

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146862.7 - phase: 0
(198 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
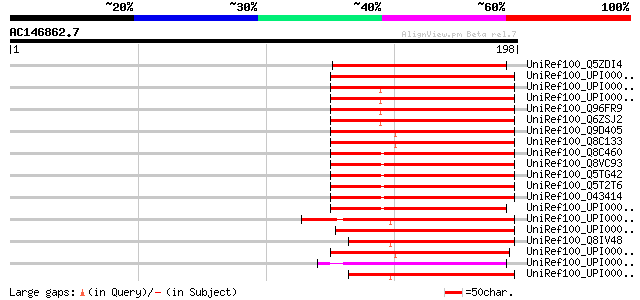
Sequences producing significant alignments: (bits) Value
UniRef100_Q5ZDI4 Hypothetical protein P0686E09.33 [Oryza sativa] 88 1e-16
UniRef100_UPI0000019BBC UPI0000019BBC UniRef100 entry 82 6e-15
UniRef100_UPI00001410D2 UPI00001410D2 UniRef100 entry 79 9e-14
UniRef100_UPI000036A6A9 UPI000036A6A9 UniRef100 entry 79 9e-14
UniRef100_Q96FR9 Hypothetical protein MGC16943 [Homo sapiens] 79 9e-14
UniRef100_Q6ZSJ2 Hypothetical protein FLJ45480 [Homo sapiens] 79 9e-14
UniRef100_Q9D405 Mus musculus adult male testis cDNA, RIKEN full... 72 8e-12
UniRef100_Q8C133 Mus musculus 10 days neonate skin cDNA, RIKEN f... 72 8e-12
UniRef100_Q8C460 Mus musculus 12 days embryo spinal cord cDNA, R... 72 1e-11
UniRef100_Q8VC93 Prnpip1 protein [Mus musculus] 72 1e-11
UniRef100_Q5TG42 OTTHUMP00000046440 [Homo sapiens] 72 1e-11
UniRef100_Q5T2T6 Prion protein interacting protein [Homo sapiens] 72 1e-11
UniRef100_O43414 Hypothetical protein [Homo sapiens] 72 1e-11
UniRef100_UPI00003AEACE UPI00003AEACE UniRef100 entry 70 2e-11
UniRef100_UPI000036E1C0 UPI000036E1C0 UniRef100 entry 70 4e-11
UniRef100_UPI00003AF359 UPI00003AF359 UniRef100 entry 70 4e-11
UniRef100_Q8IV48 3'-5' exonuclease ERI1 [Homo sapiens] 69 5e-11
UniRef100_UPI00001CEA21 UPI00001CEA21 UniRef100 entry 69 7e-11
UniRef100_UPI000031FBF7 UPI000031FBF7 UniRef100 entry 69 9e-11
UniRef100_UPI00003AD7F4 UPI00003AD7F4 UniRef100 entry 67 3e-10
>UniRef100_Q5ZDI4 Hypothetical protein P0686E09.33 [Oryza sativa]
Length = 304
Score = 88.2 bits (217), Expect = 1e-16
Identities = 38/68 (55%), Positives = 54/68 (78%)
Query: 127 EFQNFVVIDFEATCDKDKNPHPQEIIEFPSVIVSSVTGQLEACFQTYVRPTCNQHLSDFC 186
EF +FVV+DFEATC++ + +PQEIIEFP+V+V + TG+L + F+ YVRP + L+DFC
Sbjct: 26 EFDHFVVVDFEATCERGRRIYPQEIIEFPAVLVDAATGRLVSAFRAYVRPRHHPRLTDFC 85
Query: 187 KDLTGIQQ 194
++LTGI Q
Sbjct: 86 RELTGIAQ 93
>UniRef100_UPI0000019BBC UPI0000019BBC UniRef100 entry
Length = 215
Score = 82.4 bits (202), Expect = 6e-15
Identities = 38/72 (52%), Positives = 56/72 (77%)
Query: 126 QEFQNFVVIDFEATCDKDKNPHPQEIIEFPSVIVSSVTGQLEACFQTYVRPTCNQHLSDF 185
Q F + +VIDFE+TC ++KN + QEIIEFP+V++++ TG++E+ F TYV+P + LS F
Sbjct: 1 QIFSHLIVIDFESTCWREKNNYSQEIIEFPAVLLNTCTGEVESEFHTYVQPQEHPTLSGF 60
Query: 186 CKDLTGIQQIQV 197
C +LTGI Q+QV
Sbjct: 61 CTELTGITQMQV 72
>UniRef100_UPI00001410D2 UPI00001410D2 UniRef100 entry
Length = 328
Score = 78.6 bits (192), Expect = 9e-14
Identities = 39/73 (53%), Positives = 55/73 (74%), Gaps = 1/73 (1%)
Query: 126 QEFQNFVVIDFEATCDKD-KNPHPQEIIEFPSVIVSSVTGQLEACFQTYVRPTCNQHLSD 184
Q F +VIDFE+TC D K+ H QEIIEFP+V++++ TGQ+++ FQ YV+P + LS+
Sbjct: 32 QLFDYLIVIDFESTCWNDGKHHHSQEIIEFPAVLLNTSTGQIDSEFQAYVQPQEHPILSE 91
Query: 185 FCKDLTGIQQIQV 197
FC +LTGI+Q QV
Sbjct: 92 FCMELTGIKQAQV 104
>UniRef100_UPI000036A6A9 UPI000036A6A9 UniRef100 entry
Length = 328
Score = 78.6 bits (192), Expect = 9e-14
Identities = 39/73 (53%), Positives = 55/73 (74%), Gaps = 1/73 (1%)
Query: 126 QEFQNFVVIDFEATCDKD-KNPHPQEIIEFPSVIVSSVTGQLEACFQTYVRPTCNQHLSD 184
Q F +VIDFE+TC D K+ H QEIIEFP+V++++ TGQ+++ FQ YV+P + LS+
Sbjct: 32 QLFDYLIVIDFESTCWNDGKHHHSQEIIEFPAVLLNTSTGQIDSEFQAYVQPQEHPILSE 91
Query: 185 FCKDLTGIQQIQV 197
FC +LTGI+Q QV
Sbjct: 92 FCMELTGIKQAQV 104
>UniRef100_Q96FR9 Hypothetical protein MGC16943 [Homo sapiens]
Length = 328
Score = 78.6 bits (192), Expect = 9e-14
Identities = 39/73 (53%), Positives = 55/73 (74%), Gaps = 1/73 (1%)
Query: 126 QEFQNFVVIDFEATCDKD-KNPHPQEIIEFPSVIVSSVTGQLEACFQTYVRPTCNQHLSD 184
Q F +VIDFE+TC D K+ H QEIIEFP+V++++ TGQ+++ FQ YV+P + LS+
Sbjct: 32 QLFDYLIVIDFESTCWNDGKHHHSQEIIEFPAVLLNTSTGQIDSEFQAYVQPQEHPILSE 91
Query: 185 FCKDLTGIQQIQV 197
FC +LTGI+Q QV
Sbjct: 92 FCMELTGIKQAQV 104
>UniRef100_Q6ZSJ2 Hypothetical protein FLJ45480 [Homo sapiens]
Length = 279
Score = 78.6 bits (192), Expect = 9e-14
Identities = 39/73 (53%), Positives = 55/73 (74%), Gaps = 1/73 (1%)
Query: 126 QEFQNFVVIDFEATCDKD-KNPHPQEIIEFPSVIVSSVTGQLEACFQTYVRPTCNQHLSD 184
Q F +VIDFE+TC D K+ H QEIIEFP+V++++ TGQ+++ FQ YV+P + LS+
Sbjct: 32 QLFDYLIVIDFESTCWNDGKHHHSQEIIEFPAVLLNTSTGQIDSEFQAYVQPQEHPILSE 91
Query: 185 FCKDLTGIQQIQV 197
FC +LTGI+Q QV
Sbjct: 92 FCMELTGIKQAQV 104
>UniRef100_Q9D405 Mus musculus adult male testis cDNA, RIKEN full-length enriched
library, clone:4933424N09 product:hypothetical
Exonuclease containing protein, full insert sequence
[Mus musculus]
Length = 274
Score = 72.0 bits (175), Expect = 8e-12
Identities = 34/73 (46%), Positives = 52/73 (70%), Gaps = 1/73 (1%)
Query: 126 QEFQNFVVIDFEATCDKDKNPHPQ-EIIEFPSVIVSSVTGQLEACFQTYVRPTCNQHLSD 184
Q + +V+DFE+TC D H EIIEFP+V++++ TG++E+ F YV+P + LS+
Sbjct: 32 QLYAYLIVVDFESTCWNDGKHHSSPEIIEFPAVLLNTATGEIESEFHAYVQPQEHPILSE 91
Query: 185 FCKDLTGIQQIQV 197
FC +LTGI+Q+QV
Sbjct: 92 FCTELTGIKQVQV 104
>UniRef100_Q8C133 Mus musculus 10 days neonate skin cDNA, RIKEN full-length enriched
library, clone:4732486H02 product:hypothetical
Exonuclease containing protein, full insert sequence
[Mus musculus]
Length = 688
Score = 72.0 bits (175), Expect = 8e-12
Identities = 34/73 (46%), Positives = 52/73 (70%), Gaps = 1/73 (1%)
Query: 126 QEFQNFVVIDFEATCDKDKNPHPQ-EIIEFPSVIVSSVTGQLEACFQTYVRPTCNQHLSD 184
Q + +V+DFE+TC D H EIIEFP+V++++ TG++E+ F YV+P + LS+
Sbjct: 32 QLYAYLIVVDFESTCWNDGKHHSSPEIIEFPAVLLNTATGEIESEFHAYVQPQEHPILSE 91
Query: 185 FCKDLTGIQQIQV 197
FC +LTGI+Q+QV
Sbjct: 92 FCTELTGIKQVQV 104
>UniRef100_Q8C460 Mus musculus 12 days embryo spinal cord cDNA, RIKEN full-length
enriched library, clone:C530033L24 product:prion protein
interacting protein 1, full insert sequence [Mus
musculus]
Length = 337
Score = 71.6 bits (174), Expect = 1e-11
Identities = 35/72 (48%), Positives = 49/72 (67%), Gaps = 1/72 (1%)
Query: 126 QEFQNFVVIDFEATCDKDKNPHPQEIIEFPSVIVSSVTGQLEACFQTYVRPTCNQHLSDF 185
Q + F+V+DFEATCDK + HPQEIIEFP + ++ T ++E+ F YV+P + L+ F
Sbjct: 141 QRYHYFLVLDFEATCDKPQI-HPQEIIEFPILKLNGRTMEIESTFHMYVQPVVHPQLTPF 199
Query: 186 CKDLTGIQQIQV 197
C +LTGI Q V
Sbjct: 200 CTELTGIIQAMV 211
>UniRef100_Q8VC93 Prnpip1 protein [Mus musculus]
Length = 264
Score = 71.6 bits (174), Expect = 1e-11
Identities = 35/72 (48%), Positives = 49/72 (67%), Gaps = 1/72 (1%)
Query: 126 QEFQNFVVIDFEATCDKDKNPHPQEIIEFPSVIVSSVTGQLEACFQTYVRPTCNQHLSDF 185
Q + F+V+DFEATCDK + HPQEIIEFP + ++ T ++E+ F YV+P + L+ F
Sbjct: 68 QRYHYFLVLDFEATCDKPQI-HPQEIIEFPILKLNGRTMEIESTFHMYVQPVVHPQLTPF 126
Query: 186 CKDLTGIQQIQV 197
C +LTGI Q V
Sbjct: 127 CTELTGIIQAMV 138
>UniRef100_Q5TG42 OTTHUMP00000046440 [Homo sapiens]
Length = 337
Score = 71.6 bits (174), Expect = 1e-11
Identities = 35/72 (48%), Positives = 49/72 (67%), Gaps = 1/72 (1%)
Query: 126 QEFQNFVVIDFEATCDKDKNPHPQEIIEFPSVIVSSVTGQLEACFQTYVRPTCNQHLSDF 185
Q + F+V+DFEATCDK + HPQEIIEFP + ++ T ++E+ F YV+P + L+ F
Sbjct: 141 QRYHYFLVLDFEATCDKPQI-HPQEIIEFPILKLNGRTMEIESTFHMYVQPVVHPQLTPF 199
Query: 186 CKDLTGIQQIQV 197
C +LTGI Q V
Sbjct: 200 CTELTGIIQAMV 211
>UniRef100_Q5T2T6 Prion protein interacting protein [Homo sapiens]
Length = 250
Score = 71.6 bits (174), Expect = 1e-11
Identities = 35/72 (48%), Positives = 49/72 (67%), Gaps = 1/72 (1%)
Query: 126 QEFQNFVVIDFEATCDKDKNPHPQEIIEFPSVIVSSVTGQLEACFQTYVRPTCNQHLSDF 185
Q + F+V+DFEATCDK + HPQEIIEFP + ++ T ++E+ F YV+P + L+ F
Sbjct: 139 QRYHYFLVLDFEATCDKPQI-HPQEIIEFPILKLNGRTMEIESTFHMYVQPVVHPQLTPF 197
Query: 186 CKDLTGIQQIQV 197
C +LTGI Q V
Sbjct: 198 CTELTGIIQAMV 209
>UniRef100_O43414 Hypothetical protein [Homo sapiens]
Length = 397
Score = 71.6 bits (174), Expect = 1e-11
Identities = 35/72 (48%), Positives = 49/72 (67%), Gaps = 1/72 (1%)
Query: 126 QEFQNFVVIDFEATCDKDKNPHPQEIIEFPSVIVSSVTGQLEACFQTYVRPTCNQHLSDF 185
Q + F+V+DFEATCDK + HPQEIIEFP + ++ T ++E+ F YV+P + L+ F
Sbjct: 201 QRYHYFLVLDFEATCDKPQI-HPQEIIEFPILKLNGRTMEIESTFHMYVQPVVHPQLTPF 259
Query: 186 CKDLTGIQQIQV 197
C +LTGI Q V
Sbjct: 260 CTELTGIIQAMV 271
>UniRef100_UPI00003AEACE UPI00003AEACE UniRef100 entry
Length = 259
Score = 70.5 bits (171), Expect = 2e-11
Identities = 34/69 (49%), Positives = 48/69 (69%), Gaps = 1/69 (1%)
Query: 126 QEFQNFVVIDFEATCDKDKNPHPQEIIEFPSVIVSSVTGQLEACFQTYVRPTCNQHLSDF 185
Q + F+V+DFEATCDK + HPQEIIEFP + ++ T ++E+ F YV+P + L+ F
Sbjct: 63 QRYHYFLVLDFEATCDKPQI-HPQEIIEFPILKLNGRTMEIESTFHMYVQPVVHPQLTPF 121
Query: 186 CKDLTGIQQ 194
C +LTGI Q
Sbjct: 122 CTELTGIIQ 130
>UniRef100_UPI000036E1C0 UPI000036E1C0 UniRef100 entry
Length = 313
Score = 69.7 bits (169), Expect = 4e-11
Identities = 38/84 (45%), Positives = 52/84 (61%), Gaps = 3/84 (3%)
Query: 115 MVSQGYPREQYQEFQNFVVIDFEATCDKDKNPH-PQEIIEFPSVIVSSVTGQLEACFQTY 173
M+ + + Y ++ +IDFEATC++ P EIIEFP V++++ T ++E FQ Y
Sbjct: 80 MLKESNCADSYYDY--ICIIDFEATCEEGNPPEFVHEIIEFPVVLLNTHTLEIEDTFQQY 137
Query: 174 VRPTCNQHLSDFCKDLTGIQQIQV 197
VRP N LSDFC LTGI Q QV
Sbjct: 138 VRPEINTQLSDFCISLTGITQDQV 161
>UniRef100_UPI00003AF359 UPI00003AF359 UniRef100 entry
Length = 594
Score = 69.7 bits (169), Expect = 4e-11
Identities = 33/70 (47%), Positives = 48/70 (68%)
Query: 128 FQNFVVIDFEATCDKDKNPHPQEIIEFPSVIVSSVTGQLEACFQTYVRPTCNQHLSDFCK 187
F +V+DFE+TC +D EIIEFP+V++++ TG++EA F T+V+P LS+FC
Sbjct: 1 FDFLLVLDFESTCWRDARQRRPEIIEFPAVLLNAATGRIEAEFHTFVQPQEQPVLSEFCT 60
Query: 188 DLTGIQQIQV 197
LTG+ Q QV
Sbjct: 61 TLTGVTQKQV 70
>UniRef100_Q8IV48 3'-5' exonuclease ERI1 [Homo sapiens]
Length = 348
Score = 69.3 bits (168), Expect = 5e-11
Identities = 36/66 (54%), Positives = 45/66 (67%), Gaps = 1/66 (1%)
Query: 133 VIDFEATCDKDKNPH-PQEIIEFPSVIVSSVTGQLEACFQTYVRPTCNQHLSDFCKDLTG 191
+IDFEATC++ P EIIEFP V++++ T ++E FQ YVRP N LSDFC LTG
Sbjct: 131 IIDFEATCEEGNPPEFVHEIIEFPVVLLNTHTLEIEDTFQQYVRPEINTQLSDFCISLTG 190
Query: 192 IQQIQV 197
I Q QV
Sbjct: 191 ITQDQV 196
>UniRef100_UPI00001CEA21 UPI00001CEA21 UniRef100 entry
Length = 729
Score = 68.9 bits (167), Expect = 7e-11
Identities = 32/71 (45%), Positives = 50/71 (70%), Gaps = 1/71 (1%)
Query: 126 QEFQNFVVIDFEATCDKDKNPHPQ-EIIEFPSVIVSSVTGQLEACFQTYVRPTCNQHLSD 184
Q + +V+DFE+TC D H EIIEFP+V++++ TG++E+ F YV+P + LS+
Sbjct: 35 QLYDYLIVVDFESTCWNDGKHHSSPEIIEFPAVLLNTATGEIESEFHAYVQPQEHPILSE 94
Query: 185 FCKDLTGIQQI 195
FC +LTGI+Q+
Sbjct: 95 FCTELTGIKQV 105
>UniRef100_UPI000031FBF7 UPI000031FBF7 UniRef100 entry
Length = 266
Score = 68.6 bits (166), Expect = 9e-11
Identities = 36/74 (48%), Positives = 44/74 (58%), Gaps = 5/74 (6%)
Query: 121 PREQYQEFQNFVVIDFEATCDKDKNPHPQEIIEFPSVIVSSVTGQLEACFQTYVRPTCNQ 180
PR Y VV+DFEATC++ + EIIEFP+V+V + L F YVRPT N
Sbjct: 17 PRVDY-----LVVVDFEATCERGDKAYEHEIIEFPAVLVRTADATLAGEFHAYVRPTENA 71
Query: 181 HLSDFCKDLTGIQQ 194
LS FC +LTGI Q
Sbjct: 72 TLSAFCTELTGIAQ 85
>UniRef100_UPI00003AD7F4 UPI00003AD7F4 UniRef100 entry
Length = 179
Score = 66.6 bits (161), Expect = 3e-10
Identities = 33/66 (50%), Positives = 46/66 (69%), Gaps = 1/66 (1%)
Query: 133 VIDFEATCDKDKNPH-PQEIIEFPSVIVSSVTGQLEACFQTYVRPTCNQHLSDFCKDLTG 191
V+DFEATC++ P EIIEFP V++++ T ++E FQ YV+P N LS+FC LTG
Sbjct: 6 VVDFEATCEEGNPPEFIHEIIEFPVVLLNTHTLEIEDTFQQYVKPEINPKLSEFCVGLTG 65
Query: 192 IQQIQV 197
I Q+Q+
Sbjct: 66 ITQLQL 71
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.320 0.135 0.441
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 383,715,670
Number of Sequences: 2790947
Number of extensions: 17067093
Number of successful extensions: 30704
Number of sequences better than 10.0: 112
Number of HSP's better than 10.0 without gapping: 57
Number of HSP's successfully gapped in prelim test: 55
Number of HSP's that attempted gapping in prelim test: 30557
Number of HSP's gapped (non-prelim): 143
length of query: 198
length of database: 848,049,833
effective HSP length: 121
effective length of query: 77
effective length of database: 510,345,246
effective search space: 39296583942
effective search space used: 39296583942
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 71 (32.0 bits)
Medicago: description of AC146862.7


