

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146864.6 - phase: 0 /pseudo
(180 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
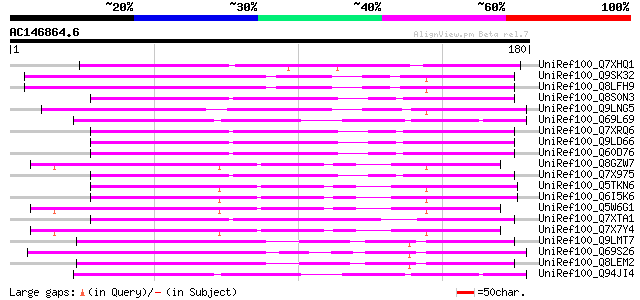
UniRef100_Q7XHQ1 Calcineurin-like phosphoesterase-like [Oryza sa... 75 8e-13
UniRef100_Q9SK32 Expressed protein [Arabidopsis thaliana] 71 1e-11
UniRef100_Q8LFH9 Hypothetical protein [Arabidopsis thaliana] 71 1e-11
UniRef100_Q8S0N3 Putative mutator-like transposase [Oryza sativa] 68 1e-10
UniRef100_Q9LNG5 F21D18.16 [Arabidopsis thaliana] 66 4e-10
UniRef100_Q69L69 Calcineurin-like phosphoesterase-like protein [... 66 5e-10
UniRef100_Q7XRQ6 OSJNBb0096E05.18 protein [Oryza sativa] 65 6e-10
UniRef100_Q9LD66 Similar to maize transposon MuDR mudrA protein ... 65 6e-10
UniRef100_Q60D76 Putative polyprotein [Oryza sativa] 65 6e-10
UniRef100_Q8GZW7 Putative transposable element [Oryza sativa] 65 6e-10
UniRef100_Q7X975 Putative Mutator protein [Oryza sativa] 65 6e-10
UniRef100_Q5TKN6 Hypothetical protein OJ1058_F05.1 [Oryza sativa] 65 8e-10
UniRef100_Q6I5K6 Hypothetical protein OSJNBb0088F07.13 [Oryza sa... 65 8e-10
UniRef100_Q5W6G1 Putative polyprotein [Oryza sativa] 65 1e-09
UniRef100_Q7XTA1 OSJNBa0008A08.9 protein [Oryza sativa] 64 1e-09
UniRef100_Q7X7Y4 OSJNBa0014F04.28 protein [Oryza sativa] 64 2e-09
UniRef100_Q9LMT7 F2H15.15 protein [Arabidopsis thaliana] 63 3e-09
UniRef100_Q69S26 Serine/threonine phosphatase PP7-related-like [... 63 3e-09
UniRef100_Q8LEM2 Hypothetical protein [Arabidopsis thaliana] 63 3e-09
UniRef100_Q94JI4 P0638D12.5 protein [Oryza sativa] 63 4e-09
>UniRef100_Q7XHQ1 Calcineurin-like phosphoesterase-like [Oryza sativa]
Length = 327
Score = 75.1 bits (183), Expect = 8e-13
Identities = 51/156 (32%), Positives = 80/156 (50%), Gaps = 9/156 (5%)
Query: 25 DHGLLIAFSERWHSETSSFHLPVGEMTITLDDVSCLLHIPVGGNLLFHESLSIHQGTQYL 84
D L+ A ERW +T++FHLPVGEMTITL DVSCL +P+ G + ++ +
Sbjct: 67 DKYLISALVERWRPKTNTFHLPVGEMTITLQDVSCLWGLPIHGRPITGQADG--SWVDMI 124
Query: 85 VNYLGLEFEES-AAQTKRLRSAHITYDTL--LSIYTSYLTEAKSYANQPEEEDSMEWYRT 141
LG+ EE Q KR + +T + SI S L + + E + WY
Sbjct: 125 ERLLGIPMEEQHMKQKKRKKEDDMTMVSYSRYSISLSKLRDRFRVMPKNATEREINWY-- 182
Query: 142 RCIRAFLLYLVGCTLFSDKAGNSCCVVYLKYFDDLT 177
RA +L ++G +F+D +G+ +YL++ +L+
Sbjct: 183 --TRALVLDIIGSMVFTDTSGDGVPAMYLQFMMNLS 216
>UniRef100_Q9SK32 Expressed protein [Arabidopsis thaliana]
Length = 509
Score = 71.2 bits (173), Expect = 1e-11
Identities = 47/171 (27%), Positives = 85/171 (49%), Gaps = 17/171 (9%)
Query: 6 NVIRASGLYLLLETNYGQVDHGLLIAFSERWHSETSSFHLPVGEMTITLDDVSCLLHIPV 65
N++ +G + +++ L+ A ERW ET++FHLP+GEMTITLD+V+ +L + +
Sbjct: 53 NLVDKAGFGYFRKIGPMSLNNSLISALVERWRRETNTFHLPLGEMTITLDEVALVLGLEI 112
Query: 66 GGNLLFHESLSIHQGTQYLVNYLGLEFEESAAQTKRLRSAHITYDTLLSIYTSYLTEAKS 125
G+ + + LG + +A K + + + + L ++
Sbjct: 113 DGDPIVGSKVGDEVAMDMCGRLLG---KLPSAANKEVNCSRVKLNWL----------KRT 159
Query: 126 YANQPEEEDSMEWYRTRC-IRAFLLYLVGCTLFSDKAGNSCCVVYLKYFDD 175
++ PE+ + +C RA+LLYL+G T+F+ G+ V YL F+D
Sbjct: 160 FSECPED---ASFDVVKCHTRAYLLYLIGSTIFATTDGDKVSVKYLPLFED 207
>UniRef100_Q8LFH9 Hypothetical protein [Arabidopsis thaliana]
Length = 509
Score = 71.2 bits (173), Expect = 1e-11
Identities = 47/171 (27%), Positives = 85/171 (49%), Gaps = 17/171 (9%)
Query: 6 NVIRASGLYLLLETNYGQVDHGLLIAFSERWHSETSSFHLPVGEMTITLDDVSCLLHIPV 65
N++ +G + +++ L+ A ERW ET++FHLP+GEMTITLD+V+ +L + +
Sbjct: 53 NLVDKAGFGYFRKIGPMSLNNSLISALVERWRRETNTFHLPLGEMTITLDEVALVLGLEI 112
Query: 66 GGNLLFHESLSIHQGTQYLVNYLGLEFEESAAQTKRLRSAHITYDTLLSIYTSYLTEAKS 125
G+ + + LG + +A K + + + + L ++
Sbjct: 113 DGDPIVGSKVDDEVAMDMCGRLLG---KLPSAANKEVNCSRVKLNWL----------KRT 159
Query: 126 YANQPEEEDSMEWYRTRC-IRAFLLYLVGCTLFSDKAGNSCCVVYLKYFDD 175
++ PE+ + +C RA+LLYL+G T+F+ G+ V YL F+D
Sbjct: 160 FSECPED---ASFDVVKCHTRAYLLYLIGSTIFATTDGDKVSVKYLPLFED 207
>UniRef100_Q8S0N3 Putative mutator-like transposase [Oryza sativa]
Length = 1556
Score = 67.8 bits (164), Expect = 1e-10
Identities = 44/147 (29%), Positives = 73/147 (48%), Gaps = 13/147 (8%)
Query: 29 LIAFSERWHSETSSFHLPVGEMTITLDDVSCLLHIPVGGNLLFHESLSIHQGTQYLVNYL 88
L A +RW ET +FHLP GE+T+TL+DV+ +L +P+ G + ++ S + + YL
Sbjct: 966 LTALVDRWRPETHTFHLPCGELTVTLEDVAMILGLPIRGQAVTGDTAS-GNWRERVEEYL 1024
Query: 89 GLEFEESAAQTKRLRSAHITYDTLLSIYTSYLTEAKSYANQPEEEDSMEWYRTRCIRAFL 148
GLE + ++ +++ + L + ++ P E D E R RA++
Sbjct: 1025 GLEPPVAPDGQRQTKTSGVPLSWLRA----------NFGQCPAEAD--EAIVQRYCRAYV 1072
Query: 149 LYLVGCTLFSDKAGNSCCVVYLKYFDD 175
LY+ G LF D GN ++L D
Sbjct: 1073 LYIFGSILFPDSGGNMASWMWLPLLAD 1099
>UniRef100_Q9LNG5 F21D18.16 [Arabidopsis thaliana]
Length = 1340
Score = 66.2 bits (160), Expect = 4e-10
Identities = 49/169 (28%), Positives = 80/169 (46%), Gaps = 21/169 (12%)
Query: 12 GLYLLLETNYGQVDHGLLIAFSERWHSETSSFHLPVGEMTITLDDVSCLLHIPVGGNLLF 71
GLY + + + Q+D+ L+ A ERW ET +FHLP GE+T+TL DV+ LL + V G
Sbjct: 68 GLYGVYKVAFIQLDYALITALVERWRPETHTFHLPAGEITVTLQDVNILLGLRVDGP--- 124
Query: 72 HESLSIHQGTQYLVNYLGLEFEESAAQTKRLRSAHITYDTLLSIYTSYLTEAKSYANQPE 131
++ T+Y L + K L +H++ L +++ N P
Sbjct: 125 ----AVTGSTKYNWADLCEDLLGHRPGPKDLHGSHVSLAWL----------RENFRNLPA 170
Query: 132 EEDSMEWYRTRC-IRAFLLYLVGCTLFSDKAGNSCCVVYLKYFDDLTTV 179
+ D + +C RAF+L L+ L+ DK+ + + +L D V
Sbjct: 171 DPDEV---TLKCHTRAFVLALMSGFLYGDKSKHDVALTFLPLLRDFDEV 216
>UniRef100_Q69L69 Calcineurin-like phosphoesterase-like protein [Oryza sativa]
Length = 766
Score = 65.9 bits (159), Expect = 5e-10
Identities = 51/157 (32%), Positives = 72/157 (45%), Gaps = 19/157 (12%)
Query: 23 QVDHGLLIAFSERWHSETSSFHLPVGEMTITLDDVSCLLHIPVGGNLLFHESLSIHQGTQ 82
Q+D L+ A ERW ET SFHL GEMT+TL DVS LL +P+ G + S + H Q
Sbjct: 128 QLDKALITALVERWRPETHSFHLASGEMTVTLQDVSMLLALPIDGRPVC--STTDHDYAQ 185
Query: 83 YLVNYLGLEFEESAAQTKRLRSAHITYDTLLSIYTSYLTEAKSYANQPEEEDSMEWYRTR 142
+++ LG + + K + Y K + PE D R
Sbjct: 186 MVIDCLGHDPRGPSMPGKS--------------FLHYKWLKKHFYELPEGTDDQT--VER 229
Query: 143 CIRAFLLYLVGCTLFSDKAGNSCCVVYLKYFDDLTTV 179
+RA++L L+ LF D G ++YL DL+ V
Sbjct: 230 HVRAYILSLLCGVLFPDGTGR-MSLIYLPLIADLSLV 265
>UniRef100_Q7XRQ6 OSJNBb0096E05.18 protein [Oryza sativa]
Length = 567
Score = 65.5 bits (158), Expect = 6e-10
Identities = 43/147 (29%), Positives = 73/147 (49%), Gaps = 13/147 (8%)
Query: 29 LIAFSERWHSETSSFHLPVGEMTITLDDVSCLLHIPVGGNLLFHESLSIHQGTQYLVNYL 88
L A +RW ET +FHLP GE+T+TL+DV+ +L +P+ G + ++ S + + YL
Sbjct: 14 LTALVDRWRPETHTFHLPCGELTVTLEDVAMILGLPIRGQAVTGDTAS-GNWRERVEEYL 72
Query: 89 GLEFEESAAQTKRLRSAHITYDTLLSIYTSYLTEAKSYANQPEEEDSMEWYRTRCIRAFL 148
GLE + ++ +++ + L + ++ P E D E R RA++
Sbjct: 73 GLEPPVAPDGQRQTKTSGVPLSWLRA----------NFGQCPAEAD--EATVQRYCRAYV 120
Query: 149 LYLVGCTLFSDKAGNSCCVVYLKYFDD 175
LY+ G LF D G+ ++L D
Sbjct: 121 LYIFGSILFPDSGGDMASWMWLPLLAD 147
>UniRef100_Q9LD66 Similar to maize transposon MuDR mudrA protein [Oryza sativa]
Length = 1230
Score = 65.5 bits (158), Expect = 6e-10
Identities = 43/147 (29%), Positives = 73/147 (49%), Gaps = 13/147 (8%)
Query: 29 LIAFSERWHSETSSFHLPVGEMTITLDDVSCLLHIPVGGNLLFHESLSIHQGTQYLVNYL 88
L A +RW ET +FHLP GE+T+TL+DV+ +L +P+ G + ++ S + + YL
Sbjct: 576 LTALVDRWRPETHTFHLPCGELTVTLEDVAMILGLPIRGQAVTGDTAS-GNWRERVEEYL 634
Query: 89 GLEFEESAAQTKRLRSAHITYDTLLSIYTSYLTEAKSYANQPEEEDSMEWYRTRCIRAFL 148
GLE + ++ +++ + L + ++ P E D E R RA++
Sbjct: 635 GLEPPVAPDGQRQTKTSGVPLSWLRA----------NFGQCPAEAD--EATVQRYCRAYV 682
Query: 149 LYLVGCTLFSDKAGNSCCVVYLKYFDD 175
LY+ G LF D G+ ++L D
Sbjct: 683 LYIFGSILFPDSGGDMASWMWLPLLAD 709
>UniRef100_Q60D76 Putative polyprotein [Oryza sativa]
Length = 1754
Score = 65.5 bits (158), Expect = 6e-10
Identities = 43/147 (29%), Positives = 73/147 (49%), Gaps = 13/147 (8%)
Query: 29 LIAFSERWHSETSSFHLPVGEMTITLDDVSCLLHIPVGGNLLFHESLSIHQGTQYLVNYL 88
L A +RW ET +FHLP GE+T+TL+DV+ +L +P+ G + ++ S + + YL
Sbjct: 958 LTALVDRWRPETHTFHLPCGELTVTLEDVAMILGLPIRGQAVTGDTAS-GNWRERVEEYL 1016
Query: 89 GLEFEESAAQTKRLRSAHITYDTLLSIYTSYLTEAKSYANQPEEEDSMEWYRTRCIRAFL 148
GLE + ++ +++ + L + ++ P E D E R RA++
Sbjct: 1017 GLEPPVAPDGQRQTKTSGVPLSWLRA----------NFGQCPAEAD--EATVQRYCRAYV 1064
Query: 149 LYLVGCTLFSDKAGNSCCVVYLKYFDD 175
LY+ G LF D G+ ++L D
Sbjct: 1065 LYIFGSILFPDSGGDMASWMWLPLLAD 1091
>UniRef100_Q8GZW7 Putative transposable element [Oryza sativa]
Length = 663
Score = 65.5 bits (158), Expect = 6e-10
Identities = 52/169 (30%), Positives = 82/169 (47%), Gaps = 22/169 (13%)
Query: 8 IRASGLY---LLLETNYGQVDHGLLIAFSERWHSETSSFHLPVGEMTITLDDVSCLLHIP 64
+RA+GL L++ + L A +RW ET SFHLP GEMTITL DV+ +L +P
Sbjct: 52 LRAAGLLGVALVVSRGMPVFNAPALTALVDRWRPETHSFHLPSGEMTITLQDVAMILALP 111
Query: 65 VGGNLLF--HESLSIHQGTQYLVNYLGLEFEESAAQTKRLRSAHITYDTLLSIYTSYLTE 122
+ G+ + E+ H Q L + L E+ K+ +++ LS S+L
Sbjct: 112 LRGHAVTGRTETPGWHAQVQQLFG-IPLNIEQGQGGKKKQNGIPLSW---LSQNFSHL-- 165
Query: 123 AKSYANQPEEEDSMEWYRTRC-IRAFLLYLVGCTLFSDKAGNSCCVVYL 170
+D E +R C RA++L+L+G LF D G+ +++
Sbjct: 166 ----------DDDAEPWRVECYARAYILHLLGGVLFPDAGGDIASAIWI 204
>UniRef100_Q7X975 Putative Mutator protein [Oryza sativa]
Length = 1456
Score = 65.5 bits (158), Expect = 6e-10
Identities = 43/147 (29%), Positives = 73/147 (49%), Gaps = 13/147 (8%)
Query: 29 LIAFSERWHSETSSFHLPVGEMTITLDDVSCLLHIPVGGNLLFHESLSIHQGTQYLVNYL 88
L A +RW ET +FHLP GE+T+TL+DV+ +L +P+ G + ++ S + + YL
Sbjct: 897 LTALVDRWRPETHTFHLPCGELTVTLEDVAMILGLPIRGQAVTGDTAS-GNWRERVEEYL 955
Query: 89 GLEFEESAAQTKRLRSAHITYDTLLSIYTSYLTEAKSYANQPEEEDSMEWYRTRCIRAFL 148
GLE + ++ +++ + L + ++ P E D E R RA++
Sbjct: 956 GLEPPVAPDGQRQTKTSGVPLSWLRA----------NFGQCPAEAD--EATVQRYCRAYV 1003
Query: 149 LYLVGCTLFSDKAGNSCCVVYLKYFDD 175
LY+ G LF D G+ ++L D
Sbjct: 1004 LYIFGSILFPDSGGDMASWMWLPLLAD 1030
>UniRef100_Q5TKN6 Hypothetical protein OJ1058_F05.1 [Oryza sativa]
Length = 725
Score = 65.1 bits (157), Expect = 8e-10
Identities = 48/151 (31%), Positives = 74/151 (48%), Gaps = 19/151 (12%)
Query: 29 LIAFSERWHSETSSFHLPVGEMTITLDDVSCLLHIPVGGNLLF--HESLSIHQGTQYLVN 86
L A +RW ET SFHLP GEMTITL DV+ +L +P+ G+ + E+ H Q L
Sbjct: 124 LTALVDRWRPETHSFHLPSGEMTITLQDVAMILALPLRGHAVTGRTETPGWHAQVQQLFG 183
Query: 87 YLGLEFEESAAQTKRLRSAHITYDTLLSIYTSYLTEAKSYANQPEEEDSMEWYRTRC-IR 145
+ L E+ K+ +++ LS S+L +D E +R C R
Sbjct: 184 -IPLNIEQGQGGKKKQNGIPLSW---LSQNFSHL------------DDDAEPWRVECYAR 227
Query: 146 AFLLYLVGCTLFSDKAGNSCCVVYLKYFDDL 176
A++L+L+G LF D G+ +++ +L
Sbjct: 228 AYILHLLGGVLFPDAGGDIASAIWIPLVTNL 258
>UniRef100_Q6I5K6 Hypothetical protein OSJNBb0088F07.13 [Oryza sativa]
Length = 1564
Score = 65.1 bits (157), Expect = 8e-10
Identities = 48/151 (31%), Positives = 74/151 (48%), Gaps = 19/151 (12%)
Query: 29 LIAFSERWHSETSSFHLPVGEMTITLDDVSCLLHIPVGGNLLF--HESLSIHQGTQYLVN 86
L A +RW ET SFHLP GEMTITL DV+ +L +P+ G+ + E+ H Q L
Sbjct: 963 LTALVDRWRPETHSFHLPSGEMTITLQDVAMILALPLRGHAVTGRTETPGWHAQVQQLFG 1022
Query: 87 YLGLEFEESAAQTKRLRSAHITYDTLLSIYTSYLTEAKSYANQPEEEDSMEWYRTRC-IR 145
+ L E+ K+ +++ LS S+L +D E +R C R
Sbjct: 1023 -IPLNIEQGQGGKKKQNGIPLSW---LSQNFSHL------------DDDAEPWRVECYAR 1066
Query: 146 AFLLYLVGCTLFSDKAGNSCCVVYLKYFDDL 176
A++L+L+G LF D G+ +++ +L
Sbjct: 1067 AYILHLLGGVLFPDAGGDIASAIWIPLVTNL 1097
>UniRef100_Q5W6G1 Putative polyprotein [Oryza sativa]
Length = 1567
Score = 64.7 bits (156), Expect = 1e-09
Identities = 52/169 (30%), Positives = 81/169 (47%), Gaps = 22/169 (13%)
Query: 8 IRASGLY---LLLETNYGQVDHGLLIAFSERWHSETSSFHLPVGEMTITLDDVSCLLHIP 64
+RA+GL L++ + L A RW ET SFHLP GEMTITL DV+ +L +P
Sbjct: 1002 LRAAGLLRVALVVSRGMPVFNAPALTALVNRWRPETHSFHLPSGEMTITLQDVAMILALP 1061
Query: 65 VGGNLLF--HESLSIHQGTQYLVNYLGLEFEESAAQTKRLRSAHITYDTLLSIYTSYLTE 122
+ G+ + E+ H Q L + L E+ K+ +++ LS S+L
Sbjct: 1062 LRGHAVTGRTETPGWHAQVQQLFG-IPLNIEQGQGGKKKQNGIPLSW---LSQNFSHL-- 1115
Query: 123 AKSYANQPEEEDSMEWYRTRC-IRAFLLYLVGCTLFSDKAGNSCCVVYL 170
+D E +R C RA++L+L+G LF D G+ +++
Sbjct: 1116 ----------DDDAEPWRAECYARAYILHLLGGVLFPDAGGDIASAIWI 1154
>UniRef100_Q7XTA1 OSJNBa0008A08.9 protein [Oryza sativa]
Length = 1560
Score = 64.3 bits (155), Expect = 1e-09
Identities = 41/147 (27%), Positives = 71/147 (47%), Gaps = 13/147 (8%)
Query: 29 LIAFSERWHSETSSFHLPVGEMTITLDDVSCLLHIPVGGNLLFHESLSIHQGTQYLVNYL 88
L A +RW ET +FHLP GE+T+TL+DV+ +L +P+ G + ++ S + + YL
Sbjct: 877 LTALVDRWRPETHTFHLPCGELTVTLEDVAMILGLPIRGQAVTSDTAS-GNWRERVEEYL 935
Query: 89 GLEFEESAAQTKRLRSAHITYDTLLSIYTSYLTEAKSYANQPEEEDSMEWYRTRCIRAFL 148
G+E + ++ +++ + L + + EA Q R RA++
Sbjct: 936 GVEPPVAPDGQRQTKTSGVPLSWLRANFGQCPAEADKATVQ------------RYCRAYV 983
Query: 149 LYLVGCTLFSDKAGNSCCVVYLKYFDD 175
LY+ G LF D G+ ++L D
Sbjct: 984 LYIFGSILFPDSGGDMASWMWLPLLAD 1010
>UniRef100_Q7X7Y4 OSJNBa0014F04.28 protein [Oryza sativa]
Length = 1575
Score = 63.9 bits (154), Expect = 2e-09
Identities = 51/169 (30%), Positives = 82/169 (48%), Gaps = 22/169 (13%)
Query: 8 IRASGLY---LLLETNYGQVDHGLLIAFSERWHSETSSFHLPVGEMTITLDDVSCLLHIP 64
+RA+GL L++ + L A +RW ET SFHLP GEMTITL DV+ +L +P
Sbjct: 944 LRAAGLLGVALVVSRGMPVFNAPALTALVDRWRPETHSFHLPSGEMTITLQDVAMILALP 1003
Query: 65 VGGNLLF--HESLSIHQGTQYLVNYLGLEFEESAAQTKRLRSAHITYDTLLSIYTSYLTE 122
+ G+ + ++ H Q L + L E+ K+ +++ LS S+L
Sbjct: 1004 LRGHAVTGRTDTPGWHAQVQQLFG-IPLNIEQGQGGKKKQNGIPLSW---LSQNFSHL-- 1057
Query: 123 AKSYANQPEEEDSMEWYRTRC-IRAFLLYLVGCTLFSDKAGNSCCVVYL 170
+D E +R C RA++L+L+G LF D G+ +++
Sbjct: 1058 ----------DDDAEPWRVECYARAYILHLLGEVLFPDAGGDIASAIWI 1096
>UniRef100_Q9LMT7 F2H15.15 protein [Arabidopsis thaliana]
Length = 478
Score = 63.2 bits (152), Expect = 3e-09
Identities = 46/153 (30%), Positives = 75/153 (48%), Gaps = 20/153 (13%)
Query: 24 VDHGLLIAFSERWHSETSSFHLPVGEMTITLDDVSCLLHIPVGGNLLFHESLSIHQGTQY 83
+++ L+ A ERW ET++FH P GEMTITLD+VS +L + V G + +Q
Sbjct: 62 LNNSLISALVERWRRETNTFHFPCGEMTITLDEVSLILGLAVDGKPVVGVKEKDEDPSQV 121
Query: 84 LVNYLGLEFEESAAQTKRLRSAHITYDTLLSIYTSYLTEAKSYANQPEEEDSME-WYRTR 142
+ LG +L ++ + + + + +S+A P+ E Y T
Sbjct: 122 CLRLLG-----------KLPKGELSGNRVTAKWLK-----ESFAECPKGATMKEIEYHT- 164
Query: 143 CIRAFLLYLVGCTLFSDKAGNSCCVVYLKYFDD 175
RA+L+Y+VG T+F+ + V YL F+D
Sbjct: 165 --RAYLIYIVGSTIFATTDPSKISVDYLILFED 195
>UniRef100_Q69S26 Serine/threonine phosphatase PP7-related-like [Oryza sativa]
Length = 619
Score = 63.2 bits (152), Expect = 3e-09
Identities = 56/174 (32%), Positives = 81/174 (46%), Gaps = 19/174 (10%)
Query: 7 VIRASGLYLLLETNYGQVDHGLLIAFSERWHSETSSFHLPVGEMTITLDDVSCLLHIPVG 66
++ ASGL L T +D LL AF ERW++ET++ H EM +L DVS +L IPV
Sbjct: 54 LVNASGLGHLALTTGFTIDRSLLTAFCERWNNETNTAHFMGFEMAPSLRDVSYILGIPVT 113
Query: 67 GNLLFHESLSIHQGTQYLVNYLGLEFEESAAQTKRLRSAHITYDTLLSIYTSYLTEAKSY 126
G+++ E + + +++LG ES ++L L+ + Y + +
Sbjct: 114 GHVVTAEPIGDEAVRRMCLHFLG----ESPGNGEQLCG-------LIRLTWLY----RKF 158
Query: 127 ANQPEEEDSME-WYRTRCIRAFLLYLVGCTLFSDKAGNSCCVVYLKYFDDLTTV 179
PE E Y T RA+LLYLVG TLF D YL D +
Sbjct: 159 HQLPENPTINEIAYST---RAYLLYLVGSTLFPDTMRGFVSPRYLPLLADFRKI 209
>UniRef100_Q8LEM2 Hypothetical protein [Arabidopsis thaliana]
Length = 478
Score = 63.2 bits (152), Expect = 3e-09
Identities = 46/153 (30%), Positives = 75/153 (48%), Gaps = 20/153 (13%)
Query: 24 VDHGLLIAFSERWHSETSSFHLPVGEMTITLDDVSCLLHIPVGGNLLFHESLSIHQGTQY 83
+++ L+ A ERW ET++FH P GEMTITLD+VS +L + V G + +Q
Sbjct: 62 LNNSLISALVERWRRETNTFHFPCGEMTITLDEVSLILGLAVDGKPVVGVKEKDEDPSQV 121
Query: 84 LVNYLGLEFEESAAQTKRLRSAHITYDTLLSIYTSYLTEAKSYANQPEEEDSME-WYRTR 142
+ LG +L ++ + + + + +S+A P+ E Y T
Sbjct: 122 CLKLLG-----------KLPKGELSGNRVTAKWLK-----ESFAECPKGATMKEIEYHT- 164
Query: 143 CIRAFLLYLVGCTLFSDKAGNSCCVVYLKYFDD 175
RA+L+Y+VG T+F+ + V YL F+D
Sbjct: 165 --RAYLIYIVGSTIFATTDPSKISVDYLILFED 195
>UniRef100_Q94JI4 P0638D12.5 protein [Oryza sativa]
Length = 761
Score = 62.8 bits (151), Expect = 4e-09
Identities = 49/157 (31%), Positives = 71/157 (45%), Gaps = 19/157 (12%)
Query: 23 QVDHGLLIAFSERWHSETSSFHLPVGEMTITLDDVSCLLHIPVGGNLLFHESLSIHQGTQ 82
Q+D L+ A ERW ET SFHL GEM +TL DV+ LL +P+ G + S + H Q
Sbjct: 145 QLDKALITALVERWRPETHSFHLASGEMAVTLQDVAMLLALPIDGRPVC--STTDHDYAQ 202
Query: 83 YLVNYLGLEFEESAAQTKRLRSAHITYDTLLSIYTSYLTEAKSYANQPEEEDSMEWYRTR 142
+++ LG + + K + Y K + PE D R
Sbjct: 203 MVIDCLGHDPRGPSMPGKS--------------FLHYKWLKKHFYELPEGADDQT--VER 246
Query: 143 CIRAFLLYLVGCTLFSDKAGNSCCVVYLKYFDDLTTV 179
+RA++L L+ LF D G ++YL DL+ V
Sbjct: 247 HVRAYILSLLCGVLFPDGTGR-MSLIYLPLIADLSRV 282
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.322 0.137 0.426
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 300,845,515
Number of Sequences: 2790947
Number of extensions: 11641697
Number of successful extensions: 21929
Number of sequences better than 10.0: 143
Number of HSP's better than 10.0 without gapping: 128
Number of HSP's successfully gapped in prelim test: 15
Number of HSP's that attempted gapping in prelim test: 21749
Number of HSP's gapped (non-prelim): 156
length of query: 180
length of database: 848,049,833
effective HSP length: 119
effective length of query: 61
effective length of database: 515,927,140
effective search space: 31471555540
effective search space used: 31471555540
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 70 (31.6 bits)
Medicago: description of AC146864.6


