

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146866.2 + phase: 0
(504 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
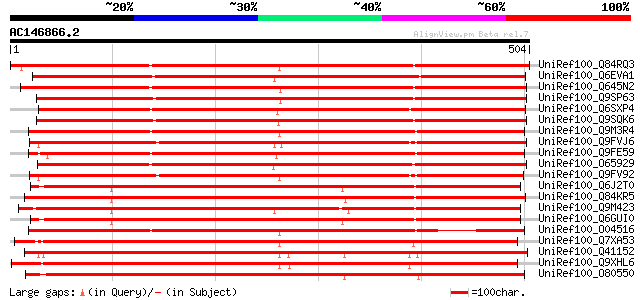
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q84RQ3 Sucrose transporter 4 protein [Lotus japonicus] 829 0.0
UniRef100_Q6EVA1 Putative sucrose-H+ symporter [Datisca glomerata] 726 0.0
UniRef100_Q645N2 Putative sucrose carrier [Ricinus communis] 710 0.0
UniRef100_Q9SP63 Sucrose transporter [Vitis vinifera] 707 0.0
UniRef100_Q6SXP4 Sucrose transporter [Malus domestica] 704 0.0
UniRef100_Q9SQK6 Putative sucrose transporter [Vitis vinifera] 701 0.0
UniRef100_Q9M3R4 Sucrose transporter [Arabidopsis thaliana] 700 0.0
UniRef100_Q9FVJ6 Sucrose transporter [Lycopersicon esculentum] 698 0.0
UniRef100_Q9FE59 Sucrose transporter SUT4 [Arabidopsis thaliana] 697 0.0
UniRef100_O65929 Sucrose/H+ symporter [Daucus carota] 686 0.0
UniRef100_Q9FV92 Sucrose transporter SUT4 [Solanum tuberosum] 686 0.0
UniRef100_Q6J2T0 Sucrose transporter SUT4 [Zea mays] 646 0.0
UniRef100_Q84KR5 Sucrose transporter [Oryza sativa] 645 0.0
UniRef100_Q9M423 Sucrose transporter 2 [Hordeum vulgare] 643 0.0
UniRef100_Q6GUI0 Sucrose transport protein [Zea mays] 642 0.0
UniRef100_O04516 F21M12.35 protein [Arabidopsis thaliana] 623 e-177
UniRef100_Q7XA53 Sucrose transporter [Glycine max] 537 e-151
UniRef100_Q41152 Sucrose carrier [Ricinus communis] 537 e-151
UniRef100_Q9XHL6 Sucrose transport protein SUT1 [Pisum sativum] 530 e-149
UniRef100_O80550 T22J18.12 protein [Arabidopsis thaliana] 530 e-149
>UniRef100_Q84RQ3 Sucrose transporter 4 protein [Lotus japonicus]
Length = 511
Score = 829 bits (2142), Expect = 0.0
Identities = 416/514 (80%), Positives = 453/514 (87%), Gaps = 13/514 (2%)
Query: 1 MPNPTTTNPH---RSRTRSSTSTSTSRPVQPVQPRTPLRQLLRVASVASGIQFGWALQLS 57
MP H RS+ R STS++ P +P R PLRQLLRVASVASGIQFGWALQLS
Sbjct: 1 MPGDPDNRHHHHPRSKNRPSTSSARPPPSRPPPARVPLRQLLRVASVASGIQFGWALQLS 60
Query: 58 LLTPYVQQLGIPHKWASIIWLCGPVSGLFVQPLVGHLSDRCSSRFGRRRPFILVGAASIV 117
LLTPYVQQLGIPH+WASIIWLCGPVSGLFVQPLVGHLSD+C+SRFGRRRPFIL GAASIV
Sbjct: 61 LLTPYVQQLGIPHQWASIIWLCGPVSGLFVQPLVGHLSDKCTSRFGRRRPFILAGAASIV 120
Query: 118 VAVVIIGYAADIGYLIGDDITQNYRPFAIVVFVIGFWILDVANNVTQGPCRALLADLTCN 177
VAV+IIGYAADIG+++GD T+++RP AI VFVIGFWILDVANNVTQGPCRALLADLT
Sbjct: 121 VAVLIIGYAADIGWMLGD--TESFRPAAITVFVIGFWILDVANNVTQGPCRALLADLTSK 178
Query: 178 DARRTRVANAYFSLFMAVGNILGYATGSYSGWYKIFTFTLTPACSISCANLKSAFFLDVA 237
D RRTRVANAYFSLFMA+GNILGYATG+YSGWY+IFTFTL+PAC+ISCANLKSAFFLDVA
Sbjct: 179 DNRRTRVANAYFSLFMAIGNILGYATGAYSGWYRIFTFTLSPACTISCANLKSAFFLDVA 238
Query: 238 FIVVTTYLSIVSAHEVPLSSSGA-------GESGSAEEAFMWELFGTFKYFSMPVWIVLS 290
FI VTTY+SI +AHEVPL+SSGA GESGS EEAFMWELFGTFKYFS VWI+LS
Sbjct: 239 FIAVTTYVSITAAHEVPLNSSGAAHAGEGAGESGSTEEAFMWELFGTFKYFSSTVWIILS 298
Query: 291 VTALTWIGWFPFNLFDTDWMGREIYGGDPEGGLIYDTGVRMGALGLLLNSVVLAVTSLLM 350
VTAL W GWFPF LFDTDWMGREIYG DP GG YD GVRMGALGL+LNSVVL VTSLLM
Sbjct: 299 VTALNWTGWFPFILFDTDWMGREIYGADPNGGPNYDAGVRMGALGLMLNSVVLGVTSLLM 358
Query: 351 ERLCRKRGAGFVWGISNIFMAICFIAMLVLTYAANSIGYVSKGQPPPTGIVIAALAIFTI 410
E+LCRKRGAGFVWGISNI MA+CF+AMLV+TY AN+IGYV K PPTGIVIAAL IFTI
Sbjct: 359 EKLCRKRGAGFVWGISNILMAVCFLAMLVVTYVANTIGYVGK-DLPPTGIVIAALIIFTI 417
Query: 411 LGFPMAITYSVPYALISTHIEPLGLGQGLSMGVLNLAIVVPQIVVSLGSGPWDQLFGGGN 470
LGFP+AITYSVPYALIS H EPLGLGQGLSMGVLNLAIV+PQIVVSLGSGPWDQLFGGGN
Sbjct: 418 LGFPLAITYSVPYALISKHTEPLGLGQGLSMGVLNLAIVIPQIVVSLGSGPWDQLFGGGN 477
Query: 471 SPAFAVAAVAALLSGLLALLAIPRTRTQKPRVRI 504
S AFAV AVAA++SGLLA+LAIPRT TQKP++R+
Sbjct: 478 SAAFAVGAVAAIMSGLLAVLAIPRTGTQKPQIRV 511
>UniRef100_Q6EVA1 Putative sucrose-H+ symporter [Datisca glomerata]
Length = 498
Score = 726 bits (1875), Expect = 0.0
Identities = 360/488 (73%), Positives = 406/488 (82%), Gaps = 12/488 (2%)
Query: 23 SRPVQPVQPRTPLRQLLRVASVASGIQFGWALQLSLLTPYVQQLGIPHKWASIIWLCGPV 82
+R PVQ R LR+LLRV+SVA GIQFGWALQLSLLTPYVQ+LGIPH WAS+IWLCGP+
Sbjct: 11 ARARPPVQARVSLRKLLRVSSVACGIQFGWALQLSLLTPYVQELGIPHAWASVIWLCGPL 70
Query: 83 SGLFVQPLVGHLSDRCSSRFGRRRPFILVGAASIVVAVVIIGYAADIGYLIGDDITQNYR 142
SGLFVQPLVGH+SDRC+SRFGRRRPFI+VGA SI VAV+IIGY+ADIG LIGD T +
Sbjct: 71 SGLFVQPLVGHMSDRCTSRFGRRRPFIVVGALSITVAVLIIGYSADIGSLIGDRGT--VK 128
Query: 143 PFAIVVFVIGFWILDVANNVTQGPCRALLADLTCNDARRTRVANAYFSLFMAVGNILGYA 202
P AI FV+GFWILDVANN+TQGPCRALLADLT D RRTRVANAYFSLFMAVGN+LGYA
Sbjct: 129 PGAIATFVVGFWILDVANNMTQGPCRALLADLTGKDHRRTRVANAYFSLFMAVGNVLGYA 188
Query: 203 TGSYSGWYKIFTFTLTPACSISCANLKSAFFLDVAFIVVTTYLSIVSAHEVPL------- 255
TGSYSGW+KIF TLT AC+++CANLKSAF LD+ FI +TTYLSI +A E PL
Sbjct: 189 TGSYSGWFKIFPLTLTSACNVNCANLKSAFLLDIVFIAITTYLSISAAQESPLDPTDRSA 248
Query: 256 --SSSGAGESGSAEEAFMWELFGTFKYFSMPVWIVLSVTALTWIGWFPFNLFDTDWMGRE 313
+ G G S EEAF+WELFG F+YFS +W++ VTALTWIGWFPF LFDTDWMGRE
Sbjct: 249 NITEEGPGPSSHTEEAFLWELFGAFRYFSASIWVIFFVTALTWIGWFPFLLFDTDWMGRE 308
Query: 314 IYGGDPEGGLIYDTGVRMGALGLLLNSVVLAVTSLLMERLCRKRGAGFVWGISNIFMAIC 373
IYGG P G Y TGVRMGALGL+LNSVVL +TS+LME+LCR GAGFVWG+SNI M++C
Sbjct: 309 IYGGKPNEGQNYSTGVRMGALGLMLNSVVLGITSVLMEKLCRYWGAGFVWGVSNILMSLC 368
Query: 374 FIAMLVLTYAANSIGYVSKGQPPPTGIVIAALAIFTILGFPMAITYSVPYALISTHIEPL 433
F+AMLV+T+ A I Y+ PP IV+AAL IF ILG P+AITYSVPYALIST IE L
Sbjct: 369 FLAMLVVTFVAKRIDYIGHKLPPDV-IVVAALVIFAILGIPLAITYSVPYALISTRIESL 427
Query: 434 GLGQGLSMGVLNLAIVVPQIVVSLGSGPWDQLFGGGNSPAFAVAAVAALLSGLLALLAIP 493
GLGQGLSMGVLNLAIV+PQ+VVSLGSGPWDQLFGGGNSPAFAVAAVAA SGL+A+LAIP
Sbjct: 428 GLGQGLSMGVLNLAIVIPQVVVSLGSGPWDQLFGGGNSPAFAVAAVAAFASGLVAILAIP 487
Query: 494 RTRTQKPR 501
R+R QKPR
Sbjct: 488 RSRAQKPR 495
>UniRef100_Q645N2 Putative sucrose carrier [Ricinus communis]
Length = 509
Score = 710 bits (1832), Expect = 0.0
Identities = 344/501 (68%), Positives = 410/501 (81%), Gaps = 12/501 (2%)
Query: 11 RSRTRSSTSTSTSRPVQPVQPRTPLRQLLRVASVASGIQFGWALQLSLLTPYVQQLGIPH 70
R+R +++T+ +RP V R LR+LLRV S+A GIQFGWALQLSLLTPYVQ+LGIPH
Sbjct: 10 RARAARTSATAAARPPAAVVKRVSLRKLLRVTSIAGGIQFGWALQLSLLTPYVQELGIPH 69
Query: 71 KWASIIWLCGPVSGLFVQPLVGHLSDRCSSRFGRRRPFILVGAASIVVAVVIIGYAADIG 130
WASIIWLCGP+SGL VQPLVGH+SDRC+SRFGRRRPFI VGA I +V+IIG++ADIG
Sbjct: 70 AWASIIWLCGPLSGLVVQPLVGHMSDRCTSRFGRRRPFIFVGAGLICCSVLIIGHSADIG 129
Query: 131 YLIGDDITQNYRPFAIVVFVIGFWILDVANNVTQGPCRALLADLTCNDARRTRVANAYFS 190
+L+GD RP AI VF+IGFWILDVANN+TQGPCRALLADLT D RRTRVANAYFS
Sbjct: 130 WLLGD--RGETRPRAIAVFIIGFWILDVANNMTQGPCRALLADLTGKDHRRTRVANAYFS 187
Query: 191 LFMAVGNILGYATGSYSGWYKIFTFTLTPACSISCANLKSAFFLDVAFIVVTTYLSIVSA 250
LFMAVGN+LGYATGS+S W+K+F FT+T AC+I CANLKSAF+LD+ F+V+TTY+SI +
Sbjct: 188 LFMAVGNVLGYATGSFSNWFKVFPFTVTSACNIDCANLKSAFYLDIVFMVITTYMSITAT 247
Query: 251 HEVPLSSSGAG---------ESGSAEEAFMWELFGTFKYFSMPVWIVLSVTALTWIGWFP 301
E P+ S +SG A+EAF+WEL GTF+YF PVW +L VTAL WIGWFP
Sbjct: 248 KESPIGLSDRSSLITEEISEQSGHAQEAFLWELLGTFRYFPWPVWTILLVTALNWIGWFP 307
Query: 302 FNLFDTDWMGREIYGGDPEGGLIYDTGVRMGALGLLLNSVVLAVTSLLMERLCRKRGAGF 361
F LFDTDWMGREIYGG P G Y++GVRMGA L++NSV+L +TS+LME+LCRK GAGF
Sbjct: 308 FLLFDTDWMGREIYGGAPNDGHNYNSGVRMGAFALMVNSVILGLTSVLMEKLCRKWGAGF 367
Query: 362 VWGISNIFMAICFIAMLVLTYAANSIGYVSKGQPPPTGIVIAALAIFTILGFPMAITYSV 421
+WGISNI MA+CF+AML+ +Y AN IGY+ PP+GIVIAA+ IF +LGFP+AITYSV
Sbjct: 368 MWGISNILMALCFLAMLITSYIANHIGYLGH-DLPPSGIVIAAIIIFAVLGFPLAITYSV 426
Query: 422 PYALISTHIEPLGLGQGLSMGVLNLAIVVPQIVVSLGSGPWDQLFGGGNSPAFAVAAVAA 481
PYALIS+ IEPLGLGQGLSMGVLNLAIV+PQ++VSLGSGPWDQLFGGGNSPAF V A+AA
Sbjct: 427 PYALISSRIEPLGLGQGLSMGVLNLAIVIPQVIVSLGSGPWDQLFGGGNSPAFVVGALAA 486
Query: 482 LLSGLLALLAIPRTRTQKPRV 502
+G++A+L IPR+ KPRV
Sbjct: 487 FAAGVIAILGIPRSGAPKPRV 507
>UniRef100_Q9SP63 Sucrose transporter [Vitis vinifera]
Length = 501
Score = 707 bits (1825), Expect = 0.0
Identities = 346/484 (71%), Positives = 405/484 (83%), Gaps = 11/484 (2%)
Query: 27 QPVQPRTPLRQLLRVASVASGIQFGWALQLSLLTPYVQQLGIPHKWASIIWLCGPVSGLF 86
+PV+PR PLR+LLRVASVA GIQFGWALQLSLLTPYVQ+LGIPH W+SIIWLCGP+SGL
Sbjct: 19 EPVRPRVPLRRLLRVASVACGIQFGWALQLSLLTPYVQELGIPHAWSSIIWLCGPLSGLL 78
Query: 87 VQPLVGHLSDRCSSRFGRRRPFILVGAASIVVAVVIIGYAADIGYLIGDDITQNYRPFAI 146
VQPLVGHLSDRC+SRFGRRRPFI+ GA SIVVAV+IIG++ DIG L+GD + RP A+
Sbjct: 79 VQPLVGHLSDRCNSRFGRRRPFIVAGATSIVVAVLIIGFSTDIGGLLGDGADR--RPRAV 136
Query: 147 VVFVIGFWILDVANNVTQGPCRALLADLTCNDARRTRVANAYFSLFMAVGNILGYATGSY 206
FV+GFW+LDVANNVTQGPCRALLADLT D RRTRVANAYFSLF+AVGN+LG+ATGSY
Sbjct: 137 ATFVVGFWLLDVANNVTQGPCRALLADLTEKDHRRTRVANAYFSLFIAVGNVLGFATGSY 196
Query: 207 SGWYKIFTFTLTPACSISCANLKSAFFLDVAFIVVTTYLSIVSAHEVPLSSSGAG----- 261
SGW++IF FT T +C+ CANLKSAF LD+ FI +TTY+SI +A E+PLSSS
Sbjct: 197 SGWFRIFWFTSTSSCNADCANLKSAFLLDIIFIAITTYISITAAQELPLSSSSRSTHISE 256
Query: 262 ---ESGSAEEAFMWELFGTFKYFSMPVWIVLSVTALTWIGWFPFNLFDTDWMGREIYGGD 318
ES A+EAF+WELFGT +Y S +WI+L VTALTWIGWFPF LFDTDWMGREIYGG
Sbjct: 257 EMAESTHAQEAFLWELFGTLRYLSGSIWIILFVTALTWIGWFPFLLFDTDWMGREIYGGK 316
Query: 319 PEGGLIYDTGVRMGALGLLLNSVVLAVTSLLMERLCRKRGAGFVWGISNIFMAICFIAML 378
P G Y+TGVRMGALGL+LNSVVL +TS+LME+LCRK GAGFVWG+SNI M++CF+ ML
Sbjct: 317 PNEGQNYNTGVRMGALGLMLNSVVLGITSVLMEKLCRKWGAGFVWGLSNILMSLCFLLML 376
Query: 379 VLTYAANSIGYVSKGQPPPTGIVIAALAIFTILGFPMAITYSVPYALISTHIEPLGLGQG 438
+L+ + ++ PP+G+VIAAL +F+ILG P+AITYSVPYALIST IE LGLGQG
Sbjct: 377 ILSAVVKHMDFLGH-DLPPSGVVIAALIVFSILGIPLAITYSVPYALISTRIESLGLGQG 435
Query: 439 LSMGVLNLAIVVPQIVVSLGSGPWDQLFGGGNSPAFAVAAVAALLSGLLALLAIPRTRTQ 498
LSMGVLNLAIV+PQ++VSLGSGPWDQLFGGGNSP+ AVAAVAA SGL+A+LAIPR+
Sbjct: 436 LSMGVLNLAIVIPQVIVSLGSGPWDQLFGGGNSPSLAVAAVAAFASGLVAILAIPRSSAD 495
Query: 499 KPRV 502
K RV
Sbjct: 496 KSRV 499
>UniRef100_Q6SXP4 Sucrose transporter [Malus domestica]
Length = 499
Score = 704 bits (1817), Expect = 0.0
Identities = 348/482 (72%), Positives = 402/482 (83%), Gaps = 11/482 (2%)
Query: 29 VQPRTPLRQLLRVASVASGIQFGWALQLSLLTPYVQQLGIPHKWASIIWLCGPVSGLFVQ 88
V+ R PLRQLLRVASVA GIQFGWALQLSLLTPYVQ+LGIPH WAS+IWLCGP+SGL VQ
Sbjct: 17 VRTRVPLRQLLRVASVACGIQFGWALQLSLLTPYVQELGIPHAWASVIWLCGPLSGLVVQ 76
Query: 89 PLVGHLSDRCSSRFGRRRPFILVGAASIVVAVVIIGYAADIGYLIGDDITQNYRPFAIVV 148
PLVGH+SDRC+SR+GRRRPFI+VGAA I V+V+IIG++ADIG+L+GD RP AI V
Sbjct: 77 PLVGHMSDRCTSRYGRRRPFIVVGAACIAVSVLIIGFSADIGWLLGDR-GGGVRPRAIAV 135
Query: 149 FVIGFWILDVANNVTQGPCRALLADLTCNDARRTRVANAYFSLFMAVGNILGYATGSYSG 208
FV GFWILDVANNVTQGPCRALLADLT D RRTRVANAYFSLFMAVGN+LGYATGS S
Sbjct: 136 FVFGFWILDVANNVTQGPCRALLADLTEKDYRRTRVANAYFSLFMAVGNVLGYATGSISY 195
Query: 209 WYKIFTFTLTPACSISCANLKSAFFLDVAFIVVTTYLSIVSAHEVPLSSS---------G 259
+K+F F++TPAC+++CANLKSAFF+D AFI +TT++SI +A PL SS G
Sbjct: 196 LFKVFPFSITPACNVNCANLKSAFFVDTAFIAITTWISISAAQVTPLGSSNRTTPFADEG 255
Query: 260 AGESGSAEEAFMWELFGTFKYFSMPVWIVLSVTALTWIGWFPFNLFDTDWMGREIYGGDP 319
G+S EEAF+WELFGTF+YF VW++L V AL WIGWFPF LFDTDWMGREIYGG P
Sbjct: 256 PGQSSHIEEAFLWELFGTFRYFPGSVWLILLVIALNWIGWFPFLLFDTDWMGREIYGGKP 315
Query: 320 EGGLIYDTGVRMGALGLLLNSVVLAVTSLLMERLCRKRGAGFVWGISNIFMAICFIAMLV 379
G+ Y TGVRMGALGL+LNSV+L +TS+LME+LCRK GAGFVWGIS+I M +CF AMLV
Sbjct: 316 NEGINYSTGVRMGALGLMLNSVILGITSVLMEKLCRKWGAGFVWGISSILMTLCFFAMLV 375
Query: 380 LTYAANSIGYVSKGQPPPTGIVIAALAIFTILGFPMAITYSVPYALISTHIEPLGLGQGL 439
+T+ SIG V PP GIVIAAL +F +LG P+AITYSVPYAL+S+ IE LGLGQGL
Sbjct: 376 ITFVNKSIG-VRGHDLPPDGIVIAALVVFAVLGIPLAITYSVPYALVSSRIESLGLGQGL 434
Query: 440 SMGVLNLAIVVPQIVVSLGSGPWDQLFGGGNSPAFAVAAVAALLSGLLALLAIPRTRTQK 499
SMGVLNLAIV+PQ++VSLGSGPWDQLFGGGN PAFAVAAVA+L SGL+A+LAIPR+ K
Sbjct: 435 SMGVLNLAIVIPQVIVSLGSGPWDQLFGGGNVPAFAVAAVASLASGLVAILAIPRSAAPK 494
Query: 500 PR 501
PR
Sbjct: 495 PR 496
>UniRef100_Q9SQK6 Putative sucrose transporter [Vitis vinifera]
Length = 501
Score = 701 bits (1810), Expect = 0.0
Identities = 345/484 (71%), Positives = 404/484 (83%), Gaps = 11/484 (2%)
Query: 27 QPVQPRTPLRQLLRVASVASGIQFGWALQLSLLTPYVQQLGIPHKWASIIWLCGPVSGLF 86
+PV+PR PLR+LLRVASVA GIQFGWALQLSLLTPYVQ+LGIPH W+SIIWLCGP+SGL
Sbjct: 19 EPVRPRVPLRRLLRVASVACGIQFGWALQLSLLTPYVQELGIPHAWSSIIWLCGPLSGLL 78
Query: 87 VQPLVGHLSDRCSSRFGRRRPFILVGAASIVVAVVIIGYAADIGYLIGDDITQNYRPFAI 146
VQPLVGHLSDRC+SRFGRRRPFI+ GA SIVVAV+IIG++ADIG L+GD + RP A+
Sbjct: 79 VQPLVGHLSDRCNSRFGRRRPFIVAGATSIVVAVLIIGFSADIGGLLGDGADR--RPRAV 136
Query: 147 VVFVIGFWILDVANNVTQGPCRALLADLTCNDARRTRVANAYFSLFMAVGNILGYATGSY 206
FV+GFW+LDVANNVTQGPCRALLADLT D RRTRVANAYFSLF+AVGN+LG+ATGSY
Sbjct: 137 ATFVVGFWLLDVANNVTQGPCRALLADLTEKDHRRTRVANAYFSLFIAVGNVLGFATGSY 196
Query: 207 SGWYKIFTFTLTPACSISCANLKSAFFLDVAFIVVTTYLSIVSAHEVPLSSSG------- 259
SGW++IF FT T +C+ CANLKSAF LD+ FI +TTY+SI +A E+PLSSS
Sbjct: 197 SGWFRIFWFTSTSSCNADCANLKSAFLLDIIFIAITTYISITAAQELPLSSSSRSTHISE 256
Query: 260 -AGESGSAEEAFMWELFGTFKYFSMPVWIVLSVTALTWIGWFPFNLFDTDWMGREIYGGD 318
ES A+EAF+WELFGT +Y S +WI+L VTALTWIG PF LFDTDWMGREIYGG
Sbjct: 257 EMAESTHAQEAFLWELFGTLRYLSGSIWIILFVTALTWIGLLPFLLFDTDWMGREIYGGK 316
Query: 319 PEGGLIYDTGVRMGALGLLLNSVVLAVTSLLMERLCRKRGAGFVWGISNIFMAICFIAML 378
P G Y+TGVRMGALGL+LNSVVL +TS+LME+LCRK GAGFVWG+SNI M++CF+ ML
Sbjct: 317 PNEGQNYNTGVRMGALGLMLNSVVLGITSVLMEKLCRKWGAGFVWGLSNILMSLCFLLML 376
Query: 379 VLTYAANSIGYVSKGQPPPTGIVIAALAIFTILGFPMAITYSVPYALISTHIEPLGLGQG 438
+L+ + ++ PP+G+VIAAL +F+ILG P+AITYSVPYALIST IE LGLGQG
Sbjct: 377 ILSAVVKHMDFLGH-DLPPSGVVIAALIVFSILGIPLAITYSVPYALISTRIESLGLGQG 435
Query: 439 LSMGVLNLAIVVPQIVVSLGSGPWDQLFGGGNSPAFAVAAVAALLSGLLALLAIPRTRTQ 498
LSMGVLNLAIV+PQ++VSLGSGPWDQLFGGGNSP+ AVAAVAA SGL+A+LAIPR+
Sbjct: 436 LSMGVLNLAIVIPQVIVSLGSGPWDQLFGGGNSPSLAVAAVAAFASGLVAILAIPRSSAD 495
Query: 499 KPRV 502
K RV
Sbjct: 496 KSRV 499
>UniRef100_Q9M3R4 Sucrose transporter [Arabidopsis thaliana]
Length = 510
Score = 700 bits (1806), Expect = 0.0
Identities = 340/486 (69%), Positives = 403/486 (81%), Gaps = 6/486 (1%)
Query: 19 STSTSRPV-QPVQPRTPLRQLLRVASVASGIQFGWALQLSLLTPYVQQLGIPHKWASIIW 77
STS+SRPV P + + R LLRVASVA GIQFGWALQLSLLTPYVQ+LGIPH WAS+IW
Sbjct: 23 STSSSRPVVSPPRSKVSKRVLLRVASVACGIQFGWALQLSLLTPYVQELGIPHAWASVIW 82
Query: 78 LCGPVSGLFVQPLVGHLSDRCSSRFGRRRPFILVGAASIVVAVVIIGYAADIGYLIGDDI 137
LCGP+SGLFVQPLVGH SDRC+S++GRRRPFI+ GA +I ++V++IG+AADIG+ GD
Sbjct: 83 LCGPLSGLFVQPLVGHSSDRCTSKYGRRRPFIVAGAVAISISVMVIGHAADIGWAFGDR- 141
Query: 138 TQNYRPFAIVVFVIGFWILDVANNVTQGPCRALLADLTCNDARRTRVANAYFSLFMAVGN 197
+P AIV FV+GFWILDVANN+TQGPCRALLADLT ND RRTRVAN YFSLFMAVGN
Sbjct: 142 EGKIKPRAIVAFVLGFWILDVANNMTQGPCRALLADLTENDNRRTRVANGYFSLFMAVGN 201
Query: 198 ILGYATGSYSGWYKIFTFTLTPACSISCANLKSAFFLDVAFIVVTTYLSIVSAHEVPLSS 257
+LGYATGSY+GWYKIFTFT T AC++ CANLKSAF++DV FI +TT LS+ +AHEVPL+S
Sbjct: 202 VLGYATGSYNGWYKIFTFTKTVACNVECANLKSAFYIDVVFIAITTILSVSAAHEVPLAS 261
Query: 258 SGA---GESGSAEEAFMWELFGTFKYFSMPVWIVLSVTALTWIGWFPFNLFDTDWMGREI 314
+ G++ +EAF+ E+FGTF+YF VWI+L VTALTWIGWFPF LFDTDWMGREI
Sbjct: 262 LASEAHGQTSGTDEAFLSEIFGTFRYFPGNVWIILLVTALTWIGWFPFILFDTDWMGREI 321
Query: 315 YGGDPEGGLIYDTGVRMGALGLLLNSVVLAVTSLLMERLCRKRGAGFVWGISNIFMAICF 374
YGG+P G Y GV MGALGL+LNSV L +TS+LME+LCRK GAGFVWGISNI MAICF
Sbjct: 322 YGGEPNIGTSYSAGVSMGALGLMLNSVFLGITSVLMEKLCRKWGAGFVWGISNILMAICF 381
Query: 375 IAMLVLTYAANSIGYVSKGQPPPTGIVIAALAIFTILGFPMAITYSVPYALISTHIEPLG 434
+ M++ ++ A+ +GY+ Q PP IV AA+ IFTILG P+AITYSVPYALIS IE LG
Sbjct: 382 LGMIITSFVASHLGYIGHEQ-PPASIVFAAVLIFTILGIPLAITYSVPYALISIRIESLG 440
Query: 435 LGQGLSMGVLNLAIVVPQIVVSLGSGPWDQLFGGGNSPAFAVAAVAALLSGLLALLAIPR 494
LGQGLS+GVLNLAIV+PQ++VS+GSGPWDQLFGGGNSPA AV A + G++A+LA+PR
Sbjct: 441 LGQGLSLGVLNLAIVIPQVIVSVGSGPWDQLFGGGNSPALAVGAATGFIGGIVAILALPR 500
Query: 495 TRTQKP 500
TR QKP
Sbjct: 501 TRIQKP 506
>UniRef100_Q9FVJ6 Sucrose transporter [Lycopersicon esculentum]
Length = 500
Score = 698 bits (1801), Expect = 0.0
Identities = 347/494 (70%), Positives = 409/494 (82%), Gaps = 15/494 (3%)
Query: 20 TSTSRPV--QPVQPRTPLRQLLRVASVASGIQFGWALQLSLLTPYVQQLGIPHKWASIIW 77
T +RP +PV+PR PLR LLRVASVA GIQFGWALQLSLLTPYVQ+LGIPH WASIIW
Sbjct: 9 TRHNRPAIREPVKPRVPLRLLLRVASVAGGIQFGWALQLSLLTPYVQELGIPHAWASIIW 68
Query: 78 LCGPVSGLFVQPLVGHLSDRCSSRFGRRRPFILVGAASIVVAVVIIGYAADIGYLIGDDI 137
LCGP+SGL VQPLVGH+SD+C+SRFGRRRPFI+ GAASI++AV+IIG++ADIG+L+GD
Sbjct: 69 LCGPLSGLLVQPLVGHMSDKCTSRFGRRRPFIVAGAASIMIAVLIIGFSADIGWLLGDRG 128
Query: 138 TQNYRPFAIVVFVIGFWILDVANNVTQGPCRALLADLTCNDARRTRVANAYFSLFMAVGN 197
R AI FV+GFW+LDVANN+TQGPCRALLADLT D RRTRVANAYFSLFMA+GN
Sbjct: 129 EIKVR--AIAAFVVGFWLLDVANNMTQGPCRALLADLTQKDHRRTRVANAYFSLFMAIGN 186
Query: 198 ILGYATGSYSGWYKIFTFTLTPACSISCANLKSAFFLDVAFIVVTTYLSIVSAHEVPL-- 255
ILG+ATGSYSGWYKIF FTL AC+I+CANLK+AF LD+ FI TT +SI +A+E PL
Sbjct: 187 ILGFATGSYSGWYKIFLFTLNTACTINCANLKAAFILDIIFIATTTCISISAANEQPLDP 246
Query: 256 --SSSGAGE-----SGSAEEAFMWELFGTFKYFSMPVWIVLSVTALTWIGWFPFNLFDTD 308
SS GE S EEAF+WELFG FKYF VW++L VTALTWIGWFPF LFDTD
Sbjct: 247 SRGSSHTGEEIDESSHGQEEAFLWELFGIFKYFPGVVWVILLVTALTWIGWFPFLLFDTD 306
Query: 309 WMGREIYGGDPEGGLIYDTGVRMGALGLLLNSVVLAVTSLLMERLCRKRGAGFVWGISNI 368
W GREIYGG+P G Y GVRMG+LGL+LNSV+L +TSL ME+LCRK GAGF WG+SN+
Sbjct: 307 WFGREIYGGEPNDGKNYSAGVRMGSLGLMLNSVLLGLTSLFMEKLCRKWGAGFTWGVSNV 366
Query: 369 FMAICFIAMLVLTYAANSIGYVSKGQPPPTGIVIAALAIFTILGFPMAITYSVPYALIST 428
M++CFIAML++T ++I + +G PP GIVIAAL +F+ILG P+AITYSVPYAL+S+
Sbjct: 367 VMSLCFIAMLIITAVRSNID-IGQGL-PPDGIVIAALVVFSILGIPLAITYSVPYALVSS 424
Query: 429 HIEPLGLGQGLSMGVLNLAIVVPQIVVSLGSGPWDQLFGGGNSPAFAVAAVAALLSGLLA 488
IE LGLGQGLSMGVLNLAIV PQIVVSLGSGPWD+LFGGGNSPAF VAA++A +GL+A
Sbjct: 425 RIEALGLGQGLSMGVLNLAIVFPQIVVSLGSGPWDELFGGGNSPAFVVAALSAFAAGLIA 484
Query: 489 LLAIPRTRTQKPRV 502
+LAIPRTR ++P++
Sbjct: 485 ILAIPRTRVERPKI 498
>UniRef100_Q9FE59 Sucrose transporter SUT4 [Arabidopsis thaliana]
Length = 510
Score = 697 bits (1800), Expect = 0.0
Identities = 341/487 (70%), Positives = 403/487 (82%), Gaps = 8/487 (1%)
Query: 19 STSTSRPVQPVQPRTPL--RQLLRVASVASGIQFGWALQLSLLTPYVQQLGIPHKWASII 76
S S+SRPV P PR+ + R LLRVASVA GIQFGWALQLSLLTPYVQ+LGIPH WAS+I
Sbjct: 23 SNSSSRPVVP-PPRSKVSKRVLLRVASVACGIQFGWALQLSLLTPYVQELGIPHAWASVI 81
Query: 77 WLCGPVSGLFVQPLVGHLSDRCSSRFGRRRPFILVGAASIVVAVVIIGYAADIGYLIGDD 136
WLCGP+SGLFVQPLVGH SDRC+S++GRRRPFI+ GA +I ++V++IG+AADIG+ GD
Sbjct: 82 WLCGPLSGLFVQPLVGHSSDRCTSKYGRRRPFIVAGAVAISISVMVIGHAADIGWAFGDR 141
Query: 137 ITQNYRPFAIVVFVIGFWILDVANNVTQGPCRALLADLTCNDARRTRVANAYFSLFMAVG 196
+P AIV FV+GFWILDVANN+TQGPCRALLADLT ND RRTRVAN YFSLFMAVG
Sbjct: 142 -EGKIKPRAIVAFVLGFWILDVANNMTQGPCRALLADLTENDNRRTRVANGYFSLFMAVG 200
Query: 197 NILGYATGSYSGWYKIFTFTLTPACSISCANLKSAFFLDVAFIVVTTYLSIVSAHEVPLS 256
N+LGYATGSY+GWYKIFTFT T AC++ CANLKSAF++DV FI +TT LS+ +AHEVPL+
Sbjct: 201 NVLGYATGSYNGWYKIFTFTKTVACNVECANLKSAFYIDVVFIAITTILSVSAAHEVPLA 260
Query: 257 S---SGAGESGSAEEAFMWELFGTFKYFSMPVWIVLSVTALTWIGWFPFNLFDTDWMGRE 313
S G++ +EAF+ E+FGTF+YF VWI+L VTALTWIGWFPF LFDTDWMGRE
Sbjct: 261 SLTSEAHGQTSGTDEAFLSEIFGTFRYFPGNVWIILLVTALTWIGWFPFILFDTDWMGRE 320
Query: 314 IYGGDPEGGLIYDTGVRMGALGLLLNSVVLAVTSLLMERLCRKRGAGFVWGISNIFMAIC 373
IYGG+P G Y GV MGALGL+LNSV L +TS+LME+LCRK GAGFVWGISNI MAIC
Sbjct: 321 IYGGEPNIGTSYSAGVSMGALGLMLNSVFLGITSVLMEKLCRKWGAGFVWGISNILMAIC 380
Query: 374 FIAMLVLTYAANSIGYVSKGQPPPTGIVIAALAIFTILGFPMAITYSVPYALISTHIEPL 433
F+ M++ ++ A+ +GY+ Q PP IV AA+ IFTILG P+AITYSVPYALIS IE L
Sbjct: 381 FLGMIITSFVASHLGYIGHEQ-PPASIVFAAVLIFTILGIPLAITYSVPYALISIRIESL 439
Query: 434 GLGQGLSMGVLNLAIVVPQIVVSLGSGPWDQLFGGGNSPAFAVAAVAALLSGLLALLAIP 493
GLGQGLS+GVLNLAIV+PQ++VS+GSGPWDQLFGGGNSPA AV A + G++A+LA+P
Sbjct: 440 GLGQGLSLGVLNLAIVIPQVIVSVGSGPWDQLFGGGNSPALAVGAATGFIGGIVAILALP 499
Query: 494 RTRTQKP 500
RTR QKP
Sbjct: 500 RTRIQKP 506
>UniRef100_O65929 Sucrose/H+ symporter [Daucus carota]
Length = 501
Score = 686 bits (1771), Expect = 0.0
Identities = 330/484 (68%), Positives = 398/484 (82%), Gaps = 13/484 (2%)
Query: 28 PVQPRTPLRQLLRVASVASGIQFGWALQLSLLTPYVQQLGIPHKWASIIWLCGPVSGLFV 87
P + R LR LLRVASVA GIQFGWALQLSLLTPYVQ+LGIPH W+SIIWLCGP+SGL V
Sbjct: 20 PPRSRVSLRLLLRVASVACGIQFGWALQLSLLTPYVQELGIPHAWSSIIWLCGPLSGLLV 79
Query: 88 QPLVGHLSDRCSSRFGRRRPFILVGAASIVVAVVIIGYAADIGYLIGDDITQNYRPFAIV 147
QP+VGH+SD+C+S++GRRRPFI+ G +I++AV+II ++ADIG L+GD T + + AIV
Sbjct: 80 QPIVGHMSDQCTSKYGRRRPFIVAGGTAIILAVIIIAHSADIGGLLGD--TADNKTMAIV 137
Query: 148 VFVIGFWILDVANNVTQGPCRALLADLTCNDARRTRVANAYFSLFMAVGNILGYATGSYS 207
FVIGFWILDVANN+TQGPCRALLADLT NDARRTRVANAYFSLFMA+GN+LGYATG+YS
Sbjct: 138 AFVIGFWILDVANNMTQGPCRALLADLTGNDARRTRVANAYFSLFMAIGNVLGYATGAYS 197
Query: 208 GWYKIFTFTLTPACSISCANLKSAFFLDVAFIVVTTYLSIVSAHEVP---------LSSS 258
GWYK+F F+LT +C+I+CANLKSAF++D+ FI++TTY+SI +A E P S
Sbjct: 198 GWYKVFPFSLTSSCTINCANLKSAFYIDIIFIIITTYISISAAKERPRISSQDGPQFSED 257
Query: 259 GAGESGSAEEAFMWELFGTFKYFSMPVWIVLSVTALTWIGWFPFNLFDTDWMGREIYGGD 318
G +SG EEAF+WELFGTF+ VW++L VT L WIGWFPF LFDTDWMGREIYGG+
Sbjct: 258 GTAQSGHIEEAFLWELFGTFRLLPGSVWVILLVTCLNWIGWFPFILFDTDWMGREIYGGE 317
Query: 319 PEGGLIYDTGVRMGALGLLLNSVVLAVTSLLMERLCRKRGAGFVWGISNIFMAICFIAML 378
P G Y GVRMGA GL++NSVVL +TS+LME+LCR G+GF+WG+SNI M ICF AML
Sbjct: 318 PNQGQSYSDGVRMGAFGLMMNSVVLGITSVLMEKLCRIWGSGFMWGLSNILMTICFFAML 377
Query: 379 VLTYAANSIGYVSKGQPPPTGIVIAALAIFTILGFPMAITYSVPYALISTHIEPLGLGQG 438
++T+ A ++ Y + PPP GIVI+AL +F ILG P+AITYSVPYAL+ST IE LGLGQG
Sbjct: 378 LITFIAKNMDYGT--NPPPNGIVISALIVFAILGIPLAITYSVPYALVSTRIESLGLGQG 435
Query: 439 LSMGVLNLAIVVPQIVVSLGSGPWDQLFGGGNSPAFAVAAVAALLSGLLALLAIPRTRTQ 498
LSMGVLNLAIVVPQ++VSLGSGPWDQLFGGGNSPAF VAA++A +GL+AL+AI R R
Sbjct: 436 LSMGVLNLAIVVPQVIVSLGSGPWDQLFGGGNSPAFVVAALSAFAAGLIALIAIRRPRVD 495
Query: 499 KPRV 502
K R+
Sbjct: 496 KSRL 499
>UniRef100_Q9FV92 Sucrose transporter SUT4 [Solanum tuberosum]
Length = 488
Score = 686 bits (1771), Expect = 0.0
Identities = 341/491 (69%), Positives = 402/491 (81%), Gaps = 15/491 (3%)
Query: 20 TSTSRPV--QPVQPRTPLRQLLRVASVASGIQFGWALQLSLLTPYVQQLGIPHKWASIIW 77
T +RP +PV+PR PLR L RVASVA GIQFGWALQLSLLTPYVQ+LGIPH WASIIW
Sbjct: 2 TRHNRPAIREPVKPRVPLRLLFRVASVAGGIQFGWALQLSLLTPYVQELGIPHAWASIIW 61
Query: 78 LCGPVSGLFVQPLVGHLSDRCSSRFGRRRPFILVGAASIVVAVVIIGYAADIGYLIGDDI 137
LCGP+SGL VQPLVGH+SD+C+SRFGRRRPFI+ GA SI++AV+IIG++ADIG+L+GD
Sbjct: 62 LCGPLSGLLVQPLVGHMSDKCTSRFGRRRPFIVAGAVSIMIAVLIIGFSADIGWLLGDRG 121
Query: 138 TQNYRPFAIVVFVIGFWILDVANNVTQGPCRALLADLTCNDARRTRVANAYFSLFMAVGN 197
R AI FV+GFW+LDVANN+TQGPCRALLADLT D RRTRVANAYFSLFMA+GN
Sbjct: 122 EIKVR--AIAAFVVGFWLLDVANNMTQGPCRALLADLTQKDHRRTRVANAYFSLFMAIGN 179
Query: 198 ILGYATGSYSGWYKIFTFTLTPACSISCANLKSAFFLDVAFIVVTTYLSIVSAHEVPLSS 257
ILG+ATGSYSGW+KIF FTL AC+I+CANLK+AF +D+ FI TT +SI +A+E PL
Sbjct: 180 ILGFATGSYSGWFKIFPFTLNTACTINCANLKAAFIIDIIFIATTTCISISAANEQPLDP 239
Query: 258 SGA--------GESG-SAEEAFMWELFGTFKYFSMPVWIVLSVTALTWIGWFPFNLFDTD 308
S GES EEAF+WELFG FKYF VW++L VTALTWIGWFPF LFDTD
Sbjct: 240 SRGSSHTREEIGESSHGQEEAFLWELFGIFKYFPGVVWVILLVTALTWIGWFPFLLFDTD 299
Query: 309 WMGREIYGGDPEGGLIYDTGVRMGALGLLLNSVVLAVTSLLMERLCRKRGAGFVWGISNI 368
W GREIYGG+P G Y GVRMG+LGL+LNSV+L +TSL ME+LCRK GAGF WG+SN+
Sbjct: 300 WFGREIYGGEPNDGKNYSAGVRMGSLGLMLNSVLLGLTSLFMEKLCRKWGAGFTWGVSNV 359
Query: 369 FMAICFIAMLVLTYAANSIGYVSKGQPPPTGIVIAALAIFTILGFPMAITYSVPYALIST 428
M++CFIAML++T ++I + +G PP GIVIAAL +F+ILG P+AITYSVPYAL+S+
Sbjct: 360 VMSLCFIAMLIITAVRSNID-IGQGL-PPDGIVIAALVVFSILGIPLAITYSVPYALVSS 417
Query: 429 HIEPLGLGQGLSMGVLNLAIVVPQIVVSLGSGPWDQLFGGGNSPAFAVAAVAALLSGLLA 488
I+ LGLGQGLSMGVLNLAIV PQIVVSLGSGPWD+LFGGGNSPAF VAA++A GL+A
Sbjct: 418 RIDALGLGQGLSMGVLNLAIVFPQIVVSLGSGPWDELFGGGNSPAFVVAALSAFAGGLIA 477
Query: 489 LLAIPRTRTQK 499
+LAIPRTR +K
Sbjct: 478 ILAIPRTRVEK 488
>UniRef100_Q6J2T0 Sucrose transporter SUT4 [Zea mays]
Length = 501
Score = 646 bits (1667), Expect = 0.0
Identities = 314/484 (64%), Positives = 388/484 (79%), Gaps = 13/484 (2%)
Query: 21 STSRPVQPVQPRTPLRQLLRVASVASGIQFGWALQLSLLTPYVQQLGIPHKWASIIWLCG 80
+TS P + V PLR+LLR ASVA G+QFGWALQLSLLTPYVQ+LGIPH +AS++WLCG
Sbjct: 9 ATSTPPRKV----PLRKLLRAASVACGVQFGWALQLSLLTPYVQELGIPHAFASLVWLCG 64
Query: 81 PVSGLFVQPLVGHLSDR---CSSRFGRRRPFILVGAASIVVAVVIIGYAADIGYLIGDDI 137
P+SGL VQPLVGHLSDR +S GRRRPFI GAA I AV+ +G++AD+G L GDD+
Sbjct: 65 PLSGLLVQPLVGHLSDRIGPAASPLGRRRPFIAAGAACIAAAVLTVGFSADLGRLFGDDV 124
Query: 138 TQ-NYRPFAIVVFVIGFWILDVANNVTQGPCRALLADLTCNDARRTRVANAYFSLFMAVG 196
T + R AI V+++GFW+LDV NN TQGPCRA LADLT ND RRTR+ANAYFSLFMA+G
Sbjct: 125 TPGSTRLGAICVYLVGFWLLDVGNNGTQGPCRAFLADLTENDPRRTRIANAYFSLFMALG 184
Query: 197 NILGYATGSYSGWYKIFTFTLTPACSISCANLKSAFFLDVAFIVVTTYLSIVSAHE-VPL 255
NILGYATG+YSGWY IF FT+T +C ISCANLKSAF LD+ +V+TTY ++ S E
Sbjct: 185 NILGYATGAYSGWYSIFPFTVTESCGISCANLKSAFLLDIIVLVITTYTTVTSVQEPQTF 244
Query: 256 SSSGAGESGSAEEAFMWELFGTFKYFSMPVWIVLSVTALTWIGWFPFNLFDTDWMGREIY 315
S A SG+ +EAF+WELFG+ +YF++P+W+VL VTALTW+ WFPF LFDTDWMGREIY
Sbjct: 245 GSDEAQNSGAEQEAFLWELFGSLRYFTLPIWMVLIVTALTWMAWFPFTLFDTDWMGREIY 304
Query: 316 GGDPEG---GLIYDTGVRMGALGLLLNSVVLAVTSLLMERLCRKRGAGFVWGISNIFMAI 372
G P+ Y GVRMG+ GL+LNSVVL TS+++E+LCRK GAG VWG+SNI M +
Sbjct: 305 RGSPDNPGETQRYHDGVRMGSFGLMLNSVVLGFTSVVLEKLCRKWGAGLVWGVSNILMTL 364
Query: 373 CFIAMLVLTYAANSIGYVSKGQPPPTGIVIAALAIFTILGFPMAITYSVPYALISTHIEP 432
CF+AMLV+TY A ++ Y S G PPTGIV+A+L +FTILG P+AITYS+PYA+ ++ +E
Sbjct: 365 CFLAMLVITYVAKNMDYPSSG-APPTGIVVASLVVFTILGAPLAITYSIPYAMAASRVEN 423
Query: 433 LGLGQGLSMGVLNLAIVVPQIVVSLGSGPWDQLFGGGNSPAFAVAAVAALLSGLLALLAI 492
LGLGQGL+MG+LNLAIV+PQ++VSLGSGPWDQLFGGGN+PAFAVAA A+ + GL+A+L +
Sbjct: 424 LGLGQGLAMGILNLAIVIPQVIVSLGSGPWDQLFGGGNAPAFAVAAGASFIGGLVAILGL 483
Query: 493 PRTR 496
PR R
Sbjct: 484 PRAR 487
>UniRef100_Q84KR5 Sucrose transporter [Oryza sativa]
Length = 501
Score = 645 bits (1665), Expect = 0.0
Identities = 315/495 (63%), Positives = 388/495 (77%), Gaps = 9/495 (1%)
Query: 15 RSSTSTSTSRPVQPVQPRTPLRQLLRVASVASGIQFGWALQLSLLTPYVQQLGIPHKWAS 74
R S + P + PLR+LLR ASVA G+QFGWA QLSLLTPYVQ+LGIPH +AS
Sbjct: 4 RPSGGGGGAGPAAAAVRKVPLRKLLRAASVACGVQFGWAPQLSLLTPYVQELGIPHAFAS 63
Query: 75 IIWLCGPVSGLFVQPLVGHLSDR---CSSRFGRRRPFILVGAASIVVAVVIIGYAADIGY 131
++WLCGP+SGL VQPLVGHLSDR +S GRRRPFI GAASI AV+ + ++AD+G
Sbjct: 64 LVWLCGPLSGLLVQPLVGHLSDRIAPAASPLGRRRPFIAAGAASIAAAVLTVRFSADLGR 123
Query: 132 LIGDDITQ-NYRPFAIVVFVIGFWILDVANNVTQGPCRALLADLTCNDARRTRVANAYFS 190
+ GD IT + R AI +++GFW+LDV NN TQGPCRA ADLT ND +RTR+ANAYFS
Sbjct: 124 IFGDSITPGSTRLGAITAYLVGFWLLDVGNNATQGPCRAFPADLTENDPKRTRIANAYFS 183
Query: 191 LFMAVGNILGYATGSYSGWYKIFTFTLTPACSISCANLKSAFFLDVAFIVVTTYLSIVSA 250
LFMA+GNILGYATG+YSGWYKIF FT+TP+CSISCANLKSAF LD+ +VVTT +++ S
Sbjct: 184 LFMALGNILGYATGAYSGWYKIFPFTVTPSCSISCANLKSAFLLDIIILVVTTCITVASV 243
Query: 251 HE-VPLSSSGAGESGSAEEAFMWELFGTFKYFSMPVWIVLSVTALTWIGWFPFNLFDTDW 309
E S A + +EAF+WELFG+F+YF++PVW+VL VTALTWIGWFPF LFDTDW
Sbjct: 244 QEPQSFGSDEADHPSTEQEAFLWELFGSFRYFTLPVWMVLIVTALTWIGWFPFILFDTDW 303
Query: 310 MGREIYGGDPEGGLI---YDTGVRMGALGLLLNSVVLAVTSLLMERLCRKRGAGFVWGIS 366
MGREIY G P+ I Y GVRMG+ GL+LNSV+L TS+++E+LCRK GAG VWG+S
Sbjct: 304 MGREIYRGSPDDPSITQSYHDGVRMGSFGLMLNSVLLGFTSIVLEKLCRKWGAGLVWGVS 363
Query: 367 NIFMAICFIAMLVLTYAANSIGYVSKGQPPPTGIVIAALAIFTILGFPMAITYSVPYALI 426
NI MA+CF+AMLV+TY A ++ Y G PPTGIVIA+L +FTILG P+AITYS+PYA+
Sbjct: 364 NILMALCFVAMLVITYVAKNMDYPPSG-VPPTGIVIASLVVFTILGAPLAITYSIPYAMA 422
Query: 427 STHIEPLGLGQGLSMGVLNLAIVVPQIVVSLGSGPWDQLFGGGNSPAFAVAAVAALLSGL 486
++ +E LGLGQGL+MG+LNLAIV+PQ++VSLGSGPWDQLFGGGN+PAFAVAA A+ + GL
Sbjct: 423 ASRVENLGLGQGLAMGILNLAIVIPQVIVSLGSGPWDQLFGGGNAPAFAVAAAASFIGGL 482
Query: 487 LALLAIPRTRTQKPR 501
+A+L +PR R R
Sbjct: 483 VAILGLPRARIASRR 497
>UniRef100_Q9M423 Sucrose transporter 2 [Hordeum vulgare]
Length = 506
Score = 643 bits (1659), Expect = 0.0
Identities = 314/499 (62%), Positives = 392/499 (77%), Gaps = 16/499 (3%)
Query: 9 PHRSRTRSSTSTSTSRPVQPVQPRTPLRQLLRVASVASGIQFGWALQLSLLTPYVQQLGI 68
P R T TS+S P + PLR LLR ASVA G+QFGWALQLSLLTPYVQ+LGI
Sbjct: 3 PRRPNTGGGGGTSSS--AAPAPRKVPLRSLLRAASVACGVQFGWALQLSLLTPYVQELGI 60
Query: 69 PHKWASIIWLCGPVSGLFVQPLVGHLSDR---CSSRFGRRRPFILVGAASIVVAVVIIGY 125
PH +AS++WLCGP+SGL VQPLVGHLSDR +S GRRRPFI GAASI AV+ +G+
Sbjct: 61 PHAFASLVWLCGPLSGLLVQPLVGHLSDRITPANSPLGRRRPFIAAGAASIAFAVLTVGF 120
Query: 126 AADIGYLIGDDITQ-NYRPFAIVVFVIGFWILDVANNVTQGPCRALLADLTCNDARRTRV 184
+AD+G L GD++ + R AI+V+++GFW+LDV NN TQGPCRA LADLT ND RRTR+
Sbjct: 121 SADLGRLFGDNVVPGSTRIGAIIVYLVGFWLLDVGNNATQGPCRAFLADLTENDPRRTRI 180
Query: 185 ANAYFSLFMAVGNILGYATGSYSGWYKIFTFTLTPACSISCANLKSAFFLDVAFIVVTTY 244
ANAYFSLFMA+GNILGYATG+Y+GWYKIF FT+T +C +SCANL SAF LD+ + +TTY
Sbjct: 181 ANAYFSLFMALGNILGYATGAYNGWYKIFPFTITGSCGVSCANLNSAFLLDIIILAITTY 240
Query: 245 LSIVSAHEVPL--SSSGAGESGSAEEAFMWELFGTFKYFSMPVWIVLSVTALTWIGWFPF 302
+S+ + + P S A S EEAF++ELFG+FKYF+MPVW+VL VT+LTW+GWF F
Sbjct: 241 ISVATVQDNPTFGSDEAAPPSSHEEEAFLFELFGSFKYFTMPVWMVLIVTSLTWVGWFLF 300
Query: 303 NLFDTDWMGREIYGGDPEGGLIYDT-----GVRMGALGLLLNSVVLAVTSLLMERLCRKR 357
LFDTDWMGREIY G PE ++ DT GVRMG+ GL+LNSVVL +TS+ ME+LCRK
Sbjct: 301 ILFDTDWMGREIYRGSPE--IVADTQKYHDGVRMGSFGLMLNSVVLGITSIGMEKLCRKW 358
Query: 358 GAGFVWGISNIFMAICFIAMLVLTYAANSIGYVSKGQPPPTGIVIAALAIFTILGFPMAI 417
GAG VWG+SNI MA+CF+AML++TY A ++ Y G PPTGIV A+L +FTILG P++I
Sbjct: 359 GAGLVWGVSNIIMALCFVAMLIITYVAQNLDYGPSG-APPTGIVAASLIVFTILGAPLSI 417
Query: 418 TYSVPYALISTHIEPLGLGQGLSMGVLNLAIVVPQIVVSLGSGPWDQLFGGGNSPAFAVA 477
TYS+PYA+ ++ +E LGLGQGL+MG+LNL+IV+PQI+VSLGSGPWDQLFGGGN+P+F VA
Sbjct: 418 TYSIPYAMAASRVENLGLGQGLAMGILNLSIVIPQIIVSLGSGPWDQLFGGGNAPSFWVA 477
Query: 478 AVAALLSGLLALLAIPRTR 496
A A+ + GL+A+L +PR R
Sbjct: 478 AAASFVGGLVAILGLPRAR 496
>UniRef100_Q6GUI0 Sucrose transport protein [Zea mays]
Length = 501
Score = 642 bits (1655), Expect = 0.0
Identities = 312/484 (64%), Positives = 386/484 (79%), Gaps = 13/484 (2%)
Query: 21 STSRPVQPVQPRTPLRQLLRVASVASGIQFGWALQLSLLTPYVQQLGIPHKWASIIWLCG 80
+TS P + V PLR+LLR ASVA G+QFGWALQLSLLTPYVQ+LGIPH +AS++WLCG
Sbjct: 9 ATSTPPRKV----PLRKLLRAASVACGVQFGWALQLSLLTPYVQELGIPHAFASLVWLCG 64
Query: 81 PVSGLFVQPLVGHLSDR---CSSRFGRRRPFILVGAASIVVAVVIIGYAADIGYLIGDDI 137
P+SGL VQPLVGHLSDR +S GRRRPFI GAA I AV+ +G++AD+G L GDD+
Sbjct: 65 PLSGLLVQPLVGHLSDRIGPAASPLGRRRPFIAAGAACIAAAVLTVGFSADLGRLFGDDV 124
Query: 138 TQ-NYRPFAIVVFVIGFWILDVANNVTQGPCRALLADLTCNDARRTRVANAYFSLFMAVG 196
T + R AI V+++GFW+LDV NN TQGPCRA LADLT ND RRTR+ANAYFSLFMA+G
Sbjct: 125 TPGSTRLGAICVYLVGFWLLDVGNNGTQGPCRAFLADLTENDPRRTRIANAYFSLFMALG 184
Query: 197 NILGYATGSYSGWYKIFTFTLTPACSISCANLKSAFFLDVAFIVVTTYLSIVSAHE-VPL 255
NILGYATG+YSGWY IF FT+T +C ISCANLKSAF LD+ +V+TTY ++ S E
Sbjct: 185 NILGYATGAYSGWYSIFPFTVTESCGISCANLKSAFLLDIIVLVITTYTTVTSVQEPQTF 244
Query: 256 SSSGAGESGSAEEAFMWELFGTFKYFSMPVWIVLSVTALTWIGWFPFNLFDTDWMGREIY 315
S A G+ +EAF+WELFG+ +YF++P+W+VL VTALTW+ WFPF LFDTDWMGREIY
Sbjct: 245 GSDEAQNPGAEQEAFLWELFGSLRYFTLPIWMVLIVTALTWMAWFPFTLFDTDWMGREIY 304
Query: 316 GGDPEG---GLIYDTGVRMGALGLLLNSVVLAVTSLLMERLCRKRGAGFVWGISNIFMAI 372
G P+ Y GVRMG+ GL+LNSVVL TS+++E+LCRK GAG VWG+SNI M +
Sbjct: 305 RGSPDNPGETQRYHDGVRMGSFGLMLNSVVLGFTSVVLEKLCRKWGAGLVWGVSNILMTL 364
Query: 373 CFIAMLVLTYAANSIGYVSKGQPPPTGIVIAALAIFTILGFPMAITYSVPYALISTHIEP 432
CF+AMLV+TY A ++ Y S G PPTGIV+A+L +FTILG P+AITYS+PYA+ ++ +E
Sbjct: 365 CFLAMLVITYVAKNMDYPSSG-APPTGIVVASLVVFTILGAPLAITYSIPYAMAASRVEN 423
Query: 433 LGLGQGLSMGVLNLAIVVPQIVVSLGSGPWDQLFGGGNSPAFAVAAVAALLSGLLALLAI 492
LG GQGL+MG+LNLAIV+PQ++VSLGSGPWDQLFGGGN+PAFAVAA A+ + GL+A+L +
Sbjct: 424 LGPGQGLAMGILNLAIVIPQVIVSLGSGPWDQLFGGGNAPAFAVAAGASFIGGLVAILGL 483
Query: 493 PRTR 496
PR R
Sbjct: 484 PRAR 487
>UniRef100_O04516 F21M12.35 protein [Arabidopsis thaliana]
Length = 474
Score = 623 bits (1606), Expect = e-177
Identities = 309/486 (63%), Positives = 370/486 (75%), Gaps = 42/486 (8%)
Query: 19 STSTSRPV-QPVQPRTPLRQLLRVASVASGIQFGWALQLSLLTPYVQQLGIPHKWASIIW 77
STS+SRPV P + + R LLRVASVA GIQFGWALQLSLLTPYVQ+LGIPH WAS+IW
Sbjct: 23 STSSSRPVVSPPRSKVSKRVLLRVASVACGIQFGWALQLSLLTPYVQELGIPHAWASVIW 82
Query: 78 LCGPVSGLFVQPLVGHLSDRCSSRFGRRRPFILVGAASIVVAVVIIGYAADIGYLIGDDI 137
LCGP+SGLFVQPLVGH SDRC+S++GRRRPFI+ GA +I ++V++IG+AADIG+ GD
Sbjct: 83 LCGPLSGLFVQPLVGHSSDRCTSKYGRRRPFIVAGAVAISISVMVIGHAADIGWAFGDR- 141
Query: 138 TQNYRPFAIVVFVIGFWILDVANNVTQGPCRALLADLTCNDARRTRVANAYFSLFMAVGN 197
+P AIV FV+GFWILDVANN+TQGPCRALLADLT ND RRTRVAN YFSLFMAVGN
Sbjct: 142 EGKIKPRAIVAFVLGFWILDVANNMTQGPCRALLADLTENDNRRTRVANGYFSLFMAVGN 201
Query: 198 ILGYATGSYSGWYKIFTFTLTPACSISCANLKSAFFLDVAFIVVTTYLSIVSAHEVPLSS 257
+LGYATGSY+GWYKIFTFT T AC++ CANLKSAF++DV FI +TT LS+ +AHEVPL+S
Sbjct: 202 VLGYATGSYNGWYKIFTFTKTVACNVECANLKSAFYIDVVFIAITTILSVSAAHEVPLAS 261
Query: 258 SGA---GESGSAEEAFMWELFGTFKYFSMPVWIVLSVTALTWIGWFPFNLFDTDWMGREI 314
+ G++ +EAF+ E+FGTF+YF VWI+L VTALTWIGWFPF LFDTDWMGREI
Sbjct: 262 LASEAHGQTSGTDEAFLSEIFGTFRYFPGNVWIILLVTALTWIGWFPFILFDTDWMGREI 321
Query: 315 YGGDPEGGLIYDTGVRMGALGLLLNSVVLAVTSLLMERLCRKRGAGFVWGISNIFMAICF 374
YGG+P G Y GV MGALGL+LNSV L +TS+LME+LCRK GAGFVWGISNI MAICF
Sbjct: 322 YGGEPNIGTSYSAGVSMGALGLMLNSVFLGITSVLMEKLCRKWGAGFVWGISNILMAICF 381
Query: 375 IAMLVLTYAANSIGYVSKGQPPPTGIVIAALAIFTILGFPMAITYSVPYALISTHIEPLG 434
+ M++ ++ A+ +GY+ Q PP IV AA+ IFTILG P+A
Sbjct: 382 LGMIITSFVASHLGYIGHEQ-PPASIVFAAVLIFTILGIPLA------------------ 422
Query: 435 LGQGLSMGVLNLAIVVPQIVVSLGSGPWDQLFGGGNSPAFAVAAVAALLSGLLALLAIPR 494
++VS+GSGPWDQLFGGGNSPA AV A + G++A+LA+PR
Sbjct: 423 ------------------VIVSVGSGPWDQLFGGGNSPALAVGAATGFIGGIVAILALPR 464
Query: 495 TRTQKP 500
TR QKP
Sbjct: 465 TRIQKP 470
>UniRef100_Q7XA53 Sucrose transporter [Glycine max]
Length = 520
Score = 537 bits (1384), Expect = e-151
Identities = 273/499 (54%), Positives = 347/499 (68%), Gaps = 13/499 (2%)
Query: 5 TTTNPHRSRTRSSTSTSTSRPVQPVQPRTPLRQLLRVASVASGIQFGWALQLSLLTPYVQ 64
T N + + T + S P P++P +PLR+++ VAS+A+G+QFGWALQLSLLTPYVQ
Sbjct: 7 TKQNHNNNNTLTKPSLHVESP--PLEP-SPLRKIIVVASIAAGVQFGWALQLSLLTPYVQ 63
Query: 65 QLGIPHKWASIIWLCGPVSGLFVQPLVGHLSDRCSSRFGRRRPFILVGAASIVVAVVIIG 124
LGIPH WA+ IWLCGP+SG+ VQP+VG+ SDRC+SRFGRRRPFI G+ ++ +AV +IG
Sbjct: 64 LLGIPHTWAAYIWLCGPISGMLVQPIVGYHSDRCTSRFGRRRPFIAAGSFAVAIAVFLIG 123
Query: 125 YAADIGYLIGDDITQNYRPFAIVVFVIGFWILDVANNVTQGPCRALLADLTCNDARRTRV 184
YAAD+G+ GDD+++ RP AI +FV+GFWILDVANN+ QGPCRALL DL + ++TR
Sbjct: 124 YAADLGHSFGDDLSKKVRPRAIGIFVVGFWILDVANNMLQGPCRALLGDLCAGNHQKTRN 183
Query: 185 ANAYFSLFMAVGNILGYATGSYSGWYKIFTFTLTPACSISCANLKSAFFLDVAFIVVTTY 244
ANA+FS FMAVGN+LGYA G+YS Y +F FT T AC + CANLKS FFL +A +
Sbjct: 184 ANAFFSFFMAVGNVLGYAAGAYSKLYHVFPFTKTTACDVYCANLKSCFFLSIALLTTLAT 243
Query: 245 LSIVSAHEVPLSSSGA--GESGSAEEAFMWELFGTFKYFSMPVWIVLSVTALTWIGWFPF 302
++V EVPLS A + +LFG F+ P+WI+L VT L WI WFPF
Sbjct: 244 AALVYVKEVPLSPEKAVIDSDDNGGMPCFGQLFGAFRELKRPMWILLLVTCLNWIAWFPF 303
Query: 303 NLFDTDWMGREIYGGDPEGGLIYDTGVRMGALGLLLNSVVLAVTSLLMERLCR-KRGAGF 361
LFDTDWMGRE+Y G G YD GVR GALGL+LNSVVL TSL +E L R G
Sbjct: 304 LLFDTDWMGREVYEGTVGEGKAYDRGVRAGALGLMLNSVVLGATSLGVEVLARGVGGVKR 363
Query: 362 VWGISNIFMAICF-IAMLVLTYAANSIGYV------SKGQPPPTGIVIAALAIFTILGFP 414
+WGI N +A+C + +LV A +S Y + PPP + ALA+F++LG P
Sbjct: 364 LWGIVNFLLAVCLAMTVLVTKMAQHSRQYTLLPNAHQEPLPPPAAVKAGALALFSLLGIP 423
Query: 415 MAITYSVPYALISTHIEPLGLGQGLSMGVLNLAIVVPQIVVSLGSGPWDQLFGGGNSPAF 474
+AITYS+P+AL S G GQGLS+GVLNLAIV+PQ+VVS+ SGPWD LFGGGN PAF
Sbjct: 424 LAITYSIPFALASIFSSTSGAGQGLSLGVLNLAIVIPQMVVSVISGPWDALFGGGNLPAF 483
Query: 475 AVAAVAALLSGLLALLAIP 493
V AVAA SG+L+++ +P
Sbjct: 484 VVGAVAAAASGILSIILLP 502
>UniRef100_Q41152 Sucrose carrier [Ricinus communis]
Length = 533
Score = 537 bits (1384), Expect = e-151
Identities = 280/525 (53%), Positives = 353/525 (66%), Gaps = 36/525 (6%)
Query: 15 RSSTSTSTSRPV--QPVQP-----------RTPLRQLLRVASVASGIQFGWALQLSLLTP 61
+SSTS +P QP P +PLR+++ VAS+A+GIQFGWALQLSLLTP
Sbjct: 2 QSSTSKENKQPPSSQPHPPPLMVAGAAEPNSSPLRKVVMVASIAAGIQFGWALQLSLLTP 61
Query: 62 YVQQLGIPHKWASIIWLCGPVSGLFVQPLVGHLSDRCSSRFGRRRPFILVGAASIVVAVV 121
YVQ LGIPH WA+ IWLCGP+SG+ VQP+VG+ SDRC+SRFGRRRPFI GAA + +AV
Sbjct: 62 YVQLLGIPHTWAAFIWLCGPISGMLVQPIVGYHSDRCTSRFGRRRPFIASGAAFVAIAVF 121
Query: 122 IIGYAADIGYLIGDDITQNYRPFAIVVFVIGFWILDVANNVTQGPCRALLADLTCNDARR 181
+IGYAAD+G+L GD + ++ + AI +FV+GFWILDVANN+ QGPCRALLADL+ ++
Sbjct: 122 LIGYAADLGHLSGDSLDKSPKTRAIAIFVVGFWILDVANNMLQGPCRALLADLSGTSQKK 181
Query: 182 TRVANAYFSLFMAVGNILGYATGSYSGWYKIFTFTLTPACSISCANLKSAFFLDVAFIVV 241
TR ANA FS FMAVGN+LGYA G+Y+ YK+F FT T AC + CANLKS FF+ + ++
Sbjct: 182 TRTANALFSFFMAVGNVLGYAAGAYTHLYKLFPFTKTTACDVYCANLKSCFFISIVLLLS 241
Query: 242 TTYLSIVSAHEVPLSSSGA---GESGSAEEA--------FMWELFGTFKYFSMPVWIVLS 290
T L++ E P S A E +A +A F E+ G FK P+WI+L
Sbjct: 242 LTVLALSYVKEKPWSPDQAVDNAEDDTASQASSSAQPMPFFGEILGAFKNLKRPMWILLL 301
Query: 291 VTALTWIGWFPFNLFDTDWMGREIYGGDPEGGL----IYDTGVRMGALGLLLNSVVLAVT 346
VT L WI WFPF LFDTDWMGRE+YGGD G +YD GVR GALGL+LNSVVL T
Sbjct: 302 VTCLNWIAWFPFLLFDTDWMGREVYGGDSSGSAEQLKLYDRGVRAGALGLMLNSVVLGFT 361
Query: 347 SLLMERLCR-KRGAGFVWGISNIFMAICFIAMLVLTYAANS---IGYVSKGQ----PPPT 398
SL +E L R G +WGI N +A+C +++T A S VS G PPP+
Sbjct: 362 SLGVEVLARGVGGVKRLWGIVNFVLAVCLAMTVLVTKQAESTRRFATVSGGAKVPLPPPS 421
Query: 399 GIVIAALAIFTILGFPMAITYSVPYALISTHIEPLGLGQGLSMGVLNLAIVVPQIVVSLG 458
G+ ALA+F ++G P AITYS+P+AL S G GQGLS+GVLNL+IV+PQ++VS+
Sbjct: 422 GVKAGALALFAVMGVPQAITYSIPFALASIFSNTSGAGQGLSLGVLNLSIVIPQMIVSVA 481
Query: 459 SGPWDQLFGGGNSPAFAVAAVAALLSGLLALLAIPRTRTQKPRVR 503
+GPWD LFGGGN PAF V AVAAL SG+ AL +P + P +
Sbjct: 482 AGPWDALFGGGNLPAFVVGAVAALASGIFALTMLPSPQPDMPSAK 526
>UniRef100_Q9XHL6 Sucrose transport protein SUT1 [Pisum sativum]
Length = 524
Score = 530 bits (1365), Expect = e-149
Identities = 271/505 (53%), Positives = 348/505 (68%), Gaps = 14/505 (2%)
Query: 2 PNPTTTNPHRSRTRSSTSTSTSRPVQPVQPRTPLRQLLRVASVASGIQFGWALQLSLLTP 61
P +T + + + +S QP++P +PLR+++ VAS+A+G+QFGWALQLSLLTP
Sbjct: 3 PLSSTKQINNNNNNLAKPSSLHVETQPLEP-SPLRKIMVVASIAAGVQFGWALQLSLLTP 61
Query: 62 YVQQLGIPHKWASIIWLCGPVSGLFVQPLVGHLSDRCSSRFGRRRPFILVGAASIVVAVV 121
YVQ LGI H WA+ IWLCGP+SG+ VQP+VG+ SDRC+SRFGRRRPFI G+ ++ +AV
Sbjct: 62 YVQLLGIHHTWAAYIWLCGPISGMLVQPVVGYHSDRCTSRFGRRRPFIAAGSIAVAIAVF 121
Query: 122 IIGYAADIGYLIGDDITQNYRPFAIVVFVIGFWILDVANNVTQGPCRALLADLTCNDARR 181
+IGYAAD+G+ GD++ + RP AI +FV+GFWILDVANN+ QGPCRALL DL + R+
Sbjct: 122 LIGYAADLGHSFGDNLDKKVRPRAIGIFVVGFWILDVANNMLQGPCRALLGDLCAGNQRK 181
Query: 182 TRVANAYFSLFMAVGNILGYATGSYSGWYKIFTFTLTPACSISCANLKSAFFLDVAFIVV 241
TR ANA+FS FMAVGN+LGYA G+YS Y +F FT T AC++ CANLKS FFL +A + V
Sbjct: 182 TRNANAFFSFFMAVGNVLGYAAGAYSKLYHVFPFTKTEACNVYCANLKSCFFLSIALLTV 241
Query: 242 TTYLSIVSAHEVPLSSSGA---GESGSAEEAF--MWELFGTFKYFSMPVWIVLSVTALTW 296
+++ E PL + A E G + +L G FK P+WI+L VT L W
Sbjct: 242 LATAALIYVKETPLIAEKAVVTAEDGGSNGGMPCFGQLSGAFKELKRPMWILLLVTCLNW 301
Query: 297 IGWFPFNLFDTDWMGREIYGGDPEGGLIYDTGVRMGALGLLLNSVVLAVTSLLMERLCR- 355
I WFPF LFDTDWMG+E+YGG G YD GVR GALGL+LNSVVL TSL ++ L R
Sbjct: 302 IAWFPFLLFDTDWMGKEVYGGTVGEGHAYDMGVRAGALGLMLNSVVLGATSLGVDILARG 361
Query: 356 KRGAGFVWGISNIFMAICF-IAMLVLTYAANS------IGYVSKGQPPPTGIVIAALAIF 408
G +WGI N +AIC + +LV A +S G + PP GI AL +F
Sbjct: 362 VGGVKRLWGIVNFLLAICLGLTVLVTKLAQHSRQYAPGTGGLQDPLPPSGGIKAGALTLF 421
Query: 409 TILGFPMAITYSVPYALISTHIEPLGLGQGLSMGVLNLAIVVPQIVVSLGSGPWDQLFGG 468
++LG P+AITYS+P+AL S G GQGLS+GVLNLAIV+PQ+ VS+ SGPWD LFGG
Sbjct: 422 SVLGIPLAITYSIPFALASIFSSTSGAGQGLSLGVLNLAIVIPQMFVSVLSGPWDALFGG 481
Query: 469 GNSPAFAVAAVAALLSGLLALLAIP 493
GN PAF V AVAAL SG+L+++ +P
Sbjct: 482 GNLPAFVVGAVAALASGILSMILLP 506
>UniRef100_O80550 T22J18.12 protein [Arabidopsis thaliana]
Length = 512
Score = 530 bits (1364), Expect = e-149
Identities = 262/494 (53%), Positives = 344/494 (69%), Gaps = 13/494 (2%)
Query: 16 SSTSTSTSRPVQPVQPRTPLRQLLRVASVASGIQFGWALQLSLLTPYVQQLGIPHKWASI 75
S+ T T QP + LR+++ V+S+A+G+QFGWALQLSLLTPYVQ LGIPHKWAS+
Sbjct: 14 SALETQTGELDQPER----LRKIISVSSIAAGVQFGWALQLSLLTPYVQLLGIPHKWASL 69
Query: 76 IWLCGPVSGLFVQPLVGHLSDRCSSRFGRRRPFILVGAASIVVAVVIIGYAADIGYLIGD 135
IWLCGP+SG+ VQP+VG+ SDRC+SRFGRRRPFI+ GA + VAV +IGYAADIG+ +GD
Sbjct: 70 IWLCGPISGMLVQPIVGYHSDRCTSRFGRRRPFIVAGAGLVTVAVFLIGYAADIGHSMGD 129
Query: 136 DITQNYRPFAIVVFVIGFWILDVANNVTQGPCRALLADLTCNDARRTRVANAYFSLFMAV 195
+ + + AI +F +GFWILDVANN QGPCRA LADL+ +A++TR ANA+FS FMAV
Sbjct: 130 QLDKPPKTRAIAIFALGFWILDVANNTLQGPCRAFLADLSAGNAKKTRTANAFFSFFMAV 189
Query: 196 GNILGYATGSYSGWYKIFTFTLTPACSISCANLKSAFFLDVAFIVVTTYLSIVSAHEVPL 255
GN+LGYA GSY YK+ FT+T +C + CANLK+ FFL + +++ T++S+ E P
Sbjct: 190 GNVLGYAAGSYRNLYKVVPFTMTESCDLYCANLKTCFFLSITLLLIVTFVSLCYVKEKPW 249
Query: 256 SSSGAGESGSAEEAFMWELFGTFKYFSMPVWIVLSVTALTWIGWFPFNLFDTDWMGREIY 315
+ + ++ F E+FG FK P+W++L VTAL WI WFPF LFDTDWMGRE+Y
Sbjct: 250 TPEPTADGKASNVPFFGEIFGAFKELKRPMWMLLIVTALNWIAWFPFLLFDTDWMGREVY 309
Query: 316 GGDPEGGL------IYDTGVRMGALGLLLNSVVLAVTSLLMERLCRK-RGAGFVWGISNI 368
GG+ + +Y+ GVR GALGL+LN++VL SL +E + RK GA +WGI N
Sbjct: 310 GGNSDATATAASKKLYNDGVRAGALGLMLNAIVLGFMSLGVEWIGRKLGGAKRLWGIVNF 369
Query: 369 FMAICFIAMLVLTYAANSIGYVSKGQP--PPTGIVIAALAIFTILGFPMAITYSVPYALI 426
+AIC +V+T A + G PP + AL +F ILG P AIT+S+P+AL
Sbjct: 370 ILAICLAMTVVVTKQAENHRRDHGGAKTGPPGNVTAGALTLFAILGIPQAITFSIPFALA 429
Query: 427 STHIEPLGLGQGLSMGVLNLAIVVPQIVVSLGSGPWDQLFGGGNSPAFAVAAVAALLSGL 486
S G GQGLS+GVLNLAIVVPQ+V+S+G GP+D+LFGGGN PAF + A+AA +SG+
Sbjct: 430 SIFSTNSGAGQGLSLGVLNLAIVVPQMVISVGGGPFDELFGGGNIPAFVLGAIAAAVSGV 489
Query: 487 LALLAIPRTRTQKP 500
LAL +P P
Sbjct: 490 LALTVLPSPPPDAP 503
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.325 0.140 0.437
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 839,551,816
Number of Sequences: 2790947
Number of extensions: 35944094
Number of successful extensions: 138114
Number of sequences better than 10.0: 665
Number of HSP's better than 10.0 without gapping: 386
Number of HSP's successfully gapped in prelim test: 279
Number of HSP's that attempted gapping in prelim test: 136830
Number of HSP's gapped (non-prelim): 917
length of query: 504
length of database: 848,049,833
effective HSP length: 132
effective length of query: 372
effective length of database: 479,644,829
effective search space: 178427876388
effective search space used: 178427876388
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 77 (34.3 bits)
Medicago: description of AC146866.2


