

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146792.8 - phase: 0
(266 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
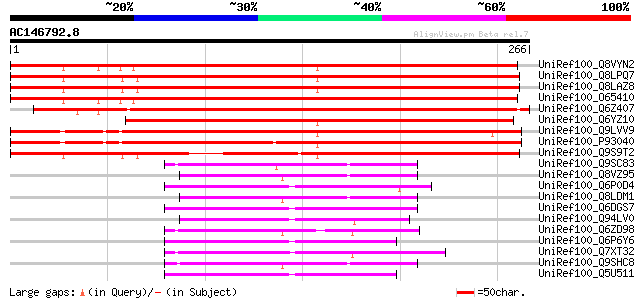
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q8VYN2 Putative membrane associated protein [Arabidops... 363 3e-99
UniRef100_Q8LPQ7 AT4g05060/T32N4_12 [Arabidopsis thaliana] 340 2e-92
UniRef100_Q8LAZ8 Membrane associated protein-like protein [Arabi... 338 8e-92
UniRef100_O65410 Putative membrane associated protein [Arabidops... 335 5e-91
UniRef100_Q6Z407 Putative 27k vesicle-associated membrane protei... 325 7e-88
UniRef100_Q6YZ10 27k vesicle-associated membrane protein-associa... 316 3e-85
UniRef100_Q9LVV9 Membrane associated protein [Arabidopsis thaliana] 306 5e-82
UniRef100_P93040 Membrane associated protein [Arabidopsis thaliana] 292 7e-78
UniRef100_Q9S9T2 T32N4.12 protein [Arabidopsis thaliana] 228 1e-58
UniRef100_Q9SC83 VAP27 [Nicotiana plumbaginifolia] 62 1e-08
UniRef100_Q8VZ95 Hypothetical protein At3g60600; T4C21_10 [Arabi... 60 8e-08
UniRef100_Q6P0D4 Hypothetical protein vapa [Brachydanio rerio] 59 2e-07
UniRef100_Q8LDM1 Putative VAMP-associated protein [Arabidopsis t... 59 2e-07
UniRef100_Q6DGS7 VAMP (Vesicle-associated membrane protein)-asso... 58 3e-07
UniRef100_Q94LV0 Putative membrane protein [Oryza sativa] 57 4e-07
UniRef100_Q6ZD98 Putative vesicle-associated membrane protein-as... 57 4e-07
UniRef100_Q6P6Y6 Vapa protein [Brachydanio rerio] 57 5e-07
UniRef100_Q7XT32 OSJNBa0010H02.20 protein [Oryza sativa] 57 6e-07
UniRef100_Q9SHC8 Putative VAMP-associated protein (Putative VAMP... 56 1e-06
UniRef100_Q5U511 Hypothetical protein [Xenopus laevis] 55 1e-06
>UniRef100_Q8VYN2 Putative membrane associated protein [Arabidopsis thaliana]
Length = 295
Score = 363 bits (931), Expect = 3e-99
Identities = 196/290 (67%), Positives = 223/290 (76%), Gaps = 30/290 (10%)
Query: 1 MAVAEPKLHSEPKV-WNFFKLPFRNSN-----------------TSSTNTTSSSVNNNLH 42
M + E K S+ K W FFK+PFRNS+ SS++T+S N++ H
Sbjct: 1 MTMTEEKPTSDGKGGWGFFKIPFRNSSGHRNAASSAATSPFPSGASSSSTSSHLHNHHQH 60
Query: 43 HH-----HHHNPNSNTPL---EGSTSHT--SNSVSSVARSLLPTRRRLKLDPSNKLYFPY 92
HH HHH N P G H S SVSSVA+S LPT+RRLKLDPS KLYFPY
Sbjct: 61 HHQHHHQHHHQLGYNGPHGDGSGQNQHPTPSPSVSSVAKSFLPTKRRLKLDPSEKLYFPY 120
Query: 93 EPGKQVRSAIRIKNTSKSNVAFKFQTTAPKSCFMRPPGAILAPGESIIATVFKFVEQPEN 152
EPGKQVRSAI+IKNTSKS+VAFKFQTTAPKSCFMRPPGAILAPGE+IIATVFKFVE PEN
Sbjct: 121 EPGKQVRSAIKIKNTSKSHVAFKFQTTAPKSCFMRPPGAILAPGETIIATVFKFVEPPEN 180
Query: 153 NEKP--EKSGLKFKIMSLKVKGSIDYVPELFDEQKDQVAVEQILRVVFLDPERPSPILEK 210
NEKP ++S +KFKIMSLKVKG +DYVPELFDEQKD V+ EQILRV+FLDPER +P LEK
Sbjct: 181 NEKPMDQRSRVKFKIMSLKVKGPMDYVPELFDEQKDDVSKEQILRVIFLDPERSNPALEK 240
Query: 211 LKRQLADADAALEERKKPAEDAGPKIIGEGLVIDEWKERRERYLAKQQGD 260
LKRQLA+ADAA+E RKKP E+ GPK+IGEGLVIDEWKERRERYLA+QQG+
Sbjct: 241 LKRQLAEADAAVEARKKPPEETGPKMIGEGLVIDEWKERRERYLAQQQGE 290
>UniRef100_Q8LPQ7 AT4g05060/T32N4_12 [Arabidopsis thaliana]
Length = 287
Score = 340 bits (873), Expect = 2e-92
Identities = 187/284 (65%), Positives = 213/284 (74%), Gaps = 23/284 (8%)
Query: 1 MAVAEPKLHSEPKVWNFFKLPFRNSN-------------TSSTNTTSSSVNNNLHHHHHH 47
MA+ E K S+ + W FKLPFRNSN TSS++ TSS +N N H H
Sbjct: 1 MALTEDKSDSDGRRWGKFKLPFRNSNSQAPSASSSSSMATSSSSVTSSHLNQNYIHQSRH 60
Query: 48 NPNSNTPLE---GSTSHTSN----SVSSVARSLLPTRRRLKLDPSNKLYFPYEPGKQVRS 100
P+ G H S S+SSVARSLLPT+RRLKLDPS KLYFPYEPGKQVRS
Sbjct: 61 FQYHGPPVVEGLGQNHHQSAATIPSMSSVARSLLPTKRRLKLDPSAKLYFPYEPGKQVRS 120
Query: 101 AIRIKNTSKSNVAFKFQTTAPKSCFMRPPGAILAPGESIIATVFKFVEQPENNEKP--EK 158
AI+IKNTSKS+VAFKFQTT PKSCFMRP GAILAPGE IIATVFKFVE PENNEKP +K
Sbjct: 121 AIKIKNTSKSHVAFKFQTTVPKSCFMRPAGAILAPGEEIIATVFKFVEPPENNEKPMEQK 180
Query: 159 SGLKFKIMSLKVKGSIDYVPELFDEQKDQVAVEQILRVVFLDPERPSPILEKLKRQLADA 218
SG+KFKIMSLK+K DY+PELF+EQKD V+ EQ++RVVFLDPE P+ ++EKLK QLA+A
Sbjct: 181 SGVKFKIMSLKMKVPTDYMPELFEEQKDHVSEEQVMRVVFLDPENPNSMMEKLKSQLAEA 240
Query: 219 DAALEERKKPAED-AGPKIIGEGLVIDEWKERRERYLAKQQGDV 261
DAA E RKK +E GPK IGEGLVIDEWK+RRERYLA+QQG V
Sbjct: 241 DAADEARKKASEGIVGPKPIGEGLVIDEWKQRRERYLAQQQGGV 284
>UniRef100_Q8LAZ8 Membrane associated protein-like protein [Arabidopsis thaliana]
Length = 287
Score = 338 bits (867), Expect = 8e-92
Identities = 186/284 (65%), Positives = 212/284 (74%), Gaps = 23/284 (8%)
Query: 1 MAVAEPKLHSEPKVWNFFKLPFRNSN-------------TSSTNTTSSSVNNNLHHHHHH 47
MA+ E K S+ + W FKLPFRNSN TSS++ TSS +N N H H
Sbjct: 1 MALTEDKSDSDGRRWGKFKLPFRNSNSQAPSASSSSSMATSSSSVTSSHLNQNYIHQSRH 60
Query: 48 NPNSNTPLE---GSTSHTSN----SVSSVARSLLPTRRRLKLDPSNKLYFPYEPGKQVRS 100
P+ G H S S+SSVARSLLPT+RRLKLDPS KLYFPYEPGKQVRS
Sbjct: 61 FQYHGPPVVEGLGQNHHQSAATIPSMSSVARSLLPTKRRLKLDPSAKLYFPYEPGKQVRS 120
Query: 101 AIRIKNTSKSNVAFKFQTTAPKSCFMRPPGAILAPGESIIATVFKFVEQPENNEKP--EK 158
AI+IKNTSKS+VAFKFQTT PKSCFMRP GAILAPGE IIATVFKFVE PENNEKP +K
Sbjct: 121 AIKIKNTSKSHVAFKFQTTVPKSCFMRPAGAILAPGEEIIATVFKFVEPPENNEKPMEQK 180
Query: 159 SGLKFKIMSLKVKGSIDYVPELFDEQKDQVAVEQILRVVFLDPERPSPILEKLKRQLADA 218
SG+KFKIMSLK+K DY+PELF+EQKD V+ EQ++RVVFLDPE P+ ++EKL QLA+A
Sbjct: 181 SGVKFKIMSLKMKVPTDYMPELFEEQKDHVSEEQVMRVVFLDPENPNSMMEKLXSQLAEA 240
Query: 219 DAALEERKKPAED-AGPKIIGEGLVIDEWKERRERYLAKQQGDV 261
DAA E RKK +E GPK IGEGLVIDEWK+RRERYLA+QQG V
Sbjct: 241 DAADEARKKASEGIVGPKPIGEGLVIDEWKQRRERYLAQQQGGV 284
>UniRef100_O65410 Putative membrane associated protein [Arabidopsis thaliana]
Length = 293
Score = 335 bits (860), Expect = 5e-91
Identities = 184/288 (63%), Positives = 212/288 (72%), Gaps = 28/288 (9%)
Query: 1 MAVAEPKLHSEPKV-WNFFKLPFRNSN-----------------TSSTNTTSSSVNNNLH 42
M + E K S+ K W FFK+PFRNS+ SS++T+S N++ H
Sbjct: 1 MTMTEEKPTSDGKGGWGFFKIPFRNSSGHRNAASSAATSPFPSGASSSSTSSHLHNHHQH 60
Query: 43 HH-----HHHNPNSNTPL---EGSTSHT--SNSVSSVARSLLPTRRRLKLDPSNKLYFPY 92
HH HHH N P G H S SVSSVA+S LPT+RRLKLDPS KLYFPY
Sbjct: 61 HHQHHHQHHHQLGYNGPHGDGSGQNQHPTPSPSVSSVAKSFLPTKRRLKLDPSEKLYFPY 120
Query: 93 EPGKQVRSAIRIKNTSKSNVAFKFQTTAPKSCFMRPPGAILAPGESIIATVFKFVEQPEN 152
EPGKQVRSAI+IKNTSKS+VAFKFQTTAPKSCFMRPPGAILAPGE+IIAT F +
Sbjct: 121 EPGKQVRSAIKIKNTSKSHVAFKFQTTAPKSCFMRPPGAILAPGETIIATGICFFCFLAS 180
Query: 153 NEKPEKSGLKFKIMSLKVKGSIDYVPELFDEQKDQVAVEQILRVVFLDPERPSPILEKLK 212
++S +KFKIMSLKVKG +DYVPELFDEQKD V+ EQILRV+FLDPER +P LEKLK
Sbjct: 181 KPMDQRSRVKFKIMSLKVKGPMDYVPELFDEQKDDVSKEQILRVIFLDPERSNPALEKLK 240
Query: 213 RQLADADAALEERKKPAEDAGPKIIGEGLVIDEWKERRERYLAKQQGD 260
RQLA+ADAA+E RKKP E+ GPK+IGEGLVIDEWKERRERYLA+QQG+
Sbjct: 241 RQLAEADAAVEARKKPPEETGPKMIGEGLVIDEWKERRERYLAQQQGE 288
>UniRef100_Q6Z407 Putative 27k vesicle-associated membrane protein-associated protein
[Oryza sativa]
Length = 284
Score = 325 bits (833), Expect = 7e-88
Identities = 172/265 (64%), Positives = 204/265 (76%), Gaps = 13/265 (4%)
Query: 13 KVWNFFKLPFRNSNTSSTNTT------SSSVNNNLHHH----HHHNPNSNTPLEGSTSHT 62
K+WN ++PF + + SSS + +HHH + H + G +
Sbjct: 22 KLWNLCRMPFWQPGGAPATASAPPPPSSSSSSAGIHHHSAGRYGHEGGGGGAVAGDGA-P 80
Query: 63 SNSVSSVARSLLPTRRRLKLDPSNKLYFPYEPGKQVRSAIRIKNTSKSNVAFKFQTTAPK 122
+ S+SSVA+SLLP RRRL+LDP NKLYFPY+PG+QVRSAIRIKNTSKS+VAFKFQTTAPK
Sbjct: 81 AGSISSVAKSLLPARRRLRLDPPNKLYFPYQPGQQVRSAIRIKNTSKSHVAFKFQTTAPK 140
Query: 123 SCFMRPPGAILAPGESIIATVFKFVEQPENNEKP-EKSGLKFKIMSLKVKGSIDYVPELF 181
SCFMRPPGAILAPGE+IIATVFKFVE PENNE +K +KFKI+SLKVKG +DY PE+F
Sbjct: 141 SCFMRPPGAILAPGETIIATVFKFVEHPENNENVLQKCKVKFKILSLKVKGPMDYAPEMF 200
Query: 182 DEQKDQVAVEQILRVVFLDPERPSPILEKLKRQLADADAALEERKKPAEDAGPKIIGEGL 241
DEQ+DQ VE+ILRVVFL+ E P P LEKL QLA+A+AALE RKKP E+ GPKI+GEGL
Sbjct: 201 DEQRDQAVVEKILRVVFLNVENPGPQLEKLNNQLAEAEAALEARKKPPEENGPKIVGEGL 260
Query: 242 VIDEWKERRERYLAKQQGDVVVDSV 266
VIDEWKERRERYLA+QQ + VVDSV
Sbjct: 261 VIDEWKERRERYLAQQQVE-VVDSV 284
>UniRef100_Q6YZ10 27k vesicle-associated membrane protein-associated protein-like
[Oryza sativa]
Length = 233
Score = 316 bits (810), Expect = 3e-85
Identities = 157/201 (78%), Positives = 182/201 (90%), Gaps = 2/201 (0%)
Query: 60 SHTSNSVSSVARSLLPTRRRLKLDPSNKLYFPYEPGKQVRSAIRIKNTSKSNVAFKFQTT 119
+ +S+ ++V RSLLPTRRRL+LDP +KL+FPYEPGKQVRSA++IKN SKS+VAFKFQTT
Sbjct: 26 ARSSSGGAAVIRSLLPTRRRLRLDPPSKLFFPYEPGKQVRSAVKIKNISKSHVAFKFQTT 85
Query: 120 APKSCFMRPPGAILAPGESIIATVFKFVEQPENNEKP--EKSGLKFKIMSLKVKGSIDYV 177
APKSCFMRPPG ILAPGESIIATVFKFVE PENNEKP +K +KFKI+SLKVKG ++YV
Sbjct: 86 APKSCFMRPPGGILAPGESIIATVFKFVEHPENNEKPLDQKCKVKFKIVSLKVKGPMEYV 145
Query: 178 PELFDEQKDQVAVEQILRVVFLDPERPSPILEKLKRQLADADAALEERKKPAEDAGPKII 237
PELFDEQK+QVAVEQILRVVFLD ER +P ++KLKRQLA+A+AALE RKKP ED GP+I+
Sbjct: 146 PELFDEQKNQVAVEQILRVVFLDAERQTPQMDKLKRQLAEAEAALEARKKPPEDTGPRIV 205
Query: 238 GEGLVIDEWKERRERYLAKQQ 258
GEGLVIDEWKERRERYLA+QQ
Sbjct: 206 GEGLVIDEWKERRERYLARQQ 226
>UniRef100_Q9LVV9 Membrane associated protein [Arabidopsis thaliana]
Length = 295
Score = 306 bits (783), Expect = 5e-82
Identities = 169/295 (57%), Positives = 208/295 (70%), Gaps = 37/295 (12%)
Query: 1 MAVAEPKLHSEPKVWNFFKL-PFRNSNTSSTNTTSSSVNNNLHHHHHHNPNSNTPLEGST 59
M + + + S K N F+L PF ST ++SS+ N N ++ H N NT + +
Sbjct: 1 MPIGDRQNPSVEKKKNLFRLCPFWQRR--STTSSSSTQNPNQNYRSRHG-NRNTDIS-AV 56
Query: 60 SHTSNSVSSVARSLLPTRRRLKLDPSNKLYFPYEPGKQVRSAIRIKNTSKSNVAFKFQTT 119
S ++SSVARSLLP RRRL+LDPS+ LYFPYEPGKQVRSAI++KNTSKS+ AFKFQTT
Sbjct: 57 SKPPLTMSSVARSLLPARRRLRLDPSSYLYFPYEPGKQVRSAIKLKNTSKSHTAFKFQTT 116
Query: 120 APKSCFMRPPGAILAPGESIIATVFKFVEQPENNEKP---EKSGLKFKIMSLKVKGSIDY 176
APKSC+MRPPG +LAPGES+ ATVFKFVE PENNEK +KS +KFKIMSLKVK ++Y
Sbjct: 117 APKSCYMRPPGGVLAPGESVFATVFKFVEHPENNEKQPLNQKSKVKFKIMSLKVKPGVEY 176
Query: 177 VPELFDEQKDQVAVEQILRVVFLDPERPSPILEKLKRQLADADAALEERKKPAEDAGPKI 236
VPELFDEQKDQVAVEQ+LRV+F+D +RPS LEKLKRQL +A+AA+E RKKP + GP++
Sbjct: 177 VPELFDEQKDQVAVEQVLRVIFIDADRPSAALEKLKRQLDEAEAAVEARKKPPPETGPRV 236
Query: 237 IGEGLVIDEW-----------------------------KERRERYLAKQQGDVV 262
+GEGLVIDEW KERRE+YLA+QQ + V
Sbjct: 237 VGEGLVIDEWVTYNYKIFGPFIVVPSFSHIFTKVCVIYQKERREKYLARQQVESV 291
>UniRef100_P93040 Membrane associated protein [Arabidopsis thaliana]
Length = 265
Score = 292 bits (747), Expect = 7e-78
Identities = 158/266 (59%), Positives = 198/266 (74%), Gaps = 9/266 (3%)
Query: 1 MAVAEPKLHSEPKVWNFFKL-PFRNSNTSSTNTTSSSVNNNLHHHHHHNPNSNTPLEGST 59
M + + + S K N F+L PF ST ++SS+ N N ++ H N NT + +
Sbjct: 1 MPIGDRQNPSVEKKKNLFRLCPFWQRR--STTSSSSTQNPNQNYRSRHG-NRNTDIS-AV 56
Query: 60 SHTSNSVSSVARSLLPTRRRLKLDPSNKLYFPYEPGKQVRSAIRIKNTSKSNVAFKFQTT 119
S ++SSVARSLLP RRRL+LDPS+ LYFPYEPGKQVRSAI++KNTSKS+ AFKFQTT
Sbjct: 57 SKPPLTMSSVARSLLPARRRLRLDPSSYLYFPYEPGKQVRSAIKLKNTSKSHTAFKFQTT 116
Query: 120 APKSCFMRPPGAILAPGESIIATVFKFVEQPENNEKP---EKSGLKFKIMSLKVKGSIDY 176
APKSC+MRPPG + E + + VE PENNEK +KS +KFKIMSLKVK ++Y
Sbjct: 117 APKSCYMRPPGGVWLQ-ERVFCYCVQVVEHPENNEKQPLNQKSKVKFKIMSLKVKPGVEY 175
Query: 177 VPELFDEQKDQVAVEQILRVVFLDPERPSPILEKLKRQLADADAALEERKKPAEDAGPKI 236
VPELFDEQKDQVAVEQ+LRV+F+D +RPS LEKLKRQL +A+AA+E RKKP + GP++
Sbjct: 176 VPELFDEQKDQVAVEQVLRVIFIDADRPSAALEKLKRQLDEAEAAVEARKKPPPETGPRV 235
Query: 237 IGEGLVIDEWKERRERYLAKQQGDVV 262
+GEGLVIDEWKERRE+YLA+QQ + V
Sbjct: 236 VGEGLVIDEWKERREKYLARQQVESV 261
>UniRef100_Q9S9T2 T32N4.12 protein [Arabidopsis thaliana]
Length = 269
Score = 228 bits (581), Expect = 1e-58
Identities = 144/284 (50%), Positives = 173/284 (60%), Gaps = 41/284 (14%)
Query: 1 MAVAEPKLHSEPKVWNFFKLPFRNSN-------------TSSTNTTSSSVNNNLHHHHHH 47
MA+ E K S+ + W FKLPFRNSN TSS++ TSS +N N H H
Sbjct: 1 MALTEDKSDSDGRRWGKFKLPFRNSNSQAPSASSSSSMATSSSSVTSSHLNQNYIHQSRH 60
Query: 48 NPNSNTPLE---GSTSHTSN----SVSSVARSLLPTRRRLKLDPSNKLYFPYEPGKQVRS 100
P+ G H S S+SSVARSLLPT+RRLKLDPS KLYFP+
Sbjct: 61 FQYHGPPVVEGLGQNHHQSAATIPSMSSVARSLLPTKRRLKLDPSAKLYFPF-------- 112
Query: 101 AIRIKNTSKSNVAFKFQTTAPKSCFMRPPGAILAPGESIIATVFKFVEQPENNEKP--EK 158
SN + K AP C+ G I + +V PENNEKP +K
Sbjct: 113 ---------SNNSTKELLHAPGWCYSCSRGRNYRNWVLFIIGLDDYVP-PENNEKPMEQK 162
Query: 159 SGLKFKIMSLKVKGSIDYVPELFDEQKDQVAVEQILRVVFLDPERPSPILEKLKRQLADA 218
SG+KFKIMSLK+K DY+PELF+EQKD V+ EQ++RVVFLDPE P+ ++EKLK QLA+A
Sbjct: 163 SGVKFKIMSLKMKVPTDYMPELFEEQKDHVSEEQVMRVVFLDPENPNSMMEKLKSQLAEA 222
Query: 219 DAALEERKKPAED-AGPKIIGEGLVIDEWKERRERYLAKQQGDV 261
DAA E RKK +E GPK IGEGLVIDEWK+RRERYLA+QQG V
Sbjct: 223 DAADEARKKASEGIVGPKPIGEGLVIDEWKQRRERYLAQQQGGV 266
>UniRef100_Q9SC83 VAP27 [Nicotiana plumbaginifolia]
Length = 240
Score = 62.4 bits (150), Expect = 1e-08
Identities = 41/133 (30%), Positives = 74/133 (54%), Gaps = 5/133 (3%)
Query: 80 LKLDPSNKLYFPYEPGKQVRSAIRIKNTSKSNVAFKFQTTAPKSCFMRPPGAILAP--GE 137
L+++P +L FP+E KQ+ +I++ N S + VAFK +TT PK +RP ++ P
Sbjct: 9 LQIEPL-ELQFPFELKKQISCSIQLTNKSDNYVAFKVKTTNPKKYCVRPNTGVVMPHSAS 67
Query: 138 SIIATVFKFVEQPENNEKPEKSGLKFKIMSLKVKGSIDYVPELFDEQKDQVAVEQILRVV 197
+I T+ E P + + +K L+ + S D PE+F++++ + LRV+
Sbjct: 68 DVIVTMQAQKEAPADMQCKDKFLLQSVVASPGATAK-DITPEMFNKEEGNHVEDCKLRVI 126
Query: 198 FLDPER-PSPILE 209
++ P++ PSP+ E
Sbjct: 127 YVPPQQPPSPVQE 139
>UniRef100_Q8VZ95 Hypothetical protein At3g60600; T4C21_10 [Arabidopsis thaliana]
Length = 256
Score = 59.7 bits (143), Expect = 8e-08
Identities = 40/125 (32%), Positives = 66/125 (52%), Gaps = 4/125 (3%)
Query: 88 LYFPYEPGKQVRSAIRIKNTSKSNVAFKFQTTAPKSCFMRPPGAILAPGES--IIATVFK 145
L FP+E KQ+ ++ + N + +NVAFK +TT PK +RP ++ P + ++ T+
Sbjct: 30 LQFPFELKKQISCSLYLTNKTDNNVAFKVKTTNPKKYCVRPNTGVVLPRSTCEVLVTMQA 89
Query: 146 FVEQPENNEKPEKSGLKFKIMSLKVKGSIDYVPELFDEQKDQVAVEQILRVVFLDPER-P 204
E P + + +K L+ I S V + PE+F ++ E LRV ++ P R P
Sbjct: 90 QKEAPSDMQCKDKFLLQGVIASPGVTAK-EVTPEMFSKEAGHRVEETKLRVTYVAPPRPP 148
Query: 205 SPILE 209
SP+ E
Sbjct: 149 SPVHE 153
>UniRef100_Q6P0D4 Hypothetical protein vapa [Brachydanio rerio]
Length = 246
Score = 58.5 bits (140), Expect = 2e-07
Identities = 41/148 (27%), Positives = 72/148 (47%), Gaps = 14/148 (9%)
Query: 80 LKLDPSNKLYFPYEPGKQVRSAIRIKNTSKSNVAFKFQTTAPKSCFMRPPGAILAPGESI 139
L L+PSN L F V + ++++N S V FK +TTAP+ +RP I+ PG ++
Sbjct: 8 LILEPSNDLRFRGPFTDVVTTNLKLRNPSDRRVCFKVKTTAPRRYCVRPNSGIIDPGATV 67
Query: 140 IATVFKFVEQPENNEKPEKSGLKFKIMSLKVKGSIDYVPELFDEQKDQVAVEQILRVVF- 198
+V + QP + + EKS KF + ++ S+ + ++ + K ++ LR VF
Sbjct: 68 TISV---MLQPFDYDPNEKSKHKFMVQTIFAPSSVSDLEAMWKDAKPDDLMDSKLRCVFD 124
Query: 199 ----------LDPERPSPILEKLKRQLA 216
L+ +P+ +L K + A
Sbjct: 125 MPSDNDKMSELESSKPACVLNSSKAEAA 152
>UniRef100_Q8LDM1 Putative VAMP-associated protein [Arabidopsis thaliana]
Length = 240
Score = 58.5 bits (140), Expect = 2e-07
Identities = 39/125 (31%), Positives = 66/125 (52%), Gaps = 4/125 (3%)
Query: 88 LYFPYEPGKQVRSAIRIKNTSKSNVAFKFQTTAPKSCFMRPPGAILAPGES--IIATVFK 145
L FP+E KQ+ ++ + N + +NVAFK +TT PK +RP ++ P + ++ T+
Sbjct: 14 LQFPFELKKQISCSLYLTNKTDNNVAFKVKTTNPKKYCVRPNTGVVLPRSTCEVLVTMQA 73
Query: 146 FVEQPENNEKPEKSGLKFKIMSLKVKGSIDYVPELFDEQKDQVAVEQILRVVFL-DPERP 204
E P + + +K L+ I S V + PE+F ++ E LRV ++ P+ P
Sbjct: 74 QKEAPSDMQCKDKFLLQGVIASPGVTAK-EVTPEMFSKEAGHRVEETKLRVTYVAPPQPP 132
Query: 205 SPILE 209
SP+ E
Sbjct: 133 SPVHE 137
>UniRef100_Q6DGS7 VAMP (Vesicle-associated membrane protein)-associated protein A,
like [Brachydanio rerio]
Length = 246
Score = 57.8 bits (138), Expect = 3e-07
Identities = 38/130 (29%), Positives = 64/130 (49%), Gaps = 3/130 (2%)
Query: 80 LKLDPSNKLYFPYEPGKQVRSAIRIKNTSKSNVAFKFQTTAPKSCFMRPPGAILAPGESI 139
L LDP N L F V + +++KN S+ V FK +TTAP+ +RP I+ PG ++
Sbjct: 8 LVLDPPNDLKFKGPFTDVVTANLKLKNPSERKVCFKVKTTAPRRYCVRPNSGIIDPGATL 67
Query: 140 IATVFKFVEQPENNEKPEKSGLKFKIMSLKVKGSIDYVPELFDEQKDQVAVEQILRVVFL 199
+V + QP + + EKS KF + ++ ++ ++ + K ++ LR VF
Sbjct: 68 TISV---MLQPFDYDPNEKSKHKFMVQTIFAPPAVSDTEAMWKDAKPDELMDSKLRCVFE 124
Query: 200 DPERPSPILE 209
P + E
Sbjct: 125 LPSENDKVNE 134
>UniRef100_Q94LV0 Putative membrane protein [Oryza sativa]
Length = 233
Score = 57.4 bits (137), Expect = 4e-07
Identities = 40/122 (32%), Positives = 63/122 (50%), Gaps = 7/122 (5%)
Query: 88 LYFPYEPGKQVRSAIRIKNTSKSNVAFKFQTTAPKSCFMRPPGAILAPGESIIATVFKFV 147
L F +E K +++ N S+ +VAFK +TT+P+ F+RP +I+ P +S T+
Sbjct: 15 LTFLFELDKPCYCNLKVVNNSEHHVAFKVKTTSPRKYFVRPNASIIQPWDSCTITI---T 71
Query: 148 EQPENNEKPE-KSGLKFKIMSLKVKGSID---YVPELFDEQKDQVAVEQILRVVFLDPER 203
Q + P+ + KF I S KV S D P F+++ D+V E L+VV+ P
Sbjct: 72 LQAQKEYPPDMQCKDKFLIQSTKVAASTDMDEIPPNTFNKEVDKVIEEMKLKVVYTVPSG 131
Query: 204 PS 205
S
Sbjct: 132 SS 133
>UniRef100_Q6ZD98 Putative vesicle-associated membrane protein-associated protein
[Oryza sativa]
Length = 243
Score = 57.4 bits (137), Expect = 4e-07
Identities = 39/137 (28%), Positives = 72/137 (52%), Gaps = 11/137 (8%)
Query: 80 LKLDPSNKLYFPYEPGKQVRSAIRIKNTSKSNVAFKFQTTAPKSCFMRPPGAILAPGES- 138
L++DP +L FP+E KQ+ ++++ N S +AFK +TT+PK +RP ++ P +
Sbjct: 13 LRIDPV-ELRFPFELKKQISCSMQLSNLSDDYIAFKVKTTSPKKYSVRPNTGVVLPRSTC 71
Query: 139 -IIATVFKFVEQPENNEKPEKSGLKFKIMSLKVKGSI---DYVPELFDEQKDQVAVEQIL 194
++ T+ E P + + + KF + S+ + D E+F ++ E L
Sbjct: 72 DVVVTMQAQREAPPDMQCKD----KFLVQSVIAPSGVTVKDITGEMFTKESGNKVEEVKL 127
Query: 195 RVVFL-DPERPSPILEK 210
RV ++ P+ PSP+ E+
Sbjct: 128 RVTYIAPPQPPSPVPEE 144
>UniRef100_Q6P6Y6 Vapa protein [Brachydanio rerio]
Length = 156
Score = 57.0 bits (136), Expect = 5e-07
Identities = 36/119 (30%), Positives = 62/119 (51%), Gaps = 3/119 (2%)
Query: 80 LKLDPSNKLYFPYEPGKQVRSAIRIKNTSKSNVAFKFQTTAPKSCFMRPPGAILAPGESI 139
L L+PSN L F V + ++++N S V FK +TTAP+ +RP I+ PG ++
Sbjct: 8 LILEPSNDLRFRGPFTDVVTTNLKLRNPSDRRVCFKVKTTAPRRYCVRPNSGIIDPGATV 67
Query: 140 IATVFKFVEQPENNEKPEKSGLKFKIMSLKVKGSIDYVPELFDEQKDQVAVEQILRVVF 198
+V + QP + + EKS KF + ++ S+ + ++ + K ++ LR VF
Sbjct: 68 TISV---MLQPFDYDPNEKSKHKFMVQTIFAPSSVSDLEAMWKDAKPDDLMDSKLRCVF 123
>UniRef100_Q7XT32 OSJNBa0010H02.20 protein [Oryza sativa]
Length = 244
Score = 56.6 bits (135), Expect = 6e-07
Identities = 46/147 (31%), Positives = 71/147 (48%), Gaps = 6/147 (4%)
Query: 80 LKLDPSNKLYFPYEPGKQVRSAIRIKNTSKSNVAFKFQTTAPKSCFMRPPGAILAPGESI 139
L++ PS +L PYE ++ +++ N + VAFK +TT P+ +R IL P S
Sbjct: 6 LRVHPS-ELKIPYEYKRKRSCCMQLTNKTNQYVAFKVKTTNPRKYSVRHACGILPPRSSC 64
Query: 140 IATVFKFVEQPENNEKPEKSGLKFKIMSLKVKGSI---DYVPELFDEQKDQVAVEQILRV 196
TV ++ P KF + S+ V D+VPELF + +V E LRV
Sbjct: 65 DITVT--MQAPVEMLSDYHCKDKFLVQSVAVGYGATMRDFVPELFTKAPGRVIEEFKLRV 122
Query: 197 VFLDPERPSPILEKLKRQLADADAALE 223
V++ PSP+ E+ + + DA E
Sbjct: 123 VYVAANPPSPVPEEEEEEEEDASPQSE 149
>UniRef100_Q9SHC8 Putative VAMP-associated protein (Putative VAMP (Vesicle-associated
membrane protein)-associated protein) [Arabidopsis
thaliana]
Length = 239
Score = 55.8 bits (133), Expect = 1e-06
Identities = 42/133 (31%), Positives = 69/133 (51%), Gaps = 5/133 (3%)
Query: 80 LKLDPSNKLYFPYEPGKQVRSAIRIKNTSKSNVAFKFQTTAPKSCFMRPPGAILAPGES- 138
L +DP + L FP+E KQ+ ++ + N + + VAFK +TT PK +RP ++ P S
Sbjct: 6 LTIDPVD-LQFPFELKKQISCSLYLGNKTDNYVAFKVKTTNPKKYCVRPNTGVVHPRSSS 64
Query: 139 -IIATVFKFVEQPENNEKPEKSGLKFKIMSLKVKGSIDYVPELFDEQKDQVAVEQILRVV 197
++ T+ E P + + +K L+ + S D E+F ++ E LRVV
Sbjct: 65 EVLVTMQAQKEAPADLQCKDKFLLQCVVASPGATPK-DVTHEMFSKEAGHRVEETKLRVV 123
Query: 198 FLDPER-PSPILE 209
++ P R PSP+ E
Sbjct: 124 YVAPPRPPSPVRE 136
>UniRef100_Q5U511 Hypothetical protein [Xenopus laevis]
Length = 243
Score = 55.5 bits (132), Expect = 1e-06
Identities = 36/119 (30%), Positives = 60/119 (50%), Gaps = 3/119 (2%)
Query: 80 LKLDPSNKLYFPYEPGKQVRSAIRIKNTSKSNVAFKFQTTAPKSCFMRPPGAILAPGESI 139
L+L+P +L F V + +++ N + NV FK +TTAP+ +RP ++ G SI
Sbjct: 8 LQLEPQQELKFQGPFTDVVTTNLKLGNPTDKNVCFKVKTTAPRRYCVRPNSGVIDAGSSI 67
Query: 140 IATVFKFVEQPENNEKPEKSGLKFKIMSLKVKGSIDYVPELFDEQKDQVAVEQILRVVF 198
I +V + QP + + EKS KF + S+ + ++ E K ++ LR VF
Sbjct: 68 IVSV---MLQPFDYDPNEKSKHKFMVQSIIAPSDTSDMEAVWKEAKPDDLMDSKLRCVF 123
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.313 0.130 0.369
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 458,472,838
Number of Sequences: 2790947
Number of extensions: 19390558
Number of successful extensions: 153182
Number of sequences better than 10.0: 1029
Number of HSP's better than 10.0 without gapping: 670
Number of HSP's successfully gapped in prelim test: 388
Number of HSP's that attempted gapping in prelim test: 123271
Number of HSP's gapped (non-prelim): 13647
length of query: 266
length of database: 848,049,833
effective HSP length: 125
effective length of query: 141
effective length of database: 499,181,458
effective search space: 70384585578
effective search space used: 70384585578
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 73 (32.7 bits)
Medicago: description of AC146792.8


