
Identities = 69/113 (61%), Positives = 84/113 (74%), Gaps = 15/113 (13%)
Query: 90 LISPYQKDRDACRALLPEGDFIEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGIDDPY 149
LISPY+ +R ACRALLP+GDFIEVF+DVPLHVCEARDPKGLYK ARAGKIKGFTG+DDPY
Sbjct: 183 LISPYRIERAACRALLPQGDFIEVFMDVPLHVCEARDPKGLYKRARAGKIKGFTGVDDPY 242
Query: 150 EPPCCCEIILQ--------QKGSDCKSPK-------DMAETVISYLEKSGHLQ 187
E P CEI++Q S SP +MA+ V+SYL+++G+L+
Sbjct: 243 EAPLDCEIVIQNSRDKGLSSSSSSSSSPSSSSSSLCEMADIVVSYLDQNGYLK 295
>UniRef100_Q6ZL22 Putative adenosine-5'-phosphosulfate kinase [Oryza sativa]
Length = 345
Score = 135 bits (339), Expect = 7e-31
Identities = 62/96 (64%), Positives = 78/96 (80%)
Query: 91 ISPYQKDRDACRALLPEGDFIEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGIDDPYE 150
ISPY++DR++CRALL +G FIEVF+++PL +CE+RDPKGLYKLARAGKIKGFTGIDDPYE
Sbjct: 248 ISPYRRDRESCRALLSDGSFIEVFLNMPLELCESRDPKGLYKLARAGKIKGFTGIDDPYE 307
Query: 151 PPCCCEIILQQKGSDCKSPKDMAETVISYLEKSGHL 186
P EI +++ C SP DMA V++YLE+ G L
Sbjct: 308 SPLNSEIEIKEVDGVCPSPSDMAGQVVTYLEEKGFL 343
>UniRef100_Q9SE92 Adenosine-5'-phosphosulfate kinase [Zea mays]
Length = 288
Score = 130 bits (327), Expect = 2e-29
Identities = 60/97 (61%), Positives = 77/97 (78%)
Query: 90 LISPYQKDRDACRALLPEGDFIEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGIDDPY 149
LISP+++DR++CRALL + FIEVF+++ L +CEARDPKGLYKLARAGKIKGFTGIDDPY
Sbjct: 190 LISPHRRDRESCRALLSDSSFIEVFLNMSLELCEARDPKGLYKLARAGKIKGFTGIDDPY 249
Query: 150 EPPCCCEIILQQKGSDCKSPKDMAETVISYLEKSGHL 186
E P CEI +++ C P +MA V++YLE+ G L
Sbjct: 250 EAPLNCEIEIKEVDGVCPPPAEMAGQVVTYLEEKGFL 286
>UniRef100_UPI00002E654A UPI00002E654A UniRef100 entry
Length = 214
Score = 114 bits (285), Expect = 1e-24
Identities = 55/98 (56%), Positives = 72/98 (73%), Gaps = 3/98 (3%)
Query: 86 TKDCLISPYQKDRDACRALLPEGDFIEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGI 145
T ISPY+ DR+A R +LPE DFIEV++ PL +CE+RDPKGLYK ARAG+IKGFTGI
Sbjct: 114 TLTAFISPYRSDREAVRKMLPEADFIEVYVKAPLDICESRDPKGLYKKARAGEIKGFTGI 173
Query: 146 DDPYEPPCCCEIILQQKGSDCKSPKDMAETVISYLEKS 183
DDPYE P EI ++ S SP+ +A+ V+ +LE++
Sbjct: 174 DDPYEEPESAEITVE---STVNSPQVLAQQVLVWLERA 208
>UniRef100_Q73WR1 Hypothetical protein [Mycobacterium paratuberculosis]
Length = 230
Score = 112 bits (281), Expect = 4e-24
Identities = 56/94 (59%), Positives = 69/94 (72%), Gaps = 3/94 (3%)
Query: 91 ISPYQKDRDACRALLPEGDFIEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGIDDPYE 150
ISPY +DRDA RA L +GDF E+FID P+ +CE RDPKGLYK ARAG+IKGFTGIDDPYE
Sbjct: 122 ISPYVRDRDAIRATLDDGDFQEIFIDTPIEICEKRDPKGLYKKARAGEIKGFTGIDDPYE 181
Query: 151 PPCCCEIILQQKGSDCKSPKDMAETVISYLEKSG 184
P E+ L D ++ +AE VI++LE+ G
Sbjct: 182 APPRPELRLDGAAKDAET---LAEEVIAHLERVG 212
>UniRef100_UPI0000278622 UPI0000278622 UniRef100 entry
Length = 233
Score = 109 bits (272), Expect = 4e-23
Identities = 52/98 (53%), Positives = 71/98 (72%), Gaps = 3/98 (3%)
Query: 86 TKDCLISPYQKDRDACRALLPEGDFIEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGI 145
T ISPY+ DR+A R +LPE DF+EV++ PL +CE+RDPKGLYK ARAG+I GFTGI
Sbjct: 133 TLTAFISPYRSDREAVRKMLPEADFVEVYVKAPLDICESRDPKGLYKKARAGEISGFTGI 192
Query: 146 DDPYEPPCCCEIILQQKGSDCKSPKDMAETVISYLEKS 183
DDPYE P EI ++ S +P+ +A+ V+ +LE++
Sbjct: 193 DDPYEEPEKAEITVE---SAVHNPQVLAQQVLVWLERA 227
>UniRef100_UPI000026030C UPI000026030C UniRef100 entry
Length = 195
Score = 108 bits (271), Expect = 5e-23
Identities = 53/98 (54%), Positives = 70/98 (71%), Gaps = 3/98 (3%)
Query: 86 TKDCLISPYQKDRDACRALLPEGDFIEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGI 145
T ISPY+ DR+A R +LP DFIEV++ PL +CE+RDPKGLYK ARAG+I GFTGI
Sbjct: 95 TLTAFISPYRSDREAVRKMLPAADFIEVYVKAPLDICESRDPKGLYKKARAGEISGFTGI 154
Query: 146 DDPYEPPCCCEIILQQKGSDCKSPKDMAETVISYLEKS 183
D PYE P EI ++ S KSP+ +AE ++ +LE++
Sbjct: 155 DAPYEEPEKAEITVE---STTKSPQVLAEQILVWLERA 189
>UniRef100_UPI000031C323 UPI000031C323 UniRef100 entry
Length = 199
Score = 106 bits (265), Expect = 3e-22
Identities = 52/90 (57%), Positives = 67/90 (73%), Gaps = 3/90 (3%)
Query: 91 ISPYQKDRDACRALLPEGDFIEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGIDDPYE 150
+SPY++DRDA R L EGDF+EVF+D PL +CE RDPKGLY+ ARAG+I+G TGIDDPYE
Sbjct: 103 VSPYRRDRDAVRNRLEEGDFVEVFVDAPLEICEQRDPKGLYRKARAGEIQGMTGIDDPYE 162
Query: 151 PPCCCEIILQQKGSDCKSPKDMAETVISYL 180
P E++L G D S + +A+ V+ YL
Sbjct: 163 SPERPELVL-PSGQD--SAESLADEVLRYL 189
>UniRef100_O06735 Probable adenylyl-sulfate kinase [Bacillus subtilis]
Length = 199
Score = 103 bits (257), Expect = 2e-21
Identities = 50/94 (53%), Positives = 66/94 (70%), Gaps = 3/94 (3%)
Query: 91 ISPYQKDRDACRALLPEGDFIEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGIDDPYE 150
ISP+++DRD RAL P+G+F E+++ PLHVCE RDPKGLYK AR G+IK FTGID PYE
Sbjct: 107 ISPFREDRDMVRALFPKGEFFEIYVKCPLHVCEQRDPKGLYKKARNGEIKHFTGIDSPYE 166
Query: 151 PPCCCEIILQQKGSDCKSPKDMAETVISYLEKSG 184
P + I++ SD S D A+ +I+ L+ G
Sbjct: 167 APLSPDFIIE---SDQTSISDGADLIINALQNRG 197
>UniRef100_O67174 Probable bifunctional SAT/APS kinase [Includes: Sulfate
adenylyltransferase (EC 2.7.7.4) (Sulfate adenylate
transferase) (SAT) (ATP-sulfurylase); Adenylyl-sulfate
kinase (EC 2.7.1.25) (APS kinase)
(Adenosine-5'phosphosulfate kinase) (ATP
adenosine-5'-phosphosulfate 3'-phosphotransferase)]
[Aquifex aeolicus]
Length = 546
Score = 99.0 bits (245), Expect = 5e-20
Identities = 46/98 (46%), Positives = 67/98 (67%), Gaps = 3/98 (3%)
Query: 90 LISPYQKDRDACRALLPEGDFIEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGIDDPY 149
L+SPY+ R+ R ++ EG FIEVF+D P+ VCE RD KGLYK A+ G IKGFTG+DDPY
Sbjct: 451 LVSPYRSARNQVRNMMEEGKFIEVFVDAPVEVCEERDVKGLYKKAKEGLIKGFTGVDDPY 510
Query: 150 EPPCCCEIILQQKGSDCKSPKDMAETVISYLEKSGHLQ 187
EPP E+ + + +P++ A ++ +L+K G ++
Sbjct: 511 EPPVAPEV---RVDTTKLTPEESALKILEFLKKEGFIK 545
>UniRef100_UPI00002F3BB3 UPI00002F3BB3 UniRef100 entry
Length = 189
Score = 98.6 bits (244), Expect = 7e-20
Identities = 52/93 (55%), Positives = 66/93 (70%), Gaps = 3/93 (3%)
Query: 91 ISPYQKDRDACRALLPEGDFIEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGIDDPYE 150
ISP++K RD R L +FIEVF+D P+ +CE RDPKGLYK AR+G IK FTGID YE
Sbjct: 96 ISPFKKQRDNTRNLFERKEFIEVFLDTPIGICEKRDPKGLYKKARSGLIKDFTGIDSAYE 155
Query: 151 PPCCCEIILQQKGSDCKSPKDMAETVISYLEKS 183
P EIIL GS K+P+D++E +I+YL K+
Sbjct: 156 KPENPEIILD--GSQ-KTPEDLSEEIINYLLKN 185
>UniRef100_UPI0000340E6D UPI0000340E6D UniRef100 entry
Length = 160
Score = 98.2 bits (243), Expect = 9e-20
Identities = 48/94 (51%), Positives = 65/94 (69%), Gaps = 3/94 (3%)
Query: 91 ISPYQKDRDACRALLPEGDFIEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGIDDPYE 150
ISP++ DRD CR+L+ +G+F+EVFID PL VCE RDPKGLYK AR+G+IK FTGID YE
Sbjct: 68 ISPFKADRDYCRSLMEDGEFVEVFIDTPLEVCEKRDPKGLYKKARSGEIKDFTGIDSAYE 127
Query: 151 PPCCCEIILQQKGSDCKSPKDMAETVISYLEKSG 184
P E+ L + + + AE + + L++ G
Sbjct: 128 APEAPEVHLTYQD---EPAEQTAERLYALLQEKG 158
>UniRef100_UPI00002F681C UPI00002F681C UniRef100 entry
Length = 198
Score = 98.2 bits (243), Expect = 9e-20
Identities = 48/94 (51%), Positives = 65/94 (69%), Gaps = 3/94 (3%)
Query: 91 ISPYQKDRDACRALLPEGDFIEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGIDDPYE 150
ISP++ DRD CR+L+ +G+F+EVFID PL VCE RDPKGLYK AR+G+IK FTGID YE
Sbjct: 106 ISPFKADRDYCRSLMEDGEFVEVFIDTPLEVCEKRDPKGLYKKARSGEIKDFTGIDSAYE 165
Query: 151 PPCCCEIILQQKGSDCKSPKDMAETVISYLEKSG 184
P E+ L + + + AE + + L++ G
Sbjct: 166 APEAPEVHLTYQD---EPAEQTAERLYALLQEKG 196
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.340 0.150 0.512
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 299,517,841
Number of Sequences: 2790947
Number of extensions: 11353563
Number of successful extensions: 26186
Number of sequences better than 10.0: 341
Number of HSP's better than 10.0 without gapping: 334
Number of HSP's successfully gapped in prelim test: 7
Number of HSP's that attempted gapping in prelim test: 25779
Number of HSP's gapped (non-prelim): 344
length of query: 187
length of database: 848,049,833
effective HSP length: 120
effective length of query: 67
effective length of database: 513,136,193
effective search space: 34380124931
effective search space used: 34380124931
T: 11
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 71 (32.0 bits)
Medicago: description of AC147000.4


