

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146971.7 - phase: 0 /pseudo
(2360 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
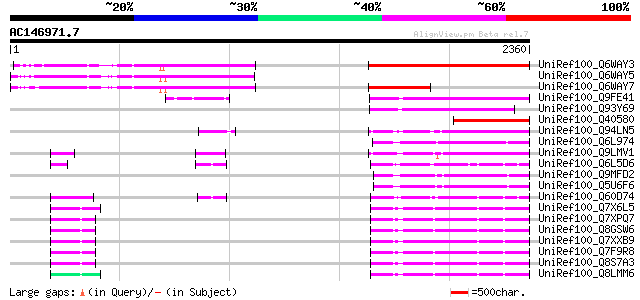
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q6WAY3 Gag/pol polyprotein [Pisum sativum] 1202 0.0
UniRef100_Q6WAY5 Gag/pol polyprotein [Pisum sativum] 589 e-166
UniRef100_Q6WAY7 Gag/pol polyprotein [Pisum sativum] 588 e-166
UniRef100_Q9FE41 Oryza sativa (japonica cultivar-group) genomic ... 520 e-145
UniRef100_Q93Y69 Putative gag-pol [Oryza sativa] 478 e-132
UniRef100_Q40580 Integrase [Nicotiana tabacum] 449 e-124
UniRef100_Q94LN5 Putative retroelement pol polyprotein [Oryza sa... 448 e-124
UniRef100_Q6L974 GAG-POL [Vitis vinifera] 430 e-118
UniRef100_Q9LMV1 F5M15.26 [Arabidopsis thaliana] 399 e-109
UniRef100_Q6L5D6 Putative polyprotein [Oryza sativa] 383 e-104
UniRef100_Q9MFD2 Orf764 protein [Beta vulgaris subsp. vulgaris] 383 e-104
UniRef100_Q5U6F6 Orf764 protein [Beta vulgaris subsp. vulgaris] 383 e-104
UniRef100_Q60D74 Putative polyprotein [Oryza sativa] 382 e-104
UniRef100_Q7X6L5 OSJNBb0093G06.3 protein [Oryza sativa] 375 e-102
UniRef100_Q7XPQ7 OSJNBa0053K19.16 protein [Oryza sativa] 375 e-102
UniRef100_Q8GSW6 Putative gag-pol polyprotein [Oryza sativa] 375 e-101
UniRef100_Q7XXB9 OSJNBa0027O01.4 protein [Oryza sativa] 374 e-101
UniRef100_Q7F9R8 OSJNBb0054B09.2 protein [Oryza sativa] 374 e-101
UniRef100_Q8S7A3 Putative retroelement [Oryza sativa] 374 e-101
UniRef100_Q8LMM6 Putative gag-pol [Oryza sativa] 373 e-101
>UniRef100_Q6WAY3 Gag/pol polyprotein [Pisum sativum]
Length = 2262
Score = 1202 bits (3109), Expect = 0.0
Identities = 560/730 (76%), Positives = 652/730 (88%), Gaps = 2/730 (0%)
Query: 1633 ENSMGCVLGQQDETGRKEHAIYYLSKKFTECESRYSMLEKTCCALAWAAKRLRHYMINHT 1692
ENSMGCVLG+ DE+GRKEHAIYYLSKKFT+CE+RYS+LEKTCCALAWAA+RLR YM+NHT
Sbjct: 1532 ENSMGCVLGRHDESGRKEHAIYYLSKKFTDCETRYSLLEKTCCALAWAARRLRQYMLNHT 1591
Query: 1693 TWLISKMDPIKYIFEKPALIGRIARWQMLLSEYDIEYRSQKAIKGSILADHLAHQPLEDY 1752
T LISKMDP+KYIFEKPAL GR+ARWQM+L+EYDI+Y SQKAIKGSIL+D+LA QP+EDY
Sbjct: 1592 TLLISKMDPVKYIFEKPALTGRVARWQMILTEYDIQYTSQKAIKGSILSDYLAEQPIEDY 1651
Query: 1753 RPIKFDFPDEEIMYLKMKDCDEPLFGEGPDPDSVWGLIFDGAVNVYGNGIGAVLLTPKGT 1812
+P+ F+FPDE+IMYLKMKDC EPL EGPDPD W L+FDGAVN+ GNG+GAVL+ PKG
Sbjct: 1652 QPMMFEFPDEDIMYLKMKDCKEPLVEEGPDPDDKWTLMFDGAVNMNGNGVGAVLINPKGA 1711
Query: 1813 HIPFTARLRFDCTNNIAEYEACIMGIEEAIDLRIKNIEIYGDSALVINQIKGKWETLHAG 1872
H+PF+ARL FD TNN AEYEACIMGIEEAIDLRIK ++I+GDSALV+NQ+ G W T
Sbjct: 1712 HMPFSARLTFDVTNNEAEYEACIMGIEEAIDLRIKTLDIFGDSALVVNQVNGDWNTNQPH 1771
Query: 1873 LIPYRDYARRLLTFFNKVELHHIPRDENQMADALATLSSMIKVNHHNDVPLISVKFLDRP 1932
LIPYRDY RR+LTFF KV+L+H+PRDENQMADALATLSSMIKVN N VP ++V L+RP
Sbjct: 1772 LIPYRDYTRRILTFFKKVKLYHVPRDENQMADALATLSSMIKVNWWNHVPHVAVNRLERP 1831
Query: 1933 AYVFAAE-VVFDDKPWFHDIKVFLQTREYPPGASNKDKKTLRRLSSNFFLN-GDILYKRN 1990
AYVFAAE VV D+KPW++DIK FL+T+EYP GAS DKKTLRRL+ +F+LN D+LYKRN
Sbjct: 1832 AYVFAAESVVIDEKPWYYDIKNFLKTQEYPEGASKNDKKTLRRLAGSFYLNQDDVLYKRN 1891
Query: 1991 FDTVLLRCVDKYEADLLIHEIHEGSFGIHPNGHTMAKKILRAGYYWMTMESDCYKHTRKC 2050
FD VLLRC+D+ EAD+L+ E+HEGSFG H GH MAKK+LRAGYYWMTMESDC+K+ RKC
Sbjct: 1892 FDMVLLRCMDRPEADMLMQEVHEGSFGTHAGGHAMAKKLLRAGYYWMTMESDCFKYARKC 1951
Query: 2051 HKCQIYADKIHMPPTTLNLLSSPWPFSMWGIDMIGRIEPKASNGHRFILVAIDYFTKWVE 2110
HKCQIYAD++H+PP+ LN+++SPWPF+MWGIDMIG+IEP ASNGHRFILVAIDYFTKWVE
Sbjct: 1952 HKCQIYADRVHVPPSPLNVMNSPWPFAMWGIDMIGKIEPTASNGHRFILVAIDYFTKWVE 2011
Query: 2111 AASYANVTKQVVVKFIKNHIICRYGIPNRIITDNGTNLNNKMMKELCDDFKIEHHNSSPY 2170
AASYAN+TKQVV +FIK IICRYG+P RIITDNG+NLNNKMMKELC DFKIEHHNSSPY
Sbjct: 2012 AASYANITKQVVTRFIKKEIICRYGVPERIITDNGSNLNNKMMKELCKDFKIEHHNSSPY 2071
Query: 2171 RPQMNGAVEAANKNIKRIVQKMVVTYKDWHEMLPFALHGYRTSVRTSIGATPFSLVYGME 2230
RP+MNGAVEAANKNIK+IV+KMVVTYKDWHEMLPFALHGYRTSVRTS GATP+SLVYGME
Sbjct: 2072 RPKMNGAVEAANKNIKKIVRKMVVTYKDWHEMLPFALHGYRTSVRTSTGATPYSLVYGME 2131
Query: 2231 AVLPVEVEIPSLRVLMEVDLSEAEWVQNRYDQLNLIEEKRMAALCHGQLYQKRMKQAFDK 2290
AVLPVEVEIPSLRVL++V L EAEW++ R+++L+LIEE+R+A +CHGQLYQ+RMK+AFD+
Sbjct: 2132 AVLPVEVEIPSLRVLLDVKLDEAEWIRTRFNELSLIEERRLAVVCHGQLYQRRMKRAFDQ 2191
Query: 2291 KVRPREFKEGDLVLKKIFSFQPDSRGKWAPNYEGPYVVKKAFSGGAMTLQTMDGEELPRP 2350
KVRPR ++ GDLVLK+I D+RGKW PNYEGPYVVKK FSGGA+ L TMDGE+ P P
Sbjct: 2192 KVRPRSYQIGDLVLKRILPPGTDNRGKWTPNYEGPYVVKKVFSGGALMLTTMDGEDFPSP 2251
Query: 2351 VNTDTVKKYF 2360
VN+D VKKYF
Sbjct: 2252 VNSDVVKKYF 2261
Score = 592 bits (1527), Expect = e-167
Identities = 404/1170 (34%), Positives = 601/1170 (50%), Gaps = 148/1170 (12%)
Query: 16 AQIQAQAQALAQAQTQAQAVTEAQARSQAPPPPPPVRTQAEASSSWTLCADTPTRSAPQR 75
AQ Q +AQ Q + +A+ + R +A PPP +A DTP + P
Sbjct: 16 AQFMHMMQGVAQGQEELRALVQ---RQEAVTPPPN-----QALPEGNPVHDTPAAAIPVN 67
Query: 76 SAPWFPPFTAGEIFRPITCEAQMPTHQYTGQVPLPAMRVTPATMTYSAPVIHTIP--QTE 133
+ + GE I + Q P AP + IP +
Sbjct: 68 N------YAVGEELMGIRVDGQ------------PIAPDAANARVIHAPARNRIPIVDRQ 109
Query: 134 EPIF----HSGNVEAYEEVSDLQEKYDELRRDMKALRDKGKFGKTAYDLCLVPSVQVPHK 189
E +F ++ + D K D L ++A+ + G ++ LV +++P+K
Sbjct: 110 EDLFTMFSEDEDIPGRNDARD--RKVDALAEKIRAMECQNSLGFDVTNMGLVEGLRIPYK 167
Query: 190 FKIPDFEKYKGSSCPEEHLKMYVRRMPAYAQDDQILIYYFQESLTGPASKWYTNLDKTRV 249
FK P F+KY G+SCP H++ Y R++ AY D+++ +Y+FQ+SL+G + WY L +
Sbjct: 168 FKAPSFDKYNGTSCPRTHVQAYYRKISAYTDDEKMWMYFFQDSLSGASLDWYMELKSDSI 227
Query: 250 QTFRDLCEAFVEQYSYNVDMTPDRSDLQAMTQGDKETFKEYAQRWRDTAAQVSPRIEEKE 309
+ +RDL EAF+ QY +N+DM P R+ LQ++ Q E+FKEYAQRWR+ AA+V P + E+E
Sbjct: 228 RCWRDLGEAFLRQYKHNMDMAPSRTQLQSLCQKSNESFKEYAQRWRELAARVQPPMLERE 287
Query: 310 MTKLFLKTLNHFYYKKMVGSTP-KSFAEMVGMGVQLEEGVREGRLVKNATPANGTKKTGN 368
+T +F+ TL + +M GS P SF+++V G + E ++ G++ ++ +KK
Sbjct: 288 LTDMFIGTLQGVFMDRM-GSCPFGSFSDVVICGERTESLIKTGKI--QDVGSSSSKKPFA 344
Query: 369 HFPRKKEQEVGMVTHGMPQQTYPAYQHIAAITPTSHPFQQTNNHSQIPQYPQIPQYPQIP 428
PR++E E V H Q Q A P P QQ
Sbjct: 345 GAPRRREGETNAVQHRRDQNRIEYRQAAAVTIPAPQPRQQ-------------------- 384
Query: 429 QYPQFPQNPSPQNTQQQNFQQQPYQQFPYQQYPQQNFQQQPYQQRPQQPRPPRMPINPIP 488
QQQ QQ PQQ QQ+PYQ P+Q P R + +P
Sbjct: 385 --------------QQQRVQQ-----------PQQQQQQRPYQ--PRQRMPDRR-FDSLP 416
Query: 489 VTYAELLPGLLKKNLVQTRTAPPIPEKLPSWYRLDQTCDFHEGGRGHNIETCYAFKSTVQ 548
++YAELLP LL+ LV+ T P P LP Y + CDFH G GH+ E C A + VQ
Sbjct: 417 MSYAELLPELLRLGLVELCTMAP-PTVLPPGYDANVRCDFHSGAPGHHTEKCRALQHKVQ 475
Query: 549 RLINDGKITFTDSAPNVQTNPLPNHGAATVNMIEDCQKTRPILNVQHIRTPLVPLHAKLC 608
LI+ I F PNV NP+P HG VN IE + + NV ++T L+ + ++L
Sbjct: 476 DLIDANAINFA-PVPNVVNNPMPQHGGHRVNNIEGKEAEDLVDNVDDVQTSLLVVKSRLL 534
Query: 609 KVDLFEHDHDLCEICLMNSGGCQKVRNDIQGLLNRGELVVERRCDD---VCVIT-----P 660
++ + C C + GC ++R IQG+++ G L R D V IT
Sbjct: 535 NEGVYSGCDEDCLGCAESENGCDQLRAGIQGMMDEGCLQFSRAVKDRGTVSTITIYFKPS 594
Query: 661 EGPLEVFYDSRKSTITPLVICLP------------GPLPYASEKAIPYKY---------- 698
EG + + + TP+ I +P G + +A+P+KY
Sbjct: 595 EGHGQRVVSAPATNGTPVTIPVPVTISAPTTIVASGRRAVENSRAVPWKYDNAYRSNRRV 654
Query: 699 -------NATMIEEG--REVPIPPLSSVDNIVEDSRVLRNGRVVPIVFPKKIDATINKEV 749
N T + G VP +VDN+ R+GR+ + +A +
Sbjct: 655 ESQTKPVNQTPVTIGLANRVPATVGPAVDNVGGPGGFTRSGRLFAPQPLRDNNAEALAKA 714
Query: 750 RTKDAGIAKEVDQPNGAGTSAEFD--EILKLIKKSEYKVVDQLMQTPSKISIMSLLLNSE 807
+ K A + +E Q S E D E +K+IKKS+YK+VDQL QTPSKISI+SLLL SE
Sbjct: 715 KGKQAVVEEEPVQKEAPEGSFEKDVEEFMKMIKKSDYKIVDQLNQTPSKISILSLLLCSE 774
Query: 808 AHKDALMKVLEQAFVDYDVTVGQFGGIVGNITACNNLSFSDEELPAEGRNHNRALHISVN 867
AH++AL+K+L A+V +++V Q G++ N++ + + F++ +LP EGRNHN+ALHI++
Sbjct: 775 AHRNALLKMLNLAYVPQEISVNQLEGVMANVSTRHGVGFTNLDLPPEGRNHNKALHITME 834
Query: 868 CKTDALSKVLVDTGSSLNVLSKTTYTQLTYQDAPLRRSGVMVKAFDGSRKDVLGEVVLPI 927
CK LS VLVDTGSSLNVL K ++ + L S ++V+AFD S++ V GEV LP+
Sbjct: 835 CKGAVLSHVLVDTGSSLNVLPKQILKKIDVEGFVLTPSDLIVRAFDRSKRSVCGEVTLPV 894
Query: 928 TVGPQVFQVNFQVMDIQASYSCLLGRPWIHEAGAVTSTLHQKLKFVKNGKLVTVNGEEAL 987
+GP+VF + F VMDIQ +YSCLLGRPWIH AGAV+STLHQKLK+V NG++VTV GEE +
Sbjct: 895 KIGPEVFDIIFYVMDIQPAYSCLLGRPWIHAAGAVSSTLHQKLKYVWNGQIVTVCGEEEI 954
Query: 988 LVSHLSSFSFIGAD-DVEGTPFQGF------TIEDKNTKRNEASISSLKDAQKVIQAGGS 1040
LVSHLSSF ++ D ++ T Q F + ++ SI+S K A++V+ +G +
Sbjct: 955 LVSHLSSFKYVEVDGEIHETLCQVFETVALEKVAYAEQRKPGVSITSYKQAKEVVDSGKA 1014
Query: 1041 TSWGKLIELPENKHREGLGFFPSTGLSTAKKG--TFHSSGFI-HAII-----ED-DPESV 1091
WGK+++LP + + G+G+ P + G TF S+G + H + ED D +
Sbjct: 1015 EGWGKMVDLPVKEDKFGVGYEPLQAEQNGQAGPSTFTSAGLMNHGDVSATGSEDCDSDCD 1074
Query: 1092 PRGFI---TPGVSSHNWVAVDVPFVAHLSK 1118
++ PG S +NW A +V V L++
Sbjct: 1075 LDNWVRPCAPGGSINNWTAEEVVQVTLLTE 1104
>UniRef100_Q6WAY5 Gag/pol polyprotein [Pisum sativum]
Length = 1105
Score = 589 bits (1518), Expect = e-166
Identities = 395/1176 (33%), Positives = 598/1176 (50%), Gaps = 152/1176 (12%)
Query: 1 AELIKTMAETQTQAQAQIQAQAQALAQAQTQAQAVTEAQARSQAPPPPPPVRTQAEASSS 60
AE+ MA+ Q Q Q + A Q Q A+ + APP PV
Sbjct: 9 AEMKANMAQFMNMMQGVAQGQEELRAMVQRQEAAIPPV---NHAPPEGGPVNDN------ 59
Query: 61 WTLCADTPTRSAPQRSAPWFPPFTAGEIFRPITCEAQMPTHQYTGQVPLPAMRVTPATMT 120
+ A P + + G+ I +Q P + T
Sbjct: 60 -NVAAAVPINN-----------YDVGDELGGIRINSQ------------PIVPDAANART 95
Query: 121 YSAPVIHTIP--QTEEPIFH--SGNVEAYEEVSDLQEKYDELRRDMKALRDKGKFGKTAY 176
AP + +P +E +F S + + V + K D L ++A+ +
Sbjct: 96 IRAPARNPVPIVDRQEDMFTLLSEDEDVLGRVDERDRKVDALAEKIRAMECQNSLSFDVT 155
Query: 177 DLCLVPSVQVPHKFKIPDFEKYKGSSCPEEHLKMYVRRMPAYAQDDQILIYYFQESLTGP 236
++ LV +++P+KFK P F+KY +SCP H++ Y R++ AY D+++ +Y+FQ+SL+G
Sbjct: 156 NMGLVEGLRIPYKFKAPSFDKYNDTSCPRTHVQAYYRKISAYTDDEKMWMYFFQDSLSGA 215
Query: 237 ASKWYTNLDKTRVQTFRDLCEAFVEQYSYNVDMTPDRSDLQAMTQGDKETFKEYAQRWRD 296
+ WY L + ++ +RDL EAF+ QY +N+DM P R+ LQ++ Q E+FKEYAQRWR+
Sbjct: 216 SLDWYMELKRDSIRCWRDLGEAFLRQYKHNMDMAPSRTQLQSLCQKSGESFKEYAQRWRE 275
Query: 297 TAAQVSPRIEEKEMTKLFLKTLNHFYYKKMVGSTP-KSFAEMVGMGVQLEEGVREGRLVK 355
AA+V P + E+E+T +F+ TL + +M GS P SF+++V G + E ++ G++
Sbjct: 276 LAARVQPPMLERELTDMFIGTLQGVFMDRM-GSCPFGSFSDVVICGERTESLIKTGKI-- 332
Query: 356 NATPANGTKKTGNHFPRKKEQEVGMVTHGMPQQTYPAYQHIAAITPTSHPFQQTNNHSQI 415
++ +KK PR++E E +V H Q Q A P P QQ Q
Sbjct: 333 QDVGSSSSKKPFAGAPRRREGETNVVQHRRDQNRIEYRQAAAVTIPAPQPRQQQQQRVQQ 392
Query: 416 PQYPQIPQYPQIPQYPQFPQNPSPQNTQQQNFQQQPYQQFPYQQYPQQNFQQQPYQQRPQ 475
PQ QQ QQ+PYQ P Q+ P + F
Sbjct: 393 PQ--------------------------QQQQQQRPYQ--PRQRMPDRRF---------- 414
Query: 476 QPRPPRMPINPIPVTYAELLPGLLKKNLVQTRTAPPIPEKLPSWYRLDQTCDFHEGGRGH 535
+ +P++YAELLP LL+ +V+ RT P P LP Y + CDFH G GH
Sbjct: 415 ---------DSLPMSYAELLPELLRLGMVELRTMAP-PTVLPPGYDANVRCDFHSGAPGH 464
Query: 536 NIETCYAFKSTVQRLINDGKITFTDSAPNVQTNPLPNHGAATVNMIEDCQKTRPILNVQH 595
+ E C A + VQ LI+ I F PNV NP+P HG VN IE + ++NV
Sbjct: 465 HTEKCRALQHKVQDLIDAKAINFA-PVPNVVNNPMPQHGGHRVNNIEGKEAEDLVVNVDD 523
Query: 596 IRTPLVPLHAKLCKVDLFEHDHDLCEICLMNSGGCQKVRNDIQGLLNRGELVVERRCDD- 654
++T L+ + +L ++ + C C + GC ++R IQG+++ G L R D
Sbjct: 524 VQTSLLVVKGRLLNGGVYSGCDEDCLGCAESENGCDQLRAGIQGMMDEGCLQFSRAVKDR 583
Query: 655 --VCVIT-----PEGPLEVFYDSRKSTITPLVICLP------------GPLPYASEKAIP 695
V IT EG + + + TP+ I +P G + + +P
Sbjct: 584 GTVSTITIYFKPTEGRGQRVVSAPATNDTPVTISVPVTISAPTTIVASGRRAVENSRVVP 643
Query: 696 YKYNATMIEEGR-------------------EVPIPPLSSVDNIVEDSRVLRNGRVVPIV 736
+KY+ R VP +VDN+ R+GR+
Sbjct: 644 WKYDNAYRSNRRAESQTRPVNQAPVTIGLPNRVPATVGPAVDNVGGPGGFTRSGRLFAPQ 703
Query: 737 FPKKIDATINKEVRTKDAGIAKE-VDQPNGAGT-SAEFDEILKLIKKSEYKVVDQLMQTP 794
+ +A + + K A + +E V + GT + +E +K+IKKS+YK+VDQL QTP
Sbjct: 704 PLRDNNAEALAKAKGKQAVVEEEPVQKEAPEGTFEKDVEEFMKIIKKSDYKIVDQLNQTP 763
Query: 795 SKISIMSLLLNSEAHKDALMKVLEQAFVDYDVTVGQFGGIVGNITACNNLSFSDEELPAE 854
SKISI+SLL+ SEAH++AL+K+L A+V +++V Q G++ N++ + + F++ +LP E
Sbjct: 764 SKISILSLLMCSEAHRNALLKMLNLAYVPQEISVNQLEGVMANVSTRHGVGFTNLDLPPE 823
Query: 855 GRNHNRALHISVNCKTDALSKVLVDTGSSLNVLSKTTYTQLTYQDAPLRRSGVMVKAFDG 914
GRNHN+ALHI++ CK LS VLVDTGSSLNVL ++ + L S ++V+AFDG
Sbjct: 824 GRNHNKALHITMECKGAVLSHVLVDTGSSLNVLPNQILKKIDVEGFVLTPSDLIVRAFDG 883
Query: 915 SRKDVLGEVVLPITVGPQVFQVNFQVMDIQASYSCLLGRPWIHEAGAVTSTLHQKLKFVK 974
S++ V GEV LP+ +GP+VF + F VMDIQ +YSCLLGRPWIH AGAV+STLHQKLK+V
Sbjct: 884 SKRSVCGEVTLPVKIGPEVFDIIFYVMDIQPAYSCLLGRPWIHAAGAVSSTLHQKLKYVW 943
Query: 975 NGKLVTVNGEEALLVSHLSSFSFIGAD-DVEGTPFQGF------TIEDKNTKRNEASISS 1027
NG++VTV GEE +LVSHLSSF ++ D ++ T Q F + ++ ASI+S
Sbjct: 944 NGQIVTVCGEEEILVSHLSSFKYVEVDGEIHETLCQAFETVALEKVAYAEQRKPGASITS 1003
Query: 1028 LKDAQKVIQAGGSTSWGKLIELPENKHREGLGFFPSTGLSTAKKG--TFHSSGFI-HAII 1084
K A++V+ +G + WGK+++LP + + G+G+ P + G TF S+G + H +
Sbjct: 1004 YKQAKEVVDSGKAEGWGKMVDLPVKEDKFGVGYEPLRAEQNGQAGPSTFTSAGLMNHGDV 1063
Query: 1085 -----ED-----DPESVPRGFITPGVSSHNWVAVDV 1110
ED D ++ R PG S +NW A +V
Sbjct: 1064 SATGSEDCDSDCDLDNWVRS-CAPGESINNWTAEEV 1098
>UniRef100_Q6WAY7 Gag/pol polyprotein [Pisum sativum]
Length = 1814
Score = 588 bits (1515), Expect = e-166
Identities = 400/1183 (33%), Positives = 598/1183 (49%), Gaps = 150/1183 (12%)
Query: 1 AELIKTMAETQTQAQAQIQAQAQALAQAQTQAQAVTEAQARSQAPPPPPPVRTQAEASSS 60
AE+ MA+ Q Q Q + A Q Q A+ + APP PV
Sbjct: 9 AEMKANMAQFMNMMQGVAQGQEELRALVQRQEAAIPPV---NHAPPEGGPVNGN------ 59
Query: 61 WTLCADTPTRSAPQRSAPWFPPFTAGEIFRPITCEAQ--MPTHQYTGQVPLPAMRVTPAT 118
+ A P + + G+ I Q P V PA P
Sbjct: 60 -NVAAAVPINN-----------YAVGDELGGIRINGQPIAPDVANARAVRAPARNPAP-- 105
Query: 119 MTYSAPVIHTIPQTEEPIFH--SGNVEAYEEVSDLQEKYDELRRDMKALRDKGKFGKTAY 176
I +E +F S + + V + K D L ++A+ + G
Sbjct: 106 ----------IVDRQEDMFSLLSEDEDVLGRVDERDRKVDALAEKIRAMECQNSLGFDVT 155
Query: 177 DLCLVPSVQVPHKFKIPDFEKYKGSSCPEEHLKMYVRRMPAYAQDDQILIYYFQESLTGP 236
++ LV +++P+KFK P F+KY G+SCP H++ Y R++ AY D+++ +Y+FQ+SL+G
Sbjct: 156 NMGLVEGLRIPYKFKAPSFDKYNGTSCPRTHVQAYYRKISAYTDDEKMWMYFFQDSLSGA 215
Query: 237 ASKWYTNLDKTRVQTFRDLCEAFVEQYSYNVDMTPDRSDLQAMTQGDKETFKEYAQRWRD 296
+ WY L + ++ ++DL EAF+ QY +N+DM P R+ LQ++ Q E+FKEYAQRWR+
Sbjct: 216 SLDWYMELKRDSIRCWKDLGEAFLRQYKHNMDMAPSRTQLQSLCQKSGESFKEYAQRWRE 275
Query: 297 TAAQVSPRIEEKEMTKLFLKTLNHFYYKKMVGSTP-KSFAEMVGMGVQLEEGVREGRLVK 355
AA+V P + E+E+T +F+ TL + +M GS P SF+++V G + E ++ G++
Sbjct: 276 LAARVQPPMLERELTDMFIGTLQGVFMDRM-GSCPFVSFSDVVICGERTESLIKTGKI-- 332
Query: 356 NATPANGTKKTGNHFPRKKEQEVGMVTHGMPQQTYPAYQHIAAITPTSHPFQQTNNHSQI 415
++ +KK PR++E E V + Q Q A P P QQ Q
Sbjct: 333 QDAGSSSSKKPFAGAPRRREGETNAVQYRRDQNRSQRCQVAAVTIPAPQPRQQQQQRVQQ 392
Query: 416 PQYPQIPQYPQIPQYPQFPQNPSPQNTQQQNFQQQPYQQFPYQQYPQQNFQQQPYQQRPQ 475
P QQQ QQ+PYQ P Q+ P + F
Sbjct: 393 P--------------------------QQQQQQQRPYQ--PRQRMPDRRF---------- 414
Query: 476 QPRPPRMPINPIPVTYAELLPGLLKKNLVQTRTAPPIPEKLPSWYRLDQTCDFHEGGRGH 535
+ +P++YAELLP LL+ +V+ RT P P LP Y + CDFH G GH
Sbjct: 415 ---------DSLPMSYAELLPELLRLGMVELRTMAP-PTVLPPGYDANVRCDFHSGAPGH 464
Query: 536 NIETCYAFKSTVQRLINDGKITFTDSAPNVQTNPLPNHGAATVNMIEDCQKTRPILNVQH 595
+ E C A + VQ LI+ I F PNV NP+P HG VN IE + ++NV
Sbjct: 465 HTEKCRALQHKVQDLIDAKAINFA-PVPNVVNNPMPQHGGHRVNNIEGKEAEDLVVNVDD 523
Query: 596 IRTPLVPLHAKLCKVDLFEHDHDLCEICLMNSGGCQKVRNDIQGLLNRGELVVERRCDD- 654
++T L+ + +L ++ + C C + GC ++R IQG+++ G L R D
Sbjct: 524 VQTSLLVVKGRLLNGGVYSGCDEDCLGCAESENGCDQLRTGIQGMMDEGCLQFSRAVKDR 583
Query: 655 --VCVIT-----PEGPLEVFYDSRKSTITPLVICLP------------GPLPYASEKAIP 695
V IT EG + + + TP+ I +P G + +A+P
Sbjct: 584 GTVSTITIYFKPSEGRGQRVVSAPATNGTPVTISVPVTISAPTTIAASGRRAVENSRAVP 643
Query: 696 YKYNATMIEEGR-------------------EVPIPPLSSVDNIVEDSRVLRNGRVVPIV 736
+KY+ R VP +VDN+ R+GR+
Sbjct: 644 WKYDNAYRSNRRAESQTRPVNQAPVTIGLPNRVPATVGPAVDNVGGPGGFTRSGRLFAPQ 703
Query: 737 FPKKIDATINKEVRTKDAGIAKEVDQPNGAGTSAEFD--EILKLIKKSEYKVVDQLMQTP 794
+ +A + + K A + +E Q S E D E +K+IKKS+YK+VDQL QTP
Sbjct: 704 PLRDNNAEALAKAKGKQAVVEEEPVQKEAPEGSFEKDVEEFMKIIKKSDYKIVDQLNQTP 763
Query: 795 SKISIMSLLLNSEAHKDALMKVLEQAFVDYDVTVGQFGGIVGNITACNNLSFSDEELPAE 854
SKISI+SLLL SEAH++AL+K+L A+V +++V Q G++ N++ + + F++ +L E
Sbjct: 764 SKISILSLLLCSEAHRNALLKMLNLAYVPQEISVNQLEGVMANVSTRHGVGFTNLDLTPE 823
Query: 855 GRNHNRALHISVNCKTDALSKVLVDTGSSLNVLSKTTYTQLTYQDAPLRRSGVMVKAFDG 914
GRNHN+ALHI++ CK LS VLVDTGSSLNVL K ++ + L S ++V+AFDG
Sbjct: 824 GRNHNKALHITMECKGAVLSHVLVDTGSSLNVLPKQILKKIDVEGFVLTPSDLIVRAFDG 883
Query: 915 SRKDVLGEVVLPITVGPQVFQVNFQVMDIQASYSCLLGRPWIHEAGAVTSTLHQKLKFVK 974
S++ V GEV LP+ +GP+VF + F VMDIQ +YSCLLGRPWIH AGAV+STLHQKLK+V
Sbjct: 884 SKRSVCGEVTLPVKIGPEVFDIIFYVMDIQPAYSCLLGRPWIHAAGAVSSTLHQKLKYVW 943
Query: 975 NGKLVTVNGEEALLVSHLSSFSFIGAD-DVEGTPFQGF------TIEDKNTKRNEASISS 1027
NG++VTV GEE +LVSHLSSF ++ D ++ T Q F + ++ ASI+S
Sbjct: 944 NGQIVTVCGEEEILVSHLSSFKYVEVDGEIHETLCQAFETVALEKVAYAEQRKPGASITS 1003
Query: 1028 LKDAQKVIQAGGSTSWGKLIELPENKHREGLGFFPSTGLSTAKKG--TFHSSGFI-HAII 1084
K A++V+ +G + WGK++ LP + + G+G+ P + G TF S+G + H +
Sbjct: 1004 YKQAKEVVDSGKAEGWGKMVYLPVKEDKFGVGYEPLQAEQNGQAGPSTFTSAGLMNHGDV 1063
Query: 1085 -----ED-DPESVPRGFI---TPGVSSHNWVAVDVPFVAHLSK 1118
ED D + ++ PG S +NW A +V V L++
Sbjct: 1064 SATGSEDCDSDCDLDNWVRPCAPGGSINNWTAEEVVQVTLLTE 1106
Score = 474 bits (1221), Expect = e-131
Identities = 220/281 (78%), Positives = 252/281 (89%)
Query: 1633 ENSMGCVLGQQDETGRKEHAIYYLSKKFTECESRYSMLEKTCCALAWAAKRLRHYMINHT 1692
ENSMGCVLGQ DE+GRKEHAIYYLSKKFT+CE+RYS+LEKTCCALAWAA+RLR YM+NHT
Sbjct: 1534 ENSMGCVLGQHDESGRKEHAIYYLSKKFTDCETRYSLLEKTCCALAWAARRLRQYMLNHT 1593
Query: 1693 TWLISKMDPIKYIFEKPALIGRIARWQMLLSEYDIEYRSQKAIKGSILADHLAHQPLEDY 1752
T LISKMDP+KYIFEKPAL GR+ARWQM+L+EYDI+Y SQKAIKGSIL+D+LA QP+EDY
Sbjct: 1594 TLLISKMDPVKYIFEKPALTGRVARWQMILTEYDIQYTSQKAIKGSILSDYLAEQPIEDY 1653
Query: 1753 RPIKFDFPDEEIMYLKMKDCDEPLFGEGPDPDSVWGLIFDGAVNVYGNGIGAVLLTPKGT 1812
+P+ F+FPDE+IMYLK+KDC+EPL EGPDPD W L+FDGAVN+ GNG+GAVL+ PKG
Sbjct: 1654 QPMMFEFPDEDIMYLKVKDCEEPLVEEGPDPDDKWTLMFDGAVNMNGNGVGAVLINPKGA 1713
Query: 1813 HIPFTARLRFDCTNNIAEYEACIMGIEEAIDLRIKNIEIYGDSALVINQIKGKWETLHAG 1872
HIPF+ARL FD TNN AEYEACIMGIEEAIDLRIK ++IYGDSALV+NQ+ G W T
Sbjct: 1714 HIPFSARLTFDVTNNEAEYEACIMGIEEAIDLRIKTLDIYGDSALVVNQVNGDWNTNQPH 1773
Query: 1873 LIPYRDYARRLLTFFNKVELHHIPRDENQMADALATLSSMI 1913
LIPYRDY RR+LTFF KV L+H+PRDENQMADALATLSSMI
Sbjct: 1774 LIPYRDYTRRILTFFKKVRLYHVPRDENQMADALATLSSMI 1814
>UniRef100_Q9FE41 Oryza sativa (japonica cultivar-group) genomic DNA, chromosome 1, PAC
clone:P0433F09 [Oryza sativa]
Length = 2876
Score = 520 bits (1340), Expect = e-145
Identities = 288/748 (38%), Positives = 436/748 (57%), Gaps = 39/748 (5%)
Query: 1635 SMGCVLGQQDETGRKEHAIYYLSKKFTECESRYSMLEKTCCALAWAAKRLRHYMINHTTW 1694
S+G +L Q ++ G KE A YYLS+ E YS +EK C AL +A K+LRHYM+ H
Sbjct: 2145 SIGALLAQHNDEG-KEVACYYLSRTMVGAEQNYSPIEKLCLALIFALKKLRHYMLAHQIQ 2203
Query: 1695 LISKMDPIKYIFEKPALIGRIARWQMLLSEYDIEYRSQKAIKGSILADHLAHQPLEDYRP 1754
LI++ DPI+Y+ +P L GR+ +W +L+ EYDI + QKAIKG LA+ LA P+ D P
Sbjct: 2204 LIARADPIRYVLSQPVLTGRLGKWALLMMEYDITFVPQKAIKGQALAEFLATHPMPDDSP 2263
Query: 1755 IKFDFPDEEIMYLKMKDCDEPLFGEGPDPDSVWGLIFDGA----VNVYGN-----GIGAV 1805
+ + PDEEI ++++ W L FDGA +N G G G V
Sbjct: 2264 LIANLPDEEIFTAELQE--------------QWELYFDGASRKDINPDGTPRRRAGAGLV 2309
Query: 1806 LLTPKGTHIPFT-ARLRFDCTNNIAEYEACIMGIEEAIDLRIKNIEIYGDSALVINQIKG 1864
TP+G I + + L+ +C+NN AEYEA I G+ A+ + ++++ +GDS L+I QI
Sbjct: 2310 FKTPQGGVIYHSFSLLKEECSNNEAEYEALIFGLLLALSMEVRSLRAHGDSRLIIRQINN 2369
Query: 1865 KWETLHAGLIPYRDYARRLLTFFNKVELHHIPRDENQMADALATLSSMIKVNHHNDVPLI 1924
+E L+PY ARRL+ F +E+ H+PR +N ADALA L++ + N ++
Sbjct: 2370 IYEVRKPELVPYYTVARRLMDKFEHIEVIHVPRSKNAPADALAKLAAALVFQGDNPAQIV 2429
Query: 1925 SVKFLDRPAYV--FAAEVVF------DDKPWFHDIKVFLQTREYPPGASNKDKKTLRRLS 1976
+ PA + EV +++ W + + P +++ L+R
Sbjct: 2430 VEERWLLPAVLELIPEEVNIIITNSAEEEDWRQPFLDYFKHGSLPEDPV--ERRQLQRRL 2487
Query: 1977 SNFFLNGDILYKRNF-DTVLLRCVDKYEADLLIHEIHEGSFGIHPNGHTMAKKILRAGYY 2035
++ +LYKR++ VLLRCVD+ EA+ ++ E+H G G H +G M I GYY
Sbjct: 2488 PSYIYKAGVLYKRSYGQEVLLRCVDRSEANRVLQEVHHGVCGGHQSGPKMYHSIRLVGYY 2547
Query: 2036 WMTMESDCYKHTRKCHKCQIYADKIHMPPTTLNLLSSPWPFSMWGIDMIGRIEPKASNGH 2095
W + +DC K + CH CQI+ + H PP L+ WPF WGID+IG I P +S GH
Sbjct: 2548 WPGIMADCLKTAKTCHGCQIHDNFKHQPPAPLHPTVPSWPFDAWGIDVIGLINPPSSRGH 2607
Query: 2096 RFILVAIDYFTKWVEAASYANVTKQVVVKFIKNHIICRYGIPNRIITDNGTNLNNKMMKE 2155
RFIL A DYF+KW EA V V+ F++ HII R+G+P+RI +DN ++ +
Sbjct: 2608 RFILTATDYFSKWAEAVPLREVKSSDVINFLERHIIYRFGVPHRITSDNAKAFKSQKIYR 2667
Query: 2156 LCDDFKIEHHNSSPYRPQMNGAVEAANKNIKRIVQKMVVTY-KDWHEMLPFALHGYRTSV 2214
+ +KI+ + S+ Y PQ NG EA NK + +I++K V + +DWH+ L AL YR +V
Sbjct: 2668 FMEKYKIKWNYSTGYYPQANGMAEAFNKTLGKILKKTVDKHRRDWHDRLYEALWAYRVTV 2727
Query: 2215 RTSIGATPFSLVYGMEAVLPVEVEIPSLRVLMEVDLSEAEWVQNRYDQLNLIEEKRMAAL 2274
RT ATP+SLVYG EAVLP+E+++PSLRV + +L++ E ++ R+ +L+ +EE+R+ AL
Sbjct: 2728 RTPTQATPYSLVYGNEAVLPLEIQLPSLRVAIHDELTKDEQIRLRFQELDAVEEERLGAL 2787
Query: 2275 CHGQLYQKRMKQAFDKKVRPREFKEGD--LVLKKIFSFQPDSRGKWAPNYEGPYVVKKAF 2332
+ +LY++ M +A+DK V+ R F++G+ LVL++ +GK+ P +EGPYV+++A+
Sbjct: 2788 QNLELYRQNMVRAYDKLVKQRVFRKGELVLVLRRPIVVTHKMKGKFEPKWEGPYVIEQAY 2847
Query: 2333 SGGAMTLQTMDGEELPRPVNTDTVKKYF 2360
GGA L G + P+N +KKYF
Sbjct: 2848 DGGAYQLIDHQGSQPMPPINGRFLKKYF 2875
Score = 119 bits (299), Expect = 9e-25
Identities = 87/291 (29%), Positives = 144/291 (48%), Gaps = 18/291 (6%)
Query: 707 REVPIPPLSSVDNIVEDSRV-LRNGRVVPIVFPKKIDATINKEVRTKDAGIAKEVDQPNG 765
RE+ L VE+ + LR G+ +P K+ ++K AK+ P
Sbjct: 1237 REIFHAELDPEKTKVEEVNISLRGGKTLPDPHKSKVP-NVDKP--------AKKASPPGE 1287
Query: 766 AGTSAEFDEILKLIKKSEYKVVDQLMQTPSKISIMSLLLNSEAHKDALMKVLEQAFVDYD 825
A + E K +YKV+ L + P+ +S+ L+ ++AL+K L+ V Y+
Sbjct: 1288 APEAPETKTGSKEKPAVDYKVLAHLKRIPALLSVYDALMMVPDLREALIKALQAPEV-YE 1346
Query: 826 VTVGQFGGIVGNITACNNLSFSDEELPAEGRNHNRALHISVNCKTDALSKVLVDTGSSLN 885
V + + + N N ++F+DE+ +G +HNR L+I N + L ++L+D GS++N
Sbjct: 1347 VDMAKHR-LYDNPLFVNEITFADEDNIIKGGDHNRPLYIEGNIGSAHLRRILIDLGSAVN 1405
Query: 886 VLSKTTYTQLTYQDAPLRRSGVMVKAFDGSRKDVLGEVVLPITVGPQVFQVNFQVMDIQA 945
+L + T+ + L V++ FD K LG + + I + F+V F V++
Sbjct: 1406 ILPVRSLTRAGFTTKDLEPIDVVICGFDNQGKPTLGAITIKIQMSTFSFKVRFFVIEANT 1465
Query: 946 SYSCLLGRPWIHEAGAVTSTLHQKLKFVKNGKLVTVNGEEALLVSHLSSFS 996
SYS LLGRPWIH+ V STLHQ LKF+ NG + + S+ S ++
Sbjct: 1466 SYSALLGRPWIHKYRVVPSTLHQCLKFLDG------NGVQQRITSNFSPYT 1510
>UniRef100_Q93Y69 Putative gag-pol [Oryza sativa]
Length = 1262
Score = 478 bits (1229), Expect = e-132
Identities = 269/683 (39%), Positives = 396/683 (57%), Gaps = 43/683 (6%)
Query: 1635 SMGCVLGQQDETGRKEHAIYYLSKKFTECESRYSMLEKTCCALAWAAKRLRHYMINHTTW 1694
S+G +L Q ++ G KE A YYLS+ E YS +EK C AL +A K+LRHYM+ H
Sbjct: 599 SVGALLAQHNDEG-KEVACYYLSRTMVGAERNYSPIEKLCLALIFALKKLRHYMLTHQIQ 657
Query: 1695 LISKMDPIKYIFEKPALIGRIARWQMLLSEYDIEYRSQKAIKGSILADHLAHQPLEDYRP 1754
LI+ +DPI+Y+ +P L GR+ +W +L+ E+DI Y QKA+KG LA+ LA P+ D P
Sbjct: 658 LIATVDPIRYVLSQPLLAGRLGKWALLMMEFDITYVPQKAVKGQALAEFLAAHPVPDDSP 717
Query: 1755 IKFDFPDEEIMYLKMKDCDEPLFGEGPDPDSVWGLIFDGAVNVYGN---------GIGAV 1805
+ + PDE++ + + + W L FDGA + G G V
Sbjct: 718 LITELPDEDVFTI--------------ETEPSWELCFDGASRTENDRDGTPRKRAGAGLV 763
Query: 1806 LLTPKGTHIPFT-ARLRFDCTNNIAEYEACIMGIEEAIDLRIKNIEIYGDSALVINQIKG 1864
TP+G I + + L+ +C+NN AEYEA I G+ A+ + +++I +YGDS L++ QI
Sbjct: 764 FKTPQGGVIYHSFSLLKEECSNNEAEYEALIFGLLLALSMEVRSIRVYGDSQLIVQQIND 823
Query: 1865 KWETLHAGLIPYRDYARRLLTFFNKVELHHIPRDENQMADALATLSSMI--------KVN 1916
+E L L+PY ARRL+ F +E+ H+PR N ADALA L++ + +VN
Sbjct: 824 IYEVLKQELVPYYSAARRLMEMFGHIEVMHVPRSRNAPADALAKLAAALVLPQGGPTQVN 883
Query: 1917 HHNDVPLISVKFLDRPAY----VFAAEVVFDDKPWFHDIKVFLQTREYPPGASNKDKKTL 1972
L +V L Y V AA DD W + + P + + ++
Sbjct: 884 VEERWLLPAVLELLPNEYEVDTVMAAAAKEDD--WRVPFLNYFRHGSLPDNSVER-RQLQ 940
Query: 1973 RRLSSNFFLNGDILYKRNF-DTVLLRCVDKYEADLLIHEIHEGSFGIHPNGHTMAKKILR 2031
RRL S + +G + Y+R++ VLLRCVD+ EAD + E+H G G H +G M I
Sbjct: 941 RRLPSYVYKSG-VSYRRSYGQEVLLRCVDRLEADKALQEVHHGVCGGHQSGPKMYHSIRL 999
Query: 2032 AGYYWMTMESDCYKHTRKCHKCQIYADKIHMPPTTLNLLSSPWPFSMWGIDMIGRIEPKA 2091
AGYYW + +DC K + CH CQI+ D H+PP L+ WPF WGID+IG I+P +
Sbjct: 1000 AGYYWPEIMADCLKVAKSCHGCQIHGDFKHLPPVPLHPTVPAWPFEAWGIDVIGPIDPPS 1059
Query: 2092 SNGHRFILVAIDYFTKWVEAASYANVTKQVVVKFIKNHIICRYGIPNRIITDNGTNLNNK 2151
S GHRFI DYF+KW EA S V V+ F++ HII R+G+P+RI +DNG +
Sbjct: 1060 SRGHRFIFAITDYFSKWAEAVSLREVKTDNVISFLERHIIYRFGVPHRISSDNGKAFKSH 1119
Query: 2152 MMKELCDDFKIEHHNSSPYRPQMNGAVEAANKNIKRIVQKMVVTY-KDWHEMLPFALHGY 2210
M+ +KI + S+ Y PQ NG +EA NK + +I++K+V + +DWH+ L AL Y
Sbjct: 1120 KMQRFIAKYKIRWNYSTGYYPQANGMIEAFNKTLGKILKKVVNRHRRDWHDHLFEALWAY 1179
Query: 2211 RTSVRTSIGATPFSLVYGMEAVLPVEVEIPSLRVLMEVDLSEAEWVQNRYDQLNLIEEKR 2270
R +VRT TP+SLVYG EAVLP+EVE+PSLRV + ++++ E V+ R+ +L+ +EE R
Sbjct: 1180 RVTVRTPTQCTPYSLVYGSEAVLPLEVEVPSLRVAIHEEITQDEQVRLRFQELDTLEEGR 1239
Query: 2271 MAALCHGQLYQKRMKQAFDKKVR 2293
+ A+ + +LY++ M +A++K V+
Sbjct: 1240 LQAVQNLELYRQNMVRAYNKLVK 1262
>UniRef100_Q40580 Integrase [Nicotiana tabacum]
Length = 382
Score = 449 bits (1154), Expect = e-124
Identities = 204/342 (59%), Positives = 273/342 (79%), Gaps = 2/342 (0%)
Query: 2019 HPNGHTMAKKILRAGYYWMTMESDCYKHTRKCHKCQIYADKIHMPPTTLNLLSSPWPFSM 2078
H NG ++KKILRAGY+WMTME+DC ++ +KCH+CQIYAD + +PP LN S+PWPF+
Sbjct: 26 HMNGFVLSKKILRAGYFWMTMETDCIRYVQKCHQCQIYADMMRVPPNKLNATSAPWPFAA 85
Query: 2079 WGIDMIGRIEPKASNGHRFILVAIDYFTKWVEAASYANVTKQVVVKFIKNHIICRYGIPN 2138
WG+D+IG IEP ASN H+FILVAIDYFTKWVEAASY VTK+VV F+++ I+CR+ +P
Sbjct: 86 WGMDIIGPIEPTASNWHKFILVAIDYFTKWVEAASYKVVTKKVVTDFVRDCIVCRFEVPE 145
Query: 2139 RIITDNGTNLNNKMMKELCDDFKIEHHNSSPYRPQMNGAVEAANKNIKRIVQKMVVTYKD 2198
IITDN NL++ +MK +CD FKI+H NS+ Y PQMNG VEAANKNIK+I++KMV +K+
Sbjct: 146 SIITDNAANLHSALMKAMCDTFKIKHKNSTAYMPQMNGVVEAANKNIKKILRKMVDNHKE 205
Query: 2199 WHEMLPFALHGYRTSVRTSIGATPFSLVYGMEAVLPVEVEIPSLRVLMEVDLSEAEWVQN 2258
WHE LPFAL GY T V TSIGATP+ LVY EAV+ VEVEI SLR++ E LS+AEW+++
Sbjct: 206 WHEKLPFALLGYCTIVCTSIGATPYLLVYSTEAVIHVEVEITSLRIIQEDVLSDAEWIRS 265
Query: 2259 RYDQLNLIEEKRMAALCHGQLYQKRMKQAFDKKVRPREFKEGDLVLKKIFSFQPDSRGKW 2318
RY+QL+LI+ KR+ +CHGQLY RM +AF+K+V+PR+F G LVLK+IF Q +++GK+
Sbjct: 266 RYEQLSLIDGKRV--VCHGQLYHNRMSRAFNKRVKPRQFASGQLVLKRIFPHQDEAKGKF 323
Query: 2319 APNYEGPYVVKKAFSGGAMTLQTMDGEELPRPVNTDTVKKYF 2360
+PN++GPY+V + +GG + L +DGE P+P+N+D VK+Y+
Sbjct: 324 SPNWQGPYMVHRVLTGGTLILAEIDGEVWPKPINSDVVKRYY 365
>UniRef100_Q94LN5 Putative retroelement pol polyprotein [Oryza sativa]
Length = 1580
Score = 448 bits (1153), Expect = e-124
Identities = 262/731 (35%), Positives = 395/731 (53%), Gaps = 53/731 (7%)
Query: 1633 ENSMGCVLGQQDETGRKEHAIYYLSKKFTECESRYSMLEKTCCALAWAAKRLRHYMINHT 1692
+ S+ VL Q+ E RKE ++YLS++ E+RYS +EK C L ++ RLRHY++++
Sbjct: 893 DKSISSVLIQELE--RKERVVFYLSRRLLNAETRYSPVEKLCLCLYFSCTRLRHYLLSNE 950
Query: 1693 TWLISKMDPIKYIFEKPALIGRIARWQMLLSEYDIEYRSQKAIKGSILADHLAHQPLEDY 1752
+I K D +KY+ P L GRI +W L+E+D+ Y S KAIKG +A+ +
Sbjct: 951 CTVICKADVVKYMLSAPILKGRIGKWIFSLTEFDLRYESPKAIKGQAIANFIV------- 1003
Query: 1753 RPIKFDFPDEEIMYLKMKDCDEPLFGEGPDPDSVWGLIFDGAVNVYGNGIGAVLLTPKGT 1812
D+ D+ I +++ W L FDG+V +G GIG V+++P+G
Sbjct: 1004 -----DYRDDSIGSVEVVP---------------WTLFFDGSVCTHGCGIGLVIISPRGA 1043
Query: 1813 HIPFTARLRFDCTNNIAEYEACIMGIEEAIDLRIKNIEIYGDSALVINQIKGKWETLHAG 1872
F ++ TNN AEYEA + G++ ++ IEI GDS LVI+Q+ G++E +
Sbjct: 1044 CFEFAYTIKPYATNNQAEYEAVLKGLQLLKEVEADTIEIMGDSLLVISQLAGEYECKNDT 1103
Query: 1873 LIPYRDYARRLLTFFNKVELHHIPRDENQMADALATLSSMIKVNHHNDVPLISVKFLDRP 1932
LI Y + + L+ F V L H+ R++N A+ LA +S K + +
Sbjct: 1104 LIVYNEKCQELMREFRLVTLKHVSREQNIEANDLAQGAS-------------GYKLMIKH 1150
Query: 1933 AYVFAAEVVFDDKPWFHDIKVFLQTREYPPGASNKDKKTLRRLSSNFFLNGDILYKRNFD 1992
+ A + DD W + + +LQ S + LR + + L D LY R D
Sbjct: 1151 VQIEVAVITADD--WRYYVYQYLQD------PSQSASRKLRYKALKYILLDDELYYRTID 1202
Query: 1993 TVLLRCVDKYEADLLIHEIHEGSFGIHPNGHTMAKKILRAGYYWMTMESDCYKHTRKCHK 2052
VLL+C+ +A + I E+HEG G H + H M + AGY+W TM DC+++ + C
Sbjct: 1203 GVLLKCLSTDQAKVAIGEVHEGICGTHQSAHKMKWLLRCAGYFWPTMLEDCFRYYKGCQD 1262
Query: 2053 CQIYADKIHMPPTTLNLLSSPWPFSMWGIDMIGRIEPKASNGHRFILVAIDYFTKWVEAA 2112
CQ + P + +N + PWPF WGIDMIG I P +S GH+FILVA DYFTKWVEA
Sbjct: 1263 CQKFGAIQRAPASAMNPIIKPWPFRGWGIDMIGMINPPSSKGHKFILVATDYFTKWVEAI 1322
Query: 2113 SYANVTKQVVVKFIKNHIICRYGIPNRIITDNGTNLNNKMMKELCDDFKIEHHNSSPYRP 2172
V ++F++ HII R GIP I TD G+ + + D I+ NSSPY
Sbjct: 1323 PLKKVDSGDAIQFVQEHIIYRVGIPQTITTDQGSIFVSDEFIQFADSMGIKLLNSSPYYA 1382
Query: 2173 QMNGAVEAANKNIKRIVQKMVVTY-KDWHEMLPFALHGYRTSVRTSIGATPFSLVYGMEA 2231
Q NG EA+NK++ +++++ + Y + WH L AL YR + SI P+ LVY +A
Sbjct: 1383 QANGQAEASNKSLIKLIKRKISDYPRQWHTRLAEALWSYRMACHGSIQVPPYKLVYRHKA 1442
Query: 2232 VLPVEVEIPSLRVLMEVDLSEAEWVQNRYDQLNLIEEKRMAALCHGQLYQKRMKQAFDKK 2291
+LP EV I S R ++ DL+ E+ D+ + + R+ AL ++R+ + ++KK
Sbjct: 1443 ILPWEVRIGSRRTELQNDLTADEYSNLMADEREDLVQSRLRALAKVIKDKERVTRHYNKK 1502
Query: 2292 VRPREFKEGDLVLKKIFSF-QPDSR-GKWAPNYEGPYVVKKAFSGGAMTLQTMDGEELPR 2349
V P++F EG+L+ K I DS+ GKW+PN+EGP+ + K S GA LQ +DGE R
Sbjct: 1503 VVPKDFSEGELIWKLILPIGTRDSKFGKWSPNWEGPFQIHKVVSKGAYMLQGLDGEVYDR 1562
Query: 2350 PVNTDTVKKYF 2360
+N +KKY+
Sbjct: 1563 ALNGKYLKKYY 1573
Score = 77.8 bits (190), Expect = 4e-12
Identities = 49/171 (28%), Positives = 94/171 (54%), Gaps = 4/171 (2%)
Query: 858 HNRALHISVNCKTDALSKVLVDTGSSLNVLSKTTYTQLTYQDAPLRRSGVMVKAFDGSRK 917
H + L+I+ +SK++VD G+++N++ T+ +L L ++ +++K F G+
Sbjct: 360 HLKPLYINGYVNGKPMSKMMVDGGAAVNLMPYATFRKLGRNAEDLIKTNMVLKDFGGNPS 419
Query: 918 DVLGEVVLPITVGPQVFQVNFQVMDIQASYSCLLGRPWIHEAGAVTSTLHQKLKFVKNGK 977
+ +G + + +TVG + F V+D + SYS LLGR WIH + ST+HQ L + K
Sbjct: 420 ETMGVLNVELTVGSKTIPTTFFVIDGKGSYSLLLGRDWIHANCCIPSTIHQCLIQWQGDK 479
Query: 978 LVTVNGEEALLVSHLSSFSFIGADDVEGT-PFQGFTIEDKNTKRNEASISS 1027
+ V + L + + S+ F G +EG+ + T++D + K+ + +S+
Sbjct: 480 IEIVPADSQLKMEN-PSYCFEGV--MEGSNVYTKDTVDDLDDKQGQGLMSA 527
Score = 37.7 bits (86), Expect = 4.3
Identities = 30/94 (31%), Positives = 39/94 (40%), Gaps = 15/94 (15%)
Query: 498 LLKKNLVQTRTAPPIPEKLPSWYRLDQT--CDFHEGGRGHNIETCYAFKSTVQRLINDGK 555
LL++ +Q P +PS L + C +H G H C FK +Q I GK
Sbjct: 148 LLREKQIQM----PASHTIPSAEELGKKRYCKWHNSG-SHTTNDCKVFKQQIQTAIEGGK 202
Query: 556 ITFTDS--APNVQTNPLPNHGAATVNMIEDCQKT 587
I F DS V NP P VNM+ +T
Sbjct: 203 IKFDDSKRPMKVDGNPFP------VNMVHTAGRT 230
>UniRef100_Q6L974 GAG-POL [Vitis vinifera]
Length = 1027
Score = 430 bits (1106), Expect = e-118
Identities = 255/720 (35%), Positives = 388/720 (53%), Gaps = 40/720 (5%)
Query: 1648 RKEHAIYYLSKKFTECESRYSMLEKTCCALAWAAKRLRHYMINHTTWLISKMDPIKYIFE 1707
+++ IYY+S+ + E+RYS +E AL AA++LR Y H +++ P++ I
Sbjct: 340 KEQKPIYYVSRALADVETRYSKMELISLALRSAAQKLRPYFQAHPVIVLTDQ-PLRNILH 398
Query: 1708 KPALIGRIARWQMLLSEYDIEYRSQKAIKGSILADHLAHQPLEDYRPIKFDFPDEEIMYL 1767
KP L GR+ +W + LSE+ IE++ + ++KG ++AD + LE R
Sbjct: 399 KPDLTGRMLQWAIELSEFGIEFQPRLSMKGQVMADFV----LEYSR-------------- 440
Query: 1768 KMKDCDEPLFGEGPDPDSVWGLIFDGAVNVYGNGIGAVLLTPKGTHIPFTARLRFDCTNN 1827
+P EG W L DGA G+G+G +L +P G H+ RL F +NN
Sbjct: 441 ------KPGQHEGSRKKEWWTLRVDGASRSSGSGVGLLLQSPTGEHLEQAIRLGFSASNN 494
Query: 1828 IAEYEACIMGIEEAIDLRIKNIEIYGDSALVINQIKGKWETLHAGLIPYRDYARRLLTFF 1887
AEYEA + G++ A+ L + + I+ DS LV+ ++ ++E A + Y R L F
Sbjct: 495 EAEYEAILSGLDLALALSVSKLRIFSDSQLVVKHVQEEYEAKDARMARYLAKVRNTLQQF 554
Query: 1888 NKVELHHIPRDENQMADALATLSSMIKVNHHNDVPLI-----SVKFLDRPAYVFAAEVVF 1942
+ + I R +N+ ADALA +++ + + +P+ SV + + A +
Sbjct: 555 TEWTIEKIKRADNRRADALAGIAASLSIKEAILLPIHVQTNPSVSEISICSTTEAPQA-- 612
Query: 1943 DDKPWFHDIKVFLQTREYPPGASNKDKKTLRRLSSNFFLNGDILYKRNFDTVLLRCVDKY 2002
DD+ W +DI +++T P K +R ++ F L G LYKR+F LRC+
Sbjct: 613 DDQEWMNDITEYIRTGTLP--GDPKQAHKVRVQAARFTLIGGHLYKRSFTGPYLRCLGHS 670
Query: 2003 EADLLIHEIHEGSFGIHPNGHTMAKKILRAGYYWMTMESDCYKHTRKCHKCQIYADKIHM 2062
EA ++ E+HEG +G H G ++A + GYYW TM+ + + ++C KCQ YA HM
Sbjct: 671 EAQYVLAELHEGIYGNHSGGRSLAHRAHSQGYYWPTMKKEAAAYVKRCDKCQRYAPIPHM 730
Query: 2063 PPTTLNLLSSPWPFSMWGIDMIGRIEPKASNGHRFILVAIDYFTKWVEAASYANVTKQVV 2122
P TTL +S PWPF+ WG+D++ R P A +F+LVA DYF+KWVEA +YA+ + V
Sbjct: 731 PSTTLKSISGPWPFAQWGMDIV-RPLPTAPAQKKFLLVATDYFSKWVEAEAYASTKDKDV 789
Query: 2123 VKFIKNHIICRYGIPNRIITDNGTNLNNKMMKELCDDFKIEHHNSSPYRPQMNGAVEAAN 2182
KF+ +IICR+GIP II DNG ++ + C + I + S+P PQ NG EA N
Sbjct: 790 TKFVWKNIICRFGIPQTIIADNGPQFDSIAFRNFCSELNIRNSYSTPRYPQSNGQAEATN 849
Query: 2183 KNIKRIVQKMVVTYK-DWHEMLPFALHGYRTSVRTSIGATPFSLVYGMEAVLPVEVEIPS 2241
K + ++K + K W E LP L YRT+ G TPF+L YGM+AV+P+E+ +P+
Sbjct: 850 KTLITALKKRLEQAKGKWVEELPGVLWAYRTTPGRPTGNTPFALAYGMDAVIPIEIGLPT 909
Query: 2242 LRVLMEVDLSEAEWVQNRYDQLNLIEEKRMAALCHGQLYQKRMKQAFDKKVRPREFKEGD 2301
+ S+A R L+ +E R +A YQ+R +++KVRPR K G
Sbjct: 910 IWT-NAAKQSDANMQLGR--NLDWTDEVRESASIRMADYQQRASAHYNRKVRPRSLKNGT 966
Query: 2302 LVLKKIFSFQPD-SRGKWAPNYEGPYVVKKAFSGGAMTLQTMDGEELPRPVNTDTVKKYF 2360
LVL+K F + GK+ N+EGPY+V KA GA LQ +DG L RP N +K+Y+
Sbjct: 967 LVLRKFFENTTEVGAGKFQANWEGPYIVSKASDNGAYHLQKLDGTPLLRPWNVSNLKQYY 1026
>UniRef100_Q9LMV1 F5M15.26 [Arabidopsis thaliana]
Length = 1838
Score = 399 bits (1026), Expect = e-109
Identities = 258/746 (34%), Positives = 383/746 (50%), Gaps = 55/746 (7%)
Query: 1633 ENSMGCVLGQQDETGRKEHAIYYLSKKFTECESRYSMLEKTCCALAWAAKRLRHYMINHT 1692
++++ VL ++D ++ I+Y SK E E+RY ++EK A+ +A++LR Y +HT
Sbjct: 1129 DHAVSSVLVREDRG--EQRPIFYTSKSLVEAETRYPVIEKAALAVVTSARKLRPYFQSHT 1186
Query: 1693 TWLISKMDPIKYIFEKPALIGRIARWQMLLSEYDIEYRSQKAIKGSILADHLAHQPLEDY 1752
+++ P++ P+ GR+ +W + LSEYDI++R + A+K +LAD L PL+
Sbjct: 1187 IAVLTDQ-PLRVALHSPSQSGRMTKWAVELSEYDIDFRPRPAMKSQVLADFLIELPLQSA 1245
Query: 1753 RPIKFDFPDEEIMYLKMKDCDEPLFGEGPDPDSVWGLIFDGAVNVYGNGIGAVLLTPKGT 1812
EE W L DG+ + G+GIG L++P
Sbjct: 1246 ERAVSGNRGEE-----------------------WSLYVDGSSSARGSGIGIRLVSPTAE 1282
Query: 1813 HIPFTARLRFDCTNNIAEYEACIMGIEEAIDLRIKNIEIYGDSALVINQIKGKWETLHAG 1872
+ + RLRF TNN+AEYE I G+ A ++I I + DS L+ Q+ G++E +
Sbjct: 1283 VLEQSFRLRFVATNNVAEYEVLIAGLRLAAGMQITTIHAFTDSQLIAGQLSGEYEAKNEK 1342
Query: 1873 LIPYRDYARRLLTFFNKVELHHIPRDENQMADALATLSSMIKVNHHNDVPLISVKFLDRP 1932
+ Y + + F +L IPR +N ADALA L+ + +P+ S+ D+P
Sbjct: 1343 MDAYLKIVQLMTKDFENFKLSKIPRGDNAPADALAALALTSDSDLRRIIPVESI---DKP 1399
Query: 1933 AY--VFAAEVV-----------FDDKPWFHDIKVFLQTREYPPGASNKDKKTLRRL---S 1976
+ A E+V D W +I+ +L P DK T RRL +
Sbjct: 1400 SIDSTDAVEIVNTIRSSNAPDPADPTDWRVEIRDYLSDGTLP-----SDKWTARRLRIKA 1454
Query: 1977 SNFFLNGDILYKRNFDTVLLRCVDKYEADLLIHEIHEGSFGIHPNGHTMAKKILRAGYYW 2036
+ + L + L K + +L C+ E + ++ E HEG+ G H G +A K+ + G+YW
Sbjct: 1455 AKYTLMKEHLLKVSAFGAMLNCLHGTEINEIMKETHEGAAGNHSGGRALALKLKKLGFYW 1514
Query: 2037 MTMESDCYKHTRKCHKCQIYADKIHMPPTTLNLLSSPWPFSMWGIDMIGRIEPKASNGHR 2096
TM SDC T KC +CQ +A IH P L +P+PF W +D++G + AS R
Sbjct: 1515 PTMISDCKTFTAKCEQCQRHAPTIHQPTELLRAGVAPYPFMRWAMDIVGPMP--ASRQKR 1572
Query: 2097 FILVAIDYFTKWVEAASYANVTKQVVVKFIKNHIICRYGIPNRIITDNGTNLNNKMMKEL 2156
FILV DYFTKWVEA SYA + V F+ IICR+G+P IITDNG+ + +
Sbjct: 1573 FILVMTDYFTKWVEAESYATIRANDVQNFVWKFIICRHGLPYEIITDNGSQFISLSFENF 1632
Query: 2157 CDDFKIEHHNSSPYRPQMNGAVEAANKNIKRIVQKMVVTYKD-WHEMLPFALHGYRTSVR 2215
C +KI + S+P PQ NG EA NK I ++K + K W + L L YRT+ R
Sbjct: 1633 CASWKIRLNKSTPRYPQGNGQAEATNKTILSGLKKRLDEKKGAWADELDGVLWSYRTTPR 1692
Query: 2216 TSIGATPFSLVYGMEAVLPVEVEIPSLRVLMEVDLSEAEWVQNRYDQLNLIEEKRMAALC 2275
++ TPF+ YGMEA+ P EV SLR M V E + D+L+ +EE R AALC
Sbjct: 1693 SATDQTPFAHAYGMEAMAPAEVGYSSLRRSMMVKNPELN-DRMMLDRLDDLEEIRNAALC 1751
Query: 2276 HGQLYQKRMKQAFDKKVRPREFKEGDLVLKKIFSFQPD-SRGKWAPNYEGPYVVKKAFSG 2334
Q YQ + +++KV R F GDLVL+K+F + + GK N+EG Y V K
Sbjct: 1752 RIQNYQLAAAKHYNQKVHNRHFDVGDLVLRKVFENTAEINAGKLGANWEGSYQVSKIVRP 1811
Query: 2335 GAMTLQTMDGEELPRPVNTDTVKKYF 2360
G L TM G +PR N+ +K+Y+
Sbjct: 1812 GDYELLTMSGTAVPRTWNSMHLKRYY 1837
Score = 84.3 bits (207), Expect = 4e-14
Identities = 45/138 (32%), Positives = 74/138 (53%)
Query: 844 LSFSDEELPAEGRNHNRALHISVNCKTDALSKVLVDTGSSLNVLSKTTYTQLTYQDAPLR 903
+SF+D +L HN L + + +++VL+DTGSS++++ K T + D ++
Sbjct: 600 ISFTDVDLEGLDTPHNDPLVVELIISDSRVTRVLIDTGSSVDLIFKDVLTAMNITDRQIK 659
Query: 904 RSGVMVKAFDGSRKDVLGEVVLPITVGPQVFQVNFQVMDIQASYSCLLGRPWIHEAGAVT 963
+ FDG +G + LPI VG + V F V+ A Y+ +LG PWIH+ A+
Sbjct: 660 PVSKPLAGFDGDFVMTIGTIKLPIFVGGLIAWVKFVVIGKPAVYNVILGTPWIHQMQAIP 719
Query: 964 STLHQKLKFVKNGKLVTV 981
ST HQ +KF + + T+
Sbjct: 720 STYHQCVKFPTHNGIFTL 737
Score = 52.8 bits (125), Expect = 1e-04
Identities = 31/117 (26%), Positives = 58/117 (49%), Gaps = 7/117 (5%)
Query: 183 SVQVPHKFKIPDFEKYKGSSCPEEHLKMY---VRRMPAYAQD-DQILIYYFQESLTGPAS 238
S++ K K+ E Y G + P+E L + + R + D F E LTGPA
Sbjct: 212 SIRGAQKIKL---ESYNGRNDPKEFLTSFNVAINRAELTIDNFDAGRCQIFIEHLTGPAH 268
Query: 239 KWYTNLDKTRVQTFRDLCEAFVEQYSYNVDMTPDRSDLQAMTQGDKETFKEYAQRWR 295
W++ L + +F L +F++ Y+ ++ +DL +++QG KE+ + + R++
Sbjct: 269 NWFSRLKPNSIDSFHQLTSSFLKHYAPLIENQTSNADLWSISQGAKESLRSFVDRFK 325
>UniRef100_Q6L5D6 Putative polyprotein [Oryza sativa]
Length = 1514
Score = 383 bits (983), Expect = e-104
Identities = 234/729 (32%), Positives = 370/729 (50%), Gaps = 55/729 (7%)
Query: 1641 GQQDETGRK-EHAIYYLSKKFTECESRYSMLEKTCCALAWAAKRLRHYMINHTTWLISKM 1699
G++D RK + +Y++S+ + ++RY +K A+ A+++LRHY H +++
Sbjct: 831 GEEDRPRRKVQRPVYFVSEALRDAKTRYPQAQKMLYAILMASRKLRHYFQAHRVTVVTSY 890
Query: 1700 DPIKYIFEKPALIGRIARWQMLLSEYDIEYRSQKAIKGSILADHLAHQPLEDYRPIKFDF 1759
P+ I GR+ RW + LSE+D+ + + AIK LAD +A ++ P
Sbjct: 891 -PLGQILHNREGTGRVVRWAIELSEFDLRFEPRHAIKSQALADFVA-----EWTPAP--- 941
Query: 1760 PDEEIMYLKMKDCDEPLFGEGPDP---DSVWGLIFDGAVNVYGNGIGAVLLTPKGTHIPF 1816
+ P GP + W + FDG++++ G G G L +P G + +
Sbjct: 942 ----------EPVSAPEASSGPTQLPHTAYWVMQFDGSLSLQGAGAGVTLTSPSGDVLRY 991
Query: 1817 TARLRFDCTNNIAEYEACIMGIEEAIDLRIKNIEIYGDSALVINQIKGKWETLHAGLIPY 1876
RL F TNN+AEYE + G+ A L I+ + + GDS LV+NQ+ +++ + Y
Sbjct: 992 LVRLDFRATNNMAEYEGLLAGLRVAAGLGIRRLLVLGDSQLVVNQVCKEYQCSDPQMDAY 1051
Query: 1877 RDYARRLLTFFNKVELHHIPRDENQMADALATLSSMIKVNHHNDVPLISVKFLDRPAYVF 1936
RR+ F+++EL H+PR +N +AD L+ L+S P F
Sbjct: 1052 VRQVRRMERHFDRIELRHVPRRDNAVADELSRLAS---------------SRAQTPPGAF 1096
Query: 1937 AAEVVFDDKPWFHDIKVFLQTREYPPGASNKDKKTLRRLSSNFFLNGDILYKRNFDTVLL 1996
+ W +I+ +L + P ++ +RR+S + L LY+R + VLL
Sbjct: 1097 EERLAQPLIAWITEIQAYLADKTLPEDREGSER--VRRISKRYVLVEGTLYRRAANGVLL 1154
Query: 1997 RCVDKYEADLLIHEIHEGSFGIHPNGHTMAKKILRAGYYWMTMESDCYKHTRKCHKCQIY 2056
+C+ + + L+ ++HEG G H T+ K R G+YW T +D R+C CQ +
Sbjct: 1155 KCIPREQGVELLADVHEGECGAHSVSRTLVGKAFRQGFYWPTALNDAVDLVRRCRACQFH 1214
Query: 2057 ADKIHMPPTTLNLLSSPWPFSMWGIDMIGRIEPKASNGHRFILVAIDYFTKWVEAASYAN 2116
A + H P L + WPF++WG+D++GR A G ++ VA+D FTKW EA
Sbjct: 1215 ARQTHQPAQALQTIPLSWPFAVWGLDILGR----APGGFEYLYVAVDKFTKWPEAYPVIK 1270
Query: 2117 VTKQVVVKFIKNHIICRYGIPNRIITDNGTNLNNKMMKELCDDFKIEHHNSSPYRPQMNG 2176
+ K +KFI+ I R+G+PNRIITDNGT +++ + C+D I+ +SP P+ NG
Sbjct: 1271 IDKHSALKFIRG-ITARFGVPNRIITDNGTQFTSELFGDYCEDMGIKLCFASPTHPRSNG 1329
Query: 2177 AVEAAN----KNIKRIVQKMVVTYKD-WHEMLPFALHGYRTSVRTSIGATPFSLVYGMEA 2231
VE AN K +K ++ + D W E LP L RT+ + G TPF LVYG EA
Sbjct: 1330 QVERANAEILKGLKTKTFNILKKHGDSWIEELPAVLWANRTTPSRATGETPFFLVYGAEA 1389
Query: 2232 VLPVEVEIPSLRVLMEVDLSEAEWVQNRYDQLNLIEEKRMAALCHGQLYQKRMKQAFDKK 2291
VLP E+ + S RV M EA+ Q R D L+ +EE+R A YQ+ +++ +
Sbjct: 1390 VLPSELTLRSPRVTM---YCEADQDQLRRDDLDYLEERRRRAALRAARYQQSLRRYHQRH 1446
Query: 2292 VRPREFKEGDLVLKKIFSFQPDSRGKWAPNYEGPYVVKKAFSGGAMTLQTMDGEELPRPV 2351
VR R DLVL+++ + K +P +EGPY V G++ L T DG ELP P
Sbjct: 1447 VRARSLCVDDLVLRRVQT--RAGLSKLSPMWEGPYRVIGVPRPGSVRLATGDGTELPNPW 1504
Query: 2352 NTDTVKKYF 2360
N + + +++
Sbjct: 1505 NIEHLCRFY 1513
Score = 53.5 bits (127), Expect = 8e-05
Identities = 26/81 (32%), Positives = 43/81 (52%)
Query: 183 SVQVPHKFKIPDFEKYKGSSCPEEHLKMYVRRMPAYAQDDQILIYYFQESLTGPASKWYT 242
+V+ P +F+ +KY GS P E L++Y + A DD+++ +F +L G A W
Sbjct: 230 NVRWPPRFRPTITKKYDGSVNPAEFLQIYTTGIEAAGGDDRVMANFFPMALKGQARGWLM 289
Query: 243 NLDKTRVQTFRDLCEAFVEQY 263
NL V ++ DLC+ F +
Sbjct: 290 NLPPASVHSWEDLCQQFTTNF 310
Score = 52.4 bits (124), Expect = 2e-04
Identities = 40/146 (27%), Positives = 70/146 (47%), Gaps = 6/146 (4%)
Query: 844 LSFSDEELPA-EGRNHNRALHISVNCKTDALSKVLVDTGSSLNVLSKTTYTQLTYQDAPL 902
L+F + P+ R A+ +S L +VL+D G++LN+LS + + L
Sbjct: 580 LTFDQTDHPSCVARGGQIAMVVSPTVCNVKLGRVLIDGGAALNILSPAAFDAIKAPGMVL 639
Query: 903 RRSGVMVKAFDGSRKDVLGEVVLPITVGP----QVFQVNFQVMDIQASYSCLLGRPWIHE 958
R S ++ G LG + LP+T G + +V+F V D+ Y+ +LGRP + +
Sbjct: 640 RPSQPIIGVTPGHTWP-LGHIDLPVTFGGSANFRTERVDFDVADLSLPYNAVLGRPALVK 698
Query: 959 AGAVTSTLHQKLKFVKNGKLVTVNGE 984
A + ++K G +TV+G+
Sbjct: 699 FMAAVHYAYLQMKMPGPGGPITVHGD 724
>UniRef100_Q9MFD2 Orf764 protein [Beta vulgaris subsp. vulgaris]
Length = 764
Score = 383 bits (983), Expect = e-104
Identities = 235/712 (33%), Positives = 375/712 (52%), Gaps = 44/712 (6%)
Query: 1653 IYYLSKKFTECESRYSMLEKTCCALAWAAKRLRHYMINHTTWLISKMDPIKYIFEKPALI 1712
IY++S E E RY +LEK A+ AA++LR Y HT ++S P++ +K
Sbjct: 92 IYFVSHVLNEAEQRYPLLEKMSLAVMIAARKLRPYFDAHTIQVLSN-HPLEKALQKLDTS 150
Query: 1713 GRIARWQMLLSEYDIEYRSQKAIKGSILADHLAHQPLEDYRPIKFDFPDEEIMYLKMKDC 1772
GR+ RW + LSE+++EY+ + AIK LAD +
Sbjct: 151 GRLLRWAVELSEFEVEYKPRTAIKAQALADFIV--------------------------- 183
Query: 1773 DEPLFGEGPDPDSVWGLIFDGAVNVYGNGIGAVLLTPKGTHIPFTARLRFDCTNNIAEYE 1832
E + E +P VW L DG+ G+G G ++ +P+G F ++F +NN AEYE
Sbjct: 184 -EASYEEEEEPVGVWKLAVDGSAAQTGSGAGIIMTSPEGN--VFEYAIKFKASNNEAEYE 240
Query: 1833 ACIMGIEEAIDLRIKNIEIYGDSALVINQIKGKWETLHAGLIPYRDYARRLLTFFNKVEL 1892
A I GI+ + K I + DS LV +QI+G++E + Y + L +E+
Sbjct: 241 AAIAGIKMCMAADAKKIRLQTDSQLVASQIRGEYEAREPAMQKYLQMIKDLAAQLISLEV 300
Query: 1893 HHIPRDENQMADALATLSS--MIKVNHHNDVPLISVKFLDRPAYVFAAEVVFDDKPWFHD 1950
+PR +N ADAL+ L+S + +N V ++ + +D+ + V K W+ D
Sbjct: 301 VLVPRADNSQADALSKLASSTLQDLNRTVMVEVMEQRSIDKAERI---NCVTAQKEWYDD 357
Query: 1951 IKVFLQTREYPPGASNKDKKTLRRLSSNFFLNGDILYKRNFDTVLLRCVDKYEADLLIHE 2010
I+ + T + P + K ++R S + + LYKR+ LLRC + E + E
Sbjct: 358 IQEYKLTGKLPDDQTIA--KAVKRDSHWYIIYQGQLYKRSDSYPLLRCTSESEKLPAMEE 415
Query: 2011 IHEGSFGIHPNGHTMAKKILRAGYYWMTMESDCYKHTRKCHKCQIYADKIHMPPTTLNLL 2070
EG G H G ++ +++R G YW TM DC ++ +KC KCQ +A ++ P L +
Sbjct: 416 APEGICGNHIGGKALSHEVMRRGNYWPTMVKDCLRYVKKCDKCQKFAPVMNQPANDLCPI 475
Query: 2071 SSPWPFSMWGIDMIGRIEPKASNGHRFILVAIDYFTKWVEAASYANVTKQVVVKFIKNHI 2130
+P PF+ WG+D++G AS G ++++VA+DYFTKW+EA + + V FI +I
Sbjct: 476 LNPIPFAQWGMDILGPFTT-ASGGRKYLIVAVDYFTKWIEAEPTKFIKARQVRSFIWKNI 534
Query: 2131 ICRYGIPNRIITDNGTNLNNKMMKELCDDFKIEHHNSSPYRPQMNGAVEAANKNIKRIVQ 2190
I R+GIP I+ D+GT + +K+ C +FKI+ +++ PQ NG EAANK I +Q
Sbjct: 535 ITRFGIPQCIVFDHGTQFDCDTIKDYCAEFKIKFASAAVCHPQSNGQAEAANKQILLALQ 594
Query: 2191 KMVVTYKD-WHEMLPFALHGYRTSVRTSIGATPFSLVYGMEAVLPVEVEIPSLRVLMEVD 2249
K + +K W +++P L RT+ + + G +PFSL +G EAV+P+EV +PS RV
Sbjct: 595 KKLEEHKGLWADLVPEVLWANRTTEKGATGKSPFSLAFGAEAVVPIEVGLPSFRVQHYDP 654
Query: 2250 LSEAEWVQNRYDQLNLIEEKRMAALCHGQLYQKRMKQAFDKKVRPREFKEGDLVLKKIFS 2309
+ ++ DQL E R+ A + RM + ++++VR R EGDLVL++ +
Sbjct: 655 EVNEQLMREALDQL---PEMRLQAALKLAAQKNRMSRIYNRRVRHRPLTEGDLVLRRTAA 711
Query: 2310 F-QPDSRGKWAPNYEGPYVVKKAFSGGAMTLQTMDGEELPRPVNTDTVKKYF 2360
+ + GK+ N+EGPY + + + G+ L T++GE L N D +KKYF
Sbjct: 712 VGKGNVHGKFTANWEGPYQICEQVAPGSYKLMTVEGEPLKNSFNADVLKKYF 763
>UniRef100_Q5U6F6 Orf764 protein [Beta vulgaris subsp. vulgaris]
Length = 764
Score = 383 bits (983), Expect = e-104
Identities = 235/712 (33%), Positives = 375/712 (52%), Gaps = 44/712 (6%)
Query: 1653 IYYLSKKFTECESRYSMLEKTCCALAWAAKRLRHYMINHTTWLISKMDPIKYIFEKPALI 1712
IY++S E E RY +LEK A+ AA++LR Y HT ++S P++ +K
Sbjct: 92 IYFVSHVLNEAEQRYPLLEKMSLAVMIAARKLRPYFDAHTIQVLSN-HPLEKALQKLDTS 150
Query: 1713 GRIARWQMLLSEYDIEYRSQKAIKGSILADHLAHQPLEDYRPIKFDFPDEEIMYLKMKDC 1772
GR+ RW + LSE+++EY+ + AIK LAD +
Sbjct: 151 GRLLRWAVELSEFEVEYKPRTAIKAQALADFIV--------------------------- 183
Query: 1773 DEPLFGEGPDPDSVWGLIFDGAVNVYGNGIGAVLLTPKGTHIPFTARLRFDCTNNIAEYE 1832
E + E +P VW L DG+ G+G G ++ +P+G F ++F +NN AEYE
Sbjct: 184 -EASYEEEEEPVGVWKLAVDGSAAQTGSGAGIIMTSPEGN--VFEYAIKFKASNNEAEYE 240
Query: 1833 ACIMGIEEAIDLRIKNIEIYGDSALVINQIKGKWETLHAGLIPYRDYARRLLTFFNKVEL 1892
A I GI+ + K I + DS LV +QI+G++E + Y + L +E+
Sbjct: 241 AAIAGIKMCMAADAKKIRLQTDSQLVASQIRGEYEAREPAMQKYLQMIKDLAAQLISLEV 300
Query: 1893 HHIPRDENQMADALATLSS--MIKVNHHNDVPLISVKFLDRPAYVFAAEVVFDDKPWFHD 1950
+PR +N ADAL+ L+S + +N V ++ + +D+ + V K W+ D
Sbjct: 301 VLVPRADNSQADALSKLASSTLQDLNRTVMVEVMEQRSIDKAERI---NCVTAQKEWYDD 357
Query: 1951 IKVFLQTREYPPGASNKDKKTLRRLSSNFFLNGDILYKRNFDTVLLRCVDKYEADLLIHE 2010
I+ + T + P + K ++R S + + LYKR+ LLRC + E + E
Sbjct: 358 IQEYKLTGKLPDDQTIA--KAVKRDSHWYIIYQGQLYKRSDSYPLLRCTSESEKLPAMEE 415
Query: 2011 IHEGSFGIHPNGHTMAKKILRAGYYWMTMESDCYKHTRKCHKCQIYADKIHMPPTTLNLL 2070
EG G H G ++ +++R G YW TM DC ++ +KC KCQ +A ++ P L +
Sbjct: 416 APEGICGNHIGGKALSHEVMRRGNYWPTMVKDCLRYVKKCDKCQKFAPVMNQPANDLCPI 475
Query: 2071 SSPWPFSMWGIDMIGRIEPKASNGHRFILVAIDYFTKWVEAASYANVTKQVVVKFIKNHI 2130
+P PF+ WG+D++G AS G ++++VA+DYFTKW+EA + + V FI +I
Sbjct: 476 LNPIPFAQWGMDILGPFTT-ASGGRKYLIVAVDYFTKWIEAEPTKFIKARQVRSFIWKNI 534
Query: 2131 ICRYGIPNRIITDNGTNLNNKMMKELCDDFKIEHHNSSPYRPQMNGAVEAANKNIKRIVQ 2190
I R+GIP I+ D+GT + +K+ C +FKI+ +++ PQ NG EAANK I +Q
Sbjct: 535 ITRFGIPQCIVFDHGTQFDCDTIKDYCAEFKIKFASAAVCHPQSNGQAEAANKQILLALQ 594
Query: 2191 KMVVTYKD-WHEMLPFALHGYRTSVRTSIGATPFSLVYGMEAVLPVEVEIPSLRVLMEVD 2249
K + +K W +++P L RT+ + + G +PFSL +G EAV+P+EV +PS RV
Sbjct: 595 KKLEEHKGLWADLVPEVLWANRTTEKGATGKSPFSLAFGAEAVVPIEVGLPSFRVQHYDP 654
Query: 2250 LSEAEWVQNRYDQLNLIEEKRMAALCHGQLYQKRMKQAFDKKVRPREFKEGDLVLKKIFS 2309
+ ++ DQL E R+ A + RM + ++++VR R EGDLVL++ +
Sbjct: 655 EVNEQLMREALDQL---PEMRLQAALKLAAQKNRMSRIYNRRVRHRPLTEGDLVLRRTAA 711
Query: 2310 F-QPDSRGKWAPNYEGPYVVKKAFSGGAMTLQTMDGEELPRPVNTDTVKKYF 2360
+ + GK+ N+EGPY + + + G+ L T++GE L N D +KKYF
Sbjct: 712 VGKGNVHGKFTANWEGPYQICEQVAPGSYKLMTVEGEPLKNSFNADVLKKYF 763
>UniRef100_Q60D74 Putative polyprotein [Oryza sativa]
Length = 1869
Score = 382 bits (980), Expect = e-104
Identities = 235/728 (32%), Positives = 373/728 (50%), Gaps = 50/728 (6%)
Query: 1642 QQDETGRK-EHAIYYLSKKFTECESRYSMLEKTCCALAWAAKRLRHYMINHTTWLISKMD 1700
++D RK + +Y++S+ + ++RY +K A+ A+++LRHY H +++
Sbjct: 1182 EEDRPHRKVQRPVYFVSEALRDAKTRYPQAQKMLYAILMASRKLRHYFQAHRVTVVTSY- 1240
Query: 1701 PIKYIFEKPALIGRIARWQMLLSEYDIEYRSQKAIKGSILADHLAHQPLEDYRPIKFDFP 1760
P+ I GR+ +W + LSE+D+ + + AIK LAD +A P
Sbjct: 1241 PLGQILHNREGTGRVVKWAIELSEFDLHFEPRHAIKSQALADFVAEWT-----------P 1289
Query: 1761 DEEIMYLKMKDCDEPLFGEGPDP---DSVWGLIFDGAVNVYGNGIGAVLLTPKGTHIPFT 1817
E++ + P GP + W + FDG++++ G G G L +P G + +
Sbjct: 1290 APELVSV-------PEASSGPSQLPHTAHWVMQFDGSLSLQGAGAGVTLTSPSGDVLRYL 1342
Query: 1818 ARLRFDCTNNIAEYEACIMGIEEAIDLRIKNIEIYGDSALVINQIKGKWETLHAGLIPYR 1877
RL F TNN+AEYE + G+ A L I+ + + GDS LV+NQ+ ++ + Y
Sbjct: 1343 VRLDFRATNNMAEYEGLLAGLRVAAGLGIRRLLVLGDSQLVVNQVCKEYRCSDPQMDAYV 1402
Query: 1878 DYARRLLTFFNKVELHHIPRDENQMADALATLSSMIKVNHHNDVPLISVKFLDRPAYVFA 1937
RR+ F+ +EL H+PR +N MAD L+ L+S + P + L +P+ + A
Sbjct: 1403 RQVRRMERHFDGIELRHVPRRDNMMADELSRLAS----SRAQTPPGAFEERLAQPSALIA 1458
Query: 1938 AEVVFDDKPWFHDIKVFLQTREYPPGASNKDKKTLRRLSSNFFLNGDILYKRNFDTVLLR 1997
W +I+ +L + P ++ + R+S + L LY+R + +LL+
Sbjct: 1459 ---------WIAEIQAYLTDKTLPEDREGSER--VHRISKRYVLVEGTLYRRAANGILLK 1507
Query: 1998 CVDKYEADLLIHEIHEGSFGIHPNGHTMAKKILRAGYYWMTMESDCYKHTRKCHKCQIYA 2057
C+ + + +L+ +IHEG G H T+ K R G+YW T +D R+C CQ +A
Sbjct: 1508 CIPREQGVVLLADIHEGECGAHSASRTLVGKAFRQGFYWPTALNDAVDLVRRCRACQFHA 1567
Query: 2058 DKIHMPPTTLNLLSSPWPFSMWGIDMIGRIEPKASNGHRFILVAIDYFTKWVEAASYANV 2117
+IH P L + WPF++WG+ ++G +A G ++ VAID FTKW EA +
Sbjct: 1568 KQIHQPAQALQTIPLSWPFAVWGLHILGPFR-RAPGGFEYLYVAIDKFTKWPEAYPVVKI 1626
Query: 2118 TKQVVVKFIKNHIICRYGIPNRIITDNGTNLNNKMMKELCDDFKIEHHNSSPYRPQMNGA 2177
K +KFIK I R+G+PNRIITDNGT +++ + C+D I+ +SP P+ NG
Sbjct: 1627 DKHSALKFIKG-ITARFGVPNRIITDNGTQFTSELFGDYCEDMGIKLCFASPAHPRSNGQ 1685
Query: 2178 VEAAN----KNIKRIVQKMVVTYKD-WHEMLPFALHGYRTSVRTSIGATPFSLVYGMEAV 2232
VE AN K +K ++ + D W E LP L RT+ + G TPF LVYG EAV
Sbjct: 1686 VERANAEILKGLKTKTFNILKKHGDSWIEELPAVLWANRTTPSRATGETPFFLVYGAEAV 1745
Query: 2233 LPVEVEIPSLRVLMEVDLSEAEWVQNRYDQLNLIEEKRMAALCHGQLYQKRMKQAFDKKV 2292
LP E+ + S R M EA+ Q R D L+ +EE+R A YQ+ +++ V
Sbjct: 1746 LPSELTLKSPRATM---YCEADQDQLRRDDLDYLEERRRRAALRAARYQQSLRRYHQCHV 1802
Query: 2293 RPREFKEGDLVLKKIFSFQPDSRGKWAPNYEGPYVVKKAFSGGAMTLQTMDGEELPRPVN 2352
R R DLVL+++ + K +P +EGPY V G++ L T DG ELP P N
Sbjct: 1803 RARSLCVDDLVLRRVQT--RAGLSKLSPMWEGPYRVVGVPRPGSIRLATGDGTELPNPWN 1860
Query: 2353 TDTVKKYF 2360
+ +++++
Sbjct: 1861 IEHLRRFY 1868
Score = 71.6 bits (174), Expect = 3e-10
Identities = 51/203 (25%), Positives = 90/203 (44%), Gaps = 8/203 (3%)
Query: 183 SVQVPHKFKIPDFEKYKGSSCPEEHLKMYVRRMPAYAQDDQILIYYFQESLTGPASKWYT 242
+V+ P +F+ EKY GS P E L++Y + A DD+++ +F +L G A W
Sbjct: 103 NVRWPPRFRPTIAEKYDGSVNPAEFLQVYTTGIEAAGGDDRVMANFFPMALKGQARGWLM 162
Query: 243 NLDKTRVQTFRDLCEAFVEQYSYNVDMTPDRSDLQAMTQGDKETFKEYAQRWRDTAAQVS 302
NL V ++ DLC+ F + + +DL A+ + D E+ + Y QR+ +
Sbjct: 163 NLPPASVHSWEDLCQQFTMNFQGTYPRPGEEADLHAVQRRDDESLRSYIQRFCQVRNTI- 221
Query: 303 PRIEEKEMTKLFLKTLNH-FYYKKMVGSTPKSFAEMVGMGVQLEEGVR------EGRLVK 355
P I + F + H +K+ P++ AE+ + ++ G V
Sbjct: 222 PCIPAHAVIYAFRGGVRHNRMLEKIASKEPQTTAELFQLADRVARKEEAWTWNPSGSGVA 281
Query: 356 NATPANGTKKTGNHFPRKKEQEV 378
+ T +TG R+K++ V
Sbjct: 282 ASAAPGSTAQTGRRDRRRKKRSV 304
Score = 51.2 bits (121), Expect = 4e-04
Identities = 39/137 (28%), Positives = 65/137 (46%), Gaps = 5/137 (3%)
Query: 852 PAEGRNHNRALHISVNCKTDALSKVLVDTGSSLNVLSKTTYTQLTYQDAPLRRSGVMVKA 911
P R A+ +S L +VL+D G++LN+LS + + LR S ++
Sbjct: 476 PCVARGGQIAMVVSPTVCNVKLGRVLIDGGAALNILSPAAFDAIKALGMMLRPSQPIIGV 535
Query: 912 FDGSRKDVLGEVVLPITVG-PQVFQ---VNFQVMDIQASYSCLLGRPWIHEAGAVTSTLH 967
G LG + LP+T G P F+ VNF V D+ Y+ +LGRP + + A +
Sbjct: 536 TPGHTWP-LGHIDLPVTFGGPANFRTERVNFDVADLSPPYNAVLGRPALVKFMAAVHYAY 594
Query: 968 QKLKFVKNGKLVTVNGE 984
++K ++V+G+
Sbjct: 595 LQMKMPGPDGPISVHGD 611
>UniRef100_Q7X6L5 OSJNBb0093G06.3 protein [Oryza sativa]
Length = 1986
Score = 375 bits (964), Expect = e-102
Identities = 230/731 (31%), Positives = 370/731 (50%), Gaps = 50/731 (6%)
Query: 1642 QQDETGRKEHAIYYLSKKFTECESRYSMLEKTCCALAWAAKRLRHYMINHTTWLISKMDP 1701
++ + + IY++S+ + ++RY ++K + ++L HY H+ +++ P
Sbjct: 1293 EEGHVQKVQRPIYFVSEVLADSKTRYPQVQKLLYGILITTRKLSHYFQGHSVTVVTSF-P 1351
Query: 1702 IKYIFEKPALIGRIARWQMLLSEYDIEYRSQKAIKGSILADHLAHQPLEDYRPIKFDFPD 1761
+ + GRIA+W + L DI ++ + +IK LAD +A ++ + D P
Sbjct: 1352 LGDVLHNREANGRIAKWALELMSLDISFKPRTSIKSQALADFVA-----EWTECQEDTPV 1406
Query: 1762 EEIMYLKMKDCDEPLFGEGPDPDSVWGLIFDGAVNVYGNGIGAVLLTPKGTHIPFTARLR 1821
E+I + W + FDG+ + G G G VL++P G + + +
Sbjct: 1407 EKIEH--------------------WTMHFDGSKRLSGTGAGVVLISPTGERLSYVLWIH 1446
Query: 1822 FDCTNNIAEYEACIMGIEEAIDLRIKNIEIYGDSALVINQIKGKWETLHAGLIPYRDYAR 1881
F ++N+AEYEA + G+ AI L IK + + GDS LV+NQ+ +W + ++ YR R
Sbjct: 1447 FSASHNVAEYEALLHGLRIAISLGIKRLIVRGDSQLVVNQVMKEWSCIDDNMMAYRQEVR 1506
Query: 1882 RLLTFFNKVELHHIPRDENQMADALATLSSMIKVN------HHNDVPLISVKFLDRPAYV 1935
+L F+ +EL H+ R N+ AD LA S + H P + K D
Sbjct: 1507 KLEDKFDGLELSHVLRHNNEAADRLANFGSKREAAPSDVFVEHLYTPTVPHK--DTTQDA 1564
Query: 1936 FAAEVVFDDKPWFHDIKVFLQTREYPPGASNKDK-KTLRRLSSNFFLNGDILYKRNFDTV 1994
+VV + W FL ++E P +KD+ + + R S + ++ LYK++ +
Sbjct: 1565 DTHDVVMVEADWREPFIRFLSSQELP---QDKDEAERISRRSKLYVMHESELYKKSPSGI 1621
Query: 1995 LLRCVDKYEADLLIHEIHEGSFGIHPNGHTMAKKILRAGYYWMTMESDCYKHTRKCHKCQ 2054
L RCV E L+ +IH G G H T+ K R G++W T SD K R C CQ
Sbjct: 1622 LQRCVSLEEGRQLLKDIHSGICGNHAAARTIVGKAYRQGFFWPTAVSDADKIVRTCEGCQ 1681
Query: 2055 IYADKIHMPPTTLNLLSSPWPFSMWGIDMIGRIEPKASNGHRFILVAIDYFTKWVEAASY 2114
+A +IH+P L + WPF++WG+DM+G + KA G+ + VAID F+KW+EA
Sbjct: 1682 FFARQIHLPAQELQTIPLSWPFAVWGLDMVGPFK-KAVGGYTHLFVAIDKFSKWIEAKPV 1740
Query: 2115 ANVTKQVVVKFIKNHIICRYGIPNRIITDNGTNLNNKMMKELCDDFKIEHHNSSPYRPQM 2174
+T F N I+ R+G+PNRIITDNGT + K+ C+DF I+ +S P
Sbjct: 1741 VTITADNARDFFIN-IVHRFGVPNRIITDNGTQFTGGVFKDFCEDFGIKICYASVAHPMS 1799
Query: 2175 NGAVEAANKNI-----KRIVQKMVVTYKDWHEMLPFALHGYRTSVRTSIGATPFSLVYGM 2229
NG VE AN I R+ ++ W + LP L RT+ + G +PF LVYG
Sbjct: 1800 NGQVERANGMILQGIKARVFDRLKPYAGKWVQQLPSVLWSLRTTPSRATGQSPFFLVYGA 1859
Query: 2230 EAVLPVEVEIPSLRVLMEVDLSEAEWVQNRYDQLNLIEEKRMAALCHGQLYQKRMKQAFD 2289
EA+LP EVE SLR + E + ++R D L+ +EE R AAL Y + +++ +
Sbjct: 1860 EAMLPSEVEFESLRFR---NFREERYEEDRVDDLHRLEEAREAALIQSARYLQGLRRYHN 1916
Query: 2290 KKVRPREFKEGDLVLKKIFSFQPDSRGKWAPNYEGPYVVKKAFSGGAMTLQTMDGEELPR 2349
+ VR R F GDLVL+KI + + R K +P +EGP+++ + G+ L+ DG +
Sbjct: 1917 RNVRSRAFLVGDLVLRKIQTTR--DRHKLSPLWEGPFIISEVTRPGSYRLKREDGTLVDN 1974
Query: 2350 PVNTDTVKKYF 2360
N + +++++
Sbjct: 1975 SWNIEHLRRFY 1985
Score = 60.8 bits (146), Expect = 5e-07
Identities = 56/242 (23%), Positives = 98/242 (40%), Gaps = 18/242 (7%)
Query: 184 VQVPHKFKIPDFEKYKGSSCPEEHLKMYVRRMPAYAQDDQILIYYFQESLTGPASKWYTN 243
V P FK EKY G++ PE L +Y + A D++ + Y +L A W
Sbjct: 428 VDWPAGFKPTGIEKYDGTTNPESWLTVYGLAIRAAGGDNKAMANYLPVALADSARSWLHG 487
Query: 244 LDKTRVQTFRDLCEAFVEQYSYNVDMTPDRSDLQAMTQGDKETFKEYAQRWRDTAAQVSP 303
L + + ++ +L + F+ + + + DL + Q E+ +EY +R+ + ++S
Sbjct: 488 LPRGTIGSWAELRDHFIANFQGTFERPGTQYDLYNVIQKSGESLREYIRRFSEQRNKISD 547
Query: 304 RIEEKEMTKLFLKTLNHFYYKKMVGS-------TPKSFAEMVGMGVQLEEGVREGR---- 352
I + + F K + H +++VG T K E + E+ V +
Sbjct: 548 -ISDDVIIAAFTKGIRH---EELVGKFGRKPPRTVKLMFEKANEYAKAEDAVTASKQSGP 603
Query: 353 --LVKNATPANGTKKTGNHFPRKKEQEVGMVTHGMPQQTYPAYQHIAAITPTSHPFQQTN 410
KN TPA G + NH RK++ E + T + I + P +
Sbjct: 604 SWKPKN-TPATGGGGSNNHKDRKRKPEELVATAAHSSRQRSRVNTFDKIMNSQCPHHPNS 662
Query: 411 NH 412
NH
Sbjct: 663 NH 664
Score = 46.6 bits (109), Expect = 0.009
Identities = 29/104 (27%), Positives = 50/104 (47%), Gaps = 5/104 (4%)
Query: 873 LSKVLVDTGSSLNVLSKTTYTQLTYQDAPLRRSGVMVKA-FDGSRKDVLGEVVLPITVGP 931
L + L+D GS+LN+L T + + L+ S G LG++ LP+T G
Sbjct: 784 LRRTLIDGGSALNILFAKTLDDMQIPRSELKPSNAPFHGVIPGLSATPLGQITLPVTFGT 843
Query: 932 Q----VFQVNFQVMDIQASYSCLLGRPWIHEAGAVTSTLHQKLK 971
+ ++F+V D + +Y +LGRP + + AV + +K
Sbjct: 844 RENFRTENISFEVADFETAYHAILGRPALAKFMAVPHYTYMMMK 887
>UniRef100_Q7XPQ7 OSJNBa0053K19.16 protein [Oryza sativa]
Length = 2010
Score = 375 bits (963), Expect = e-102
Identities = 231/731 (31%), Positives = 370/731 (50%), Gaps = 50/731 (6%)
Query: 1642 QQDETGRKEHAIYYLSKKFTECESRYSMLEKTCCALAWAAKRLRHYMINHTTWLISKMDP 1701
++ + + IY++S+ + ++RY ++K + ++L HY H+ +++ P
Sbjct: 1317 EEGHVQKVQRPIYFVSEVLADSKTRYPQVQKLLYGILITTRKLSHYFQGHSVTVVTSF-P 1375
Query: 1702 IKYIFEKPALIGRIARWQMLLSEYDIEYRSQKAIKGSILADHLAHQPLEDYRPIKFDFPD 1761
+ I GRIA+W + L DI ++ + +IK LAD +A ++ + D P
Sbjct: 1376 LGDILHNREANGRIAKWALELMSLDISFKPRISIKSQALADFVA-----EWTECQEDTPA 1430
Query: 1762 EEIMYLKMKDCDEPLFGEGPDPDSVWGLIFDGAVNVYGNGIGAVLLTPKGTHIPFTARLR 1821
E + + W + FDG+ + G G G VL++P G + + +
Sbjct: 1431 ENMEH--------------------WTMHFDGSKRLSGTGAGVVLISPTGERLSYVLWIH 1470
Query: 1822 FDCTNNIAEYEACIMGIEEAIDLRIKNIEIYGDSALVINQIKGKWETLHAGLIPYRDYAR 1881
F ++N+AEYEA + G+ AI L IK + + GDS LV+NQ+ +W L ++ YR R
Sbjct: 1471 FSASHNVAEYEALLHGLRIAISLGIKRLIVRGDSQLVVNQVMKEWSCLDDNMMAYRQEVR 1530
Query: 1882 RLLTFFNKVELHHIPRDENQMADALATLSSMIKVN------HHNDVPLISVKFLDRPAYV 1935
+L F+ +EL H+ R N+ AD LA S +V H P + K D
Sbjct: 1531 KLEDKFDGLELSHVLRHNNEAADRLANFGSKREVAPSDVFVEHLYTPTVPHK--DTTQVA 1588
Query: 1936 FAAEVVFDDKPWFHDIKVFLQTREYPPGASNKDK-KTLRRLSSNFFLNGDILYKRNFDTV 1994
+V + W + FL ++E P +KD+ + + R S + ++ LYK++ +
Sbjct: 1589 GTHDVAMVETDWREPLIRFLTSQELP---QDKDEAERISRRSKLYVMHEAELYKKSPSGI 1645
Query: 1995 LLRCVDKYEADLLIHEIHEGSFGIHPNGHTMAKKILRAGYYWMTMESDCYKHTRKCHKCQ 2054
L RCV E L+ +IH G G H T+ K R G++W T SD K R C CQ
Sbjct: 1646 LQRCVSLEEGRQLLQDIHSGICGNHAAARTIVGKAYRQGFFWPTAVSDADKIVRTCEGCQ 1705
Query: 2055 IYADKIHMPPTTLNLLSSPWPFSMWGIDMIGRIEPKASNGHRFILVAIDYFTKWVEAASY 2114
+A +IH+P L + WPF++WG+DM+G + KA G+ + VAID F+KW+EA
Sbjct: 1706 FFARQIHLPAQELQTIPLSWPFAVWGLDMVGPFK-KAVGGYTHLFVAIDKFSKWIEAKPV 1764
Query: 2115 ANVTKQVVVKFIKNHIICRYGIPNRIITDNGTNLNNKMMKELCDDFKIEHHNSSPYRPQM 2174
+T F N I+ R+G+PNRIITDNGT + K+ C+DF I+ +S P
Sbjct: 1765 VTITADNARDFFIN-IVHRFGVPNRIITDNGTQFTGGVFKDFCEDFGIKICYASVAHPMS 1823
Query: 2175 NGAVEAANKNI-----KRIVQKMVVTYKDWHEMLPFALHGYRTSVRTSIGATPFSLVYGM 2229
NG VE AN I R+ ++ W + LP L RT+ + G +PF LVYG
Sbjct: 1824 NGQVERANGMILQGIKARVFDRLKPYAGKWVQQLPSVLWSLRTTPSRATGQSPFFLVYGA 1883
Query: 2230 EAVLPVEVEIPSLRVLMEVDLSEAEWVQNRYDQLNLIEEKRMAALCHGQLYQKRMKQAFD 2289
EA+LP EVE SLR + E + ++R D L+ +EE R AAL Y + +++ +
Sbjct: 1884 EAMLPSEVEFESLRFR---NFREERYEEDRVDDLHRLEEVREAALIRSARYLQGLRRYHN 1940
Query: 2290 KKVRPREFKEGDLVLKKIFSFQPDSRGKWAPNYEGPYVVKKAFSGGAMTLQTMDGEELPR 2349
+ VR R F GDLVL+KI + + R K +P +EGP+++ + G+ L+ DG +
Sbjct: 1941 RNVRSRAFLVGDLVLRKIQTTR--DRHKLSPLWEGPFIISEVTRPGSYRLKREDGTLVDN 1998
Query: 2350 PVNTDTVKKYF 2360
N + +++++
Sbjct: 1999 SWNIEHLRRFY 2009
Score = 57.4 bits (137), Expect = 5e-06
Identities = 50/221 (22%), Positives = 93/221 (41%), Gaps = 19/221 (8%)
Query: 184 VQVPHKFKIPDFEKYKGSSCPEEHLKMYVRRMPAYAQDDQILIYYFQESLTGPASKWYTN 243
V P FK EKY G++ PE L +Y + A D++ + Y +L A W
Sbjct: 425 VDWPAGFKPTGIEKYDGTTNPESWLTVYGLAIRAAGGDNKAMANYLPVALADSARSWLHG 484
Query: 244 LDKTRVQTFRDLCEAFVEQYSYNVDMTPDRSDLQAMTQGDKETFKEYAQRWRDTAAQVSP 303
L + + ++ +L + F+ + + + DL + Q E+ ++Y +R+ + ++S
Sbjct: 485 LPRGTIGSWAELRDHFIANFQGTFERPGTQYDLYNVIQKSGESLRDYIRRFSEQRNKISD 544
Query: 304 RIEEKEMTKLFLKTLNHFYYKKMVGS-------TPKSFAEMVGMGVQLEEGVREGR---- 352
I + + F K + H +++VG T K E + E+ V +
Sbjct: 545 -ITDDVIIAAFTKGIRH---EELVGKFGRKPPMTVKLMFEKANEYAKAEDAVTASKQSGP 600
Query: 353 --LVKNATPANGTKKTGNHFPRKKE--QEVGMVTHGMPQQT 389
TPA G + NH RK++ + V +H Q++
Sbjct: 601 SWKQNKGTPATGGGGSNNHKDRKRKPAELVATASHSSRQRS 641
Score = 47.0 bits (110), Expect = 0.007
Identities = 29/104 (27%), Positives = 50/104 (47%), Gaps = 5/104 (4%)
Query: 873 LSKVLVDTGSSLNVLSKTTYTQLTYQDAPLRRSGVMVKA-FDGSRKDVLGEVVLPITVGP 931
L + L+D GS+LN+L T + + L+ S G LG++ LP+T G
Sbjct: 782 LRRTLIDGGSALNILFAKTLDDMQIPHSELKPSNAPFHGVIPGLSATPLGQITLPVTFGT 841
Query: 932 Q----VFQVNFQVMDIQASYSCLLGRPWIHEAGAVTSTLHQKLK 971
+ ++F+V D + +Y +LGRP + + AV + +K
Sbjct: 842 RENFRTENISFEVADFETAYHAILGRPALAKFMAVPHYTYMMMK 885
>UniRef100_Q8GSW6 Putative gag-pol polyprotein [Oryza sativa]
Length = 2012
Score = 375 bits (962), Expect = e-101
Identities = 232/731 (31%), Positives = 370/731 (49%), Gaps = 50/731 (6%)
Query: 1642 QQDETGRKEHAIYYLSKKFTECESRYSMLEKTCCALAWAAKRLRHYMINHTTWLISKMDP 1701
++ + + IY++S+ + ++RY ++K + ++L HY H+ +++ P
Sbjct: 1319 EEGHVQKVQRPIYFVSEVLADSKTRYPQVQKLLYGILITTRKLSHYFQGHSVTVVTSF-P 1377
Query: 1702 IKYIFEKPALIGRIARWQMLLSEYDIEYRSQKAIKGSILADHLAHQPLEDYRPIKFDFPD 1761
+ I GRIA+W + L DI ++ + +IK LAD +A ++ + D P
Sbjct: 1378 LGDILHNREANGRIAKWALELMSLDISFKPRISIKSQALADFVA-----EWTECQEDTPA 1432
Query: 1762 EEIMYLKMKDCDEPLFGEGPDPDSVWGLIFDGAVNVYGNGIGAVLLTPKGTHIPFTARLR 1821
E + + W + FDG+ + G G G VL++P G + + +
Sbjct: 1433 ENMEH--------------------WTMHFDGSKRLSGTGAGVVLISPTGERLSYVLWIH 1472
Query: 1822 FDCTNNIAEYEACIMGIEEAIDLRIKNIEIYGDSALVINQIKGKWETLHAGLIPYRDYAR 1881
F ++N+AEYEA + G+ AI L IK + + GDS LV+NQ+ +W L ++ YR R
Sbjct: 1473 FSASHNVAEYEALLHGLRIAISLGIKRLIVRGDSQLVVNQVMKEWSCLDDNMMAYRQEVR 1532
Query: 1882 RLLTFFNKVELHHIPRDENQMADALATLSSMIKVN------HHNDVPLISVKFLDRPAYV 1935
+L F+ +EL H+ R N+ AD LA S + H P + K D
Sbjct: 1533 KLEDKFDGLELSHVLRHNNEAADRLANFGSKREAAPSDVFVEHLYSPTVPHK--DATQAA 1590
Query: 1936 FAAEVVFDDKPWFHDIKVFLQTREYPPGASNKDK-KTLRRLSSNFFLNGDILYKRNFDTV 1994
A +V + W + FL ++E P +KD+ + + R S + L+ LYK++ +
Sbjct: 1591 GAHDVAMVEADWREPLIRFLTSQELP---QDKDEAERISRRSKLYVLHEAELYKKSPSGI 1647
Query: 1995 LLRCVDKYEADLLIHEIHEGSFGIHPNGHTMAKKILRAGYYWMTMESDCYKHTRKCHKCQ 2054
L RCV E L+ +IH G G H T+ K R G++W T SD K R C CQ
Sbjct: 1648 LQRCVSLEEGRQLLKDIHSGICGNHAAARTIVGKAYRQGFFWPTAVSDADKIVRTCEGCQ 1707
Query: 2055 IYADKIHMPPTTLNLLSSPWPFSMWGIDMIGRIEPKASNGHRFILVAIDYFTKWVEAASY 2114
+A +IH+P L + WPF++WG+DM+G + KA G+ + VAID F+KW+EA
Sbjct: 1708 FFARQIHLPAQELQTIPLSWPFAVWGLDMVGPFK-KAVGGYTHLFVAIDKFSKWIEAKPV 1766
Query: 2115 ANVTKQVVVKFIKNHIICRYGIPNRIITDNGTNLNNKMMKELCDDFKIEHHNSSPYRPQM 2174
+T F N I+ R+G+PNRIITDNGT + K+ C+DF I+ +S P
Sbjct: 1767 VTITADNARDFFIN-IVHRFGVPNRIITDNGTQFTGGVFKDFCEDFGIKICYASVAHPMS 1825
Query: 2175 NGAVEAANKNI-----KRIVQKMVVTYKDWHEMLPFALHGYRTSVRTSIGATPFSLVYGM 2229
NG VE AN I R+ ++ W + LP L RT+ + G +PF LVYG
Sbjct: 1826 NGQVERANGMILQGIKARVFDRLKPYAGKWVQQLPSVLWSLRTTSSRATGQSPFFLVYGA 1885
Query: 2230 EAVLPVEVEIPSLRVLMEVDLSEAEWVQNRYDQLNLIEEKRMAALCHGQLYQKRMKQAFD 2289
EA+LP EVE SLR + E + ++R D L+ +EE R AAL Y + +++ +
Sbjct: 1886 EAMLPSEVEFESLRFR---NFREERYEEDRVDDLHRLEEAREAALIRSARYLQGLRRYHN 1942
Query: 2290 KKVRPREFKEGDLVLKKIFSFQPDSRGKWAPNYEGPYVVKKAFSGGAMTLQTMDGEELPR 2349
+ VR R F GDLVL+KI + + R K +P +EGP+++ + G+ L+ DG +
Sbjct: 1943 RNVRSRAFLVGDLVLRKIQTTR--DRHKLSPLWEGPFIISEVTRPGSYRLKREDGTLVDN 2000
Query: 2350 PVNTDTVKKYF 2360
N + +++++
Sbjct: 2001 SWNIEHLRRFY 2011
Score = 57.4 bits (137), Expect = 5e-06
Identities = 50/221 (22%), Positives = 93/221 (41%), Gaps = 19/221 (8%)
Query: 184 VQVPHKFKIPDFEKYKGSSCPEEHLKMYVRRMPAYAQDDQILIYYFQESLTGPASKWYTN 243
V P FK EKY G++ PE L +Y + A D++ + Y +L A W
Sbjct: 427 VDWPAGFKPTGIEKYDGTTNPESWLTVYGLAIRAAGGDNKAMANYLPVALADSARSWLHG 486
Query: 244 LDKTRVQTFRDLCEAFVEQYSYNVDMTPDRSDLQAMTQGDKETFKEYAQRWRDTAAQVSP 303
L + + ++ +L + F+ + + + DL + Q E+ ++Y +R+ + ++S
Sbjct: 487 LPRGTIGSWAELRDHFIANFQGTFERPGTQYDLYNVIQKSGESLRDYIRRFSEQRNKISD 546
Query: 304 RIEEKEMTKLFLKTLNHFYYKKMVGS-------TPKSFAEMVGMGVQLEEGVREGR---- 352
I + + F K + H +++VG T K E + E+ V +
Sbjct: 547 -ITDDVIIAAFTKGIRH---EELVGKFGRKPPRTVKLMFEKANEYAKAEDAVTASKQSGP 602
Query: 353 --LVKNATPANGTKKTGNHFPRKKE--QEVGMVTHGMPQQT 389
TPA G + NH RK++ + V +H Q++
Sbjct: 603 SWKQNKGTPATGGGGSNNHKDRKRKPAELVATASHSSRQRS 643
Score = 46.6 bits (109), Expect = 0.009
Identities = 29/104 (27%), Positives = 50/104 (47%), Gaps = 5/104 (4%)
Query: 873 LSKVLVDTGSSLNVLSKTTYTQLTYQDAPLRRSGVMVKA-FDGSRKDVLGEVVLPITVGP 931
L + L+D GS+LN+L T + + L+ S G LG++ LP+T G
Sbjct: 784 LRRTLIDGGSALNILFAKTLDDMQIPRSELKPSNAPFHGVIPGLSATPLGQITLPVTFGT 843
Query: 932 Q----VFQVNFQVMDIQASYSCLLGRPWIHEAGAVTSTLHQKLK 971
+ ++F+V D + +Y +LGRP + + AV + +K
Sbjct: 844 RENFRTENISFEVADFETAYHAILGRPALAKFMAVPHYTYMMMK 887
>UniRef100_Q7XXB9 OSJNBa0027O01.4 protein [Oryza sativa]
Length = 2013
Score = 374 bits (961), Expect = e-101
Identities = 233/731 (31%), Positives = 372/731 (50%), Gaps = 50/731 (6%)
Query: 1642 QQDETGRKEHAIYYLSKKFTECESRYSMLEKTCCALAWAAKRLRHYMINHTTWLISKMDP 1701
++ + + IY++S+ + ++RY ++K + ++L HY H+ +++ P
Sbjct: 1320 EEGHVQKVQRPIYFVSEVLADSKTRYPQVQKLLYGILITTRKLSHYFQGHSVTVVTSF-P 1378
Query: 1702 IKYIFEKPALIGRIARWQMLLSEYDIEYRSQKAIKGSILADHLAHQPLEDYRPIKFDFPD 1761
+ I + GRIA+W + L DI ++ + +IK LAD +A ++ + D P
Sbjct: 1379 LGDILHNREVNGRIAKWALELMSLDISFKPRISIKSQALADFVA-----EWTECQEDTPA 1433
Query: 1762 EEIMYLKMKDCDEPLFGEGPDPDSVWGLIFDGAVNVYGNGIGAVLLTPKGTHIPFTARLR 1821
E + + W + FDG+ + G G G VL++P G + + +
Sbjct: 1434 ENMEH--------------------WTMHFDGSKRLSGTGAGVVLISPTGERLSYVLWIH 1473
Query: 1822 FDCTNNIAEYEACIMGIEEAIDLRIKNIEIYGDSALVINQIKGKWETLHAGLIPYRDYAR 1881
F ++N+AEYEA + G+ AI L IK + + GDS LV+NQ+ +W L ++ YR R
Sbjct: 1474 FSASHNVAEYEALLHGLRIAISLGIKRLIVRGDSQLVVNQVMKEWSYLDDNMMAYRQEVR 1533
Query: 1882 RLLTFFNKVELHHIPRDENQMADALATLSSMIKVN------HHNDVPLISVKFLDRPAYV 1935
+L F+ +EL H+ R N+ AD LA S + H P + K + A
Sbjct: 1534 KLEDKFDGLELSHVLRHNNEAADRLANFGSKREAAPSDVFVEHLYTPTVPHKDTTQVAGT 1593
Query: 1936 FAAEVVFDDKPWFHDIKVFLQTREYPPGASNKDK-KTLRRLSSNFFLNGDILYKRNFDTV 1994
A +V D W + FL ++E P +KD+ + + R S + L+ LYK++ +
Sbjct: 1594 HDAAMVEVD--WREPLIRFLTSQELP---QDKDEAERISRRSKLYVLHEAELYKKSPSGI 1648
Query: 1995 LLRCVDKYEADLLIHEIHEGSFGIHPNGHTMAKKILRAGYYWMTMESDCYKHTRKCHKCQ 2054
L RCV E L+ +IH G G H T+ K R G++W T SD K R C CQ
Sbjct: 1649 LQRCVSLEEGRQLLKDIHSGICGNHAAARTIVGKAYRQGFFWPTAVSDADKIVRTCEGCQ 1708
Query: 2055 IYADKIHMPPTTLNLLSSPWPFSMWGIDMIGRIEPKASNGHRFILVAIDYFTKWVEAASY 2114
+A +IH+P L + WPF++WG+DM+G + KA G+ + VAID F+KW+EA
Sbjct: 1709 FFARQIHLPAQELQTIPLSWPFAVWGLDMVGPFK-KAVGGYTHLFVAIDKFSKWIEAKPV 1767
Query: 2115 ANVTKQVVVKFIKNHIICRYGIPNRIITDNGTNLNNKMMKELCDDFKIEHHNSSPYRPQM 2174
+T F N I+ R+G+PNRIITDNGT + K+ C+DF I+ +S P
Sbjct: 1768 VTITADNARDFFIN-IVHRFGVPNRIITDNGTQFTGGVFKDFCEDFGIKICYASVAHPMS 1826
Query: 2175 NGAVEAANKNI-----KRIVQKMVVTYKDWHEMLPFALHGYRTSVRTSIGATPFSLVYGM 2229
NG VE AN I R+ ++ W + LP L RT+ + G +PF LVYG
Sbjct: 1827 NGQVERANGMILQGIKARVFDRLKPYAGKWVQQLPSVLWSLRTTPSRATGQSPFFLVYGA 1886
Query: 2230 EAVLPVEVEIPSLRVLMEVDLSEAEWVQNRYDQLNLIEEKRMAALCHGQLYQKRMKQAFD 2289
EA+LP EVE SLR + E + ++R D L+ +EE R AAL Y + +++ +
Sbjct: 1887 EAMLPSEVEFESLRFR---NFREERYEEDRVDDLHRLEEVREAALIRSARYLQGLRRYHN 1943
Query: 2290 KKVRPREFKEGDLVLKKIFSFQPDSRGKWAPNYEGPYVVKKAFSGGAMTLQTMDGEELPR 2349
+ VR R F GDLVL+KI + + R K +P +EGP+++ + G+ L+ DG +
Sbjct: 1944 RNVRSRAFLVGDLVLRKIQTTR--DRHKLSPLWEGPFIISEVTRPGSYRLKREDGTLVDN 2001
Query: 2350 PVNTDTVKKYF 2360
N + +++++
Sbjct: 2002 SWNIEHLRRFY 2012
Score = 57.4 bits (137), Expect = 5e-06
Identities = 50/221 (22%), Positives = 93/221 (41%), Gaps = 19/221 (8%)
Query: 184 VQVPHKFKIPDFEKYKGSSCPEEHLKMYVRRMPAYAQDDQILIYYFQESLTGPASKWYTN 243
V P FK EKY G++ PE L +Y + A D++ + Y +L A W
Sbjct: 428 VDWPAGFKPTGIEKYDGTTNPESWLTVYGLAIRAAGGDNKAMANYLPVALADSARSWLHG 487
Query: 244 LDKTRVQTFRDLCEAFVEQYSYNVDMTPDRSDLQAMTQGDKETFKEYAQRWRDTAAQVSP 303
L + + ++ +L + F+ + + + DL + Q E+ ++Y +R+ + ++S
Sbjct: 488 LPRGTIGSWAELRDHFIANFQGTFERPGTQYDLYNVIQKSGESLRDYIRRFSEQRNKISD 547
Query: 304 RIEEKEMTKLFLKTLNHFYYKKMVGS-------TPKSFAEMVGMGVQLEEGVREGR---- 352
I + + F K + H +++VG T K E + E+ V +
Sbjct: 548 -ITDDVIIAAFTKGIRH---EELVGKFGRKPPRTVKLMFEKANEYAKAEDAVTASKQSGP 603
Query: 353 --LVKNATPANGTKKTGNHFPRKKE--QEVGMVTHGMPQQT 389
TPA G + NH RK++ + V +H Q++
Sbjct: 604 SWKQNKGTPATGGGGSNNHKDRKRKPAELVATASHSSRQRS 644
Score = 46.6 bits (109), Expect = 0.009
Identities = 29/104 (27%), Positives = 50/104 (47%), Gaps = 5/104 (4%)
Query: 873 LSKVLVDTGSSLNVLSKTTYTQLTYQDAPLRRSGVMVKA-FDGSRKDVLGEVVLPITVGP 931
L + L+D GS+LN+L T + + L+ S G LG++ LP+T G
Sbjct: 785 LRRTLIDGGSALNILFAKTLDDMQIPRSELKPSNAPFHGVIPGLSATPLGQITLPVTFGT 844
Query: 932 Q----VFQVNFQVMDIQASYSCLLGRPWIHEAGAVTSTLHQKLK 971
+ ++F+V D + +Y +LGRP + + AV + +K
Sbjct: 845 RENFRTENISFEVADFETAYHAILGRPALAKFMAVPHYTYMMMK 888
>UniRef100_Q7F9R8 OSJNBb0054B09.2 protein [Oryza sativa]
Length = 1992
Score = 374 bits (960), Expect = e-101
Identities = 230/731 (31%), Positives = 371/731 (50%), Gaps = 50/731 (6%)
Query: 1642 QQDETGRKEHAIYYLSKKFTECESRYSMLEKTCCALAWAAKRLRHYMINHTTWLISKMDP 1701
++ + + IY++S+ + ++RY ++K + ++L HY H+ +++ P
Sbjct: 1299 EEGHVQKVQRPIYFVSEVLADSKTRYPQVQKLLYGILITTRKLSHYFQGHSVTVVTSF-P 1357
Query: 1702 IKYIFEKPALIGRIARWQMLLSEYDIEYRSQKAIKGSILADHLAHQPLEDYRPIKFDFPD 1761
+ I GRIA+W + L DI ++ + +IK LAD +A ++ + D P
Sbjct: 1358 LGDILHNREANGRIAKWALELMSLDISFKPRISIKSQALADFVA-----EWTECQEDTPA 1412
Query: 1762 EEIMYLKMKDCDEPLFGEGPDPDSVWGLIFDGAVNVYGNGIGAVLLTPKGTHIPFTARLR 1821
E++ + W + FDG+ + G G G VL++P G + + +
Sbjct: 1413 EKMEH--------------------WTMHFDGSKRLSGTGAGVVLISPTGERLSYVLWIH 1452
Query: 1822 FDCTNNIAEYEACIMGIEEAIDLRIKNIEIYGDSALVINQIKGKWETLHAGLIPYRDYAR 1881
F ++N+AEYEA + G+ AI L IK + + GDS LV+NQ+ +W L ++ YR R
Sbjct: 1453 FSASHNVAEYEALLHGLRIAISLGIKRLIVRGDSQLVVNQVMKEWSCLDDNMMAYRQEVR 1512
Query: 1882 RLLTFFNKVELHHIPRDENQMADALATLSSMIKVN------HHNDVPLISVKFLDRPAYV 1935
+L F+ +EL H+ R N+ AD LA S ++ H P + K D
Sbjct: 1513 KLEDKFDGLELSHVLRHNNEAADRLANFGSKREMAPSDVFVEHLYAPTVPHK--DTTQVA 1570
Query: 1936 FAAEVVFDDKPWFHDIKVFLQTREYPPGASNKDK-KTLRRLSSNFFLNGDILYKRNFDTV 1994
+V + W + FL ++E P +KD+ + + R S + ++ LYK++ +
Sbjct: 1571 GTHDVAMVEADWREPLIRFLTSQELP---QDKDEAERISRRSKLYVMHEAELYKKSPSGI 1627
Query: 1995 LLRCVDKYEADLLIHEIHEGSFGIHPNGHTMAKKILRAGYYWMTMESDCYKHTRKCHKCQ 2054
L RCV E L+ +IH G G H T+ K R G++W T SD K R C CQ
Sbjct: 1628 LQRCVSLEEGRQLLKDIHSGICGNHAAARTIVGKAYRQGFFWPTAVSDADKIVRTCEGCQ 1687
Query: 2055 IYADKIHMPPTTLNLLSSPWPFSMWGIDMIGRIEPKASNGHRFILVAIDYFTKWVEAASY 2114
+A +IH+P L + WPF++WG+DM+G + KA G+ + VAID F+KW+EA
Sbjct: 1688 FFARQIHLPAQELQTIPLSWPFAVWGLDMVGPFK-KAVGGYTHLFVAIDKFSKWIEAKPV 1746
Query: 2115 ANVTKQVVVKFIKNHIICRYGIPNRIITDNGTNLNNKMMKELCDDFKIEHHNSSPYRPQM 2174
+T F N I+ R+G+PNRIITDNGT + K+ C+DF I+ +S P
Sbjct: 1747 VTITADNARDFFIN-IVHRFGVPNRIITDNGTQFTGGVFKDFCEDFGIKICYASVAHPMS 1805
Query: 2175 NGAVEAANKNI-----KRIVQKMVVTYKDWHEMLPFALHGYRTSVRTSIGATPFSLVYGM 2229
NG VE AN I R+ ++ W + LP L RT+ + G +PF LVYG
Sbjct: 1806 NGQVERANGMILQGIKARVFDRLKPYAGKWVQQLPSVLWSLRTTPSRATGQSPFFLVYGA 1865
Query: 2230 EAVLPVEVEIPSLRVLMEVDLSEAEWVQNRYDQLNLIEEKRMAALCHGQLYQKRMKQAFD 2289
EA+LP EVE SLR + E + ++R D L+ +EE R AAL Y + +++ +
Sbjct: 1866 EAMLPSEVEFESLRFR---NFREERYEEDRVDDLHRLEEVREAALIRSARYLQGLRRYHN 1922
Query: 2290 KKVRPREFKEGDLVLKKIFSFQPDSRGKWAPNYEGPYVVKKAFSGGAMTLQTMDGEELPR 2349
+ VR R F GDLVL+KI + + R K +P +EGP+++ + G+ L+ DG +
Sbjct: 1923 RNVRSRAFLVGDLVLRKIQTTR--DRHKLSPLWEGPFIISEVTRPGSYRLKREDGTLVDN 1980
Query: 2350 PVNTDTVKKYF 2360
N + +++++
Sbjct: 1981 SWNIEHLRRFY 1991
Score = 57.4 bits (137), Expect = 5e-06
Identities = 51/221 (23%), Positives = 93/221 (42%), Gaps = 19/221 (8%)
Query: 184 VQVPHKFKIPDFEKYKGSSCPEEHLKMYVRRMPAYAQDDQILIYYFQESLTGPASKWYTN 243
V P FK EKY G++ PE L +Y + A D + + Y +L A W
Sbjct: 427 VDWPAGFKPTGIEKYDGTTNPESWLTVYGLAIRAVGGDSKAMANYLPVALEDSARSWLHG 486
Query: 244 LDKTRVQTFRDLCEAFVEQYSYNVDMTPDRSDLQAMTQGDKETFKEYAQRWRDTAAQVSP 303
L + + ++ +L + F+ + + + DL + Q E+ ++Y +R+ + ++S
Sbjct: 487 LPRGTIGSWAELRDHFIANFQGTFERPGTQYDLYNVIQKSGESLRDYIRRFSEQRNKISD 546
Query: 304 RIEEKEMTKLFLKTLNHFYYKKMVGS-------TPKSFAEMVGMGVQLEEGVREGR---- 352
I + + F K + H +++VG T K E + E+ V +
Sbjct: 547 -ITDDVIIAAFTKGIRH---EELVGKFGRKPPRTVKLMFEKANEYAKAEDAVIASKQSGP 602
Query: 353 --LVKNATPANGTKKTGNHFPRKKE--QEVGMVTHGMPQQT 389
K TPA G + NH RK++ + V +H Q++
Sbjct: 603 SWKPKKDTPATGGGGSNNHKDRKRKPAELVATASHSSRQRS 643
Score = 46.6 bits (109), Expect = 0.009
Identities = 29/104 (27%), Positives = 50/104 (47%), Gaps = 5/104 (4%)
Query: 873 LSKVLVDTGSSLNVLSKTTYTQLTYQDAPLRRSGVMVKA-FDGSRKDVLGEVVLPITVGP 931
L + L+D GS+LN+L T + + L+ S G LG++ LP+T G
Sbjct: 784 LRRTLIDGGSALNILFAKTLDDMQIPRSELKPSNAPFHGVIPGLSATPLGQITLPVTFGT 843
Query: 932 Q----VFQVNFQVMDIQASYSCLLGRPWIHEAGAVTSTLHQKLK 971
+ ++F+V D + +Y +LGRP + + AV + +K
Sbjct: 844 RENFRTENISFEVADFETAYHAILGRPALAKFMAVPHYTYMMMK 887
>UniRef100_Q8S7A3 Putative retroelement [Oryza sativa]
Length = 2017
Score = 374 bits (959), Expect = e-101
Identities = 233/734 (31%), Positives = 372/734 (49%), Gaps = 53/734 (7%)
Query: 1642 QQDETG---RKEHAIYYLSKKFTECESRYSMLEKTCCALAWAAKRLRHYMINHTTWLISK 1698
+++E G + + IY++S+ + ++RY ++K + ++L HY H +++
Sbjct: 1321 EREEDGHVQKVQRPIYFVSEVLADSKTRYPQVQKLLYGILITTRKLSHYFQGHLVTVVTS 1380
Query: 1699 MDPIKYIFEKPALIGRIARWQMLLSEYDIEYRSQKAIKGSILADHLAHQPLEDYRPIKFD 1758
P+ I GRIA+W + L DI ++ + +IK LAD +A ++ + D
Sbjct: 1381 F-PLGDILHNREANGRIAKWALELMSLDISFKPRISIKSQALADFVA-----EWTECQED 1434
Query: 1759 FPDEEIMYLKMKDCDEPLFGEGPDPDSVWGLIFDGAVNVYGNGIGAVLLTPKGTHIPFTA 1818
P E++ + W + FDG+ + G G G VL++P G + +
Sbjct: 1435 TPAEKVEH--------------------WTMHFDGSKRLSGTGAGVVLISPTGERLSYVL 1474
Query: 1819 RLRFDCTNNIAEYEACIMGIEEAIDLRIKNIEIYGDSALVINQIKGKWETLHAGLIPYRD 1878
+ F ++N+AEYEA + G+ AI L IK + + GDS LV+NQ+ +W L ++ YR
Sbjct: 1475 WIHFSASHNVAEYEALLHGLRIAISLGIKRLIVRGDSQLVVNQVMKEWSCLDDNMMAYRQ 1534
Query: 1879 YARRLLTFFNKVELHHIPRDENQMADALATLSSMIKVN------HHNDVPLISVKFLDRP 1932
R+L F+ +EL H+ R N+ AD LA S +V H P + K D
Sbjct: 1535 EVRKLEDKFDGLELSHVLRHNNKAADRLANFGSKREVAPSDVFVEHLYAPTVPHK--DTT 1592
Query: 1933 AYVFAAEVVFDDKPWFHDIKVFLQTREYPPGASNKDK-KTLRRLSSNFFLNGDILYKRNF 1991
+V + W + FL ++E P +KD+ + + R S + ++ LYK++
Sbjct: 1593 QVAGTHDVAMVEADWREPLIRFLTSQELP---QDKDEAERISRRSRLYVMHEAELYKKSP 1649
Query: 1992 DTVLLRCVDKYEADLLIHEIHEGSFGIHPNGHTMAKKILRAGYYWMTMESDCYKHTRKCH 2051
+L RCV E L+ +IH G G H T+ K R G++W T SD K R C
Sbjct: 1650 SGILQRCVSLEEGRQLLKDIHSGICGNHAAARTIVGKAYRQGFFWPTAVSDADKIVRTCE 1709
Query: 2052 KCQIYADKIHMPPTTLNLLSSPWPFSMWGIDMIGRIEPKASNGHRFILVAIDYFTKWVEA 2111
CQ +A +IH+P L + WPF++WG+DM+G + KA G+ + VAID F+KW+EA
Sbjct: 1710 GCQFFARQIHLPAQELQTIPLSWPFAVWGLDMVGPFK-KAFGGYTHLFVAIDKFSKWIEA 1768
Query: 2112 ASYANVTKQVVVKFIKNHIICRYGIPNRIITDNGTNLNNKMMKELCDDFKIEHHNSSPYR 2171
+T F N I+ R+G+PNRIITDNG + K+ C+DF I+ +S
Sbjct: 1769 KPVVTITADNARDFFIN-IVHRFGVPNRIITDNGRQFTGGVFKDFCEDFGIKICYASVAH 1827
Query: 2172 PQMNGAVEAANKNI-----KRIVQKMVVTYKDWHEMLPFALHGYRTSVRTSIGATPFSLV 2226
P NG VE AN I R+ ++ W E LP L RT+ + G +PF LV
Sbjct: 1828 PMSNGQVERANGMILQGIKARVFDRLKPYAGKWVEQLPSVLWSLRTTPSRATGQSPFFLV 1887
Query: 2227 YGMEAVLPVEVEIPSLRVLMEVDLSEAEWVQNRYDQLNLIEEKRMAALCHGQLYQKRMKQ 2286
YG EA+LP EVE SLR + E + ++R D L+ +EE R AAL Y + +++
Sbjct: 1888 YGAEAMLPSEVEFESLRFR---NFREERYEEDRVDDLHRLEEVREAALIQSARYLQGLRR 1944
Query: 2287 AFDKKVRPREFKEGDLVLKKIFSFQPDSRGKWAPNYEGPYVVKKAFSGGAMTLQTMDGEE 2346
++ VR R F GDLVL+KI + + R K +P +EGP+++ + G+ L+ DG
Sbjct: 1945 YHNRNVRSRAFLVGDLVLRKIQTTR--DRHKLSPLWEGPFIISEVTRPGSYRLKREDGTL 2002
Query: 2347 LPRPVNTDTVKKYF 2360
+ N + +++++
Sbjct: 2003 IDNSWNIEHLRRFY 2016
Score = 62.0 bits (149), Expect = 2e-07
Identities = 52/221 (23%), Positives = 93/221 (41%), Gaps = 19/221 (8%)
Query: 184 VQVPHKFKIPDFEKYKGSSCPEEHLKMYVRRMPAYAQDDQILIYYFQESLTGPASKWYTN 243
V P FK EKY G++ PE L +Y + A D + + Y +L A W
Sbjct: 432 VDWPAGFKPTGIEKYDGTTNPESWLTVYGLAIRAAGGDSKAMANYLPVALADSARSWLHG 491
Query: 244 LDKTRVQTFRDLCEAFVEQYSYNVDMTPDRSDLQAMTQGDKETFKEYAQRWRDTAAQVSP 303
L + ++++ +L + F+ + + + DL + Q E+ ++Y +R+ + ++S
Sbjct: 492 LPRGTIESWAELRDHFIANFQGTFERPGTQYDLYNVIQKSGESLRDYIRRFSEQRNKISD 551
Query: 304 RIEEKEMTKLFLKTLNHFYYKKMVGS-------TPKSFAEMVGMGVQLEEGVREGR---- 352
I + + F K + H +++VG T K E + E+ V +
Sbjct: 552 -ITDDVIIAAFTKGIRH---EELVGKFGRKPPRTVKLMFEKANEYAKAEDAVTASKQSGP 607
Query: 353 --LVKNATPANGTKKTGNHFPRKKEQE--VGMVTHGMPQQT 389
TPA G + NH RK++ E V TH Q++
Sbjct: 608 SWKPNKGTPATGGGGSNNHKDRKRKPEELVATATHSSRQRS 648
Score = 46.6 bits (109), Expect = 0.009
Identities = 29/104 (27%), Positives = 50/104 (47%), Gaps = 5/104 (4%)
Query: 873 LSKVLVDTGSSLNVLSKTTYTQLTYQDAPLRRSGVMVKA-FDGSRKDVLGEVVLPITVGP 931
L + L+D GS+LN+L T + + L+ S G LG++ LP+T G
Sbjct: 789 LRRTLIDGGSALNILFAKTLDDMQIPRSELKPSNAPFHGVIPGLSATPLGQITLPVTFGT 848
Query: 932 Q----VFQVNFQVMDIQASYSCLLGRPWIHEAGAVTSTLHQKLK 971
+ ++F+V D + +Y +LGRP + + AV + +K
Sbjct: 849 RENFRTENISFEVADFETAYHAILGRPALAKFMAVPHYTYMMMK 892
>UniRef100_Q8LMM6 Putative gag-pol [Oryza sativa]
Length = 1986
Score = 373 bits (958), Expect = e-101
Identities = 229/731 (31%), Positives = 368/731 (50%), Gaps = 50/731 (6%)
Query: 1642 QQDETGRKEHAIYYLSKKFTECESRYSMLEKTCCALAWAAKRLRHYMINHTTWLISKMDP 1701
++ + + IY++S+ + ++RY ++K + ++L HY H+ +++ P
Sbjct: 1293 EEGHVQKVQRPIYFVSEVLADSKARYPQVQKLLYGILITTRKLSHYFQGHSVTVVTSF-P 1351
Query: 1702 IKYIFEKPALIGRIARWQMLLSEYDIEYRSQKAIKGSILADHLAHQPLEDYRPIKFDFPD 1761
+ + GRIA+W + L DI ++ + +IK LAD +A ++ + D P
Sbjct: 1352 LGDVLHNREANGRIAKWALELMSLDISFKPRTSIKSQALADFVA-----EWTECQEDTPV 1406
Query: 1762 EEIMYLKMKDCDEPLFGEGPDPDSVWGLIFDGAVNVYGNGIGAVLLTPKGTHIPFTARLR 1821
E+I + W + FDG+ + G G G VL++P G + + +
Sbjct: 1407 EKIEH--------------------WTMHFDGSKRLSGTGAGVVLISPTGERLSYVLWIH 1446
Query: 1822 FDCTNNIAEYEACIMGIEEAIDLRIKNIEIYGDSALVINQIKGKWETLHAGLIPYRDYAR 1881
F ++N+AEYEA + G+ AI L IK + + GDS LV+NQ+ +W + ++ YR R
Sbjct: 1447 FSASHNVAEYEALLHGLRIAISLGIKRLIVRGDSQLVVNQVMKEWSCIDDNMMAYRQEVR 1506
Query: 1882 RLLTFFNKVELHHIPRDENQMADALATLSSMIKVN------HHNDVPLISVKFLDRPAYV 1935
+L F+ +EL H+ R N+ AD LA S + H P + K D
Sbjct: 1507 KLEDKFDGLELSHVLRHNNEAADRLANFGSKRETAPSDVFVEHLYTPTVPHK--DTTQDA 1564
Query: 1936 FAAEVVFDDKPWFHDIKVFLQTREYPPGASNKDK-KTLRRLSSNFFLNGDILYKRNFDTV 1994
+ + W FL ++E P +KD+ + + R S + L+ LYK++ +
Sbjct: 1565 DTRNIAMVEADWREPFIRFLTSQELP---QDKDEAERISRRSKLYVLHESELYKKSPSGI 1621
Query: 1995 LLRCVDKYEADLLIHEIHEGSFGIHPNGHTMAKKILRAGYYWMTMESDCYKHTRKCHKCQ 2054
L RCV E L+ +IH G G H T+ K R G++W T SD K R C CQ
Sbjct: 1622 LQRCVSLEEGRQLLKDIHSGICGNHAAARTIVGKAYRQGFFWPTAVSDADKIVRTCEGCQ 1681
Query: 2055 IYADKIHMPPTTLNLLSSPWPFSMWGIDMIGRIEPKASNGHRFILVAIDYFTKWVEAASY 2114
+A +IH+P L + WPF++WG+DM+G + KA G+ + VAID F+KW+EA
Sbjct: 1682 FFARQIHLPAQELQTIPLSWPFAVWGLDMVGPFK-KAVGGYTHLFVAIDKFSKWIEAKPV 1740
Query: 2115 ANVTKQVVVKFIKNHIICRYGIPNRIITDNGTNLNNKMMKELCDDFKIEHHNSSPYRPQM 2174
+T F N I+ R+G+PNRIITDNGT + K+ C+DF I+ +S P
Sbjct: 1741 VTITADNARDFFIN-IVHRFGVPNRIITDNGTQFTGGVFKDFCEDFGIKICYASVAHPMS 1799
Query: 2175 NGAVEAANKNI-----KRIVQKMVVTYKDWHEMLPFALHGYRTSVRTSIGATPFSLVYGM 2229
NG VE AN I R+ ++ W + LP L RT+ + G +PF LVYG
Sbjct: 1800 NGQVERANGMILQGIKARVFDRLKPYAGKWVQQLPSVLWSLRTTPSRATGQSPFFLVYGA 1859
Query: 2230 EAVLPVEVEIPSLRVLMEVDLSEAEWVQNRYDQLNLIEEKRMAALCHGQLYQKRMKQAFD 2289
EA+LP EVE SLR + E + ++R D L+ +EE R AAL Y + +++ +
Sbjct: 1860 EAMLPSEVEFESLRFR---NFREERYEEDRVDDLHRLEEVREAALIQSARYLQGLRRYHN 1916
Query: 2290 KKVRPREFKEGDLVLKKIFSFQPDSRGKWAPNYEGPYVVKKAFSGGAMTLQTMDGEELPR 2349
+ VR R F GDLVL+KI + + R K +P +EGP+++ + G+ L+ DG +
Sbjct: 1917 RNVRSRAFLVGDLVLRKIQTTR--DRHKLSPLWEGPFIISEVTRPGSYRLKREDGTLVDN 1974
Query: 2350 PVNTDTVKKYF 2360
N + +++++
Sbjct: 1975 SWNIEHLRRFY 1985
Score = 59.3 bits (142), Expect = 1e-06
Identities = 55/242 (22%), Positives = 98/242 (39%), Gaps = 18/242 (7%)
Query: 184 VQVPHKFKIPDFEKYKGSSCPEEHLKMYVRRMPAYAQDDQILIYYFQESLTGPASKWYTN 243
V P FK EKY G++ PE L +Y + A D++ + Y +L A W
Sbjct: 428 VDWPAGFKPTGIEKYDGTTNPESWLTVYGLAIRAAGGDNKAMANYLPVALADSARSWLHG 487
Query: 244 LDKTRVQTFRDLCEAFVEQYSYNVDMTPDRSDLQAMTQGDKETFKEYAQRWRDTAAQVSP 303
L + + ++ +L + F+ + + + DL + Q E+ ++Y +R+ + ++S
Sbjct: 488 LPRGTIGSWAELRDHFIANFQGTFERPGTQYDLYNVIQKSGESLRDYIRRFSEQRNKISD 547
Query: 304 RIEEKEMTKLFLKTLNHFYYKKMVGS-------TPKSFAEMVGMGVQLEEGVREGR---- 352
I + + F K + H +++VG T K E + E+ V +
Sbjct: 548 -ITDDVIIAAFTKGIRH---EELVGKFGRKPPRTVKLMFEKANEYAKAEDAVTASKQSGP 603
Query: 353 --LVKNATPANGTKKTGNHFPRKKEQEVGMVTHGMPQQTYPAYQHIAAITPTSHPFQQTN 410
KN TPA G + NH RK++ E + T + I + P +
Sbjct: 604 SWKPKN-TPATGGGGSNNHKDRKRKPEELVATAAHSSRQRSRVNTFDKIMNSQCPHHPNS 662
Query: 411 NH 412
NH
Sbjct: 663 NH 664
Score = 45.1 bits (105), Expect = 0.027
Identities = 28/104 (26%), Positives = 49/104 (46%), Gaps = 5/104 (4%)
Query: 873 LSKVLVDTGSSLNVLSKTTYTQLTYQDAPLRRSGVMVKA-FDGSRKDVLGEVVLPITVGP 931
L + L+D GS+LN+L T + + L+ S G LG++ LP+T G
Sbjct: 784 LRRTLIDGGSALNILFAKTLDDMQIPRSELKSSNAPFHGVIPGLSATPLGQITLPVTFGT 843
Query: 932 Q----VFQVNFQVMDIQASYSCLLGRPWIHEAGAVTSTLHQKLK 971
+ ++F+V D + +Y +LG P + + AV + +K
Sbjct: 844 RENFRTENISFEVADFETAYHAILGHPALAKFMAVPHYTYMMMK 887
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.327 0.140 0.434
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,907,772,628
Number of Sequences: 2790947
Number of extensions: 170459688
Number of successful extensions: 824590
Number of sequences better than 10.0: 7866
Number of HSP's better than 10.0 without gapping: 5097
Number of HSP's successfully gapped in prelim test: 2980
Number of HSP's that attempted gapping in prelim test: 675889
Number of HSP's gapped (non-prelim): 50231
length of query: 2360
length of database: 848,049,833
effective HSP length: 144
effective length of query: 2216
effective length of database: 446,153,465
effective search space: 988676078440
effective search space used: 988676078440
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 83 (36.6 bits)
Medicago: description of AC146971.7


