

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146790.10 - phase: 0
(406 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
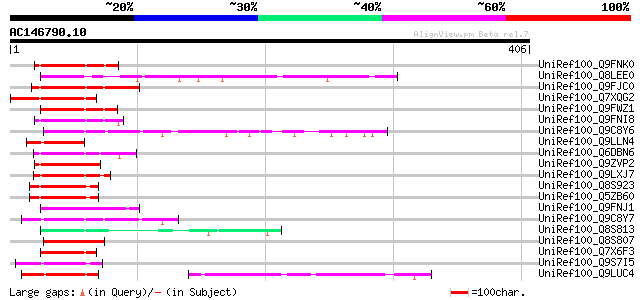
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9FNK0 Arabidopsis thaliana genomic DNA, chromosome 5,... 58 4e-07
UniRef100_Q8LEE0 Hypothetical protein [Arabidopsis thaliana] 57 9e-07
UniRef100_Q9FJC0 Heat shock transcription factor HSF30-like prot... 56 2e-06
UniRef100_Q7XQG2 OJ000114_01.15 protein [Oryza sativa] 55 3e-06
UniRef100_Q9FWZ1 Hypothetical protein F24J1.28 [Arabidopsis thal... 54 8e-06
UniRef100_Q9FNI8 Arabidopsis thaliana genomic DNA, chromosome 5,... 54 8e-06
UniRef100_Q9C8Y6 Hypothetical protein T27F4.6 [Arabidopsis thali... 54 8e-06
UniRef100_Q9LLN4 Hypothetical protein DUPR11.22 [Oryza sativa] 53 1e-05
UniRef100_Q6DBN6 At1g51370 [Arabidopsis thaliana] 52 3e-05
UniRef100_Q9ZVP2 T7A14.5 [Arabidopsis thaliana] 52 4e-05
UniRef100_Q9LXJ7 Hypothetical protein F3C22_70 [Arabidopsis thal... 51 5e-05
UniRef100_Q8S923 RNA apurinic site specific lyase [Oryza sativa] 51 7e-05
UniRef100_Q5ZB60 Putative ribosomal RNA apurinic site specific l... 51 7e-05
UniRef100_Q9FNJ1 Arabidopsis thaliana genomic DNA, chromosome 5,... 50 9e-05
UniRef100_Q9C8Y7 Hypothetical protein T27F4.5 [Arabidopsis thali... 50 9e-05
UniRef100_Q8S813 Hypothetical protein OSJNBa0020E23.11 [Oryza sa... 50 9e-05
UniRef100_Q8S807 Hypothetical protein OSJNBa0020E23.17 [Oryza sa... 50 9e-05
UniRef100_Q7X6F3 P0076O17.4 protein [Oryza sativa] 50 9e-05
UniRef100_Q9S7I5 Ribosomal RNA apurinic site specific lyase [Tri... 50 1e-04
UniRef100_Q9LUC4 Gb|AAC31233.1 [Arabidopsis thaliana] 50 1e-04
>UniRef100_Q9FNK0 Arabidopsis thaliana genomic DNA, chromosome 5, P1 clone:MDJ22
[Arabidopsis thaliana]
Length = 502
Score = 58.2 bits (139), Expect = 4e-07
Identities = 33/66 (50%), Positives = 44/66 (66%), Gaps = 3/66 (4%)
Query: 20 EDTINSKLPESLINRVLSFLPTKDVVRTCVLSKRWMNRWTSSITKLDLDDIDLSYNYNPL 79
ED I SKLPE L++++LS+LPTKD+VRT VLSKRW + W I LDLD + + Y+
Sbjct: 17 EDLI-SKLPEVLLSQILSYLPTKDIVRTSVLSKRWKSVWL-LIPGLDLDSSEFPH-YDTF 73
Query: 80 CSYCEE 85
+ E
Sbjct: 74 VDFMNE 79
>UniRef100_Q8LEE0 Hypothetical protein [Arabidopsis thaliana]
Length = 459
Score = 57.0 bits (136), Expect = 9e-07
Identities = 79/298 (26%), Positives = 129/298 (42%), Gaps = 47/298 (15%)
Query: 25 SKLPESLINRVLSFLPTKDVVRTCVLSKRWMNRWTSSITKLDLDDIDLSYNYNPLCSYCE 84
S+LPESLI ++L LPTKD V+T VLS RW N W L++ +DL+
Sbjct: 8 SELPESLITQILLCLPTKDSVKTSVLSTRWKNLW------LNVPGLDLT----------- 50
Query: 85 EICTDYCYDYDKYK---LTVCPRCHVCGNHKANLRDAEV-YERRVEIDFES--GMKIESG 138
C+D+ ++ ++Y+ + R + N ++ L+ EV Y+RR F+ G I G
Sbjct: 51 --CSDFPFEDEEYEQVFINFIDR-FLEFNPESRLQTFEVDYKRREIRGFKDRIGTAINRG 107
Query: 139 KKEQQFVN----FVGRALLLTSISSMELERF---SLLINNKRDISLQNTWISSILNRRVK 191
+ V+ + ++ + M L F +L+ RD +L + + S+ +
Sbjct: 108 IRLLDAVSSTMYWEEECIMYPYLEFMPLNLFTSKTLVTLKLRDSALNDPGLVSMPCLKFM 167
Query: 192 ILRIHSSFYQLPFSALTSHYLFNCTSLEELELVLHVSSTIKFPSISVHFGHLKLLK---- 247
ILR + L S C LEEL LV + I ++V + LK
Sbjct: 168 ILREVRWSGTMSLEKLVS----GCPVLEELTLVRTLICDIDDDELAVTRVRSRSLKTFYV 223
Query: 248 --LYGIFFKIDTSSDCLTLNLPLLRKFDIKNCNWSGGKDLIVEAPLLEIVSIEQDIEF 303
YG+ + L ++ P L +K ++ G IV L + IE +I+F
Sbjct: 224 PLAYGVGCRSRVPDKVLEIDAPGLENMTLKEDHFDG----IVVKNLTSLFMIELNIKF 277
>UniRef100_Q9FJC0 Heat shock transcription factor HSF30-like protein [Arabidopsis
thaliana]
Length = 454
Score = 55.8 bits (133), Expect = 2e-06
Identities = 34/85 (40%), Positives = 54/85 (63%), Gaps = 4/85 (4%)
Query: 18 IEEDTINSKLPESLINRVLSFLPTKDVVRTCVLSKRWMNRWTSSITKLDLDDIDLSYNYN 77
++ED I S+ ESL+ ++L++LPTKDVV+T VLS RW + W + L+LD D S ++N
Sbjct: 19 LKEDRI-SQFYESLLCQILNYLPTKDVVKTSVLSTRWRSLWL-LVPSLELDSRDFS-DFN 75
Query: 78 PLCSYCEE-ICTDYCYDYDKYKLTV 101
S+C+ ++ +K KLT+
Sbjct: 76 TFVSFCDRYFDSNRVLCINKLKLTI 100
>UniRef100_Q7XQG2 OJ000114_01.15 protein [Oryza sativa]
Length = 543
Score = 55.5 bits (132), Expect = 3e-06
Identities = 29/68 (42%), Positives = 44/68 (64%), Gaps = 2/68 (2%)
Query: 1 MEAGSSKVHKSTQMASVIEEDTINSKLPESLINRVLSFLPTKDVVRTCVLSKRWMNRWTS 60
ME S+ K T A+ ED + S+LP+ L++ +LS L + VV+TCVLS+RW + W
Sbjct: 1 MEIEGSRKRKRTPAAAAAAEDRL-SELPDCLLHDILSHLKARQVVQTCVLSRRWRHLW-R 58
Query: 61 SITKLDLD 68
S+ +LD+D
Sbjct: 59 SVPRLDVD 66
>UniRef100_Q9FWZ1 Hypothetical protein F24J1.28 [Arabidopsis thaliana]
Length = 451
Score = 53.9 bits (128), Expect = 8e-06
Identities = 29/60 (48%), Positives = 38/60 (63%), Gaps = 2/60 (3%)
Query: 25 SKLPESLINRVLSFLPTKDVVRTCVLSKRWMNRWTSSITKLDLDDIDLSYNYNPLCSYCE 84
SKLP+ L+ VL LPTKDVV+T VLS+RW N W + LDLD+ D +N S+ +
Sbjct: 21 SKLPDCLLCEVLLNLPTKDVVKTSVLSRRWRNLW-KHVPGLDLDNTDFQ-EFNTFLSFVD 78
>UniRef100_Q9FNI8 Arabidopsis thaliana genomic DNA, chromosome 5, P1 clone:MDJ22
[Arabidopsis thaliana]
Length = 457
Score = 53.9 bits (128), Expect = 8e-06
Identities = 34/79 (43%), Positives = 45/79 (56%), Gaps = 12/79 (15%)
Query: 20 EDTINSKLPESLINRVLSFLPTKDVVRTCVLSKRWMNRWTSSITKLDLDDIDLSYNYNPL 79
ED I SKLP+SLI ++L +LP KD+VRT LS RW + W I +LDLD + +YN
Sbjct: 18 EDLI-SKLPDSLITQILLYLPIKDIVRTSSLSSRWKSLWL-LIPRLDLDSEEFQ-DYNAF 74
Query: 80 CSYC---------EEICTD 89
+ E+IC D
Sbjct: 75 VGFMNKFIDFSGEEKICLD 93
>UniRef100_Q9C8Y6 Hypothetical protein T27F4.6 [Arabidopsis thaliana]
Length = 442
Score = 53.9 bits (128), Expect = 8e-06
Identities = 88/317 (27%), Positives = 139/317 (43%), Gaps = 77/317 (24%)
Query: 27 LPESLINRVLSFLPTKDVVRTCVLSKRWMNRWTSSITKLDLDDIDLSYNYNPLCSYCEEI 86
LPESL+ +L LPTKDVV+T VLS +W N W + LDLD D + N N S+ +
Sbjct: 24 LPESLLCHILLNLPTKDVVKTSVLSSKWRNLW-RLVPGLDLDSSDFTEN-NTFVSFIDRF 81
Query: 87 CTDYCYDY-DKYKLTVCPRCHVCGNH-KANLRDA----EVYERRVEIDFESGMKIESGKK 140
+ + Y K+KL C++ G+ N A +V +R+V + + G
Sbjct: 82 MSFHSDLYLKKFKLRFF--CNLNGDEVSENAHIARWINDVVKRKVH-----NLDLTWGAV 134
Query: 141 EQQFVNFVGRALLLTSISSMELERFSLL------------INNKRDISLQNTWISSIL-- 186
E + ++ +L+ + + L L + D++L+ IS L
Sbjct: 135 EIPPILYLCNSLVSLKLCGVTLPNLELTSLPCVKVIVLEWVKFANDLALE-MLISGCLVL 193
Query: 187 ---------NRRVKILRIHSSFYQLPFSALTSHYLFNCTSLEEL--ELVLHVSSTIKFPS 235
N VKILR+ SS L FS +N +S + L +LVL +++
Sbjct: 194 ESLTLCRRPNDNVKILRV-SSQSLLRFS-------YNGSSYKGLHDDLVLEINAP----- 240
Query: 236 ISVHFGHLKLLKLYG----IFFKIDTSSDCL--TLNLPLLRKFDIKN-------CNWSGG 282
LK+LKL+ F +TSS + +N+ L +KFD K+ CN+ G
Sbjct: 241 ------KLKILKLFSHQLTTSFIRNTSSSIVEADINIGLGKKFDPKDLPKRNVICNFLAG 294
Query: 283 ----KDLIVEAPLLEIV 295
K+L + LE++
Sbjct: 295 ISSVKNLFIAPCTLEVI 311
>UniRef100_Q9LLN4 Hypothetical protein DUPR11.22 [Oryza sativa]
Length = 431
Score = 53.1 bits (126), Expect = 1e-05
Identities = 24/45 (53%), Positives = 33/45 (73%), Gaps = 1/45 (2%)
Query: 14 MASVIEEDTINSKLPESLINRVLSFLPTKDVVRTCVLSKRWMNRW 58
MA EED I+ LP++L+ +LSFLP +D V+TCVLS+RW + W
Sbjct: 1 MAEASEEDGIDV-LPDALLQHILSFLPAEDAVKTCVLSRRWRHLW 44
>UniRef100_Q6DBN6 At1g51370 [Arabidopsis thaliana]
Length = 435
Score = 52.0 bits (123), Expect = 3e-05
Identities = 38/93 (40%), Positives = 48/93 (50%), Gaps = 15/93 (16%)
Query: 19 EEDTINSKLPESLINRVLSFLPTKDVVRTCVLSKRWMNRWTSSITKLDLDDIDLSYNYNP 78
EED I S+LPE LI+ +L L TKD VRT LS +W W S+ LDLD S N N
Sbjct: 17 EEDRI-SQLPEPLISEILFHLSTKDSVRTSALSTKWRYLW-QSVPGLDLDPY-ASSNTNT 73
Query: 79 LCSYCE------------EICTDYCYDYDKYKL 99
+ S+ E ++ D Y +DKY L
Sbjct: 74 IVSFVESFFDSHRDSWIRKLRLDLGYHHDKYDL 106
>UniRef100_Q9ZVP2 T7A14.5 [Arabidopsis thaliana]
Length = 467
Score = 51.6 bits (122), Expect = 4e-05
Identities = 28/52 (53%), Positives = 34/52 (64%), Gaps = 1/52 (1%)
Query: 20 EDTINSKLPESLINRVLSFLPTKDVVRTCVLSKRWMNRWTSSITKLDLDDID 71
ED I S LPE L+ +L LPTKDVV T +LSKRW++ WT T DD+D
Sbjct: 12 EDRI-SVLPEDLLVVILDLLPTKDVVATMILSKRWLSIWTMVRTLEYTDDMD 62
>UniRef100_Q9LXJ7 Hypothetical protein F3C22_70 [Arabidopsis thaliana]
Length = 384
Score = 51.2 bits (121), Expect = 5e-05
Identities = 29/61 (47%), Positives = 40/61 (65%), Gaps = 4/61 (6%)
Query: 19 EEDTINSKLPESLINRVLSFLPTKDVVRTCVLSKRWMNRWTSSITKLDLDDIDLSYNYNP 78
+ED +N +LPE LI R+LSFLPT+ V+ T VLSK+W + W + L+ D D Y NP
Sbjct: 8 KEDRMN-QLPEDLILRILSFLPTELVIATSVLSKQWRSLW-KLVPNLEFDSDD--YEINP 63
Query: 79 L 79
+
Sbjct: 64 V 64
>UniRef100_Q8S923 RNA apurinic site specific lyase [Oryza sativa]
Length = 475
Score = 50.8 bits (120), Expect = 7e-05
Identities = 28/54 (51%), Positives = 35/54 (63%), Gaps = 3/54 (5%)
Query: 16 SVIEEDTINSKLPESLINRVLSFLPTKDVVRTCVLSKRWMNRWTSSITKLDLDD 69
S+ EED I S+LP++L+ RV+S LPTKD RT +S RW W S L LDD
Sbjct: 37 SLDEEDRI-SRLPDALLRRVVSRLPTKDAARTAAISSRWRRVWAS--IPLHLDD 87
>UniRef100_Q5ZB60 Putative ribosomal RNA apurinic site specific lyase [Oryza
sativa]
Length = 433
Score = 50.8 bits (120), Expect = 7e-05
Identities = 28/54 (51%), Positives = 35/54 (63%), Gaps = 3/54 (5%)
Query: 16 SVIEEDTINSKLPESLINRVLSFLPTKDVVRTCVLSKRWMNRWTSSITKLDLDD 69
S+ EED I S+LP++L+ RV+S LPTKD RT +S RW W S L LDD
Sbjct: 37 SLDEEDRI-SRLPDALLRRVVSRLPTKDAARTAAISSRWRRVWAS--IPLHLDD 87
>UniRef100_Q9FNJ1 Arabidopsis thaliana genomic DNA, chromosome 5, P1 clone:MDJ22
[Arabidopsis thaliana]
Length = 452
Score = 50.4 bits (119), Expect = 9e-05
Identities = 29/78 (37%), Positives = 44/78 (56%), Gaps = 3/78 (3%)
Query: 25 SKLPESLINRVLSFLPTKDVVRTCVLSKRWMNRWTSS-ITKLDLDDIDLSYNYNPLCSYC 83
S LP+ L+ ++LS LPTK+ V T +LS RW + W S+ + +D+D D + + S
Sbjct: 24 SSLPDELLCQILSNLPTKNAVTTSILSTRWRSIWLSTPVLDIDIDAFDDATTFVSFASRF 83
Query: 84 EEICTDYCYDYDKYKLTV 101
E D C K+KL+V
Sbjct: 84 LEFSKDSC--LHKFKLSV 99
>UniRef100_Q9C8Y7 Hypothetical protein T27F4.5 [Arabidopsis thaliana]
Length = 456
Score = 50.4 bits (119), Expect = 9e-05
Identities = 45/130 (34%), Positives = 67/130 (50%), Gaps = 12/130 (9%)
Query: 10 KSTQMASVIEEDTINSKLPESLINRVLSFLPTKDVVRTCVLSKRWMNRWTSSITKLDLDD 69
+S+ S E D + LPESLI VL L TKDV++ CVLS +W W + LDLD
Sbjct: 18 ESSDKKSGYEVDWVRD-LPESLICHVLLNLSTKDVIKNCVLSTKWRYLW-RYVPGLDLDC 75
Query: 70 IDLSYNYNPLCSYCEEICTDYCYDY-DKYKLTVCPRCHVCGNHK-ANLRDA----EVYER 123
D + YN S+ + + Y +K+KL C + G+ + N + A +V +R
Sbjct: 76 SDFT-EYNTFVSFVDRFLSTNTESYLNKFKLGF--DCDLVGDEETGNAQMARWINDVVKR 132
Query: 124 RVE-IDFESG 132
+V+ +D E G
Sbjct: 133 KVQHLDLEWG 142
>UniRef100_Q8S813 Hypothetical protein OSJNBa0020E23.11 [Oryza sativa]
Length = 454
Score = 50.4 bits (119), Expect = 9e-05
Identities = 55/194 (28%), Positives = 75/194 (38%), Gaps = 57/194 (29%)
Query: 25 SKLPESLINRVLSFLPTKDVVRTCVLSKRWMNRWTSSITKLDLDDIDLSYNYNPLCSYCE 84
S LPE +++RV+SFL ++ VRTCVLS+RW + W ++ ++ D D + N+
Sbjct: 65 SDLPEGVLHRVMSFLDSRQAVRTCVLSRRWRDVW-RTVPRVHADFCDFTLNWTS------ 117
Query: 85 EICTDYCYDYDKYKLTVCPRCHVCGNHKANLRDAEVYERRVEIDFESGMKIESGKKEQQF 144
D EV E V D E F
Sbjct: 118 -------------------------------DDDEVDEAAVAED------------EVVF 134
Query: 145 VNFVGRALLL----TSISSMELERFSLLINNKRDISLQNTWISSILNRRVKILRIHSSFY 200
FV R L L SI S L RF + N WIS L + V+IL + Y
Sbjct: 135 NRFVNRLLELRDPNASIRSFFL-RFCRSDGGDDGSAEGNRWISYALQKNVRILEVSVLSY 193
Query: 201 --QLPFSALTSHYL 212
+L S +S YL
Sbjct: 194 ALELDHSVFSSRYL 207
>UniRef100_Q8S807 Hypothetical protein OSJNBa0020E23.17 [Oryza sativa]
Length = 450
Score = 50.4 bits (119), Expect = 9e-05
Identities = 22/49 (44%), Positives = 31/49 (62%), Gaps = 1/49 (2%)
Query: 27 LPESLINRVLSFLPTKDVVRTCVLSKRWMNRW-TSSITKLDLDDIDLSY 74
LPE +++ ++SFL + VRTCVLS+RW N W T D D+ DL +
Sbjct: 21 LPEEVLHHIMSFLDARQAVRTCVLSRRWRNLWRTVPCINADFDEFDLVF 69
>UniRef100_Q7X6F3 P0076O17.4 protein [Oryza sativa]
Length = 457
Score = 50.4 bits (119), Expect = 9e-05
Identities = 22/44 (50%), Positives = 33/44 (75%), Gaps = 1/44 (2%)
Query: 25 SKLPESLINRVLSFLPTKDVVRTCVLSKRWMNRWTSSITKLDLD 68
S LP+ L++ V+SFLP + + +TCVLSKRW++ W S+ L+LD
Sbjct: 5 SALPDGLLHAVMSFLPARQMAQTCVLSKRWVHLW-RSVPSLNLD 47
>UniRef100_Q9S7I5 Ribosomal RNA apurinic site specific lyase [Triticum aestivum]
Length = 456
Score = 50.1 bits (118), Expect = 1e-04
Identities = 28/68 (41%), Positives = 39/68 (57%), Gaps = 3/68 (4%)
Query: 5 SSKVHKSTQMASVIEEDTINSKLPESLINRVLSFLPTKDVVRTCVLSKRWMNRWTSSITK 64
S+ S+ + ++D IN LP+ L+ ++S LPTKD RT +LS RW + W S T
Sbjct: 18 STTASPSSSSTASDDDDRINC-LPDVLLRNIISRLPTKDAARTTILSSRWRSLWAS--TP 74
Query: 65 LDLDDIDL 72
L LDD L
Sbjct: 75 LRLDDAGL 82
>UniRef100_Q9LUC4 Gb|AAC31233.1 [Arabidopsis thaliana]
Length = 442
Score = 50.1 bits (118), Expect = 1e-04
Identities = 53/196 (27%), Positives = 89/196 (45%), Gaps = 21/196 (10%)
Query: 141 EQQFVNFVGRALLLTSISSMELERFSLLINNKRDISLQNTWISSILNRRVKILRIHSSFY 200
+ +F +FV R ++ I S + F L N D +L +W+S +L R ++ L + +
Sbjct: 80 DSRFTDFVER--VVGDIGSPHISSFYLCSVNTFDETLLFSWLSKVLKRNLQRLVVTCNDL 137
Query: 201 QL-PFSALTSHYLFNCTSLEELELVLHVSSTIKFPSISVHFGHLKLLKLYGI-FFKIDTS 258
++ FS+L ++ SL EL L I P+I +LK L L F + +
Sbjct: 138 EIISFSSLFPSFV----SLVELRLRTKSILDISAPAI---LPNLKFLSLEDARIFNMSSV 190
Query: 259 SDCLTLNLPLLRKFDIKNCNWSGGKDLIVEAPLLEIVSIEQDIEFYNAASHDLHSQS--- 315
S L LN P+L F+ C +I+++PLL I E + S + + S
Sbjct: 191 SKNLVLNFPVLETFEASYCRCFRTDTVIIDSPLLRI------FEMFKCTSEHVPNPSQVC 244
Query: 316 -IKFNALHLKQFTYSG 330
I+ A L++ T+ G
Sbjct: 245 KIRVLASKLEKITFCG 260
Score = 48.5 bits (114), Expect = 3e-04
Identities = 29/60 (48%), Positives = 38/60 (63%), Gaps = 2/60 (3%)
Query: 10 KSTQMASVIEEDTINSKLPESLINRVLSFLPTKDVVRTCVLSKRWMNRWTSSITKLDLDD 69
K T++ S ED +S L ES+++ +LS LPT + V T VLSK W N WT +IT L DD
Sbjct: 15 KKTKLCSENFEDKFSSLL-ESVVSIILSQLPTAEAVSTSVLSKSWKNIWT-NITDLHFDD 72
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.324 0.138 0.413
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 656,887,187
Number of Sequences: 2790947
Number of extensions: 27126209
Number of successful extensions: 83349
Number of sequences better than 10.0: 315
Number of HSP's better than 10.0 without gapping: 272
Number of HSP's successfully gapped in prelim test: 43
Number of HSP's that attempted gapping in prelim test: 82959
Number of HSP's gapped (non-prelim): 421
length of query: 406
length of database: 848,049,833
effective HSP length: 130
effective length of query: 276
effective length of database: 485,226,723
effective search space: 133922575548
effective search space used: 133922575548
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.5 bits)
S2: 76 (33.9 bits)
Medicago: description of AC146790.10


