

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146853.5 - phase: 0 /pseudo
(235 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
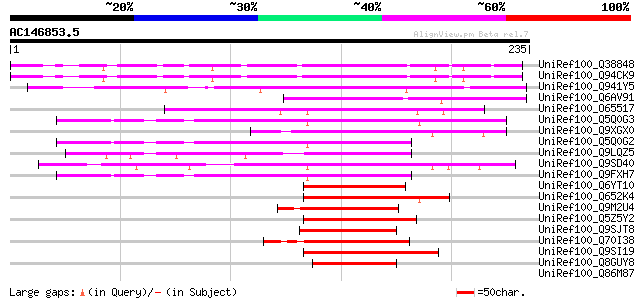
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q38848 LRP1 [Arabidopsis thaliana] 141 2e-32
UniRef100_Q94CK9 Lateral root primordia [Arabidopsis thaliana] 138 1e-31
UniRef100_Q941Y5 Putative LRP (Lateral root primordia) 1 [Oryza ... 124 2e-27
UniRef100_Q6AV91 Hypothetical protein OJ1286_E05.15 [Oryza sativa] 116 5e-25
UniRef100_O65517 Hypothetical protein F23E13.150 [Arabidopsis th... 103 5e-21
UniRef100_Q5Q0G3 Hypothetical protein [Arabidopsis thaliana] 93 5e-18
UniRef100_Q9XGX0 Zinc finger protein SHI-like [Arabidopsis thali... 93 5e-18
UniRef100_Q5Q0G2 Hypothetical protein [Arabidopsis thaliana] 92 1e-17
UniRef100_Q9LQZ5 F10A5.26 [Arabidopsis thaliana] 92 1e-17
UniRef100_Q9SD40 Hypothetical protein F24M12.100 [Arabidopsis th... 92 1e-17
UniRef100_Q9FXH7 F6F9.16 protein [Arabidopsis thaliana] 92 1e-17
UniRef100_Q6YT10 Putative lateral root primordia [Oryza sativa] 89 9e-17
UniRef100_Q652K4 Putative LRP1 [Oryza sativa] 89 1e-16
UniRef100_Q9M2U4 Hypothetical protein T12E18_120 [Arabidopsis th... 88 2e-16
UniRef100_Q5Z5Y2 Putative LRP1 [Oryza sativa] 86 8e-16
UniRef100_Q9SJT8 Hypothetical protein At2g21400 [Arabidopsis tha... 80 4e-14
UniRef100_Q70I38 Hypothetical protein [Lotus japonicus] 79 1e-13
UniRef100_Q9SI19 Hypothetical protein At2g18120 [Arabidopsis tha... 78 2e-13
UniRef100_Q8GUY8 Hypothetical protein F3K23.16 [Arabidopsis thal... 67 3e-10
UniRef100_Q86M87 ComB [Dictyostelium discoideum] 45 0.002
>UniRef100_Q38848 LRP1 [Arabidopsis thaliana]
Length = 320
Score = 141 bits (355), Expect = 2e-32
Identities = 101/241 (41%), Positives = 134/241 (54%), Gaps = 36/241 (14%)
Query: 1 MSMLGLRDLVLIAPSPSSLQHHHQQNQNQNQNQNQPISHDH-NSNHSLPSSASLSVGFGI 59
M M+GLRD+ L+AP+ +HHQ N DH NSN ++A+ ++G G+
Sbjct: 1 MGMVGLRDVFLVAPA-----YHHQ-------NAGVISGSDHMNSN----AAAAAALGVGV 44
Query: 60 FPLLTATPCMPQQQSQNNEVQENPSNNNNFW-NLRMCPEVVNLPKKGVITEEENHGKIAM 118
PLLTA P PQQ +++++ N NN W N E L K T + G +
Sbjct: 45 IPLLTAGP--PQQNVEDSDI--NFLGNNRRWQNNNNNHETQYLHFKS--TNQTTVGTSSN 98
Query: 119 MESEENGVYGSEYRVCQDCGNRAKKDCVFRRCRTCCKGRGYDCSTHLKSTWIPSTRRRER 178
+G G+ CQDCGN+AKK+C RRCRTCCK RG+DCSTH+KSTW+ + RRRER
Sbjct: 99 NSGSGSGASGTA--TCQDCGNQAKKECKQRRCRTCCKSRGFDCSTHVKSTWVSAARRRER 156
Query: 179 EVEMFAGGGDGEG-----CSGVKRQKGLLGS--SQNAAATSHSSNSNGTTPKSFATSSCH 231
+V M G G SG K+ + ++GS Q ATSH+S SN T P+SF TSS
Sbjct: 157 QV-MPTGANPTAGSSLSTSSGTKKPR-IVGSQQQQQQQATSHTSTSN-TPPQSFETSSSR 213
Query: 232 Q 232
Q
Sbjct: 214 Q 214
>UniRef100_Q94CK9 Lateral root primordia [Arabidopsis thaliana]
Length = 320
Score = 138 bits (347), Expect = 1e-31
Identities = 100/241 (41%), Positives = 133/241 (54%), Gaps = 36/241 (14%)
Query: 1 MSMLGLRDLVLIAPSPSSLQHHHQQNQNQNQNQNQPISHDH-NSNHSLPSSASLSVGFGI 59
M M+GLRD+ L+AP+ +HHQ N DH NSN ++A+ ++G G+
Sbjct: 1 MGMVGLRDVFLVAPA-----YHHQ-------NAGVISGSDHMNSN----AAAAAALGVGV 44
Query: 60 FPLLTATPCMPQQQSQNNEVQENPSNNNNFW-NLRMCPEVVNLPKKGVITEEENHGKIAM 118
PLLTA PQQ +++++ N NN W N E L K T + G +
Sbjct: 45 IPLLTAGT--PQQNVEDSDI--NFLGNNRRWQNNNNNHETQYLHFKS--TNQTTVGTSSN 98
Query: 119 MESEENGVYGSEYRVCQDCGNRAKKDCVFRRCRTCCKGRGYDCSTHLKSTWIPSTRRRER 178
+G G+ CQDCGN+AKK+C RRCRTCCK RG+DCSTH+KSTW+ + RRRER
Sbjct: 99 NSGSGSGASGTA--TCQDCGNQAKKECKQRRCRTCCKSRGFDCSTHVKSTWVSAARRRER 156
Query: 179 EVEMFAGGGDGEG-----CSGVKRQKGLLGS--SQNAAATSHSSNSNGTTPKSFATSSCH 231
+V M G G SG K+ + ++GS Q ATSH+S SN T P+SF TSS
Sbjct: 157 QV-MPTGANPTAGSSLSTSSGTKKPR-IVGSQQQQQQQATSHTSTSN-TPPQSFETSSSR 213
Query: 232 Q 232
Q
Sbjct: 214 Q 214
>UniRef100_Q941Y5 Putative LRP (Lateral root primordia) 1 [Oryza sativa]
Length = 340
Score = 124 bits (311), Expect = 2e-27
Identities = 78/234 (33%), Positives = 117/234 (49%), Gaps = 33/234 (14%)
Query: 9 LVLIAPSPSSLQHHHQQNQNQNQNQNQPISHDHNSNHSLPSSASLSVGFGIFPLLTATPC 68
+V++ P+ S HH H H+ + ++A+ + IFPLL+A PC
Sbjct: 6 MVVVTPAASFHHTHHH--------------HHHHEAAAAAAAAAAAAADPIFPLLSAGPC 51
Query: 69 M--PQQQSQNNEVQENPSNNNNFWNLRMCPEVVNLPKKGVITEEEN-----HGKIAMMES 121
+ P + + + + FW + P++ + G + + M+++
Sbjct: 52 VLDPDKSAASGSAIQ-------FW--QPPPQLPSSAAGGNPNPSSSAFPYLKKPLPMLDT 102
Query: 122 EENGVYGSEYRVCQDCGNRAKKDCVFRRCRTCCKGRGYDCSTHLKSTWIPSTRRRER-EV 180
CQDCGN+AKKDC +RCRTCCK RG+DCSTH+KSTW+P+ RRRER ++
Sbjct: 103 GGGSSGSGGAATCQDCGNQAKKDCGHQRCRTCCKSRGFDCSTHVKSTWVPAARRRERQQL 162
Query: 181 EMFAGGGDGEGCSGVKRQKGLLGSSQNAAATSHSSNSNGTTPKSFATSSCHQGA 234
A + +K L +SQ TSH+S SN TTP+SF T+S HQ A
Sbjct: 163 TGSASSSPATASAAAASKKPRLLTSQ--TTTSHTSTSNATTPRSFDTTSSHQDA 214
>UniRef100_Q6AV91 Hypothetical protein OJ1286_E05.15 [Oryza sativa]
Length = 453
Score = 116 bits (290), Expect = 5e-25
Identities = 59/117 (50%), Positives = 69/117 (58%), Gaps = 9/117 (7%)
Query: 125 GVYGSEYRVCQDCGNRAKKDCVFRRCRTCCKGRGYDCSTHLKSTWIPSTRRREREVEMFA 184
G GS C DCGN+AKKDCV RCRTCCK RG+DC TH++STW+P+ RRRER + A
Sbjct: 223 GPGGSGAATCHDCGNQAKKDCVHHRCRTCCKSRGFDCPTHVRSTWVPAARRRER--QQLA 280
Query: 185 GGGDGEGCSG-------VKRQKGLLGSSQNAAATSHSSNSNGTTPKSFATSSCHQGA 234
G S +K L SQ TS +S SN TTP+SF TSS HQ A
Sbjct: 281 GAASSPPTSSAFPAATTASAKKPRLLGSQTTTTTSRTSTSNATTPRSFDTSSSHQVA 337
>UniRef100_O65517 Hypothetical protein F23E13.150 [Arabidopsis thaliana]
Length = 322
Score = 103 bits (256), Expect = 5e-21
Identities = 59/167 (35%), Positives = 87/167 (51%), Gaps = 22/167 (13%)
Query: 71 QQQSQNNEVQENPSNNNNFWNLRMCPEVVNLPKKGVITEEENHGKIAMMES------EEN 124
QQQ + N V + N N N +V +P + ++ +H + ++ + +
Sbjct: 22 QQQQKTNWVWYRSNANTNNINPSSSQQVWQIPPEQMLMHHHSHPQQQSLDLYPGHQIDVS 81
Query: 125 GVYGSEYRV---CQDCGNRAKKDCVFRRCRTCCKGRGYDCSTHLKSTWIPSTRRREREVE 181
+ S + C+DCGN+AKKDC RCRTCCK RG+DCSTH++STWIP RRRER+ +
Sbjct: 82 DLATSSRSITISCRDCGNQAKKDCTHMRCRTCCKSRGFDCSTHVRSTWIPVARRRERQQQ 141
Query: 182 MF---AGGGDGEGCSGV----------KRQKGLLGSSQNAAATSHSS 215
+ +GGG G G G R L G+S ++ SHS+
Sbjct: 142 LHMSTSGGGGGSGSGGAGGGGSSIPKRHRDPTLPGTSSSSRLPSHSA 188
>UniRef100_Q5Q0G3 Hypothetical protein [Arabidopsis thaliana]
Length = 234
Score = 93.2 bits (230), Expect = 5e-18
Identities = 64/221 (28%), Positives = 97/221 (42%), Gaps = 27/221 (12%)
Query: 22 HHQQNQNQNQNQNQPISHDHNSNHSLPSSASLSVGFGIFPLLTATPCMPQQQSQNNEVQE 81
H++Q+ +Q ++ N+ S+++ + + + GF IFP P QQQ Q N
Sbjct: 12 HNKQDHHQEKDHNEDKSNNYLYLYK-DEIYNNNKGFEIFP-----PQYFQQQQQQNHA-- 63
Query: 82 NPSNNNNFWNLRMCPEVVNLPKKGVITEEENHGKIAMMESEENGVYGSEYRV-----CQD 136
+ N ++ M P N+ ++ E + G CQD
Sbjct: 64 --AAPTNLYSFGMVPSGGNINNNRSTNRSLYFNVVSDHEPVRSSTGGFTVTRQGNMNCQD 121
Query: 137 CGNRAKKDCVFRRCRTCCKGRGYDCSTHLKSTWIPSTRRREREVEMFA------------ 184
CGN+AKKDC RCRTCCK RG+DC TH+KSTW+ + +RRER+ ++
Sbjct: 122 CGNQAKKDCPHMRCRTCCKSRGFDCQTHVKSTWVSAAKRRERQAQLAVLPAKRIRDANSR 181
Query: 185 GGGDGEGCSGVKRQKGLLGSSQNAAATSHSSNSNGTTPKSF 225
GGGD + + G A T + S+ SF
Sbjct: 182 GGGDDDDDDKEDEKNDSCGGGSALACTRVVNASSSGMTMSF 222
>UniRef100_Q9XGX0 Zinc finger protein SHI-like [Arabidopsis thaliana]
Length = 331
Score = 93.2 bits (230), Expect = 5e-18
Identities = 50/119 (42%), Positives = 71/119 (59%), Gaps = 7/119 (5%)
Query: 110 EENHGKIAMMESEENGVYGSEYRVCQDCGNRAKKDCVFRRCRTCCKGRGYDCSTHLKSTW 169
E + G + MM S GS CQDCGN++KKDC RCRTCCK RG DC TH+KSTW
Sbjct: 100 ETSGGALMMMRSGS----GSGGPSCQDCGNQSKKDCSHMRCRTCCKSRGLDCPTHVKSTW 155
Query: 170 IPSTRRREREVEMFAGGGDGE-GCSGVKRQKGLLGSSQNAAATSH--SSNSNGTTPKSF 225
+P+ +RRER+ ++ G + G S KRQ+ + + + A + S+N++G +F
Sbjct: 156 VPAAKRRERQQQLSTGQQPQQLGGSVPKRQRERIPARSTSMAYTRIPSNNTSGLEVGNF 214
>UniRef100_Q5Q0G2 Hypothetical protein [Arabidopsis thaliana]
Length = 345
Score = 92.0 bits (227), Expect = 1e-17
Identities = 54/166 (32%), Positives = 83/166 (49%), Gaps = 15/166 (9%)
Query: 22 HHQQNQNQNQNQNQPISHDHNSNHSLPSSASLSVGFGIFPLLTATPCMPQQQSQNNEVQE 81
H++Q+ +Q ++ N+ S+++ + + + GF IFP P QQQ Q N
Sbjct: 12 HNKQDHHQEKDHNEDKSNNYLYLYK-DEIYNNNKGFEIFP-----PQYFQQQQQQNHA-- 63
Query: 82 NPSNNNNFWNLRMCPEVVNLPKKGVITEEENHGKIAMMESEENGVYGSEYRV-----CQD 136
+ N ++ M P N+ ++ E + G CQD
Sbjct: 64 --AAPTNLYSFGMVPSGGNINNNRSTNRSLYFNVVSDHEPVRSSTGGFTVTRQGNMNCQD 121
Query: 137 CGNRAKKDCVFRRCRTCCKGRGYDCSTHLKSTWIPSTRRREREVEM 182
CGN+AKKDC RCRTCCK RG+DC TH+KSTW+ + +RRER+ ++
Sbjct: 122 CGNQAKKDCPHMRCRTCCKSRGFDCQTHVKSTWVSAAKRRERQAQL 167
>UniRef100_Q9LQZ5 F10A5.26 [Arabidopsis thaliana]
Length = 346
Score = 92.0 bits (227), Expect = 1e-17
Identities = 58/175 (33%), Positives = 84/175 (47%), Gaps = 32/175 (18%)
Query: 26 NQNQNQNQNQPISHDHN----SNHSLPSSASL---SVGFGIFPLLTATPCMPQQQS--QN 76
N N N+ + + DH+ SN+ + + GF I+P P QQQ Q
Sbjct: 13 NSNNNKQDHHQVDKDHHHQDKSNYLYLYKDEIYNNNKGFEIWP-----PQYFQQQEHQQQ 67
Query: 77 NEVQENPSNNNNFWNLRMCPEVVNLPKKG---------VITEEENHGKIAMMESEENGVY 127
+ Q++ S NF++ M P + V+++ E G + N
Sbjct: 68 QQQQQHASAPANFYSFGMVPSGSSSNNNNNRSRSLYFNVVSDHEPGGFTVTRQGGMN--- 124
Query: 128 GSEYRVCQDCGNRAKKDCVFRRCRTCCKGRGYDCSTHLKSTWIPSTRRREREVEM 182
CQDCGN+AKKDC RCRTCCK RG+ C TH+KSTW+P+ +RRER ++
Sbjct: 125 ------CQDCGNQAKKDCPHMRCRTCCKSRGFHCQTHVKSTWVPAAKRRERLAQL 173
>UniRef100_Q9SD40 Hypothetical protein F24M12.100 [Arabidopsis thaliana]
Length = 252
Score = 92.0 bits (227), Expect = 1e-17
Identities = 74/234 (31%), Positives = 104/234 (43%), Gaps = 36/234 (15%)
Query: 14 PSPSSLQHHHQQNQNQNQNQNQPISHDHNSNHSLPSSASLSVG-FGIFPLLTATPCMPQQ 72
P P S +N N N N N N N + PSS++ ++G ++ M Q
Sbjct: 31 PPPVSEAWLWYRNPNVNANANT------NVNANAPSSSNAALGTLELWQNHNQQEIMFQH 84
Query: 73 QSQNNEVQ---------ENPSNNNNFWNLRMCPEVVNLPKKGVITEEENHGKIAMMESEE 123
Q + PSN+N F G + AMM
Sbjct: 85 QQHQQRLDLYSSAAGLGVGPSNHNQF------------DISGETSTAGAGRAAAMMMIRS 132
Query: 124 NGVYGSEYRV-CQDCGNRAKKDCVFRRCRTCCKGRGYDCSTHLKSTWIPSTRRREREVEM 182
G G V CQDCGN+AKKDC RCRTCCK RG++CSTH++STW+P+ +RRER+ ++
Sbjct: 133 GGSGGGSGGVSCQDCGNQAKKDCSHMRCRTCCKSRGFECSTHVRSTWVPAAKRRERQQQL 192
Query: 183 FAGGGDGE---GCSGVKR-QKGLLGSSQNAAAT---SHSSNSNGTTPKSFATSS 229
+ G S KR ++ L +S + T SHS G P ++S+
Sbjct: 193 ATVQPQTQLPRGESVPKRHRENLPATSSSLVCTRIPSHSGLEVGNFPAEVSSSA 246
>UniRef100_Q9FXH7 F6F9.16 protein [Arabidopsis thaliana]
Length = 345
Score = 92.0 bits (227), Expect = 1e-17
Identities = 54/166 (32%), Positives = 83/166 (49%), Gaps = 15/166 (9%)
Query: 22 HHQQNQNQNQNQNQPISHDHNSNHSLPSSASLSVGFGIFPLLTATPCMPQQQSQNNEVQE 81
H++Q+ +Q ++ N+ S+++ + + + GF IFP P QQQ Q N
Sbjct: 12 HNKQDHHQEKDHNEDKSNNYLYLYK-DEIYNNNKGFEIFP-----PQYFQQQQQQNHA-- 63
Query: 82 NPSNNNNFWNLRMCPEVVNLPKKGVITEEENHGKIAMMESEENGVYGSEYRV-----CQD 136
+ N ++ M P N+ ++ E + G CQD
Sbjct: 64 --AAPTNLYSFGMVPSGGNINNNRSTNRSLYFNVVSDHEPVRSSTGGFTVTRQGNMNCQD 121
Query: 137 CGNRAKKDCVFRRCRTCCKGRGYDCSTHLKSTWIPSTRRREREVEM 182
CGN+AKKDC RCRTCCK RG+DC TH+KSTW+ + +RRER+ ++
Sbjct: 122 CGNQAKKDCPHMRCRTCCKSRGFDCQTHVKSTWVSAAKRRERQAQL 167
>UniRef100_Q6YT10 Putative lateral root primordia [Oryza sativa]
Length = 324
Score = 89.0 bits (219), Expect = 9e-17
Identities = 33/46 (71%), Positives = 41/46 (88%)
Query: 134 CQDCGNRAKKDCVFRRCRTCCKGRGYDCSTHLKSTWIPSTRRRERE 179
CQDCGN+AKKDC RCRTCCK RG+DC+TH+KSTW+P+ +RRER+
Sbjct: 116 CQDCGNQAKKDCTHLRCRTCCKSRGFDCATHVKSTWVPAAKRRERQ 161
>UniRef100_Q652K4 Putative LRP1 [Oryza sativa]
Length = 315
Score = 88.6 bits (218), Expect = 1e-16
Identities = 38/74 (51%), Positives = 49/74 (65%), Gaps = 8/74 (10%)
Query: 134 CQDCGNRAKKDCVFRRCRTCCKGRGYDCSTHLKSTWIPSTRRREREVEMFA--------G 185
CQDCGN+AKKDC RCRTCCK RG+ C+TH+KSTW+P+ +RRER+ ++ A
Sbjct: 97 CQDCGNQAKKDCTHMRCRTCCKSRGFACATHVKSTWVPAAKRRERQQQLAALAASAAATA 156
Query: 186 GGDGEGCSGVKRQK 199
GG G KR +
Sbjct: 157 GGAGPSRDPTKRPR 170
>UniRef100_Q9M2U4 Hypothetical protein T12E18_120 [Arabidopsis thaliana]
Length = 183
Score = 87.8 bits (216), Expect = 2e-16
Identities = 36/55 (65%), Positives = 44/55 (79%), Gaps = 2/55 (3%)
Query: 122 EENGVYGSEYRVCQDCGNRAKKDCVFRRCRTCCKGRGYDCSTHLKSTWIPSTRRR 176
E+N G +VC+DCGNRAKK+C+F RCRTCCK RGY+C TH+KSTWIPS+ R
Sbjct: 31 EDNNTVGE--KVCRDCGNRAKKECLFERCRTCCKSRGYNCVTHVKSTWIPSSATR 83
>UniRef100_Q5Z5Y2 Putative LRP1 [Oryza sativa]
Length = 384
Score = 85.9 bits (211), Expect = 8e-16
Identities = 32/51 (62%), Positives = 42/51 (81%)
Query: 134 CQDCGNRAKKDCVFRRCRTCCKGRGYDCSTHLKSTWIPSTRRREREVEMFA 184
CQDCGN AKKDC RCRTCC+ RG+ C+TH+KSTW+P+ +RRER+ ++ A
Sbjct: 133 CQDCGNNAKKDCSHLRCRTCCRSRGFSCATHVKSTWVPAAKRRERQQQLAA 183
>UniRef100_Q9SJT8 Hypothetical protein At2g21400 [Arabidopsis thaliana]
Length = 174
Score = 80.1 bits (196), Expect = 4e-14
Identities = 29/44 (65%), Positives = 37/44 (83%)
Query: 132 RVCQDCGNRAKKDCVFRRCRTCCKGRGYDCSTHLKSTWIPSTRR 175
R C+DCGN+AKKDCV+ RCRTCCK + + C TH+KSTW+P+ RR
Sbjct: 7 RKCEDCGNQAKKDCVYMRCRTCCKSKAFHCQTHIKSTWVPAYRR 50
>UniRef100_Q70I38 Hypothetical protein [Lotus japonicus]
Length = 246
Score = 78.6 bits (192), Expect = 1e-13
Identities = 35/66 (53%), Positives = 46/66 (69%), Gaps = 5/66 (7%)
Query: 116 IAMMESEENGVYGSEYRVCQDCGNRAKKDCVFRRCRTCCKGRGYDCSTHLKSTWIPSTRR 175
++MM +E V GS CQ+CGN+AKK C + RCRTCC +G+ C TH++STWIP RR
Sbjct: 1 MSMMLGQE--VKGSR---CQECGNQAKKGCAYSRCRTCCNNKGFQCQTHVRSTWIPVDRR 55
Query: 176 REREVE 181
R R +E
Sbjct: 56 RHRLME 61
>UniRef100_Q9SI19 Hypothetical protein At2g18120 [Arabidopsis thaliana]
Length = 222
Score = 78.2 bits (191), Expect = 2e-13
Identities = 32/61 (52%), Positives = 41/61 (66%)
Query: 134 CQDCGNRAKKDCVFRRCRTCCKGRGYDCSTHLKSTWIPSTRRREREVEMFAGGGDGEGCS 193
CQ+CGN+AKK C RCRTCCK G C TH++STWIP +RRER+ ++ + G S
Sbjct: 72 CQECGNQAKKGCTHGRCRTCCKSNGLHCPTHVRSTWIPIAKRRERQQQLQTPTSNPTGGS 131
Query: 194 G 194
G
Sbjct: 132 G 132
>UniRef100_Q8GUY8 Hypothetical protein F3K23.16 [Arabidopsis thaliana]
Length = 162
Score = 67.4 bits (163), Expect = 3e-10
Identities = 24/38 (63%), Positives = 32/38 (84%)
Query: 138 GNRAKKDCVFRRCRTCCKGRGYDCSTHLKSTWIPSTRR 175
GN+AKKDCV+ RCRTCCK + + C TH++STW+P+ RR
Sbjct: 1 GNQAKKDCVYMRCRTCCKSKAFHCQTHIESTWVPAYRR 38
>UniRef100_Q86M87 ComB [Dictyostelium discoideum]
Length = 2107
Score = 44.7 bits (104), Expect = 0.002
Identities = 44/206 (21%), Positives = 79/206 (37%), Gaps = 26/206 (12%)
Query: 17 SSLQHHHQQNQNQNQNQNQPISHDHNSNHSLPSSASLSVGFGIFPLLTATPCMPQQQSQN 76
+S+ +++ N N N N N ++++N+N+S S+ + + N
Sbjct: 486 NSINNNNNNNNNNNNNNNNNNNNNNNNNNSSISNNN------------------NNNNNN 527
Query: 77 NEVQENPSNNNNFWNLRMCPEVVNLPKKGVITE-EENHGKIAMMESEENGVYGSEYRVCQ 135
N N +NNNN N N+ + T+ ++ M + N G+
Sbjct: 528 NNNNNNNNNNNNNNNNNSSSSNNNINNNNINTDNNSSNNNNNNMNNNNNNNSGNNKNDKN 587
Query: 136 DCGNRAKKDCVFRRCRTCCKGRGYDCSTHLKSTWIPSTRRREREVEMFAGGGDGEGCSGV 195
D KK VF+ + G K + +ERE E + E GV
Sbjct: 588 DSNGSIKKKGVFKNLKN--SGDKEKDKEKEKEKEKEKEKEKEREKEK-----EKENAKGV 640
Query: 196 KRQKGLLGSSQNAAATSHSSNSNGTT 221
K +L + N++ +++ +NGTT
Sbjct: 641 TSPKNILPINLNSSFLPNNNGNNGTT 666
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.313 0.128 0.389
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 427,364,322
Number of Sequences: 2790947
Number of extensions: 18874541
Number of successful extensions: 108572
Number of sequences better than 10.0: 417
Number of HSP's better than 10.0 without gapping: 123
Number of HSP's successfully gapped in prelim test: 323
Number of HSP's that attempted gapping in prelim test: 100325
Number of HSP's gapped (non-prelim): 3631
length of query: 235
length of database: 848,049,833
effective HSP length: 123
effective length of query: 112
effective length of database: 504,763,352
effective search space: 56533495424
effective search space used: 56533495424
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 73 (32.7 bits)
Medicago: description of AC146853.5


