

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146972.10 - phase: 0
(106 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
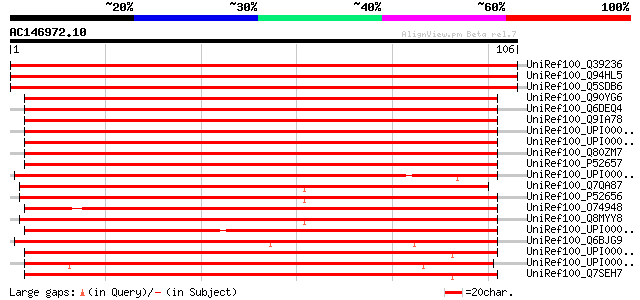
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q39236 Transcription initiation factor IIA gamma chain... 198 3e-50
UniRef100_Q94HL5 Transcription initiation factor IIA gamma chain... 191 5e-48
UniRef100_Q5SDB6 Transcription factor IIA gamma subunit [Oryza s... 188 2e-47
UniRef100_Q90YG6 Transcription initiation factor IIA gamma chain... 114 5e-25
UniRef100_Q6DEQ4 MGC89923 protein [Xenopus tropicalis] 113 9e-25
UniRef100_Q9IA78 Transcription initiation factor IIA gamma chain... 113 9e-25
UniRef100_UPI000019D145 UPI000019D145 UniRef100 entry 113 1e-24
UniRef100_UPI000036074D UPI000036074D UniRef100 entry 113 1e-24
UniRef100_Q80ZM7 Putative transcription initiation factor IIA ga... 112 2e-24
UniRef100_P52657 Transcription initiation factor IIA gamma chain... 112 2e-24
UniRef100_UPI0000235579 UPI0000235579 UniRef100 entry 110 1e-23
UniRef100_Q7QA87 ENSANGP00000013128 [Anopheles gambiae str. PEST] 110 1e-23
UniRef100_P52656 Transcription initiation factor IIA gamma chain... 107 7e-23
UniRef100_O74948 Probable transcription initiation factor IIA sm... 107 9e-23
UniRef100_Q8MYY8 RE44302p [Drosophila melanogaster] 105 3e-22
UniRef100_UPI00001815AC UPI00001815AC UniRef100 entry 104 4e-22
UniRef100_Q6BJG9 Similar to CA5187|CaTOA2 Candida albicans CaTOA... 103 7e-22
UniRef100_UPI000021B7D1 UPI000021B7D1 UniRef100 entry 102 2e-21
UniRef100_UPI000042DF18 UPI000042DF18 UniRef100 entry 101 4e-21
UniRef100_Q7SEH7 Hypothetical protein [Neurospora crassa] 99 2e-20
>UniRef100_Q39236 Transcription initiation factor IIA gamma chain [Arabidopsis
thaliana]
Length = 106
Score = 198 bits (503), Expect = 3e-50
Identities = 96/106 (90%), Positives = 104/106 (97%)
Query: 1 MATFELYRRSTIGMCLTETLDEMVQNGTLSPEIAIQVLVQFDKSMTEALETQVKSKVSIK 60
MATFELYRRSTIGMCLTETLDEMVQ+GTLSPE+AIQVLVQFDKSMTEALE+QVK+KVSIK
Sbjct: 1 MATFELYRRSTIGMCLTETLDEMVQSGTLSPELAIQVLVQFDKSMTEALESQVKTKVSIK 60
Query: 61 GHLHTYRFCDNVWTFILQDALFKNEDNQENVGRVKIVACDSKLLSQ 106
GHLHTYRFCDNVWTFILQDA+FK++D QENV RVKIVACDSKLL+Q
Sbjct: 61 GHLHTYRFCDNVWTFILQDAMFKSDDRQENVSRVKIVACDSKLLTQ 106
>UniRef100_Q94HL5 Transcription initiation factor IIA gamma chain [Oryza sativa]
Length = 106
Score = 191 bits (484), Expect = 5e-48
Identities = 94/106 (88%), Positives = 99/106 (92%)
Query: 1 MATFELYRRSTIGMCLTETLDEMVQNGTLSPEIAIQVLVQFDKSMTEALETQVKSKVSIK 60
MATFELYRRSTIGMCLTETLDEMV +GTLSPE+AIQVLVQFDKSMTEALE QVKSKVSIK
Sbjct: 1 MATFELYRRSTIGMCLTETLDEMVSSGTLSPELAIQVLVQFDKSMTEALENQVKSKVSIK 60
Query: 61 GHLHTYRFCDNVWTFILQDALFKNEDNQENVGRVKIVACDSKLLSQ 106
GHLHTYRFCDNVWTFIL +A FKNE+ E VG+VKIVACDSKLLSQ
Sbjct: 61 GHLHTYRFCDNVWTFILTEASFKNEETTEQVGKVKIVACDSKLLSQ 106
>UniRef100_Q5SDB6 Transcription factor IIA gamma subunit [Oryza sativa]
Length = 106
Score = 188 bits (478), Expect = 2e-47
Identities = 93/106 (87%), Positives = 98/106 (91%)
Query: 1 MATFELYRRSTIGMCLTETLDEMVQNGTLSPEIAIQVLVQFDKSMTEALETQVKSKVSIK 60
MATFELYRRSTIGMCLTETLDEMV +GTLSPE+AIQVL QFDKSMTEALE QVKSKVSIK
Sbjct: 1 MATFELYRRSTIGMCLTETLDEMVSSGTLSPELAIQVLEQFDKSMTEALENQVKSKVSIK 60
Query: 61 GHLHTYRFCDNVWTFILQDALFKNEDNQENVGRVKIVACDSKLLSQ 106
GHLHTYRFCDNVWTFIL +A FKNE+ E VG+VKIVACDSKLLSQ
Sbjct: 61 GHLHTYRFCDNVWTFILTEASFKNEETTEQVGKVKIVACDSKLLSQ 106
>UniRef100_Q90YG6 Transcription initiation factor IIA gamma chain [Oncorhynchus
mykiss]
Length = 108
Score = 114 bits (285), Expect = 5e-25
Identities = 50/99 (50%), Positives = 75/99 (75%)
Query: 4 FELYRRSTIGMCLTETLDEMVQNGTLSPEIAIQVLVQFDKSMTEALETQVKSKVSIKGHL 63
++LYR +T+G L E+LDE++Q ++P++A+QVL+QFDK++ AL +V+++V+ KG L
Sbjct: 3 YQLYRNTTLGNSLQESLDELIQTQQITPQLALQVLLQFDKAINTALANRVRNRVNFKGSL 62
Query: 64 HTYRFCDNVWTFILQDALFKNEDNQENVGRVKIVACDSK 102
+TYRFCDNVWTF+L D F+ + V +VKIVACD K
Sbjct: 63 NTYRFCDNVWTFVLNDVEFREVTDLVKVDKVKIVACDGK 101
>UniRef100_Q6DEQ4 MGC89923 protein [Xenopus tropicalis]
Length = 109
Score = 113 bits (283), Expect = 9e-25
Identities = 49/99 (49%), Positives = 76/99 (76%)
Query: 4 FELYRRSTIGMCLTETLDEMVQNGTLSPEIAIQVLVQFDKSMTEALETQVKSKVSIKGHL 63
++LYR +T+G L E+LDE++Q+ ++P++A+QVL+QFDK++ AL +V+++V+ +G L
Sbjct: 3 YQLYRNTTLGNSLQESLDELIQSQQINPQLALQVLLQFDKAINSALAQRVRNRVNFRGSL 62
Query: 64 HTYRFCDNVWTFILQDALFKNEDNQENVGRVKIVACDSK 102
+TYRFCDNVWTF+L D F+ + V +VKIVACD K
Sbjct: 63 NTYRFCDNVWTFVLNDVEFREVTDLVKVDKVKIVACDGK 101
>UniRef100_Q9IA78 Transcription initiation factor IIA gamma chain [Paralichthys
olivaceus]
Length = 111
Score = 113 bits (283), Expect = 9e-25
Identities = 49/99 (49%), Positives = 76/99 (76%)
Query: 4 FELYRRSTIGMCLTETLDEMVQNGTLSPEIAIQVLVQFDKSMTEALETQVKSKVSIKGHL 63
++LYR +T+G L E+LDE++Q ++P++A+QVL+QFDK++ AL ++V+++V+ +G L
Sbjct: 3 YQLYRNTTLGNSLQESLDELIQTQQITPQLALQVLLQFDKAINTALASRVRNRVNFRGSL 62
Query: 64 HTYRFCDNVWTFILQDALFKNEDNQENVGRVKIVACDSK 102
+TYRFCDNVWTF+L D F+ + V +VKIVACD K
Sbjct: 63 NTYRFCDNVWTFVLNDVEFREVTDLVKVDKVKIVACDGK 101
>UniRef100_UPI000019D145 UPI000019D145 UniRef100 entry
Length = 109
Score = 113 bits (282), Expect = 1e-24
Identities = 49/99 (49%), Positives = 75/99 (75%)
Query: 4 FELYRRSTIGMCLTETLDEMVQNGTLSPEIAIQVLVQFDKSMTEALETQVKSKVSIKGHL 63
++LYR +T+G L E+LDE++Q ++P++A+QVL+QFDK++ AL +V+++V+ +G L
Sbjct: 3 YQLYRNTTLGNSLQESLDELIQTQQITPQLALQVLLQFDKAINTALANRVRNRVNFRGSL 62
Query: 64 HTYRFCDNVWTFILQDALFKNEDNQENVGRVKIVACDSK 102
+TYRFCDNVWTF+L D F+ + V +VKIVACD K
Sbjct: 63 NTYRFCDNVWTFVLNDVEFREVTDLVKVDKVKIVACDGK 101
>UniRef100_UPI000036074D UPI000036074D UniRef100 entry
Length = 111
Score = 113 bits (282), Expect = 1e-24
Identities = 49/99 (49%), Positives = 75/99 (75%)
Query: 4 FELYRRSTIGMCLTETLDEMVQNGTLSPEIAIQVLVQFDKSMTEALETQVKSKVSIKGHL 63
++LYR +T+G L E+LDE++Q ++P++A+QVL+QFDK++ AL +V+++V+ +G L
Sbjct: 3 YQLYRNTTLGNSLQESLDELIQTQQITPQLALQVLLQFDKAINTALANRVRNRVNFRGSL 62
Query: 64 HTYRFCDNVWTFILQDALFKNEDNQENVGRVKIVACDSK 102
+TYRFCDNVWTF+L D F+ + V +VKIVACD K
Sbjct: 63 NTYRFCDNVWTFVLNDVEFREVTDLVKVDKVKIVACDGK 101
>UniRef100_Q80ZM7 Putative transcription initiation factor IIA gamma chain [Mus
musculus]
Length = 109
Score = 112 bits (281), Expect = 2e-24
Identities = 49/99 (49%), Positives = 75/99 (75%)
Query: 4 FELYRRSTIGMCLTETLDEMVQNGTLSPEIAIQVLVQFDKSMTEALETQVKSKVSIKGHL 63
++LYR +T+G L E+LDE++Q+ ++P++A+QVL+QFDK++ AL +V+++V+ +G L
Sbjct: 3 YQLYRNTTLGNSLQESLDELIQSQQITPQLALQVLLQFDKAINSALAQRVRNRVNFRGSL 62
Query: 64 HTYRFCDNVWTFILQDALFKNEDNQENVGRVKIVACDSK 102
+TYRFCDNVWTF+L D F+ V +VKIVACD K
Sbjct: 63 NTYRFCDNVWTFVLNDVEFREVTELIKVDKVKIVACDGK 101
>UniRef100_P52657 Transcription initiation factor IIA gamma chain [Homo sapiens]
Length = 109
Score = 112 bits (280), Expect = 2e-24
Identities = 49/99 (49%), Positives = 75/99 (75%)
Query: 4 FELYRRSTIGMCLTETLDEMVQNGTLSPEIAIQVLVQFDKSMTEALETQVKSKVSIKGHL 63
++LYR +T+G L E+LDE++Q+ ++P++A+QVL+QFDK++ AL +V+++V+ +G L
Sbjct: 3 YQLYRNTTLGNSLQESLDELIQSQQITPQLALQVLLQFDKAINAALAQRVRNRVNFRGSL 62
Query: 64 HTYRFCDNVWTFILQDALFKNEDNQENVGRVKIVACDSK 102
+TYRFCDNVWTF+L D F+ V +VKIVACD K
Sbjct: 63 NTYRFCDNVWTFVLNDVEFREVTELIKVDKVKIVACDGK 101
>UniRef100_UPI0000235579 UPI0000235579 UniRef100 entry
Length = 111
Score = 110 bits (274), Expect = 1e-23
Identities = 49/103 (47%), Positives = 79/103 (76%), Gaps = 3/103 (2%)
Query: 2 ATFELYRRSTIGMCLTETLDEMVQNGTLSPEIAIQVLVQFDKSMTEALETQVKSKVSIKG 61
A +ELYR S++G+ LT+TLD+++ G + P++A+++L FD+ +TE L +V+++++ KG
Sbjct: 5 AYYELYRGSSLGLSLTDTLDDLINEGRIEPQLAMKILSTFDRVITEVLADKVRTRLTFKG 64
Query: 62 HLHTYRFCDNVWTFILQDALFKNEDNQENVG--RVKIVACDSK 102
HL TYRFCD VWTF+++D FK DNQ+ + +VKIV+C+SK
Sbjct: 65 HLDTYRFCDEVWTFLIKDVNFK-LDNQQTISADKVKIVSCNSK 106
>UniRef100_Q7QA87 ENSANGP00000013128 [Anopheles gambiae str. PEST]
Length = 112
Score = 110 bits (274), Expect = 1e-23
Identities = 52/99 (52%), Positives = 73/99 (73%), Gaps = 1/99 (1%)
Query: 3 TFELYRRSTIGMCLTETLDEMVQNGTLSPEIAIQVLVQFDKSMTEALETQVKSKVSIK-G 61
T++LYR +T+G L E+LDE++Q G ++P++A++VLVQFDKS+ AL +VKS+V+ K
Sbjct: 2 TYQLYRNTTLGNTLQESLDELIQYGQITPQLAVRVLVQFDKSINAALSNRVKSRVTFKAA 61
Query: 62 HLHTYRFCDNVWTFILQDALFKNEDNQENVGRVKIVACD 100
L+TYRFCDNVWT +L D F+ V +VKIVACD
Sbjct: 62 KLNTYRFCDNVWTLMLNDVEFREVHEFARVDKVKIVACD 100
>UniRef100_P52656 Transcription initiation factor IIA gamma chain [Drosophila
melanogaster]
Length = 106
Score = 107 bits (267), Expect = 7e-23
Identities = 51/101 (50%), Positives = 73/101 (71%), Gaps = 1/101 (0%)
Query: 3 TFELYRRSTIGMCLTETLDEMVQNGTLSPEIAIQVLVQFDKSMTEALETQVKSKVSIK-G 61
+++LYR +T+G L E+LDE++Q G ++P +A +VL+QFDKS+ AL +VK++V+ K G
Sbjct: 2 SYQLYRNTTLGNTLQESLDELIQYGQITPGLAFKVLLQFDKSINNALNQRVKARVTFKAG 61
Query: 62 HLHTYRFCDNVWTFILQDALFKNEDNQENVGRVKIVACDSK 102
L+TYRFCDNVWT +L D F+ V +VKIVACD K
Sbjct: 62 KLNTYRFCDNVWTLMLNDVEFREVHEIVKVDKVKIVACDGK 102
>UniRef100_O74948 Probable transcription initiation factor IIA small chain
[Schizosaccharomyces pombe]
Length = 107
Score = 107 bits (266), Expect = 9e-23
Identities = 48/99 (48%), Positives = 74/99 (74%), Gaps = 2/99 (2%)
Query: 4 FELYRRSTIGMCLTETLDEMVQNGTLSPEIAIQVLVQFDKSMTEALETQVKSKVSIKGHL 63
+ELYRRS I LT+ LD+++ G +SP++A++VL FDKSMTEAL +V+S+++ KGHL
Sbjct: 5 YELYRRSRIS--LTDALDDLISQGKISPQLAMKVLFNFDKSMTEALAEKVRSRLTFKGHL 62
Query: 64 HTYRFCDNVWTFILQDALFKNEDNQENVGRVKIVACDSK 102
TYRFCD VWTFI+++ F+ ++ +++IVAC ++
Sbjct: 63 DTYRFCDEVWTFIIKNPSFRFDNETVTSNKIRIVACATR 101
>UniRef100_Q8MYY8 RE44302p [Drosophila melanogaster]
Length = 106
Score = 105 bits (261), Expect = 3e-22
Identities = 50/101 (49%), Positives = 72/101 (70%), Gaps = 1/101 (0%)
Query: 3 TFELYRRSTIGMCLTETLDEMVQNGTLSPEIAIQVLVQFDKSMTEALETQVKSKVSIK-G 61
+++LYR +T+G L E+LDE++Q G ++P +A +VL+Q DKS+ AL +VK++V+ K G
Sbjct: 2 SYQLYRNTTLGNTLQESLDELIQYGQITPGLAFKVLLQLDKSINNALNQRVKARVTFKAG 61
Query: 62 HLHTYRFCDNVWTFILQDALFKNEDNQENVGRVKIVACDSK 102
L+TYRFCDNVWT +L D F+ V +VKIVACD K
Sbjct: 62 KLNTYRFCDNVWTLMLNDVEFREVHEIVKVDKVKIVACDGK 102
>UniRef100_UPI00001815AC UPI00001815AC UniRef100 entry
Length = 108
Score = 104 bits (260), Expect = 4e-22
Identities = 48/99 (48%), Positives = 72/99 (72%), Gaps = 1/99 (1%)
Query: 4 FELYRRSTIGMCLTETLDEMVQNGTLSPEIAIQVLVQFDKSMTEALETQVKSKVSIKGHL 63
++LYR + +G L E LDE++Q+ ++P++A+QVL+QFDK+ AL +V+++V+ +G L
Sbjct: 3 YQLYRNTILGNSLQERLDELIQSQQITPQLALQVLLQFDKA-NSALAQRVRNRVNFRGSL 61
Query: 64 HTYRFCDNVWTFILQDALFKNEDNQENVGRVKIVACDSK 102
+TYRFCDNVWTF+L D F+ V +VKIVACD K
Sbjct: 62 NTYRFCDNVWTFVLNDVEFREVTELIKVDKVKIVACDGK 100
>UniRef100_Q6BJG9 Similar to CA5187|CaTOA2 Candida albicans CaTOA2 TFIIA subunit 13.5
kD [Debaryomyces hansenii]
Length = 124
Score = 103 bits (258), Expect = 7e-22
Identities = 52/115 (45%), Positives = 78/115 (67%), Gaps = 14/115 (12%)
Query: 2 ATFELYRRSTIGMCLTETLDEMVQNGTLSPEIAIQVLVQFDKSMTEALETQV---KSKVS 58
A +ELYRRSTIG LT+ LD ++ + + P++A+++L FD+ ++E L+ ++ KSK++
Sbjct: 5 AYYELYRRSTIGTTLTDALDTLISDEKIQPQLAMRILNNFDRIISELLKNELGISKSKLT 64
Query: 59 IKGHLHTYRFCDNVWTFILQDALFK-----------NEDNQENVGRVKIVACDSK 102
KG LHTYRFCD+VWTFI+++ L K DN+ NV + KIVAC+SK
Sbjct: 65 FKGDLHTYRFCDDVWTFIIKNVLIKLTDVSNSGNNDLNDNEINVDKFKIVACNSK 119
>UniRef100_UPI000021B7D1 UPI000021B7D1 UniRef100 entry
Length = 116
Score = 102 bits (255), Expect = 2e-21
Identities = 41/101 (40%), Positives = 76/101 (74%), Gaps = 2/101 (1%)
Query: 4 FELYRRSTIGMCLTETLDEMVQNGTLSPEIAIQVLVQFDKSMTEALETQVKSKVSIKGHL 63
+ELYRR+++G+CLT+ LD+++ N ++P++A+++L FD+ + E L+ +VK+++ KG L
Sbjct: 10 YELYRRTSLGICLTDALDDLITNDRINPQLAMKILANFDRVVAETLQEKVKARLQFKGAL 69
Query: 64 HTYRFCDNVWTFILQDALFKNEDNQENV--GRVKIVACDSK 102
YRFCD+VWTF++++ FK + + + +VKIV+C++K
Sbjct: 70 DNYRFCDDVWTFVIKNINFKLDGGNQTIQADKVKIVSCNAK 110
>UniRef100_UPI000042DF18 UPI000042DF18 UniRef100 entry
Length = 127
Score = 101 bits (252), Expect = 4e-21
Identities = 51/106 (48%), Positives = 70/106 (65%), Gaps = 8/106 (7%)
Query: 4 FELYRRST---IGMCLTETLDEMVQNGTLSPEIAIQVLVQFDKSMTEALETQVKSKVSIK 60
+E YR S IG LT+ LDE++ G + P++A++VL QFDKS+TE L+ VK+K +IK
Sbjct: 8 YEFYRGSRHVHIGTALTDALDELITQGDIPPQLAMRVLQQFDKSLTECLQKGVKNKTTIK 67
Query: 61 GHLHTYRFCDNVWTFILQDALFKNE-----DNQENVGRVKIVACDS 101
GHL TYR CD+VWTF+++D FK E ++KIVAC S
Sbjct: 68 GHLSTYRLCDDVWTFVVKDPQFKMEGVGAGSEMVTGSKIKIVACKS 113
>UniRef100_Q7SEH7 Hypothetical protein [Neurospora crassa]
Length = 115
Score = 99.4 bits (246), Expect = 2e-20
Identities = 44/101 (43%), Positives = 73/101 (71%), Gaps = 2/101 (1%)
Query: 4 FELYRRSTIGMCLTETLDEMVQNGTLSPEIAIQVLVQFDKSMTEALETQVKSKVSIKGHL 63
++LYR ++G LT+ LD+++ + P++A++VL+QFD+ +TEAL +VK++++ KG L
Sbjct: 10 YDLYRHGSLGSTLTDALDDLIGAERIDPQLAMKVLMQFDRVITEALSEKVKARLTFKGSL 69
Query: 64 HTYRFCDNVWTFILQDALFKNEDNQENV--GRVKIVACDSK 102
TYRFCD VWTF++++ FK + Q V +VKIV+C +K
Sbjct: 70 DTYRFCDEVWTFLIKNVTFKMDGGQSVVTADKVKIVSCSAK 110
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.320 0.133 0.377
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 153,693,893
Number of Sequences: 2790947
Number of extensions: 4764728
Number of successful extensions: 12055
Number of sequences better than 10.0: 43
Number of HSP's better than 10.0 without gapping: 39
Number of HSP's successfully gapped in prelim test: 4
Number of HSP's that attempted gapping in prelim test: 11997
Number of HSP's gapped (non-prelim): 45
length of query: 106
length of database: 848,049,833
effective HSP length: 82
effective length of query: 24
effective length of database: 619,192,179
effective search space: 14860612296
effective search space used: 14860612296
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 68 (30.8 bits)
Medicago: description of AC146972.10


