

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
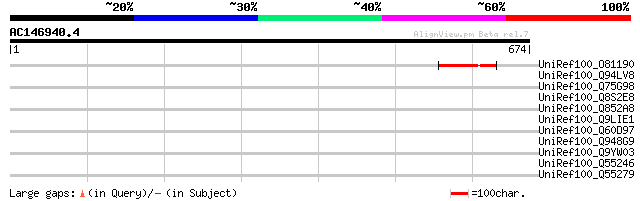
Query= AC146940.4 + phase: 1 /pseudo
(674 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_O81190 Putative transposase [Arabidopsis thaliana] 67 1e-09
UniRef100_Q94LV8 Hypothetical protein OSJNBa0082M15.8 [Oryza sat... 44 0.016
UniRef100_Q75G98 Hypothetical protein OSJNBa0042F15.17 [Oryza sa... 44 0.016
UniRef100_Q8S2E8 P0022F10.3 protein [Oryza sativa] 44 0.020
UniRef100_Q852A8 Hypothetical protein OSJNBb0081B07.17 [Oryza sa... 40 0.23
UniRef100_Q9LIE1 Transposase-like protein [Arabidopsis thaliana] 37 1.5
UniRef100_Q60D97 Putative polyprotein [Oryza sativa] 37 2.5
UniRef100_Q948G9 Putative transposable element [Oryza sativa] 36 3.3
UniRef100_Q9YW03 ORF MSV089 Putative NTPase, Rabbit fibroma viru... 36 4.3
UniRef100_Q55246 M protein [Streptococcus sp] 36 4.3
UniRef100_Q55279 MLG72 precursor [Streptococcus sp.] 36 4.3
>UniRef100_O81190 Putative transposase [Arabidopsis thaliana]
Length = 729
Score = 67.4 bits (163), Expect = 1e-09
Identities = 38/77 (49%), Positives = 52/77 (67%), Gaps = 2/77 (2%)
Query: 557 EKLECV*AYSLKKKRNRLEHQRLNDLVFVRYNLMLENRNNKIRNYDPINDELLDDHHDNW 616
E+ CV KKRNRLEHQRLNDLVFV YNL L++R+ + R+YDP++ E + D + W
Sbjct: 612 ERNWCVFERIHTKKRNRLEHQRLNDLVFVHYNLRLQHRSKRKRSYDPVDYESI-DKTEFW 670
Query: 617 VLEDSPP-FLTVEELES 632
V+E+ L +ELE+
Sbjct: 671 VVEEEEAGELEYDELEN 687
>UniRef100_Q94LV8 Hypothetical protein OSJNBa0082M15.8 [Oryza sativa]
Length = 645
Score = 43.9 bits (102), Expect = 0.016
Identities = 25/43 (58%), Positives = 30/43 (69%), Gaps = 3/43 (6%)
Query: 569 KKRNRLEHQRLNDLVFVRYNLMLEN-RNNKIRNYDPINDELLD 610
KKRNRL H+R+ DLVFV++N L N R NK R DPI E+ D
Sbjct: 511 KKRNRLLHERMRDLVFVKFNSKLRNKRENKGR--DPIEKEVDD 551
>UniRef100_Q75G98 Hypothetical protein OSJNBa0042F15.17 [Oryza sativa]
Length = 1005
Score = 43.9 bits (102), Expect = 0.016
Identities = 25/43 (58%), Positives = 30/43 (69%), Gaps = 3/43 (6%)
Query: 569 KKRNRLEHQRLNDLVFVRYNLMLEN-RNNKIRNYDPINDELLD 610
KKRNRL H+R+ DLVFV++N L N R NK R DPI E+ D
Sbjct: 866 KKRNRLLHERMRDLVFVKFNSKLRNKRENKGR--DPIEKEVDD 906
>UniRef100_Q8S2E8 P0022F10.3 protein [Oryza sativa]
Length = 965
Score = 43.5 bits (101), Expect = 0.020
Identities = 24/43 (55%), Positives = 30/43 (68%), Gaps = 3/43 (6%)
Query: 569 KKRNRLEHQRLNDLVFVRYNLMLEN-RNNKIRNYDPINDELLD 610
KKRNRL H+R+ DLVF+++N L N R NK R DPI E+ D
Sbjct: 822 KKRNRLLHERMRDLVFIKFNSKLRNKRENKGR--DPIEKEVDD 862
>UniRef100_Q852A8 Hypothetical protein OSJNBb0081B07.17 [Oryza sativa]
Length = 779
Score = 40.0 bits (92), Expect = 0.23
Identities = 20/54 (37%), Positives = 37/54 (68%), Gaps = 2/54 (3%)
Query: 569 KKRNRLEHQRLNDLVFVRYNLMLENR-NNKIRNYDPINDELLDDHHDNWVLEDS 621
++ N E QR++ L FV+YNL L++R +K + +DP++ + + D D+WV++ S
Sbjct: 670 QRMNLFERQRMHHLTFVQYNLRLQHRQQHKTKAFDPVSVDNI-DIVDDWVVDRS 722
>UniRef100_Q9LIE1 Transposase-like protein [Arabidopsis thaliana]
Length = 761
Score = 37.4 bits (85), Expect = 1.5
Identities = 22/52 (42%), Positives = 31/52 (59%), Gaps = 3/52 (5%)
Query: 569 KKRNRLEHQRLNDLVFVRYNLMLENRNNKIRNYDPINDELLDDHHDNWVLED 620
+ +N +E QRLNDLVFV+YN+ L ++ D + D L H + VLED
Sbjct: 646 ESKNSIERQRLNDLVFVQYNMRLRRIGSESSGDDTV-DPL--SHSNMEVLED 694
>UniRef100_Q60D97 Putative polyprotein [Oryza sativa]
Length = 1086
Score = 36.6 bits (83), Expect = 2.5
Identities = 19/45 (42%), Positives = 27/45 (59%), Gaps = 4/45 (8%)
Query: 564 AYSLKKKRNRLEHQRLNDLVFVRYNLMLE----NRNNKIRNYDPI 604
A+ K RNRL H++LN LV+V YNL L N ++ + DP+
Sbjct: 772 AFVHTKVRNRLTHKKLNKLVYVNYNLRLRIQQANAQIRVEDDDPL 816
>UniRef100_Q948G9 Putative transposable element [Oryza sativa]
Length = 891
Score = 36.2 bits (82), Expect = 3.3
Identities = 20/41 (48%), Positives = 26/41 (62%), Gaps = 2/41 (4%)
Query: 564 AYSLKKKRNRLEHQRLNDLVFVRYNLML--ENRNNKIRNYD 602
A+ K RNRL H++LN LV+V YNL L + N +IR D
Sbjct: 660 AFVHTKVRNRLTHKKLNKLVYVNYNLRLRIQQANAQIRCLD 700
>UniRef100_Q9YW03 ORF MSV089 Putative NTPase, Rabbit fibroma virus C5 (Vaccinia D5R)
homolog, similar to PIR:G41700 [Melanoplus sanguinipes
entomopoxvirus]
Length = 834
Score = 35.8 bits (81), Expect = 4.3
Identities = 24/92 (26%), Positives = 47/92 (51%), Gaps = 8/92 (8%)
Query: 567 LKKKRNRLEHQRLNDLVFVRYNLMLENRNNKIRNYDPI---NDELLDDHHDNWVLEDSPP 623
L KK+ LE + +N + + + +LE NN I N D I ++ +L+++HD + L
Sbjct: 16 LDKKKCILEEEYVNQIDKLTLDELLEKINNAIENNDSIEEYHEIILNEYHDRFKL----- 70
Query: 624 FLTVEELESLRNDLANMTIQPISNDIGMYVFS 655
F +E + +D + T + ++I ++ S
Sbjct: 71 FFDIELKKKYNSDYFDSTFENFKDEISSFLAS 102
>UniRef100_Q55246 M protein [Streptococcus sp]
Length = 441
Score = 35.8 bits (81), Expect = 4.3
Identities = 21/74 (28%), Positives = 41/74 (55%), Gaps = 8/74 (10%)
Query: 567 LKKKRNRLEHQRLNDLV---FVRYNLMLENRNNKIRNYDPINDELLDDHHDNWVLEDSPP 623
++K+R R E+++L++LV YN +L+ N+ ++ + +ND L N +LE+
Sbjct: 67 IEKERLRKENEQLDELVRKEVAEYNSLLDKHNSLVKKMEVVNDSLQATERANELLENK-- 124
Query: 624 FLTVEELESLRNDL 637
++E + L DL
Sbjct: 125 ---LKENQDLNQDL 135
>UniRef100_Q55279 MLG72 precursor [Streptococcus sp.]
Length = 472
Score = 35.8 bits (81), Expect = 4.3
Identities = 21/74 (28%), Positives = 41/74 (55%), Gaps = 8/74 (10%)
Query: 567 LKKKRNRLEHQRLNDLV---FVRYNLMLENRNNKIRNYDPINDELLDDHHDNWVLEDSPP 623
++K+R R E+++L++LV YN +L+ N+ ++ + +ND L N +LE+
Sbjct: 55 IEKERLRKENEQLDELVRKEVAEYNSLLDKHNSLVKKMEVVNDSLQATERANELLENK-- 112
Query: 624 FLTVEELESLRNDL 637
++E + L DL
Sbjct: 113 ---LKENQDLNQDL 123
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.366 0.162 0.645
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 905,977,530
Number of Sequences: 2790947
Number of extensions: 32440194
Number of successful extensions: 182817
Number of sequences better than 10.0: 11
Number of HSP's better than 10.0 without gapping: 7
Number of HSP's successfully gapped in prelim test: 4
Number of HSP's that attempted gapping in prelim test: 182809
Number of HSP's gapped (non-prelim): 12
length of query: 674
length of database: 848,049,833
effective HSP length: 134
effective length of query: 540
effective length of database: 474,062,935
effective search space: 255993984900
effective search space used: 255993984900
T: 11
A: 40
X1: 14 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.6 bits)
S2: 78 (34.7 bits)
Medicago: description of AC146940.4


