

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146818.6 + phase: 0 /pseudo
(642 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
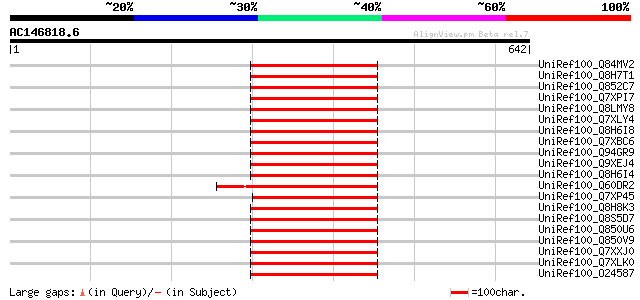
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q84MV2 Putative gag-pol polyprotein [Oryza sativa] 184 7e-45
UniRef100_Q8H7T1 Putative Zea mays retrotransposon Opie-2 [Oryza... 183 1e-44
UniRef100_Q852C7 Putative gag-pol polyprotein [Oryza sativa] 183 1e-44
UniRef100_Q7XPI7 OSJNBb0004A17.2 protein [Oryza sativa] 183 1e-44
UniRef100_Q8LMY8 Putative polyprotein [Oryza sativa] 183 2e-44
UniRef100_Q7XLY4 OSJNBa0042I15.6 protein [Oryza sativa] 183 2e-44
UniRef100_Q8H6I8 Putative gag-pol polyprotein [Zea mays] 182 2e-44
UniRef100_Q7XBC6 Putative copia-type pol polyprotein [Zea mays] 182 2e-44
UniRef100_Q94GR9 Putative gag-pol polyprotein [Oryza sativa] 182 3e-44
UniRef100_Q9XEJ4 Copia-type pol polyprotein [Zea mays] 181 5e-44
UniRef100_Q8H6I4 Putative gag-pol polyprotein [Zea mays] 181 5e-44
UniRef100_Q60DR2 Putative polyprotein [Oryza sativa] 181 6e-44
UniRef100_Q7XP45 OSJNBa0063G07.6 protein [Oryza sativa] 181 6e-44
UniRef100_Q8H8K3 Putative retroelement [Oryza sativa] 180 1e-43
UniRef100_Q8S5D7 Putative gag-pol polyprotein [Oryza sativa] 180 1e-43
UniRef100_Q850U6 Putative polyprotein [Oryza sativa] 179 2e-43
UniRef100_Q850V9 Putative polyprotein [Oryza sativa] 179 2e-43
UniRef100_Q7XXJ0 OSJNBa0094O15.12 protein [Oryza sativa] 179 3e-43
UniRef100_Q7XLK0 OSJNBa0009K15.10 protein [Oryza sativa] 178 4e-43
UniRef100_O24587 Pol protein [Zea mays] 177 1e-42
>UniRef100_Q84MV2 Putative gag-pol polyprotein [Oryza sativa]
Length = 1975
Score = 184 bits (467), Expect = 7e-45
Identities = 96/158 (60%), Positives = 112/158 (70%)
Query: 298 LIKNDFKRGQVDTTLFRGTLKKYILVVQIYVDDIIFGSTNASLCKEFSK*MQDEFEKSMM 357
L+ DFK G+VDTTL + +V QIYVDDIIFGSTN CKEF M EFE SM+
Sbjct: 1629 LLPKDFKIGKVDTTLSTKIIDDDFVVCQIYVDDIIFGSTNEVFCKEFGDMMSREFEMSMV 1688
Query: 358 GELKFFLGIQINQRKDGVYVHQTKYTKELLKKFKLEDCKVMNTPMHPTCT*SKEDIGTKV 417
GEL FFLG+QI Q KDG +V QTKY K+LLK+F LED K + TPM ++ G V
Sbjct: 1689 GELSFFLGLQIKQLKDGTFVSQTKYIKDLLKRFGLEDAKPIKTPMATNGHLDLDEGGKPV 1748
Query: 418 DQKLYIGMIGSLLYLTASRPDILFSVCLCARFQSNPKR 455
D KLY MIGSLLYLTASRPDI+FSVC+CARFQ+ PK+
Sbjct: 1749 DLKLYHSMIGSLLYLTASRPDIMFSVCMCARFQAAPKK 1786
>UniRef100_Q8H7T1 Putative Zea mays retrotransposon Opie-2 [Oryza sativa]
Length = 2145
Score = 183 bits (465), Expect = 1e-44
Identities = 94/157 (59%), Positives = 117/157 (73%)
Query: 298 LIKNDFKRGQVDTTLFRGTLKKYILVVQIYVDDIIFGSTNASLCKEFSK*MQDEFEKSMM 357
L++N F+ G VD TLF L+VQIYVDDIIFG ++ +L +FS M EFE SMM
Sbjct: 1450 LLQNGFEMGAVDKTLFTLHSGIDFLLVQIYVDDIIFGGSSHALVAQFSDVMSREFEMSMM 1509
Query: 358 GELKFFLGIQINQRKDGVYVHQTKYTKELLKKFKLEDCKVMNTPMHPTCT*SKEDIGTKV 417
GEL FFLG+QI Q K+G++VHQTKY+KELLKKF + DCK + TPM T + ++ G +V
Sbjct: 1510 GELTFFLGLQIKQTKEGIFVHQTKYSKELLKKFDMADCKPIATPMATTSSLGPDEDGEEV 1569
Query: 418 DQKLYIGMIGSLLYLTASRPDILFSVCLCARFQSNPK 454
DQ+ Y MIGSLLYLTASRPDI FSVCLCARFQ++P+
Sbjct: 1570 DQREYRSMIGSLLYLTASRPDIHFSVCLCARFQASPR 1606
>UniRef100_Q852C7 Putative gag-pol polyprotein [Oryza sativa]
Length = 1969
Score = 183 bits (465), Expect = 1e-44
Identities = 94/157 (59%), Positives = 117/157 (73%)
Query: 298 LIKNDFKRGQVDTTLFRGTLKKYILVVQIYVDDIIFGSTNASLCKEFSK*MQDEFEKSMM 357
L++N F+ G VD TLF L+VQIYVDDIIFG ++ +L +FS M EFE SMM
Sbjct: 1450 LLQNGFEMGAVDKTLFTLHSGIDFLLVQIYVDDIIFGGSSHALVAQFSDVMSREFEMSMM 1509
Query: 358 GELKFFLGIQINQRKDGVYVHQTKYTKELLKKFKLEDCKVMNTPMHPTCT*SKEDIGTKV 417
GEL FFLG+QI Q K+G++VHQTKY+KELLKKF + DCK + TPM T + ++ G +V
Sbjct: 1510 GELTFFLGLQIKQTKEGIFVHQTKYSKELLKKFDMADCKPIATPMATTSSLGPDEDGEEV 1569
Query: 418 DQKLYIGMIGSLLYLTASRPDILFSVCLCARFQSNPK 454
DQ+ Y MIGSLLYLTASRPDI FSVCLCARFQ++P+
Sbjct: 1570 DQREYRSMIGSLLYLTASRPDIHFSVCLCARFQASPR 1606
>UniRef100_Q7XPI7 OSJNBb0004A17.2 protein [Oryza sativa]
Length = 1877
Score = 183 bits (465), Expect = 1e-44
Identities = 94/157 (59%), Positives = 117/157 (73%)
Query: 298 LIKNDFKRGQVDTTLFRGTLKKYILVVQIYVDDIIFGSTNASLCKEFSK*MQDEFEKSMM 357
L++N F+ G VD TLF L+VQIYVDDIIFG ++ +L +FS M EFE SMM
Sbjct: 1532 LLQNGFEMGAVDKTLFTLHSGIDFLLVQIYVDDIIFGGSSHALVAQFSDVMSREFEMSMM 1591
Query: 358 GELKFFLGIQINQRKDGVYVHQTKYTKELLKKFKLEDCKVMNTPMHPTCT*SKEDIGTKV 417
GEL FFLG+QI Q K+G++VHQTKY+KELLKKF + DCK + TPM T + ++ G +V
Sbjct: 1592 GELTFFLGLQIKQTKEGIFVHQTKYSKELLKKFDMADCKPIATPMATTSSLGPDEDGEEV 1651
Query: 418 DQKLYIGMIGSLLYLTASRPDILFSVCLCARFQSNPK 454
DQ+ Y MIGSLLYLTASRPDI FSVCLCARFQ++P+
Sbjct: 1652 DQREYRSMIGSLLYLTASRPDIHFSVCLCARFQASPR 1688
>UniRef100_Q8LMY8 Putative polyprotein [Oryza sativa]
Length = 1584
Score = 183 bits (464), Expect = 2e-44
Identities = 96/157 (61%), Positives = 110/157 (69%)
Query: 298 LIKNDFKRGQVDTTLFRGTLKKYILVVQIYVDDIIFGSTNASLCKEFSK*MQDEFEKSMM 357
L+ DFK G+V+TTLF + V QIYVDDIIFGSTN CKEF M EFE SM+
Sbjct: 1239 LLSKDFKIGKVNTTLFTKIIGDDFFVCQIYVDDIIFGSTNEVFCKEFGDMMSREFEMSMI 1298
Query: 358 GELKFFLGIQINQRKDGVYVHQTKYTKELLKKFKLEDCKVMNTPMHPTCT*SKEDIGTKV 417
GEL FFLG+QI Q KDG +V QTKY K+LLK+F LED K + TPM + G V
Sbjct: 1299 GELNFFLGLQIKQLKDGTFVSQTKYIKDLLKRFGLEDAKPIKTPMATNGHLDLDKGGKPV 1358
Query: 418 DQKLYIGMIGSLLYLTASRPDILFSVCLCARFQSNPK 454
D KLY MIGSLLYLTASRPDI+FSVC+CARFQ+ PK
Sbjct: 1359 DLKLYRSMIGSLLYLTASRPDIMFSVCMCARFQAAPK 1395
>UniRef100_Q7XLY4 OSJNBa0042I15.6 protein [Oryza sativa]
Length = 1510
Score = 183 bits (464), Expect = 2e-44
Identities = 91/157 (57%), Positives = 111/157 (69%)
Query: 298 LIKNDFKRGQVDTTLFRGTLKKYILVVQIYVDDIIFGSTNASLCKEFSK*MQDEFEKSMM 357
L+KN F+ G DTTLF K + + QIYVDDIIFGSTNAS C+EFS M FE SMM
Sbjct: 1160 LLKNGFEIGNADTTLFTKKFKSDLFICQIYVDDIIFGSTNASFCEEFSSIMTKRFEMSMM 1219
Query: 358 GELKFFLGIQINQRKDGVYVHQTKYTKELLKKFKLEDCKVMNTPMHPTCT*SKEDIGTKV 417
GEL FFL +Q+ Q ++G ++ QTKY K++LKKF +ED K + TPM +D G V
Sbjct: 1220 GELTFFLWLQVKQAQEGTFISQTKYVKDILKKFGMEDAKPIKTPMPTNGHLDLDDNGKCV 1279
Query: 418 DQKLYIGMIGSLLYLTASRPDILFSVCLCARFQSNPK 454
DQK+Y MIGSLLYL ASRPDI+ SVC+CARFQ+ PK
Sbjct: 1280 DQKVYRSMIGSLLYLCASRPDIMLSVCMCARFQAEPK 1316
>UniRef100_Q8H6I8 Putative gag-pol polyprotein [Zea mays]
Length = 1892
Score = 182 bits (463), Expect = 2e-44
Identities = 91/157 (57%), Positives = 114/157 (71%)
Query: 298 LIKNDFKRGQVDTTLFRGTLKKYILVVQIYVDDIIFGSTNASLCKEFSK*MQDEFEKSMM 357
LI N FK G+ D TLF TL+ + V QIYVDDIIFGSTN S C+EFS+ M +FE SMM
Sbjct: 1546 LIANGFKVGKADPTLFTKTLENDLFVCQIYVDDIIFGSTNESTCEEFSRIMTQKFEMSMM 1605
Query: 358 GELKFFLGIQINQRKDGVYVHQTKYTKELLKKFKLEDCKVMNTPMHPTCT*SKEDIGTKV 417
GELK+FLG Q+ Q ++G ++ QTKYT+++L KF ++D K + TPM + G V
Sbjct: 1606 GELKYFLGFQVKQLREGTFISQTKYTQDILAKFGMKDAKPIKTPMGTNGHLDLDTGGKSV 1665
Query: 418 DQKLYIGMIGSLLYLTASRPDILFSVCLCARFQSNPK 454
DQK+Y MIGSLLYL ASRPDI+ SVC+CARFQS+PK
Sbjct: 1666 DQKVYRSMIGSLLYLCASRPDIMLSVCMCARFQSDPK 1702
>UniRef100_Q7XBC6 Putative copia-type pol polyprotein [Zea mays]
Length = 1760
Score = 182 bits (463), Expect = 2e-44
Identities = 91/157 (57%), Positives = 114/157 (71%)
Query: 298 LIKNDFKRGQVDTTLFRGTLKKYILVVQIYVDDIIFGSTNASLCKEFSK*MQDEFEKSMM 357
LI N FK G+ D TLF TL+ + V QIYVDDIIFGSTN S C+EFS+ M +FE SMM
Sbjct: 1414 LIANGFKVGKADPTLFTKTLENDLFVCQIYVDDIIFGSTNESTCEEFSRIMTQKFEMSMM 1473
Query: 358 GELKFFLGIQINQRKDGVYVHQTKYTKELLKKFKLEDCKVMNTPMHPTCT*SKEDIGTKV 417
GELK+FLG Q+ Q ++G ++ QTKYT+++L KF ++D K + TPM + G V
Sbjct: 1474 GELKYFLGFQVKQLQEGTFISQTKYTQDILSKFGMKDAKPIKTPMGTNGHLDLDTGGKSV 1533
Query: 418 DQKLYIGMIGSLLYLTASRPDILFSVCLCARFQSNPK 454
DQK+Y MIGSLLYL ASRPDI+ SVC+CARFQS+PK
Sbjct: 1534 DQKVYRSMIGSLLYLCASRPDIMLSVCMCARFQSDPK 1570
>UniRef100_Q94GR9 Putative gag-pol polyprotein [Oryza sativa]
Length = 1700
Score = 182 bits (461), Expect = 3e-44
Identities = 95/157 (60%), Positives = 110/157 (69%)
Query: 298 LIKNDFKRGQVDTTLFRGTLKKYILVVQIYVDDIIFGSTNASLCKEFSK*MQDEFEKSMM 357
L+ DFK G+VDTTLF + V QIYVDDIIF STN CKEF M EFE SM+
Sbjct: 1113 LLSKDFKIGKVDTTLFTKIIGDDFFVCQIYVDDIIFSSTNEVFCKEFGDMMSREFEMSMI 1172
Query: 358 GELKFFLGIQINQRKDGVYVHQTKYTKELLKKFKLEDCKVMNTPMHPTCT*SKEDIGTKV 417
GEL FFLG+QI Q KDG +V QTKY K+LLK+F LED K + TPM ++ G V
Sbjct: 1173 GELSFFLGLQIKQLKDGTFVSQTKYIKDLLKRFGLEDAKPIKTPMATNGHLDLDEGGKLV 1232
Query: 418 DQKLYIGMIGSLLYLTASRPDILFSVCLCARFQSNPK 454
D KLY MIGSLLYLTASRPDI+FS+C+CARFQ+ PK
Sbjct: 1233 DLKLYRFMIGSLLYLTASRPDIMFSICMCARFQAAPK 1269
>UniRef100_Q9XEJ4 Copia-type pol polyprotein [Zea mays]
Length = 1063
Score = 181 bits (460), Expect = 5e-44
Identities = 91/157 (57%), Positives = 114/157 (71%)
Query: 298 LIKNDFKRGQVDTTLFRGTLKKYILVVQIYVDDIIFGSTNASLCKEFSK*MQDEFEKSMM 357
LI N FK G+ D TLF TL+ + V QIYVDDIIFGSTN S C+EFS+ M +FE SMM
Sbjct: 717 LIANGFKVGKADPTLFTKTLENDLFVCQIYVDDIIFGSTNKSTCEEFSRIMTQKFEMSMM 776
Query: 358 GELKFFLGIQINQRKDGVYVHQTKYTKELLKKFKLEDCKVMNTPMHPTCT*SKEDIGTKV 417
GELK+FLG Q+ Q ++G ++ QTKYT+++L KF ++D K + TPM + G V
Sbjct: 777 GELKYFLGFQVKQLQEGTFICQTKYTQDILTKFGMKDAKPIKTPMGTNGHLDLDTGGKSV 836
Query: 418 DQKLYIGMIGSLLYLTASRPDILFSVCLCARFQSNPK 454
DQK+Y MIGSLLYL ASRPDI+ SVC+CARFQS+PK
Sbjct: 837 DQKVYRSMIGSLLYLCASRPDIMLSVCMCARFQSDPK 873
>UniRef100_Q8H6I4 Putative gag-pol polyprotein [Zea mays]
Length = 2319
Score = 181 bits (460), Expect = 5e-44
Identities = 92/157 (58%), Positives = 111/157 (70%)
Query: 298 LIKNDFKRGQVDTTLFRGTLKKYILVVQIYVDDIIFGSTNASLCKEFSK*MQDEFEKSMM 357
LIKN F G+ D+TLF + + V QIYVDDIIFGSTN C+EFSK M + FE SMM
Sbjct: 1477 LIKNGFTIGKADSTLFTRKVDNELFVCQIYVDDIIFGSTNEKFCEEFSKVMTNRFEMSMM 1536
Query: 358 GELKFFLGIQINQRKDGVYVHQTKYTKELLKKFKLEDCKVMNTPMHPTCT*SKEDIGTKV 417
GELK+FLG Q+ Q K+G ++ QTKYT+++LKKF +E K TPM + G V
Sbjct: 1537 GELKYFLGFQVKQLKEGTFLCQTKYTQDMLKKFGMEKAKHAKTPMPSNGHLDLNEEGKPV 1596
Query: 418 DQKLYIGMIGSLLYLTASRPDILFSVCLCARFQSNPK 454
DQKLY MIGSLLYL ASRPDI+ SVC+CARFQ+NPK
Sbjct: 1597 DQKLYRSMIGSLLYLCASRPDIMLSVCMCARFQANPK 1633
>UniRef100_Q60DR2 Putative polyprotein [Oryza sativa]
Length = 1577
Score = 181 bits (459), Expect = 6e-44
Identities = 102/198 (51%), Positives = 123/198 (61%), Gaps = 1/198 (0%)
Query: 257 ETKLYLLNKVTVNKKVLSTLKNLLQLQDWKQSGYFYPMQ*ILIKNDFKRGQVDTTLFRGT 316
+ K LN V K L L+ ++ Y ++ L+ DFK G+VD TLF
Sbjct: 1192 DVKSAFLNDTKYPNHVYKLSKALYGLRQAPRAWY-ERLRDFLLSKDFKIGKVDITLFTKI 1250
Query: 317 LKKYILVVQIYVDDIIFGSTNASLCKEFSK*MQDEFEKSMMGELKFFLGIQINQRKDGVY 376
+ V QIYVDDIIFGSTN CKEF M EFE SM+GEL FFLG+QI Q K+G +
Sbjct: 1251 IGDDFFVYQIYVDDIIFGSTNEVFCKEFGDMMSREFEMSMIGELSFFLGLQIKQLKNGTF 1310
Query: 377 VHQTKYTKELLKKFKLEDCKVMNTPMHPTCT*SKEDIGTKVDQKLYIGMIGSLLYLTASR 436
V QTKY K+LLK+F LED K + TPM ++ G VD KLY MIGSLLYLT SR
Sbjct: 1311 VSQTKYIKDLLKRFGLEDAKPIKTPMATNGHLDLDEGGKPVDLKLYRSMIGSLLYLTVSR 1370
Query: 437 PDILFSVCLCARFQSNPK 454
PDI+FSVC+CARFQ+ PK
Sbjct: 1371 PDIMFSVCMCARFQAAPK 1388
>UniRef100_Q7XP45 OSJNBa0063G07.6 protein [Oryza sativa]
Length = 1539
Score = 181 bits (459), Expect = 6e-44
Identities = 95/154 (61%), Positives = 109/154 (70%)
Query: 301 NDFKRGQVDTTLFRGTLKKYILVVQIYVDDIIFGSTNASLCKEFSK*MQDEFEKSMMGEL 360
+DFK G+VDTTLF + V QIYVDDIIFGSTN CKEF M EFE SM+GEL
Sbjct: 1197 HDFKIGKVDTTLFTKIIGDDFFVCQIYVDDIIFGSTNEVFCKEFGDMMSREFEMSMIGEL 1256
Query: 361 KFFLGIQINQRKDGVYVHQTKYTKELLKKFKLEDCKVMNTPMHPTCT*SKEDIGTKVDQK 420
FFLG+QI Q KDG +V QTKY K+LLK+F LED K + TPM ++ G VD K
Sbjct: 1257 SFFLGLQIKQLKDGTFVSQTKYIKDLLKRFGLEDAKPIKTPMATNGHLDLDEGGKPVDLK 1316
Query: 421 LYIGMIGSLLYLTASRPDILFSVCLCARFQSNPK 454
LY MIGSLLYLTASRPDI+FSVC+CA FQ+ PK
Sbjct: 1317 LYRSMIGSLLYLTASRPDIMFSVCMCAWFQAAPK 1350
>UniRef100_Q8H8K3 Putative retroelement [Oryza sativa]
Length = 1299
Score = 180 bits (457), Expect = 1e-43
Identities = 95/157 (60%), Positives = 109/157 (68%)
Query: 298 LIKNDFKRGQVDTTLFRGTLKKYILVVQIYVDDIIFGSTNASLCKEFSK*MQDEFEKSMM 357
L+ DFK G+VDTTLF + V QIYVDDIIFGSTN CKEF M EFE SM+
Sbjct: 954 LLSKDFKIGKVDTTLFTKIIGDDFFVCQIYVDDIIFGSTNEVFCKEFGDMMSREFEMSMI 1013
Query: 358 GELKFFLGIQINQRKDGVYVHQTKYTKELLKKFKLEDCKVMNTPMHPTCT*SKEDIGTKV 417
GEL FFL +QI Q KDG ++ QTKY K+LLK+F LED K + TPM ++ G V
Sbjct: 1014 GELSFFLRLQIKQLKDGTFLSQTKYIKDLLKRFGLEDAKPIKTPMVTNGHLDLDEGGKPV 1073
Query: 418 DQKLYIGMIGSLLYLTASRPDILFSVCLCARFQSNPK 454
D KLY MIGSLLYLTASRPDI+FSVC+CARFQ PK
Sbjct: 1074 DLKLYHSMIGSLLYLTASRPDIMFSVCMCARFQVAPK 1110
>UniRef100_Q8S5D7 Putative gag-pol polyprotein [Oryza sativa]
Length = 1627
Score = 180 bits (457), Expect = 1e-43
Identities = 94/157 (59%), Positives = 110/157 (69%)
Query: 298 LIKNDFKRGQVDTTLFRGTLKKYILVVQIYVDDIIFGSTNASLCKEFSK*MQDEFEKSMM 357
L+ FK G+VDTTLF + V QIYVDDIIFGSTN CKEF M EFE SM+
Sbjct: 1110 LLSKYFKIGKVDTTLFTKIIGDDFFVCQIYVDDIIFGSTNEVFCKEFGNMMSREFEMSMI 1169
Query: 358 GELKFFLGIQINQRKDGVYVHQTKYTKELLKKFKLEDCKVMNTPMHPTCT*SKEDIGTKV 417
GEL FFLG+QI Q KDG +V QTKY K+LLK+F LED K + TP+ ++ G V
Sbjct: 1170 GELSFFLGVQIKQLKDGTFVSQTKYIKDLLKRFGLEDAKSIKTPIATNGHLDLDEGGKPV 1229
Query: 418 DQKLYIGMIGSLLYLTASRPDILFSVCLCARFQSNPK 454
D KLY MIGSLL+LTASRPDI+FSVC+CARFQ+ PK
Sbjct: 1230 DLKLYRSMIGSLLFLTASRPDIIFSVCMCARFQAAPK 1266
>UniRef100_Q850U6 Putative polyprotein [Oryza sativa]
Length = 374
Score = 179 bits (455), Expect = 2e-43
Identities = 90/156 (57%), Positives = 111/156 (70%)
Query: 299 IKNDFKRGQVDTTLFRGTLKKYILVVQIYVDDIIFGSTNASLCKEFSK*MQDEFEKSMMG 358
+KN F+ G+ DTTLF +K + + QIYVDDIIFGSTNAS C+EFS M FE SMM
Sbjct: 151 LKNGFEIGKGDTTLFTKKIKSDLFICQIYVDDIIFGSTNASFCEEFSSIMTKRFEMSMMD 210
Query: 359 ELKFFLGIQINQRKDGVYVHQTKYTKELLKKFKLEDCKVMNTPMHPTCT*SKEDIGTKVD 418
EL FFLG+Q+ Q ++G + QTKY K++LKKF++ED K + TPM D G VD
Sbjct: 211 ELTFFLGLQVKQAQEGTCISQTKYVKDILKKFEMEDAKPIRTPMPINGHLDLNDNGKCVD 270
Query: 419 QKLYIGMIGSLLYLTASRPDILFSVCLCARFQSNPK 454
QK+Y MIGSLLYL ASRPDI+ SVC+CARFQ+ PK
Sbjct: 271 QKVYRSMIGSLLYLCASRPDIMLSVCMCARFQAEPK 306
>UniRef100_Q850V9 Putative polyprotein [Oryza sativa]
Length = 1128
Score = 179 bits (454), Expect = 2e-43
Identities = 95/157 (60%), Positives = 109/157 (68%)
Query: 298 LIKNDFKRGQVDTTLFRGTLKKYILVVQIYVDDIIFGSTNASLCKEFSK*MQDEFEKSMM 357
L+ DFK G+VDTTLF + V QIYVDDIIFGSTN CKEF M EFE SM+
Sbjct: 781 LLSKDFKIGKVDTTLFTKIIGDDFFVCQIYVDDIIFGSTNEVFCKEFGDMMSREFEMSMI 840
Query: 358 GELKFFLGIQINQRKDGVYVHQTKYTKELLKKFKLEDCKVMNTPMHPTCT*SKEDIGTKV 417
EL FFLG+QI Q KDG +V QTKY K+LLK+F LED K + TPM ++ G V
Sbjct: 841 EELSFFLGLQIKQLKDGTFVSQTKYIKDLLKRFGLEDAKPIKTPMATNWHLDLDEGGKPV 900
Query: 418 DQKLYIGMIGSLLYLTASRPDILFSVCLCARFQSNPK 454
D KLY MIGSLLYLTASRPDI+FSVC+ ARFQ+ PK
Sbjct: 901 DLKLYRSMIGSLLYLTASRPDIMFSVCMYARFQAAPK 937
>UniRef100_Q7XXJ0 OSJNBa0094O15.12 protein [Oryza sativa]
Length = 1739
Score = 179 bits (453), Expect = 3e-43
Identities = 94/157 (59%), Positives = 108/157 (67%)
Query: 298 LIKNDFKRGQVDTTLFRGTLKKYILVVQIYVDDIIFGSTNASLCKEFSK*MQDEFEKSMM 357
L+ DFK G+VDTTLF + V QIYVDD IFGSTN CK F M EFE SM+
Sbjct: 1024 LLSKDFKIGKVDTTLFTKIIGDDFFVCQIYVDDFIFGSTNKVFCKGFGDMMSREFEMSMI 1083
Query: 358 GELKFFLGIQINQRKDGVYVHQTKYTKELLKKFKLEDCKVMNTPMHPTCT*SKEDIGTKV 417
EL FFLG+QI Q KDG +V QTKY K+LLK+F LED K + TPM ++ G V
Sbjct: 1084 EELSFFLGLQIKQLKDGTFVSQTKYIKDLLKRFGLEDAKPIKTPMATNGHLDLDEGGKPV 1143
Query: 418 DQKLYIGMIGSLLYLTASRPDILFSVCLCARFQSNPK 454
D KLY MIGSLLYLTASRPDI+FSVC+CARFQ+ PK
Sbjct: 1144 DLKLYRSMIGSLLYLTASRPDIMFSVCMCARFQAAPK 1180
>UniRef100_Q7XLK0 OSJNBa0009K15.10 protein [Oryza sativa]
Length = 845
Score = 178 bits (452), Expect = 4e-43
Identities = 95/157 (60%), Positives = 109/157 (68%)
Query: 298 LIKNDFKRGQVDTTLFRGTLKKYILVVQIYVDDIIFGSTNASLCKEFSK*MQDEFEKSMM 357
L+ DFK G+VDTTLF + V QIYVDDIIFGSTN CKEF + EFE SM+
Sbjct: 500 LLSKDFKIGKVDTTLFTKIIGDDFFVCQIYVDDIIFGSTNEVFCKEFGDMISREFEISMI 559
Query: 358 GELKFFLGIQINQRKDGVYVHQTKYTKELLKKFKLEDCKVMNTPMHPTCT*SKEDIGTKV 417
GEL FFL +QI Q KDG++V QTKY K LLK+F LED K + TPM + G V
Sbjct: 560 GELSFFLTLQIKQLKDGMFVSQTKYIKYLLKRFGLEDAKPIKTPMATNGHLDLDGGGKPV 619
Query: 418 DQKLYIGMIGSLLYLTASRPDILFSVCLCARFQSNPK 454
D KLY MIGSLLYLTASRPDI+FSVC+CARFQ+ PK
Sbjct: 620 DLKLYRSMIGSLLYLTASRPDIMFSVCMCARFQAAPK 656
>UniRef100_O24587 Pol protein [Zea mays]
Length = 1068
Score = 177 bits (448), Expect = 1e-42
Identities = 91/157 (57%), Positives = 108/157 (67%)
Query: 298 LIKNDFKRGQVDTTLFRGTLKKYILVVQIYVDDIIFGSTNASLCKEFSK*MQDEFEKSMM 357
LI N FK G+ D TLF T + V QIYVDDIIFGSTN C+EFS+ M +FE SMM
Sbjct: 723 LIANAFKVGKADPTLFTKTCDGDLFVCQIYVDDIIFGSTNQKSCEEFSRVMTQKFEMSMM 782
Query: 358 GELKFFLGIQINQRKDGVYVHQTKYTKELLKKFKLEDCKVMNTPMHPTCT*SKEDIGTKV 417
GEL +FLG Q+ Q KDG ++ QTKYT++LLK+F ++D K TPM G V
Sbjct: 783 GELNYFLGFQVKQLKDGTFISQTKYTQDLLKRFGMKDAKPAKTPMGTDGHTDLNKGGKSV 842
Query: 418 DQKLYIGMIGSLLYLTASRPDILFSVCLCARFQSNPK 454
DQK Y MIGSLLYL ASRPDI+ SVC+CARFQS+PK
Sbjct: 843 DQKAYRSMIGSLLYLCASRPDIMLSVCMCARFQSDPK 879
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.352 0.155 0.517
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 890,366,965
Number of Sequences: 2790947
Number of extensions: 32937373
Number of successful extensions: 117023
Number of sequences better than 10.0: 635
Number of HSP's better than 10.0 without gapping: 483
Number of HSP's successfully gapped in prelim test: 152
Number of HSP's that attempted gapping in prelim test: 115838
Number of HSP's gapped (non-prelim): 758
length of query: 642
length of database: 848,049,833
effective HSP length: 134
effective length of query: 508
effective length of database: 474,062,935
effective search space: 240823970980
effective search space used: 240823970980
T: 11
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (22.0 bits)
S2: 78 (34.7 bits)
Medicago: description of AC146818.6


