

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146776.6 - phase: 0
(388 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
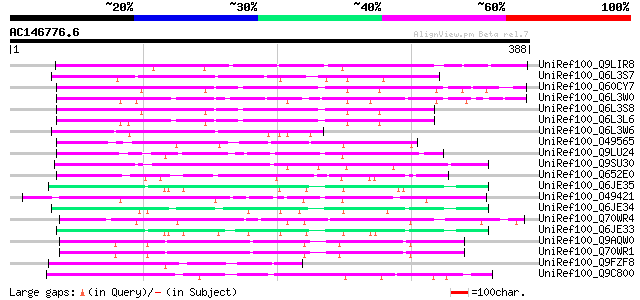
Sequences producing significant alignments: (bits) Value
UniRef100_Q9LIR8 Gb|AAF30317.1 [Arabidopsis thaliana] 151 3e-35
UniRef100_Q6L3S7 Putative F-Box protein [Solanum demissum] 136 1e-30
UniRef100_Q60CY7 Putative F-box protein [Solanum demissum] 119 2e-25
UniRef100_Q6L3W0 Putative F-box protein [Solanum demissum] 114 5e-24
UniRef100_Q6L3S8 Putative F-Box protein [Solanum demissum] 109 1e-22
UniRef100_Q6L3L6 Putative F-box-like protein [Solanum demissum] 104 4e-21
UniRef100_Q6L3W6 Putative S haplotype-specific F-box protein [So... 103 8e-21
UniRef100_O49565 Hypothetical protein F7J7.180 [Arabidopsis thal... 92 2e-17
UniRef100_Q9LU24 Gb|AAF25964.1 [Arabidopsis thaliana] 92 2e-17
UniRef100_Q9SU30 Hypothetical protein AT4g12560 [Arabidopsis tha... 91 7e-17
UniRef100_Q652E0 Hypothetical protein P0463G11.10 [Oryza sativa] 88 5e-16
UniRef100_Q6JE35 S1-locus linked F-box protein [Petunia integrif... 87 1e-15
UniRef100_O49421 Hypothetical protein AT4g19940 [Arabidopsis tha... 84 5e-15
UniRef100_Q6JE34 S2-locus linked F-box protein [Petunia integrif... 82 2e-14
UniRef100_Q70WR4 S locus F-box (SLF)-S4A protein [Antirrhinum hi... 82 2e-14
UniRef100_Q6JE33 S3-locus linked F-box protein [Petunia integrif... 82 2e-14
UniRef100_Q9AQW0 SLF-S2 protein (S locus F-box (SLF)-S2 protein)... 81 4e-14
UniRef100_Q70WR1 S locus F-box (SLF)-S5 protein [Antirrhinum his... 81 4e-14
UniRef100_Q9FZF8 T2E6.11 [Arabidopsis thaliana] 81 6e-14
UniRef100_Q9C800 Hypothetical protein F10C21.17 [Arabidopsis tha... 80 7e-14
>UniRef100_Q9LIR8 Gb|AAF30317.1 [Arabidopsis thaliana]
Length = 364
Score = 151 bits (382), Expect = 3e-35
Identities = 114/364 (31%), Positives = 185/364 (50%), Gaps = 25/364 (6%)
Query: 35 DMPEEIIVEILLRLPVRSLLQFRCVCKLWKTLISDPQFAKKHVSISTAYPQLVSVFVSIA 94
++P E++ EILLRLPV+SL +F+CVC W++LIS+ FA KH I S
Sbjct: 13 NLPLEMMEEILLRLPVKSLTRFKCVCSSWRSLISETLFALKHALILETSKATTSTKSPYG 72
Query: 95 KCNLVSYPLKPL----LDNPSAHRVEPADFEMIHTTSMTIIGSCNGLLCLSDFY--QFTL 148
Y LK L N S V D E++ ++G+C+GL+C Y L
Sbjct: 73 VITTSRYHLKSCCIHSLYNASTVYVSEHDGELLGRDYYQVVGTCHGLVCFHVDYDKSLYL 132
Query: 149 WNPSIKLKSKPSPTIIAFDSFDSKRFLYRGFGYDQVNDRYKVLAVVQNCYNLDETKTLIY 208
WNP+IKL+ + S + + ++ D + + GFGYD+ D YKV+A++Q + + + +T IY
Sbjct: 133 WNPTIKLQQRLSSSDL--ETSDDECVVTYGFGYDESEDDYKVVALLQQRHQV-KIETKIY 189
Query: 209 TFGGKDWTTIQKFPCDPSRCDLGRLGVGKFVSGNLNWIV----SKKVIVFFDIEKETYGE 264
+ K W + FP D R G+ +++G LNW S I+ +D+ ++ + E
Sbjct: 190 STRQKLWRSNTSFPSGVVVADKSRSGI--YINGTLNWAATSSSSSWTIISYDMSRDEFKE 247
Query: 265 MSLPQDYGDKNTVLYVSSNRIYVSF-DHSNKTHWVVWMMKEYGVVESWTKLMIIPQDKLT 323
+ P G + + R +S + + VW+MKE+G V SW+KL+ I
Sbjct: 248 LPGPVCCGRGCFTMTLGDLRGCLSMVCYCKGANADVWVMKEFGEVYSWSKLLSI------ 301
Query: 324 SPGPYCLSDALFISEHGVLLMRPQHSKLSVYNLNNDGGLDYCTTISGQFARYLHIYNESL 383
PG L+IS+ G++++ S L++YN +N G Y + S R +Y +++
Sbjct: 302 -PGLTDFVRPLWISD-GLVVLLEFRSGLALYNCSN-GRFHYPVSNSISGCRDAKVYLKTM 358
Query: 384 VSPH 387
VSP+
Sbjct: 359 VSPN 362
>UniRef100_Q6L3S7 Putative F-Box protein [Solanum demissum]
Length = 372
Score = 136 bits (342), Expect = 1e-30
Identities = 107/312 (34%), Positives = 150/312 (47%), Gaps = 34/312 (10%)
Query: 32 ITADMPEEIIVEILLRLPVRSLLQFRCVCKLWKTLISDPQFAKKHVSI---STAYPQLVS 88
I + +P EII+EILL++P +SLL+F CV K W LIS +F K H+ + Y
Sbjct: 5 IISVLPHEIIIEILLKVPPKSLLKFMCVSKTWLELISSAKFIKTHLELIANDKEYSHHRI 64
Query: 89 VFVSIAKCNLVSYPLKPLLDNPSAHRVEPADFEMIHTTSMT-IIGSCNGLLCL-SDFYQF 146
+F A CN L +L+ + + M + T T I+GS NGL+CL S +
Sbjct: 65 IFQESA-CNFKVCCLPSMLNKERSTELFDIGSPMENPTIYTWIVGSVNGLICLYSKIEET 123
Query: 147 TLWNPSIKLKSKPSPTIIAFDSFDSKRFLYRGFGYDQVNDRYKVLAVVQNCYNLD----E 202
LWNP++K KSK PT+ A +L GFGYD+ D YKV VV C D +
Sbjct: 124 VLWNPAVK-KSKKLPTLGAKLRNGCSYYLKYGFGYDETRDDYKV--VVIQCIYEDSGSCD 180
Query: 203 TKTLIYTFGGKDWTTIQKFPCDPSRCDLGRLGV---GKFVSGNLNWIVSKKV-------I 252
+ IY+ W TI KF G V GKFV+G + W +S V I
Sbjct: 181 SVVNIYSLKADSWRTINKFQ--------GNFLVNSPGKFVNGKIYWALSADVDTFNMCNI 232
Query: 253 VFFDIEKETYGEMSLPQDYGDKNTVL---YVSSNRIYVSFDHSNKTHWVVWMMKEYGVVE 309
+ D+ ET+ + LP YG + L V S+ + + T+ VW+ K+ GV
Sbjct: 233 ISLDLADETWRRLELPDSYGKGSYPLALGVVGSHLSVLCLNCIEGTNSDVWIRKDCGVEV 292
Query: 310 SWTKLMIIPQDK 321
SWTK+ + K
Sbjct: 293 SWTKIFTVDHPK 304
>UniRef100_Q60CY7 Putative F-box protein [Solanum demissum]
Length = 383
Score = 119 bits (297), Expect = 2e-25
Identities = 117/376 (31%), Positives = 167/376 (44%), Gaps = 47/376 (12%)
Query: 36 MPEEIIVEILLRLPVRSLLQFRCVCKLWKTLISDPQFAKKHVSISTAYPQLVSVFVSIAK 95
+P+E+I EILLRLP++SL +F CV K W LIS P F KKH+ ++ + +
Sbjct: 11 LPDELITEILLRLPIKSLSKFMCVSKSWLQLISSPAFVKKHIKLTANDKGYIYHRLIFRN 70
Query: 96 CN--LVSYPLKPLLDNPS-AHRVEPADFEMIHTTSMT-IIGSCNGLLCLSDFYQ--FTLW 149
N PL PL N + D + TT T I+GS NGL+C++ Q +W
Sbjct: 71 TNNDFKFCPLPPLFTNQQLIEEILHIDSPIERTTLSTHIVGSVNGLICVAHVRQREAYIW 130
Query: 150 NPSIKLKSKPSPTIIAFDSFDSKRFLYRGFGYDQVNDRYKVLAVVQNCYNLDETKTLIYT 209
NP+I KSK P + D + GFGYD+ D YKV+ + + T IY+
Sbjct: 131 NPAI-TKSKELPKSTSNLCSDG---IKCGFGYDESRDDYKVVFIDYPIRHNHRTVVNIYS 186
Query: 210 FGGKDWTTIQKFPCDPSRCDLGRLGVGKFVSGNLNWIVSKKV-------IVFFDIEKETY 262
WTT+ +L G+FV+G L W S + I FD+ T+
Sbjct: 187 LRTNSWTTLHDQLQGIFLLNLH----GRFVNGKLYWTSSTCINNYKVCNITSFDLADGTW 242
Query: 263 GEMSLPQDYGDKN--TVLYVSSNRIYVSFDHSNKTHWVVWMMKEYGVVESWTKLMII--P 318
G + LP D + V V S+ + VW+MK GV SWTKL I P
Sbjct: 243 GSLELPSCGKDNSYINVGVVGSDLSLLYTCQLGAATSDVWIMKHSGVNVSWTKLFTIKYP 302
Query: 319 QDKLTSPGPYCLSDALFIS---EHGVLLMRPQHSKLSVYN-----LNNDGGLDYCTTISG 370
Q+ C++ A S HG +L+ S + +Y+ L + ++ C
Sbjct: 303 QNIKIH---RCVAPAFTFSIHIRHGEILLL-LDSAIMIYDGSTRQLKHTFHVNQCE---- 354
Query: 371 QFARYLHIYNESLVSP 386
IY ESLV+P
Sbjct: 355 ------EIYVESLVNP 364
>UniRef100_Q6L3W0 Putative F-box protein [Solanum demissum]
Length = 427
Score = 114 bits (285), Expect = 5e-24
Identities = 121/381 (31%), Positives = 168/381 (43%), Gaps = 55/381 (14%)
Query: 36 MPEEIIVEILLRLPVRSLLQFRCVCKLWKTLISDPQFAKKHVSIST---AYPQLVSVFVS 92
+P+E+I EILL+LPV+SL +F CV K W LIS P F K H+ ++ Y +F +
Sbjct: 53 LPDELITEILLKLPVKSLSKFMCVSKSWLQLISSPTFVKNHIKLTADDKGYIHHRLIFRN 112
Query: 93 I----AKCNLVSYPLKPLLDNPSAHRVEPADFEMIHTTSMTIIGSCNGLLC-LSDFYQFT 147
I C+L K H P + T S I+GS NGL+C + +
Sbjct: 113 IDGNFKFCSLPPLFTKQQHTEELFHIDSPIERS---TLSTHIVGSVNGLICVVHGQKEAY 169
Query: 148 LWNPSIKLKSKPSPTIIAFDSFDSKRFLYRGFGYDQVNDRYKVLAVVQNCYNLDETKTL- 206
+WNP+I KSK P F S + GFGYD+ D YKV+ + YN + +
Sbjct: 170 IWNPTI-TKSKELP---KFTSNMCSSSIKYGFGYDESRDDYKVV-FIHYPYNHSSSSNMT 224
Query: 207 ----IYTFGGKDWTTIQKFPCDPSRCDLGRLGVGKFVSGNLNWIVSKKV-------IVFF 255
IY+ WTT + D +C L G+FV+G L W S + I F
Sbjct: 225 TVVHIYSLRNNSWTTFR----DQLQCFLVN-HYGRFVNGKLYWTSSTCINKYKVCNITSF 279
Query: 256 DIEKETYGEMSLP---QDYGDKNTVLYVSSNRIYVSFDHSNKTHWVVWMMKEYGVVESWT 312
D+ T+G + LP +D D N + S + + T VW+MK V SWT
Sbjct: 280 DLADGTWGSLDLPICGKDNFDINLGVVGSDLSLLYTCQRGAATS-DVWIMKHSAVNVSWT 338
Query: 313 KLMII--PQDKLTSPGPYCLSDALFISEHG-----VLLMRPQHSKLSVYNLNNDGGLDYC 365
KL I PQ+ T C + S H +LL+R S + +Y DG
Sbjct: 339 KLFTIKYPQNIKTH---RCFAPVFTFSIHFRHSEILLLLR---SAIMIY----DGSTRQL 388
Query: 366 TTISGQFARYLHIYNESLVSP 386
S + IY ESLV+P
Sbjct: 389 KHTS-DVMQCEEIYVESLVNP 408
>UniRef100_Q6L3S8 Putative F-Box protein [Solanum demissum]
Length = 327
Score = 109 bits (273), Expect = 1e-22
Identities = 96/298 (32%), Positives = 135/298 (45%), Gaps = 24/298 (8%)
Query: 36 MPEEIIVEILLRLPVRSLLQFRCVCKLWKTLISDPQFAKKHVSISTAYPQLVSVFVSIAK 95
+P+E+I EILL+LP++SLL+F CV K W LIS P F K H+ ++ + +
Sbjct: 12 LPDELITEILLKLPIKSLLKFMCVSKSWLQLISSPAFVKNHIKLTADDKGYIYHRLIFRN 71
Query: 96 CN--LVSYPLKPLLDNPS-AHRVEPADFEMIHTTSMT-IIGSCNGLLCLSDFYQ--FTLW 149
N PL PL + D + TT T I+GS NGL+C + Q +W
Sbjct: 72 TNDDFKFCPLPPLFTQQQLIKELYHIDSPIERTTLSTHIVGSVNGLICAAHVRQREAYIW 131
Query: 150 NPSIKLKSKPSPTIIAFDSFDSKRFLYRGFGYDQVNDRYKVLAVVQNCYNLD-ETKTLIY 208
NP+I KSK P + D + GFGYD+ D YKV+ + + + T IY
Sbjct: 132 NPTI-TKSKELPKSRSNLCSDG---IKCGFGYDESRDDYKVVFIDYPIHRHNHRTVVNIY 187
Query: 209 TFGGKDWTTIQKFPCDPSRCDLGRLGVGKFVSGNLNWIVSKKV-------IVFFDIEKET 261
+ K WTT+ +L G+FV+G L W S + I FD+ T
Sbjct: 188 SLRTKSWTTLHDQLQGFFLLNLH----GRFVNGKLYWTSSSCINNYKVCNITSFDLADGT 243
Query: 262 YGEMSLPQDYGDKN--TVLYVSSNRIYVSFDHSNKTHWVVWMMKEYGVVESWTKLMII 317
+ + LP D + V V S+ + VW+MK GV SWTKL I
Sbjct: 244 WERLELPSCGKDNSYINVGVVGSDLSLLYTCQRGAATSDVWIMKHSGVNVSWTKLFTI 301
>UniRef100_Q6L3L6 Putative F-box-like protein [Solanum demissum]
Length = 384
Score = 104 bits (260), Expect = 4e-21
Identities = 94/300 (31%), Positives = 134/300 (44%), Gaps = 29/300 (9%)
Query: 36 MPEEIIVEILLRLPVRSLLQFRCVCKLWKTLISDPQFAKKHVSIST-----AYPQLV--S 88
+P+E+I EILLRLP++SL +F CV K W LIS P F K H+ ++ Y +L+ +
Sbjct: 12 LPDELITEILLRLPIKSLSKFMCVSKSWLQLISSPAFVKNHIKLTANGKGYIYHRLIFRN 71
Query: 89 VFVSIAKCNLVSYPLKPLLDNPSAHRVEPADFEMIHTTSMTIIGSCNGLLCLSDFYQ--F 146
C L S K L V P + T S I+GS NGL+C + Q
Sbjct: 72 TNDDFKFCPLPSLFTKQQLIEELFDIVSPIERT---TLSTHIVGSVNGLICAAHVRQREA 128
Query: 147 TLWNPSIKLKSKPSPTIIAFDSFDSKRFLYRGFGYDQVNDRYKVLAVVQNCYNLDETKTL 206
+WNP+I KSK P + D + GFGYD+ +D YKV+ + ++ +
Sbjct: 129 YIWNPTI-TKSKELPKSRSNLCSDG---IKCGFGYDESHDDYKVVFINYPSHHNHRSVVN 184
Query: 207 IYTFGGKDWTTIQKFPCDPSRCDLGRLGVGKFVSGNLNWIVSKKV-------IVFFDIEK 259
IY+ WTT+ +L +FV L W S + I FD+
Sbjct: 185 IYSLRTNSWTTLHDQLQGIFLLNLH----CRFVKEKLYWTSSTCINNYKVCNITSFDLAD 240
Query: 260 ETYGEMSLPQDYGDKN--TVLYVSSNRIYVSFDHSNKTHWVVWMMKEYGVVESWTKLMII 317
T+ + LP D + V V S+ + + VW+MK GV SWTKL I
Sbjct: 241 GTWESLELPSCGKDNSYINVGVVGSDLSLLYTCQRGAANSDVWIMKHSGVNVSWTKLFTI 300
>UniRef100_Q6L3W6 Putative S haplotype-specific F-box protein [Solanum demissum]
Length = 240
Score = 103 bits (257), Expect = 8e-21
Identities = 81/223 (36%), Positives = 113/223 (50%), Gaps = 22/223 (9%)
Query: 32 ITADMPEEIIVEILLRLPVRSLLQFRCVCKLWKTLISDPQFAKKHVSISTAYPQLVS--- 88
I + +P EII EILL LP +SLL+FRCV K W LIS +F K H+ TA + S
Sbjct: 5 IISVLPHEIIKEILLNLPPKSLLKFRCVSKSWLELISSAKFIKNHLK-QTANDKEYSHHR 63
Query: 89 VFVSIAKCNLVSYPLKPLLDNPSAHRVEPADFEMIHTTSMT-IIGSCNGLLCL-SDFYQF 146
+ + CN L+ +L+ + + M + + T I+GS NGL+CL S +
Sbjct: 64 IIFQESACNFKVCCLRSMLNKEQSTELFDIGSPMENPSIYTWIVGSVNGLICLYSKIEET 123
Query: 147 TLWNPSIKLKSKPSPTIIAFDSFDSKRFLYRGFGYDQVNDRYKVLA---VVQNCYNL--D 201
LWNP++K KSK PT+ A +L GFGYD+ D YKV++ +V N Y+L D
Sbjct: 124 VLWNPAVK-KSKKLPTLGAKLRNGCSYYLKYGFGYDETRDDYKVMSNMRIVVNIYSLRTD 182
Query: 202 ETKTL--------IYTFGGKDWTTIQKFPC--DPSRCDLGRLG 234
TL I + W ++ F C D S LG +G
Sbjct: 183 SWTTLHDQLHGIFIINHSDETWGCLELFVCGEDNSNIKLGVVG 225
>UniRef100_O49565 Hypothetical protein F7J7.180 [Arabidopsis thaliana]
Length = 417
Score = 92.4 bits (228), Expect = 2e-17
Identities = 76/303 (25%), Positives = 132/303 (43%), Gaps = 55/303 (18%)
Query: 36 MPEEIIVEILLRLPVRSLLQFRCVCKLWKTLISDPQFAKKHVSISTAYPQLVSVFVSIAK 95
+P ++I +ILLRLP +S ++FR V KLW ++ + P F + S A+P S
Sbjct: 36 IPLDMIPDILLRLPAKSAVRFRIVSKLWLSITTRPYFIR-----SFAFPS------STRL 84
Query: 96 CNLVSYPLKPLLDNPSAHRVEPADFEMI---------HTTSMTIIGSCNGLLCLSDFYQF 146
C + + + S H+ + + + H S NGL+C DFY
Sbjct: 85 CLMACVKARDMRLFISLHQHDDGSYAHVDRCEIKSPKHDYYNPSSESVNGLVCFGDFYNI 144
Query: 147 TLWNPSIKL-----KSKPSPTIIAFDSFDSKRFLYRGFGYDQVNDRYKVLAVVQNCYNLD 201
+WNPS++ + KP T+ + F+ GYD V D+YKVL++ + Y+
Sbjct: 145 VVWNPSMRQHVTLPEPKPHSTV--------RYFIRSCLGYDPVEDKYKVLSI--SGYHNG 194
Query: 202 ETKTLIYTFGGKD-WTTIQKFPCDPSRCDLGRLGVGKFVSGNLNWIVS-----------K 249
L++T G ++ W IQ P D G VG ++G++ + +
Sbjct: 195 NHDPLVFTLGPQESWRVIQNSPLD-IPLPTGGSRVGTCINGHVYYEAQIRFKVDDIFNFE 253
Query: 250 KVIVFFDIEKETYGEMSLPQDYGDKNTVLYVSSNRIYVSFDHSNKTHWVV-------WMM 302
+++ FD+ E + + P D +N +L + D+S+ WV+ W +
Sbjct: 254 NILMSFDVRYEKFNTIKKPADPTLRNFMLNYQGKLAWFCSDYSSIRFWVLDDGDKQEWSL 313
Query: 303 KEY 305
K +
Sbjct: 314 KNF 316
>UniRef100_Q9LU24 Gb|AAF25964.1 [Arabidopsis thaliana]
Length = 360
Score = 92.0 bits (227), Expect = 2e-17
Identities = 85/302 (28%), Positives = 139/302 (45%), Gaps = 43/302 (14%)
Query: 36 MPEEIIVEILLRLPVRSLLQFRCVCKLWKTLISDPQFAKKHVSISTAYPQLVSVFVSIAK 95
+PEE+ +EIL+RL ++ L +FRCVCK W+ LI+DP F + + +S A FVS
Sbjct: 5 LPEELAIEILVRLSMKDLARFRCVCKTWRDLINDPGFTETYRDMSPA------KFVSFYD 58
Query: 96 CNLVSYPLKPLLDNPSAHRV------EPADFEMIHTTSMTIIGSCNGLLCLS-DFYQFTL 148
N +LD H V P D MI + T + C+G LC++ + +
Sbjct: 59 KNFY------MLDVEGKHPVITNKLDFPLDQSMIDES--TCVLHCDGTLCVTLKNHTLMV 110
Query: 149 WNP-SIKLKSKPSPTIIAFDSFDSKRFLYRGFGYDQVNDRYKVLAVVQNCYNLDETKTLI 207
WNP S + K P+P I + L GFGYD V+D YKV+ + LD + +
Sbjct: 111 WNPFSKQFKIVPNPGI-----YQDSNIL--GFGYDPVHDDYKVVTFID---RLDVSTAHV 160
Query: 208 YTFGGKDWTTIQKFPCDPSRCDLGRLGVGKFVSGNLNWIV----SKKVIVFFDIEKETYG 263
+ F W + R G F+ L WI + + I+ F++ Y
Sbjct: 161 FEFRTGSWGESLRISYPDWHYRDRR---GTFLDQYLYWIAYRSSADRFILCFNLSTHEYR 217
Query: 264 EMSLP-QDYGDKNTVLYVSSNRIYVSFDHSNKTHWVVWMMKEYGVVESWTKLMIIPQDKL 322
++ LP + G ++ L V+S ++ ++ K + +M++ G SW+K++ +
Sbjct: 218 KLPLPVYNQGVTSSWLGVTSQKLCITEYEMCKKEIRISVMEKTG---SWSKIISLSMSSF 274
Query: 323 TS 324
S
Sbjct: 275 IS 276
>UniRef100_Q9SU30 Hypothetical protein AT4g12560 [Arabidopsis thaliana]
Length = 408
Score = 90.5 bits (223), Expect = 7e-17
Identities = 83/352 (23%), Positives = 157/352 (44%), Gaps = 37/352 (10%)
Query: 34 ADMPEEIIVEILLRLPVRSLLQFRCVCKLWKTLISDPQFAKKHVSISTAYPQLVSVFVSI 93
A +P +I+ +I LRLP ++L++ R + K LI+DP F + H+ + + +
Sbjct: 2 ATIPMDIVNDIFLRLPAKTLVRCRALSKPCYHLINDPDFIESHLHRVLQTGDHLMILLRG 61
Query: 94 AKCNLVSYPLKPLLDNPSAHRVEPADFEMIHTTSMTIIGSCNGLLCLSDF-YQFTLWNPS 152
A L S +D S V + M + GS NGL+ LS+ ++NPS
Sbjct: 62 A-LRLYS------VDLDSLDSVSDVEHPMKRGGPTEVFGSSNGLIGLSNSPTDLAVFNPS 114
Query: 153 IK-LKSKPSPTIIAFDSFDSKRFLYRGFGYDQVNDRYKVLAVVQNCYNLDETKTL----- 206
+ + P +I D ++ +++ G GYD V+D YKV+ +VQ + +D L
Sbjct: 115 TRQIHRLPPSSIDLPDGSSTRGYVFYGLGYDSVSDDYKVVRMVQ--FKIDSEDELGCSFP 172
Query: 207 ----IYTFGGKDWTTIQKFPCDPSRCD------LGRLGVGKFVSGNLNWIVSKK------ 250
+++ W I+ L R G G +L+W++ ++
Sbjct: 173 YEVKVFSLKKNSWKRIESVASSIQLLFYFYYHLLYRRGYGVLAGNSLHWVLPRRPGLIAF 232
Query: 251 -VIVFFDIEKETYGEMSLPQDYGDKNTVLYVSSNRI---YVSFDHSNKTHWVVWMMKEYG 306
+IV FD+ E + + P+ + N + + + + ++++ VWMMKEY
Sbjct: 233 NLIVRFDLALEEFEIVRFPEAVANGNVDIQMDIGVLDGCLCLMCNYDQSYVDVWMMKEYN 292
Query: 307 VVESWTKLMIIPQDKLTSPGPYCLSDALFISEHGVLLMRPQHSKLSVYNLNN 358
V +SWTK+ + + K Y + ++ + +L+ ++KL ++L +
Sbjct: 293 VRDSWTKVFTVQKPKSVKSFSY-MRPLVYSKDKKKVLLELNNTKLVWFDLES 343
>UniRef100_Q652E0 Hypothetical protein P0463G11.10 [Oryza sativa]
Length = 503
Score = 87.8 bits (216), Expect = 5e-16
Identities = 88/319 (27%), Positives = 145/319 (44%), Gaps = 48/319 (15%)
Query: 36 MPEEIIVEILLRLPVRSLLQFRCVCKLWKTLISDPQFAKKHVSISTAYPQLVSVFVSIAK 95
+P++I+ ILLRLPV +LL+ R VCK W +I DPQFA H+ + P +
Sbjct: 20 LPQDIVELILLRLPVSTLLRCRGVCKQWDGIIRDPQFAMVHIQRAPRRP-----LFFFQR 74
Query: 96 CNLVS--YPLKPLLDNPS-------AHRVEPADFEMIHTTSMTIIGSCNGLLCL-SDFYQ 145
NLV YP + +L + + +EP DF + SCNGL+CL SD
Sbjct: 75 ENLVHLLYPSEAILFDEAWSPSKWVVPVIEPDDF---------LCASCNGLICLHSDKST 125
Query: 146 FTLWNPSIKLKSKPSPTIIAFDSFDSKRFLYRGFGYDQVNDRYKVLAVVQNCY---NLDE 202
+ N + + + + + F Y FG+ V +YKV+ +++ +
Sbjct: 126 IKIANLA---TGECMHLVKPVRNSKTDHFSYYSFGFHPVTKQYKVMHFLRDEHLHVGTSF 182
Query: 203 TKTLIYTFGGKDWTTIQKFPCDPSRCDLGRLGVGKFVSGNLNWI------VSKKVIVFFD 256
+ +YT G + W ++ RC + R GV V G + W+ V K +V FD
Sbjct: 183 SIIQVYTLGDEKWRDVRTPQALSLRC-VERSGVVN-VGGAMYWLTEDEESVWKHAVVTFD 240
Query: 257 IEKETYGEMSL----PQDY--GDKNTVLYVS-SNRIYVSFDHSNKTHWVVWMMKEYGVVE 309
+ +E + + L P +Y GD + L + + VS+ + K H +W + + + +
Sbjct: 241 LSEELFQWLQLPAVDPANYVLGDPDQWLITEVDSNVSVSYYETGKLH--IWTI-DSKIEQ 297
Query: 310 SWTKLMIIPQDKLTSPGPY 328
SW++ I L PGP+
Sbjct: 298 SWSQKYNIRLSMLEVPGPH 316
>UniRef100_Q6JE35 S1-locus linked F-box protein [Petunia integrifolia subsp. inflata]
Length = 389
Score = 86.7 bits (213), Expect = 1e-15
Identities = 89/368 (24%), Positives = 147/368 (39%), Gaps = 60/368 (16%)
Query: 30 SEITADMPEEIIVEILLRLPVRSLLQFRCVCKLWKTLISDPQFAKKHVSISTAYPQLVSV 89
++I +PE+++ +LL PV+SLL+F+C+ K W LI F +HV+ T +
Sbjct: 3 NDILMKLPEDLVFLVLLTFPVKSLLRFKCISKAWSILIQSTTFINRHVNRKTNTKDEFII 62
Query: 90 FVSIAKCNLVSYPLKPLLDNPSAHR--VEP--ADFEMIHTTSM------TIIGSCNGLLC 139
F K + K +L S H + P D E+ + TS +IG C+GL+
Sbjct: 63 FKRSIKDEQEGF--KDILSFFSGHDDVLNPLFPDVEVSYMTSKCNCTFNPLIGPCDGLIA 120
Query: 140 LSDFYQFTLWNPSIKLKSKPSPTIIAFDSFDSKRFLYRGFGYDQVNDRYKVLAVVQ-NCY 198
L+D L NP+ + P+ + G G D +++ YKV+ + + C
Sbjct: 121 LTDSIITILLNPATRNFRLLPPSPFGCPKGYHRSVEGVGLGLDTISNYYKVVRISEVYCE 180
Query: 199 NL------DETKTLIYTFGGKDWTTIQK--------FPCDPSRCDLGRLGVGKFVSGNLN 244
++K + G W + PC G ++
Sbjct: 181 EAGGYPGPKDSKIDVCDLGTDSWRELDHVQLPLIYWVPCS-----------GMLYKEMVH 229
Query: 245 WIVS---KKVIVFFDIEKETYGEMSLPQDYGDKNTVLYVSSNRIYVSF---DHSN----- 293
W + VI+ FD+ E + M +P LY + SF +SN
Sbjct: 230 WFATTDESMVILCFDMSTEMFRNMEMPDSCSPITHELYYGLVILCESFTLIGYSNPISSI 289
Query: 294 ---KTHWVVWMMKEYGVVESWTKLMIIPQDKLTSPGPYCLSDALFISEHGVLLMRPQHSK 350
K +W+M EYGV ESW I + SP L + + +LL++ + +
Sbjct: 290 DPVKDKMHIWVMMEYGVSESWIMKYTIKPLSIESP--------LAVWKKNILLLQSRSGR 341
Query: 351 LSVYNLNN 358
L Y+LN+
Sbjct: 342 LISYDLNS 349
>UniRef100_O49421 Hypothetical protein AT4g19940 [Arabidopsis thaliana]
Length = 411
Score = 84.3 bits (207), Expect = 5e-15
Identities = 94/375 (25%), Positives = 163/375 (43%), Gaps = 45/375 (12%)
Query: 10 KTRRRLRQHSQNVTVFTQSVSEITADMPEEIIVEILLRLPVRSLLQFRCVCKLWKTLISD 69
+ RRR ++ +++ Q + EI P ++++EIL RLP +SL++F+ V KLW +LI
Sbjct: 13 RERRRTKRRRRDLCKSRQPIPEI----PFDLVIEILTRLPAKSLMRFKSVSKLWSSLICS 68
Query: 70 PQFAKKHVSIST---AYPQLVSVFVSIAKCNLVSYPLKPLLDNPSAHRVEPADFEMIHTT 126
F + + +S+ + L S S K L+S P D + V D M
Sbjct: 69 RNFTNRLLKLSSPPRLFMCLSSSDNSHLKTVLLSLSSPPDSDITMSSSVIDQDLTMPGMK 128
Query: 127 SMTIIGSCNGLLCLSDFYQFTLWNPS----IKLKSKPSPTIIAFDSFDSKRFLYRGFGYD 182
I GL+CL ++N + + L TI+A + SK+ +Y G+D
Sbjct: 129 GYQISHVFRGLMCLVKKSSAQIYNTTTRQLVVLPDIEESTILA-EEHKSKKIMYH-IGHD 186
Query: 183 QVNDRYKVLAVVQNCYNLDETKTLIYTFGGKDWTTI---------QKFPCD-PSRCDLGR 232
V D+YKV+ +V DE + YTF + W + +K C P LG+
Sbjct: 187 PVYDQYKVVCIVSRA--SDEVEE--YTFLSEHWVLLLEGEGSRRWRKISCKYPPHVPLGQ 242
Query: 233 LGVGKFVSGNLNWIV-----SKKVIVFFDIEKETYGEMSLPQDYGDKNTVLYVSSNRIYV 287
G +SG ++++ +V+V FD E + + +P D K L +I +
Sbjct: 243 ---GLTLSGRMHYLAWVRVSDNRVLVIFDTHSEEFSMLQVPGDIFWKYNGLLEYGGKIAI 299
Query: 288 SFDHSNKTHWVVWMMKEYGVVES-----W-TKLMIIPQDKLTSPGPYCLSDALFISEHGV 341
N T + + E VVE W +K++++ +L L + +G
Sbjct: 300 ----LNYTKVDIEGVMELWVVEDEEKNLWSSKILVVNPLQLQMVNSIISLTVLGTTRNGE 355
Query: 342 LLMRPQHSKLSVYNL 356
+++ P +V+N+
Sbjct: 356 VILVPGPEDKTVFNI 370
>UniRef100_Q6JE34 S2-locus linked F-box protein [Petunia integrifolia subsp. inflata]
Length = 389
Score = 82.4 bits (202), Expect = 2e-14
Identities = 89/376 (23%), Positives = 151/376 (39%), Gaps = 80/376 (21%)
Query: 32 ITADMPEEIIVEILLRLPVRSLLQFRCVCKLWKTLISDPQFAKKHVSISTAYPQLVSVFV 91
I +PE+++ ILL PV+SL++F+C+ K W LI F +H++ T +F
Sbjct: 5 ILKKLPEDLVFLILLTFPVKSLMRFKCISKAWSILIQSTTFINRHINRKTNTKDEFILFK 64
Query: 92 SIAK------CNLVSY------PLKPLL-DNPSAHRVEPADFEMIHTTSMTIIGSCNGLL 138
K N++S+ L PL D ++ D T +IG C+GL+
Sbjct: 65 RAIKDDEEEFINILSFFSGHVDVLNPLFPDMDVSYMTSKCD-----CTFNPLIGPCDGLI 119
Query: 139 CLSDFYQFTLWNPSIKLKSKPSPTIIAFDSFDSKRFLYR-----GFGYDQVNDRYKVLAV 193
L+D + NP+ + + ++ F + +R GFG D +++ YKV+ +
Sbjct: 120 ALTDTIITIVLNPATR-----NFRVLPASPFGCPKGYHRSVEGVGFGLDTISNYYKVVRI 174
Query: 194 VQ-NCYNLD------ETKTLIYTFGGKDW--------TTIQKFPCDPSRCDLGRLGVGKF 238
+ C D ++K + W +I PC G
Sbjct: 175 SEVYCEEADGYPGPKDSKIDVCDLSTDSWRELDHVQLPSIYWVPC-----------AGML 223
Query: 239 VSGNLNWIVS---KKVIVFFDIEKETYGEMSLPQDYGDKNTVLYVS-------------S 282
++W + VI+ FD+ E + +M +P LY S
Sbjct: 224 YKEMVHWFATTDMSMVILCFDMSTEMFHDMKMPDTCSRITHELYYGLVILCESFTLIGYS 283
Query: 283 NRIYVSFDHSNKTHWVVWMMKEYGVVESWTKLMIIPQDKLTSPGPYCLSDALFISEHGVL 342
N I + +K H +W+M EYGV ESW I + SP L + ++ +L
Sbjct: 284 NPISSTDPAHDKMH--IWVMMEYGVSESWIMKYTIRPLSIESP--------LAVWKNHIL 333
Query: 343 LMRPQHSKLSVYNLNN 358
L++ + L Y+LN+
Sbjct: 334 LLQCRSGLLISYDLNS 349
>UniRef100_Q70WR4 S locus F-box (SLF)-S4A protein [Antirrhinum hispanicum]
Length = 391
Score = 82.4 bits (202), Expect = 2e-14
Identities = 100/384 (26%), Positives = 163/384 (42%), Gaps = 55/384 (14%)
Query: 38 EEIIVEILLRLPVRSLLQFRCVCKLWKTLISDPQFAKKHVSISTAYPQLVSVFVSI---- 93
E++ ++IL++L VRSL++FRC+ K W LI F H+ + Y ++ V +
Sbjct: 9 EDVFIQILVKLSVRSLMRFRCISKSWCALIKSSTF---HLLRNQKYDNVLLVRRYLPPPE 65
Query: 94 -----AKCNLVSYPLKPLLDNPSAHRVEPADFEMIH---TTSMTIIGSCNGLLCLSDFYQ 145
+ NL S +K +L N S ++ F+ H + ++G +GLLC++
Sbjct: 66 DEDVFSFYNLNSLEVKQVLPNLSIPLLKDLRFKYDHPYCPEAAYLLGPDSGLLCIACIGN 125
Query: 146 FTLWNPSIK-LKSKPSPTIIAFDSFDSKRFLYRGFGYDQVNDRYKVLAVVQNCYNLDETK 204
+ L NP+++ K P + F ++ F GFG ND +K++ + + D
Sbjct: 126 YYLCNPALREFKQLPPCPFVCPKGFSTEIFA-EGFGCTCTND-FKIVLIRRVTLYDDYDP 183
Query: 205 TL-----IYTFGGKDWTTIQ------KFPCDPSRCDLGRLGVGKFVSGNLNWIVSKKVIV 253
L +YT W T K C+ + +L GV + + N S I+
Sbjct: 184 DLYIMVHLYTSNTNLWRTFAGDIISVKNLCNYACSELLFNGVCHW-NANSTGFSSPDTIL 242
Query: 254 FFDIEKETYGEMSLPQDYGDKNTVLYVSSNRIYVSFDHSNKTHWV-------VWMMKEYG 306
F+I E +G++ D+ ++ VS I W+ +W+M EYG
Sbjct: 243 TFNIGTEVFGQLEFIPDWEEEVYGYCVSLIAIDNCLGMIRYEGWLEEPQLIDIWVMNEYG 302
Query: 307 VVESWTKLMIIPQDKLTSPGPY-CLSDALFISEHGVLLMRPQHSKL--SVYNLNNDGGLD 363
V ESWTK +I GPY + +F + G LL+ +L + N L
Sbjct: 303 VGESWTKSFVI--------GPYEVICPFMFWNNDGWLLVESTEGQLVACALHTNEARRLQ 354
Query: 364 YCTTISGQFARYLH--IYNESLVS 385
C R L IY ESL+S
Sbjct: 355 ICGV-----ERTLRSVIYKESLIS 373
>UniRef100_Q6JE33 S3-locus linked F-box protein [Petunia integrifolia subsp. inflata]
Length = 388
Score = 82.4 bits (202), Expect = 2e-14
Identities = 91/370 (24%), Positives = 150/370 (39%), Gaps = 77/370 (20%)
Query: 36 MPEEIIVEILLRLPVRSLLQFRCVCKLWKTLISDPQFAKKHVSISTAYPQLVSVFVSIAK 95
+PE+++ ILL PV+SL++F+C+ K W LI F +HV+ T +F K
Sbjct: 9 LPEDLVCLILLTFPVKSLMRFKCISKTWSILIQSTTFINRHVNRKTNTKDEFILFKRAIK 68
Query: 96 CNLVSYPLKPLLDNPSAHR--VEP--ADFEMIHTTSM------TIIGSCNGLLCLSDFYQ 145
+ + +L S H + P AD ++ + TS +IG C+GL+ L+D
Sbjct: 69 DEQEEF--RDILSFLSGHDDVLNPLFADIDVSYMTSKCNCAFNPLIGPCDGLIALTDTII 126
Query: 146 FTLWNPSIK--LKSKPSPTIIAFDSFDSKRFLYR-----GFGYDQVNDRYKVLAVVQNCY 198
+ NP+ + PSP F S + +R GFG D +++ YKV+ + +
Sbjct: 127 TIILNPATRNFRLLPPSP-------FGSPKGYHRSVEGVGFGLDTISNYYKVVRISEVYC 179
Query: 199 NLD-------ETKTLIYTFGGKDWTTIQK--------FPCDPSRCDLGRLGVGKFVSGNL 243
D ++K + W + PC G +
Sbjct: 180 EEDGGYPGPKDSKIDAFDLSTDSWRELDHVQLPLIYWLPCS-----------GMLYKEMV 228
Query: 244 NWIVS---KKVIVFFDIEKETYGEMSLP--------QDYG----DKNTVLYVSSNRIYVS 288
+W + VI+ FD+ E + M +P Q YG ++ L N +
Sbjct: 229 HWFATTDMSTVILCFDMSTEMFRNMKMPDTCSVTHKQYYGLVILCESFTLIGYPNPVSPI 288
Query: 289 FDHSNKTHWVVWMMKEYGVVESWTKLMIIPQDKLTSPGPYCLSDALFISEHGVLLMRPQH 348
+K H +W+M EYGV ESW I + SP L + + +LL++
Sbjct: 289 DPAHDKMH--IWVMMEYGVSESWIMKYTIRPLSIESP--------LAVWKKNILLLQSSS 338
Query: 349 SKLSVYNLNN 358
L Y+LN+
Sbjct: 339 GLLISYDLNS 348
>UniRef100_Q9AQW0 SLF-S2 protein (S locus F-box (SLF)-S2 protein) [Antirrhinum
hispanicum]
Length = 376
Score = 81.3 bits (199), Expect = 4e-14
Identities = 86/339 (25%), Positives = 137/339 (40%), Gaps = 51/339 (15%)
Query: 38 EEIIVEILLRLPVRSLLQFRCVCKLWKTLISDPQFAKKHV--SISTAYPQLVSVFVSIAK 95
+++I EILL V+SLL+FRCV K W +LI F H+ + +V +V +
Sbjct: 9 QDVISEILLFSSVKSLLRFRCVSKSWCSLIKSNDFIDNHLLRRQTNGNVMVVKRYVRTPE 68
Query: 96 CNLVSY---------PLKPLLDNPSAHRVEPADFEMIHTTS-MTIIGSCNGLLCLSDFYQ 145
++ S+ L P L NP ++ D++ + + ++G CNGL+CL+
Sbjct: 69 RDMFSFYNINSPELDELLPDLPNPYFKNIK-FDYDYFYLPQRVNLMGPCNGLICLAYGDC 127
Query: 146 FTLWNPSIKLKSKPSPTIIAFDSFDSKRFLYRGFGYDQVNDRYKVLAVVQNCYNLDETKT 205
L NP+++ + PT A + GFG + ND YKV+ +
Sbjct: 128 VLLSNPALREIKRLPPTPFANPEGHCTDIIGYGFG-NTCNDCYKVVLIESVGPEDHHINI 186
Query: 206 LIYTFGGKDWTTIQ----------KFPCDPSRCDLGRLGVGKFVSGNLNW------IVSK 249
+Y W I+ FPC+ F G +W I
Sbjct: 187 YVYYSDTNSWKHIEDDSTPIKYICHFPCNE-----------LFFKGAFHWNANSTDIFYA 235
Query: 250 KVIVFFDIEKETYGEMSLPQDYGD-KNTVLYVSSNRIYVSFDHSNKTHWV-------VWM 301
I+ FDI E + EM+ P N+ L + S ++ + W+ +W+
Sbjct: 236 DFILTFDIITEVFKEMAYPHCLAQFSNSFLSLMSLNECLAMVRYKE--WMEDPELFDIWV 293
Query: 302 MKEYGVVESWTKLMIIPQDKLTSPGPYCLSDALFISEHG 340
M +YGV ESWTK +I + +D I E G
Sbjct: 294 MNQYGVRESWTKQYVIGPQVVVCSHVCWKNDECLIVEDG 332
>UniRef100_Q70WR1 S locus F-box (SLF)-S5 protein [Antirrhinum hispanicum]
Length = 376
Score = 81.3 bits (199), Expect = 4e-14
Identities = 86/339 (25%), Positives = 137/339 (40%), Gaps = 51/339 (15%)
Query: 38 EEIIVEILLRLPVRSLLQFRCVCKLWKTLISDPQFAKKHV--SISTAYPQLVSVFVSIAK 95
+++I EILL V+SLL+FRCV K W +LI F H+ + +V +V +
Sbjct: 9 QDVISEILLFSSVKSLLRFRCVSKSWCSLIKSNDFIDNHLLRRQTNGNVMVVKRYVRTPE 68
Query: 96 CNLVSY---------PLKPLLDNPSAHRVEPADFEMIHTTS-MTIIGSCNGLLCLSDFYQ 145
++ S+ L P L NP ++ D++ + + ++G CNGL+CL+
Sbjct: 69 RDMFSFYNINSPKLEELLPDLPNPYFKNIK-FDYDYFYLPQRVNLMGPCNGLICLAYGDC 127
Query: 146 FTLWNPSIKLKSKPSPTIIAFDSFDSKRFLYRGFGYDQVNDRYKVLAVVQNCYNLDETKT 205
L NP+++ + PT A + GFG + ND YKV+ +
Sbjct: 128 VLLSNPALREIKRLPPTPFANPEGHCTDIIGYGFG-NTCNDCYKVVLIESVGPEDHHINI 186
Query: 206 LIYTFGGKDWTTIQ----------KFPCDPSRCDLGRLGVGKFVSGNLNW------IVSK 249
+Y W I+ FPC+ F G +W I
Sbjct: 187 YVYYSDTNSWKHIEDDSTPIKYICHFPCNE-----------LFFKGAFHWNANSTDIFYA 235
Query: 250 KVIVFFDIEKETYGEMSLPQDYGD-KNTVLYVSSNRIYVSFDHSNKTHWV-------VWM 301
I+ FDI E + EM+ P N+ L + S ++ + W+ +W+
Sbjct: 236 DFILTFDIITEVFKEMAYPHCLAQFSNSFLSLMSLNECLAMVRYKE--WMEEPELFDIWV 293
Query: 302 MKEYGVVESWTKLMIIPQDKLTSPGPYCLSDALFISEHG 340
M +YGV ESWTK +I + +D I E G
Sbjct: 294 MNQYGVRESWTKQYVIGPQVVVCSHVCWKNDECLIVEDG 332
>UniRef100_Q9FZF8 T2E6.11 [Arabidopsis thaliana]
Length = 389
Score = 80.9 bits (198), Expect = 6e-14
Identities = 59/198 (29%), Positives = 93/198 (46%), Gaps = 21/198 (10%)
Query: 30 SEITADMPEEIIVEILLRLPVRSLLQFRCVCKLWKTLISDPQFAKKHVSISTAYPQLVSV 89
S+ T+ P ++ EILLRLPV+S+++FRCV KLW ++I+DP F K + + S+ L+
Sbjct: 19 SKPTSSFPLDLASEILLRLPVKSVVRFRCVSKLWSSIITDPYFIKTYETQSSTRQSLLFC 78
Query: 90 FVSIAKCNLVSYPLKPLLDNPSAHRV-------EPADFEMIHTTSMTIIGSCNGLLCLSD 142
F K + S P N S+ P +F T S +GL+C
Sbjct: 79 FKQSDKLFVFSIPKHHYDSNSSSQAAIDRFQVKLPQEFSYPSPTE-----SVHGLICFHV 133
Query: 143 FYQFTLWNPSIK-LKSKPSPTIIAFDSFDSKRFLYRGFGYDQVNDRYKVLAVVQNCYNLD 201
+WNPS++ + P P S + L GYD + ++KV+ + +N D
Sbjct: 134 LATVIVWNPSMRQFLTLPKPR-------KSWKELTVFLGYDPIEGKHKVVCLPRN-RTCD 185
Query: 202 ETKTLIYTFGGKDWTTIQ 219
E + L K W T++
Sbjct: 186 ECQVLTLGSAQKSWRTVK 203
>UniRef100_Q9C800 Hypothetical protein F10C21.17 [Arabidopsis thaliana]
Length = 441
Score = 80.5 bits (197), Expect = 7e-14
Identities = 82/357 (22%), Positives = 147/357 (40%), Gaps = 46/357 (12%)
Query: 28 SVSEITADMPEEIIVEILLRLPVRSLLQFRCVCKLWKTLISDPQFAKKHVSISTAYPQLV 87
S + + ++P+ ++ EIL RLPV+ L++ + + K WK+LI A+KH+ + L
Sbjct: 89 SETTLAVELPDVLVEEILQRLPVKYLVRLKSISKGWKSLIESDHLAEKHLRLLEKKYGLK 148
Query: 88 SVFVSIAKCNLVSYPLKPLLDNPSAHRVEPADFEMIHTTSMTIIGSCNGLLCL--SDFYQ 145
+ +++ + S +K + + +++ + GSCNGL+C+ D
Sbjct: 149 EIKITVERSTSKSICIKFFSRRSGMNAINSDSDDLLR-----VPGSCNGLVCVYELDSVY 203
Query: 146 FTLWNPSIKLKSKPSPTIIAFDSFDSKRFLYRGFGYDQVNDRYKVLAVVQNCYNLDETKT 205
L NP + +P L GFG D V YKV+ + Y D T
Sbjct: 204 IYLLNPMTGVTRTLTP--------PRGTKLSVGFGIDVVTGTYKVMVL----YGFDRVGT 251
Query: 206 LIYTFGGKDWTTIQKFPCD-PSRCDLGRLGVGKFVSGNLNWIVSK--KVIVFFDIEKETY 262
+++ W K P C FV+G+L W+++ I+ D+ E +
Sbjct: 252 VVFDLDTNKWRQRYKTAGPMPLSCIPTPERNPVFVNGSLFWLLASDFSEILVMDLHTEKF 311
Query: 263 GEMSLPQDYGDKNT-----VLYVSSNRIYVSFDHSNKTHWVVWMMKEYGVVESWTKLM-- 315
+S P D D + ++ +R+ VS + H VW++ + + E W +
Sbjct: 312 RTLSQPNDMDDVDVSSGYIYMWSLEDRLCVS-NVRQGLHSYVWVLVQDELSEKWERTRFN 370
Query: 316 ----IIPQDKLTSP-------GPYCLSDALFISEHGVLLMRPQHSKLSVYNLNNDGG 361
+ P L S PY LS + I + Q+S ++++ N D G
Sbjct: 371 LLGHVFPPLSLNSAWFSQTLVSPYQLSSSTCIGSR-----QRQNSTSALFSRNGDTG 422
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.323 0.138 0.430
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 677,205,412
Number of Sequences: 2790947
Number of extensions: 28707529
Number of successful extensions: 56618
Number of sequences better than 10.0: 507
Number of HSP's better than 10.0 without gapping: 378
Number of HSP's successfully gapped in prelim test: 129
Number of HSP's that attempted gapping in prelim test: 55884
Number of HSP's gapped (non-prelim): 593
length of query: 388
length of database: 848,049,833
effective HSP length: 129
effective length of query: 259
effective length of database: 488,017,670
effective search space: 126396576530
effective search space used: 126396576530
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 76 (33.9 bits)
Medicago: description of AC146776.6


