

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146758.8 - phase: 0 /pseudo
(35 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
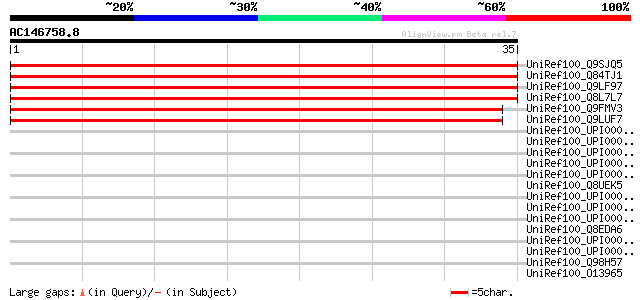
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9SJQ5 Hypothetical protein At2g36500 [Arabidopsis tha... 56 2e-07
UniRef100_Q84TJ1 Hypothetical protein At2g36500 [Arabidopsis tha... 56 2e-07
UniRef100_Q9LF97 Hypothetical protein F8J2_120 [Arabidopsis thal... 55 7e-07
UniRef100_Q8L7L7 Hypothetical protein At3g52950 [Arabidopsis tha... 55 7e-07
UniRef100_Q9FMV3 Emb|CAB86899.1 [Arabidopsis thaliana] 52 3e-06
UniRef100_Q9LUF7 Emb|CAB86899.1 [Arabidopsis thaliana] 51 7e-06
UniRef100_UPI000023E7CF UPI000023E7CF UniRef100 entry 36 0.25
UniRef100_UPI0000234E63 UPI0000234E63 UniRef100 entry 36 0.25
UniRef100_UPI000021B39B UPI000021B39B UniRef100 entry 36 0.25
UniRef100_UPI00003C1641 UPI00003C1641 UniRef100 entry 35 0.56
UniRef100_UPI000042E7F9 UPI000042E7F9 UniRef100 entry 34 1.2
UniRef100_Q8UEK5 Hypothetical protein Atu1752 [Agrobacterium tum... 33 1.6
UniRef100_UPI000042CD55 UPI000042CD55 UniRef100 entry 33 2.1
UniRef100_UPI0000342CE6 UPI0000342CE6 UniRef100 entry 32 4.7
UniRef100_UPI00002AD132 UPI00002AD132 UniRef100 entry 32 4.7
UniRef100_Q8EDA6 CBS domain protein [Shewanella oneidensis] 32 4.7
UniRef100_UPI0000311ECC UPI0000311ECC UniRef100 entry 32 6.1
UniRef100_UPI000029351E UPI000029351E UniRef100 entry 32 6.1
UniRef100_Q98H57 Mll3023 protein [Rhizobium loti] 32 6.1
UniRef100_O13965 Hypothetical protein C24C9.05c in chromosome I ... 32 6.1
>UniRef100_Q9SJQ5 Hypothetical protein At2g36500 [Arabidopsis thaliana]
Length = 536
Score = 56.2 bits (134), Expect = 2e-07
Identities = 28/35 (80%), Positives = 32/35 (91%)
Query: 1 MAARRVDAVLLTDANGLLFGIMTDKDTATRVISEG 35
MAARRVDAVLLTD++ LL GI+TDKD ATRVI+EG
Sbjct: 86 MAARRVDAVLLTDSSALLSGIVTDKDIATRVIAEG 120
>UniRef100_Q84TJ1 Hypothetical protein At2g36500 [Arabidopsis thaliana]
Length = 536
Score = 56.2 bits (134), Expect = 2e-07
Identities = 28/35 (80%), Positives = 32/35 (91%)
Query: 1 MAARRVDAVLLTDANGLLFGIMTDKDTATRVISEG 35
MAARRVDAVLLTD++ LL GI+TDKD ATRVI+EG
Sbjct: 86 MAARRVDAVLLTDSSALLSGIVTDKDIATRVIAEG 120
>UniRef100_Q9LF97 Hypothetical protein F8J2_120 [Arabidopsis thaliana]
Length = 556
Score = 54.7 bits (130), Expect = 7e-07
Identities = 27/35 (77%), Positives = 31/35 (88%)
Query: 1 MAARRVDAVLLTDANGLLFGIMTDKDTATRVISEG 35
MAARRVDA LLTD++ LL GI+TDKD ATRVI+EG
Sbjct: 88 MAARRVDACLLTDSSALLSGIVTDKDVATRVIAEG 122
>UniRef100_Q8L7L7 Hypothetical protein At3g52950 [Arabidopsis thaliana]
Length = 469
Score = 54.7 bits (130), Expect = 7e-07
Identities = 27/35 (77%), Positives = 31/35 (88%)
Query: 1 MAARRVDAVLLTDANGLLFGIMTDKDTATRVISEG 35
MAARRVDA LLTD++ LL GI+TDKD ATRVI+EG
Sbjct: 1 MAARRVDACLLTDSSALLSGIVTDKDVATRVIAEG 35
>UniRef100_Q9FMV3 Emb|CAB86899.1 [Arabidopsis thaliana]
Length = 543
Score = 52.4 bits (124), Expect = 3e-06
Identities = 25/34 (73%), Positives = 31/34 (90%)
Query: 1 MAARRVDAVLLTDANGLLFGIMTDKDTATRVISE 34
MA+RRVDA+LLTD+N +L GI+TDKD ATRVIS+
Sbjct: 79 MASRRVDALLLTDSNEMLCGILTDKDIATRVISQ 112
>UniRef100_Q9LUF7 Emb|CAB86899.1 [Arabidopsis thaliana]
Length = 548
Score = 51.2 bits (121), Expect = 7e-06
Identities = 24/34 (70%), Positives = 31/34 (90%)
Query: 1 MAARRVDAVLLTDANGLLFGIMTDKDTATRVISE 34
MAARRVDA+LLTD+N LL GI+TD+D AT+VI++
Sbjct: 87 MAARRVDALLLTDSNALLCGILTDRDIATKVIAK 120
>UniRef100_UPI000023E7CF UPI000023E7CF UniRef100 entry
Length = 680
Score = 36.2 bits (82), Expect = 0.25
Identities = 18/35 (51%), Positives = 23/35 (65%)
Query: 1 MAARRVDAVLLTDANGLLFGIMTDKDTATRVISEG 35
MAA+R D VL+TD + + GI T KD A RV+ G
Sbjct: 121 MAAKREDCVLVTDDDDRIAGIFTAKDLAFRVVGAG 155
>UniRef100_UPI0000234E63 UPI0000234E63 UniRef100 entry
Length = 666
Score = 36.2 bits (82), Expect = 0.25
Identities = 18/35 (51%), Positives = 23/35 (65%)
Query: 1 MAARRVDAVLLTDANGLLFGIMTDKDTATRVISEG 35
MAA+R D VL+TD + + GI T KD A RV+ G
Sbjct: 132 MAAKREDCVLVTDDDDRIAGIFTAKDLAFRVVGAG 166
>UniRef100_UPI000021B39B UPI000021B39B UniRef100 entry
Length = 733
Score = 36.2 bits (82), Expect = 0.25
Identities = 18/35 (51%), Positives = 23/35 (65%)
Query: 1 MAARRVDAVLLTDANGLLFGIMTDKDTATRVISEG 35
MAA+R D VL+TD + + GI T KD A RV+ G
Sbjct: 161 MAAKREDCVLVTDDDDRIAGIFTAKDLAFRVVGAG 195
>UniRef100_UPI00003C1641 UPI00003C1641 UniRef100 entry
Length = 708
Score = 35.0 bits (79), Expect = 0.56
Identities = 18/34 (52%), Positives = 22/34 (63%)
Query: 2 AARRVDAVLLTDANGLLFGIMTDKDTATRVISEG 35
AA+R D VL+ D + L GI T KD A RV+S G
Sbjct: 87 AAKRTDCVLVVDEDEHLAGIFTAKDLAFRVVSAG 120
>UniRef100_UPI000042E7F9 UPI000042E7F9 UniRef100 entry
Length = 831
Score = 33.9 bits (76), Expect = 1.2
Identities = 18/34 (52%), Positives = 21/34 (60%)
Query: 2 AARRVDAVLLTDANGLLFGIMTDKDTATRVISEG 35
AA+R D VL+ D L GI T KD A RV +EG
Sbjct: 235 AAKRADCVLVVDEEEGLSGIFTAKDLAFRVTAEG 268
>UniRef100_Q8UEK5 Hypothetical protein Atu1752 [Agrobacterium tumefaciens]
Length = 144
Score = 33.5 bits (75), Expect = 1.6
Identities = 13/33 (39%), Positives = 23/33 (69%)
Query: 3 ARRVDAVLLTDANGLLFGIMTDKDTATRVISEG 35
A ++ AV++TDA+G++ GI T++D V +G
Sbjct: 34 AHKIGAVVVTDADGVVLGIFTERDLVKAVAGQG 66
>UniRef100_UPI000042CD55 UPI000042CD55 UniRef100 entry
Length = 605
Score = 33.1 bits (74), Expect = 2.1
Identities = 16/35 (45%), Positives = 21/35 (59%)
Query: 1 MAARRVDAVLLTDANGLLFGIMTDKDTATRVISEG 35
M ARR + VL+ + G L GI T KD A R++ G
Sbjct: 73 MTARRENCVLVVNEIGELLGIFTAKDVAFRIVGSG 107
Score = 31.6 bits (70), Expect = 6.1
Identities = 16/35 (45%), Positives = 19/35 (53%)
Query: 1 MAARRVDAVLLTDANGLLFGIMTDKDTATRVISEG 35
M R AVL+ D N + GI T KD RVI+ G
Sbjct: 245 MKENRTTAVLVKDTNEQVAGIFTSKDVVLRVIAAG 279
>UniRef100_UPI0000342CE6 UPI0000342CE6 UniRef100 entry
Length = 360
Score = 32.0 bits (71), Expect = 4.7
Identities = 14/33 (42%), Positives = 22/33 (66%)
Query: 1 MAARRVDAVLLTDANGLLFGIMTDKDTATRVIS 33
M RR++ +L+TD G L G++T KDT V++
Sbjct: 50 MKERRIEKLLITDKRGRLTGLLTLKDTEQAVLN 82
>UniRef100_UPI00002AD132 UPI00002AD132 UniRef100 entry
Length = 317
Score = 32.0 bits (71), Expect = 4.7
Identities = 14/33 (42%), Positives = 22/33 (66%)
Query: 1 MAARRVDAVLLTDANGLLFGIMTDKDTATRVIS 33
M RR++ +L+TD G L G++T KDT V++
Sbjct: 183 MKERRIEKLLITDKQGRLTGLLTLKDTEQAVLN 215
>UniRef100_Q8EDA6 CBS domain protein [Shewanella oneidensis]
Length = 620
Score = 32.0 bits (71), Expect = 4.7
Identities = 17/35 (48%), Positives = 23/35 (65%), Gaps = 1/35 (2%)
Query: 1 MAARRVDAVLLTDANGLLFGIMTDKDTATRVISEG 35
M RV ++L+TD N L GI+TDKD RV++ G
Sbjct: 176 MRNSRVSSLLVTD-NHKLVGILTDKDLRNRVLASG 209
>UniRef100_UPI0000311ECC UPI0000311ECC UniRef100 entry
Length = 620
Score = 31.6 bits (70), Expect = 6.1
Identities = 17/35 (48%), Positives = 23/35 (65%), Gaps = 1/35 (2%)
Query: 1 MAARRVDAVLLTDANGLLFGIMTDKDTATRVISEG 35
M RV ++L+TD N L GI+TDKD RV++ G
Sbjct: 176 MRNSRVSSLLVTD-NRKLVGILTDKDLRNRVLAAG 209
>UniRef100_UPI000029351E UPI000029351E UniRef100 entry
Length = 620
Score = 31.6 bits (70), Expect = 6.1
Identities = 17/35 (48%), Positives = 23/35 (65%), Gaps = 1/35 (2%)
Query: 1 MAARRVDAVLLTDANGLLFGIMTDKDTATRVISEG 35
M RV ++L+TD N L GI+TDKD RV++ G
Sbjct: 176 MRNSRVSSLLVTD-NHKLVGILTDKDLRNRVLAAG 209
>UniRef100_Q98H57 Mll3023 protein [Rhizobium loti]
Length = 333
Score = 31.6 bits (70), Expect = 6.1
Identities = 14/28 (50%), Positives = 20/28 (71%)
Query: 1 MAARRVDAVLLTDANGLLFGIMTDKDTA 28
++ ++V VL+ DANG L GI+TD D A
Sbjct: 239 LSQKKVGCVLIVDANGELAGIITDGDVA 266
>UniRef100_O13965 Hypothetical protein C24C9.05c in chromosome I [Schizosaccharomyces
pombe]
Length = 730
Score = 31.6 bits (70), Expect = 6.1
Identities = 16/35 (45%), Positives = 22/35 (62%)
Query: 1 MAARRVDAVLLTDANGLLFGIMTDKDTATRVISEG 35
MAA+R + VL+ D + L GI+T D ATR + G
Sbjct: 89 MAAKRQNCVLVVDDDEQLAGIVTATDIATRCVGAG 123
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.322 0.136 0.360
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 41,858,264
Number of Sequences: 2790947
Number of extensions: 655776
Number of successful extensions: 1531
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 18
Number of HSP's successfully gapped in prelim test: 4
Number of HSP's that attempted gapping in prelim test: 1502
Number of HSP's gapped (non-prelim): 33
length of query: 35
length of database: 848,049,833
effective HSP length: 11
effective length of query: 24
effective length of database: 817,349,416
effective search space: 19616385984
effective search space used: 19616385984
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 69 (31.2 bits)
Medicago: description of AC146758.8


