

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146794.13 + phase: 0 /pseudo
(818 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
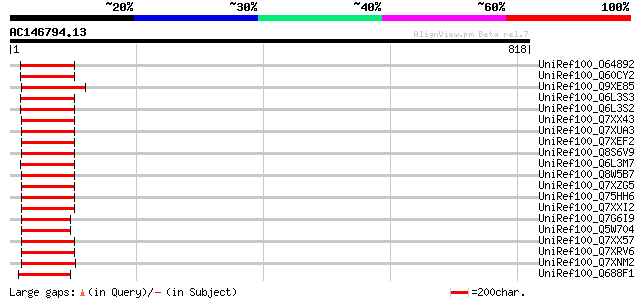
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_O64892 Polyprotein [Ananas comosus] 119 5e-25
UniRef100_Q60CY2 Putative polyprotein [Solanum tuberosum] 117 2e-24
UniRef100_Q9XE85 Polyprotein [Sorghum bicolor] 117 2e-24
UniRef100_Q6L3S3 Putative gag-pol polyprotein [Solanum demissum] 116 2e-24
UniRef100_Q6L3S2 Putative retrotransposon protein [Solanum demis... 116 2e-24
UniRef100_Q7XX43 OSJNBa0059D20.6 protein [Oryza sativa] 115 4e-24
UniRef100_Q7XUA3 OSJNBa0019D11.8 protein [Oryza sativa] 115 5e-24
UniRef100_Q7XEF2 Putative gag-pol protein [Oryza sativa] 115 5e-24
UniRef100_Q8S6V9 Putative retroelement [Oryza sativa] 114 9e-24
UniRef100_Q6L3M7 Putative gag-pol protein [Solanum demissum] 114 1e-23
UniRef100_Q8W5B7 Putative polyprotein [Oryza sativa] 114 1e-23
UniRef100_Q7XZG5 Putative gag-pol polyprotein [Oryza sativa] 114 2e-23
UniRef100_Q75HH6 Putative polyprotein [Oryza sativa] 114 2e-23
UniRef100_Q7XXI2 OSJNBa0059H15.7 protein [Oryza sativa] 113 2e-23
UniRef100_Q7G6I9 Putative polyprotein [Oryza sativa] 113 2e-23
UniRef100_Q5W704 Hypothetical protein OSJNBa0037H06.5 [Oryza sat... 113 3e-23
UniRef100_Q7XX57 OSJNBb0049I21.6 protein [Oryza sativa] 113 3e-23
UniRef100_Q7XRV6 OSJNBb0049I21.5 protein [Oryza sativa] 113 3e-23
UniRef100_Q7XNM2 OSJNBb0068N06.8 protein [Oryza sativa] 113 3e-23
UniRef100_Q688F1 Hypothetical protein OSJNBa0035I01.9 [Oryza sat... 113 3e-23
>UniRef100_O64892 Polyprotein [Ananas comosus]
Length = 871
Score = 119 bits (297), Expect = 5e-25
Identities = 55/84 (65%), Positives = 69/84 (81%)
Query: 18 AFRCYMNKIFHAYLDWFVVVFIDDILIYSRSEEEHAEHLKIMLQVLKEKKMYAKLSKCEF 77
AF MN++F YLD FVVVFIDDIL+YSRS+ +H EHL+I+LQVL+EK++Y KL KCEF
Sbjct: 149 AFMDLMNRVFKPYLDRFVVVFIDDILVYSRSDADHEEHLRIVLQVLREKELYVKLKKCEF 208
Query: 78 WLSEVSFLGHIISDSGIVVTHQKL 101
WL EV+FLGH+IS SGI V +K+
Sbjct: 209 WLREVAFLGHLISGSGIAVDPKKI 232
>UniRef100_Q60CY2 Putative polyprotein [Solanum tuberosum]
Length = 1487
Score = 117 bits (292), Expect = 2e-24
Identities = 53/84 (63%), Positives = 68/84 (80%)
Query: 18 AFRCYMNKIFHAYLDWFVVVFIDDILIYSRSEEEHAEHLKIMLQVLKEKKMYAKLSKCEF 77
AF MN++F YLD FV+VFIDDILIYSR++E+HA HL+I+LQ LK+K++YAK SKCEF
Sbjct: 713 AFMDLMNRVFKPYLDMFVIVFIDDILIYSRNKEDHASHLRIVLQTLKDKELYAKFSKCEF 772
Query: 78 WLSEVSFLGHIISDSGIVVTHQKL 101
WL V+FLGHI+S GI V +K+
Sbjct: 773 WLKSVAFLGHIVSGDGIKVDTRKI 796
>UniRef100_Q9XE85 Polyprotein [Sorghum bicolor]
Length = 1484
Score = 117 bits (292), Expect = 2e-24
Identities = 55/101 (54%), Positives = 74/101 (72%)
Query: 19 FRCYMNKIFHAYLDWFVVVFIDDILIYSRSEEEHAEHLKIMLQVLKEKKMYAKLSKCEFW 78
F MNK+F YLD FVVVFIDDIL++S+++EEHAEHL+++LQ L+E K+YAK SKCEFW
Sbjct: 693 FMYMMNKVFMEYLDKFVVVFIDDILVFSKTKEEHAEHLRLVLQKLREHKLYAKRSKCEFW 752
Query: 79 LSEVSFLGHIISDSGIVVTHQKLMQ*HNGRLRSQLQRLEVF 119
L EVSFLGH++S+ GI V K+ N + + + + F
Sbjct: 753 LEEVSFLGHVVSNGGIAVDPSKVKDVLNWKPPTDVSEIRSF 793
>UniRef100_Q6L3S3 Putative gag-pol polyprotein [Solanum demissum]
Length = 1515
Score = 116 bits (291), Expect = 2e-24
Identities = 52/84 (61%), Positives = 67/84 (78%)
Query: 18 AFRCYMNKIFHAYLDWFVVVFIDDILIYSRSEEEHAEHLKIMLQVLKEKKMYAKLSKCEF 77
AF MN++F YLD FV++FIDDILIYSR+EE+HA HL+ +LQ LK+K++YAK SKCEF
Sbjct: 790 AFMDLMNRVFRPYLDMFVIIFIDDILIYSRNEEDHASHLRTVLQTLKDKELYAKFSKCEF 849
Query: 78 WLSEVSFLGHIISDSGIVVTHQKL 101
WL V+FLGHI+S GI V +K+
Sbjct: 850 WLKSVAFLGHIVSGDGIKVDTRKI 873
>UniRef100_Q6L3S2 Putative retrotransposon protein [Solanum demissum]
Length = 1602
Score = 116 bits (291), Expect = 2e-24
Identities = 52/84 (61%), Positives = 67/84 (78%)
Query: 18 AFRCYMNKIFHAYLDWFVVVFIDDILIYSRSEEEHAEHLKIMLQVLKEKKMYAKLSKCEF 77
AF MN++F YLD FV++FIDDILIYSR+EE+HA HL+ +LQ LK+K++YAK SKCEF
Sbjct: 796 AFMDLMNRVFRPYLDMFVIIFIDDILIYSRNEEDHASHLRTVLQTLKDKELYAKFSKCEF 855
Query: 78 WLSEVSFLGHIISDSGIVVTHQKL 101
WL V+FLGHI+S GI V +K+
Sbjct: 856 WLKSVAFLGHIVSGDGIKVDTRKI 879
>UniRef100_Q7XX43 OSJNBa0059D20.6 protein [Oryza sativa]
Length = 1470
Score = 115 bits (289), Expect = 4e-24
Identities = 54/83 (65%), Positives = 66/83 (79%)
Query: 19 FRCYMNKIFHAYLDWFVVVFIDDILIYSRSEEEHAEHLKIMLQVLKEKKMYAKLSKCEFW 78
F MNK+F YLD FVVVFIDDILIYS+ EEEHAEHL+++LQ L++ K+YAK SKCEFW
Sbjct: 675 FMNLMNKVFMEYLDKFVVVFIDDILIYSKDEEEHAEHLRLVLQKLRKHKLYAKFSKCEFW 734
Query: 79 LSEVSFLGHIISDSGIVVTHQKL 101
L EV+FLGH+IS G+ V K+
Sbjct: 735 LKEVAFLGHVISAGGVAVDPAKV 757
>UniRef100_Q7XUA3 OSJNBa0019D11.8 protein [Oryza sativa]
Length = 1470
Score = 115 bits (288), Expect = 5e-24
Identities = 54/83 (65%), Positives = 66/83 (79%)
Query: 19 FRCYMNKIFHAYLDWFVVVFIDDILIYSRSEEEHAEHLKIMLQVLKEKKMYAKLSKCEFW 78
F MNK+F YLD FVVVFIDDILIYS+ EEEHAEHL+++LQ L++ K+YAK SKCEFW
Sbjct: 675 FMNLMNKVFMDYLDKFVVVFIDDILIYSKDEEEHAEHLRLVLQKLRKHKLYAKFSKCEFW 734
Query: 79 LSEVSFLGHIISDSGIVVTHQKL 101
L EV+FLGH+IS G+ V K+
Sbjct: 735 LKEVAFLGHVISAGGVAVDPAKV 757
>UniRef100_Q7XEF2 Putative gag-pol protein [Oryza sativa]
Length = 1470
Score = 115 bits (288), Expect = 5e-24
Identities = 54/83 (65%), Positives = 66/83 (79%)
Query: 19 FRCYMNKIFHAYLDWFVVVFIDDILIYSRSEEEHAEHLKIMLQVLKEKKMYAKLSKCEFW 78
F MNK+F YLD FVVVFIDDILIYS+ EEEHAEHL+++LQ L++ K+YAK SKCEFW
Sbjct: 675 FMNLMNKVFMDYLDKFVVVFIDDILIYSKDEEEHAEHLRLVLQKLRKHKLYAKFSKCEFW 734
Query: 79 LSEVSFLGHIISDSGIVVTHQKL 101
L EV+FLGH+IS G+ V K+
Sbjct: 735 LKEVAFLGHVISAGGVAVDPAKV 757
>UniRef100_Q8S6V9 Putative retroelement [Oryza sativa]
Length = 986
Score = 114 bits (286), Expect = 9e-24
Identities = 54/83 (65%), Positives = 67/83 (80%)
Query: 19 FRCYMNKIFHAYLDWFVVVFIDDILIYSRSEEEHAEHLKIMLQVLKEKKMYAKLSKCEFW 78
F MNK+F YLD FVVVFIDDILIYS+ EEEHA+HL+++L+ L++ K+YAK SKCEFW
Sbjct: 191 FMNLMNKVFMDYLDKFVVVFIDDILIYSKDEEEHAKHLRLVLEKLRKHKLYAKFSKCEFW 250
Query: 79 LSEVSFLGHIISDSGIVVTHQKL 101
L EV+FLGHIIS G+VV K+
Sbjct: 251 LKEVAFLGHIISAGGVVVDPAKV 273
>UniRef100_Q6L3M7 Putative gag-pol protein [Solanum demissum]
Length = 1438
Score = 114 bits (285), Expect = 1e-23
Identities = 52/84 (61%), Positives = 65/84 (76%)
Query: 18 AFRCYMNKIFHAYLDWFVVVFIDDILIYSRSEEEHAEHLKIMLQVLKEKKMYAKLSKCEF 77
AF MN++F YLD FV+VFIDDIL+YSR+EE+HA HL+ +LQ LK+ K+YAK SKCEF
Sbjct: 739 AFMDLMNRVFKPYLDTFVIVFIDDILVYSRNEEDHASHLRTVLQTLKDNKLYAKFSKCEF 798
Query: 78 WLSEVSFLGHIISDSGIVVTHQKL 101
WL V+FLGHI+S GI V K+
Sbjct: 799 WLKSVAFLGHIVSGDGIKVDTGKI 822
>UniRef100_Q8W5B7 Putative polyprotein [Oryza sativa]
Length = 962
Score = 114 bits (285), Expect = 1e-23
Identities = 52/83 (62%), Positives = 68/83 (81%)
Query: 19 FRCYMNKIFHAYLDWFVVVFIDDILIYSRSEEEHAEHLKIMLQVLKEKKMYAKLSKCEFW 78
F MNKIF YLD FVVVFIDDILIYS++EEEH +HL+++L+ L++ ++YAKLSKCEFW
Sbjct: 457 FMNLMNKIFMEYLDQFVVVFIDDILIYSKNEEEHKQHLRLVLEKLRQHQLYAKLSKCEFW 516
Query: 79 LSEVSFLGHIISDSGIVVTHQKL 101
L +V+FLGHII+ G+ V QK+
Sbjct: 517 LDQVAFLGHIITKEGVAVDPQKI 539
>UniRef100_Q7XZG5 Putative gag-pol polyprotein [Oryza sativa]
Length = 823
Score = 114 bits (284), Expect = 2e-23
Identities = 53/83 (63%), Positives = 66/83 (78%)
Query: 19 FRCYMNKIFHAYLDWFVVVFIDDILIYSRSEEEHAEHLKIMLQVLKEKKMYAKLSKCEFW 78
F MNK+F YLD FVVVFIDDILIYS+ EEEHAEHL+++L+ L++ K+YAK SKCEFW
Sbjct: 126 FMNLMNKVFVDYLDKFVVVFIDDILIYSKDEEEHAEHLRLVLEKLRKHKLYAKFSKCEFW 185
Query: 79 LSEVSFLGHIISDSGIVVTHQKL 101
L EV+FLGH+IS G+ V K+
Sbjct: 186 LKEVAFLGHVISAGGVAVDPAKV 208
>UniRef100_Q75HH6 Putative polyprotein [Oryza sativa]
Length = 1312
Score = 114 bits (284), Expect = 2e-23
Identities = 53/83 (63%), Positives = 66/83 (78%)
Query: 19 FRCYMNKIFHAYLDWFVVVFIDDILIYSRSEEEHAEHLKIMLQVLKEKKMYAKLSKCEFW 78
F MNK+F YLD FVVVFIDDILIYS+ EEEHAEHL+++L+ L++ K+YAK SKCEFW
Sbjct: 590 FMNLMNKVFVDYLDKFVVVFIDDILIYSKDEEEHAEHLRLVLEKLRKHKLYAKFSKCEFW 649
Query: 79 LSEVSFLGHIISDSGIVVTHQKL 101
L EV+FLGH+IS G+ V K+
Sbjct: 650 LKEVAFLGHVISAGGVAVDPAKV 672
>UniRef100_Q7XXI2 OSJNBa0059H15.7 protein [Oryza sativa]
Length = 920
Score = 113 bits (283), Expect = 2e-23
Identities = 53/83 (63%), Positives = 66/83 (78%)
Query: 19 FRCYMNKIFHAYLDWFVVVFIDDILIYSRSEEEHAEHLKIMLQVLKEKKMYAKLSKCEFW 78
F MNK+F YLD FVVVFIDDILIYS+ EEEHAEHL+++L+ L++ K+YAK SKCEFW
Sbjct: 256 FMNLMNKVFMDYLDKFVVVFIDDILIYSKDEEEHAEHLRLVLEKLQKHKLYAKFSKCEFW 315
Query: 79 LSEVSFLGHIISDSGIVVTHQKL 101
L EV+FLGH+IS G+ V K+
Sbjct: 316 LKEVAFLGHVISAGGVAVDPVKV 338
>UniRef100_Q7G6I9 Putative polyprotein [Oryza sativa]
Length = 1443
Score = 113 bits (283), Expect = 2e-23
Identities = 52/78 (66%), Positives = 64/78 (81%)
Query: 19 FRCYMNKIFHAYLDWFVVVFIDDILIYSRSEEEHAEHLKIMLQVLKEKKMYAKLSKCEFW 78
F MNK+F YLD FVVVFIDDILIYSR++EEH EHL++ L++L+E ++YAK SKCEFW
Sbjct: 639 FMNLMNKVFMEYLDKFVVVFIDDILIYSRTKEEHEEHLRLALEMLREHQLYAKFSKCEFW 698
Query: 79 LSEVSFLGHIISDSGIVV 96
LSEV FLGH+IS G+ V
Sbjct: 699 LSEVKFLGHVISAGGVAV 716
>UniRef100_Q5W704 Hypothetical protein OSJNBa0037H06.5 [Oryza sativa]
Length = 846
Score = 113 bits (282), Expect = 3e-23
Identities = 52/78 (66%), Positives = 63/78 (80%)
Query: 19 FRCYMNKIFHAYLDWFVVVFIDDILIYSRSEEEHAEHLKIMLQVLKEKKMYAKLSKCEFW 78
F MNK+F YLD FVVVFIDDILIYSR++EEH EHL++ L+ L+E ++YAK SKCEFW
Sbjct: 48 FMNLMNKVFMEYLDMFVVVFIDDILIYSRTKEEHEEHLRLALEKLREHQLYAKFSKCEFW 107
Query: 79 LSEVSFLGHIISDSGIVV 96
LSEV FLGH+IS G+ V
Sbjct: 108 LSEVKFLGHVISAGGVAV 125
>UniRef100_Q7XX57 OSJNBb0049I21.6 protein [Oryza sativa]
Length = 1606
Score = 113 bits (282), Expect = 3e-23
Identities = 52/83 (62%), Positives = 66/83 (78%)
Query: 19 FRCYMNKIFHAYLDWFVVVFIDDILIYSRSEEEHAEHLKIMLQVLKEKKMYAKLSKCEFW 78
F MNK+F YLD FVVVFIDDILIYS+ EEEHAEHL+++L+ L++ K+YAK SKCEFW
Sbjct: 608 FMNLMNKVFMDYLDKFVVVFIDDILIYSKDEEEHAEHLRLVLEKLRKHKLYAKFSKCEFW 667
Query: 79 LSEVSFLGHIISDSGIVVTHQKL 101
L E++FLGH+IS G+ V K+
Sbjct: 668 LKELAFLGHVISAGGVAVDPAKV 690
>UniRef100_Q7XRV6 OSJNBb0049I21.5 protein [Oryza sativa]
Length = 1644
Score = 113 bits (282), Expect = 3e-23
Identities = 52/83 (62%), Positives = 66/83 (78%)
Query: 19 FRCYMNKIFHAYLDWFVVVFIDDILIYSRSEEEHAEHLKIMLQVLKEKKMYAKLSKCEFW 78
F MNK+F YLD F+VVFIDDILIYS+ EEEHAEHL+++L+ L++ K+YAK SKCEFW
Sbjct: 765 FMNLMNKVFMDYLDKFMVVFIDDILIYSKDEEEHAEHLRLVLEKLRKHKLYAKFSKCEFW 824
Query: 79 LSEVSFLGHIISDSGIVVTHQKL 101
L EV+FLGH+IS G+ V K+
Sbjct: 825 LKEVAFLGHVISAGGVAVDPTKV 847
>UniRef100_Q7XNM2 OSJNBb0068N06.8 protein [Oryza sativa]
Length = 947
Score = 113 bits (282), Expect = 3e-23
Identities = 52/85 (61%), Positives = 68/85 (79%)
Query: 19 FRCYMNKIFHAYLDWFVVVFIDDILIYSRSEEEHAEHLKIMLQVLKEKKMYAKLSKCEFW 78
F MNK+F YLD FVVVFIDDIL+YS+SEE+H +HL+++L L+E ++YAKLSKCEFW
Sbjct: 537 FMNLMNKVFMEYLDKFVVVFIDDILVYSQSEEDHQQHLRLVLGKLREHQLYAKLSKCEFW 596
Query: 79 LSEVSFLGHIISDSGIVVTHQKLMQ 103
L EV FLGH+IS G+VV + L++
Sbjct: 597 LPEVKFLGHVISAKGVVVDPETLLK 621
>UniRef100_Q688F1 Hypothetical protein OSJNBa0035I01.9 [Oryza sativa]
Length = 666
Score = 113 bits (282), Expect = 3e-23
Identities = 54/82 (65%), Positives = 66/82 (79%)
Query: 15 FSDAFRCYMNKIFHAYLDWFVVVFIDDILIYSRSEEEHAEHLKIMLQVLKEKKMYAKLSK 74
F+ F MNK+F YLD FVVVFIDDILIYS+++EEH EHL++ L+ L+E ++YAK SK
Sbjct: 75 FTAFFMNLMNKVFLEYLDKFVVVFIDDILIYSQTKEEHEEHLRLALEKLREHQLYAKFSK 134
Query: 75 CEFWLSEVSFLGHIISDSGIVV 96
CEFWLSEV FLGHIIS G+VV
Sbjct: 135 CEFWLSEVKFLGHIISVGGVVV 156
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.372 0.167 0.652
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,005,862,332
Number of Sequences: 2790947
Number of extensions: 32206525
Number of successful extensions: 162471
Number of sequences better than 10.0: 1671
Number of HSP's better than 10.0 without gapping: 1599
Number of HSP's successfully gapped in prelim test: 72
Number of HSP's that attempted gapping in prelim test: 160398
Number of HSP's gapped (non-prelim): 2053
length of query: 818
length of database: 848,049,833
effective HSP length: 136
effective length of query: 682
effective length of database: 468,481,041
effective search space: 319504069962
effective search space used: 319504069962
T: 11
A: 40
X1: 14 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.9 bits)
S2: 79 (35.0 bits)
Medicago: description of AC146794.13


