

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146784.10 + phase: 0
(168 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
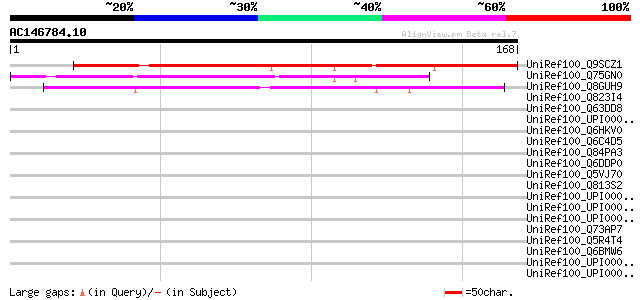
Sequences producing significant alignments: (bits) Value
UniRef100_Q9SCZ1 Hypothetical protein F26O13.220 [Arabidopsis th... 141 7e-33
UniRef100_Q75GN0 Hypothetical protein OSJNBa0018K15.9 [Oryza sat... 88 7e-17
UniRef100_Q8GUH9 Hypothetical protein At1g64385 [Arabidopsis tha... 65 5e-10
UniRef100_Q823I4 Hypothetical protein [Chlamydophila caviae] 39 0.039
UniRef100_Q63DD8 Hypothetical protein [Bacillus cereus] 36 0.43
UniRef100_UPI0000437598 UPI0000437598 UniRef100 entry 35 0.73
UniRef100_Q6HKV0 Hypothetical protein [Bacillus thuringiensis] 35 0.73
UniRef100_Q6C4D5 Similar to tr|Q9HEA4 Neurospora crassa [Yarrowi... 35 0.73
UniRef100_Q84PA3 ADP ribosylation GTPase-like protein [Oryza sat... 35 0.96
UniRef100_Q6DDP0 Snag1-prov protein [Xenopus laevis] 34 1.3
UniRef100_Q5VJ70 Cell adhesion molecule JCAM [Rattus norvegicus] 34 1.3
UniRef100_Q813S2 Hypothetical Membrane Spanning Protein [Bacillu... 34 1.3
UniRef100_UPI00002D1719 UPI00002D1719 UniRef100 entry 33 2.1
UniRef100_UPI0000361174 UPI0000361174 UniRef100 entry 33 2.8
UniRef100_UPI0000361173 UPI0000361173 UniRef100 entry 33 2.8
UniRef100_Q73AP7 Hypothetical protein [Bacillus cereus] 33 2.8
UniRef100_Q5R4T4 Hypothetical protein DKFZp459N1416 [Pongo pygma... 33 2.8
UniRef100_Q6BMW6 Similarity [Debaryomyces hansenii] 33 2.8
UniRef100_UPI00003AFDCB UPI00003AFDCB UniRef100 entry 33 3.6
UniRef100_UPI00003AFDCA UPI00003AFDCA UniRef100 entry 33 3.6
>UniRef100_Q9SCZ1 Hypothetical protein F26O13.220 [Arabidopsis thaliana]
Length = 390
Score = 141 bits (355), Expect = 7e-33
Identities = 70/153 (45%), Positives = 101/153 (65%), Gaps = 10/153 (6%)
Query: 22 IKSESTELTLDAGKGDCVLHVTVVTPVPEASFFLRLPSFDKILTPVNGAYFLIFTVIVFA 81
I ++ ++ LD GKG C LH+ P E++ PS++K++TP+NGAYFLI +VI+F
Sbjct: 242 ISGDTNKIILDTGKGQCALHMY---PSEESTLPFHFPSYEKLVTPINGAYFLIVSVIIFG 298
Query: 82 VTWA-CCCIFKKKPRDEIPYQELEMA----LPESASATVVESAEGWDQGWDDDWDDNVAV 136
WA C C ++ +PY+ELE++ L + VE+A+ WD+GWDDDWD+N AV
Sbjct: 299 GIWAFCLCRKNRRAGSGVPYRELELSGGPGLENESGVHDVETAD-WDEGWDDDWDENNAV 357
Query: 137 KSP-VVRHAGSISANGLTSRSSNKDGWEDNWDD 168
KSP + SISANGLT+R+ N+DGW+ +WDD
Sbjct: 358 KSPGSAAKSVSISANGLTARAPNRDGWDHDWDD 390
>UniRef100_Q75GN0 Hypothetical protein OSJNBa0018K15.9 [Oryza sativa]
Length = 515
Score = 88.2 bits (217), Expect = 7e-17
Identities = 53/142 (37%), Positives = 71/142 (49%), Gaps = 8/142 (5%)
Query: 1 MSDLQLAYCFYNIVFQVTIKQIKSESTELTLDAGKGDCVLHVTVVTPVPEASFFLRLPSF 60
M LQLA F QV I E+T+ +GKG C LH T T F + ++
Sbjct: 307 MLPLQLAKGFSR---QVNITYSNPNGVEITVKSGKGQCSLH-TKQTVFDWQQQFQQFAAY 362
Query: 61 DKILTPVNGAYFLIFTVIVFAVTWACCCIFKKKPRDEIPYQELEMA--LPESASA-TVVE 117
P+ GA FL+FTV++ V ACC F ++ +PYQ+LEM P S+
Sbjct: 363 ATRANPIYGASFLVFTVVLVGVVCACCK-FARRRASGVPYQQLEMGDQAPNSSGVENTTS 421
Query: 118 SAEGWDQGWDDDWDDNVAVKSP 139
+ +GW+ GWDDDWDD A P
Sbjct: 422 TVDGWEDGWDDDWDDEEAAAKP 443
>UniRef100_Q8GUH9 Hypothetical protein At1g64385 [Arabidopsis thaliana]
Length = 351
Score = 65.5 bits (158), Expect = 5e-10
Identities = 50/173 (28%), Positives = 77/173 (43%), Gaps = 23/173 (13%)
Query: 12 NIVFQVTIKQIKSESTELTLDAGKGDCVL---------HVTVVTPVPEASFFLRLPSFDK 62
+I +V+IK+ S + + L + KG C L H T S L +
Sbjct: 182 DIKVKVSIKKGGSNDSAIVLASSKGRCRLELKDLAAAAHETESDDTVSVSRPSILNISSR 241
Query: 63 ILTPVNGAYFLIFTVIVFAVTWACCCIFKKKPRDEIPYQELEMALPESASATVVESAE-- 120
L + FL+ ++++ V ++K K R YQ L+M LP S A V +S +
Sbjct: 242 TLIVIIMISFLVLSLVIIPVI---IHVYKNKSRGNNKYQRLDMELPVSNPALVTKSDQES 298
Query: 121 ---GWDQGWDDDWD------DNVAVKSPVVRHAGSISANGLTSRSSNKDGWED 164
GW+ W DDWD D +PV+ S+S+ GL R +K+GW+D
Sbjct: 299 GDDGWNNNWGDDWDDENGGGDEEQPNTPVLPLTPSLSSRGLAPRRLSKEGWKD 351
>UniRef100_Q823I4 Hypothetical protein [Chlamydophila caviae]
Length = 379
Score = 39.3 bits (90), Expect = 0.039
Identities = 19/46 (41%), Positives = 26/46 (56%), Gaps = 1/46 (2%)
Query: 74 IFTVIVFAVTWACCCIFKKKP-RDEIPYQELEMALPESASATVVES 118
+ TVI + ACC + KKP RD PY LE +PE+ T+ E+
Sbjct: 65 VATVIFSIIAVACCVLLNKKPQRDATPYPPLEEGMPENPRLTIPET 110
>UniRef100_Q63DD8 Hypothetical protein [Bacillus cereus]
Length = 113
Score = 35.8 bits (81), Expect = 0.43
Identities = 31/93 (33%), Positives = 45/93 (48%), Gaps = 13/93 (13%)
Query: 20 KQIKSESTELTLDAGKGDCVLHVTVVTPVPEASFFLRLP---SFDKIL-TPVNG--AYFL 73
K ++SE +L L+ K V + SFF+ LP S+ K+L TP G +
Sbjct: 19 KVVQSEEFQLLLNKKK-----KFIVPMSIFFLSFFIALPILTSYSKVLNTPAFGDVTWAW 73
Query: 74 IFTVIVFAVTWACCCIFKKKPR--DEIPYQELE 104
+F F +TWA C I+ KK DEI + L+
Sbjct: 74 VFAFAQFIMTWALCMIYSKKAESFDEISQKILQ 106
>UniRef100_UPI0000437598 UPI0000437598 UniRef100 entry
Length = 172
Score = 35.0 bits (79), Expect = 0.73
Identities = 14/47 (29%), Positives = 18/47 (37%)
Query: 120 EGWDQGWDDDWDDNVAVKSPVVRHAGSISANGLTSRSSNKDGWEDNW 166
+GW GW + W D + S R G + DGW D W
Sbjct: 125 DGWMDGWMNGWMDGSKIGSKEARKEGRKEGRKEGRKEGRMDGWMDGW 171
>UniRef100_Q6HKV0 Hypothetical protein [Bacillus thuringiensis]
Length = 113
Score = 35.0 bits (79), Expect = 0.73
Identities = 24/61 (39%), Positives = 33/61 (53%), Gaps = 8/61 (13%)
Query: 52 SFFLRLP---SFDKIL-TPVNG--AYFLIFTVIVFAVTWACCCIFKKKPR--DEIPYQEL 103
SFF+ LP S+ K+L TP G + +F F +TWA C I+ KK DEI + L
Sbjct: 46 SFFIALPILTSYSKVLNTPAFGDVTWAWVFAFAQFIMTWALCMIYSKKAESFDEISQKIL 105
Query: 104 E 104
+
Sbjct: 106 Q 106
>UniRef100_Q6C4D5 Similar to tr|Q9HEA4 Neurospora crassa [Yarrowia lipolytica]
Length = 749
Score = 35.0 bits (79), Expect = 0.73
Identities = 18/67 (26%), Positives = 31/67 (45%), Gaps = 3/67 (4%)
Query: 105 MALPESASATVVESAEGW--DQGWDDDWDDNVAVK-SPVVRHAGSISANGLTSRSSNKDG 161
+A + A+ ++ +GW D WDD D V +K +P + A ++DG
Sbjct: 682 LATKKPAAPPTLQDNDGWGLDDNWDDGSDGEVVIKEAPKTKPAPKTKKPAKLDLDDDEDG 741
Query: 162 WEDNWDD 168
+D WD+
Sbjct: 742 DDDGWDN 748
>UniRef100_Q84PA3 ADP ribosylation GTPase-like protein [Oryza sativa]
Length = 308
Score = 34.7 bits (78), Expect = 0.96
Identities = 21/61 (34%), Positives = 27/61 (43%), Gaps = 3/61 (4%)
Query: 111 ASATVVESAEGWDQGWDDDWDDNVAVKSPVVRHAGSISANGLTSRS---SNKDGWEDNWD 167
AS V E A+ GW DDW P R + NG S S S+K+ ++WD
Sbjct: 188 ASQKVEEYAKEGGNGWGDDWQRREQGSEPYHRFERETNGNGWNSSSHDGSSKNYNSNSWD 247
Query: 168 D 168
D
Sbjct: 248 D 248
>UniRef100_Q6DDP0 Snag1-prov protein [Xenopus laevis]
Length = 591
Score = 34.3 bits (77), Expect = 1.3
Identities = 18/59 (30%), Positives = 29/59 (48%), Gaps = 2/59 (3%)
Query: 93 KPRDEIPYQELEMALPESASATVVESAEGWDQGWDDDWDD--NVAVKSPVVRHAGSISA 149
+P + L +A A + ++++ D WDD+WDD + A P + AGS SA
Sbjct: 91 QPFQSVAAPHLGLAFQPRAPSLGYQTSQPSDDDWDDEWDDSSSTAADEPAGQPAGSYSA 149
>UniRef100_Q5VJ70 Cell adhesion molecule JCAM [Rattus norvegicus]
Length = 369
Score = 34.3 bits (77), Expect = 1.3
Identities = 27/107 (25%), Positives = 48/107 (44%), Gaps = 10/107 (9%)
Query: 40 LHVTVVTPVPEA--SFFLRLPSFDKILTPVNGAYFLIFTVIVFAVTWACCCIFKKKPRDE 97
+++TVV P P++ LP++ IL V A+ L+ +I+ + CCC ++ ++E
Sbjct: 216 VNLTVVQPPPDSIGGEGQALPTWAIILLAV--AFSLLLILIIALIIIFCCCCVSRREKEE 273
Query: 98 IPYQELEMALPESASATVVESAEGWDQGWDDDWDDNVAVKSPVVRHA 144
YQ E + V + + + W +N KS R A
Sbjct: 274 STYQN------EIRKSANVRTGKADPETWLKSGKENYGYKSDEARAA 314
>UniRef100_Q813S2 Hypothetical Membrane Spanning Protein [Bacillus cereus]
Length = 113
Score = 34.3 bits (77), Expect = 1.3
Identities = 24/61 (39%), Positives = 33/61 (53%), Gaps = 8/61 (13%)
Query: 52 SFFLRLP---SFDKIL-TPVNG--AYFLIFTVIVFAVTWACCCIFKKKPR--DEIPYQEL 103
SFF+ LP S+ K+L TP G + +F F +TWA C I+ KK DEI + L
Sbjct: 46 SFFIALPILTSYSKVLNTPAFGDVTWAWVFAFSQFIMTWALCMIYSKKAESFDEISRKIL 105
Query: 104 E 104
+
Sbjct: 106 Q 106
>UniRef100_UPI00002D1719 UPI00002D1719 UniRef100 entry
Length = 665
Score = 33.5 bits (75), Expect = 2.1
Identities = 16/49 (32%), Positives = 24/49 (48%), Gaps = 8/49 (16%)
Query: 120 EGWDQGWDDDWDDNVAVKSPVVRHAGSISANGLTSRSSNKDGWEDNWDD 168
+ WD DDDWDD GS G+ SR ++++G ++N D
Sbjct: 595 DDWDDEGDDDWDDG--------DDGGSEDMGGIGSRDNHENGTDENGTD 635
>UniRef100_UPI0000361174 UPI0000361174 UniRef100 entry
Length = 584
Score = 33.1 bits (74), Expect = 2.8
Identities = 11/20 (55%), Positives = 15/20 (75%)
Query: 117 ESAEGWDQGWDDDWDDNVAV 136
++++G D WDDDWDDN V
Sbjct: 127 QASQGSDDDWDDDWDDNSTV 146
>UniRef100_UPI0000361173 UPI0000361173 UniRef100 entry
Length = 586
Score = 33.1 bits (74), Expect = 2.8
Identities = 11/20 (55%), Positives = 15/20 (75%)
Query: 117 ESAEGWDQGWDDDWDDNVAV 136
++++G D WDDDWDDN V
Sbjct: 127 QASQGSDDDWDDDWDDNSTV 146
>UniRef100_Q73AP7 Hypothetical protein [Bacillus cereus]
Length = 113
Score = 33.1 bits (74), Expect = 2.8
Identities = 20/48 (41%), Positives = 27/48 (55%), Gaps = 6/48 (12%)
Query: 52 SFFLRLP---SFDKIL-TPVNG--AYFLIFTVIVFAVTWACCCIFKKK 93
SFF+ LP S+ K+L TP G + IF F +TW+ C I+ KK
Sbjct: 46 SFFIALPILTSYSKVLNTPAFGDVTWAWIFAFAQFIMTWSLCMIYSKK 93
>UniRef100_Q5R4T4 Hypothetical protein DKFZp459N1416 [Pongo pygmaeus]
Length = 549
Score = 33.1 bits (74), Expect = 2.8
Identities = 23/71 (32%), Positives = 36/71 (50%), Gaps = 5/71 (7%)
Query: 93 KPRDEIPYQELEMALPESASATVV-ESAEGWDQGWDDDWDDNVAV----KSPVVRHAGSI 147
K +D+I +++L+ + E+ + E EG+D DDDWD + V K V +
Sbjct: 33 KEKDDILFEDLQDNVNENGEGEIEDEEEEGYDDDDDDDWDWDEGVGKLTKGYVWNGGSNP 92
Query: 148 SANGLTSRSSN 158
AN TS SS+
Sbjct: 93 QANRQTSDSSS 103
>UniRef100_Q6BMW6 Similarity [Debaryomyces hansenii]
Length = 413
Score = 33.1 bits (74), Expect = 2.8
Identities = 27/104 (25%), Positives = 46/104 (43%), Gaps = 8/104 (7%)
Query: 63 ILTPVNGAYFLIFTVIVFAVTWACCCIFKKKPRDEIPYQELEMALPESASATV--VESAE 120
I+ V G FL+ V++ W CC I+ K P+DE +L + + T+ + A
Sbjct: 11 IIGAVIGTCFLLSFVMILYCYW-CCVIYVKSPQDEYNVSDLHFRDKQKTTHTLRHMFKAM 69
Query: 121 GWDQGW--DDDWDDNVAVKSP---VVRHAGSISANGLTSRSSNK 159
W G D +D + + P V ++++N SR + K
Sbjct: 70 SWIYGGLRSKDLEDQIDIDKPPSSVTNCDTNLNSNSTFSRLALK 113
>UniRef100_UPI00003AFDCB UPI00003AFDCB UniRef100 entry
Length = 544
Score = 32.7 bits (73), Expect = 3.6
Identities = 16/56 (28%), Positives = 27/56 (47%), Gaps = 4/56 (7%)
Query: 95 RDEIPYQELEMALPESASATVVESAEGWDQGWDDDWDDNVAVKSPVVRHAGSISAN 150
R P ++ P + + ++G D WDDDWDD+ S V G++S++
Sbjct: 41 RGLFPASYVQYPFPPAPYGGSYQPSQGSDDDWDDDWDDS----STVADEPGALSSS 92
>UniRef100_UPI00003AFDCA UPI00003AFDCA UniRef100 entry
Length = 541
Score = 32.7 bits (73), Expect = 3.6
Identities = 16/56 (28%), Positives = 27/56 (47%), Gaps = 4/56 (7%)
Query: 95 RDEIPYQELEMALPESASATVVESAEGWDQGWDDDWDDNVAVKSPVVRHAGSISAN 150
R P ++ P + + ++G D WDDDWDD+ S V G++S++
Sbjct: 41 RGLFPASYVQYPFPPAPYGGSYQPSQGSDDDWDDDWDDS----STVADEPGALSSS 92
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.319 0.135 0.427
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 277,020,542
Number of Sequences: 2790947
Number of extensions: 10761915
Number of successful extensions: 26021
Number of sequences better than 10.0: 32
Number of HSP's better than 10.0 without gapping: 10
Number of HSP's successfully gapped in prelim test: 22
Number of HSP's that attempted gapping in prelim test: 25950
Number of HSP's gapped (non-prelim): 58
length of query: 168
length of database: 848,049,833
effective HSP length: 118
effective length of query: 50
effective length of database: 518,718,087
effective search space: 25935904350
effective search space used: 25935904350
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 70 (31.6 bits)
Medicago: description of AC146784.10


