

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146862.24 - phase: 0
(580 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
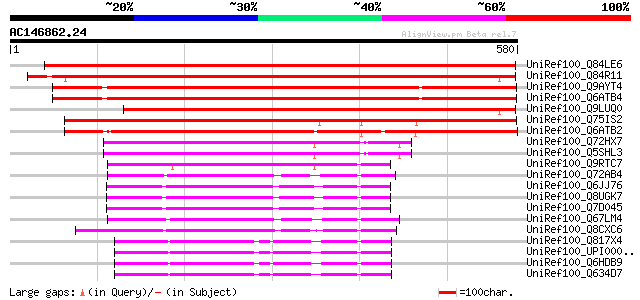
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q84LE6 RelA-SpoT like protein RSH4 [Nicotiana tabacum] 751 0.0
UniRef100_Q84R11 Hypothetical protein At3g17470 [Arabidopsis tha... 713 0.0
UniRef100_Q9AYT4 Chloroplast RelA homologue 2 [Oryza sativa] 665 0.0
UniRef100_Q6ATB4 Putative chloroplast RelA [Oryza sativa] 665 0.0
UniRef100_Q9LUQ0 Similarity to [Arabidopsis thaliana] 615 e-174
UniRef100_Q75IS2 Hypothetical protein OSJNBb0099P06.1 [Oryza sat... 573 e-162
UniRef100_Q6ATB2 Hypothetical protein OSJNBa0034O12.17 [Oryza sa... 535 e-150
UniRef100_Q72HX7 Guanosine-3',5'-bis(Diphosphate) 3'-pyrophospho... 206 2e-51
UniRef100_Q5SHL3 Guanosine-3',5'-bis(Diphosphate) 3'-pyrophospho... 205 3e-51
UniRef100_Q9RTC7 GTP pyrophosphokinase [Deinococcus radiodurans] 184 5e-45
UniRef100_Q72AB4 GTP pyrophosphokinase [Desulfovibrio vulgaris] 174 8e-42
UniRef100_Q6JJ76 RelA [Agrobacterium tumefaciens] 165 3e-39
UniRef100_Q8UGK7 GTP pyrophosphohydrolases/synthetases, RelA/Spo... 164 9e-39
UniRef100_Q7D045 AGR_C_1896p [Agrobacterium tumefaciens] 164 9e-39
UniRef100_Q67LM4 GTP pyrophosphokinase [Symbiobacterium thermoph... 162 3e-38
UniRef100_Q8CXC6 GTP pyrophosphokinase [Oceanobacillus iheyensis] 161 6e-38
UniRef100_Q817X4 GTP pyrophosphokinase [Bacillus cereus] 160 7e-38
UniRef100_UPI00003CB978 UPI00003CB978 UniRef100 entry 160 1e-37
UniRef100_Q6HDB9 GTP diphosphokinase [Bacillus thuringiensis] 160 1e-37
UniRef100_Q634D7 GTP diphosphokinase [Bacillus cereus] 160 1e-37
>UniRef100_Q84LE6 RelA-SpoT like protein RSH4 [Nicotiana tabacum]
Length = 552
Score = 751 bits (1938), Expect = 0.0
Identities = 374/543 (68%), Positives = 467/543 (85%), Gaps = 6/543 (1%)
Query: 41 PTRSTTLRWSSTPRA--CSVEVPGGGKMVIELVGAFNDLTERM--KVLSTSSSGLLFKSL 96
PT+ T+R S++ A + E PGG KMV+ELVGAFN+LTERM KVLSTSSS LLFK+L
Sbjct: 11 PTKIATVRVSASAEALVATPEQPGG-KMVVELVGAFNELTERMDNKVLSTSSSRLLFKAL 69
Query: 97 KLSIPVLQTSPLTPDGRSPLSKALSIAMLLADLQMDAEVISAGILREVLEVGELNLHEIR 156
KL IP+LQ+ PL PDGR+PLS+ALS+A++LADL+MDAEVIS G+LREVLE G ++++++R
Sbjct: 70 KLCIPILQSLPLAPDGRAPLSRALSVAVILADLRMDAEVISTGLLREVLEAGAISIYDVR 129
Query: 157 SQIGSATAHLLHESLRVKNFASRVDILDDENAAALRKFCLTYYDIRALILDLALKLDMMR 216
+IG++TAHLLHESLRVK+ + +V++LDD++A ALRKFCLTYYD+RAL+LDLA+KLDMMR
Sbjct: 130 DRIGTSTAHLLHESLRVKHMSLKVEVLDDDSATALRKFCLTYYDVRALVLDLAIKLDMMR 189
Query: 217 HLGHLPRYQQQIISLQVMKIYAPLAHAVGTNYISLELEDLSFQYLFPYSYLYVDTWLRSQ 276
HL +LPRY+QQ+ISL+VMK++APLAHA+GTN +SLELEDLSF+YLFPYSYLY+D WLRS
Sbjct: 190 HLDYLPRYRQQMISLEVMKLHAPLAHAIGTNLLSLELEDLSFRYLFPYSYLYLDAWLRSH 249
Query: 277 ETGGISLIDVYKDELLESLKSDPILAELVDDISVKGRYKSRYSTMKKLLKDGRRPEDVND 336
E+G LID+ K++LL+SL SDP+L ++V ISV+GRYKSRYSTMKKLL+DGR+ ++VND
Sbjct: 250 ESGNKPLIDICKEQLLQSLNSDPLLMQMVSKISVEGRYKSRYSTMKKLLRDGRKLDEVND 309
Query: 337 VLGLRVVLNPKSR-ENALEAGERACYRAHQIIQSMWKEIPSRTKDYISRPKGNGYRSLHM 395
+LGLRVVL P S + +E GE+ACYRA +++ S+WKEIPSR+KDYI RPK NGY+SLHM
Sbjct: 310 ILGLRVVLTPLSAGVDEMEIGEKACYRAREVVLSLWKEIPSRSKDYILRPKANGYKSLHM 369
Query: 396 AVDVSEIGRTRPLMEIQIRTTEMDRLAVGGMASHSLYKAGLTNPEEAKRLKTIMLAAAEL 455
AVD SE RTRPLMEIQIRT EMD LA GG ASH+LYK G+T+PEEAK LK IM+AAAE
Sbjct: 370 AVDTSERDRTRPLMEIQIRTAEMDILAAGGTASHALYKGGVTDPEEAKHLKAIMMAAAEF 429
Query: 456 AALRLKDFPSANHKGIEFDQRDRVFRLLDKNGDGKISIEELTEVIEELGAPGEDAHDMML 515
AALRLKD PS N KG+E D+R RVFRLLDKNGDGK+SI+EL EV+EELGAPG+DA +MM
Sbjct: 430 AALRLKDLPSGNPKGLETDKRGRVFRLLDKNGDGKLSIDELMEVMEELGAPGDDAREMMQ 489
Query: 516 LLDSNSDGSLSSDEFQMFQKQVEMVRNLEDRDDEYKKILDEKLHMADDSGLIQVYNKEFG 575
LLDSNSDG LSSDEF +FQ Q+E +RN+EDRDD+Y+ +L+EKL MA+++GLI VY+KE G
Sbjct: 490 LLDSNSDGFLSSDEFDLFQDQIEFIRNIEDRDDQYQTLLNEKLQMANETGLIHVYSKEQG 549
Query: 576 NRL 578
+ L
Sbjct: 550 DTL 552
>UniRef100_Q84R11 Hypothetical protein At3g17470 [Arabidopsis thaliana]
Length = 583
Score = 713 bits (1840), Expect = 0.0
Identities = 374/573 (65%), Positives = 447/573 (77%), Gaps = 19/573 (3%)
Query: 21 PTTFSTTTHRRLLSLFFPTRPTRSTTLRWSSTPRACSVEVPG--------GGKMVIELVG 72
P S HRRL S+ R R RW+ + GGKMV+ELVG
Sbjct: 12 PRCRSQVLHRRLYSIQLIQRRRR----RWNPRSEVEDTAIESTARSPEAAGGKMVVELVG 67
Query: 73 AFNDLTERMKV--LSTSSSGLLFKSLKLSIPVLQTSPLTPDGRSPLSKALSIAMLLADLQ 130
AFN++TERM LSTSSS LLFK+LKLSIP+LQ+ PL DGRSPLSKALS++++LADLQ
Sbjct: 68 AFNEVTERMNSVWLSTSSSRLLFKALKLSIPILQSLPLASDGRSPLSKALSLSIILADLQ 127
Query: 131 MDAEVISAGILREVLEVGELNLHEIRSQIGSATAHLLHESLRVKNFASRVDILDDENAAA 190
MDAEVISA IL EV++ ++++E+R IG+ TAHLLHE RVKN +VD+LDDE AA+
Sbjct: 128 MDAEVISASILSEVVDANAISIYEVRDHIGTGTAHLLHEIFRVKNIPFKVDVLDDETAAS 187
Query: 191 LRKFCLTYYDIRALILDLALKLDMMRHLGHLPRYQQQIISLQVMKIYAPLAHAVGTNYIS 250
LRKF LTYYDIRA+I+DL KLD MRHL HLPRY+QQI+SL+V+KIY+PLAHAVG N++S
Sbjct: 188 LRKFYLTYYDIRAVIMDLVSKLDEMRHLDHLPRYRQQILSLEVLKIYSPLAHAVGANHLS 247
Query: 251 LELEDLSFQYLFPYSYLYVDTWLRSQETGGISLIDVYKDELLESLKSDPILAELVDDISV 310
LELED+SF+YLFP SY+Y+D+WLR E G LIDVYK++L SLK D +LAE+V+D+ +
Sbjct: 248 LELEDISFRYLFPCSYIYLDSWLRGHENGSKPLIDVYKEQLHRSLKDDLVLAEMVNDVYI 307
Query: 311 KGRYKSRYSTMKKLLKDGRRPEDVNDVLGLRVVLNPKSRENALEAGERACYRAHQIIQSM 370
KGRYKSRYS MKKLL+DGR+PE+VNDVLGLRV+L P S N +E GE+ACYR +II+S+
Sbjct: 308 KGRYKSRYSMMKKLLRDGRKPEEVNDVLGLRVILMPNSVVNDVEVGEKACYRTSEIIRSL 367
Query: 371 WKEIPSRTKDYISRPKGNGYRSLHMAVDVSEIGRTRPLMEIQIRTTEMDRLAVGGMASHS 430
WKEIP RTKDYI+RPK NGYRSLHMAVDVS+ + RPLMEIQIRT +MD A G ASHS
Sbjct: 368 WKEIPHRTKDYIARPKENGYRSLHMAVDVSDSDQIRPLMEIQIRTMDMDGSANAGTASHS 427
Query: 431 LYKAGLTNPEEAKRLKTIMLAAAELAALRLKDFPSANHKGIE--FDQRDRVFRLLDKNGD 488
LYK GLT+P+EAKRLK IMLAAA+LAA+RLKD S H+ + +QRDRVF LLDKNGD
Sbjct: 428 LYKGGLTDPKEAKRLKAIMLAAADLAAIRLKDISSNKHQSFKTTTNQRDRVFCLLDKNGD 487
Query: 489 GKISIEELTEVIEELGAPGEDAHDMMLLLDSNSDGSLSSDEFQMFQKQVEMVRNLEDRDD 548
G ISIEEL EV+EELGAPGEDA +MM LLDSNSDGSLSSDEF FQKQVE +R EDRD+
Sbjct: 488 GMISIEELMEVMEELGAPGEDAEEMMQLLDSNSDGSLSSDEFDTFQKQVEFMRKWEDRDN 547
Query: 549 EYKKILDEKLH---MADDSGLIQVYNKEFGNRL 578
EYK +LDEKLH D +GLIQ+YNKE +RL
Sbjct: 548 EYKSLLDEKLHDLPHQDTTGLIQLYNKELEDRL 580
>UniRef100_Q9AYT4 Chloroplast RelA homologue 2 [Oryza sativa]
Length = 583
Score = 665 bits (1716), Expect = 0.0
Identities = 335/532 (62%), Positives = 439/532 (81%), Gaps = 8/532 (1%)
Query: 50 SSTPRACSVEVPGGGKMVIELVGAFNDLTERMK--VLSTSSSGLLFKSLKLSIPVLQTSP 107
SS+ + S GGG++V ELVGAFN+LT RM + ++SSS LLF++LKL++P L+
Sbjct: 56 SSSSSSSSTPAEGGGRLVAELVGAFNELTGRMGEGLATSSSSRLLFRALKLALPALRDG- 114
Query: 108 LTPDGRSPLSKALSIAMLLADLQMDAEVISAGILREVLEVGELNLHEIRSQIGSATAHLL 167
DG L++AL+IA LADLQMDAEVISAGILRE L+ G +++ +++S+IG +TAHLL
Sbjct: 115 ---DGGRALARALAIAASLADLQMDAEVISAGILREALDAGAISMRDVKSEIGISTAHLL 171
Query: 168 HESLRVKNFASRVDILDDENAAALRKFCLTYYDIRALILDLALKLDMMRHLGHLPRYQQQ 227
HESLR+K+ S++D+LDDE+A+ALRKFCL+YYDIRA+IL+LALKLDMMRHL LPRY Q+
Sbjct: 172 HESLRLKHAPSKLDVLDDESASALRKFCLSYYDIRAVILELALKLDMMRHLNCLPRYLQR 231
Query: 228 IISLQVMKIYAPLAHAVGTNYISLELEDLSFQYLFPYSYLYVDTWLRSQETGGISLIDVY 287
I SL+V+KIYAPLAHAVG +SLELEDLSF+YLFP+SY ++D WLRSQET LID Y
Sbjct: 232 IKSLEVLKIYAPLAHAVGAGNLSLELEDLSFRYLFPHSYDHIDQWLRSQETENKLLIDSY 291
Query: 288 KDELLESLKSDPILAELVDDISVKGRYKSRYSTMKKLLKDGRRPEDVNDVLGLRVVLNPK 347
K++LL++LK D L+++V DIS++GRYKSR+STMKKL+KDGR+PE+VND+L LRV+L P+
Sbjct: 292 KEQLLQALKDDDELSQIVQDISIQGRYKSRFSTMKKLVKDGRKPEEVNDILALRVILEPR 351
Query: 348 SRENALEAGERACYRAHQIIQSMWKEIPSRTKDYISRPKGNGYRSLHMAVDVSEIGRTRP 407
++L+ G RAC+R H+IIQ+MWKE+P RTK+Y++RPK NGY+SLH+A+DVSE G+ RP
Sbjct: 352 CDGSSLDWGPRACHRTHEIIQAMWKEVPGRTKNYVTRPKENGYQSLHVAIDVSEPGKMRP 411
Query: 408 LMEIQIRTTEMDRLAVGGMASHSLYKAGLTNPEEAKRLKTIMLAAAELAALRLKDFPSAN 467
LMEIQIRT EM + AVGG ASHSLYK GLT+P EAKRLK IMLAAAELAA+RL+D P+++
Sbjct: 412 LMEIQIRTKEMHKFAVGGEASHSLYKGGLTDPGEAKRLKAIMLAAAELAAMRLRDLPASD 471
Query: 468 HKGIEFDQRDRVFRLLDKNGDGKISIEELTEVIEELGAPGEDAHDMMLLLDSNSDGSLSS 527
+ + +R F LDKNGDG+ISIEELTEV+E+LGA G+DA ++M LLD+NSDGSLSS
Sbjct: 472 QG--DSNCTNRAFCQLDKNGDGRISIEELTEVMEDLGAGGKDAKELMHLLDANSDGSLSS 529
Query: 528 DEFQMFQKQVEMVRNLEDRDDEYKKILDEKLHMADDSGLIQVYNKEFGNRLV 579
DEF+ FQ+Q+E++R+L+D+DD Y+KIL EKL D +GLIQVY K+ G++L+
Sbjct: 530 DEFEAFQRQIELMRSLDDKDDRYRKILKEKLQTIDSAGLIQVYRKQLGDKLL 581
>UniRef100_Q6ATB4 Putative chloroplast RelA [Oryza sativa]
Length = 583
Score = 665 bits (1715), Expect = 0.0
Identities = 335/532 (62%), Positives = 439/532 (81%), Gaps = 8/532 (1%)
Query: 50 SSTPRACSVEVPGGGKMVIELVGAFNDLTERMK--VLSTSSSGLLFKSLKLSIPVLQTSP 107
SS+ + S GGG++V ELVGAFN+LT RM + ++SSS LLF++LKL++P L+
Sbjct: 56 SSSSSSSSTPAEGGGRLVAELVGAFNELTGRMGEGLATSSSSRLLFRALKLALPALRDG- 114
Query: 108 LTPDGRSPLSKALSIAMLLADLQMDAEVISAGILREVLEVGELNLHEIRSQIGSATAHLL 167
DG L++AL+IA LADLQMDAEVISAGILRE L+ G +++ +++S+IG +TAHLL
Sbjct: 115 ---DGGRALARALAIAASLADLQMDAEVISAGILREALDAGAISMRDVKSEIGISTAHLL 171
Query: 168 HESLRVKNFASRVDILDDENAAALRKFCLTYYDIRALILDLALKLDMMRHLGHLPRYQQQ 227
HESLR+K+ S++D+LDDE+A+ALRKFCL+YYDIRA+IL+LALKLDMMRHL LPRY Q+
Sbjct: 172 HESLRLKHAPSKLDVLDDESASALRKFCLSYYDIRAVILELALKLDMMRHLDCLPRYLQR 231
Query: 228 IISLQVMKIYAPLAHAVGTNYISLELEDLSFQYLFPYSYLYVDTWLRSQETGGISLIDVY 287
I SL+V+KIYAPLAHAVG +SLELEDLSF+YLFP+SY ++D WLRSQET LID Y
Sbjct: 232 IKSLEVLKIYAPLAHAVGAGNLSLELEDLSFRYLFPHSYDHIDQWLRSQETENKLLIDSY 291
Query: 288 KDELLESLKSDPILAELVDDISVKGRYKSRYSTMKKLLKDGRRPEDVNDVLGLRVVLNPK 347
K++LL++LK D L+++V DIS++GRYKSR+STMKKL+KDGR+PE+VND+L LRV+L P+
Sbjct: 292 KEQLLQALKDDDELSQIVQDISIQGRYKSRFSTMKKLVKDGRKPEEVNDILALRVILEPR 351
Query: 348 SRENALEAGERACYRAHQIIQSMWKEIPSRTKDYISRPKGNGYRSLHMAVDVSEIGRTRP 407
++L+ G RAC+R H+IIQ+MWKE+P RTK+Y++RPK NGY+SLH+A+DVSE G+ RP
Sbjct: 352 CDGSSLDWGPRACHRTHEIIQAMWKEVPGRTKNYVTRPKENGYQSLHVAIDVSEPGKMRP 411
Query: 408 LMEIQIRTTEMDRLAVGGMASHSLYKAGLTNPEEAKRLKTIMLAAAELAALRLKDFPSAN 467
LMEIQIRT EM + AVGG ASHSLYK GLT+P EAKRLK IMLAAAELAA+RL+D P+++
Sbjct: 412 LMEIQIRTKEMHKFAVGGEASHSLYKGGLTDPGEAKRLKAIMLAAAELAAMRLRDLPASD 471
Query: 468 HKGIEFDQRDRVFRLLDKNGDGKISIEELTEVIEELGAPGEDAHDMMLLLDSNSDGSLSS 527
+ + +R F LDKNGDG+ISIEELTEV+E+LGA G+DA ++M LLD+NSDGSLSS
Sbjct: 472 QG--DSNCTNRAFCQLDKNGDGRISIEELTEVMEDLGAGGKDAKELMHLLDANSDGSLSS 529
Query: 528 DEFQMFQKQVEMVRNLEDRDDEYKKILDEKLHMADDSGLIQVYNKEFGNRLV 579
DEF+ FQ+Q+E++R+L+D+DD Y+KIL EKL D +GLIQVY K+ G++L+
Sbjct: 530 DEFEAFQRQIELMRSLDDKDDRYRKILKEKLQTIDSAGLIQVYRKQLGDKLL 581
>UniRef100_Q9LUQ0 Similarity to [Arabidopsis thaliana]
Length = 456
Score = 615 bits (1586), Expect = e-174
Identities = 311/453 (68%), Positives = 371/453 (81%), Gaps = 5/453 (1%)
Query: 131 MDAEVISAGILREVLEVGELNLHEIRSQIGSATAHLLHESLRVKNFASRVDILDDENAAA 190
MDAEVISA IL EV++ ++++E+R IG+ TAHLLHE RVKN +VD+LDDE AA+
Sbjct: 1 MDAEVISASILSEVVDANAISIYEVRDHIGTGTAHLLHEIFRVKNIPFKVDVLDDETAAS 60
Query: 191 LRKFCLTYYDIRALILDLALKLDMMRHLGHLPRYQQQIISLQVMKIYAPLAHAVGTNYIS 250
LRKF LTYYDIRA+I+DL KLD MRHL HLPRY+QQI+SL+V+KIY+PLAHAVG N++S
Sbjct: 61 LRKFYLTYYDIRAVIMDLVSKLDEMRHLDHLPRYRQQILSLEVLKIYSPLAHAVGANHLS 120
Query: 251 LELEDLSFQYLFPYSYLYVDTWLRSQETGGISLIDVYKDELLESLKSDPILAELVDDISV 310
LELED+SF+YLFP SY+Y+D+WLR E G LIDVYK++L SLK D +LAE+V+D+ +
Sbjct: 121 LELEDISFRYLFPCSYIYLDSWLRGHENGSKPLIDVYKEQLHRSLKDDLVLAEMVNDVYI 180
Query: 311 KGRYKSRYSTMKKLLKDGRRPEDVNDVLGLRVVLNPKSRENALEAGERACYRAHQIIQSM 370
KGRYKSRYS MKKLL+DGR+PE+VNDVLGLRV+L P S N +E GE+ACYR +II+S+
Sbjct: 181 KGRYKSRYSMMKKLLRDGRKPEEVNDVLGLRVILMPNSVVNDVEVGEKACYRTSEIIRSL 240
Query: 371 WKEIPSRTKDYISRPKGNGYRSLHMAVDVSEIGRTRPLMEIQIRTTEMDRLAVGGMASHS 430
WKEIP RTKDYI+RPK NGYRSLHMAVDVS+ + RPLMEIQIRT +MD A G ASHS
Sbjct: 241 WKEIPHRTKDYIARPKENGYRSLHMAVDVSDSDQIRPLMEIQIRTMDMDGSANAGTASHS 300
Query: 431 LYKAGLTNPEEAKRLKTIMLAAAELAALRLKDFPSANHKGIE--FDQRDRVFRLLDKNGD 488
LYK GLT+P+EAKRLK IMLAAA+LAA+RLKD S H+ + +QRDRVF LLDKNGD
Sbjct: 301 LYKGGLTDPKEAKRLKAIMLAAADLAAIRLKDISSNKHQSFKTTTNQRDRVFCLLDKNGD 360
Query: 489 GKISIEELTEVIEELGAPGEDAHDMMLLLDSNSDGSLSSDEFQMFQKQVEMVRNLEDRDD 548
G ISIEEL EV+EELGAPGEDA +MM LLDSNSDGSLSSDEF FQKQVE +R EDRD+
Sbjct: 361 GMISIEELMEVMEELGAPGEDAEEMMQLLDSNSDGSLSSDEFDTFQKQVEFMRKWEDRDN 420
Query: 549 EYKKILDEKLH---MADDSGLIQVYNKEFGNRL 578
EYK +LDEKLH D +GLIQ+YNKE +RL
Sbjct: 421 EYKSLLDEKLHDLPHQDTTGLIQLYNKELEDRL 453
>UniRef100_Q75IS2 Hypothetical protein OSJNBb0099P06.1 [Oryza sativa]
Length = 578
Score = 573 bits (1478), Expect = e-162
Identities = 300/532 (56%), Positives = 398/532 (74%), Gaps = 15/532 (2%)
Query: 63 GGKMVIELVGAFNDLTERM--KVLSTSSSGLLFKSLKLSIPVLQTSPLTPDGRSPLSKAL 120
GG+++ EL+G FN LTERM V ++SSS LLF++LKL++P L+ G +S+AL
Sbjct: 45 GGRLMAELLGVFNGLTERMGEDVATSSSSRLLFRALKLALPALRDGGGDGGGGQSVSRAL 104
Query: 121 SIAMLLADLQMDAEVISAGILREVLEVGELNLHEIRSQIGSATAHLLHESLRVKNFASRV 180
+A LADLQMDAEVISAG++R L+ G L + ++ +Q+G++ A L+ ESL+VK S V
Sbjct: 105 VVAASLADLQMDAEVISAGMVRGALDTGALAMADVEAQLGASAAGLVEESLKVKRAPSEV 164
Query: 181 DILDDENAAALRKFCLTYYDIRALILDLALKLDMMRHLGHLPRYQQQIISLQVMKIYAPL 240
D+ D+E A+ALRK CL+ YDIRA+IL+LA+KLD M+HL LP++QQ+ SL+V+K++A L
Sbjct: 165 DVADEEAASALRKRCLSSYDIRAVILELAVKLDAMKHLDVLPKHQQRTTSLEVLKVFALL 224
Query: 241 AHAVGTNYISLELEDLSFQYLFPYSYLYVDTWLRSQETGGISLIDVYKDELLESLKSDPI 300
AHAVG +SLELEDLSFQ L+P +Y ++D WL SQE +I K+ELL +L +D
Sbjct: 225 AHAVGAGELSLELEDLSFQRLYPQAYAHIDQWLSSQEDDCKRVIAASKEELLRALTADDE 284
Query: 301 LAELVDDISVKGRYKSRYSTMKKLLKDGRRPEDVNDVLGLRVVLNPKSRENA---LEAGE 357
L V + V GRYKSR+STMKKL+KDGRRPEDVND+LG+RV+L+P+ + G+
Sbjct: 285 LRCTVTGVDVMGRYKSRFSTMKKLVKDGRRPEDVNDILGMRVILDPRPGGGGGGDGDGGD 344
Query: 358 RACYRAHQIIQSMWKEIPSRTKDYISRPKGNGYRSLHMAVDVSE---IGRTRPLMEIQIR 414
RAC R H++I++MWK++P+RTKDYI+RPKGNGYRSLH+AVD+SE G+ RPLMEIQ+R
Sbjct: 345 RACLRTHEVIKAMWKDVPARTKDYITRPKGNGYRSLHVAVDMSEPGPEGKKRPLMEIQVR 404
Query: 415 TTEMDRLAVGGMASHSLYKAGLTNPEEAKRLKTIMLAAAELAALRLKDFP-------SAN 467
T EMD AVGG ASH+LYK GLT+PEEAKRLK IMLAAAE+AA L+D P +A
Sbjct: 405 TREMDMAAVGGQASHALYKGGLTDPEEAKRLKAIMLAAAEVAAQHLRDEPAGDGGQTTAA 464
Query: 468 HKGIEFDQRDRVFRLLDKNGDGKISIEELTEVIEELGAPGEDAHDMMLLLDSNSDGSLSS 527
+R F+LLDKNGDG+IS+EELTE++E+LGA G DA ++M LLD+NSDGSLSS
Sbjct: 465 ASAATAGNVERAFQLLDKNGDGRISMEELTEIMEDLGAGGHDAEELMRLLDANSDGSLSS 524
Query: 528 DEFQMFQKQVEMVRNLEDRDDEYKKILDEKLHMADDSGLIQVYNKEFGNRLV 579
DEF +FQK+V++ LE++DDEYK+IL +KL DD+GLI VY K ++LV
Sbjct: 525 DEFALFQKRVKLKTKLENKDDEYKEILKQKLQKVDDTGLIHVYRKNLSDKLV 576
>UniRef100_Q6ATB2 Hypothetical protein OSJNBa0034O12.17 [Oryza sativa]
Length = 559
Score = 535 bits (1377), Expect = e-150
Identities = 289/532 (54%), Positives = 387/532 (72%), Gaps = 24/532 (4%)
Query: 63 GGKMVIELVGAFNDLTERM--KVLSTSSSGLLFKSLKLSIPVLQTSPLTPDGRSPLSKAL 120
GG+++ EL+G FN LTERM V ++SS LLF++LKL++P L+ + GRS L++AL
Sbjct: 37 GGRLMAELLGVFNGLTERMGDDVATSSSWTLLFRALKLALPALRDAA---GGRS-LARAL 92
Query: 121 SIAMLLADLQMDAEVISAGILREVLEVGELNLHEIRSQIGSATAHLLHESLRVKNFASRV 180
+A LADLQMDAEVISAGI+R+ ++ G + + + +Q+G A LL ESL VKN SRV
Sbjct: 93 IVAASLADLQMDAEVISAGIVRQAMDAGAVAMADAEAQLGPGAAALLLESLDVKNAPSRV 152
Query: 181 DILDDENAAALRKFCLTYYDIRALILDLALKLDMMRHLGHLPRYQQQIISLQVMKIYAPL 240
D+ D+E A+A+R L+ YD+RA+IL+LA++LD M+HL +P++QQ+ SL+V+K++APL
Sbjct: 153 DVADEEAASAVRNRILSGYDVRAVILELAIRLDAMKHLDGVPKHQQRTTSLEVLKVFAPL 212
Query: 241 AHAVGTNYISLELEDLSFQYLFPYSYLYVDTWLRSQETGGISLIDVYKDELLESLKSDPI 300
AHAVG +S ELEDLSF L+P +Y VD WL QE ++ KD+LL++L +D
Sbjct: 213 AHAVGAGALSKELEDLSFWRLYPQAYAQVDQWLSGQEDDCKRVLATCKDDLLQALAADDE 272
Query: 301 LAELVDDISVKGRYKSRYSTMKKLLKDGRRPEDVNDVLGLRVVLNPKSRENALEAGERAC 360
L V VKGRYKSR+S MKKL+KDGRRPEDV+D+LG+RV+L+ ++ G RAC
Sbjct: 273 LRHTVAGFDVKGRYKSRFSAMKKLVKDGRRPEDVHDILGMRVILDHRA---GAGDGHRAC 329
Query: 361 YRAHQIIQSMWKEIPSRTKDYISRPKGNGYRSLHMAVDVSE---IGRTRPLMEIQIRTTE 417
R H++I+ MWK++P+RTKDYI+RPKG+GYRSLH+AVD+SE G+ RPLME+QIRT E
Sbjct: 330 IRTHEVIKGMWKDVPARTKDYIARPKGDGYRSLHIAVDMSEPGPEGKKRPLMEVQIRTKE 389
Query: 418 MDRLAVGGMASHSLYKAGLTNPEEAKRLKTIMLAAAELAALRLKD---------FPSANH 468
M+ AV G H+LYK L +PEEAKRLK IMLAAAE+AA L+D P+A
Sbjct: 390 MNDAAVFG---HALYKGCLADPEEAKRLKDIMLAAAEVAAQHLRDEPATGDQTGVPAAAA 446
Query: 469 KGIEFDQRDRVFRLLDKNGDGKISIEELTEVIEELGAPGEDAHDMMLLLDSNSDGSLSSD 528
+R FRLLDKNGDG+IS+EELTE++E+LGA G+DA ++M LLD N+DGSLSSD
Sbjct: 447 AAASAGNIERAFRLLDKNGDGRISMEELTELMEDLGAGGKDAEELMRLLDDNNDGSLSSD 506
Query: 529 EFQMFQKQVEMVRNLEDRDDEYKKILDEKLHMADDSGLIQVYNKEFGNRLVS 580
EF +FQK+VE+ LED+DDEYK+IL +KL DD+GLI VY K ++LVS
Sbjct: 507 EFALFQKRVELKAKLEDKDDEYKEILRQKLQKVDDTGLIHVYRKNLSDKLVS 558
>UniRef100_Q72HX7 Guanosine-3',5'-bis(Diphosphate) 3'-pyrophosphohydrolase [Thermus
thermophilus]
Length = 727
Score = 206 bits (523), Expect = 2e-51
Identities = 129/361 (35%), Positives = 203/361 (55%), Gaps = 13/361 (3%)
Query: 108 LTPDGRSPLSKALSIAMLLADLQMDAEVISAGILREVLEVGELNLHEIRSQIGSATAHLL 167
L G ++ +++A +LA LQMDA+ ++AG+L + LE + E+ + G ++
Sbjct: 42 LRRSGEPYITHPVAVAEILAGLQMDADTVAAGLLHDTLEDCGVAPEELERRFGPTVRRIV 101
Query: 168 HESLRVKNFASRVDILDDENAAA-LRK-FCLTYYDIRALILDLALKLDMMRHLGHLPRYQ 225
+V ++ +E A LR+ F D+R +I+ LA +L +R L H+P +
Sbjct: 102 EGETKVSKLYKLANLEGEERRAEDLRQMFIAMAEDVRIIIVKLADRLHNLRTLEHMPPEK 161
Query: 226 QQIISLQVMKIYAPLAHAVGTNYISLELEDLSFQYLFPYSYLYVDTWLRSQETGGISLID 285
Q+ I+ + ++IYAPLAH +G + ELEDLSF+YL P ++ + +++ + LI
Sbjct: 162 QKRIAQETLEIYAPLAHRLGMGQLKWELEDLSFRYLHPEAFASLSARIQATQEARERLIQ 221
Query: 286 VYKDELLESLKSDPILAELVDDISVKGRYKSRYSTMKKLLKDGRRPEDVNDVLGLRVVLN 345
L E+L D +L + V GR K YS KK+ ++G+ E + D+L +RV+L+
Sbjct: 222 KAIHLLQETLARDELLQSQLQGFEVTGRPKHLYSIWKKMEREGKTLEQIYDLLAVRVILD 281
Query: 346 PK---SRENALEAGERACYRAHQIIQSMWKEIPSRTKDYISRPKGNGYRSLHMAVDVSEI 402
PK +RE+ ++ CY ++ ++W+ IP R KDYI+ PK NGY+SLH V E
Sbjct: 282 PKPAPTRESQALREKQVCYHVLGLVHALWQPIPGRVKDYIAVPKPNGYQSLHTTVIALE- 340
Query: 403 GRTRPLMEIQIRTTEMDRLAVGGMASHSLYKAGLTNPEEAKR----LKTIMLAAAELAAL 458
PL E+QIRT EM R+A G+A+H LYK GLT+PEE KR LK+I E ++
Sbjct: 341 --GLPL-EVQIRTREMHRVAEYGIAAHWLYKEGLTDPEEVKRRVSWLKSIQEWQKEFSSS 397
Query: 459 R 459
R
Sbjct: 398 R 398
>UniRef100_Q5SHL3 Guanosine-3',5'-bis(Diphosphate) 3'-pyrophosphohydrolase [Thermus
thermophilus HB8]
Length = 727
Score = 205 bits (522), Expect = 3e-51
Identities = 129/361 (35%), Positives = 203/361 (55%), Gaps = 13/361 (3%)
Query: 108 LTPDGRSPLSKALSIAMLLADLQMDAEVISAGILREVLEVGELNLHEIRSQIGSATAHLL 167
L G ++ +++A +LA LQMDA+ ++AG+L + LE + E+ + G ++
Sbjct: 42 LRRSGEPYITHPVAVAEILAGLQMDADTVAAGLLHDTLEDCGVAPEELERRFGPTVRRIV 101
Query: 168 HESLRVKNFASRVDILDDENAAA-LRK-FCLTYYDIRALILDLALKLDMMRHLGHLPRYQ 225
+V ++ +E A LR+ F D+R +I+ LA +L +R L H+P +
Sbjct: 102 EGETKVSKLYKLANLEGEERRAEDLRQMFIAMAEDVRIIIVKLADRLHNLRTLEHMPPEK 161
Query: 226 QQIISLQVMKIYAPLAHAVGTNYISLELEDLSFQYLFPYSYLYVDTWLRSQETGGISLID 285
Q+ I+ + ++IYAPLAH +G + ELEDLSF+YL P ++ + +++ + LI
Sbjct: 162 QKRIAQETLEIYAPLAHRLGMGQLKWELEDLSFRYLHPEAFASLSARIQATQEARERLIQ 221
Query: 286 VYKDELLESLKSDPILAELVDDISVKGRYKSRYSTMKKLLKDGRRPEDVNDVLGLRVVLN 345
L E+L D +L + V GR K YS KK+ ++G+ E + D+L +RV+L+
Sbjct: 222 RAIHLLQETLARDELLQSQLQGFEVTGRPKHLYSIWKKMEREGKTLEQIYDLLAVRVILD 281
Query: 346 PK---SRENALEAGERACYRAHQIIQSMWKEIPSRTKDYISRPKGNGYRSLHMAVDVSEI 402
PK +RE+ ++ CY ++ ++W+ IP R KDYI+ PK NGY+SLH V E
Sbjct: 282 PKPAPTRESQALREKQVCYHVLGLVHALWQPIPGRVKDYIAVPKPNGYQSLHTTVIALE- 340
Query: 403 GRTRPLMEIQIRTTEMDRLAVGGMASHSLYKAGLTNPEEAKR----LKTIMLAAAELAAL 458
PL E+QIRT EM R+A G+A+H LYK GLT+PEE KR LK+I E ++
Sbjct: 341 --GLPL-EVQIRTREMHRVAEYGIAAHWLYKEGLTDPEEVKRRVSWLKSIQEWQKEFSSS 397
Query: 459 R 459
R
Sbjct: 398 R 398
>UniRef100_Q9RTC7 GTP pyrophosphokinase [Deinococcus radiodurans]
Length = 787
Score = 184 bits (468), Expect = 5e-45
Identities = 121/345 (35%), Positives = 186/345 (53%), Gaps = 25/345 (7%)
Query: 112 GRSPLSKALSIAMLLADLQMDAEVISAGILREVLE-VGELNLHEIRSQIGSATAHLLHES 170
G ++ +++A++LA L MD + + AG+L + +E V + I G ++
Sbjct: 75 GEPYITHPVAVAIILARLGMDTDSLMAGLLHDTVEDVEGVTFEVIERNFGPDVRRIVEGE 134
Query: 171 LRVKNFASRVDILD-------DENAAALRKFCLTY-YDIRALILDLALKLDMMRHLGHLP 222
+V + + + D A LR+ + D+R +++ LA +L MR LG +
Sbjct: 135 TKVSKLSKQGNQQAEVPGDGRDMQAENLRQMLIAMTVDLRIIVVKLADRLHNMRTLGSMK 194
Query: 223 RYQQQIISLQVMKIYAPLAHAVGTNYISLELEDLSFQYLFPYSYLYVDTWLRSQETGGIS 282
+QQ I+ + M+I+APLAH +G I ELEDLSF+YL P +Y Y+ T LR+++
Sbjct: 195 PEKQQRIARETMEIFAPLAHRLGIGRIKWELEDLSFRYLHPDAYEYLQTRLRTRQEERDE 254
Query: 283 LIDVYKDELLESLKSDPILAELVDDISVKGRYKSRYSTMKKLLKDGRRPEDVNDVLGLRV 342
LI EL ++L+ D L E V+DI + GR K +S K+ K+G+ E + D+L LRV
Sbjct: 255 LIVNAVAELQDALEDDLELLEWVEDIDIAGRSKHLWSIHNKMQKEGKALEQIFDLLALRV 314
Query: 343 VLNPK-----------SRENALEAGE-RACYRAHQIIQSMWKEIPSRTKDYISRPKGNGY 390
+L P+ RE A E E R CY ++ SMW +P R KDYI+ PK NGY
Sbjct: 315 ILKPRPLTVPEGVEESRRERAEENREKRVCYHTLSVVHSMWTPLPGRVKDYIAVPKPNGY 374
Query: 391 RSLHMAVDVSEIGRTRPLMEIQIRTTEMDRLAVGGMASHSLYKAG 435
+SLH V I R+ +E+QIR+ M +A G+A+H +YK G
Sbjct: 375 QSLHTTV----ISRSGQPIEVQIRSLRMHEVAEYGVAAHWMYKQG 415
>UniRef100_Q72AB4 GTP pyrophosphokinase [Desulfovibrio vulgaris]
Length = 717
Score = 174 bits (440), Expect = 8e-42
Identities = 114/332 (34%), Positives = 175/332 (52%), Gaps = 24/332 (7%)
Query: 112 GRSPLSKALSIAMLLADLQMDAEVISAGILREVLEVGELNLHEIRSQIGSATAHLLHESL 171
G LS L++A LLA++ D ++AG+L + +E + + +I + G A ++
Sbjct: 42 GEPYLSHPLAVADLLAEMHFDEATVAAGLLHDTVEDTDAGIEDIDREFGEEVADIVDGVT 101
Query: 172 RVKNFASRVDILDDENAAALRKFCLTY-YDIRALILDLALKLDMMRHLGHLPRYQQQIIS 230
++ D ++ A +RK L +DIR L++ LA +L MR L ++QQ I+
Sbjct: 102 KISQMT--FDSKEEAQAENIRKMILAMSHDIRVLMVKLADRLHNMRTLDFQKSHKQQSIA 159
Query: 231 LQVMKIYAPLAHAVGTNYISLELEDLSFQYLFPYSYLYVDTWLRSQETGGISLIDVYKDE 290
+ + IYAPLA+ +G + I LELEDL +Y P + + WL + +LID
Sbjct: 160 QETLDIYAPLANRLGLHRIKLELEDLGLRYTKPDVFAQITDWLDENQMVERNLIDKVIAR 219
Query: 291 LLESLKSDPILAELVDDISVKGRYKSRYSTMKKLLKDGRRPEDVNDVLGLRVVLNPKSRE 350
+ E L ++ I E V+GR K + S KK+ + G ++++D++ RV++
Sbjct: 220 IREVLDANGIEGE------VRGRIKHKSSIYKKMTQQGLALDEMHDIIAFRVIV------ 267
Query: 351 NALEAGERACYRAHQIIQSMWKEIPSRTKDYISRPKGNGYRSLHMAVDVSEIGRTRPLME 410
A R CY ++ + WK + R KDYIS PK NGY+SLH V IG +E
Sbjct: 268 ----ADLRDCYAVLGLMHAQWKPVHGRFKDYISMPKANGYQSLHTTV----IGPEGERIE 319
Query: 411 IQIRTTEMDRLAVGGMASHSLYK-AGLTNPEE 441
IQIRT EM R+A G+ASH LYK AG NP +
Sbjct: 320 IQIRTREMHRMAEHGVASHWLYKDAGRVNPRD 351
>UniRef100_Q6JJ76 RelA [Agrobacterium tumefaciens]
Length = 744
Score = 165 bits (418), Expect = 3e-39
Identities = 111/328 (33%), Positives = 174/328 (52%), Gaps = 26/328 (7%)
Query: 111 DGRSPLSKALSIAMLLADLQMDAEVISAGILREVLEVGELNLHEIRSQIGSATAHLLHES 170
+G +S L +A +L ++ +D I+ +L + +E EI G L+
Sbjct: 41 NGDPYISHPLEVAAILTEMHLDESTIAVALLHDTIEDTTATRAEIDELFGEDIGRLVEGL 100
Query: 171 LRVKNFASRVDILDDE--NAAALRKFCLTYYD-IRALILDLALKLDMMRHLGHLPRYQQQ 227
++K ++D++ + A LRK L D +R L++ LA +L MR + ++P ++
Sbjct: 101 TKLK----KLDLVTRKAKQAENLRKLLLAISDDVRVLLVKLADRLHNMRTMEYMPADKRS 156
Query: 228 IISLQVMKIYAPLAHAVGTNYISLELEDLSFQYLFPYSYLYVDTWLRSQETGGISLIDVY 287
IS + M+IYAPLA +G + ELEDLSF+YL P +Y V L+ ET LI
Sbjct: 157 RISEETMEIYAPLAGRMGMQDMRDELEDLSFRYLNPEAYETVTNRLQELETRNEGLIKKI 216
Query: 288 KDELLESLKSDPILAELVDDISVKGRYKSRYSTMKKLLKDGRRPEDVNDVLGLRVVLNPK 347
+DEL E L ++ +L SVKGR K YS +K+ E ++DV G R++++
Sbjct: 217 EDELRELLVANGLLG-----ASVKGRQKKPYSGFRKMQSKSLSFEQLSDVYGFRILVDDI 271
Query: 348 SRENALEAGERACYRAHQIIQSMWKEIPSRTKDYISRPKGNGYRSLHMAVDVSEIGRTRP 407
+CYRA I+ + W+ +P R KDYIS PK N YRS+H + +G +R
Sbjct: 272 P----------SCYRALGIVHTRWRVVPGRFKDYISTPKQNDYRSIHTTI----VGPSRQ 317
Query: 408 LMEIQIRTTEMDRLAVGGMASHSLYKAG 435
+E+Q+RT M +A G+A+H+LYK G
Sbjct: 318 RIELQMRTKRMHEIAEFGIAAHALYKDG 345
>UniRef100_Q8UGK7 GTP pyrophosphohydrolases/synthetases, RelA/SpoT family
[Agrobacterium tumefaciens]
Length = 744
Score = 164 bits (414), Expect = 9e-39
Identities = 111/328 (33%), Positives = 171/328 (51%), Gaps = 26/328 (7%)
Query: 111 DGRSPLSKALSIAMLLADLQMDAEVISAGILREVLEVGELNLHEIRSQIGSATAHLLHES 170
+G +S L +A +L ++ +D I+ +L + +E EI G L+
Sbjct: 41 NGDPYISHPLEVAAILTEMHLDESTIAVALLHDTIEDTTATRAEIDELFGEDIGRLVEGL 100
Query: 171 LRVKNFASRVDILDDE--NAAALRKFCLTYYD-IRALILDLALKLDMMRHLGHLPRYQQQ 227
++K ++D++ + A LRK L D +R L++ LA +L MR + ++P ++
Sbjct: 101 TKLK----KLDLVTRKAKQAENLRKLLLAISDDVRVLLVKLADRLHNMRTMEYMPADKRS 156
Query: 228 IISLQVMKIYAPLAHAVGTNYISLELEDLSFQYLFPYSYLYVDTWLRSQETGGISLIDVY 287
IS + M+IYAPLA +G + ELEDLSF+YL P +Y V L ET LI
Sbjct: 157 RISEETMEIYAPLAGRMGMQDMRDELEDLSFRYLNPEAYETVTNRLLELETRNEGLIKKI 216
Query: 288 KDELLESLKSDPILAELVDDISVKGRYKSRYSTMKKLLKDGRRPEDVNDVLGLRVVLNPK 347
+DEL E L ++ +L VKGR K YS +K+ E ++DV G R++++
Sbjct: 217 EDELRELLVANGLLG-----THVKGRQKKPYSVFRKMQSKSLSFEQLSDVYGFRILVDDI 271
Query: 348 SRENALEAGERACYRAHQIIQSMWKEIPSRTKDYISRPKGNGYRSLHMAVDVSEIGRTRP 407
CYRA I+ + W+ +P R KDYIS PK N YRS+H + +G +R
Sbjct: 272 P----------GCYRALGIVHTRWRVVPGRFKDYISTPKQNDYRSIHTTI----VGPSRQ 317
Query: 408 LMEIQIRTTEMDRLAVGGMASHSLYKAG 435
+E+QIRT M +A G+A+H+LYK G
Sbjct: 318 RIELQIRTKRMHEIAEFGIAAHALYKDG 345
>UniRef100_Q7D045 AGR_C_1896p [Agrobacterium tumefaciens]
Length = 764
Score = 164 bits (414), Expect = 9e-39
Identities = 111/328 (33%), Positives = 171/328 (51%), Gaps = 26/328 (7%)
Query: 111 DGRSPLSKALSIAMLLADLQMDAEVISAGILREVLEVGELNLHEIRSQIGSATAHLLHES 170
+G +S L +A +L ++ +D I+ +L + +E EI G L+
Sbjct: 61 NGDPYISHPLEVAAILTEMHLDESTIAVALLHDTIEDTTATRAEIDELFGEDIGRLVEGL 120
Query: 171 LRVKNFASRVDILDDE--NAAALRKFCLTYYD-IRALILDLALKLDMMRHLGHLPRYQQQ 227
++K ++D++ + A LRK L D +R L++ LA +L MR + ++P ++
Sbjct: 121 TKLK----KLDLVTRKAKQAENLRKLLLAISDDVRVLLVKLADRLHNMRTMEYMPADKRS 176
Query: 228 IISLQVMKIYAPLAHAVGTNYISLELEDLSFQYLFPYSYLYVDTWLRSQETGGISLIDVY 287
IS + M+IYAPLA +G + ELEDLSF+YL P +Y V L ET LI
Sbjct: 177 RISEETMEIYAPLAGRMGMQDMRDELEDLSFRYLNPEAYETVTNRLLELETRNEGLIKKI 236
Query: 288 KDELLESLKSDPILAELVDDISVKGRYKSRYSTMKKLLKDGRRPEDVNDVLGLRVVLNPK 347
+DEL E L ++ +L VKGR K YS +K+ E ++DV G R++++
Sbjct: 237 EDELRELLVANGLLG-----THVKGRQKKPYSVFRKMQSKSLSFEQLSDVYGFRILVDDI 291
Query: 348 SRENALEAGERACYRAHQIIQSMWKEIPSRTKDYISRPKGNGYRSLHMAVDVSEIGRTRP 407
CYRA I+ + W+ +P R KDYIS PK N YRS+H + +G +R
Sbjct: 292 P----------GCYRALGIVHTRWRVVPGRFKDYISTPKQNDYRSIHTTI----VGPSRQ 337
Query: 408 LMEIQIRTTEMDRLAVGGMASHSLYKAG 435
+E+QIRT M +A G+A+H+LYK G
Sbjct: 338 RIELQIRTKRMHEIAEFGIAAHALYKDG 365
>UniRef100_Q67LM4 GTP pyrophosphokinase [Symbiobacterium thermophilum]
Length = 735
Score = 162 bits (410), Expect = 3e-38
Identities = 115/338 (34%), Positives = 173/338 (51%), Gaps = 27/338 (7%)
Query: 112 GRSPLSKALSIAMLLADLQMDAEVISAGILREVLEVGELNLHEIRSQIGSATAHLLH--E 169
G ++ +++A ++A L++D E I+A +L +VLE + E+ + G A L+
Sbjct: 45 GEPYITHPIAVAEIVASLELDEESIAAALLHDVLEDCGVKYEELEERFGKEVATLVDGVS 104
Query: 170 SLRVKNFASRVDILDDENAAALRKFCLTYY-DIRALILDLALKLDMMRHLGHLPRYQQQI 228
L F SR D+ LRK L D+R +++ LA +L MR L H P Q
Sbjct: 105 KLDRLQFTSR----DEAQVENLRKMFLAMAKDLRVILIKLADRLHNMRTLKHQPSDAQVR 160
Query: 229 ISLQVMKIYAPLAHAVGTNYISLELEDLSFQYLFPYSYLYVDTWLRSQETGGISLIDVYK 288
I+ + M IYAPLAH +G + I ELEDL+F+ L P Y + + + +++
Sbjct: 161 IAQETMDIYAPLAHRLGLSEIKWELEDLAFRILEPDRYQEMAYLVARKRQERMAITQDLM 220
Query: 289 DELLESLKSDPILAELVDDISVKGRYKSRYSTMKKLLKDGRRPEDVNDVLGLRVVLNPKS 348
D+L E L+ + I AE + GR K YS KK+ + G+ + D++ +R+++
Sbjct: 221 DQLREVLEKEGIQAE------ISGRPKHFYSIYKKMYRQGKDISQIYDLIAIRIIVEE-- 272
Query: 349 RENALEAGERACYRAHQIIQSMWKEIPSRTKDYISRPKGNGYRSLHMAVDVSEIGRTRPL 408
+ CY II S WK +P R KDYI+ PK N Y+SLH V IG
Sbjct: 273 --------VKDCYAVLGIIHSHWKPLPLRFKDYIATPKPNMYQSLHTTV----IGPHGEP 320
Query: 409 MEIQIRTTEMDRLAVGGMASHSLYKAGLTNPEEAKRLK 446
EIQIRT EM R A G+A+H YK G T+ E ++L+
Sbjct: 321 FEIQIRTREMHRTAEFGVAAHWSYKEGKTDKEFDRKLQ 358
>UniRef100_Q8CXC6 GTP pyrophosphokinase [Oceanobacillus iheyensis]
Length = 734
Score = 161 bits (407), Expect = 6e-38
Identities = 117/368 (31%), Positives = 185/368 (49%), Gaps = 23/368 (6%)
Query: 76 DLTERMKVLSTSSSGLLFKSLKLSIPVLQTSPLTPDGRSPLSKALSIAMLLADLQMDAEV 135
D+ E++K + L + G + + +A +LADLQMD+E
Sbjct: 11 DIVEQVKTYLSEDDVALIEQAYEFASDAHKDQFRKSGEPYIIHPVQVAGILADLQMDSET 70
Query: 136 ISAGILREVLEVGELNLHEIRSQIGSATAHLLHESLRVKNFASRVDILDDENAAALRK-F 194
ISAG L +V+E ++ + ++ A L+ ++ R + + + A RK F
Sbjct: 71 ISAGFLHDVVEDTDVTVEQLEEAFNHEIAMLVDGVTKLGKI--RYETKEAQQAENHRKMF 128
Query: 195 CLTYYDIRALILDLALKLDMMRHLGHLPRYQQQIISLQVMKIYAPLAHAVGTNYISLELE 254
DIR +++ LA +L MR L HL +Q+ IS + ++I+APLAH +G + I ELE
Sbjct: 129 VAMAKDIRVIMIKLADRLHNMRTLKHLAPEKQRRISNETLEIFAPLAHRLGISTIKWELE 188
Query: 255 DLSFQYLFPYSYLYVDTWLRSQETGGISLIDVYKDELLESLKSDPILAELVDDISVKGRY 314
D + +YL P Y + ++ + S I DEL L + I AE+ GR
Sbjct: 189 DTALRYLNPQQYYRIVQLMKQKREERESYIQEVIDELALELDTVNIEAEM------SGRP 242
Query: 315 KSRYSTMKKLLKDGRRPEDVNDVLGLRVVLNPKSRENALEAGERACYRAHQIIQSMWKEI 374
K YS +K++K ++ ++ D+L +R+++ EN + CY II + WK +
Sbjct: 243 KHLYSIYQKMVKQKKQFNEIYDLLAVRILV-----ENI-----KDCYAVLGIIHTNWKPM 292
Query: 375 PSRTKDYISRPKGNGYRSLHMAVDVSEIGRTRPLMEIQIRTTEMDRLAVGGMASHSLYKA 434
P R KDYI+ PK N Y+SLH V IG +E+QIRT EM +A G+A+H YK
Sbjct: 293 PGRFKDYIAMPKQNLYQSLHTTV----IGPKGDPLEVQIRTKEMHDIAEYGIAAHWAYKE 348
Query: 435 GLTNPEEA 442
G T+ E++
Sbjct: 349 GKTSREKS 356
>UniRef100_Q817X4 GTP pyrophosphokinase [Bacillus cereus]
Length = 727
Score = 160 bits (406), Expect = 7e-38
Identities = 106/319 (33%), Positives = 166/319 (51%), Gaps = 23/319 (7%)
Query: 120 LSIAMLLADLQMDAEVISAGILREVLEVGELNLHEIRSQIGSATAHLLHESLRVKNFASR 179
+ +A +L DL MD +SAG L +V+E E+ L +I + A L+ ++ +
Sbjct: 55 IQVAGILVDLHMDPATVSAGFLHDVVEDTEITLEDIEREFNKEIAMLVDGVTKLGKIKYK 114
Query: 180 VDILDDENAAALRK-FCLTYYDIRALILDLALKLDMMRHLGHLPRYQQQIISLQVMKIYA 238
+ + A RK F DIR +++ LA +L MR L HLP+ +Q+ I+ + ++I+A
Sbjct: 115 SH--EQQQAENHRKMFIAMAQDIRVILIKLADRLHNMRTLKHLPQEKQRRIANETLEIFA 172
Query: 239 PLAHAVGTNYISLELEDLSFQYLFPYSYLYVDTWLRSQETGGISLIDVYKDELLESLKSD 298
PLAH +G + I ELED S +YL P Y + ++ + + Y DE++ ++
Sbjct: 173 PLAHRLGISTIKWELEDTSLRYLNPQQYYRIVNLMKRKRAER----EEYLDEVMTGIREK 228
Query: 299 PILAELVDDISVKGRYKSRYSTMKKLLKDGRRPEDVNDVLGLRVVLNPKSRENALEAGER 358
L E+ + GR K YS +K+ ++ ++ D+L +RVV+N +
Sbjct: 229 --LKEVAIQPEISGRPKHIYSIYRKMALQNKQFNEIYDLLAVRVVVN----------SIK 276
Query: 359 ACYRAHQIIQSMWKEIPSRTKDYISRPKGNGYRSLHMAVDVSEIGRTRPLMEIQIRTTEM 418
CY II + WK +P R KDYI+ PK N Y+SLH V IG +E+QIRT EM
Sbjct: 277 DCYAVLGIIHTCWKPMPGRFKDYIAMPKANLYQSLHTTV----IGPKGDPLEVQIRTKEM 332
Query: 419 DRLAVGGMASHSLYKAGLT 437
+A G+A+H YK G T
Sbjct: 333 HEIAEFGIAAHWAYKEGKT 351
>UniRef100_UPI00003CB978 UPI00003CB978 UniRef100 entry
Length = 727
Score = 160 bits (405), Expect = 1e-37
Identities = 106/319 (33%), Positives = 166/319 (51%), Gaps = 23/319 (7%)
Query: 120 LSIAMLLADLQMDAEVISAGILREVLEVGELNLHEIRSQIGSATAHLLHESLRVKNFASR 179
+ +A +L DL MD +SAG L +V+E E+ L +I + A L+ ++ +
Sbjct: 55 IQVAGILVDLHMDPATVSAGFLHDVVEDTEITLEDIEREFNKEIAMLVDGVTKLGKIKYK 114
Query: 180 VDILDDENAAALRK-FCLTYYDIRALILDLALKLDMMRHLGHLPRYQQQIISLQVMKIYA 238
+ + A RK F DIR +++ LA +L MR L HLP+ +Q+ I+ + ++I+A
Sbjct: 115 SH--EQQQAENHRKMFIAMAQDIRVILIKLADRLHNMRTLKHLPQEKQRRIANETLEIFA 172
Query: 239 PLAHAVGTNYISLELEDLSFQYLFPYSYLYVDTWLRSQETGGISLIDVYKDELLESLKSD 298
PLAH +G + I ELED S +YL P Y + ++ + + Y DE++ ++
Sbjct: 173 PLAHRLGISTIKWELEDTSLRYLNPQQYYRIVNLMKRKRAER----EEYLDEVMTGIREK 228
Query: 299 PILAELVDDISVKGRYKSRYSTMKKLLKDGRRPEDVNDVLGLRVVLNPKSRENALEAGER 358
L E+ + GR K YS +K+ ++ ++ D+L +RVV+N +
Sbjct: 229 --LDEVAIQPEISGRPKHIYSIYRKMALQNKQFNEIYDLLAVRVVVN----------SIK 276
Query: 359 ACYRAHQIIQSMWKEIPSRTKDYISRPKGNGYRSLHMAVDVSEIGRTRPLMEIQIRTTEM 418
CY II + WK +P R KDYI+ PK N Y+SLH V IG +E+QIRT EM
Sbjct: 277 DCYAVLGIIHTCWKPMPGRFKDYIAMPKANLYQSLHTTV----IGPKGDPLEVQIRTKEM 332
Query: 419 DRLAVGGMASHSLYKAGLT 437
+A G+A+H YK G T
Sbjct: 333 HEIAEFGIAAHWAYKEGKT 351
>UniRef100_Q6HDB9 GTP diphosphokinase [Bacillus thuringiensis]
Length = 727
Score = 160 bits (405), Expect = 1e-37
Identities = 106/319 (33%), Positives = 166/319 (51%), Gaps = 23/319 (7%)
Query: 120 LSIAMLLADLQMDAEVISAGILREVLEVGELNLHEIRSQIGSATAHLLHESLRVKNFASR 179
+ +A +L DL MD +SAG L +V+E E+ L +I + A L+ ++ +
Sbjct: 55 IQVAGILVDLHMDPATVSAGFLHDVVEDTEITLEDIEREFNKEIAMLVDGVTKLGKIKYK 114
Query: 180 VDILDDENAAALRK-FCLTYYDIRALILDLALKLDMMRHLGHLPRYQQQIISLQVMKIYA 238
+ + A RK F DIR +++ LA +L MR L HLP+ +Q+ I+ + ++I+A
Sbjct: 115 SH--EQQQAENHRKMFIAMAQDIRVILIKLADRLHNMRTLKHLPQEKQRRIANETLEIFA 172
Query: 239 PLAHAVGTNYISLELEDLSFQYLFPYSYLYVDTWLRSQETGGISLIDVYKDELLESLKSD 298
PLAH +G + I ELED S +YL P Y + ++ + + Y DE++ ++
Sbjct: 173 PLAHRLGISTIKWELEDTSLRYLNPQQYYRIVNLMKRKRAER----EEYLDEVMTGIREK 228
Query: 299 PILAELVDDISVKGRYKSRYSTMKKLLKDGRRPEDVNDVLGLRVVLNPKSRENALEAGER 358
L E+ + GR K YS +K+ ++ ++ D+L +RVV+N +
Sbjct: 229 --LDEVAIQPEISGRPKHIYSIYRKMALQNKQFNEIYDLLAVRVVVN----------SIK 276
Query: 359 ACYRAHQIIQSMWKEIPSRTKDYISRPKGNGYRSLHMAVDVSEIGRTRPLMEIQIRTTEM 418
CY II + WK +P R KDYI+ PK N Y+SLH V IG +E+QIRT EM
Sbjct: 277 DCYAVLGIIHTCWKPMPGRFKDYIAMPKANLYQSLHTTV----IGPKGDPLEVQIRTKEM 332
Query: 419 DRLAVGGMASHSLYKAGLT 437
+A G+A+H YK G T
Sbjct: 333 HEIAEFGIAAHWAYKEGKT 351
>UniRef100_Q634D7 GTP diphosphokinase [Bacillus cereus]
Length = 727
Score = 160 bits (405), Expect = 1e-37
Identities = 106/319 (33%), Positives = 166/319 (51%), Gaps = 23/319 (7%)
Query: 120 LSIAMLLADLQMDAEVISAGILREVLEVGELNLHEIRSQIGSATAHLLHESLRVKNFASR 179
+ +A +L DL MD +SAG L +V+E E+ L +I + A L+ ++ +
Sbjct: 55 IQVAGILVDLHMDPATVSAGFLHDVVEDTEITLEDIEREFNKEIAMLVDGVTKLGKIKYK 114
Query: 180 VDILDDENAAALRK-FCLTYYDIRALILDLALKLDMMRHLGHLPRYQQQIISLQVMKIYA 238
+ + A RK F DIR +++ LA +L MR L HLP+ +Q+ I+ + ++I+A
Sbjct: 115 SH--EQQQAENHRKMFIAMAQDIRVILIKLADRLHNMRTLKHLPQEKQRRIANETLEIFA 172
Query: 239 PLAHAVGTNYISLELEDLSFQYLFPYSYLYVDTWLRSQETGGISLIDVYKDELLESLKSD 298
PLAH +G + I ELED S +YL P Y + ++ + + Y DE++ ++
Sbjct: 173 PLAHRLGISTIKWELEDTSLRYLNPQQYYRIVNLMKRKRAER----EEYLDEVMTGIREK 228
Query: 299 PILAELVDDISVKGRYKSRYSTMKKLLKDGRRPEDVNDVLGLRVVLNPKSRENALEAGER 358
L E+ + GR K YS +K+ ++ ++ D+L +RVV+N +
Sbjct: 229 --LDEVAIQPEISGRPKHIYSIYRKMALQNKQFNEIYDLLAVRVVVN----------SIK 276
Query: 359 ACYRAHQIIQSMWKEIPSRTKDYISRPKGNGYRSLHMAVDVSEIGRTRPLMEIQIRTTEM 418
CY II + WK +P R KDYI+ PK N Y+SLH V IG +E+QIRT EM
Sbjct: 277 DCYAVLGIIHTCWKPMPGRFKDYIAMPKANLYQSLHTTV----IGPKGDPLEVQIRTKEM 332
Query: 419 DRLAVGGMASHSLYKAGLT 437
+A G+A+H YK G T
Sbjct: 333 HEIAEFGIAAHWAYKEGKT 351
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.319 0.135 0.379
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 928,985,400
Number of Sequences: 2790947
Number of extensions: 38499672
Number of successful extensions: 121993
Number of sequences better than 10.0: 1891
Number of HSP's better than 10.0 without gapping: 923
Number of HSP's successfully gapped in prelim test: 974
Number of HSP's that attempted gapping in prelim test: 117596
Number of HSP's gapped (non-prelim): 3040
length of query: 580
length of database: 848,049,833
effective HSP length: 133
effective length of query: 447
effective length of database: 476,853,882
effective search space: 213153685254
effective search space used: 213153685254
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 78 (34.7 bits)
Medicago: description of AC146862.24


