

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146789.8 + phase: 0 /pseudo
(2263 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
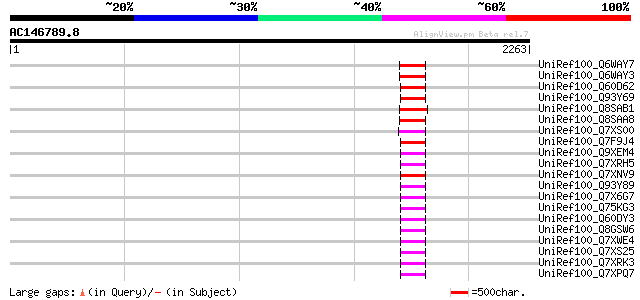
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q6WAY7 Gag/pol polyprotein [Pisum sativum] 181 2e-43
UniRef100_Q6WAY3 Gag/pol polyprotein [Pisum sativum] 179 1e-42
UniRef100_Q60D62 Putative gag/pol polyprotein, 3'-partial [Solan... 112 2e-22
UniRef100_Q93Y69 Putative gag-pol [Oryza sativa] 94 6e-17
UniRef100_Q8SAB1 Putative GAG-POL-orf2 protein [Sorghum bicolor] 93 1e-16
UniRef100_Q8SAA8 Putative GAG-POL-orf2 protein [Sorghum bicolor] 93 1e-16
UniRef100_Q7XS00 OSJNBb0002N06.6 protein [Oryza sativa] 91 5e-16
UniRef100_Q7F9J4 OSJNBa0013A04.10 protein [Oryza sativa] 88 3e-15
UniRef100_Q9XEM4 Polyprotein [Oryza sativa] 88 3e-15
UniRef100_Q7XRH5 OSJNBb0003A12.14 protein [Oryza sativa] 88 3e-15
UniRef100_Q7XNV9 OSJNBb0015G09.7 protein [Oryza sativa] 88 3e-15
UniRef100_Q93Y89 Gag-pol [Oryza sativa] 87 5e-15
UniRef100_Q7X6G7 OSJNBa0079F16.18 protein [Oryza sativa] 87 5e-15
UniRef100_Q75KG3 Putative polyprotein [Oryza sativa] 87 5e-15
UniRef100_Q60DY3 Putative polyprotein [Oryza sativa] 87 5e-15
UniRef100_Q8GSW6 Putative gag-pol polyprotein [Oryza sativa] 87 5e-15
UniRef100_Q7XWE4 OSJNBa0035O13.3 protein [Oryza sativa] 87 5e-15
UniRef100_Q7XS25 OSJNBa0080E14.11 protein [Oryza sativa] 87 5e-15
UniRef100_Q7XRK3 OSJNBa0042D13.7 protein [Oryza sativa] 87 5e-15
UniRef100_Q7XPQ7 OSJNBa0053K19.16 protein [Oryza sativa] 87 5e-15
>UniRef100_Q6WAY7 Gag/pol polyprotein [Pisum sativum]
Length = 1814
Score = 181 bits (459), Expect = 2e-43
Identities = 85/115 (73%), Positives = 98/115 (84%)
Query: 1699 LSMYTNGIGAVLLTPKGAHIPFTARLRFDCTNNIAEYEACIMGIEEAIDLRIKNIEIYGD 1758
++M NG+GAVL+ PKGAHIPF+ARL FD TNN AEYEACIMGIEEAIDLRIK ++IYGD
Sbjct: 1696 VNMNGNGVGAVLINPKGAHIPFSARLTFDVTNNEAEYEACIMGIEEAIDLRIKTLDIYGD 1755
Query: 1759 SALVINQIKGKWETLHAGLIPYRDYARRLLTFFNKVELHHIPRDENQMADALATV 1813
SALV+NQ+ G W T LIPYRDY RR+LTFF KV L+H+PRDENQMADALAT+
Sbjct: 1756 SALVVNQVNGDWNTNQPHLIPYRDYTRRILTFFKKVRLYHVPRDENQMADALATL 1810
>UniRef100_Q6WAY3 Gag/pol polyprotein [Pisum sativum]
Length = 2262
Score = 179 bits (453), Expect = 1e-42
Identities = 83/115 (72%), Positives = 99/115 (85%)
Query: 1699 LSMYTNGIGAVLLTPKGAHIPFTARLRFDCTNNIAEYEACIMGIEEAIDLRIKNIEIYGD 1758
++M NG+GAVL+ PKGAH+PF+ARL FD TNN AEYEACIMGIEEAIDLRIK ++I+GD
Sbjct: 1694 VNMNGNGVGAVLINPKGAHMPFSARLTFDVTNNEAEYEACIMGIEEAIDLRIKTLDIFGD 1753
Query: 1759 SALVINQIKGKWETLHAGLIPYRDYARRLLTFFNKVELHHIPRDENQMADALATV 1813
SALV+NQ+ G W T LIPYRDY RR+LTFF KV+L+H+PRDENQMADALAT+
Sbjct: 1754 SALVVNQVNGDWNTNQPHLIPYRDYTRRILTFFKKVKLYHVPRDENQMADALATL 1808
>UniRef100_Q60D62 Putative gag/pol polyprotein, 3'-partial [Solanum demissum]
Length = 1784
Score = 112 bits (279), Expect = 2e-22
Identities = 53/109 (48%), Positives = 77/109 (70%)
Query: 1705 GIGAVLLTPKGAHIPFTARLRFDCTNNIAEYEACIMGIEEAIDLRIKNIEIYGDSALVIN 1764
GIGAVL++ G + P TA+LRF CTNN++EYEACI+G+ A D+ I+ + + GDS L+++
Sbjct: 298 GIGAVLMSESGEYFPITAQLRFYCTNNMSEYEACILGLRLAADMGIQELLVLGDSDLLVH 357
Query: 1765 QIKGKWETLHAGLIPYRDYARRLLTFFNKVELHHIPRDENQMADALATV 1813
QI+G+WET LIPY+ + L F ++ HI R N++ADALAT+
Sbjct: 358 QIQGEWETRDPKLIPYQHCLQDLCRRFVSIKFRHISRVHNEIADALATL 406
>UniRef100_Q93Y69 Putative gag-pol [Oryza sativa]
Length = 1262
Score = 93.6 bits (231), Expect = 6e-17
Identities = 47/108 (43%), Positives = 68/108 (62%), Gaps = 1/108 (0%)
Query: 1705 GIGAVLLTPKGAHIPFT-ARLRFDCTNNIAEYEACIMGIEEAIDLRIKNIEIYGDSALVI 1763
G G V TP+G I + + L+ +C+NN AEYEA I G+ A+ + +++I +YGDS L++
Sbjct: 759 GAGLVFKTPQGGVIYHSFSLLKEECSNNEAEYEALIFGLLLALSMEVRSIRVYGDSQLIV 818
Query: 1764 NQIKGKWETLHAGLIPYRDYARRLLTFFNKVELHHIPRDENQMADALA 1811
QI +E L L+PY ARRL+ F +E+ H+PR N ADALA
Sbjct: 819 QQINDIYEVLKQELVPYYSAARRLMEMFGHIEVMHVPRSRNAPADALA 866
>UniRef100_Q8SAB1 Putative GAG-POL-orf2 protein [Sorghum bicolor]
Length = 756
Score = 92.8 bits (229), Expect = 1e-16
Identities = 48/122 (39%), Positives = 75/122 (61%), Gaps = 2/122 (1%)
Query: 1700 SMYTNGIGAVLL--TPKGAHIPFTARLRFDCTNNIAEYEACIMGIEEAIDLRIKNIEIYG 1757
S+ NG GA +L +P + + R+ F +NN+AEYEAC+ GI A++L IK + +YG
Sbjct: 288 SLNINGAGAGILFVSPNKDKLRYILRILFPASNNVAEYEACLHGIRLAVELGIKRLYVYG 347
Query: 1758 DSALVINQIKGKWETLHAGLIPYRDYARRLLTFFNKVELHHIPRDENQMADALATVFNDQ 1817
D ALVINQ+ +W+ H + Y + R+ T F +E H+ RD+NQ ADAL+ + + +
Sbjct: 348 DFALVINQLNKEWDATHEKMDLYCNEIRKWETNFYGIEYIHVVRDKNQAADALSKLGSSR 407
Query: 1818 GE 1819
+
Sbjct: 408 AQ 409
>UniRef100_Q8SAA8 Putative GAG-POL-orf2 protein [Sorghum bicolor]
Length = 678
Score = 92.8 bits (229), Expect = 1e-16
Identities = 47/114 (41%), Positives = 72/114 (62%), Gaps = 2/114 (1%)
Query: 1700 SMYTNGIGAVLL--TPKGAHIPFTARLRFDCTNNIAEYEACIMGIEEAIDLRIKNIEIYG 1757
S+ NG GA +L +P + + R+ F +NN+AEYEAC+ GI A++L +K + ++G
Sbjct: 558 SLNINGAGAGILFVSPNKDKLRYVLRILFPASNNVAEYEACLHGIRLAVELGVKRLYVHG 617
Query: 1758 DSALVINQIKGKWETLHAGLIPYRDYARRLLTFFNKVELHHIPRDENQMADALA 1811
DSALVINQ+ +W+T H + Y R+ + F +E H+ RD+NQ ADAL+
Sbjct: 618 DSALVINQLNKEWDTTHEKMDLYCKEIRKWKSNFYGIEYIHVVRDKNQAADALS 671
>UniRef100_Q7XS00 OSJNBb0002N06.6 protein [Oryza sativa]
Length = 1607
Score = 90.5 bits (223), Expect = 5e-16
Identities = 47/116 (40%), Positives = 68/116 (58%)
Query: 1696 LMGLSMYTNGIGAVLLTPKGAHIPFTARLRFDCTNNIAEYEACIMGIEEAIDLRIKNIEI 1755
L G GI AVL +PKG I +TARL+FD TNN AEYEA ++G+ +A L ++ + I
Sbjct: 1244 LQGAKTRGAGIAAVLFSPKGVPIRYTARLQFDTTNNAAEYEAVLLGLRKAKALGVRRLLI 1303
Query: 1756 YGDSALVINQIKGKWETLHAGLIPYRDYARRLLTFFNKVELHHIPRDENQMADALA 1811
DS LV + +E G+ Y + R + F + + H+P+D+N+ ADALA
Sbjct: 1304 RTDSKLVAGHVDKSFEAKEEGMKRYLEAVRSMEKCFTGITVEHLPKDQNEEADALA 1359
>UniRef100_Q7F9J4 OSJNBa0013A04.10 protein [Oryza sativa]
Length = 1230
Score = 88.2 bits (217), Expect = 3e-15
Identities = 42/107 (39%), Positives = 65/107 (60%)
Query: 1705 GIGAVLLTPKGAHIPFTARLRFDCTNNIAEYEACIMGIEEAIDLRIKNIEIYGDSALVIN 1764
G+G VL++P G + + + F ++N+AEYEA + G+ AI L I+ + + GDS LV+N
Sbjct: 1095 GVGVVLISPTGERLSYILWIHFSASHNVAEYEALLHGLRIAISLGIRRLIVRGDSQLVVN 1154
Query: 1765 QIKGKWETLHAGLIPYRDYARRLLTFFNKVELHHIPRDENQMADALA 1811
Q+ +W L +I YR R+L F+ +EL H+ R N+ AD LA
Sbjct: 1155 QVMKEWSCLDDNMIAYRQEVRKLEDKFDGLELTHVLRHNNEAADRLA 1201
>UniRef100_Q9XEM4 Polyprotein [Oryza sativa]
Length = 829
Score = 87.8 bits (216), Expect = 3e-15
Identities = 43/107 (40%), Positives = 64/107 (59%)
Query: 1705 GIGAVLLTPKGAHIPFTARLRFDCTNNIAEYEACIMGIEEAIDLRIKNIEIYGDSALVIN 1764
G G VL++P G + + + F ++NIAEYEA + G+ AI L IK + + GDS LV+N
Sbjct: 269 GAGVVLISPTGERLSYVLWIHFSASHNIAEYEALLHGLRIAISLGIKRLIVRGDSQLVVN 328
Query: 1765 QIKGKWETLHAGLIPYRDYARRLLTFFNKVELHHIPRDENQMADALA 1811
Q+ +W L ++ YR R+L F+ +EL H+ R N+ AD LA
Sbjct: 329 QVMKEWSCLDDNMMAYRQEVRKLEDKFDGLELSHVLRHNNEAADRLA 375
>UniRef100_Q7XRH5 OSJNBb0003A12.14 protein [Oryza sativa]
Length = 1863
Score = 87.8 bits (216), Expect = 3e-15
Identities = 43/107 (40%), Positives = 64/107 (59%)
Query: 1705 GIGAVLLTPKGAHIPFTARLRFDCTNNIAEYEACIMGIEEAIDLRIKNIEIYGDSALVIN 1764
G G VL++P G + + + F ++NIAEYEA + G+ AI L IK + + GDS LV+N
Sbjct: 1303 GAGVVLISPTGERLSYVLWIHFSASHNIAEYEALLHGLRIAISLGIKRLIVRGDSQLVVN 1362
Query: 1765 QIKGKWETLHAGLIPYRDYARRLLTFFNKVELHHIPRDENQMADALA 1811
Q+ +W L ++ YR R+L F+ +EL H+ R N+ AD LA
Sbjct: 1363 QVMKEWSCLDDNMMAYRQEVRKLEDKFDGLELSHVLRHNNEAADRLA 1409
>UniRef100_Q7XNV9 OSJNBb0015G09.7 protein [Oryza sativa]
Length = 1991
Score = 87.8 bits (216), Expect = 3e-15
Identities = 42/107 (39%), Positives = 65/107 (60%)
Query: 1705 GIGAVLLTPKGAHIPFTARLRFDCTNNIAEYEACIMGIEEAIDLRIKNIEIYGDSALVIN 1764
G G VL++P G + + + F ++N+AEYEA + G+ AI L IK++ + GDS LV+N
Sbjct: 1431 GAGVVLISPTGERLSYVLWIHFSASHNVAEYEALLHGLRIAISLGIKHLIVRGDSQLVVN 1490
Query: 1765 QIKGKWETLHAGLIPYRDYARRLLTFFNKVELHHIPRDENQMADALA 1811
Q+ +W L ++ YR R+L F+ +EL H+ R N+ AD LA
Sbjct: 1491 QVMKEWSCLDDNMMAYRQEVRKLEDKFDGLELSHVLRHNNEAADRLA 1537
>UniRef100_Q93Y89 Gag-pol [Oryza sativa]
Length = 2017
Score = 87.4 bits (215), Expect = 5e-15
Identities = 42/107 (39%), Positives = 64/107 (59%)
Query: 1705 GIGAVLLTPKGAHIPFTARLRFDCTNNIAEYEACIMGIEEAIDLRIKNIEIYGDSALVIN 1764
G G VL++P G + + + F ++N+AEYEA + G+ AI L IK + + GDS LV+N
Sbjct: 1457 GAGVVLISPTGERLSYVLWIHFSASHNVAEYEALLHGLRIAISLGIKRLIVRGDSQLVVN 1516
Query: 1765 QIKGKWETLHAGLIPYRDYARRLLTFFNKVELHHIPRDENQMADALA 1811
Q+ +W L ++ YR R+L F+ +EL H+ R N+ AD LA
Sbjct: 1517 QVMKEWSCLDDNMMAYRQEVRKLEDKFDGLELSHVLRHNNEAADRLA 1563
>UniRef100_Q7X6G7 OSJNBa0079F16.18 protein [Oryza sativa]
Length = 1802
Score = 87.4 bits (215), Expect = 5e-15
Identities = 42/107 (39%), Positives = 64/107 (59%)
Query: 1705 GIGAVLLTPKGAHIPFTARLRFDCTNNIAEYEACIMGIEEAIDLRIKNIEIYGDSALVIN 1764
G G VL++P G + + + F ++N+AEYEA + G+ AI L IK + + GDS LV+N
Sbjct: 1457 GAGVVLISPTGERLSYVLWIHFSASHNVAEYEALLHGLRIAISLGIKRLIVRGDSQLVVN 1516
Query: 1765 QIKGKWETLHAGLIPYRDYARRLLTFFNKVELHHIPRDENQMADALA 1811
Q+ +W L ++ YR R+L F+ +EL H+ R N+ AD LA
Sbjct: 1517 QVMKEWSCLDDNMMAYRQEVRKLEDKFDGLELSHVLRHNNEAADRLA 1563
>UniRef100_Q75KG3 Putative polyprotein [Oryza sativa]
Length = 2000
Score = 87.4 bits (215), Expect = 5e-15
Identities = 42/107 (39%), Positives = 64/107 (59%)
Query: 1705 GIGAVLLTPKGAHIPFTARLRFDCTNNIAEYEACIMGIEEAIDLRIKNIEIYGDSALVIN 1764
G G VL++P G + + + F ++N+AEYEA + G+ AI L IK + + GDS LV+N
Sbjct: 1440 GAGVVLISPTGERLSYVLWIHFSASHNVAEYEALLHGLRIAISLGIKRLIVRGDSQLVVN 1499
Query: 1765 QIKGKWETLHAGLIPYRDYARRLLTFFNKVELHHIPRDENQMADALA 1811
Q+ +W L ++ YR R+L F+ +EL H+ R N+ AD LA
Sbjct: 1500 QVMKEWSCLDDNMMAYRQEVRKLEDKFDGLELSHVLRHNNEAADRLA 1546
>UniRef100_Q60DY3 Putative polyprotein [Oryza sativa]
Length = 1799
Score = 87.4 bits (215), Expect = 5e-15
Identities = 42/107 (39%), Positives = 64/107 (59%)
Query: 1705 GIGAVLLTPKGAHIPFTARLRFDCTNNIAEYEACIMGIEEAIDLRIKNIEIYGDSALVIN 1764
G G VL++P G + + + F ++N+AEYEA + G+ AI L IK + + GDS LV+N
Sbjct: 1239 GAGVVLISPTGERLSYVLWIHFSASHNVAEYEALLHGLRIAISLGIKRLIVRGDSQLVVN 1298
Query: 1765 QIKGKWETLHAGLIPYRDYARRLLTFFNKVELHHIPRDENQMADALA 1811
Q+ +W L ++ YR R+L F+ +EL H+ R N+ AD LA
Sbjct: 1299 QVMKEWSCLDDNMMAYRQEVRKLEDKFDGLELSHVLRHNNEAADRLA 1345
>UniRef100_Q8GSW6 Putative gag-pol polyprotein [Oryza sativa]
Length = 2012
Score = 87.4 bits (215), Expect = 5e-15
Identities = 42/107 (39%), Positives = 64/107 (59%)
Query: 1705 GIGAVLLTPKGAHIPFTARLRFDCTNNIAEYEACIMGIEEAIDLRIKNIEIYGDSALVIN 1764
G G VL++P G + + + F ++N+AEYEA + G+ AI L IK + + GDS LV+N
Sbjct: 1452 GAGVVLISPTGERLSYVLWIHFSASHNVAEYEALLHGLRIAISLGIKRLIVRGDSQLVVN 1511
Query: 1765 QIKGKWETLHAGLIPYRDYARRLLTFFNKVELHHIPRDENQMADALA 1811
Q+ +W L ++ YR R+L F+ +EL H+ R N+ AD LA
Sbjct: 1512 QVMKEWSCLDDNMMAYRQEVRKLEDKFDGLELSHVLRHNNEAADRLA 1558
>UniRef100_Q7XWE4 OSJNBa0035O13.3 protein [Oryza sativa]
Length = 2008
Score = 87.4 bits (215), Expect = 5e-15
Identities = 42/107 (39%), Positives = 64/107 (59%)
Query: 1705 GIGAVLLTPKGAHIPFTARLRFDCTNNIAEYEACIMGIEEAIDLRIKNIEIYGDSALVIN 1764
G G VL++P G + + + F ++N+AEYEA + G+ AI L IK + + GDS LV+N
Sbjct: 1448 GAGVVLISPTGERLSYVLWIHFSASHNVAEYEALLHGLRIAISLGIKRLIVRGDSQLVVN 1507
Query: 1765 QIKGKWETLHAGLIPYRDYARRLLTFFNKVELHHIPRDENQMADALA 1811
Q+ +W L ++ YR R+L F+ +EL H+ R N+ AD LA
Sbjct: 1508 QVMKEWSCLDDNMMAYRQEVRKLEDKFDGLELSHVLRHNNEAADRLA 1554
>UniRef100_Q7XS25 OSJNBa0080E14.11 protein [Oryza sativa]
Length = 2001
Score = 87.4 bits (215), Expect = 5e-15
Identities = 42/107 (39%), Positives = 64/107 (59%)
Query: 1705 GIGAVLLTPKGAHIPFTARLRFDCTNNIAEYEACIMGIEEAIDLRIKNIEIYGDSALVIN 1764
G G VL++P G + + + F ++N+AEYEA + G+ AI L IK + + GDS LV+N
Sbjct: 1457 GAGVVLISPAGERLSYVLWIHFSASHNVAEYEALLHGLRIAISLGIKRLIVRGDSQLVVN 1516
Query: 1765 QIKGKWETLHAGLIPYRDYARRLLTFFNKVELHHIPRDENQMADALA 1811
Q+ +W L ++ YR R+L F+ +EL H+ R N+ AD LA
Sbjct: 1517 QVMKEWSCLDDNMMAYRQEVRKLEDKFDGLELSHVLRHNNEAADRLA 1563
>UniRef100_Q7XRK3 OSJNBa0042D13.7 protein [Oryza sativa]
Length = 909
Score = 87.4 bits (215), Expect = 5e-15
Identities = 42/107 (39%), Positives = 64/107 (59%)
Query: 1705 GIGAVLLTPKGAHIPFTARLRFDCTNNIAEYEACIMGIEEAIDLRIKNIEIYGDSALVIN 1764
G G VL++P G + + + F ++N+AEYEA + G+ AI L IK + + GDS LV+N
Sbjct: 349 GAGVVLISPTGERLSYVLWIHFSASHNVAEYEALLHGLRIAISLGIKRLIVRGDSQLVVN 408
Query: 1765 QIKGKWETLHAGLIPYRDYARRLLTFFNKVELHHIPRDENQMADALA 1811
Q+ +W L ++ YR R+L F+ +EL H+ R N+ AD LA
Sbjct: 409 QVMKEWSCLDDNMMAYRQEVRKLEDKFDGLELSHVLRHNNEAADRLA 455
>UniRef100_Q7XPQ7 OSJNBa0053K19.16 protein [Oryza sativa]
Length = 2010
Score = 87.4 bits (215), Expect = 5e-15
Identities = 42/107 (39%), Positives = 64/107 (59%)
Query: 1705 GIGAVLLTPKGAHIPFTARLRFDCTNNIAEYEACIMGIEEAIDLRIKNIEIYGDSALVIN 1764
G G VL++P G + + + F ++N+AEYEA + G+ AI L IK + + GDS LV+N
Sbjct: 1450 GAGVVLISPTGERLSYVLWIHFSASHNVAEYEALLHGLRIAISLGIKRLIVRGDSQLVVN 1509
Query: 1765 QIKGKWETLHAGLIPYRDYARRLLTFFNKVELHHIPRDENQMADALA 1811
Q+ +W L ++ YR R+L F+ +EL H+ R N+ AD LA
Sbjct: 1510 QVMKEWSCLDDNMMAYRQEVRKLEDKFDGLELSHVLRHNNEAADRLA 1556
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.357 0.157 0.557
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,053,707,743
Number of Sequences: 2790947
Number of extensions: 109831182
Number of successful extensions: 824907
Number of sequences better than 10.0: 2292
Number of HSP's better than 10.0 without gapping: 1753
Number of HSP's successfully gapped in prelim test: 622
Number of HSP's that attempted gapping in prelim test: 644849
Number of HSP's gapped (non-prelim): 51533
length of query: 2263
length of database: 848,049,833
effective HSP length: 144
effective length of query: 2119
effective length of database: 446,153,465
effective search space: 945399192335
effective search space used: 945399192335
T: 11
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.7 bits)
S2: 83 (36.6 bits)
Medicago: description of AC146789.8


