

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
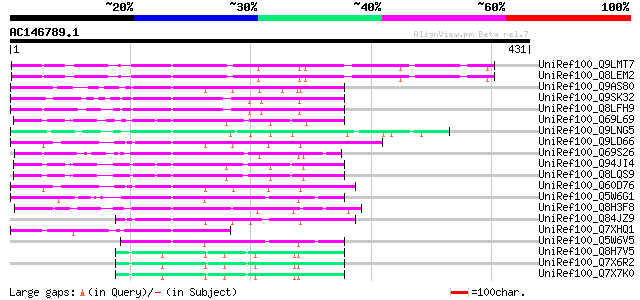
Query= AC146789.1 + phase: 0
(431 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9LMT7 F2H15.15 protein [Arabidopsis thaliana] 100 6e-20
UniRef100_Q8LEM2 Hypothetical protein [Arabidopsis thaliana] 100 6e-20
UniRef100_Q9AS80 P0028E10.12 protein [Oryza sativa] 89 2e-16
UniRef100_Q9SK32 Expressed protein [Arabidopsis thaliana] 87 7e-16
UniRef100_Q8LFH9 Hypothetical protein [Arabidopsis thaliana] 87 7e-16
UniRef100_Q69L69 Calcineurin-like phosphoesterase-like protein [... 86 2e-15
UniRef100_Q9LNG5 F21D18.16 [Arabidopsis thaliana] 83 2e-14
UniRef100_Q9LD66 Similar to maize transposon MuDR mudrA protein ... 83 2e-14
UniRef100_Q69S26 Serine/threonine phosphatase PP7-related-like [... 83 2e-14
UniRef100_Q94JI4 P0638D12.5 protein [Oryza sativa] 82 2e-14
UniRef100_Q8LQS9 P0702H08.15 protein [Oryza sativa] 82 2e-14
UniRef100_Q60D76 Putative polyprotein [Oryza sativa] 79 2e-13
UniRef100_Q5W6G1 Putative polyprotein [Oryza sativa] 72 2e-11
UniRef100_Q8H3F8 Serine/threonine phosphatase PP7-related-like p... 68 4e-10
UniRef100_Q84JZ9 Mutator-like transposase [Oryza sativa] 67 1e-09
UniRef100_Q7XHQ1 Calcineurin-like phosphoesterase-like [Oryza sa... 63 1e-08
UniRef100_Q5W6V5 Putative polyprotein [Oryza sativa] 62 4e-08
UniRef100_Q8H7V5 Putative mutator-like transposase [Oryza sativa] 55 3e-06
UniRef100_Q7X6R2 OSJNBa0001M07.4 protein [Oryza sativa] 55 3e-06
UniRef100_Q7X7K0 OSJNBa0006M15.10 protein [Oryza sativa] 55 3e-06
>UniRef100_Q9LMT7 F2H15.15 protein [Arabidopsis thaliana]
Length = 478
Score = 100 bits (250), Expect = 6e-20
Identities = 104/420 (24%), Positives = 180/420 (42%), Gaps = 61/420 (14%)
Query: 2 PMGEMTVTLDDVACLMHLPIEGRMLAHGKKMPKHEGAALLMTYLGVAQHEAEKICNQEY- 60
P GEMT+TLD+V+ ++ L ++G+ + G K + + + + LG K+ E
Sbjct: 84 PCGEMTITLDEVSLILGLAVDGKPVV-GVKEKDEDPSQVCLRLLG-------KLPKGELS 135
Query: 61 GGYISYPRLRDFYTSYLGRANVLVGTEDPEEVEELERVRTYCVRCYLLYLVGCLLFGDRS 120
G ++ L++ + A + +E+E Y R YL+Y+VG +F
Sbjct: 136 GNRVTAKWLKESFAECPKGATM-------KEIE-------YHTRAYLIYIVGSTIFATTD 181
Query: 121 NKRIELIYLTTMADGYAGMRNYSWGAMTLAYLYGELADACRPGHRALGGSVTLLTTWFLK 180
+I + YL D + Y+WGA LA+LY ++ +A + +GG +TLL W
Sbjct: 182 PSKISVDYLILFED-FEKAGEYAWGAAALAFLYRQIGNASQRSQSIIGGCLTLLQCWSYF 240
Query: 181 HFPGFFSVDLNTDYLEKYPVAARWK---LEKGHGEGITYRSLLDRIQLDDVCWRPYEEHR 237
H ++D ++P+A WK + + YR LD + +V W P+E
Sbjct: 241 H----LNIDRPKRTTRQFPLALLWKGRQQSRSKNDLFKYRKALDDLDPSNVSWCPFEGDL 296
Query: 238 EI--QDFEE--VFWYSGWIMCGVRRVYRHLPERVLRQYGYVQTIPRHPTDVRDLPPPSIV 293
+I Q F++ + S + G + V H P+R ++Q+G Q IP ++PP
Sbjct: 297 DIVPQSFKDNLLLGRSRTKLIGPKVVEWHFPDRCMKQFGLCQVIP------GEVPPRKNE 350
Query: 294 QMF-IDFRTHTLKADARGVQAGEDTRRVADG------YVLWYTRVSHPQILPPIPGDLPR 346
+ D AD ++ E+ G Y+ W+ ++ +P + D
Sbjct: 351 KNHDEDLLEDMNTADEEWMRRRENIVENGGGNGDESEYMQWFNSIT----VPKLHRDTSL 406
Query: 347 PANEEQIIAEQWQRYEARSSPDTYDMVSGAVAYADAQLGQEEVMSMTPQ----QWYEAMT 402
A+ + A Q E S+ D+ + + +E VMSM+ WYEA T
Sbjct: 407 EADIMNVQAAILQFDEVASTLSLEDLHP-----EEREAIEEAVMSMSNALRVGDWYEAST 461
>UniRef100_Q8LEM2 Hypothetical protein [Arabidopsis thaliana]
Length = 478
Score = 100 bits (250), Expect = 6e-20
Identities = 104/420 (24%), Positives = 180/420 (42%), Gaps = 61/420 (14%)
Query: 2 PMGEMTVTLDDVACLMHLPIEGRMLAHGKKMPKHEGAALLMTYLGVAQHEAEKICNQEY- 60
P GEMT+TLD+V+ ++ L ++G+ + G K + + + + LG K+ E
Sbjct: 84 PCGEMTITLDEVSLILGLAVDGKPVV-GVKEKDEDPSQVCLKLLG-------KLPKGELS 135
Query: 61 GGYISYPRLRDFYTSYLGRANVLVGTEDPEEVEELERVRTYCVRCYLLYLVGCLLFGDRS 120
G ++ L++ + A + +E+E Y R YL+Y+VG +F
Sbjct: 136 GNRVTAKWLKESFAECPKGATM-------KEIE-------YHTRAYLIYIVGSTIFATTD 181
Query: 121 NKRIELIYLTTMADGYAGMRNYSWGAMTLAYLYGELADACRPGHRALGGSVTLLTTWFLK 180
+I + YL D + Y+WGA LA+LY ++ +A + +GG +TLL W
Sbjct: 182 PSKISVDYLILFED-FEKAGEYAWGAAALAFLYRQIGNASQRSQSIIGGCLTLLQCWSYF 240
Query: 181 HFPGFFSVDLNTDYLEKYPVAARWK---LEKGHGEGITYRSLLDRIQLDDVCWRPYEEHR 237
H ++D ++P+A WK + + YR LD + +V W P+E
Sbjct: 241 H----LNIDRPKRTTRQFPLALLWKGRQQSRSKNDLFKYRKALDDLDPSNVSWCPFEGDL 296
Query: 238 EI--QDFEE--VFWYSGWIMCGVRRVYRHLPERVLRQYGYVQTIPRHPTDVRDLPPPSIV 293
+I Q F++ + S + G + V H P+R ++Q+G Q IP ++PP
Sbjct: 297 DIVPQSFKDNLLLGRSRTKLIGPKVVEWHFPDRCMKQFGLCQVIP------GEVPPRKNE 350
Query: 294 QMF-IDFRTHTLKADARGVQAGEDTRRVADG------YVLWYTRVSHPQILPPIPGDLPR 346
+ D AD ++ E+ G Y+ W+ ++ +P + D
Sbjct: 351 KNHDEDLLEDMNTADEEWMRRRENIVENGGGNGDESEYMQWFNSIT----VPKLHRDTSL 406
Query: 347 PANEEQIIAEQWQRYEARSSPDTYDMVSGAVAYADAQLGQEEVMSMTPQ----QWYEAMT 402
A+ + A Q E S+ D+ + + +E VMSM+ WYEA T
Sbjct: 407 EADIMNVQAAILQFDEVASTLSLEDLHP-----EEREAIEEAVMSMSNALRVGDWYEAST 461
>UniRef100_Q9AS80 P0028E10.12 protein [Oryza sativa]
Length = 792
Score = 89.4 bits (220), Expect = 2e-16
Identities = 90/302 (29%), Positives = 128/302 (41%), Gaps = 48/302 (15%)
Query: 1 MPMGEMTVTLDDVACLMHLPIEGRMLAHGKKMPKHEGAALLMTYLGVAQHEAEKICNQEY 60
MP GE+T+TL DVA ++ LPI G + P++E L+ YLG A
Sbjct: 278 MPCGEITITLQDVAMILGLPIAGHAVTVNPTEPQNE---LVERYLGKAPPPDR------- 327
Query: 61 GGYISYPRLRDFYTSYLGRANVLVGTEDPEEVEELERVRTYCVRCYLLYLVGCLLFGDRS 120
P LR + RA ED +E E ++ + R Y+L L+ LLF D S
Sbjct: 328 ----PRPGLRVSWV----RAEFNNCPEDADE----ETIKQH-ARAYILSLISGLLFPDAS 374
Query: 121 NKRIELIYLTTMADGYAGMRNYSWGAMTLAYLYGELADACR--PGHRALGGSVTLLTTWF 178
+AD + +YSWG+ TLAYLY + DACR L G + LL W
Sbjct: 375 GDLYTFYPFPLIAD-LENIGSYSWGSATLAYLYRAMCDACRRQSDGSNLTGCLLLLQFWS 433
Query: 179 LKHFP---------GFFSVDLNTDYLEKYPVAARWKL---EKGHGEGITYRSLLDRIQL- 225
+HFP + +V+ D ++ V RW + + Y + L
Sbjct: 434 WEHFPIGRPDLVKLKYPNVEELEDERDRPTVGLRWVVGVCARRAAPARCYEHFTNEFDLL 493
Query: 226 --DDVCWRPYEEHR--EIQ-----DFEEVFWYSGWIMCGVRRVYRHLPERVLRQYGYVQT 276
D V W PY E R E+Q + W + + V ++PERV+RQ+G Q
Sbjct: 494 TDDQVVWCPYREERMKELQLAPICTQDSHLWLTRAPLLYFFMVEIYMPERVMRQFGLHQV 553
Query: 277 IP 278
P
Sbjct: 554 CP 555
>UniRef100_Q9SK32 Expressed protein [Arabidopsis thaliana]
Length = 509
Score = 87.4 bits (215), Expect = 7e-16
Identities = 77/286 (26%), Positives = 128/286 (43%), Gaps = 33/286 (11%)
Query: 1 MPMGEMTVTLDDVACLMHLPIEGRMLAHGKKMPKHEGAALLMTYLGVAQHEAEKICNQEY 60
+P+GEMT+TLD+VA ++ L I+G + K G + M G + N+E
Sbjct: 92 LPLGEMTITLDEVALVLGLEIDGDPIVGSK-----VGDEVAMDMCGRLLGKLPSAANKE- 145
Query: 61 GGYISYPRLRDFYTSYLGRANVLVGTEDPEEVEELERVRTYCVRCYLLYLVGCLLFGDRS 120
++ R++ ++L R +E PE+ + V+ + R YLLYL+G +F
Sbjct: 146 ---VNCSRVK---LNWLKR----TFSECPEDA-SFDVVKCH-TRAYLLYLIGSTIFATTD 193
Query: 121 NKRIELIYLTTMADGYAGMRNYSWGAMTLAYLYGELADACRPGHRALGGSVTLLTTWFLK 180
++ + YL D + Y+WGA LA LY L +A + G +TLL W
Sbjct: 194 GDKVSVKYLPLFED-FDQAGRYAWGAAALACLYRALGNASLKSQSNICGCLTLLQCW--- 249
Query: 181 HFPGFFSVDLNTDYLEK--YPVAARWKLE--KGHGEGITYRSLLDRIQLDDVCWRPYEEH 236
+F +D+ + +P+A WK + + + YR LD + + W PYE
Sbjct: 250 ---SYFHLDIGRPEKSEACFPLALLWKGKGSRSKTDLSEYRRELDDLDPSKITWCPYERF 306
Query: 237 REI----QDFEEVFWYSGWIMCGVRRVYRHLPERVLRQYGYVQTIP 278
+ + + S + ++ H P+R LRQ+G Q IP
Sbjct: 307 ENLIPPHIKAKLILGRSKTTLVCFEKIELHFPDRCLRQFGKRQPIP 352
>UniRef100_Q8LFH9 Hypothetical protein [Arabidopsis thaliana]
Length = 509
Score = 87.4 bits (215), Expect = 7e-16
Identities = 77/286 (26%), Positives = 124/286 (42%), Gaps = 33/286 (11%)
Query: 1 MPMGEMTVTLDDVACLMHLPIEGRMLAHGKKMPKHEGAALLMTYLGVAQHEAEKICNQEY 60
+P+GEMT+TLD+VA ++ L I+G + G K+ + LG A K N
Sbjct: 92 LPLGEMTITLDEVALVLGLEIDGDPIV-GSKVDDEVAMDMCGRLLGKLPSAANKEVN--- 147
Query: 61 GGYISYPRLRDFYTSYLGRANVLVGTEDPEEVEELERVRTYCVRCYLLYLVGCLLFGDRS 120
S +L ++ +E PE+ + V+ + R YLLYL+G +F
Sbjct: 148 ---CSRVKLNWLKRTF---------SECPEDA-SFDVVKCH-TRAYLLYLIGSTIFATTD 193
Query: 121 NKRIELIYLTTMADGYAGMRNYSWGAMTLAYLYGELADACRPGHRALGGSVTLLTTWFLK 180
++ + YL D + Y+WGA LA LY L +A + G +TLL W
Sbjct: 194 GDKVSVKYLPLFED-FDQAGRYAWGAAALACLYRALGNASLKSQSNICGCLTLLQCW--- 249
Query: 181 HFPGFFSVDLNTDYLEK--YPVAARWKLE--KGHGEGITYRSLLDRIQLDDVCWRPYEEH 236
+F +D+ + +P+A WK + + + YR LD + + W PYE
Sbjct: 250 ---SYFHLDIGRPEKSEACFPLALLWKGKGSRSKTDLSEYRRELDDLDPSKITWCPYERF 306
Query: 237 REI----QDFEEVFWYSGWIMCGVRRVYRHLPERVLRQYGYVQTIP 278
+ + + S + ++ H P+R LRQ+G Q IP
Sbjct: 307 ENLIPPHIKAKLILGRSKTTLVCFEKIELHFPDRCLRQFGKRQPIP 352
>UniRef100_Q69L69 Calcineurin-like phosphoesterase-like protein [Oryza sativa]
Length = 766
Score = 85.9 bits (211), Expect = 2e-15
Identities = 92/296 (31%), Positives = 135/296 (45%), Gaps = 48/296 (16%)
Query: 4 GEMTVTLDDVACLMHLPIEGRMLAHGKKMPKHEGAALLMTYLGVAQHEAEKICNQEYGGY 63
GEMTVTL DV+ L+ LPI+GR + H+ A +++ LG H+ + + +
Sbjct: 153 GEMTVTLQDVSMLLALPIDGRPVC---STTDHDYAQMVIDCLG---HD-PRGPSMPGKSF 205
Query: 64 ISYPRLRDFYTSYLGRANVLVGTEDPEEVEELERVRTYCVRCYLLYLVGCLLFGDRSNKR 123
+ Y L+ + + GT+D + +ER VR Y+L L+ +LF D + R
Sbjct: 206 LHYKWLKKHF------YELPEGTDD----QTVER----HVRAYILSLLCGVLFPDGTG-R 250
Query: 124 IELIYLTTMADGYAGMRNYSWGAMTLAYLYGELAD-ACRPGHRALGGSVTLLTTWF---- 178
+ LIYL +AD + + YSWG+ LA+LY L A + +GGS+ LL W
Sbjct: 251 MSLIYLPLIAD-LSLVGTYSWGSAVLAFLYRSLCSVASSHNIKNIGGSLLLLQLWSWERS 309
Query: 179 -----LKHFPGFFSVDLNTDYLEKYPVAARWKLEKGHGEGIT-----YRSLLDRIQLDDV 228
L P D+ D L PV RW + E T YR L+ + D V
Sbjct: 310 HVGRPLVRSPVCPETDIPQDVL---PVGFRWVGARTQSENATRCLKQYRDELNLQRADQV 366
Query: 229 CWRPYEEHREIQDFEEV------FWYSGWIMCGVRRVYRHLPERVLRQYGYVQTIP 278
W PY H E + W + + V +LPERV+RQ+G Q+IP
Sbjct: 367 KWEPY-LHIESASLPLLCTKNADLWLTQAPLINFPIVEMYLPERVMRQFGLRQSIP 421
>UniRef100_Q9LNG5 F21D18.16 [Arabidopsis thaliana]
Length = 1340
Score = 82.8 bits (203), Expect = 2e-14
Identities = 106/403 (26%), Positives = 162/403 (39%), Gaps = 67/403 (16%)
Query: 1 MPMGEMTVTLDDVACLMHLPIEGRMLAHGKKMPKHEGAALLMTYLGVAQHEAEKICNQEY 60
+P GE+TVTL DV L+ L ++G + K + A L LG + +
Sbjct: 101 LPAGEITVTLQDVNILLGLRVDGPAVTGSTK---YNWADLCEDLLGHRPGPKDL-----H 152
Query: 61 GGYISYPRLRDFYTSYLGRANVLVGTEDPEEVEELERVRTYC-VRCYLLYLVGCLLFGDR 119
G ++S LR+ + + DP+EV C R ++L L+ L+GD+
Sbjct: 153 GSHVSLAWLRENFRNL---------PADPDEVT------LKCHTRAFVLALMSGFLYGDK 197
Query: 120 SNKRIELIYLTTMADGYAGMRNYSWGAMTLAYLYGELADACRPGHRALGGSVTLLTTWFL 179
S + L +L + D + + SWG+ TLA LY EL A + + G + LL W
Sbjct: 198 SKHDVALTFLPLLRD-FDEVAKLSWGSATLALLYRELCRASKRTVSTICGPLVLLQLWAW 256
Query: 180 KHF----PGFFSVDLNTDYLEKY------PVAARWKLEKGHGEGIT-----YRSLLDRIQ 224
+ PG D+ Y++ P+ RW+ H E YR D+ +
Sbjct: 257 ERLHVGRPGRLK-DVGASYMDGIDGPLPDPLGCRWRASLSHKENPRGGLDFYRDQFDQQK 315
Query: 225 LDDVCWRPY-----EEHREIQDFEEVFWYSGWIMCGVRRVYRHLPERVLRQYGYVQTIPR 279
+ V W+PY + I E W + + V H P+RVLRQ+G QTIP
Sbjct: 316 DEQVIWQPYTPDLLAKIPLICVSGENIWRTVAPLICFDVVEWHRPDRVLRQFGLHQTIPA 375
Query: 280 --------HPTDVRDLPPPSIVQMFIDFRTHTLKADAR--GVQAGE---DTRRVADGYVL 326
H D R S H +AR V +GE D Y+
Sbjct: 376 PCDNEKALHAIDKRG---KSEYDWSARHSRHIGLWEARVSSVVSGEPECSPMDYNDPYME 432
Query: 327 WYTRVSHPQILPPI----PGDLPRPANEEQIIAEQWQRYEARS 365
WY R++ +I+ P+ PG Q++ ++ ARS
Sbjct: 433 WYRRITR-RIISPMNERRPGQFLPTGFAFQVLVQRVAAIHARS 474
>UniRef100_Q9LD66 Similar to maize transposon MuDR mudrA protein [Oryza sativa]
Length = 1230
Score = 82.8 bits (203), Expect = 2e-14
Identities = 92/335 (27%), Positives = 137/335 (40%), Gaps = 47/335 (14%)
Query: 1 MPMGEMTVTLDDVACLMHLPIEGRML----AHGKKMPKHEGAALLMTYLGVAQHEAEKIC 56
+P GE+TVTL+DVA ++ LPI G+ + A G + E YLG+ A
Sbjct: 592 LPCGELTVTLEDVAMILGLPIRGQAVTGDTASGNWRERVE------EYLGLEPPVAPDGQ 645
Query: 57 NQEYGGYISYPRLRDFYTSYLGRANVLVGTEDPEEVEELERVRTYCVRCYLLYLVGCLLF 116
Q + LR + + P E +E V+ YC R Y+LY+ G +LF
Sbjct: 646 RQTKTSGVPLSWLRANFG------------QCPAEADEAT-VQRYC-RAYVLYIFGSILF 691
Query: 117 GDRSNKRIELIYLTTMADGYAGMRNYSWGAMTLAYLYGELADACR--PGHRALGGSVTLL 174
D ++L +AD + +SWG+ LA+LY +L DACR G L G V LL
Sbjct: 692 PDSGGDMASWMWLPLLAD-WDEAGTFSWGSAALAWLYRQLCDACRRQGGDANLAGCVWLL 750
Query: 175 TTWFLKHFP------GFFSVDLNTDYLEKYPVAARWKLEKGHGEG------ITYRSLLDR 222
W P + D + VA W+ G Y + +D
Sbjct: 751 QVWMWMRLPVGRPMWRTHQAWPHQDADRRPTVAHLWESVPSPVVGRRNLAYYHYTNEMDY 810
Query: 223 IQLDDVCWRPYEEHREIQ-------DFEEVFWYSGWIMCGVRRVYRHLPERVLRQYGYVQ 275
+Q + V W PY+ ++ E+ + V + RV+RQ+G +Q
Sbjct: 811 LQPEHVVWMPYQAQEVLELELNPMCHIEDALKTLRCPLICFYAVEFQMCHRVMRQFGRLQ 870
Query: 276 TIPRHPTDVRDLPPP-SIVQMFIDFRTHTLKADAR 309
TIP + DL +I +F HT + D R
Sbjct: 871 TIPHRFSTSIDLHNHFNIYYIFELDILHTCRVDRR 905
>UniRef100_Q69S26 Serine/threonine phosphatase PP7-related-like [Oryza sativa]
Length = 619
Score = 82.8 bits (203), Expect = 2e-14
Identities = 78/283 (27%), Positives = 129/283 (45%), Gaps = 37/283 (13%)
Query: 5 EMTVTLDDVACLMHLPIEGRMLAHGKKMPKHEGAALLMTYLGVAQHEAEKICNQEYGGYI 64
EM +L DV+ ++ +P+ G ++ + + + + +LG + E++C G I
Sbjct: 96 EMAPSLRDVSYILGIPVTGHVVT-AEPIGDEAVRRMCLHFLGESPGNGEQLC-----GLI 149
Query: 65 SYPRLRDFYTSYLGRANVLVGTEDPEEVEELERVRTYCVRCYLLYLVGCLLFGDRSNKRI 124
RL Y + E+P + E+ Y R YLLYLVG LF D +
Sbjct: 150 ---RLTWLYRKFHQLP------ENPT-INEI----AYSTRAYLLYLVGSTLFPDTMRGFV 195
Query: 125 ELIYLTTMADGYAGMRNYSWGAMTLAYLYGELADACRPGHRA-LGGSVTLLTTWFLKHFP 183
YL +AD + +R Y+WG+ LA+LY L+ A P GS TLL W ++ P
Sbjct: 196 SPRYLPLLAD-FRKIREYAWGSAALAHLYRGLSVAVTPNATTQFLGSATLLMAWIYEYLP 254
Query: 184 GFFSVDLNTDYLEKYPVAARWKL---EKGHGEGI-TYRSLLDRIQLDDVCWRPYEEHRE- 238
N + L P A RW +G + + +R + D +Q DV W PY++
Sbjct: 255 LTQPQQKNQNTL--LPRACRWNFGGATRGQRKKVMEWRKVFDELQFSDVNWNPYKDMNPA 312
Query: 239 -IQDF----EEVFWYSGWIMC-GVRRVYRHLPERVLRQYGYVQ 275
I ++ + + + W++ ++ VY +P+R RQ+G Q
Sbjct: 313 IIPEYCIAADNICYSRTWLISFNIKEVY--VPDRFSRQFGREQ 353
>UniRef100_Q94JI4 P0638D12.5 protein [Oryza sativa]
Length = 761
Score = 82.4 bits (202), Expect = 2e-14
Identities = 90/296 (30%), Positives = 130/296 (43%), Gaps = 48/296 (16%)
Query: 4 GEMTVTLDDVACLMHLPIEGRMLAHGKKMPKHEGAALLMTYLGVAQHEAEKICNQEYGGY 63
GEM VTL DVA L+ LPI+GR + H+ A +++ LG H+ + + +
Sbjct: 170 GEMAVTLQDVAMLLALPIDGRPVC---STTDHDYAQMVIDCLG---HD-PRGPSMPGKSF 222
Query: 64 ISYPRLRDFYTSYLGRANVLVGTEDPEEVEELERVRTYCVRCYLLYLVGCLLFGDRSNKR 123
+ Y L+ + E PE ++ R VR Y+L L+ +LF D + R
Sbjct: 223 LHYKWLKKHF------------YELPEGADDQTVERH--VRAYILSLLCGVLFPDGTG-R 267
Query: 124 IELIYLTTMADGYAGMRNYSWGAMTLAYLYGELAD-ACRPGHRALGGSVTLLTTWF---- 178
+ LIYL +AD + + YSWG+ LA+LY L A + +GGS+ LL W
Sbjct: 268 MSLIYLPLIAD-LSRVGTYSWGSAVLAFLYRSLCSVASSHNIKNIGGSLLLLQLWSWERS 326
Query: 179 -----LKHFPGFFSVDLNTDYLEKYPVAARWKLEKGHGEGIT-----YRSLLDRIQLDDV 228
L P D+ D PV RW + E T YR L+ + D V
Sbjct: 327 HVGRPLVRSPLCPETDIPQDL---PPVGFRWVGARTQSENATRCLKQYRDELNLQRADQV 383
Query: 229 CWRPYEEHREIQDF------EEVFWYSGWIMCGVRRVYRHLPERVLRQYGYVQTIP 278
W PY H E + W + + V +LPERV+RQ+G Q+IP
Sbjct: 384 KWEPY-LHIESSSLPLLCTKDADLWLTQAPLINFPIVEMYLPERVMRQFGLRQSIP 438
>UniRef100_Q8LQS9 P0702H08.15 protein [Oryza sativa]
Length = 718
Score = 82.4 bits (202), Expect = 2e-14
Identities = 90/296 (30%), Positives = 130/296 (43%), Gaps = 48/296 (16%)
Query: 4 GEMTVTLDDVACLMHLPIEGRMLAHGKKMPKHEGAALLMTYLGVAQHEAEKICNQEYGGY 63
GEM VTL DVA L+ LPI+GR + H+ A +++ LG H+ + + +
Sbjct: 184 GEMAVTLQDVAMLLALPIDGRPVC---STTDHDYAQMVIDCLG---HD-PRGPSMPGKSF 236
Query: 64 ISYPRLRDFYTSYLGRANVLVGTEDPEEVEELERVRTYCVRCYLLYLVGCLLFGDRSNKR 123
+ Y L+ + E PE ++ R VR Y+L L+ +LF D + R
Sbjct: 237 LHYKWLKKHF------------YELPEGADDQTVERH--VRAYILSLLCGVLFPDGTG-R 281
Query: 124 IELIYLTTMADGYAGMRNYSWGAMTLAYLYGELAD-ACRPGHRALGGSVTLLTTWF---- 178
+ LIYL +AD + + YSWG+ LA+LY L A + +GGS+ LL W
Sbjct: 282 MSLIYLPLIAD-LSRVGTYSWGSAVLAFLYRSLCSVASSHNIKNIGGSLLLLQLWSWERS 340
Query: 179 -----LKHFPGFFSVDLNTDYLEKYPVAARWKLEKGHGEGIT-----YRSLLDRIQLDDV 228
L P D+ D PV RW + E T YR L+ + D V
Sbjct: 341 HVGRPLVRSPLCPETDIPQDV---PPVGFRWVGARTQSENATRCLKQYRDELNLQRADQV 397
Query: 229 CWRPYEEHREIQDF------EEVFWYSGWIMCGVRRVYRHLPERVLRQYGYVQTIP 278
W PY H E + W + + V +LPERV+RQ+G Q+IP
Sbjct: 398 KWEPY-LHIESSSLPLLCTKDADLWLTQAPLINFPIVEMYLPERVMRQFGLRQSIP 452
>UniRef100_Q60D76 Putative polyprotein [Oryza sativa]
Length = 1754
Score = 79.3 bits (194), Expect = 2e-13
Identities = 86/312 (27%), Positives = 131/312 (41%), Gaps = 46/312 (14%)
Query: 1 MPMGEMTVTLDDVACLMHLPIEGRML----AHGKKMPKHEGAALLMTYLGVAQHEAEKIC 56
+P GE+TVTL+DVA ++ LPI G+ + A G + E YLG+ A
Sbjct: 974 LPCGELTVTLEDVAMILGLPIRGQAVTGDTASGNWRERVE------EYLGLEPPVAPDGQ 1027
Query: 57 NQEYGGYISYPRLRDFYTSYLGRANVLVGTEDPEEVEELERVRTYCVRCYLLYLVGCLLF 116
Q + LR + + P E +E V+ YC R Y+LY+ G +LF
Sbjct: 1028 RQTKTSGVPLSWLRANFG------------QCPAEADEAT-VQRYC-RAYVLYIFGSILF 1073
Query: 117 GDRSNKRIELIYLTTMADGYAGMRNYSWGAMTLAYLYGELADACR--PGHRALGGSVTLL 174
D ++L +AD + +SWG+ LA+LY +L DACR G L G V LL
Sbjct: 1074 PDSGGDMASWMWLPLLAD-WDEAGTFSWGSAALAWLYRQLCDACRRQGGDANLAGCVWLL 1132
Query: 175 TTW-FLKHFPG-----FFSVDLNTDYLEKYPVAARWKLEKGHGEG------ITYRSLLDR 222
W +++ G + D + VA W+ G Y + +D
Sbjct: 1133 QVWMWMRLLVGRPMWRTHQAWPHQDADRRPTVAHLWESVPSPVVGRRNLAYCHYTNEMDY 1192
Query: 223 IQLDDVCWRPYEEHREIQ-------DFEEVFWYSGWIMCGVRRVYRHLPERVLRQYGYVQ 275
+Q + V W PY+ ++ E+ + V + RV+RQ+G +Q
Sbjct: 1193 LQPEHVVWMPYQAQEVLELELNPMCHIEDALKTLRCPLICFYAVEFQMCHRVMRQFGRLQ 1252
Query: 276 TIPRHPTDVRDL 287
TIP + DL
Sbjct: 1253 TIPHRFSTSIDL 1264
>UniRef100_Q5W6G1 Putative polyprotein [Oryza sativa]
Length = 1567
Score = 72.4 bits (176), Expect = 2e-11
Identities = 73/291 (25%), Positives = 123/291 (42%), Gaps = 37/291 (12%)
Query: 1 MPMGEMTVTLDDVACLMHLPIEGRMLAHGKKMPKHEGAA--LLMTYLGVAQHEAEKICNQ 58
+P GEMT+TL DVA ++ LP+ G + + P L L + Q + K +
Sbjct: 1042 LPSGEMTITLQDVAMILALPLRGHAVTGRTETPGWHAQVQQLFGIPLNIEQGQGGK--KK 1099
Query: 59 EYGGYISYPRLRDFYTSYLGRANVLVGTEDPEEVEELERVRTYC-VRCYLLYLVGCLLFG 117
+ G +S+ ++F S+L ++ E R C R Y+L+L+G +LF
Sbjct: 1100 QNGIPLSWLS-QNF--SHLD--------------DDAEPWRAECYARAYILHLLGGVLFP 1142
Query: 118 DRSNKRIELIYLTTMADGYAGMRNYSWGAMTLAYLYGELADAC--RPGHRALGGSVTLLT 175
D I++ +A+ + +SWG+ LA+ Y +L +AC + + G V L+
Sbjct: 1143 DAGGDIASAIWIPLVAN-LGDLGRFSWGSAVLAWTYRQLCEACCRQAPSSNMSGCVLLIQ 1201
Query: 176 TWFLKHFPGFFSVDLNTDYLEKYPVAARWKLEKGHGEGITYRSLLDRIQLDDVCWRPYEE 235
W P ++ YL + A + + Y + +D +Q + W PY
Sbjct: 1202 MWMWLRLPNEPDMEKTVAYLFESTATAHAHRDVAYKH---YVNEMDYLQPQHIEWLPYHT 1258
Query: 236 HRE--------IQDFEEVFWYSGWIMCGVRRVYRHLPERVLRQYGYVQTIP 278
+ + F Y ++C V HLP RV+RQ+G Q P
Sbjct: 1259 NEASSLTLNSMCNRDSDYFMYQCPLIC-FWAVEYHLPHRVMRQFGKKQDWP 1308
>UniRef100_Q8H3F8 Serine/threonine phosphatase PP7-related-like protein [Oryza
sativa]
Length = 589
Score = 68.2 bits (165), Expect = 4e-10
Identities = 73/303 (24%), Positives = 124/303 (40%), Gaps = 41/303 (13%)
Query: 5 EMTVTLDDVACLMHLPIEGRMLAHGKKMPKHEGAALLMTYLGVAQHEAEKICNQEYGGYI 64
EM +L D A ++ +P+ GR++ G + K L YLG C G ++
Sbjct: 83 EMAPSLRDAAYILGIPVTGRVVTTGAVLNKSV-EDLCFQYLGQVPD-----CRDCRGSHV 136
Query: 65 SYPRLRDFYTSYLGRANVLVGTEDPEEVEELERVRTYCVRCYLLYLVGCLLFGDRSNKRI 124
L+ ++ E P + + Y R YLL+L+G L +R +
Sbjct: 137 KLSWLQSKFSRI---------PERPTNDQTM-----YGTRAYLLFLIGSALLPERDRGYV 182
Query: 125 ELIYLTTMADGYAGMRNYSWGAMTLAYLYGELADA-CRPGHRALGGSVTLLTTWFLKHFP 183
YL ++D + ++ Y+WGA LA+LY L+ A + L GS LL W ++ P
Sbjct: 183 SPKYLPLLSD-FDKVQEYAWGAAALAHLYKALSIAVAHSARKRLFGSAALLMGWIYEYIP 241
Query: 184 GFFSVDLNTDYLEKYPVAARW---KLEKGHGEGITYRSLLDRIQLDDVCWRPYE------ 234
D+ +P +W + + R +Q+ DV W PY+
Sbjct: 242 A-LRPDMYDPPEHIFPRVLKWTGSTISQPAKNVSDIRKAFGLLQVSDVNWEPYKGVDPAS 300
Query: 235 --EHREIQDFEEVFWYSGWIMC-GVRRVYRHLPERVLRQYGYVQTIPRH--PTDVRDLPP 289
+H D + + W++ ++ +Y P+R RQ+G Q P + P R L
Sbjct: 301 IPKHCAAPD--NLCFSRTWLVSFNLKEIY--APDRFARQFGQEQHRPLNDVPAFQRQLWN 356
Query: 290 PSI 292
P++
Sbjct: 357 PAV 359
>UniRef100_Q84JZ9 Mutator-like transposase [Oryza sativa]
Length = 1381
Score = 67.0 bits (162), Expect = 1e-09
Identities = 63/225 (28%), Positives = 93/225 (41%), Gaps = 34/225 (15%)
Query: 89 PEEVEELERVRTYCVRCYLLYLVGCLLFGDRSNKRIELIYLTTMADGYAGMRNYSWGAMT 148
P E +E V+ YC R Y+LY+ G +LF D ++L +AD + +SWG+
Sbjct: 804 PAEADEAT-VQRYC-RAYVLYIFGSILFPDSGGDMASWMWLPLLAD-WDEAGTFSWGSAA 860
Query: 149 LAYLYGELADACR--PGHRALGGSVTLLTTWFLKHFP----------GFFSVDLNTDYLE 196
LA+LY +L DACR G L G V LL W P + D +
Sbjct: 861 LAWLYRQLCDACRRQGGDANLAGCVWLLQVWMWMRLPVGRPMWRTHQAWPHQDADRCPTV 920
Query: 197 KY-------PVAARWKLEKGHGEGITYRSLLDRIQLDDVCWRPYEEHREIQ-------DF 242
+ PV R L H Y + +D +Q + V W PY+ ++
Sbjct: 921 AHLWESVPSPVVGRRNLAYYH-----YTNEMDYLQPEHVVWMPYQAQEVLELELNPMCHI 975
Query: 243 EEVFWYSGWIMCGVRRVYRHLPERVLRQYGYVQTIPRHPTDVRDL 287
E+ + V + RV+RQ+G +QTIP + DL
Sbjct: 976 EDALKTLRCPLICFYAVEFQMCHRVMRQFGRLQTIPHRFSTSIDL 1020
>UniRef100_Q7XHQ1 Calcineurin-like phosphoesterase-like [Oryza sativa]
Length = 327
Score = 63.2 bits (152), Expect = 1e-08
Identities = 50/187 (26%), Positives = 81/187 (42%), Gaps = 15/187 (8%)
Query: 1 MPMGEMTVTLDDVACLMHLPIEGRMLAHGKKMPKHEGAALLMTYLGVAQHE----AEKIC 56
+P+GEMT+TL DV+CL LPI GR + ++ LG+ E +K
Sbjct: 87 LPVGEMTITLQDVSCLWGLPIHGRPIT---GQADGSWVDMIERLLGIPMEEQHMKQKKRK 143
Query: 57 NQEYGGYISYPRLRDFYTSYLGRANVLVGTEDPEEVEELERVRTYCVRCYLLYLVGCLLF 116
++ +SY R + R V+ P+ ER + R +L ++G ++F
Sbjct: 144 KEDDMTMVSYSRYSISLSKLRDRFRVM-----PKNA--TEREINWYTRALVLDIIGSMVF 196
Query: 117 GDRSNKRIELIYLTTMADGYAGMRNYSWGAMTLAYLYGELADACRPGHRALGGSVTLLTT 176
D S + +YL M + + Y+WGA L+ LY +L+ A + + LL
Sbjct: 197 TDTSGDGVPAMYLQFMMN-LSEQTEYNWGAAALSMLYRQLSIASEKERAEISRPLLLLQL 255
Query: 177 WFLKHFP 183
W P
Sbjct: 256 WSWSRLP 262
>UniRef100_Q5W6V5 Putative polyprotein [Oryza sativa]
Length = 1525
Score = 61.6 bits (148), Expect = 4e-08
Identities = 50/197 (25%), Positives = 84/197 (42%), Gaps = 16/197 (8%)
Query: 93 EELERVRTYC-VRCYLLYLVGCLLFGDRSNKRIELIYLTTMADGYAGMRNYSWGAMTLAY 151
++ E R C R Y+L+L+G +LF D I++ +A+ + +SWG+ LA+
Sbjct: 1075 DDAEPWRAECYARAYILHLLGGVLFPDAGGDIASAIWIPLVAN-LGDLGRFSWGSAVLAW 1133
Query: 152 LYGELADAC--RPGHRALGGSVTLLTTWFLKHFPGFFSVDLNTDYLEKYPVAARWKLEKG 209
Y +L +AC + + G V L+ W P ++ YL + A +
Sbjct: 1134 TYRQLCEACCRQAPSSNMSGCVLLIQMWMWLRLPNEPDMEKTVAYLFESTATAHAHRDVA 1193
Query: 210 HGEGITYRSLLDRIQLDDVCWRPYEEHRE--------IQDFEEVFWYSGWIMCGVRRVYR 261
+ Y + +D +Q + W PY + + F Y ++C V
Sbjct: 1194 YKH---YVNEMDYLQPQHIEWLPYHTNEASSLTLNSMCNRDSDYFMYQCPLIC-FWAVEY 1249
Query: 262 HLPERVLRQYGYVQTIP 278
HLP RV+RQ+G Q P
Sbjct: 1250 HLPHRVMRQFGKKQDWP 1266
>UniRef100_Q8H7V5 Putative mutator-like transposase [Oryza sativa]
Length = 1596
Score = 55.5 bits (132), Expect = 3e-06
Identities = 58/225 (25%), Positives = 91/225 (39%), Gaps = 39/225 (17%)
Query: 89 PEEVEELERVRTYCVRCYLLYLVGCLLFGDRSNKRIE---LIYLTTMADGYAG-MRNYSW 144
PEE +E R C+ YLL+L G ++F ++ + Y ++AD G + +SW
Sbjct: 1054 PEEADEYSFSR--CLEAYLLWLFGWVMFCGSHGHSVDRGLVHYARSIADAAVGEVPQWSW 1111
Query: 145 GAMTLAYLYGELADACR---PGHRALGGSVTLLTTW----------FLKHFPGFFSVDLN 191
G+ LA Y L +AC PG GG ++ W ++H P S+ N
Sbjct: 1112 GSALLAAQYRGLCEACTRTDPG-AIFGGCPLFISLWAAERIAIGRPLVEHHPYDASLYEN 1170
Query: 192 TDYLEKYPVAA--------RWKLEKGHGEGITYRSLLDRIQLDDVCWRPYEE---HRE-- 238
++ YP RW + + DR+ DV W PY H
Sbjct: 1171 RPAVD-YPTMGTLWCRRQRRWAHVQVRRSYAEFVMEFDRLLPTDVVWEPYSALLTHSRAP 1229
Query: 239 -----IQDFEEVFWYSGWIMCGVRRVYRHLPERVLRQYGYVQTIP 278
+ ++ +W + M V H P RV+RQ+G+ Q+ P
Sbjct: 1230 LGLSTLCTRDQAYWMTTVPMVFDICVEPHAPYRVMRQFGFRQSFP 1274
>UniRef100_Q7X6R2 OSJNBa0001M07.4 protein [Oryza sativa]
Length = 1031
Score = 55.5 bits (132), Expect = 3e-06
Identities = 58/225 (25%), Positives = 91/225 (39%), Gaps = 39/225 (17%)
Query: 89 PEEVEELERVRTYCVRCYLLYLVGCLLFGDRSNKRIE---LIYLTTMADGYAG-MRNYSW 144
PEE +E R C+ YLL+L G ++F ++ + Y ++AD G + +SW
Sbjct: 524 PEEADEYSFSR--CLEAYLLWLFGWVMFCGSHGHSVDRGLVHYARSIADAAVGEVPQWSW 581
Query: 145 GAMTLAYLYGELADACR---PGHRALGGSVTLLTTW----------FLKHFPGFFSVDLN 191
G+ LA Y L +AC PG GG ++ W ++H P S+ N
Sbjct: 582 GSALLAAQYRGLCEACTRTDPG-AIFGGCPLFISLWAAERIAIGRPLVEHHPYDASLYEN 640
Query: 192 TDYLEKYPVAA--------RWKLEKGHGEGITYRSLLDRIQLDDVCWRPYEE---HRE-- 238
++ YP RW + + DR+ DV W PY H
Sbjct: 641 RPAVD-YPTMGTLWCRRQRRWAHVQVRRSYAEFVMEFDRLLPTDVVWEPYSALLTHSRAP 699
Query: 239 -----IQDFEEVFWYSGWIMCGVRRVYRHLPERVLRQYGYVQTIP 278
+ ++ +W + M V H P RV+RQ+G+ Q+ P
Sbjct: 700 LGLSTLCTRDQAYWMTTVPMVFDICVEPHAPYRVMRQFGFRQSFP 744
>UniRef100_Q7X7K0 OSJNBa0006M15.10 protein [Oryza sativa]
Length = 1620
Score = 55.5 bits (132), Expect = 3e-06
Identities = 58/225 (25%), Positives = 91/225 (39%), Gaps = 39/225 (17%)
Query: 89 PEEVEELERVRTYCVRCYLLYLVGCLLFGDRSNKRIE---LIYLTTMADGYAG-MRNYSW 144
PEE +E R C+ YLL+L G ++F ++ + Y ++AD G + +SW
Sbjct: 1054 PEEADEYSFSR--CLEAYLLWLFGWVMFCGSHGHSVDRGLVHYARSIADAAVGEVPQWSW 1111
Query: 145 GAMTLAYLYGELADACR---PGHRALGGSVTLLTTW----------FLKHFPGFFSVDLN 191
G+ LA Y L +AC PG GG ++ W ++H P S+ N
Sbjct: 1112 GSALLAAQYRGLCEACTRTDPG-AIFGGCPLFISLWAAERIAIGRPLVEHHPYDASLYEN 1170
Query: 192 TDYLEKYPVAA--------RWKLEKGHGEGITYRSLLDRIQLDDVCWRPYEE---HRE-- 238
++ YP RW + + DR+ DV W PY H
Sbjct: 1171 RPAVD-YPTMGTLWCRRQRRWAHVQVRRSYAEFVMEFDRLLPTDVVWEPYSALLTHSRAP 1229
Query: 239 -----IQDFEEVFWYSGWIMCGVRRVYRHLPERVLRQYGYVQTIP 278
+ ++ +W + M V H P RV+RQ+G+ Q+ P
Sbjct: 1230 LGLSTLCTRDQAYWMTTVPMVFDICVEPHAPYRVMRQFGFRQSFP 1274
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.323 0.139 0.442
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 772,869,390
Number of Sequences: 2790947
Number of extensions: 33673509
Number of successful extensions: 66682
Number of sequences better than 10.0: 119
Number of HSP's better than 10.0 without gapping: 24
Number of HSP's successfully gapped in prelim test: 95
Number of HSP's that attempted gapping in prelim test: 66426
Number of HSP's gapped (non-prelim): 248
length of query: 431
length of database: 848,049,833
effective HSP length: 130
effective length of query: 301
effective length of database: 485,226,723
effective search space: 146053243623
effective search space used: 146053243623
T: 11
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 76 (33.9 bits)
Medicago: description of AC146789.1


