

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146866.12 + phase: 0
(273 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
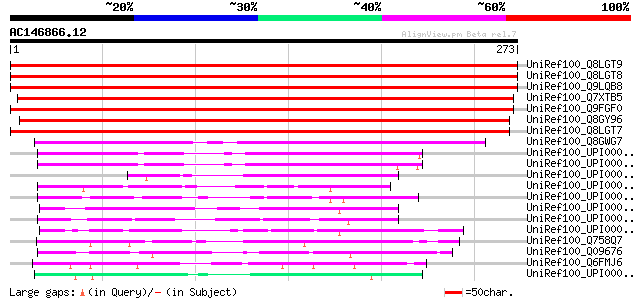
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q8LGT9 Phosphoglycerate mutase-like protein [Glycine max] 447 e-124
UniRef100_Q8LGT8 Phosphoglycerate mutase-like protein [Glycine max] 444 e-124
UniRef100_Q9LQB8 F19C14.10 protein [Arabidopsis thaliana] 382 e-105
UniRef100_Q7XTB5 OSJNBa0068L06.8 protein [Oryza sativa] 379 e-104
UniRef100_Q9FGF0 Putative ZW10 protein [Arabidopsis thaliana] 377 e-103
UniRef100_Q8GY96 Hypothetical protein At2g17280 [Arabidopsis tha... 362 7e-99
UniRef100_Q8LGT7 Hypothetical protein [Glycine max] 310 3e-83
UniRef100_Q8GWG7 Hypothetical protein At1g09932 [Arabidopsis tha... 192 8e-48
UniRef100_UPI0000274A0A UPI0000274A0A UniRef100 entry 91 3e-17
UniRef100_UPI000025C913 UPI000025C913 UniRef100 entry 88 2e-16
UniRef100_UPI000025AD06 UPI000025AD06 UniRef100 entry 75 2e-12
UniRef100_UPI000029101B UPI000029101B UniRef100 entry 67 4e-10
UniRef100_UPI0000319C2C UPI0000319C2C UniRef100 entry 65 2e-09
UniRef100_UPI000021B807 UPI000021B807 UniRef100 entry 64 5e-09
UniRef100_UPI000042F445 UPI000042F445 UniRef100 entry 62 2e-08
UniRef100_UPI000023E2A1 UPI000023E2A1 UniRef100 entry 62 2e-08
UniRef100_Q758Q7 AEL304Cp [Ashbya gossypii] 61 4e-08
UniRef100_Q09676 Hypothetical protein C5H10.03 in chromosome I [... 58 2e-07
UniRef100_Q6FMJ6 Similar to sp|P36069 Saccharomyces cerevisiae Y... 57 5e-07
UniRef100_UPI000042ECE8 UPI000042ECE8 UniRef100 entry 56 1e-06
>UniRef100_Q8LGT9 Phosphoglycerate mutase-like protein [Glycine max]
Length = 284
Score = 447 bits (1151), Expect = e-124
Identities = 209/274 (76%), Positives = 238/274 (86%), Gaps = 1/274 (0%)
Query: 1 MDTAPGQSLYPLHHSKTIHLVRHAQGVHNVEGEKNHDAYLSYDFFDANLTPLGWQQVENL 60
MDTA GQSLYPLH KT+HLVRHAQG HNVEGEKN +AY SYD FDANLTPLGW+QV+NL
Sbjct: 1 MDTAAGQSLYPLHRCKTLHLVRHAQGFHNVEGEKNFEAYKSYDLFDANLTPLGWKQVDNL 60
Query: 61 QKHVKAIGLSKKIELVVVSPLLRTMQTAVGVFGGEANTDGVNKPPLMIENVGHSDHPAVS 120
++HVKA GLSK+IELV+VSPLLRTMQTAVGVFGG+ TDG+N PPLM +NVG S PA+S
Sbjct: 61 RQHVKASGLSKRIELVIVSPLLRTMQTAVGVFGGQPYTDGINVPPLMNDNVGDSGRPAIS 120
Query: 121 SLNCPPFVAVELCREQMGLHPCDKRRTVSEYRHMFPGIDFSLIETDDDTWWKPE-REKKE 179
SLN PPF+AVELCRE +G+HPCDKRR +++YRHMFP IDFSLIE D+D WKP+ REK E
Sbjct: 121 SLNAPPFIAVELCREHLGVHPCDKRRNITDYRHMFPAIDFSLIENDEDILWKPDIREKNE 180
Query: 180 EVTGRGLKFLEWLCTRKEKEIAVVTHSSFLFNTLSAFGNDCHPNIKTEMCAHFANCELRS 239
EV RGLKFLEWL TRKEKEIAVVTHS FLF++LSAFGNDCHPN+K E+C HFANCELRS
Sbjct: 181 EVAARGLKFLEWLWTRKEKEIAVVTHSGFLFHSLSAFGNDCHPNVKNEICTHFANCELRS 240
Query: 240 MVIVDKCMIGSNNSTTNYPGKIPHGPDLPSDATD 273
MVI+D+ MIGS+ S+TNYPGK+P G DLPSD D
Sbjct: 241 MVIIDRGMIGSDESSTNYPGKVPDGLDLPSDVAD 274
>UniRef100_Q8LGT8 Phosphoglycerate mutase-like protein [Glycine max]
Length = 284
Score = 444 bits (1143), Expect = e-124
Identities = 208/274 (75%), Positives = 237/274 (85%), Gaps = 1/274 (0%)
Query: 1 MDTAPGQSLYPLHHSKTIHLVRHAQGVHNVEGEKNHDAYLSYDFFDANLTPLGWQQVENL 60
MDTA GQSLYPLH KT+HLVRHAQG HNVEGEKN +AY SYD FDANLTPLGW+QV+NL
Sbjct: 1 MDTAAGQSLYPLHRCKTLHLVRHAQGFHNVEGEKNFEAYKSYDLFDANLTPLGWKQVDNL 60
Query: 61 QKHVKAIGLSKKIELVVVSPLLRTMQTAVGVFGGEANTDGVNKPPLMIENVGHSDHPAVS 120
++HVKA GLSK+IELV+VSPLLRTMQTAVGVFGG+ TDG+N PPLM +NVG S PA+S
Sbjct: 61 RQHVKASGLSKRIELVIVSPLLRTMQTAVGVFGGQPYTDGINVPPLMNDNVGDSGRPAIS 120
Query: 121 SLNCPPFVAVELCREQMGLHPCDKRRTVSEYRHMFPGIDFSLIETDDDTWWKPE-REKKE 179
SLN PPF+AVELCRE +G+H CDKRR +++YRHMFP IDFSLIE D+D WKP+ REK E
Sbjct: 121 SLNAPPFIAVELCREHLGVHSCDKRRNITDYRHMFPAIDFSLIENDEDILWKPDIREKNE 180
Query: 180 EVTGRGLKFLEWLCTRKEKEIAVVTHSSFLFNTLSAFGNDCHPNIKTEMCAHFANCELRS 239
EV RGLKFLEWL TRKEKEIAVVTHS FLF++LSAFGNDCHPN+K E+C HFANCELRS
Sbjct: 181 EVAARGLKFLEWLWTRKEKEIAVVTHSGFLFHSLSAFGNDCHPNVKNEICTHFANCELRS 240
Query: 240 MVIVDKCMIGSNNSTTNYPGKIPHGPDLPSDATD 273
MVI+D+ MIGS+ S+TNYPGK+P G DLPSD D
Sbjct: 241 MVIIDRGMIGSDESSTNYPGKVPDGLDLPSDVAD 274
>UniRef100_Q9LQB8 F19C14.10 protein [Arabidopsis thaliana]
Length = 313
Score = 382 bits (982), Expect = e-105
Identities = 180/274 (65%), Positives = 212/274 (76%), Gaps = 1/274 (0%)
Query: 1 MDTAPGQSLYPLHHSKTIHLVRHAQGVHNVEGEKNHDAYLSYDFFDANLTPLGWQQVENL 60
M+T P Q LYPLH KTIHLVRHAQG+HNVEGEKNH AYLS D FDA+LTPLGWQQV+NL
Sbjct: 38 METKPSQGLYPLHRCKTIHLVRHAQGIHNVEGEKNHKAYLSEDLFDAHLTPLGWQQVDNL 97
Query: 61 QKHVKAIGLSKKIELVVVSPLLRTMQTAVGVFGGEANTDGVNKPPLMIENVGHSDHPAVS 120
KHV A G+S +IELVVVSPLLRT+QTAVG FGGE DGVN P LM G+SD PA+S
Sbjct: 98 HKHVNASGISNRIELVVVSPLLRTLQTAVGTFGGEGYKDGVNTPLLMTAGAGNSDRPAIS 157
Query: 121 SLNCPPFVAVELCREQMGLHPCDKRRTVSEYRHMFPGIDFSLIETDDDTWWKPE-REKKE 179
LN PPF+AVE CRE +G+HPCD+R +++YR +FP IDFSLIETD+D WKP+ RE+ +
Sbjct: 158 RLNRPPFIAVESCREHLGVHPCDRRSNITKYRELFPAIDFSLIETDEDVLWKPDIREEDK 217
Query: 180 EVTGRGLKFLEWLCTRKEKEIAVVTHSSFLFNTLSAFGNDCHPNIKTEMCAHFANCELRS 239
++ RG+KF WL TRKEKEIAVVTHS FL+ TL++FGNDC P++K E+ F NCELRS
Sbjct: 218 DIATRGVKFFNWLSTRKEKEIAVVTHSGFLYQTLNSFGNDCDPSVKNEISKKFVNCELRS 277
Query: 240 MVIVDKCMIGSNNSTTNYPGKIPHGPDLPSDATD 273
V+VDKCM S+ TNYPG I G D SD D
Sbjct: 278 FVLVDKCMSSSDPPMTNYPGTILTGEDASSDIAD 311
>UniRef100_Q7XTB5 OSJNBa0068L06.8 protein [Oryza sativa]
Length = 275
Score = 379 bits (974), Expect = e-104
Identities = 174/268 (64%), Positives = 216/268 (79%), Gaps = 1/268 (0%)
Query: 5 PGQSLYPLHHSKTIHLVRHAQGVHNVEGEKNHDAYLSYDFFDANLTPLGWQQVENLQKHV 64
P ++YPLH KTI+LVRHAQGVHNVEGEK+H AY+S FDA+LTPLGW QV+ L++HV
Sbjct: 6 PTTAMYPLHRCKTIYLVRHAQGVHNVEGEKDHSAYMSPQLFDAHLTPLGWNQVDCLREHV 65
Query: 65 KAIGLSKKIELVVVSPLLRTMQTAVGVFGGEANTDGVNKPPLMIENVGHSDHPAVSSLNC 124
K GL++KIELV+ SPLLRTMQTAVGVFGGE + DGV+ PPLM+EN GHS PA+SSLNC
Sbjct: 66 KKSGLAQKIELVITSPLLRTMQTAVGVFGGENSVDGVSAPPLMVENAGHSSRPAISSLNC 125
Query: 125 PPFVAVELCREQMGLHPCDKRRTVSEYRHMFPGIDFSLIETDDDTWWKPE-REKKEEVTG 183
PPF+A E CRE +G+HPCDKRR+++EY +FP IDFSLIE D+D W+P RE V
Sbjct: 126 PPFLAFEACREHLGVHPCDKRRSITEYHALFPAIDFSLIENDEDVLWEPNVREANSSVAA 185
Query: 184 RGLKFLEWLCTRKEKEIAVVTHSSFLFNTLSAFGNDCHPNIKTEMCAHFANCELRSMVIV 243
RG+KF++WL TR+EKEIA+V+HS FL++TLS + +CHP I+ E+ HFANCELRSMV+V
Sbjct: 186 RGMKFIDWLWTREEKEIAIVSHSGFLYHTLSMYSRECHPTIREEVGKHFANCELRSMVLV 245
Query: 244 DKCMIGSNNSTTNYPGKIPHGPDLPSDA 271
D M+GS++ + NYPG IP G DLPSDA
Sbjct: 246 DTSMLGSDSPSYNYPGSIPAGLDLPSDA 273
>UniRef100_Q9FGF0 Putative ZW10 protein [Arabidopsis thaliana]
Length = 282
Score = 377 bits (968), Expect = e-103
Identities = 174/272 (63%), Positives = 221/272 (80%), Gaps = 1/272 (0%)
Query: 1 MDTAPGQSLYPLHHSKTIHLVRHAQGVHNVEGEKNHDAYLSYDFFDANLTPLGWQQVENL 60
M+T G LYPLH KTI+LVRHAQG+HNV+GEKN+ AY+S+D+FDA LT LGW+QV++L
Sbjct: 1 METGAGIGLYPLHRCKTIYLVRHAQGIHNVDGEKNYKAYMSHDYFDAELTQLGWKQVDSL 60
Query: 61 QKHVKAIGLSKKIELVVVSPLLRTMQTAVGVFGGEANTDGVNKPPLMIENVGHSDHPAVS 120
+KHV + GL KKIELV+ SPL+RT+QTAVGVFGGE TD + PLM+ N G+S A+S
Sbjct: 61 RKHVHSSGLHKKIELVISSPLMRTLQTAVGVFGGEGYTDMSDVLPLMVANAGNSSRAAIS 120
Query: 121 SLNCPPFVAVELCREQMGLHPCDKRRTVSEYRHMFPGIDFSLIETDDDTWWKPE-REKKE 179
SLNCPP + E CRE +G+HPCD+RR++S+Y+ +FP +DFSLIE+++D WK + RE E
Sbjct: 121 SLNCPPVITEESCREHLGVHPCDQRRSISDYQFLFPAVDFSLIESEEDKLWKADVRETIE 180
Query: 180 EVTGRGLKFLEWLCTRKEKEIAVVTHSSFLFNTLSAFGNDCHPNIKTEMCAHFANCELRS 239
E+ RG KFL WL TRKEKEIA+VTHS FLF+TL+A N+CHP++K E+C HFANCELRS
Sbjct: 181 ELAARGKKFLNWLWTRKEKEIAIVTHSGFLFHTLNALQNECHPDVKKEICGHFANCELRS 240
Query: 240 MVIVDKCMIGSNNSTTNYPGKIPHGPDLPSDA 271
MVIVD+ M+GS++S T+YPGKIP G DLPSDA
Sbjct: 241 MVIVDRSMLGSDSSVTDYPGKIPKGIDLPSDA 272
>UniRef100_Q8GY96 Hypothetical protein At2g17280 [Arabidopsis thaliana]
Length = 271
Score = 362 bits (928), Expect = 7e-99
Identities = 170/266 (63%), Positives = 213/266 (79%), Gaps = 2/266 (0%)
Query: 6 GQSLYPLHHSKTIHLVRHAQGVHNVEGEKNHDAYLSYDFFDANLTPLGWQQVENLQKHVK 65
G LYPLH KTIHLVRHAQG+HNV GEK+H AY S D+FDA+LTPLGWQQV+NL+ HV+
Sbjct: 5 GIGLYPLHRCKTIHLVRHAQGIHNVAGEKDHSAYSSEDYFDAHLTPLGWQQVDNLRNHVR 64
Query: 66 AIGLSKKIELVVVSPLLRTMQTAVGVFGGEANTDGVNKPPLMIENVGHSDHPAVSSLNCP 125
A L K+ELV+VSP+LRT+QTAVG FGGE +T+G + PLM+ N G SD PA+SSLN P
Sbjct: 65 AAQLLNKVELVIVSPMLRTIQTAVGAFGGEEDTNGADATPLMVANAGSSDRPAISSLNSP 124
Query: 126 PFVAVELCREQMGLHPCDKRRTVSEYRHMFPGIDFSLIETDDDTWWKPE-REKKEEVTGR 184
PF+AVELCRE MG HPCD+RR+V+EY+ +FP IDFS+IETD+D WKP RE EEV R
Sbjct: 125 PFLAVELCRETMGDHPCDRRRSVTEYKALFPAIDFSIIETDNDVLWKPSPRESLEEVAAR 184
Query: 185 GLKFLEWLCTRKEKEIAVVTHSSFLFNTLSAFGNDCHPNIKTEMCAHFANCELRSMVIVD 244
G++F++W+ TRKEKEIA+V+HS FL LS+FG DC ++K E+ H +NCELRSMVIVD
Sbjct: 185 GVEFIKWIWTRKEKEIAIVSHSGFLHGLLSSFGKDCCDDLKKELSIHLSNCELRSMVIVD 244
Query: 245 KCMIGSNNS-TTNYPGKIPHGPDLPS 269
+ +G++++ TTNYPGK+P G D PS
Sbjct: 245 RGNLGTDSAETTNYPGKVPEGLDNPS 270
>UniRef100_Q8LGT7 Hypothetical protein [Glycine max]
Length = 313
Score = 310 bits (793), Expect = 3e-83
Identities = 165/272 (60%), Positives = 195/272 (71%), Gaps = 3/272 (1%)
Query: 1 MDTAPGQSLYPLHHSKTIHLVRHAQGVHNVEGEKNHDAYLSYDFFDANLTPLGWQQVENL 60
MDTA GQS +PLH KT+HLVRHAQG HNVEGEKN +AY SYD FDANLTPLGW QV+NL
Sbjct: 1 MDTAAGQSPHPLHRCKTLHLVRHAQGFHNVEGEKNFEAYKSYDLFDANLTPLGWNQVDNL 60
Query: 61 QKHVKAIGLSKKIELVVVSPLLRTMQTAVGVFGGEANTDGVNKPPLMIENVGHSDHPAVS 120
++HVKA GLSKKIELV+VSPLLRTMQTAVGVFGGEA TDG+N PPLM +NVG S PA+S
Sbjct: 61 REHVKASGLSKKIELVIVSPLLRTMQTAVGVFGGEAYTDGINVPPLMNDNVGDSRRPAIS 120
Query: 121 SLNCPPFVAVE-LCREQMGLHPCDKRRTVSEYRHMFPGIDFSLIETDDDTWWKPE-REKK 178
SLN PPF + L R G+ C +++ ++ + F +T P REK
Sbjct: 121 SLNVPPFNSSRALPRTFWGVSLCKEKKHHCLPTYVSQLLIFHCYKTMPTFCGNPPIREKN 180
Query: 179 EEVTGRGLKFLEWLC-TRKEKEIAVVTHSSFLFNTLSAFGNDCHPNIKTEMCAHFANCEL 237
+G + C RK+KE AVVTH FLF++L A GNDCHPN+K E+C HFANCEL
Sbjct: 181 CRSCCQGTEIFGNGCGHRKKKEKAVVTHRGFLFHSLRALGNDCHPNVKNEICTHFANCEL 240
Query: 238 RSMVIVDKCMIGSNNSTTNYPGKIPHGPDLPS 269
RSMVI+DK +IGSN S+TNY GKIP+G PS
Sbjct: 241 RSMVIIDKGVIGSNESSTNYTGKIPYGRPCPS 272
>UniRef100_Q8GWG7 Hypothetical protein At1g09932 [Arabidopsis thaliana]
Length = 260
Score = 192 bits (488), Expect = 8e-48
Identities = 105/244 (43%), Positives = 146/244 (59%), Gaps = 15/244 (6%)
Query: 14 HSKTIHLVRHAQGVHNVEGEKNHDAYLSYDFFDANLTPLGWQQVENLQKHVKAIGLSKKI 73
H KT+HLVRHAQGVHN+ E+ + S FDA+L+P G QQV + + GL +
Sbjct: 15 HFKTLHLVRHAQGVHNIALEEKGEKPESEKLFDAHLSPKGLQQVSERRNQILESGLLNTV 74
Query: 74 ELVVVSPLLRTMQTAVGVFGGEANTDGVNKPPLMIENVGHSDHPAVSSLNCPPFVAVELC 133
ELV+ SPL R M+T++G+F G+ + + E+ ++ N PP VA+E+C
Sbjct: 75 ELVITSPLCRAMETSIGIFRGQGYVN-------ISEDFAKAN-------NFPPIVALEIC 120
Query: 134 REQMGLHPCDKRRTVSEYRHMFPGIDFSLIETDDDTWWK-PEREKKEEVTGRGLKFLEWL 192
RE+MGL+PCD+R ++S R FP IDF++IE+D+D W+ EREK E+V RGL F++WL
Sbjct: 121 RERMGLYPCDRRASISTRRTFFPEIDFTMIESDEDALWQDKEREKLEDVATRGLHFVKWL 180
Query: 193 CTRKEKEIAVVTHSSFLFNTLSAFGNDCHPNIKTEMCAHFANCELRSMVIVDKCMIGSNN 252
R EKEIA+V+H FL TL A ++ + FANCELRS+ I M
Sbjct: 181 WERPEKEIAIVSHGIFLQQTLCALHGKVGIPLEDSLLTRFANCELRSIRIEKSDMEADTL 240
Query: 253 STTN 256
T N
Sbjct: 241 MTCN 244
>UniRef100_UPI0000274A0A UPI0000274A0A UniRef100 entry
Length = 194
Score = 90.9 bits (224), Expect = 3e-17
Identities = 70/217 (32%), Positives = 100/217 (45%), Gaps = 40/217 (18%)
Query: 16 KTIHLVRHAQGVHNVEG-EKNHDAYLSYDFFDANLTPLGWQQVENLQKHVKAIGLSKKIE 74
K I +RH HN++ E DAYL + FDA L G QQ ++L K + ++
Sbjct: 2 KNIFCIRHGLAFHNIKAMEIGSDAYLMEECFDAPLVEKGIQQAKDLGNQWKGLNA---VQ 58
Query: 75 LVVVSPLLRTMQTAVGVFGGEANTDGVNKPPLMIENVGHSDHPAVSSLNCPPFVAVELCR 134
+V VSPL RT+QT +F HP V +A + +
Sbjct: 59 IVFVSPLTRTLQTCQEIFHA---------------------HPGVK------IIAHDKVK 91
Query: 135 EQ-MGLHPCDKRRTVSEYRHMFPGIDFSLIETDDDTWWKPER-EKKEEVTGRGLKFLEWL 192
E GL C+KRR E FPGIDFS ++++ D W+ +R E+ EE+ R +F L
Sbjct: 92 EYPQGLQICNKRREKLELEKQFPGIDFSGLDSESDEMWRDDRLEEIEELRERIEQFKTML 151
Query: 193 CTRKEKEIAVVTHSSFLFNTLSAFGND-------CHP 222
E +IA+V+HS++L L D CHP
Sbjct: 152 QGLAETQIAIVSHSAYLNQFLYGILGDESNPLKHCHP 188
>UniRef100_UPI000025C913 UPI000025C913 UniRef100 entry
Length = 194
Score = 88.2 bits (217), Expect = 2e-16
Identities = 71/217 (32%), Positives = 100/217 (45%), Gaps = 40/217 (18%)
Query: 16 KTIHLVRHAQGVHNVEG-EKNHDAYLSYDFFDANLTPLGWQQVENLQKHVKAIGLSKKIE 74
K I +RH HN++ E DAYL + FDA L G QQ ++L K + ++
Sbjct: 2 KNIFCIRHGLAFHNIKAMEIGSDAYLMEECFDAPLVEKGIQQAKDLGNQWKGLNA---VQ 58
Query: 75 LVVVSPLLRTMQTAVGVFGGEANTDGVNKPPLMIENVGHSDHPAVSSLNCPPFVAVELCR 134
+V VSPL RT+QT +F HP V +A + +
Sbjct: 59 IVFVSPLTRTLQTCEEIFQA---------------------HPGVK------IIAHDQVK 91
Query: 135 EQ-MGLHPCDKRRTVSEYRHMFPGIDFSLIETDDDTWWKPER-EKKEEVTGRGLKFLEWL 192
E GL C+KRR E F GIDFS ++++ D W+ +R E+ EE+ R +F L
Sbjct: 92 EYPQGLQICNKRREKLELEKQFQGIDFSGLDSESDEMWRDDRLEEIEELRERIEQFKIML 151
Query: 193 CTRKEKEIAVVTHSS----FLFNTLSAFGN---DCHP 222
E IA+V+HS+ FL+ TL N CHP
Sbjct: 152 QGLAETNIAIVSHSAYLNQFLYGTLGDESNPLKHCHP 188
>UniRef100_UPI000025AD06 UPI000025AD06 UniRef100 entry
Length = 154
Score = 74.7 bits (182), Expect = 2e-12
Identities = 54/154 (35%), Positives = 78/154 (50%), Gaps = 36/154 (23%)
Query: 64 VKAIGLSKK------IELVVVSPLLRTMQTAVGVFGGEANTDGVNKPPLMIENVGHSDHP 117
++A+ L KK +ELVVVSPL+RT+QTA +F + N
Sbjct: 1 IQALELKKKWKHLKDVELVVVSPLMRTLQTADTIF-KDTNI------------------- 40
Query: 118 AVSSLNCPPFVAVELCRE-QMGLHPCDKRRTVSEYRHMFPGIDFSLIETDDDTWWKPERE 176
P +A+E RE MG C+KR + +FP IDF ++T++D W P+RE
Sbjct: 41 --------PMIALECVREFPMGKQTCNKRSDKDDLLRLFPYIDFRDLKTNEDKLWDPQRE 92
Query: 177 KK-EEVTGRGLKFLEWLCTRKEKEIAVVTHSSFL 209
+ E+ R K L +L +R E IA+V HSSF+
Sbjct: 93 ETIPELNARINKLLSFLHSRPENVIALVNHSSFI 126
>UniRef100_UPI000029101B UPI000029101B UniRef100 entry
Length = 241
Score = 67.4 bits (163), Expect = 4e-10
Identities = 67/203 (33%), Positives = 84/203 (41%), Gaps = 39/203 (19%)
Query: 16 KTIHLVRHAQGVHNVEGEKNHDA---YLSYDFFDANLTPLGWQQVENLQKHVKAIGLSKK 72
K IHLVRH + N +NH A ++ D LT +G +Q L+ A GL
Sbjct: 52 KVIHLVRHGRTEMNDYLRENHWADPDFVDPMMIDTRLTSVGERQARELR--ATARGLDPV 109
Query: 73 IELVVVSPLLRTMQTAVGVFGGEANTDGVNKPPLMIENVGHSDHPAVSSLNCPPFVAVEL 132
EL+V SPL R ++TA FG EA D V L P V L
Sbjct: 110 PELIVASPLRRALRTAELAFG-EAGEDEV--------------------LGDVPRVVCAL 148
Query: 133 CREQMGLHPCDKRRTVSEYRHMFPGIDFSLIETDDDTWW----------KPEREKKEEVT 182
RE++ H D R VSE D+ + E D WW E E E
Sbjct: 149 ARERL-FHGSDIGRLVSELSG--DHADWDMSELGDGAWWYAPEGRDPFTTAELEPAETFE 205
Query: 183 GRGLKFLEWLCTRKEKEIAVVTH 205
R +F+ WL R EK IAVV+H
Sbjct: 206 ARMEEFVAWLEDRPEKSIAVVSH 228
>UniRef100_UPI0000319C2C UPI0000319C2C UniRef100 entry
Length = 180
Score = 64.7 bits (156), Expect = 2e-09
Identities = 60/214 (28%), Positives = 89/214 (41%), Gaps = 51/214 (23%)
Query: 16 KTIHLVRHAQGVHNVEGEKNHDAYLSYDFFDANLTPLGWQQVENLQKHVKAIGLSKKIEL 75
K I+L+RHA+ N + ++ Y ++DA +T G Q + ++K I + +
Sbjct: 2 KKIYLIRHAESEANAAMDLDNPTY----YYDAKITKRGEDQAAKARDNLKNI----EFDT 53
Query: 76 VVVSPLLRTMQTAVGVFGGEANTDGVNKPPLMIENVGHSDHPAVSSLNCPPFVAVELCRE 135
+ SPL RTMQT +F NK PL+ L RE
Sbjct: 54 YICSPLPRTMQTFSIIFP--------NKKPLI----------------------QPLIRE 83
Query: 136 QMGLHPCDKRRTVSEYRHMFPGIDFSLIETDDDTWW---KPEREKK------EEVTGRGL 186
Q+ H CD R + + FP DFS + + WW +P EKK ++ R
Sbjct: 84 QL-YHSCDVGRQPNILKKEFPLFDFSKL---NQYWWNNDEPINEKKIVKENFNDIKIRLE 139
Query: 187 KFLEWLCTRKEKEIAVVTHSSFLFNTLSAFGNDC 220
KF WL IA+V+H +FL F +C
Sbjct: 140 KFKSWLMLNDSSTIAIVSHGTFLSQITGYFLENC 173
>UniRef100_UPI000021B807 UPI000021B807 UniRef100 entry
Length = 244
Score = 63.5 bits (153), Expect = 5e-09
Identities = 59/198 (29%), Positives = 86/198 (42%), Gaps = 36/198 (18%)
Query: 17 TIHLVRHAQGVHNVEGEKNHDAYLSYDFFDANLTPLGWQQVENLQKHV-KAIGLSKKIEL 75
T+ L+RHAQ HNV S + D LTPLG +Q L H+ K + S EL
Sbjct: 4 TLILIRHAQAEHNV----------SNNIPDPELTPLGKEQAAALSAHLQKRLPGSLDPEL 53
Query: 76 VVVSPLLRTMQTAVGVFGGEANTDGVNKPPLMIENVGHSDHPAVSSLNCPPFVAVELCRE 135
++VSP R +QTA F + + G S P V++ + +
Sbjct: 54 IIVSPFRRCLQTATIAFDWLIDAES-----------GRSKVPMVANASW----------Q 92
Query: 136 QMGLHPCDKRRTVSEYRHMFPGIDFSLIETDDDTWWKPERE----KKEEVTGRGLKFLEW 191
+ PCD + FP IDFS ++ P +E + GR L+
Sbjct: 93 ENADKPCDTGTDPTLVAPNFPHIDFSTLDPVYPDKTSPAASLYHYTREALLGRAQSCLKE 152
Query: 192 LCTRKEKEIAVVTHSSFL 209
L +R E+ IAVV+HS+F+
Sbjct: 153 LRSRPERVIAVVSHSAFM 170
>UniRef100_UPI000042F445 UPI000042F445 UniRef100 entry
Length = 234
Score = 62.0 bits (149), Expect = 2e-08
Identities = 59/201 (29%), Positives = 85/201 (41%), Gaps = 42/201 (20%)
Query: 16 KTIHLVRHAQGVHNVEGEKNHDAYLSYDFFDANLTPLGWQQVENLQKHVKAIGLSKKIEL 75
K IHL RHAQ HNV + Y DA LT LG +Q L + K G+ K EL
Sbjct: 10 KRIHLTRHAQAEHNVADD--------YTIADAPLTALGQEQSRQLNEATKN-GVQKTAEL 60
Query: 76 VVVSPLLRTMQTAVGVFGGEANTDGVNKPPLMIENVGHSDHPAVSSLNCPPFVAVELCRE 135
+V SPL R ++T + +G+ + + + P + +++ +E
Sbjct: 61 LVTSPLRRPLETML---------------------LGYPELKSRLEKSGKPVILLDILQE 99
Query: 136 QMGLHPCD-KRRTVSEYRHMFPGIDFSLIETDDDTWWKPEREKKEEV------TGRGLKF 188
+G +PCD +S + GI SL D + P+ KE + R
Sbjct: 100 -VGPYPCDTPTHPISALKASNGGIFSSL----DFSTLSPDYASKEGIFAPANGAARAKLA 154
Query: 189 LEWLCTRKEKEIAVVTHSSFL 209
+WL R EKEI VV H L
Sbjct: 155 RKWLRERPEKEIVVVAHGDIL 175
>UniRef100_UPI000023E2A1 UPI000023E2A1 UniRef100 entry
Length = 245
Score = 62.0 bits (149), Expect = 2e-08
Identities = 74/236 (31%), Positives = 103/236 (43%), Gaps = 55/236 (23%)
Query: 17 TIHLVRHAQGVHNVEGEKNHDAYLSYDFFDANLTPLGWQQVENLQKHVKAIGLSKKIELV 76
TIHLVRHAQGVHN+ N D D D +LTPLG +Q +L+ + KI +
Sbjct: 4 TIHLVRHAQGVHNL---PNGD-----DIPDPDLTPLGEEQCASLR---EKFPYHDKITKL 52
Query: 77 VVSPLLRTMQTAVGVFGGEANTDGVNKPPLMIENVGHSDHPAVSSLNCPPFVAVELCREQ 136
SP+ RT+ T FG LN P + + + +E
Sbjct: 53 FASPMRRTIYTCFLAFG-------------------------TKELN--PIIPLPILQEV 85
Query: 137 MGLHPCDKRRTVSEYRHMFPGI-DFSLIETDDDTWWKPERE-----KKEEVTGRGLKFLE 190
L PCD V+ + F GI D+S +E ++ T PE E +K EV GR + +
Sbjct: 86 SAL-PCDTGSPVTTVQAEFAGIADYSQVE-ENWTDKGPESEYYPTIEKLEVRGRKARNVL 143
Query: 191 WLCTRKEKEIAVVTHSSFL-FNTLSAFG-NDCHPNIKTEMCAHFANCELRSMVIVD 244
++ I VV+H FL T +G + PN ++NCE RS VD
Sbjct: 144 RDLVSGDEHIVVVSHGGFLHLLTDDWYGVPEGQPN-------SWSNCEFRSYQFVD 192
>UniRef100_Q758Q7 AEL304Cp [Ashbya gossypii]
Length = 303
Score = 60.8 bits (146), Expect = 4e-08
Identities = 64/253 (25%), Positives = 102/253 (40%), Gaps = 57/253 (22%)
Query: 15 SKTIHLVRHAQGVHNV-EGEKNHDAYLSY----------DFFDANLTPLGWQQVENLQKH 63
+K + RHA+G+HN E H+A+ Y + DA LTP G QQ +H
Sbjct: 72 AKLLIFQRHAEGLHNAAEARYGHEAWDDYWSKIDGDEYGTWVDAQLTPKGHQQASASSEH 131
Query: 64 ----VKAIGLSKKIELVVVSPLLRTMQTAVGVFGGEANTDGVNKPPLMIENVGHSDHPAV 119
V+A+G+ +++ SPL R ++T + + A +++P L
Sbjct: 132 VGALVRALGMPERL---YSSPLRRCLETFIEAWAPVAQY--ISEPVL------------- 173
Query: 120 SSLNCPPFVAVELCREQMGLHPCDKRRTVSEYRHMFPG---------IDFSLIETDDDTW 170
E RE +G+H CD+R S+ + G + + T+ DT
Sbjct: 174 ------DLYVREGLRETLGVHTCDRRVPHSQAVAAYQGHRLANSTLQLHYEPYYTEPDTL 227
Query: 171 WK-PEREKKEEVTGRGLKFLEWLCTRKEKEIAVVTHSSFLFNTLSAFGNDCHPNIKTEMC 229
W RE E+ R K L+ + R E+ I V HS + L A HP +
Sbjct: 228 WTVAHRETTPEIRNRVTKALDRILDRPERYIFVTAHSEMMDAALHALH---HPRVN---- 280
Query: 230 AHFANCELRSMVI 242
H N + +V+
Sbjct: 281 -HVPNAGILYLVV 292
>UniRef100_Q09676 Hypothetical protein C5H10.03 in chromosome I [Schizosaccharomyces
pombe]
Length = 219
Score = 58.2 bits (139), Expect = 2e-07
Identities = 60/228 (26%), Positives = 98/228 (42%), Gaps = 40/228 (17%)
Query: 16 KTIHLVRHAQGVHNVEGEKNHDAYLSYDFFDANLTPLGWQQVENLQKHVKAIGLSKKIEL 75
KT++L+RH Q HNV +++H + D LT G +Q E L K ++ SK+I +
Sbjct: 7 KTVYLIRHGQAQHNVGPDEDH------NIRDPVLTSEGIEQCEALAKELE----SKQIPI 56
Query: 76 --VVVSPLLRTMQTAVGVFGGEANTDGVNKPPLMIENVGHSDHPAVSSLNCPPFVAVELC 133
+V SP+ RT+QT G +K P+ I PF
Sbjct: 57 DGIVCSPMRRTLQTMEIALKKYLAEGGPDKVPVYIS----------------PFF----- 95
Query: 134 REQMGLHPCDKRRTVSEYRHMFPGIDFSLIETDDDTWWKPEREKKEEVT---GRGLKFLE 190
+++G PCD + + ++P +F D + + +VT R + LE
Sbjct: 96 -QEVGHLPCDIGLELDKLNKLYPKYNFQ--SCQDGIYPEKRDIYASDVTISAIRSKEALE 152
Query: 191 WLCTRKEKEIAVVTHSSFLFNTLSAFGNDCHPNIKTEMCAHFANCELR 238
+L +++IAV+THS+F+ L + + F NCE R
Sbjct: 153 YLAALPQQQIAVITHSAFIRFLLKKMVKAADIDFLPPQLS-FKNCEFR 199
>UniRef100_Q6FMJ6 Similar to sp|P36069 Saccharomyces cerevisiae YKL128c suppressor of
TPS2 mutant [Candida glabrata]
Length = 325
Score = 57.0 bits (136), Expect = 5e-07
Identities = 65/241 (26%), Positives = 102/241 (41%), Gaps = 51/241 (21%)
Query: 13 HHSKTIHLVRHAQGVHNVE----GEKNHDAYLSY-------DFFDANLTPLGWQQ-VENL 60
H K + L RH +G HN GEK + Y S + DA LTPLG +Q +E
Sbjct: 78 HSYKLLVLARHGEGYHNAAEARYGEKAWNEYWSKLEGDQYGSWLDAELTPLGKKQALEAG 137
Query: 61 QKHVKAI--GLSKKIELVVVSPLLRTMQTAVGVFGGEANTDGVNKPPLMIENVGHSDHPA 118
Q ++ + GL VSP+ R + T + + E V + P+
Sbjct: 138 QTYLTNLTDGLQLLPHKFFVSPMRRCLDTYIREW----------------EPVFSTHRPS 181
Query: 119 VSSLNCPPFVAVELCREQMGLHPCDKR----RTVSEYR-HMFPGIDFSL-------IETD 166
+++N +E RE +G+H CD+R + +SEY+ H + D ++
Sbjct: 182 NATVNVK---VIEYLRETLGVHTCDERVSHSQALSEYQDHRYNNSDVTVHFDYPGNYSEK 238
Query: 167 DDTWWKPEREKKEEVTGR---GLKFLEWLCTRKEKEIAVVTHSSFLFNTLSAFGNDCHPN 223
D W+ RE K E+ R GL+ + +K I++ HS + + L N HP
Sbjct: 239 DQLWYPDHRETKAEMDRRTRIGLREMFSSVNTTDKVISLTCHSGVIASVLR---NIKHPA 295
Query: 224 I 224
I
Sbjct: 296 I 296
>UniRef100_UPI000042ECE8 UPI000042ECE8 UniRef100 entry
Length = 315
Score = 55.8 bits (133), Expect = 1e-06
Identities = 59/223 (26%), Positives = 88/223 (39%), Gaps = 41/223 (18%)
Query: 14 HSKTIHLVRHAQGVHNVEGEK----NHDAYLSYD-------FFDANLTPLGWQQVENLQK 62
+ K L RH QG HNV + + Y + + DA LTP G QQ++NL +
Sbjct: 68 NEKLFFLQRHGQGWHNVAPSNFSRVDWNCYWAEQSGRDGVVWEDAELTPKGVQQIQNLHQ 127
Query: 63 HVKAIGLSKKIELVVVSPLLRTMQTAVGVFGGEANTDGVNKPPLMIENVGHSDHPAVSSL 122
+K + E VSPL RT+QT + G +K PL+
Sbjct: 128 RIKDTPDFPQPEKFFVSPLRRTLQTWNITWNGLP-----HKTPLI--------------- 167
Query: 123 NCPPFVAVELCREQMGLHPCDKRRTVSEYRHMFPGIDFSLIETDDDTWWKPER-EKKEEV 181
E RE G+ KR + + P +F T+ D W P++ E +
Sbjct: 168 -------KEFAREIYGIDSESKRHNKTFIHNYVPSFEFESGFTEQDENWSPDKSESDQHC 220
Query: 182 TGRGLKFLEWLC--TRKEKEIAVVTHSSFLFNTLSAFGNDCHP 222
R L+ + + EK I+VV HS ++ L G+ P
Sbjct: 221 DYRAAVLLQDIFNDSPDEKVISVVLHSGIIYCLLDVVGHRYFP 263
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.318 0.135 0.423
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 493,427,747
Number of Sequences: 2790947
Number of extensions: 20258454
Number of successful extensions: 42540
Number of sequences better than 10.0: 72
Number of HSP's better than 10.0 without gapping: 19
Number of HSP's successfully gapped in prelim test: 53
Number of HSP's that attempted gapping in prelim test: 42458
Number of HSP's gapped (non-prelim): 81
length of query: 273
length of database: 848,049,833
effective HSP length: 125
effective length of query: 148
effective length of database: 499,181,458
effective search space: 73878855784
effective search space used: 73878855784
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 74 (33.1 bits)
Medicago: description of AC146866.12


