

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146755.8 + phase: 0 /pseudo
(1711 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
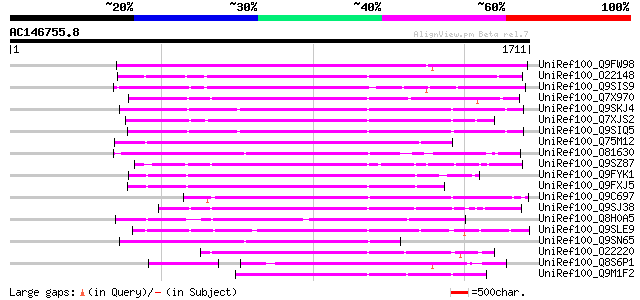
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9FW98 Putative non-LTR retroelement reverse transcrip... 803 0.0
UniRef100_O22148 Putative non-LTR retroelement reverse transcrip... 794 0.0
UniRef100_Q9SIS9 Putative non-LTR retroelement reverse transcrip... 749 0.0
UniRef100_Q7X970 BZIP-like protein [Oryza sativa] 736 0.0
UniRef100_Q9SKJ4 Putative non-LTR retroelement reverse transcrip... 734 0.0
UniRef100_Q7XJS2 Putative non-LTR retroelement reverse transcrip... 719 0.0
UniRef100_Q9SIQ5 Putative non-LTR retroelement reverse transcrip... 715 0.0
UniRef100_Q75M12 Hypothetical protein P0676G05.2 [Oryza sativa] 710 0.0
UniRef100_O81630 F8M12.22 protein [Arabidopsis thaliana] 702 0.0
UniRef100_Q9SZ87 RNA-directed DNA polymerase-like protein [Arabi... 692 0.0
UniRef100_Q9FYK1 F21J9.30 [Arabidopsis thaliana] 677 0.0
UniRef100_Q9FXJ5 F5A9.24 [Arabidopsis thaliana] 674 0.0
UniRef100_Q9C697 Reverse transcriptase, putative; 16838-20266 [A... 668 0.0
UniRef100_Q9SJ38 Putative non-LTR retroelement reverse transcrip... 653 0.0
UniRef100_Q8H0A5 Putative reverse transcriptase [Oryza sativa] 650 0.0
UniRef100_Q9SLE9 Putative non-LTR retroelement reverse transcrip... 635 e-180
UniRef100_Q9SN65 Hypothetical protein F25I24.40 [Arabidopsis tha... 627 e-178
UniRef100_O22220 Putative non-LTR retroelement reverse transcrip... 562 e-158
UniRef100_Q8S6P1 Putative reverse transcriptase [Oryza sativa] 560 e-157
UniRef100_Q9M1F2 Hypothetical protein F9K21.130 [Arabidopsis tha... 535 e-150
>UniRef100_Q9FW98 Putative non-LTR retroelement reverse transcriptase [Oryza sativa]
Length = 1382
Score = 803 bits (2073), Expect = 0.0
Identities = 468/1384 (33%), Positives = 727/1384 (51%), Gaps = 36/1384 (2%)
Query: 351 PRKMSFIVW--NCRGLGSPSTVPTLKYLVRTYKPEGIFLSETMAALNKIEELKYILSFDS 408
P K+ + W NCRGLGS +TV L++LV++ +P +FLSET + L + L F
Sbjct: 3 PNKL--VTWGRNCRGLGSAATVGELRWLVKSLRPSLVFLSETKMRDKQARNLMWSLGFSG 60
Query: 409 CFTVDRIGRGGGLAFLWKNSANCTITNFSQNHIDVEVDDLLIGKWRLTGFYGMPENGRRK 468
F V G GGLA W + ++ F+ + IDV V + WR++ YG P+ R
Sbjct: 61 SFAVSCEGLSGGLALFWTTAYTVSLRGFNSHFIDVLVSTEELPPWRISFVYGEPKRELRH 120
Query: 469 ESWAFLKNLARTSSLPWCVIGDFNDILSSEEKKGRSERAPWLIRGFRQAVLDAGLFDLHM 528
W L+ L PW GDFN++L +E G ER+ ++ FR + D GL DL
Sbjct: 121 FFWNLLRRLHDQWRGPWLCCGDFNEVLCLDEHLGMRERSEPHMQHFRSCLDDCGLIDLGF 180
Query: 529 TGYAFTWFKSLGTARAVEEKLDRALVNSDWCNMFPNAGVECLSTTSSDHYPLWLTCKPVT 588
G FTW + +LDRA+ N ++ F + VE + TTSSDHY + +
Sbjct: 181 VGPKFTWSNKQDANSNSKVRLDRAVANGEFSRYFEDCLVENVITTSSDHYAISIDLSRRN 240
Query: 589 AVNRN---PHRFKFENAWLAEPDFKQQVQQRWKLYPE-----EGITQKLSYCAEDLTDWS 640
R F+FE AWL D+++ V+ W++ G+ L A L DWS
Sbjct: 241 HGQRRIPIQQGFRFEAAWLRAEDYREVVENSWRISSAGCVGLRGVWSVLQQVAVSLKDWS 300
Query: 641 RNN-NNFRRDISKVQKKIEKLR-THVTAANVSYFNSLKNKLDKLLVQDDLFWKQRAKTFW 698
+ + + RR I K+++K++ LR + V + ++ +L +L ++++ +QR++ W
Sbjct: 301 KASFGSVRRKILKMERKLKSLRQSPVNDVVIQEEKLIEQQLCELFEKEEIMARQRSRVDW 360
Query: 699 YKEGDLNTKFFHAAATSRRKVNRIEHLENSNGTVCRKEDELQNIARNYFSHLFT-KLSST 757
+EGD NT FFHA A++RR+ NRI+ L +G+ C ++ ++ +A ++ +LF+ + +
Sbjct: 361 LREGDRNTAFFHARASARRRTNRIKELVRDDGSRCISQEGIKRMAEVFYENLFSSEPCDS 420
Query: 758 RADVISKVATSISDDDNCFLTAAFTLEEFKTAAFSMQADKCPGPDGFNPGFYQNFWNMCG 817
+V+ + + D N L +T EE KTA F M + K PGPDGF FYQ W +
Sbjct: 421 MEEVLDAIPNKVGDFINGELGKQYTNEEIKTALFQMGSTKAPGPDGFPALFYQTHWGILE 480
Query: 818 SEIHKAGCEWLENGVFPPHLNSTNIALIPKGDAQTTMKDWRPIALCNVVYKMVAKVLANR 877
I A +L P L + + LIPK + + + +RPI+LCNV+YK+ +KVLANR
Sbjct: 481 EHICNAVRGFLLGEEIPEGLCDSVVVLIPKVNNASHLSKFRPISLCNVLYKIASKVLANR 540
Query: 878 LKNVLDKCISTSQSAFVPGRSILDNALVAIELIHYMKAKTKGSQGDVALKLDISKAYDRL 937
LK L +S QSAFVPGR I D+ALVA E +H ++ K ALK+D+ KAYDR+
Sbjct: 541 LKPFLPDIVSEFQSAFVPGRLITDSALVAYECLHTIR-KQHNKNPFFALKIDMMKAYDRV 599
Query: 938 DWEYLRDIMIQMRFSEKWIRWIMLCVETVDYTVLVNGVQVGPLIPGRGIRQGDPLSPYLF 997
+W YL + ++ FS+ WI +M CV +V Y V +NG P++P RGIRQGDP+SPYLF
Sbjct: 600 EWAYLSGCLSKLGFSQDWINTVMRCVSSVRYAVKINGELTKPVVPSRGIRQGDPISPYLF 659
Query: 998 ILCAEGLSALIRDAERRGVITGTRICTSAPAISHLLFADDCFLFFRASEQEASVMKNILT 1057
+LC EGLS L+ E G + G + P ISHLLFADD F +A + +KN L
Sbjct: 660 LLCTEGLSCLLHKKEVAGELQGIKNGRHGPPISHLLFADDSIFFAKADSRNVQALKNTLR 719
Query: 1058 TYEAASGQAINLQKSEMYCSRNTPTECQNRIATTLGVKQVLGTGKYLGLPSMIGRSKKAT 1117
+Y +ASGQ INL KS ++ + P + + + L V + YLG+P+ IG +
Sbjct: 720 SYCSASGQKINLHKSSIFFGKRCPDAVKISVKSCLQVDNEVLQDSYLGMPTEIGLATTNF 779
Query: 1118 FKFVKDRIWNRINSWSSRCLSQAGREILIKSVLQSIPSYVMSIFLLPSTFIGEIEKMLNA 1177
FKF+ +RIW R+N W+ R LS+AG E ++K+V Q+IP+YVMS F +P + +++ +
Sbjct: 780 FKFLPERIWKRVNGWTDRPLSRAGMETMLKAVAQAIPNYVMSCFRIPVSICEKMKTCIAD 839
Query: 1178 FWWGHNSANSRGMHWLSWERLSVPKTFGGMGFKSLRAFNLAMIGKQAWKLISNPDSLITK 1237
WWG + MHW SW LS PK GGMGF+ FN AM+G+Q W+L+++PDSL ++
Sbjct: 840 HWWGFEDGKKK-MHWKSWSWLSTPKFLGGMGFREFTTFNQAMLGRQCWRLLTDPDSLCSR 898
Query: 1238 LLKAKYFPHSDYFSASIGHNPSYVWRSLWSVRDYLKHGLKWSIGTGETISVWNQPWLKDP 1297
+LK +YFP+S ++ A+ +PS+ WRSL R+ L G++W +G G+TI +++ W+
Sbjct: 899 VLKGRYFPNSSFWEAAQPKSPSFTWRSLLFGRELLAKGVRWGVGDGKTIKIFSDNWIPG- 957
Query: 1298 LCLQPVTEVQIMWDALTVGHLFKPNTKEWNENFIRYVFNAETSNQILQTPLLQSVQVDKA 1357
Q VT + TV L + + W+ + IR +F + + +ILQ P+ + D A
Sbjct: 958 FRPQLVTTLSPFPTDATVSCLMNEDARCWDGDLIRSLFPVDIAKEILQIPISRHGDADFA 1017
Query: 1358 TWRFEKNGLYSVRSAYREIINRNDVLLQHRVPG-------------KWNIIWNLKLPPKI 1404
+W +K GLYSVRSAY + R++ + W +W + P K+
Sbjct: 1018 SWPHDKLGLYSVRSAYN--LARSEAFFADQSNSGRGMASRLLESQKDWKGLWKINAPGKM 1075
Query: 1405 KNFLWRVCRNCLPTRMRLISKGVQCPPTCAICNNHEEDGKHLFFACSKSVGCWQRLGFWP 1464
K LWR CL T +L + + C C N ++ +H+F C + W+ +
Sbjct: 1076 KITLWRAAHECLATGFQLRRRHIPSTDGCVFC-NRDDTVEHVFLFCPFAAQIWEEIKGKC 1134
Query: 1465 SIQQIWSHNAFCADIIFSLLQQLDIAHQQIFAVTLWSLWRHRNNKVWNNIVETTEEIGDR 1524
+++ + + IF L++ + AVT W +W RNN NN + + +
Sbjct: 1135 AVKLGRNGFSTMRQWIFDFLKRGSSHANTLLAVTFWHIWEARNNTKNNNGTVHPQRVVIK 1194
Query: 1525 AVAFLNSWKAAQETRIRSSPANPHFDISKWSKPSVGRFKCNVDAAFSASLHRVGFGACIR 1584
+++++ + I +W P + N DAA +S +G GA IR
Sbjct: 1195 ILSYVDMILKHNTKTVDGQRGGNTQAIPRWQPPPASVWMINSDAAIFSSSRTMGVGALIR 1254
Query: 1585 DANGNHVISRTECFTPLLDVEMGEAIGLLHAMRWAKDLNLVNMDFETDSKVVVENIY-KG 1643
D G +++ +E + ++ E+ EA+ + A+ AK+ L ++ +D V+ I G
Sbjct: 1255 DNTGKCLVACSEMISDVVLPELAEALAIRRALGLAKEEGLEHIVMASDCLTVIRRIQTSG 1314
Query: 1644 DGVSDFMAIIHDCRHLLMTDLANSDVKFIRRQANSAAHNLAREALNHASFHYHLNIPHCI 1703
S +I D + L T + S + R +N AAH+LAR A Y IP I
Sbjct: 1315 RDRSGVGCVIEDIKKLASTFVLCS-FMHVNRLSNLAAHSLARNAELSTCTVYRSVIPDYI 1373
Query: 1704 HTLI 1707
++
Sbjct: 1374 RDIL 1377
>UniRef100_O22148 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1374
Score = 794 bits (2051), Expect = 0.0
Identities = 471/1374 (34%), Positives = 710/1374 (51%), Gaps = 63/1374 (4%)
Query: 354 MSFIVWNCRGLGSPSTVPTLKYLVRTYKPEGIFLSETMAALNKIEELKYILSFDSCFTVD 413
M + WNC+G+G+ TV L+ + Y PE IFL ET N +E + L F TV+
Sbjct: 1 MRILSWNCQGVGNTPTVRHLREIRGLYFPEVIFLCETKKRRNYLENVVGHLGFFDLHTVE 60
Query: 414 RIGRGGGLAFLWKNSANCTITNFSQNHIDVEVDDLLIGK---WRLTGFYGMPENGRRKES 470
IG+ GGLA +WK+S + + ID LLI + + LT YG P R E
Sbjct: 61 PIGKSGGLALMWKDSVQIKVLQSDKRLIDA----LLIWQDKEFYLTCIYGEPVQAERGEL 116
Query: 471 WAFLKNLARTSSLPWCVIGDFNDILSSEEKKGRSERAPWLIRGFRQAVLDAGLFDLHMTG 530
W L L + S PW + GDFN+++ EK G R FRQ + GL++++ +G
Sbjct: 117 WERLTRLGLSRSGPWMLTGDFNELVDPSEKIGGPARKESSCLEFRQMLNSCGLWEVNHSG 176
Query: 531 YAFTWFKSLGTARAVEEKLDRALVNSDWCNMFPNAGVECLSTTSSDHYPLWLTCKPVTAV 590
Y F+W+ + V+ +LDR + N W +FP A L SDH PL +
Sbjct: 177 YQFSWYGNRND-ELVQCRLDRTVANQAWMELFPQAKATYLQKICSDHSPL---INNLVGD 232
Query: 591 N-RNPHRFKFENAWLAEPDFKQQVQQRWKLYPEEG---ITQKLSYCAEDLTDWSRNNNNF 646
N R FK++ W+ FK + W + + +K++ C +++ W R
Sbjct: 233 NWRKWAGFKYDKRWVQREGFKDLLCNFWSQQSTKTNALMMEKIASCRREISKWKR----- 287
Query: 647 RRDISKVQK--KIEKLRTHVTAANVSY------FNSLKNKLDKLLVQDDLFWKQRAKTFW 698
+SK +I++L+ + AA LK +L + ++ FW+++++ W
Sbjct: 288 ---VSKPSSAVRIQELQFKLDAATKQIPFDRRELARLKKELSQEYNNEEQFWQEKSRIMW 344
Query: 699 YKEGDLNTKFFHAAATSRRKVNRIEHLENSNGTVCRKEDELQNIARNYFSHLFTKLS-ST 757
+ GD NTK+FHAA +RR NRI+ L + G +++L +A YF LF
Sbjct: 345 MRNGDRNTKYFHAATKNRRAQNRIQKLIDEEGREWTSDEDLGRVAEAYFKKLFASEDVGY 404
Query: 758 RADVISKVATSISDDDNCFLTAAFTLEEFKTAAFSMQADKCPGPDGFNPGFYQNFWNMCG 817
+ + + +SD N L A T EE + A FS+ KCPGPDG N YQ FW G
Sbjct: 405 TVEELENLTPLVSDQMNNNLLAPITKEEVQRATFSINPHKCPGPDGMNGFLYQQFWETMG 464
Query: 818 SEIHKAGCEWLENGVFPPHLNSTNIALIPKGDAQTTMKDWRPIALCNVVYKMVAKVLANR 877
+I + + +G +N TNI LIPK M D+RPI+LCNV+YK++ K++ANR
Sbjct: 465 DQITEMVQAFFRSGSIEEGMNKTNICLIPKILKAEKMTDFRPISLCNVIYKVIGKLMANR 524
Query: 878 LKNVLDKCISTSQSAFVPGRSILDNALVAIELIHYMKAKTKGSQGDVALKLDISKAYDRL 937
LK +L IS +Q+AFV GR I DN L+A EL+H + + K S+ +A+K DISKAYDR+
Sbjct: 525 LKKILPSLISETQAAFVKGRLISDNILIAHELLHALSSNNKCSEEFIAIKTDISKAYDRV 584
Query: 938 DWEYLRDIMIQMRFSEKWIRWIMLCVETVDYTVLVNGVQVGPLIPGRGIRQGDPLSPYLF 997
+W +L M + F++ WIR IM CV++V Y VL+NG G +IP RG+RQGDPLSPYLF
Sbjct: 585 EWPFLEKAMRGLGFADHWIRLIMECVKSVRYQVLINGTPHGEIIPSRGLRQGDPLSPYLF 644
Query: 998 ILCAEGLSALIRDAERRGVITGTRICTSAPAISHLLFADDCFLFFRASEQEASVMKNILT 1057
++C E L +++ AE++ ITG ++ AP ISHLLFADD + + +++ + I+
Sbjct: 645 VICTEMLVKMLQSAEQKNQITGLKVARGAPPISHLLFADDSMFYCKVNDEALGQIIRIIE 704
Query: 1058 TYEAASGQAINLQKSEMYCSRNTPTECQNRIATTLGVKQVLGTGKYLGLPSMIGRSKKAT 1117
Y ASGQ +N KS +Y ++ E + + LG+++ G G YLGLP SK AT
Sbjct: 705 EYSLASGQRVNYLKSSIYFGKHISEERRCLVKRKLGIEREGGEGVYLGLPESFQGSKVAT 764
Query: 1118 FKFVKDRIWNRINSWSSRCLSQAGREILIKSVLQSIPSYVMSIFLLPSTFIGEIEKMLNA 1177
++KDR+ ++ W S LS G+EIL+K+V ++P+Y MS F +P T +IE ++
Sbjct: 765 LSYLKDRLGKKVLGWQSNFLSPGGKEILLKAVAMALPTYTMSCFKIPKTICQQIESVMAE 824
Query: 1178 FWWGHNSANSRGMHWLSWERLSVPKTFGGMGFKSLRAFNLAMIGKQAWKLISNPDSLITK 1237
FWW N RG+HW +W LS PK GG+GFK + AFN+A++GKQ W++I+ DSL+ K
Sbjct: 825 FWW-KNKKEGRGLHWKAWCHLSRPKAVGGLGFKEIEAFNIALLGKQLWRMITEKDSLMAK 883
Query: 1238 LLKAKYFPHSDYFSASIGHNPSYVWRSLWSVRDYLKHGLKWSIGTGETISVWNQPWL--K 1295
+ K++YF SD +A +G PS+ W+S++ + +K G++ IG GETI+VW PW+ K
Sbjct: 884 VFKSRYFSKSDPLNAPLGSRPSFAWKSIYEAQVLIKQGIRAVIGNGETINVWTDPWIGAK 943
Query: 1296 DPLCLQPVTEVQIMWDAL-----TVGHLFKPNTKEWNENFIRYVFNAETSNQILQTPLLQ 1350
Q V ++ V L P+ ++WN N + +F T IL
Sbjct: 944 PAKAAQAVKRSHLVSQYAANSIHVVKDLLLPDGRDWNWNLVSLLFPDNTQENILALRPGG 1003
Query: 1351 SVQVDKATWRFEKNGLYSVRSAY---REIINRND---VLLQHRVPGKWNIIWNLKLPPKI 1404
D+ TW + ++G YSV+S Y EIIN+ + +LQ + + IW L +PPKI
Sbjct: 1004 KETRDRFTWEYSRSGHYSVKSGYWVMTEIINQRNNPQEVLQPSLDPIFQQIWKLDVPPKI 1063
Query: 1405 KNFLWRVCRNCLPTRMRLISKGVQCPPTCAICNNHEEDGKHLFFACSKSVGCWQRLGFWP 1464
+FLWR NCL L + + +C C +H E HL F C + W
Sbjct: 1064 HHFLWRCVNNCLSVASNLAYRHLAREKSCVRCPSHGETVNHLLFKCPFARLTWAISPLPA 1123
Query: 1465 SIQQIWSHNAF-----CADIIFSLLQQLDIAHQQIFAVTLWSLWRHRNNKVWNNIVETTE 1519
W+ + F + S ++ D H + LW LW++RN+ V+ T
Sbjct: 1124 PPGGEWAESLFRNMHHVLSVHKSQPEESD--HHALIPWILWRLWKNRNDLVFKGREFTAP 1181
Query: 1520 EIGDRAVAFLNSWKAAQETRIRSSPANPHFDISKWSKPSVGRFKCNVDAAFSASLHRVGF 1579
++ +A +++W +E + + + + + KW PS G KCN D A+S L G
Sbjct: 1182 QVILKATEDMDAWNNRKEPQPQVTSSTRDRCV-KWQPPSHGWVKCNTDGAWSKDLGNCGV 1240
Query: 1580 GACIRDANGNHVISRTECFTPLLDVEMGEAIGLLHAMRWA----KDLNLVNMDFETDSKV 1635
G +R+ G + V E + A+RWA N + FE+DS+
Sbjct: 1241 GWVLRNHTGRLLWLGLRALPSQQSVLETE----VEALRWAVLSLSRFNYRRVIFESDSQY 1296
Query: 1636 VVENIYKGDGVSDFMAIIHDCRHLLMTDLANSDVKFIRRQANSAAHNLAREALN 1689
+V I + I D R+LL +F RR+ N+ A ARE+L+
Sbjct: 1297 LVSLIQNEMDIPSLAPRIQDIRNLL-RHFEEVKFQFTRREGNNVADRTARESLS 1349
>UniRef100_Q9SIS9 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1715
Score = 749 bits (1934), Expect = 0.0
Identities = 457/1389 (32%), Positives = 690/1389 (48%), Gaps = 72/1389 (5%)
Query: 343 GGAWSPGPPRKMSFIVWNCRGLGSPSTVPTLKYLVRTYKPEGIFLSETMAALNKIEELKY 402
GGA + P ++ F WNC+GLG P TV L+ + R Y + +FL ET N +L
Sbjct: 354 GGAGTTTSPMRVGF--WNCQGLGQPLTVRRLEEVQRVYFLDMLFLIETKQQDNYTRDLGV 411
Query: 403 ILSFDSCFTVDRIGRGGGLAFLWKNSANCTITNFSQNHIDVEVDDLLIGKWRLTGFYGMP 462
+ F+ + G GGL WK + + + +D+ V+ + L+ YG P
Sbjct: 412 KMGFEDMCIISPRGLSGGLVVYWKKHLSIQVISHDVRLVDLYVEYKNFNFY-LSCIYGHP 470
Query: 463 ENGRRKESWAFLKNLARTSSLPWCVIGDFNDILSSEEKKGRSERAPWLIRGFRQAVLDAG 522
R W L+ ++ S PW + GDFN+IL+ EKKG R+ ++ F +
Sbjct: 471 IPSERHHLWEKLQRVSAHRSGPWMMCGDFNEILNLNEKKGGRRRSIGSLQNFTNMINCCN 530
Query: 523 LFDLHMTGYAFTWFKSLGTARAVEEKLDRALVNSDWCNMFPNAGVECLSTTSSDHYPLWL 582
+ DL G ++W +E LDR +NSDW FP E L SDH P+ +
Sbjct: 531 MKDLKSKGNPYSWVGKRQN-ETIESCLDRVFINSDWQASFPAFETEFLPIAGSDHAPVII 589
Query: 583 TCKPVTAVNRNPHRFKFENAWLAEPDFKQQVQQRWKLYPEE---GITQKLSYCAEDLTDW 639
R +F+++ DF VQ+ W + G +KL C ++L W
Sbjct: 590 DIAEEVCTKRG--QFRYDRRHFQFEDFVDSVQRGWNRGRSDSHGGYYEKLHCCRQELAKW 647
Query: 640 SRNNNNFRRDISKVQKKIEKLRTHVTAANVSY------FNSLKNKLDKLLVQDDLFWKQR 693
R R + +KIE L+ V AA + L+ L++ ++L+W +
Sbjct: 648 KR------RTKTNTAEKIETLKYRVDAAERDHTLPHQTILRLRQDLNQAYRDEELYWHLK 701
Query: 694 AKTFWYKEGDLNTKFFHAAATSRRKVNRIEHLENSNGTVCRKEDELQNIARNYFSHLFTK 753
++ W GD NT FF+A+ R+ NRI+ + ++ G ++D + +A NYF+ LFT
Sbjct: 702 SRNRWMLLGDRNTMFFYASTKLRKSRNRIKAITDAQGIENFRDDTIGKVAENYFADLFTT 761
Query: 754 L-SSTRADVISKVATSISDDDNCFLTAAFTLEEFKTAAFSMQADKCPGPDGFNPGFYQNF 812
+S ++IS +A +++ N L + T +E + A F++ AD+ PG DGF FY +
Sbjct: 762 TQTSDWEEIISGIAPKVTEQMNHELLQSVTDQEVRDAVFAIGADRAPGFDGFTAAFYHHL 821
Query: 813 WNMCGSEIHKAGCEWLENGVFPPHLNSTNIALIPKGDAQTTMKDWRPIALCNVVYKMVAK 872
W++ G+++ + E+ V +N T I LIPK M D+RPI+LC YK+++K
Sbjct: 822 WDLIGNDVCLMVRHFFESDVMDNQINQTQICLIPKIIDPKHMSDYRPISLCTASYKIISK 881
Query: 873 VLANRLKNVLDKCISTSQSAFVPGRSILDNALVAIELIHYMKAKTKGSQGDVALKLDISK 932
+L RLK L IS SQ+AFVPG++I DN LVA EL+H +K++ + G VA+K DISK
Sbjct: 882 ILIKRLKQCLGDVISDSQAAFVPGQNISDNVLVAHELLHSLKSRRECQSGYVAVKTDISK 941
Query: 933 AYDRLDWEYLRDIMIQMRFSEKWIRWIMLCVETVDYTVLVNGVQVGPLIPGRGIRQGDPL 992
AYDR++W +L +MIQ+ F+ +W++WIM CV +V Y VL+NG G + P RGIRQGDPL
Sbjct: 942 AYDRVEWNFLEKVMIQLGFAPRWVKWIMTCVTSVSYEVLINGSPYGKIFPSRGIRQGDPL 1001
Query: 993 SPYLFILCAEGLSALIRDAERRGVITGTRICTSAPAISHLLFADDCFLFFRASEQEASVM 1052
SPYLF+ CAE LS ++R AE I G +I AISHLLFADD F RAS Q +
Sbjct: 1002 SPYLFLFCAEVLSNMLRKAEVNKQIHGMKITKDCLAISHLLFADDSLFFCRASNQNIEQL 1061
Query: 1053 KNILTTYEAASGQAINLQKSEMYCSRNTPTECQNRIATTLGVKQVLGTGKYLGLPSMIGR 1112
I YE ASGQ IN KS + + PT + R+ LG+ V G GKYLGLP +GR
Sbjct: 1062 ALIFKKYEEASGQKINYAKSSIIFGQKIPTMRRQRLHRLLGIDNVRGGGKYLGLPEQLGR 1121
Query: 1113 SKKATFKFVKDRIWNRINSWSSRCLSQAGREILIKSVLQSIPSYVMSIFLLPSTFIGEIE 1172
K F+++ ++ R W+ LS AG+EI+IK++ ++P Y M+ FLLP+ EI
Sbjct: 1122 RKVELFEYIVTKVKERTEGWAYNYLSPAGKEIVIKAIAMALPVYSMNCFLLPTLICNEIN 1181
Query: 1173 KMLNAFWWGHNSANSRGMHWLSWERLSVPKTFGGMGFKSLRAFNLAMIGKQAWKLISNPD 1232
++ AFWWG + G +GFK L FN A++ KQAW++++NP
Sbjct: 1182 SLITAFWWGKENE-------------------GDLGFKDLHQFNRALLAKQAWRILTNPQ 1222
Query: 1233 SLITKLLKAKYFPHSDYFSASIGHNPSYVWRSLWSVRDYLKHGLKWSIGTGETISVWNQP 1292
SL+ +L K Y+P++ Y A+ G + SY W S+ + L+ GL+ +G G+T +W P
Sbjct: 1223 SLLARLYKGLYYPNTTYLRANKGGHASYGWNSIQEGKLLLQQGLRVRLGDGQTTKIWEDP 1282
Query: 1293 WLKDPLCLQPVTEVQIMWDALTVGHLFKPNTKEWNENFIRYVFNAETSNQILQTPLLQSV 1352
WL P I+ + + V L++ N +EW+ V N E L
Sbjct: 1283 WL--PTLPPRPARGPILDEDMKVADLWRENKREWDPVIFEGVLNPEDQQLAKSLYLSNYA 1340
Query: 1353 QVDKATWRFEKNGLYSVRSAY----------REIINRNDVLLQHRVPGKWNIIWNLKLPP 1402
D W + +N Y+VRS Y EIIN L+ VP K IW LK+ P
Sbjct: 1341 ARDSYKWAYTRNTQYTVRSGYWVATHVNLTEEEIINP----LEGDVPLKQE-IWRLKITP 1395
Query: 1403 KIKNFLWRVCRNCLPTRMRLISKGVQCPPTCAICNNHEEDGKHLFFACSKSVGCWQRLGF 1462
KIK+F+WR L T +L ++ + PTC C N +E H+ F CS + W+ F
Sbjct: 1396 KIKHFIWRCLSGALSTTTQLRNRNIPADPTCQRCCNADETINHIIFTCSYAQVVWRSANF 1455
Query: 1463 WPSIQQIWSHNAFCADIIFSLL-----QQLDIAHQQIFAVTLWSLWRHRNNKVWNNIVET 1517
S + ++ N + I +L Q L I + + +W LW+ RN ++ +
Sbjct: 1456 SGSNRLCFTDN--LEENIRLILQGKKNQNLPILNGLMPFWIMWRLWKSRNEYLFQQLDRF 1513
Query: 1518 TEEIGDRAVAFLNSW--KAAQETRIRSSPA----NPHFDISKWSKPSVGRFKCNVDAAFS 1571
++ +A W +T I + A P +WS P G KCN D+ +
Sbjct: 1514 PWKVAQKAEQEATEWVETMVNDTAISHNTAQSNDRPLSRSKQWSSPPEGFLKCNFDSGYV 1573
Query: 1572 ASLHRVGFGACIRDANGNHVISRTECFTPLLDVEMGEAIGLLHAMRWAKDLNLVNMDFET 1631
G +RD NG + S EA+G LHA++ + FE
Sbjct: 1574 QGRDYTSTGWILRDCNGRVLHSGCAKLQQSYSALQAEALGFLHALQMVWIRGYCYVWFEG 1633
Query: 1632 DSKVVVENIYKGDGVSDFMAIIHDCRHLLMTDLANSDVKFIRRQANSAAHNLAREALNHA 1691
D+ + I K + +++D R MT L S + ++ R+ N AA L + A + +
Sbjct: 1634 DNLELTNLINKTEDHHLLETLLYDIR-FWMTKLPFSSIGYVNRERNLAADKLTKYANSMS 1692
Query: 1692 SFHYHLNIP 1700
S + ++P
Sbjct: 1693 SLYETFHVP 1701
>UniRef100_Q7X970 BZIP-like protein [Oryza sativa]
Length = 2367
Score = 736 bits (1901), Expect = 0.0
Identities = 441/1336 (33%), Positives = 683/1336 (51%), Gaps = 66/1336 (4%)
Query: 393 ALNKIEELKYILSFDSCFTVDRIGRGGGLAFLWKNSANCTITNFSQNHIDVEVD-DLLIG 451
++ K+ L+ L V G GGLA W S + + + ++ +ID V
Sbjct: 769 SVEKMSRLRGRLGLRGFTGVSSEGMSGGLALYWDESVSVDVKDINKRYIDAYVQLSPEEP 828
Query: 452 KWRLTGFYGMPENGRRKESWAFLKNLARTSSLPWCVIGDFNDILSSEEKKGRSERAPWLI 511
+W +T YG P R W+ L+ + ++SSLPW VIGDFN+ + E R+ R +
Sbjct: 829 QWHVTFVYGEPRVENRHRMWSLLRTIHQSSSLPWAVIGDFNETMWQFEHFSRTPRGEPQM 888
Query: 512 RGFRQAVLDAGLFDLHMTGYAFTWFKSLGTARAVEEKLDRALVNSDWCNMFPNAGVECLS 571
+ FR + D L DL G T+ R V+ +LDR + + W +++ A V L
Sbjct: 889 QDFRDVLQDCELHDLGFKGVPHTYDNKREGWRNVKVRLDRVVADDKWRDIYSTAQVVHLV 948
Query: 572 TTSSDHYPLWLTCKPVTAVNRNPHRFK-----FENAWLAEPDFKQQVQQRWKLYPEEG-- 624
+ SDH P+ L V ++PH+ + +E W EP+ Q +++ W + E+
Sbjct: 949 SPCSDHCPILLNL-----VVKDPHQLRQKCLHYEIVWEREPEATQVIEEAWVVAGEKADL 1003
Query: 625 --ITQKLSYCAEDLTDWSRNN-NNFRRDISKVQKKIEKLRTHVTAANVSYFNSLKNKLDK 681
I + L+ L WSR N R++ K +KK+ +L + A+ + + + +++
Sbjct: 1004 GDINKALAKVMTALRSWSRAKVKNVGRELEKARKKLAELIE--SNADRTVIRNATDHMNE 1061
Query: 682 LLVQDDLFWKQRAKTFWYKEGDLNTKFFHAAATSRRKVNRIEHLENSNGTVCRKEDELQN 741
LL ++++ W QR++ W K+ D NTKFFH+ A R K N+I L ++N TV +L++
Sbjct: 1062 LLYREEMLWLQRSRVNWLKDEDRNTKFFHSRAVWRAKKNKISKLRDANETVHSSTMKLES 1121
Query: 742 IARNYFSHLFTKLSSTRADVISK-VATSISDDDNCFLTAAFTLEEFKTAAFSMQADKCPG 800
+A YF ++T + + +++ + ++D N L FT +E A F + K PG
Sbjct: 1122 MATEYFQDVYTADPNLNPETVTRLIQEKVTDIMNEKLCEDFTEDEISQAIFQIGPLKSPG 1181
Query: 801 PDGFNPGFYQNFWNMCGSEIHKAGCEWLENGVFPPHLNSTNIALIPKGDAQTTMKDWRPI 860
PDGF FYQ W ++I A + + G+ P +N T I LIPK + ++D+RPI
Sbjct: 1182 PDGFPARFYQRNWGTIKADIIGAVRRFFQTGLMPEGVNDTAIVLIPKKEQPVDLRDFRPI 1241
Query: 861 ALCNVVYKMVAKVLANRLKNVLDKCISTSQSAFVPGRSILDNALVAIELIHYMKAKTKGS 920
+LCNVVYK+V+K L NRL+ +LD +S QSAFV GR I DNAL+A E H M+ K +
Sbjct: 1242 SLCNVVYKVVSKCLVNRLRPILDDLVSVEQSAFVQGRMITDNALLAFECFHAMQKNKKAN 1301
Query: 921 QGDVALKLDISKAYDRLDWEYLRDIMIQMRFSEKWIRWIMLCVETVDYTVLVNGVQVGPL 980
A KLD+SKAYDR+DW +L M ++ F+ +W+ WIM CV +V Y V NG +
Sbjct: 1302 HAACAYKLDLSKAYDRVDWRFLEMAMNKLGFARRWVNWIMKCVTSVRYMVKFNGTLLQSF 1361
Query: 981 IPGRGIRQGDPLSPYLFILCAEGLSALIRDAERRGVITGTRICTSAPAISHLLFADDCFL 1040
P RG+RQGDPL P+LF+ A+GLS L+++ + +T ++C +AP ISHLLFADD L
Sbjct: 1362 APTRGLRQGDPLLPFLFLFVADGLSLLLKEKVAQNSLTPFKVCRAAPGISHLLFADDTLL 1421
Query: 1041 FFRASEQEASVMKNILTTYEAASGQAINLQKSEMYCSRNTPTECQNRIATTLGVKQVLGT 1100
FF+A ++EA V+K +L++Y +GQ IN K + + I+ L V++
Sbjct: 1422 FFKAHQREAEVVKEVLSSYAMGTGQLINPAKCSILMGGASTPAVSEAISEILQVERDRFE 1481
Query: 1101 GKYLGLPSMIGRSKKATFKFVKDRIWNRINSWSSRCLSQAGREILIKSVLQSIPSYVMSI 1160
+YLG P+ GR K F+ ++ +IW R+ W LS G+E+LIK+V+Q+IP YVM I
Sbjct: 1482 DRYLGFPTPEGRMHKGRFQSLQAKIWKRVIQWGENHLSTGGKEVLIKAVIQAIPVYVMGI 1541
Query: 1161 FLLPSTFIGEIEKMLNAFWWGHNSANSRGMHWLSWERLSVPKTFGGMGFKSLRAFNLAMI 1220
F LP + I ++ K+ FWW + R HW +W+ L+ PK+ GG+GF+ R FN A++
Sbjct: 1542 FKLPESVIDDLTKLTKNFWW-DSMNGQRKTHWKAWDSLTKPKSLGGLGFRDYRLFNQALL 1600
Query: 1221 GKQAWKLISNPDSLITKLLKAKYFPHSDYFSASIGHNPSYVWRSLWSVRDYLKHGLKWSI 1280
+QAW+LI+ PDSL ++LKAKYFPH S G N S WRS+ D LK G+ W +
Sbjct: 1601 ARQAWRLITYPDSLCARVLKAKYFPHGSLIDTSFGSNSSPAWRSIEYGLDLLKKGIIWRV 1660
Query: 1281 GTGETISVWNQPWLKDPLCLQPVT---EVQIMW--DALTVGHLFKPNTKEWNENFIRYVF 1335
G G +I +W WL +P+T ++ W D +T W+ I F
Sbjct: 1661 GNGNSIRIWRDSWLPRDHSRRPITGKANCRLKWVSDLIT-------EDGSWDVPKIHQYF 1713
Query: 1336 NAETSNQILQTPLLQSVQVDKATWRFEKNGLYSVRSAYR---EIINRNDVLLQ--HRVPG 1390
+ + IL + + D W +KNG++SVRSAYR +++N + + +
Sbjct: 1714 HNLDAEVILNICISSRSEEDFIAWHPDKNGMFSVRSAYRLAAQLVNIEESSSSGTNNINK 1773
Query: 1391 KWNIIWNLKLPPKIKNFLWRVCRNCLPTRMRLISKGVQCPPTCAICNNHEEDGKHLFFAC 1450
W +IW K+P K+K F WRV NCL T + + ++ C IC+ ED H C
Sbjct: 1774 AWEMIWKCKVPQKVKIFAWRVASNCLATMVNKKKRKLEQSDMCQICDRENEDDAHALCRC 1833
Query: 1451 SKSVGCWQRLGFWPSIQQIWSHNAFCADIIFSLLQQLDIAHQQIFAVTLWSLWRHRNNKV 1510
++ W + S+ + +F L+++ Q +F +TLW W RN +
Sbjct: 1834 IQASQLWSCMHKSGSVSVDIKASVLGRFWLFDCLEKIPEYEQAMFLMTLWRNWYVRNELI 1893
Query: 1511 WNNI---VETTEEIGDRAVAFLNSWKAAQETR-------IRSSP--ANPHFDISK----- 1553
ET++ V L + A + +R+ P P + +
Sbjct: 1894 HGKSAPPTETSQRFIQSYVDLLFQIRQAPQADLVKGKHVVRTVPLKGGPKYRVLNNHQPC 1953
Query: 1554 WSKPSVGRFKCNVDAAFSASLHRVGFGACIRDANGNHVISRTE----CFTPLLDVEMGEA 1609
W +P G K NVD +F AS + G G +R++ G+ + + + C PL
Sbjct: 1954 WERPKDGWMKLNVDGSFDASSGKGGLGMILRNSAGDIIFTSCKPLERCNNPLESELRACV 2013
Query: 1610 IGLLHAMRWAKDLNLVNMDFETDSKVVVENIYK-GDGVSDFMAIIHDCRHLLMTDLANSD 1668
GL A+ W L+ + ETD VV+ + G S IIH+ RHLL T +
Sbjct: 2014 EGLKLAIHW----TLLPIQVETDCASVVQLLQGIGRDFSVLANIIHEARHLLQTPSRTNT 2069
Query: 1669 VKFIRRQA---NSAAH 1681
++Q N+ AH
Sbjct: 2070 KVTTQKQTLRPNATAH 2085
>UniRef100_Q9SKJ4 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1750
Score = 734 bits (1894), Expect = 0.0
Identities = 451/1356 (33%), Positives = 694/1356 (50%), Gaps = 39/1356 (2%)
Query: 363 GLGSPSTVPTLKYLVRTYKPEGIFLSETMAALNKIEELKYILSFDSCFTVDRIGRGGGLA 422
G+G P T L L R Y + +FL ET+ +K+ +L Y L F + T GR GGLA
Sbjct: 391 GIGMPLTQSRLFRLFRMYNYDILFLVETLNQCDKVCKLAYDLGFPNVITQPPNGRSGGLA 450
Query: 423 FLWKNSANCTITNFSQNHIDVEVDDLLIGKWRLTGFYGMPENGRRKESWAFLKNLARTSS 482
+WKN+ + ++ + + ID V + L+ YG P R + W L++++ +
Sbjct: 451 LMWKNNVSLSLISQDERLIDSHVT-FNNKSFYLSCVYGHPTQSERHQLWQTLEHISDNRN 509
Query: 483 LPWCVIGDFNDILSSEEKKGRSERAPWLIRGFRQAVLDAGLFDLHMTGYAFTWFKSLGTA 542
W ++GDFN+ILS+ EK G R W R FR V + D+ G F+W T
Sbjct: 510 AEWLLVGDFNEILSNAEKIGGPMREEWTFRNFRNMVSHCDIEDMRSKGDRFSWVGERHT- 568
Query: 543 RAVEEKLDRALVNSDWCNMFPNAGVECLSTTSSDHYPLWLTCKPVTAVNRNPHRFKFENA 602
V+ LDR +NS W FP A +E L T SDH P+ + + R F+F+N
Sbjct: 569 HTVKCCLDRVFINSAWTATFPYAEIEFLDFTGSDHKPVLVHFNE--SFPRRSKLFRFDNR 626
Query: 603 WLAEPDFKQQVQQRWKLYPEEG---ITQKLSYCAEDLTDWSRNNN-NFRRDISKVQKKIE 658
+ P FK+ VQ W+ IT+++S C + + +N N + I K+Q +
Sbjct: 627 LIDIPTFKRIVQTSWRTNRNSRSTPITERISSCRQAMARLKHASNLNSEQRIKKLQSSLN 686
Query: 659 KLRTHVTAANVSYFNSLKNKLDKLLVQDDLFWKQRAKTFWYKEGDLNTKFFHAAATSRRK 718
+ + L+ L K ++++WKQ+++ W KEGD NT +FHA +R
Sbjct: 687 RAMESTRRVDRQLIPQLQESLAKAFSDEEIYWKQKSRNQWMKEGDQNTGYFHACTKTRYS 746
Query: 719 VNRIEHLENSNGTVCRKEDELQNIARNYFSHLFTKLSSTRADVISKV-----ATSISDDD 773
NR+ + + G + + E+ N A+++F+++F ST +S + +++++
Sbjct: 747 QNRVNTIMDDQGRMFTGDKEIGNHAQDFFTNIF----STNGIKVSPIDFADFKSTVTNTV 802
Query: 774 NCFLTAAFTLEEFKTAAFSMQADKCPGPDGFNPGFYQNFWNMCGSEIHKAGCEWLENGVF 833
N LT F+ E A + DK PGPDG FY+N W++ G ++ ++ E
Sbjct: 803 NLDLTKEFSDTEIYDAICQIGDDKAPGPDGLTARFYKNCWDIVGYDVILEVKKFFETSFM 862
Query: 834 PPHLNSTNIALIPKGDAQTTMKDWRPIALCNVVYKMVAKVLANRLKNVLDKCISTSQSAF 893
P +N TNI +IPK TT+ D+RPIALCNV+YK+++K L NRLK+ L+ +S SQ+AF
Sbjct: 863 KPSINHTNICMIPKITNPTTLSDYRPIALCNVLYKVISKCLVNRLKSHLNSIVSDSQAAF 922
Query: 894 VPGRSILDNALVAIELIHYMKAKTKGSQGDVALKLDISKAYDRLDWEYLRDIMIQMRFSE 953
+PGR I DN ++A E++H +K + + S+ +A+K D+SKAYDR++W++L M F
Sbjct: 923 IPGRIINDNVMIAHEVMHSLKVRKRVSKTYMAVKTDVSKAYDRVEWDFLETTMRLFGFCN 982
Query: 954 KWIRWIMLCVETVDYTVLVNGVQVGPLIPGRGIRQGDPLSPYLFILCAEGLSALIRDAER 1013
KWI WIM V++V Y+VL+NG G + P RGIRQGDPLSPYLFILC + LS LI
Sbjct: 983 KWIGWIMAAVKSVHYSVLINGSPHGYITPTRGIRQGDPLSPYLFILCGDILSHLINGRAS 1042
Query: 1014 RGVITGTRICTSAPAISHLLFADDCFLFFRASEQEASVMKNILTTYEAASGQAINLQKSE 1073
G + G RI APAI+HL FADD F +A+ + +K++ YE SGQ IN+QKS
Sbjct: 1043 SGDLRGVRIGNGAPAITHLQFADDSLFFCQANVRNCQALKDVFDVYEYYSGQKINVQKSM 1102
Query: 1074 MYCSRNTPTECQNRIATTLGVKQVLGTGKYLGLPSMIGRSKKATFKFVKDRIWNRINSWS 1133
+ Q+R+ L + G GKYLGLP GR KK F+++ DR+ R ++WS
Sbjct: 1103 ITFGSRVYGSTQSRLKQILEIPNQGGGGKYLGLPEQFGRKKKEMFEYIIDRVKKRTSTWS 1162
Query: 1134 SRCLSQAGREILIKSVLQSIPSYVMSIFLLPSTFIGEIEKMLNAFWWGHNSANSRGMHWL 1193
+R LS AG+EI++KSV ++P Y MS F LP + EIE +L FWW ++N RG+ W+
Sbjct: 1163 ARFLSPAGKEIMLKSVALAMPVYAMSCFKLPKGIVSEIESLLMNFWW-EKASNQRGIPWV 1221
Query: 1194 SWERLSVPKTFGGMGFKSLRAFNLAMIGKQAWKLISNPDSLITKLLKAKYFPHSDYFSAS 1253
+W+RL K GG+GF+ L FN A++ KQAW+LI P+SL +++KA+YF A
Sbjct: 1222 AWKRLQYSKKEGGLGFRDLAKFNDALLAKQAWRLIQYPNSLFARVMKARYFKDVSILDAK 1281
Query: 1254 IGHNPSYVWRSLWSVRDYLKHGLKWSIGTGETISVWNQPWLKDPLCLQPVTEVQIMWDAL 1313
+ SY W SL LK G + IG G+ I + + D +P+ + + +
Sbjct: 1282 VRKQQSYGWASLLDGIALLKKGTRHLIGDGQNIRI-GLDNIVDSHPPRPL-NTEETYKEM 1339
Query: 1314 TVGHLF--KPNTKEWNENFIRYVFNAETSNQILQTPLLQSVQVDKATWRFEKNGLYSVRS 1371
T+ +LF K + W+++ I + I + L +S + DK W + G Y+VRS
Sbjct: 1340 TINNLFERKGSYYFWDDSKISQFVDQSDHGFIHRIYLAKSKKPDKIIWNYNTTGEYTVRS 1399
Query: 1372 AYREIINRNDVLLQHRVPGKWNI-----IWNLKLPPKIKNFLWRVCRNCLPTRMRLISKG 1426
Y + + + P +I IWNL + PK+K+FLWR L T RL ++G
Sbjct: 1400 GYWLLTHDPSTNIPAINPPHGSIDLKTRIWNLPIMPKLKHFLWRALSQALATTERLTTRG 1459
Query: 1427 VQCPPTCAICNNHEEDGKHLFFACSKSVGCWQRLGFWPSIQQIWSHNAFCADI--IFSLL 1484
++ P+C C+ E H F C + W RL I+ N F +I I + +
Sbjct: 1460 MRIDPSCPRCHRENESINHALFTCPFATMAW-RLSDSSLIRNQLMSNDFEENISNILNFV 1518
Query: 1485 QQLDIA--HQQIFAVTLWSLWRHRNNKVWNNIVETTEEIGDRAVAFLNSW-KAAQETRIR 1541
Q ++ H+ + +W +W+ RNN V+N E+ + A A + W A Q +
Sbjct: 1519 QDTTMSDFHKLLPVWLIWRIWKARNNVVFNKFRESPSKTVLSAKAETHDWLNATQSHKKT 1578
Query: 1542 SSPANPHFDIS-KWSKPSVGRFKCNVDAAFSASLHRVGFGACIRDANGNHVISRTECFTP 1600
SP + +W P KCN DA F G IR+ G + +
Sbjct: 1579 PSPTRQIAENKIEWRNPPATYVKCNFDAGFDVQKLEATGGWIIRNHYGTPISWGSMKLAH 1638
Query: 1601 LLDVEMGEAIGLLHAMR--WAKDLNLVNMDFETDSKVVVENIYKGDGVSDFMAIIHDCRH 1658
+ E LL A++ W + V M+ + + + N+ G +A +
Sbjct: 1639 TSNPLEAETKALLAALQQTWIRGYTQVFMEGDCQTLI---NLINGISFHSSLANHLEDIS 1695
Query: 1659 LLMTDLANSDVKFIRRQANSAAHNLAREALNHASFH 1694
A+ FIR++ N AH LA+ +++F+
Sbjct: 1696 FWANKFASIQFGFIRKKGNKLAHVLAKYGCTYSTFY 1731
>UniRef100_Q7XJS2 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1344
Score = 719 bits (1855), Expect = 0.0
Identities = 416/1237 (33%), Positives = 653/1237 (52%), Gaps = 37/1237 (2%)
Query: 382 PEGIFLSETMAALNKIEELKYILSFDSCFTVDRIGRGGGLAFLWKNSANCTITNFSQNHI 441
P+ +FL ET + + + ++ L +D TV+ GR GGLA WK+ +N +
Sbjct: 7 PDILFLMETKNSQDFVYKVFCWLGYDFIHTVEPEGRSGGLAIFWKSHLEIEFLYADKNLM 66
Query: 442 DVEVDDLLIGKWRLTGFYGMPENGRRKESWAFLKNLARTSSLPWCVIGDFNDILSSEEKK 501
D++V W ++ YG+P R + W L ++ + WC+IGDFNDI S++EK
Sbjct: 67 DLQVSSRN-KVWFISCVYGLPVTHMRPKLWEHLNSIGLKRAEAWCLIGDFNDIRSNDEKL 125
Query: 502 GRSERAPWLIRGFRQAVLDAGLFDLHMTGYAFTWFKSLGTARAVEEKLDRALVNSDWCNM 561
G R+P + F +L+ + +L TG +FTW + + V+ KLDR N W ++
Sbjct: 126 GGPRRSPSSFQCFEHMLLNCSMHELGSTGNSFTWGGNRND-QWVQCKLDRCFGNPAWFSI 184
Query: 562 FPNAGVECLSTTSSDHYPLWLTCKPVTAVNRNPHRFKFENAWLAEPDFKQQVQQRWKLYP 621
FPNA L SDH P+ + + R +F+++ +P + + + W
Sbjct: 185 FPNAHQWFLEKFGSDHRPVLVKFTNDNELFRG--QFRYDKRLDDDPYCIEVIHRSWNSAM 242
Query: 622 EEGITQK---LSYCAEDLTDWSRNNNNFRRDISKVQKKIEKLRTHVTAANVSYF------ 672
+G L C ++ W +++ + Q +I++LR + A
Sbjct: 243 SQGTHSSFFSLIECRRAISVWKHSSD------TNAQSRIKRLRKDLDAEKSIQIPCWPRI 296
Query: 673 NSLKNKLDKLLVQDDLFWKQRAKTFWYKEGDLNTKFFHAAATSRRKVNRIEHLENSNGTV 732
+K++L ++LFW+Q+++ W GD NT FFHA S R N + L + N
Sbjct: 297 EYIKDQLSLAYGDEELFWRQKSRQKWLAGGDKNTGFFHATVHSERLKNELSFLLDENDQE 356
Query: 733 CRKEDELQNIARNYFSHLFTKLSS-TRADVISKVATSISDDDNCFLTAAFTLEEFKTAAF 791
+ + IA ++F +LFT T + + + ++ + N L T E A F
Sbjct: 357 FTRNSDKGKIASSFFENLFTSTYILTHNNHLEGLQAKVTSEMNHNLIQEVTELEVYNAVF 416
Query: 792 SMQADKCPGPDGFNPGFYQNFWNMCGSEIHKAGCEWLENGVFPPHLNSTNIALIPKGDAQ 851
S+ + PGPDGF F+Q W++ +I + E GV P N T+I LIPK +
Sbjct: 417 SINKESAPGPDGFTALFFQQHWDLVKHQILTEIFGFFETGVLPQDWNHTHICLIPKITSP 476
Query: 852 TTMKDWRPIALCNVVYKMVAKVLANRLKNVLDKCISTSQSAFVPGRSILDNALVAIELIH 911
M D RPI+LC+V+YK+++K+L RLK L +ST+QSAFVP R I DN LVA E+IH
Sbjct: 477 QRMSDLRPISLCSVLYKIISKILTQRLKKHLPAIVSTTQSAFVPQRLISDNILVAHEMIH 536
Query: 912 YMKAKTKGSQGDVALKLDISKAYDRLDWEYLRDIMIQMRFSEKWIRWIMLCVETVDYTVL 971
++ + S+ +A K D+SKAYDR++W +L +M + F+ KWI WIM CV +V Y+VL
Sbjct: 537 SLRTNDRISKEHMAFKTDMSKAYDRVEWPFLETMMTALGFNNKWISWIMNCVTSVSYSVL 596
Query: 972 VNGVQVGPLIPGRGIRQGDPLSPYLFILCAEGLSALIRDAERRGVITGTRICTSAPAISH 1031
+NG G +IP RGIRQGDPLSP LF+LC E L ++ AE+ G ITG + +++H
Sbjct: 597 INGQPYGHIIPTRGIRQGDPLSPALFVLCTEALIHILNKAEQAGKITGIQFQDKKVSVNH 656
Query: 1032 LLFADDCFLFFRASEQEASVMKNILTTYEAASGQAINLQKSEMYCSRNTPTECQNRIATT 1091
LLFADD L +A++QE + L+ Y SGQ INL KS + +N + ++ I +
Sbjct: 657 LLFADDTLLMCKATKQECEELMQCLSQYGQLSGQMINLNKSAITFGKNVDIQIKDWIKSR 716
Query: 1092 LGVKQVLGTGKYLGLPSMIGRSKKATFKFVKDRIWNRINSWSSRCLSQAGREILIKSVLQ 1151
G+ GTGKYLGLP + SK+ F F+K+++ +R+ W ++ LSQ G+E+L+KS+
Sbjct: 717 SGISLEGGTGKYLGLPECLSGSKRDLFGFIKEKLQSRLTGWYAKTLSQGGKEVLLKSIAL 776
Query: 1152 SIPSYVMSIFLLPSTFIGEIEKMLNAFWWGHNSANSRGMHWLSWERLSVPKTFGGMGFKS 1211
++P YVMS F LP ++ ++ FWW ++ R +HWLSW+RL++PK GG GFK
Sbjct: 777 ALPVYVMSCFKLPKNLCQKLTTVMMDFWW-NSMQQKRKIHWLSWQRLTLPKDQGGFGFKD 835
Query: 1212 LRAFNLAMIGKQAWKLISNPDSLITKLLKAKYFPHSDYFSASIGHNPSYVWRSLWSVRDY 1271
L+ FN A++ KQAW+++ SL +++ +++YF +SD+ SA+ G PSY WRS+ R+
Sbjct: 836 LQCFNQALLAKQAWRVLQEKGSLFSRVFQSRYFSNSDFLSATRGSRPSYAWRSILFGREL 895
Query: 1272 LKHGLKWSIGTGETISVWNQPWLKDPLCLQPVTEVQIMWDALTVGHLFKPNTKEWNENFI 1331
L GL+ IG G+ VW WL D +P+ + + L V L P ++ WN N +
Sbjct: 896 LMQGLRTVIGNGQKTFVWTDKWLHDGSNRRPLNRRRFINVDLKVSQLIDPTSRNWNLNML 955
Query: 1332 RYVFNAETSNQIL-QTPLLQSVQVDKATWRFEKNGLYSVRSAY----REIINR--NDVLL 1384
R +F + IL Q PL + D W NGLYSV++ Y +++ +R + +
Sbjct: 956 RDLFPWKDVEIILKQRPLF--FKEDSFCWLHSHNGLYSVKTGYEFLSKQVHHRLYQEAKV 1013
Query: 1385 QHRVPGKWNIIWNLKLPPKIKNFLWRVCRNCLPTRMRLISKGVQCPPTCAICNNHEEDGK 1444
+ V ++ IWNL PKI+ FLW+ +P RL ++G++ C +C+ E
Sbjct: 1014 KPSVNSLFDKIWNLHTAPKIRIFLWKALHGAIPVEDRLRTRGIRSDDGCLMCDTENETIN 1073
Query: 1445 HLFFACSKSVGCWQRLGFWPSIQQIWSHNAFC-ADIIFSLLQQLDIAHQQIFAV--TLWS 1501
H+ F C + W + S +S++ + + L QQ D+ H F LW
Sbjct: 1074 HILFECPLARQVW-AITHLSSAGSEFSNSVYTNMSRLIDLTQQNDLPHHLRFVSPWILWF 1132
Query: 1502 LWRHRNNKVWNNIVETTEEIGDRAVAFLNSWKAAQETRIRSSPANPHFDISKWSKPSVGR 1561
LW++RN ++ T + D+A + W +AQ T +++ H I+KW P G
Sbjct: 1133 LWKNRNALLFEGKGSITTTLVDKAYEAYHEWFSAQ-THMQND--EKHLKITKWCPPLPGE 1189
Query: 1562 FKCNVDAAFSASLHRVGFGACIRDANGNHVISRTECF 1598
KCN+ A+S H G +RD+ G ++ F
Sbjct: 1190 LKCNIGFAWSKQHHFSGASWVVRDSQGKVLLHSRRSF 1226
>UniRef100_Q9SIQ5 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1524
Score = 715 bits (1846), Expect = 0.0
Identities = 438/1334 (32%), Positives = 676/1334 (49%), Gaps = 47/1334 (3%)
Query: 389 ETMAALNKIEELKYILSFDSCFTVDRIGRGGGLAFLWKNSANCTITNFSQNHIDVEVDDL 448
ET+ +K+ +L Y L F + T GR GGLA +WKN+ + ++ + + ID V
Sbjct: 191 ETLNQCDKVCKLAYDLGFPNVITQPPNGRSGGLALMWKNNVSLSLISQDERLIDSHVT-F 249
Query: 449 LIGKWRLTGFYGMPENGRRKESWAFLKNLARTSSLPWCVIGDFNDILSSEEKKGRSERAP 508
+ L+ YG P R + W L++++ + W ++GDFN+ILS+ EK G R
Sbjct: 250 NNKSFYLSCVYGHPTQSERHQLWQTLEHISDNRNAEWLLVGDFNEILSNAEKIGGPMREE 309
Query: 509 WLIRGFRQAVLDAGLFDLHMTGYAFTWFKSLGTARAVEEKLDRALVNSDWCNMFPNAGVE 568
W R FR V + D+ G F+W T V+ LDR +NS W FP A E
Sbjct: 310 WTFRNFRNMVSHCDIEDMRSKGDRFSWVGERHT-HTVKCCLDRVFINSAWTATFPYAETE 368
Query: 569 CLSTTSSDHYPLWLTCKPVTAVNRNPHRFKFENAWLAEPDFKQQVQQRWKLYPEEG---I 625
L T SDH P+ + + R F+F+N + P FK+ VQ W+ I
Sbjct: 369 FLDFTGSDHKPVLVHFNE--SFPRRSKLFRFDNRLIDIPTFKRIVQTSWRTNRNSRSTPI 426
Query: 626 TQKLSYCAEDLTDWSRNNN-NFRRDISKVQKKIEKLRTHVTAANVSYFNSLKNKLDKLLV 684
T+++S C + + +N N + I K+Q + + + L+ L K
Sbjct: 427 TERISSCRQAMARLKHASNLNSEQRIKKLQSSLNRAMESTRRVDRQLIPQLQESLAKAFS 486
Query: 685 QDDLFWKQRAKTFWYKEGDLNTKFFHAAATSRRKVNRIEHLENSNGTVCRKEDELQNIAR 744
++++WKQ+++ W KEGD NT +FHA +R NR+ + + G + + E+ N A+
Sbjct: 487 DEEIYWKQKSRNQWMKEGDQNTGYFHACTKTRYSQNRVNTIMDDQGRMFTGDKEIGNHAQ 546
Query: 745 NYFSHLFTKLSSTRADVISKV-----ATSISDDDNCFLTAAFTLEEFKTAAFSMQADKCP 799
++F+++F ST +S + +++++ N LT F+ E A + DK P
Sbjct: 547 DFFTNIF----STNGIKVSPIDFADFKSTVTNTVNLDLTKEFSDTEIYDAICQIGDDKAP 602
Query: 800 GPDGFNPGFYQNFWNMCGSEIHKAGCEWLENGVFPPHLNSTNIALIPKGDAQTTMKDWRP 859
GPDG FY+N W++ G ++ ++ E P +N TNI +IPK TT+ D+RP
Sbjct: 603 GPDGLTARFYKNCWDIVGYDVILEVKKFFETSFMKPSINHTNICMIPKITNPTTLSDYRP 662
Query: 860 IALCNVVYKMVAKVLANRLKNVLDKCISTSQSAFVPGRSILDNALVAIELIHYMKAKTKG 919
IALCNV+YK+++K L NRLK+ L+ +S SQ+AF+PGR I DN ++A E++H +K + +
Sbjct: 663 IALCNVLYKVISKCLVNRLKSHLNSIVSDSQAAFIPGRIINDNVMIAHEVMHSLKVRKRV 722
Query: 920 SQGDVALKLDISKAYDRLDWEYLRDIMIQMRFSEKWIRWIMLCVETVDYTVLVNGVQVGP 979
S+ +A+K D+SKAYDR++W++L M F KWI WIM V++V Y+VL+NG G
Sbjct: 723 SKTYMAVKTDVSKAYDRVEWDFLETTMRLFGFCNKWIGWIMAAVKSVHYSVLINGSPHGY 782
Query: 980 LIPGRGIRQGDPLSPYLFILCAEGLSALIRDAERRGVITGTRICTSAPAISHLLFADDCF 1039
+ P RGIRQGDPLSPYLFILC + LS LI G + G RI APAI+HL FADD
Sbjct: 783 ITPTRGIRQGDPLSPYLFILCGDILSHLINGRASSGDLRGVRIGNGAPAITHLQFADDSL 842
Query: 1040 LFFRASEQEASVMKNILTTYEAASGQAINLQKSEMYCSRNTPTECQNRIATTLGVKQVLG 1099
F +A+ + +K++ YE SGQ IN+QKS + Q+++ L + G
Sbjct: 843 FFCQANVRNCQALKDVFDVYEYYSGQKINVQKSMITFGSRVYGSTQSKLKQILEIPNQGG 902
Query: 1100 TGKYLGLPSMIGRSKKATFKFVKDRIWNRINSWSSRCLSQAGREILIKSVLQSIPSYVMS 1159
GKYLGLP GR KK F+++ DR+ R ++WS+R LS AG+EI++KSV ++P Y MS
Sbjct: 903 GGKYLGLPEQFGRKKKEMFEYIIDRVKKRTSTWSARFLSPAGKEIMLKSVALAMPVYAMS 962
Query: 1160 IFLLPSTFIGEIEKMLNAFWWGHNSANSRGMHWLSWERLSVPKTFGGMGFKSLRAFNLAM 1219
F LP + EIE +L FWW ++N RG+ W++W+RL K GG+GF+ L FN A+
Sbjct: 963 CFKLPKGIVSEIESLLMNFWW-EKASNQRGIPWVAWKRLQYSKKEGGLGFRDLAKFNDAL 1021
Query: 1220 IGKQAWKLISNPDSLITKLLKAKYFPHSDYFSASIGHNPSYVWRSLWSVRDYLKHGLKWS 1279
+ KQAW+LI P+SL +++KA+YF A + SY W SL LK G +
Sbjct: 1022 LAKQAWRLIQYPNSLFARVMKARYFKDVSILDAKVRKQQSYGWASLLDGIALLKKGTRHL 1081
Query: 1280 IGTGETISVWNQPWLKDPLCLQPVTEVQIMWDALTVGHLF--KPNTKEWNENFIRYVFNA 1337
IG G+ I + + D +P+ + + +T+ +LF K + W+++ I +
Sbjct: 1082 IGDGQNIRI-GLDNIVDSHPPRPL-NTEETYKEMTINNLFERKGSYYFWDDSKISQFVDQ 1139
Query: 1338 ETSNQILQTPLLQSVQVDKATWRFEKNGLYSVRSAYREIINRNDVLLQHRVPGKWNI--- 1394
I + L +S + DK W + G Y+VRS Y + + + P +I
Sbjct: 1140 SDHGFIHRIYLAKSKKPDKIIWNYNTTGEYTVRSGYWLLTHDPSTNIPAINPPHGSIDLK 1199
Query: 1395 --IWNLKLPPKIKNFLWRVCRNCLPTRMRLISKGVQCPPTCAICNNHEEDGKHLFFACSK 1452
IWNL + PK+K+FLWR L T RL ++G++ P C C+ E H F C
Sbjct: 1200 TRIWNLPIMPKLKHFLWRALSQALATTERLTTRGMRIDPICPRCHRENESINHALFTCPF 1259
Query: 1453 SVGCWQRLGFWPSIQQIWSHNAFCADI------IFSLLQQLDIA--HQQIFAVTLWSLWR 1504
+ W W S + + D I + +Q ++ H+ + +W +W+
Sbjct: 1260 ATMAW-----WLSDSSLIRNQLMSNDFEENISNILNFVQDTTMSDFHKLLPVWLIWRIWK 1314
Query: 1505 HRNNKVWNNIVETTEEIGDRAVAFLNSW-KAAQETRIRSSPANPHFDIS-KWSKPSVGRF 1562
RNN V+N E+ + A A + W A Q + SP + +W P
Sbjct: 1315 ARNNVVFNKFRESPSKTVLSAKAETHDWLNATQSHKKTPSPTRQIAENKIEWRNPPATYV 1374
Query: 1563 KCNVDAAFSASLHRVGFGACIRDANGNHVISRTECFTPLLDVEMGEAIGLLHAMR--WAK 1620
KCN DA F G IR+ G + + + E LL A++ W +
Sbjct: 1375 KCNFDAGFDVQKLEATGGWIIRNHYGTPISWGSMKLAHTSNPLEAETKALLAALQQTWIR 1434
Query: 1621 DLNLVNMDFETDSKVVVENIYKGDGVSDFMAIIHDCRHLLMTDLANSDVKFIRRQANSAA 1680
V M+ + + + N+ G +A + A+ FIRR+ N A
Sbjct: 1435 GYTQVFMEGDCQTLI---NLINGISFHSSLANHLEDISFWANKFASIQFGFIRRKGNKLA 1491
Query: 1681 HNLAREALNHASFH 1694
H LA+ +++F+
Sbjct: 1492 HVLAKYGCTYSTFY 1505
>UniRef100_Q75M12 Hypothetical protein P0676G05.2 [Oryza sativa]
Length = 1936
Score = 710 bits (1833), Expect = 0.0
Identities = 396/1134 (34%), Positives = 603/1134 (52%), Gaps = 29/1134 (2%)
Query: 345 AWSPGPPRKMSFIVWNCRGLGSPSTVPTLKYLVRTYKPEGIFLSETMAALNKIEELKYIL 404
A SPG MS + WNCRGLG+ +TV L+ L++ + +FL ET ++ K+ L+ L
Sbjct: 627 ATSPGAAGAMSCLAWNCRGLGNTATVQDLRALIQKAGSQLVFLCETRQSVEKMSRLRRKL 686
Query: 405 SFDSCFTVDRIGRGGGLAFLWKNSANCTITNFSQNHIDVEV----DDLLIGKWRLTGFYG 460
+F V G+ GGLA W S + + + ++ +ID V D+ +W +T YG
Sbjct: 687 AFRGFVGVSSEGKSGGLALYWDESVSVDVKDINKRYIDAYVRLSPDE---PQWHITFVYG 743
Query: 461 MPENGRRKESWAFLKNLARTSSLPWCVIGDFNDILSSEEKKGRSERAPWLIRGFRQAVLD 520
P R W+ L+ + ++S+LPW VIGDFN+ L E ++ R ++ FR A+ D
Sbjct: 744 EPRVENRHRMWSLLRTIRQSSALPWMVIGDFNETLWQFEHFSKNPRCETQMQNFRDALYD 803
Query: 521 AGLFDLHMTGYAFTWFKSLGTARAVEEKLDRALVNSDWCNMFPNAGVECLSTTSSDHYPL 580
L DL G T+ R V+ +LDRA+ + W ++FP A V L + SDH P+
Sbjct: 804 CDLQDLGFKGVPHTYDNRRDGWRNVKVRLDRAVADDKWRDLFPEAQVSHLVSPCSDHSPI 863
Query: 581 WLTCKPVTAVNRNPHRFKFENAWLAEPDFKQQVQQRW---KLYPEEG-ITQKLSYCAEDL 636
L +E W EP+ Q +++ W + + G I L L
Sbjct: 864 LLEFIVKDTTRPRQKCLHYEIVWEREPESVQVIEEAWINAGVKTDLGDINIALGRVMSAL 923
Query: 637 TDWSRNN-NNFRRDISKVQKKIEKLRTHVTAANVSYFNSLKNKLDKLLVQDDLFWKQRAK 695
WS+ N +++ K +KK+E L A S + ++++L ++++ W QR++
Sbjct: 924 RSWSKTKVKNVGKELEKARKKLEDLIASNAAR--SSIRQATDHMNEMLYREEMLWLQRSR 981
Query: 696 TFWYKEGDLNTKFFHAAATSRRKVNRIEHLENSNGTVCRKEDELQNIARNYFSHLFTKLS 755
W KEGD NT+FFH+ A R K N+I L + NG + L+ +A YF ++
Sbjct: 982 VNWLKEGDRNTRFFHSRAVWRAKKNKISKLRDENGAIHSTTSVLETMATEYFQGVYKADP 1041
Query: 756 STRADVISKV-ATSISDDDNCFLTAAFTLEEFKTAAFSMQADKCPGPDGFNPGFYQNFWN 814
S + ++++ ++D N L F EE A F + K P PDGF FYQ W
Sbjct: 1042 SLNPESVTRLFQEKVTDAMNEKLCQEFKEEEIAQAIFQIGPLKSPRPDGFPARFYQRNWG 1101
Query: 815 MCGSEIHKAGCEWLENGVFPPHLNSTNIALIPKGDAQTTMKDWRPIALCNVVYKMVAKVL 874
S+I A + ++G+ P +N T I LIPK D +KD+RPI+LCNVVYK+V+K L
Sbjct: 1102 TLKSDIILAVRNFFQSGLMPKGVNDTAIVLIPKKDQPIDLKDYRPISLCNVVYKVVSKCL 1161
Query: 875 ANRLKNVLDKCISTSQSAFVPGRSILDNALVAIELIHYMKAKTKGSQGDVALKLDISKAY 934
NRL+ +LD +S QSAF+ GR I DNAL+A E H ++ K + A KLD+SKAY
Sbjct: 1162 VNRLRPILDDLVSKEQSAFIQGRMITDNALLAFECFHSIQKNKKANSAACAYKLDLSKAY 1221
Query: 935 DRLDWEYLRDIMIQMRFSEKWIRWIMLCVETVDYTVLVNGVQVGPLIPGRGIRQGDPLSP 994
DR+DW +L + ++ F+ +W+ WIMLCV TV Y+V NG + P RG+RQG+PLSP
Sbjct: 1222 DRVDWRFLELALNKLGFAHRWVSWIMLCVTTVRYSVKFNGTLLRSFAPTRGLRQGEPLSP 1281
Query: 995 YLFILCAEGLSALIRDAERRGVITGTRICTSAPAISHLLFADDCFLFFRASEQEASVMKN 1054
+LF+ A+GLS L+++ + +T +IC AP IS+LLFADD LFF+A ++EA V+K
Sbjct: 1282 FLFLFVADGLSLLLKEKVAQNSLTPLKICRQAPGISYLLFADDTLLFFKAEKKEAEVVKE 1341
Query: 1055 ILTTYEAASGQAINLQKSEMYCSRNTPTECQNRIATTLGVKQVLGTGKYLGLPSMIGRSK 1114
+LT Y +GQ IN K + +P+ I TL V++ +YLG P+ GR
Sbjct: 1342 VLTNYAQGTGQLINPAKCSILFGEASPSSVSEDIRNTLQVERDNFEDRYLGFPTPEGRMH 1401
Query: 1115 KATFKFVKDRIWNRINSWSSRCLSQAGREILIKSVLQSIPSYVMSIFLLPSTFIGEIEKM 1174
K F+ ++ +I R+ W LS G+EILIK+V+Q+IP YVM +F P + E+ KM
Sbjct: 1402 KGRFQSLQAKIAKRVIQWGENFLSSGGKEILIKAVIQAIPVYVMGLFKFPDSVYDELTKM 1461
Query: 1175 LNAFWWGHNSANSRGMHWLSWERLSVPKTFGGMGFKSLRAFNLAMIGKQAWKLISNPDSL 1234
FWWG ++ R HW +W+ L+ K GG+GF+ + FN A++ +QAW+LI P+SL
Sbjct: 1462 TRNFWWGADNGRRR-THWRAWDSLTKAKINGGLGFRDYKLFNQALLTRQAWRLIEFPNSL 1520
Query: 1235 ITKLLKAKYFPHSDYFSASIGHNPSYVWRSLWSVRDYLKHGLKWSIGTGETISVWNQPWL 1294
++LKAKYFPH + N S W + D LK G+ W IG G ++ +W PW+
Sbjct: 1521 CAQVLKAKYFPHGSLTDTTFSANASPTWHGIEYGLDLLKKGIIWRIGNGNSVRIWRDPWI 1580
Query: 1295 KDPLCLQPVT---EVQIMWDALTVGHLFKPNTKEWNENFIRYVFNAETSNQILQTPLLQS 1351
L +PV+ ++ W + + ++ + N+ F++ +A+ +I + L+
Sbjct: 1581 PRDLSRRPVSSKANCRLKWVSDLIAEDGTWDSAKINQYFLK--IDADIIQKICISARLEE 1638
Query: 1352 VQVDKATWRFEKNGLYSVRSAYREIINRNDV-----LLQHRVPGKWNIIWNLKLPPKIKN 1406
D W +K G +SVRSAY+ + D+ R+ W +IW +P K++
Sbjct: 1639 ---DFIAWHPDKTGRFSVRSAYKLALQLADMNNCSSSSSSRLNKSWELIWKCNVPQKVRI 1695
Query: 1407 FLWRVCRNCLPTRMRLISKGVQCPPTCAICNNHEEDGKHLFFACSKSVGCWQRL 1460
F WRV N L T + ++ C IC+ +ED H C + W L
Sbjct: 1696 FAWRVASNSLATMENKKKRNLERFDVCGICDREKEDAGHALCRCVHANSLWVNL 1749
Score = 48.1 bits (113), Expect = 0.002
Identities = 39/146 (26%), Positives = 63/146 (42%), Gaps = 10/146 (6%)
Query: 1547 PHFDISKWSKPSVGRFKCNVDAAFSASLHRVGFGACIRDANGNHVISR----TECFTPLL 1602
P + +W +P G K NVD +F + + G G +R+ GN + S C P L
Sbjct: 1770 PGAENRRWERPRNGWMKLNVDGSFDINSEKGGIGMILRNCLGNVIFSSCRSLDSCSGP-L 1828
Query: 1603 DVEMGEAIGLLH-AMRWAKDLNLVNMDFETDSKVVVENIYKGDGVSDFMAIIHDCRHLLM 1661
+ E+ + LH A+ W L+ + ETD V++ + D +A I LM
Sbjct: 1829 EAELHACVEGLHLALHW----TLLPIQVETDCSSVIQLLNHPDKDRSVLANIAQEAKSLM 1884
Query: 1662 TDLANSDVKFIRRQANSAAHNLAREA 1687
+ ++R N +H LA +A
Sbjct: 1885 AGDRQIAISKVQRSQNVISHFLANKA 1910
>UniRef100_O81630 F8M12.22 protein [Arabidopsis thaliana]
Length = 1662
Score = 702 bits (1812), Expect = 0.0
Identities = 434/1356 (32%), Positives = 657/1356 (48%), Gaps = 120/1356 (8%)
Query: 342 NGGAWSPGPPRKMSFIVWNCRGLGSPSTVPTLKYLVRTYKPEGIFLSETMAALNKIEELK 401
+GGA SPG +G P T L L + +K + +FL ET+ I L
Sbjct: 384 DGGAGSPG--------------IGVPLTQSQLSNLCKVFKFDVLFLIETLNKCEVISNLA 429
Query: 402 YILSFDSCFTVDRIGRGGGLAFLWKNSANCTITNFSQNHIDVEVDDLLIGKWRLTGFYGM 461
+L F + T G GGLA LWK+S + HIDV + I + L+ YG
Sbjct: 430 SVLGFPNVITQPPQGHSGGLALLWKDSVRLSNLYQDDRHIDVHISINNINFY-LSRVYGH 488
Query: 462 PENGRRKESWAFLKNLARTSSLPWCVIGDFNDILSSEEKKGRSERAPWLIRGFRQAVLDA 521
P R W +NL++T + PW +IGDFN+ILS+ EK G +R W RGFR V
Sbjct: 489 PCQSERHSLWTHFENLSKTRNDPWILIGDFNEILSNNEKIGGPQRDEWTFRGFRNMVSTC 548
Query: 522 GLFDLHMTGYAFTWFKSLGTARAVEEKLDRALVNSDWCNMFPNAGVECLSTTSSDHYPLW 581
L D+ G F+W + V+ LDRA +NS+ +FP A +E L T SDH PL+
Sbjct: 549 DLKDIRSIGDRFSWVGERHS-HTVKCCLDRAFINSEGAFLFPFAELEFLEFTGSDHKPLF 607
Query: 582 LTCKPVTAVNRNPHRFKFENAWLAEPDFKQQVQQRWKLY---PEEGITQKLSYCAEDLTD 638
L+ + P F+F+ L P FK V+ W + + ++ C + +
Sbjct: 608 LSLEKTETRKMRP--FRFDKRLLEVPHFKTYVKAGWNKAINGQRKHLPDQVRTCRQAMAK 665
Query: 639 WSRNNN-NFRRDISKVQKKIEKLRTHVTAANVSYFNSLKNKLDKLLVQDDLFWKQRAKTF 697
+N N R I+++Q ++K + V + ++ +L ++ +W+Q+++
Sbjct: 666 LKHKSNLNSRIRINQLQAALDKAMSSVNRTERRTISHIQRELTVAYRDEERYWQQKSRNQ 725
Query: 698 WYKEGDLNTKFFHAAATSRRKVNRIEHLENSNGTVCRKEDELQNIARNYFSHLFTKLSST 757
W KEGD NT+FFHA +R VNR+ +++ G + R + E+ A+ +F+ ++
Sbjct: 726 WMKEGDRNTEFFHACTKTRFSVNRLVTIKDEEGMIYRGDKEIGVHAQEFFTKVYESNGRP 785
Query: 758 RADV----ISKVATSISDDDNCFLTAAFTLEEFKTAAFSMQADKCPGPDGFNPGFYQNFW 813
+ + + T +DD LT + E A + DK PGPDG FY++ W
Sbjct: 786 VSIIDFAGFKPIVTEQINDD---LTKDLSDLEIYNAICHIGDDKAPGPDGLTARFYKSCW 842
Query: 814 NMCGSEIHKAGCEWLENGVFPPHLNSTNIALIPKGDAQTTMKDWRPIALCNVVYKMVAKV 873
+ G ++ K + +N TNI +IPK T+ D+RPIALCNV+YK+++K
Sbjct: 843 EIVGPDVIKEVKIFFRTSYMKQSINHTNICMIPKITNPETLSDYRPIALCNVLYKIISKC 902
Query: 874 LANRLKNVLDKCISTSQSAFVPGRSILDNALVAIELIHYMKAKTKGSQGDVALKLDISKA 933
L RLK LD +S SQ+AF+PGR + DN ++A E++H +K + + SQ +A+K D+SKA
Sbjct: 903 LVERLKGHLDAIVSDSQAAFIPGRLVNDNVMIAHEMMHSLKTRKRVSQSYMAVKTDVSKA 962
Query: 934 YDRLDWEYLRDIMIQMRFSEKWIRWIMLCVETVDYTVLVNGVQVGPLIPGRGIRQGDPLS 993
YDR++W +L M FSE WI+WIM V++V+Y+VLVNG+ G + P RGIRQGDPLS
Sbjct: 963 YDRVEWNFLETTMRLFGFSETWIKWIMGAVKSVNYSVLVNGIPHGTIQPQRGIRQGDPLS 1022
Query: 994 PYLFILCAEGLSALIRDAERRGVITGTRICTSAPAISHLLFADDCFLFFRASEQEASVMK 1053
PYLFILCA+ L+ LI++ G I G RI P ++HL FADD F +++ + +K
Sbjct: 1023 PYLFILCADILNHLIKNRVAEGDIRGIRIGNGVPGVTHLQFADDSLFFCQSNVRNCQALK 1082
Query: 1054 NILTTYEAASGQAINLQKSEMYCSRNTPTECQNRIATTLGVKQVLGTGKYLGLPSMIGRS 1113
++ YE SGQ IN+ KS + QNR+ LG++ G GKYLGLP GR
Sbjct: 1083 DVFDVYEYYSGQKINMSKSMITFGSRVHGTTQNRLKNILGIQSHGGGGKYLGLPEQFGRK 1142
Query: 1114 KKATFKFVKDRIWNRINSWSSRCLSQAGREILIKSVLQSIPSYVMSIFLLPSTFIGEIEK 1173
K+ F ++ +R+ R +SWS++ LS AG+EI++KSV S+P Y MS F LP + EIE
Sbjct: 1143 KRDMFNYIIERVKKRTSSWSAKYLSPAGKEIMLKSVAMSMPVYAMSCFKLPLNIVSEIEA 1202
Query: 1174 MLNAFWWGHNSANSRGMHWLSWERLSVPKTFGGMGFKSLRAFNLAMIGKQAWKLISNPDS 1233
+L FWW N A R + W++W+RL K GG+GF+ L FN A++ KQ W++I+NP+S
Sbjct: 1203 LLMNFWWEKN-AKKREIPWIAWKRLQYSKKEGGLGFRDLAKFNDALLAKQVWRMINNPNS 1261
Query: 1234 LITKLLKAKYFPHSDYFSASIGHNPSYVWRSLWSVRDYLKHGLKWSIGTGETISVWNQPW 1293
L +++KA+YF A SY W S+ + D +K G ++ +G G+T S
Sbjct: 1262 LFARIMKARYFREDSILDAKRQRYQSYGWTSMLAGLDVIKKGSRFIVGDGKTGSY----- 1316
Query: 1294 LKDPLCLQPVTEVQIMWDALTVGHLFKPNTKEWNENFIRYVFNAETSNQILQTPLLQSVQ 1353
+ WN + I + + + ++ L + V
Sbjct: 1317 ------------------------------RYWNAHLISQLVSPDDHRFVMNHHLSRIVH 1346
Query: 1354 VDKATWRFEKNGLYSVRSAYREIINRNDVLLQHRVPGKWNIIWNLKLPPKIKNFLWRVCR 1413
DK W + +G Y+ +W L + PKIK LWR
Sbjct: 1347 QDKLVWNYSSSGDYT--------------------------LWKLPIIPKIKYMLWRTIS 1380
Query: 1414 NCLPTRMRLISKGVQCPPTCAICNNHEEDGKHLFFACSKSVGCWQRLGF-WPSIQQIWSH 1472
LPTR RL+++G+ P C C EE H+ F C + W F W
Sbjct: 1381 KALPTRSRLLTRGMDIDPHCPRCPTEEETINHVLFTCPYAASIWGLSNFPWLPGHTFSQD 1440
Query: 1473 NAFCADIIFSLLQQLDIAHQQIFAV--TLWSLWRHRNNKVWNNIVETTEEIGDRAVAFLN 1530
+ + + +Q A +W LW+ RNN V+N E+ + + A +N
Sbjct: 1441 TEENISFLINSFSNNTLNTEQRLAPFWLIWRLWKARNNLVFNKFSESCSRVVTQTEAEVN 1500
Query: 1531 SWKAAQETRIRSS--PANPHFDISKWSKPSVGRFKCNVDAAFSASLHRVGFGACIRDANG 1588
W + R +S + H +W KP KCN DA F+ G
Sbjct: 1501 EWLQSVNRREDASVLTRSSHRANVRWKKPVFPLVKCNFDAGFT----------------G 1544
Query: 1589 NHVISRTECFTPLLDVEMGEAIGLLHAMRWAKDLNLVNMDFETDSKVVVENIYKGDGVSD 1648
N+ S P E LL AM+ A + FE D +++++ I +
Sbjct: 1545 NNTQSTGGWIIP-------ETKALLIAMQQAWVRGYKCVQFEGDCEILIKAINGAISRCE 1597
Query: 1649 FMAIIHDCRHLLMTDLANSDVKFIRRQANSAAHNLA 1684
+++ D + + F R N+ AH LA
Sbjct: 1598 ITSLLRDV-DFWASRFSTVVFTFTNRLCNNTAHLLA 1632
>UniRef100_Q9SZ87 RNA-directed DNA polymerase-like protein [Arabidopsis thaliana]
Length = 1274
Score = 692 bits (1785), Expect = 0.0
Identities = 413/1294 (31%), Positives = 666/1294 (50%), Gaps = 66/1294 (5%)
Query: 410 FTVDRIGRGGGLAFLWKNSANCTITNFSQNHIDVEVDDLLIGKWRLTGFYGMPENGRRKE 469
FT+ G GGLA WK + I + N ID R
Sbjct: 23 FTIPPEGLSGGLALYWKENVEVEILEAAPNFID-----------------------NRSV 59
Query: 470 SWAFLKNLARTSSLPWCVIGDFNDILSSEEKKGRSERAPWLIRGFRQAVLDAGLFDLHMT 529
W + +L S W + GDFNDIL + EK+G R FR V GL+D++ T
Sbjct: 60 FWDKISSLGAQRSSAWLLTGDFNDILDNSEKQGGPLRWEGFFLAFRSFVSQNGLWDINHT 119
Query: 530 GYAFTWFKSLGTARAVEEKLDRALVNSDWCNMFPNAGVECLSTTSSDHYPLWLTCKPVTA 589
G + +W + + ++ +LDRAL N W +FP + E L SDH PL VT
Sbjct: 120 GNSLSW-RGTRYSHFIKSRLDRALGNCSWSELFPMSKCEYLRFEGSDHRPL------VTY 172
Query: 590 VNRNPHR----FKFENAWLAEPDFKQQVQQRWKLYPEEGITQKLSYCAEDLTDWSRN-NN 644
P + F+F+ + + + V++ W+L ++ + K+S C + + W++ N+
Sbjct: 173 FGAPPLKRSKPFRFDRRLREKEEIRALVKEVWELARQDSVLYKISRCRQSIIKWTKEQNS 232
Query: 645 NFRRDISKVQKKIEKLRTHVTAANVSYFNSLKNKLDKLLVQDDLFWKQRAKTFWYKEGDL 704
N + I K Q+ +E + + S S+ +L+ Q++LFWKQ ++ W GD
Sbjct: 233 NSAKAIKKAQQALESALS-ADIPDPSLIGSITQELEAAYRQEELFWKQWSRVQWLNSGDR 291
Query: 705 NTKFFHAAATSRRKVNRIEHLENSNGTVCRKEDELQNIARNYFSHLFTKLSSTRADVISK 764
N +FHA +RR +N + +E+ +G +E+++ + +YF ++FT +++ V+ +
Sbjct: 292 NKGYFHATTRTRRMLNNLSVIEDGSGQEFHEEEQIASTISSYFQNIFTTSNNSDLQVVQE 351
Query: 765 VATSI-SDDDNCFLTAAFTLEEFKTAAFSMQADKCPGPDGFNPGFYQNFWNMCGSEIHKA 823
+ I S N L +L E K A FS+ ADK PGPDGF+ F+ +W++ +++ +
Sbjct: 352 ALSPIISSHCNEELIKISSLLEIKEALFSISADKAPGPDGFSASFFHAYWDIIEADVSRD 411
Query: 824 GCEWLENGVFPPHLNSTNIALIPKGDAQTTMKDWRPIALCNVVYKMVAKVLANRLKNVLD 883
+ + P LN T++ LIPK A + D+RPIALCNV YK+VAK+L RL+ L
Sbjct: 412 IRSFFVDSCLSPRLNETHVTLIPKISAPRKVSDYRPIALCNVQYKIVAKILTRRLQPWLS 471
Query: 884 KCISTSQSAFVPGRSILDNALVAIELIHYMKAKTKGSQGDVALKLDISKAYDRLDWEYLR 943
+ IS QSAFVPGR+I DN L+ E++H+++ +A+K D+SKAYDR+ W +L+
Sbjct: 472 ELISLHQSAFVPGRAIADNVLITHEILHFLRVSGAKKYCSMAIKTDMSKAYDRIKWNFLQ 531
Query: 944 DIMIQMRFSEKWIRWIMLCVETVDYTVLVNGVQVGPLIPGRGIRQGDPLSPYLFILCAEG 1003
++++++ F +KWIRW+M CV TV Y+ L+NG G ++P RG+RQGDPLSPYLFILC E
Sbjct: 532 EVLMRLGFHDKWIRWVMQCVCTVSYSFLINGSPQGSVVPSRGLRQGDPLSPYLFILCTEV 591
Query: 1004 LSALIRDAERRGVITGTRICTSAPAISHLLFADDCFLFFRASEQEASVMKNILTTYEAAS 1063
LS L R A+ +GV+ G R+ +P ++HLLFADD F + + + NIL YE AS
Sbjct: 592 LSGLCRKAQEKGVMVGIRVARGSPQVNHLLFADDTMFFCKTNPTCCGALSNILKKYELAS 651
Query: 1064 GQAINLQKSEMYCSRNTPTECQNRIATTLGVKQVLGTGKYLGLPSMIGRSKKATFKFVKD 1123
GQ+INL KS + S TP + + R+ +L + G GKYLGLP GR K+ F + D
Sbjct: 652 GQSINLAKSAITFSSKTPQDIKRRVKLSLRIDNEGGIGKYLGLPEHFGRRKRDIFSSIVD 711
Query: 1124 RIWNRINSWSSRCLSQAGREILIKSVLQSIPSYVMSIFLLPSTFIGEIEKMLNAFWWGHN 1183
RI R +SWS R LS AG++IL+K+VL S+PSY M F LP++ +I+ +L FWW +
Sbjct: 712 RIRQRSHSWSIRFLSSAGKQILLKAVLSSMPSYAMMCFKLPASLCKQIQSVLTRFWW-DS 770
Query: 1184 SANSRGMHWLSWERLSVPKTFGGMGFKSLRAFNLAMIGKQAWKLISNPDSLITKLLKAKY 1243
+ R M W+SW++L++P GG+GF+ + A K +W+++ P SL++++L KY
Sbjct: 771 KPDKRKMAWVSWDKLTLPINEGGLGFREIEA-------KLSWRILKEPHSLLSRVLLGKY 823
Query: 1244 FPHSDYFSASIGHN-PSYVWRSLWSVRDYLKHGLKWSIGTGETISVWNQPWLKDPLCLQP 1302
S + S + S+ WR + + RD L+ GL WSIG G++I+VW + WL P
Sbjct: 824 CNTSSFMDCSASPSFASHGWRGILAGRDLLRKGLGWSIGQGDSINVWTEAWLSPSSPQTP 883
Query: 1303 VTEVQIMWDALTVGHLFKPNTKEWNENFIRYVFNAETSNQILQTPLLQSVQVDKATWRFE 1362
+ L+V L + K WN IR + +QI + + D W
Sbjct: 884 IGPPTETNKDLSVHDLICHDVKSWNVEAIRKHL-PQYEDQIRKITINALPLQDSLVWLPV 942
Query: 1363 KNGLYSVRSAYREIINRNDVLLQHRVPGKW-NIIWNLKLPPKIKNFLWRVCRNCLPTRMR 1421
K+G Y+ ++ Y + + + ++ W IW + PK+K+FLW+ + LP
Sbjct: 943 KSGEYTTKTGY--ALAKLNSFPASQLDFNWQKNIWKIHTSPKVKHFLWKAMKGALPVGEA 1000
Query: 1422 LISKGVQCPPTCAICNNHEEDGKHLFFACSKSVGCWQ--RLGFWPSIQQIWSHNAFCADI 1479
L + ++ TC C E HL C + W+ + F PS +H++ +
Sbjct: 1001 LSRRNIEAEVTCKRC-GQTESSLHLMLLCPYAKKVWELAPVLFNPSEA---THSSVALLL 1056
Query: 1480 I----FSLLQQLDIAHQQIFAVTLWSLWRHRNNKVWNNIVETTEEIGDRAVAFLNSWKAA 1535
+ L + ++ LW LW+ RN +++N + E + +A+ +W A
Sbjct: 1057 VDAKRMVALPPTGLGSAPLYPWLLWHLWKARNRLIFDNHSCSEEGLVLKAILDARAWMEA 1116
Query: 1536 QETRIRSSPANPHFDISKWSKPSVGRFKCNVDAAFSASLHRVGFGACIRDANGNHVISRT 1595
Q SP + + P++ C VDAA++ S + G G ++D +
Sbjct: 1117 QLLIHHPSPISDY----PSPTPNLKVTSCFVDAAWTTSGY-CGMGWFLQDPYKVKIKENQ 1171
Query: 1596 ECFTPLLDVEMGEAIGLLHAMRWAKDLNLVNMDFETDSKVVVENIYKGDGVSDFMAIIHD 1655
+ + M E + + A+ A + ++ +D K ++ + G + + ++HD
Sbjct: 1172 SSSSFVGSALMAETLAVHLALVDALSTGVRQLNVFSDCKELISLLNSGKSIVELRGLLHD 1231
Query: 1656 CRHLLMTDLANSDVKFIRRQANSAAHNLAREALN 1689
R L ++ + FI R +N A +LA+ AL+
Sbjct: 1232 IRELSVS-FTHLCFFFIPRLSNVVADSLAKSALS 1264
>UniRef100_Q9FYK1 F21J9.30 [Arabidopsis thaliana]
Length = 1270
Score = 677 bits (1748), Expect = 0.0
Identities = 395/1179 (33%), Positives = 632/1179 (53%), Gaps = 61/1179 (5%)
Query: 391 MAALNKIEELKYILSFDSCFTVDRIGRGGGLAFLWKNSANCTITNFSQNHIDVEVDDLLI 450
M + + + +++ L +D +TV+ +G+ GGLA LWK+S + +N +D +V +
Sbjct: 1 MHSRDDLVDIQSWLEYDQVYTVEPVGKCGGLALLWKSSVQVDLKFVDKNLMDAQVQFGAV 60
Query: 451 GKWRLTGFYGMPENGRRKESWAFLKNLARTSSLPWCVIGDFNDILSSEEKKGRSERAPWL 510
+ ++ YG P+ +R ++W + + WC+ GDFNDIL + EK G R+
Sbjct: 61 N-FCVSCVYGDPDRSKRSQAWERISRIGVGRRDKWCMFGDFNDILHNGEKNGGPRRSDLD 119
Query: 511 IRGFRQAVLDAGLFDLHMTGYAFTWFKSLGTARAVEEKLDRALVNSDWCNMFPNAGVECL 570
+ F + + L ++ G FTW G ++ +LDRA N +W FP + L
Sbjct: 120 CKAFNEMIKGCDLVEMPAHGNGFTWAGRRGD-HWIQCRLDRAFGNKEWFCFFPVSNQTFL 178
Query: 571 STTSSDHYPLWLTCKPVTAVNRNPHRFKFENAWLAEPDFKQQVQQRW---KLYPEEGITQ 627
SDH P+ + K +++ + +F+F+ +L + D K+ + + W K +
Sbjct: 179 DFRGSDHRPVLI--KLMSSQDSYRGQFRFDKRFLFKEDVKEAIIRTWSRGKHGTNISVAD 236
Query: 628 KLSYCAEDLTDWSRNNN-NFRRDISKVQKKIEKLRTHVTAANVSYFNSLKNKLDKLLVQD 686
+L C + L+ W + NN N I++++ +EK ++ V + LK L K ++
Sbjct: 237 RLRACRKSLSSWKKQNNLNSLDKINQLEAALEKEQSLVWPI-FQRVSVLKKDLAKAYREE 295
Query: 687 DLFWKQRAKTFWYKEGDLNTKFFHAAATSRRKVNRIEHLENSNGTVCRKEDELQNIARNY 746
+ +WKQ+++ W + G+ N+K+FHAA R+ RIE L++ NG + E +A Y
Sbjct: 296 EAYWKQKSRQKWLRSGNRNSKYFHAAVKQNRQRKRIEKLKDVNGNMQTSEAAKGEVAAAY 355
Query: 747 FSHLFTKLS-STRADVISKVATSISDDDNCFLTAAFTLEEFKTAAFSMQADKCPGPDGFN 805
F +LF + S D S + +S+ N L + +E K A FS++ PGPDG +
Sbjct: 356 FGNLFKSSNPSGFTDWFSGLVPRVSEVMNESLVGEVSAQEIKEAVFSIKPASAPGPDGMS 415
Query: 806 PGFYQNFWNMCGSEIHKAGCEWLENGVFPPHLNSTNIALIPKGDAQTTMKDWRPIALCNV 865
F+Q++W+ G+++ ++ +G+ P N T++ LIPK T M D RPI+LC+V
Sbjct: 416 ALFFQHYWSTVGNQVTSEVKKFFADGIMPAEWNYTHLCLIPKTQHPTEMVDLRPISLCSV 475
Query: 866 VYKMVAKVLANRLKNVLDKCISTSQSAFVPGRSILDNALVAIELIHYMKAKTKGSQGDVA 925
+YK+++K++A RL+ L + +S +QSAFV R I DN LVA EL+H +K + S +A
Sbjct: 476 LYKIISKIMAKRLQPWLPEIVSDTQSAFVSERLITDNILVAHELVHSLKVHPRISSEFMA 535
Query: 926 LKLDISKAYDRLDWEYLRDIMIQMRFSEKWIRWIMLCVETVDYTVLVNGVQVGPLIPGRG 985
+K D+SKAYDR++W YLR +++ + F KW+ WIM+CV +V Y+VL+N G +I RG
Sbjct: 536 VKSDMSKAYDRVEWSYLRSLLLSLGFHLKWVNWIMVCVSSVTYSVLINDCPFGLIILQRG 595
Query: 986 IRQGDPLSPYLFILCAEGLSALIRDAERRGVITGTRICTSAPAISHLLFADDCFLFFRAS 1045
+RQGDPLSP+LF+LC EGL+ L+ A+ G + G + + P + HLLFADD +AS
Sbjct: 596 LRQGDPLSPFLFVLCTEGLTHLLNKAQWEGALEGIQFSENGPMVHHLLFADDSLFLCKAS 655
Query: 1046 EQEASVMKNILTTYEAASGQAINLQKSEMYCSRNTPTECQNRIATTLGVKQVLGTGKYLG 1105
+++ V++ IL Y A+GQ INL KS + + + I T LG+ G G YLG
Sbjct: 656 REQSLVLQKILKVYGNATGQTINLNKSSITFGEKVDEQLKGTIRTCLGIFTEGGAGTYLG 715
Query: 1106 LPSMIGRSKKATFKFVKDRIWNRINSWSSRCLSQAGREILIKSVLQSIPSYVMSIFLLPS 1165
LP SK ++KDR+ +++ W +RCLSQ G+E+L+KSV ++P + MS F LP
Sbjct: 716 LPECFSGSKVDMLHYLKDRLKEKLDVWFTRCLSQGGKEVLLKSVALAMPVFAMSCFKLPI 775
Query: 1166 TFIGEIEKMLNAFWWGHNSANSRGMHWLSWERLSVPKTFGGMGFKSLRAFNLAMIGKQAW 1225
T +E + +FWW + +SR +HW SWERL +PK GG+GF+ +++FN A++ KQAW
Sbjct: 776 TTCENLESAMASFWWD-SCDHSRKIHWQSWERLCLPKDSGGLGFRDIQSFNQALLAKQAW 834
Query: 1226 KLISNPDSLITKLLKAKYFPHSDYFSASIGHNPSYVWRSLWSVRDYLKHGLKWSIGTGET 1285
+L+ PD L+++LLK++YF +D+ A++ PS+ WRS+ R+ L GL+ +G G +
Sbjct: 835 RLLHFPDCLLSRLLKSRYFDATDFLDAALSQRPSFGWRSILFGRELLSKGLQKRVGDGAS 894
Query: 1286 ISVWNQPWLKDPLCLQPVTEVQIMWDALTVGHLFKPNTKEWNENFIRYVFNAETSNQILQ 1345
+ VW PW+ D P + I L V L P T W+E + +F E IL+
Sbjct: 895 LFVWIDPWIDDNGFRAPWRKNLIYDVTLKVKALLNPRTGFWDEEVLHDLFLPE---DILR 951
Query: 1346 TPLLQSV--QVDKATWRFEKNGLYSVRSAY------REIINRNDVLLQHRVPGKWNIIWN 1397
++ V Q D W+ K+G +SV+SAY + R++V +Q G +WN
Sbjct: 952 IKAIKPVISQADFFVWKLNKSGDFSVKSAYWLAYQTKSQNLRSEVSMQPSTLGLKTQVWN 1011
Query: 1398 LKLPPKIKNFLWRVCRNCLPTRMRLISKGVQCPPTCAICNNHEEDGKHLFFACSKSVGCW 1457
L+ PKIK FLW+VC E H F C S W
Sbjct: 1012 LQTDPKIKIFLWKVCGEL------------------------GESTNHTLFLCPLSRQIW 1047
Query: 1458 QRLGFWPSIQQIWSHNAFCADIIFSLLQQLD-----IAHQQIFAVTLWSLWRHRNNKVWN 1512
L +P +S+ + ++I LL+ D I ++IF LW +W++RN+ ++
Sbjct: 1048 A-LSDYPFPPDGFSNGSIYSNINH-LLENKDNKEWPINLRKIFPWILWRIWKNRNSFIFE 1105
Query: 1513 NI----VETTEEIGDRAVAFLNSWKAAQETRIRSSPANP 1547
I +T +I D V W AQ S NP
Sbjct: 1106 GISYPATDTVIKIRDDVV----EWFEAQCLDGEGSALNP 1140
>UniRef100_Q9FXJ5 F5A9.24 [Arabidopsis thaliana]
Length = 1254
Score = 674 bits (1739), Expect = 0.0
Identities = 369/1056 (34%), Positives = 591/1056 (55%), Gaps = 22/1056 (2%)
Query: 389 ETMAALNKIEELKYILSFDSCFTVDRIGRGGGLAFLWKNSANCTITNFSQNHIDVEVDDL 448
ETM + + + +++ L +D +TV+ +G+ GGLA LWK+S + +N +D +V
Sbjct: 2 ETMHSRDDLVDIQSWLEYDQVYTVEPVGKCGGLALLWKSSVQVDLKFVDKNLMDAQVQFG 61
Query: 449 LIGKWRLTGFYGMPENGRRKESWAFLKNLARTSSLPWCVIGDFNDILSSEEKKGRSERAP 508
+ + ++ YG P+ +R ++W + + WC+ GDFNDIL + EK G R+
Sbjct: 62 AVN-FCVSCVYGDPDRSKRSQAWERISRIGVGRRDKWCMFGDFNDILHNGEKNGGPRRSD 120
Query: 509 WLIRGFRQAVLDAGLFDLHMTGYAFTWFKSLGTARAVEEKLDRALVNSDWCNMFPNAGVE 568
+ F + + L ++ G FTW G ++ +LDRA N +W FP +
Sbjct: 121 LDCKAFNEMIKGCDLVEMPAHGNGFTWAGRRGD-HWIQCRLDRAFGNKEWFCFFPVSNQT 179
Query: 569 CLSTTSSDHYPLWLTCKPVTAVNRNPHRFKFENAWLAEPDFKQQVQQRW---KLYPEEGI 625
L SDH P+ + K +++ + +F+F+ +L + D K+ + + W K +
Sbjct: 180 FLDFRGSDHRPVLI--KLMSSQDSYRGQFRFDKRFLFKEDVKEAIIRTWSRGKHGTNISV 237
Query: 626 TQKLSYCAEDLTDWSRNNN-NFRRDISKVQKKIEKLRTHVTAANVSYFNSLKNKLDKLLV 684
+L C + L+ W + NN N I++++ +EK ++ V + LK L K
Sbjct: 238 ADRLRACRKSLSSWKKQNNLNSLDKINQLEAALEKEQSLVWPI-FQRVSVLKKDLAKAYR 296
Query: 685 QDDLFWKQRAKTFWYKEGDLNTKFFHAAATSRRKVNRIEHLENSNGTVCRKEDELQNIAR 744
+++ +WKQ+++ W + G+ N+K+FHAA R+ RIE L++ NG + E +A
Sbjct: 297 EEEAYWKQKSRQKWLRSGNRNSKYFHAAVKQNRQRKRIEKLKDVNGNMQTSEAAKGEVAA 356
Query: 745 NYFSHLFTKLS-STRADVISKVATSISDDDNCFLTAAFTLEEFKTAAFSMQADKCPGPDG 803
YF +LF + S D S + +S+ N L + +E K A FS++ PGPDG
Sbjct: 357 AYFGNLFKSSNPSGFTDWFSGLVPRVSEVMNESLVGEVSAQEIKEAVFSIKPASAPGPDG 416
Query: 804 FNPGFYQNFWNMCGSEIHKAGCEWLENGVFPPHLNSTNIALIPKGDAQTTMKDWRPIALC 863
+ F+Q++W+ G+++ ++ +G+ P N T++ LIPK T M D RPI+LC
Sbjct: 417 MSALFFQHYWSTVGNQVTSEVKKFFADGIMPAEWNYTHLCLIPKTQHPTEMVDLRPISLC 476
Query: 864 NVVYKMVAKVLANRLKNVLDKCISTSQSAFVPGRSILDNALVAIELIHYMKAKTKGSQGD 923
+V+YK+++K++A RL+ L + +S +QSAFV R I DN LVA EL+H +K + S
Sbjct: 477 SVLYKIISKIMAKRLQPWLPEIVSDTQSAFVSERLITDNILVAHELVHSLKVHPRISSEF 536
Query: 924 VALKLDISKAYDRLDWEYLRDIMIQMRFSEKWIRWIMLCVETVDYTVLVNGVQVGPLIPG 983
+A+K D+SKAYDR++W YLR +++ + F KW+ WIM+CV +V Y+VL+N G +I
Sbjct: 537 MAVKSDMSKAYDRVEWSYLRSLLLSLGFHLKWVNWIMVCVSSVTYSVLINDCPFGLIILQ 596
Query: 984 RGIRQGDPLSPYLFILCAEGLSALIRDAERRGVITGTRICTSAPAISHLLFADDCFLFFR 1043
RG+RQGDPLSP+LF+LC EGL+ L+ A+ G + G + + P + HLLFADD +
Sbjct: 597 RGLRQGDPLSPFLFVLCTEGLTHLLNKAQWEGALEGIQFSENGPMVHHLLFADDSLFLCK 656
Query: 1044 ASEQEASVMKNILTTYEAASGQAINLQKSEMYCSRNTPTECQNRIATTLGVKQVLGTGKY 1103
AS +++ V++ IL Y A+GQ INL KS + + + I T LG+ G G Y
Sbjct: 657 ASREQSLVLQKILKVYGNATGQTINLNKSSITFGEKVDEQLKGTIRTCLGIFTEGGAGTY 716
Query: 1104 LGLPSMIGRSKKATFKFVKDRIWNRINSWSSRCLSQAGREILIKSVLQSIPSYVMSIFLL 1163
LGLP SK ++KDR+ +++ W +RCLSQ G+E+L+KSV ++P + MS F L
Sbjct: 717 LGLPECFSGSKVDMLHYLKDRLKEKLDVWFTRCLSQGGKEVLLKSVALAMPVFAMSCFKL 776
Query: 1164 PSTFIGEIEKMLNAFWWGHNSANSRGMHWLSWERLSVPKTFGGMGFKSLRAFNLAMIGKQ 1223
P T +E + +FWW + +SR +HW SWERL +PK GG+GF+ +++FN A++ KQ
Sbjct: 777 PITTCENLESAMASFWW-DSCDHSRKIHWQSWERLCLPKDSGGLGFRDIQSFNQALLAKQ 835
Query: 1224 AWKLISNPDSLITKLLKAKYFPHSDYFSASIGHNPSYVWRSLWSVRDYLKHGLKWSIGTG 1283
AW+L+ PD L+++LLK++YF +D+ A++ PS+ WRS+ R+ L GL+ +G G
Sbjct: 836 AWRLLHFPDCLLSRLLKSRYFDATDFLDAALSQRPSFGWRSILFGRELLSKGLQKRVGDG 895
Query: 1284 ETISVWNQPWLKDPLCLQPVTEVQIMWDALTVGHLFKPNTKEWNENFIRYVFNAETSNQI 1343
++ VW PW+ D P + I L V L P T W+E + +F E I
Sbjct: 896 ASLFVWIDPWIDDNGFRAPWRKNLIYDVTLKVKALLNPRTGFWDEEVLHDLFLPE---DI 952
Query: 1344 LQTPLLQSV--QVDKATWRFEKNGLYSVRSAY------REIINRNDVLLQHRVPGKWNII 1395
L+ ++ V Q D W+ K+G +SV+SAY + R++V +Q G +
Sbjct: 953 LRIKAIKPVISQADFFVWKLNKSGDFSVKSAYWLAYQTKSQNLRSEVSMQPSTLGLKTQV 1012
Query: 1396 WNLKLPPKIKNFLWRVCRNCLPTRMRLISKGVQCPP 1431
WNL+ PKIK FLW+V LP L +G+ P
Sbjct: 1013 WNLQTDPKIKIFLWKVLSGILPVAENLNGRGMSLIP 1048
>UniRef100_Q9C697 Reverse transcriptase, putative; 16838-20266 [Arabidopsis thaliana]
Length = 1142
Score = 668 bits (1723), Expect = 0.0
Identities = 394/1157 (34%), Positives = 600/1157 (51%), Gaps = 45/1157 (3%)
Query: 574 SSDHYPLWLTCKPVTAVNRNPHRFKFENAWLAEPDFKQQVQQRWKL---YPEEGITQKLS 630
+SDH P+ T + R F+F+ W+ + + + Q W L + E +KL+
Sbjct: 3 ASDHSPVIATI--ADKIPRGKQNFRFDKRWIGKDGLLEAISQGWNLDSGFREGQFVEKLT 60
Query: 631 YCAEDLTDWSRNNNNFRR--------DISKVQKKIEKLRTHVTAANVSYFNSLKNKLDKL 682
C ++ W ++ F R ++ Q+ ++ R +T L +L +
Sbjct: 61 NCRRAISKWRKSLIPFGRQTIEDLKAELDVAQRDDDRSREEIT--------ELTLRLKEA 112
Query: 683 LVQDDLFWKQRAKTFWYKEGDLNTKFFHAAATSRRKVNRIEHLENSNGTVCRKEDELQNI 742
++ +W Q++++ W K GD N+KFFHA RR NRI L + NG ++D++QNI
Sbjct: 113 YRDEEQYWYQKSRSLWMKLGDNNSKFFHALTKQRRARNRITGLHDENGIWSIEDDDIQNI 172
Query: 743 ARNYFSHLFTKLSSTRAD-VISKVATSISDDDNCFLTAAFTLEEFKTAAFSMQADKCPGP 801
A +YF +LFT + D + +V I+D N LTA T E + A F + +K PGP
Sbjct: 173 AVSYFQNLFTTANPQVFDEALGEVQVLITDRINDLLTADATECEVRAALFMIHPEKAPGP 232
Query: 802 DGFNPGFYQNFWNMCGSEIHKAGCEWLENGVFPPHLNSTNIALIPKGDAQTTMKDWRPIA 861
DG F+Q W + S++ +L+ GVF LN+TNI LIPK + T M + RPI+
Sbjct: 233 DGMTALFFQKSWAIIKSDLLSLVNSFLQEGVFDKRLNTTNICLIPKTERPTRMTELRPIS 292
Query: 862 LCNVVYKMVAKVLANRLKNVLDKCISTSQSAFVPGRSILDNALVAIELIHYMKAKTKGSQ 921
LCNV YK+++K+L RLK VL IS +QSAFV GR I DN L+A E+ H ++ +
Sbjct: 293 LCNVGYKVISKILCQRLKTVLPNLISETQSAFVDGRLISDNILIAQEMFHGLRTNSSCKD 352
Query: 922 GDVALKLDISKAYDRLDWEYLRDIMIQMRFSEKWIRWIMLCVETVDYTVLVNGVQVGPLI 981
+A+K D+SKAYD+++W ++ ++ +M F EKWI WIM C+ TV Y VL+NG G +I
Sbjct: 353 KFMAIKTDMSKAYDQVEWNFIEALLRKMGFCEKWISWIMWCITTVQYKVLINGQPKGLII 412
Query: 982 PGRGIRQGDPLSPYLFILCAEGLSALIRDAERRGVITGTRICTSAPAISHLLFADDCFLF 1041
P RG+RQGDPLSPYLFILC E L A IR AER+ +ITG ++ T +PA+SHLLFADD F
Sbjct: 413 PERGLRQGDPLSPYLFILCTEVLIANIRKAERQNLITGIKVATPSPAVSHLLFADDSLFF 472
Query: 1042 FRASEQEASVMKNILTTYEAASGQAINLQKSEMYCSRNTPTECQNRIATTLGVKQVLGTG 1101
+A++++ ++ IL YE+ SGQ IN KS + + I LG+ + G G
Sbjct: 473 CKANKEQCGIILEILKQYESVSGQQINFSKSSIQFGHKVEDSIKADIKLILGIHNLGGMG 532
Query: 1102 KYLGLPSMIGRSKKATFKFVKDRIWNRINSWSSRCLSQAGREILIKSVLQSIPSYVMSIF 1161
YLGLP +G SK F FV+DR+ +RIN WS++ LS+ G+E++IKSV ++P YVMS F
Sbjct: 533 SYLGLPESLGGSKTKVFSFVRDRLQSRINGWSAKFLSKGGKEVMIKSVAATLPRYVMSCF 592
Query: 1162 LLPSTFIGEIEKMLNAFWWGHNSANSRGMHWLSWERLSVPKTFGGMGFKSLRAFNLAMIG 1221
LP ++ + FWW N +SRGMHW++W++L K+ GG+GF+++ FN A++
Sbjct: 593 RLPKAITSKLTSAVAKFWWSSN-GDSRGMHWMAWDKLCSSKSDGGLGFRNVDDFNSALLA 651
Query: 1222 KQAWKLISNPDSLITKLLKAKYFPHSDYFSASIGHNPSYVWRSLWSVRDYLKHGLKWSIG 1281
KQ W+LI+ PDSL K+ K +YF S+ + ++PSY WRS+ S R + GL +G
Sbjct: 652 KQLWRLITAPDSLFAKVFKGRYFRKSNPLDSIKSYSPSYGWRSMISARSLVYKGLIKRVG 711
Query: 1282 TGETISVWNQPWLKDPLCLQPVTEVQIMWDALTVGHLFKPNTKEWNENFIRYVFNAETSN 1341
+G +ISVWN PW+ I+ +L V L + WN + ++ +F+ E
Sbjct: 712 SGASISVWNDPWIPAQFPRPAKYGGSIVDPSLKVKSLIDSRSNFWNIDLLKELFDPEDVP 771
Query: 1342 QILQTPLLQSVQVDKATWRFEKNGLYSVRSAY---REIINRNDVLLQHRVPGKWNIIWNL 1398
I P+ D W F K G Y+V+S Y R +N L+ + IW +
Sbjct: 772 LISALPIGNPNMEDTLGWHFTKAGNYTVKSGYHTARLDLNEGTTLIGPDLTTLKAYIWKV 831
Query: 1399 KLPPKIKNFLWRVCRNCLPTRMRLISKGVQCPPTCAICNNHEEDGKHLFFACSKSVGCWQ 1458
+ PPK+++FLW++ C+P L +G+ C C C EE H F C + W
Sbjct: 832 QCPPKLRHFLWQILSGCVPVSENLRKRGILCDKGCVSCGASEESINHTLFQCHPARQIW- 890
Query: 1459 RLGFWPSIQQIWSHNAFCADIIFSLLQQLDIAHQQIFAVTLWSLWRHRNNKVWNNIVETT 1518
L P+ I+ N+ ++ + + +W +W+ RN KV+ N+ +
Sbjct: 891 ALSQIPTAPGIFPSNSIFTNLDHLFWRIPSGVDSAPYPWIIWYIWKARNEKVFENVDKDP 950
Query: 1519 EEIGDRAVAFLNSWKAAQ----ETRIRSSPANPHFDISKWSKPSV-GRFKCNVDAAFSAS 1573
EI AV SW+ AQ R S + + S+ + F+C +D ++ AS
Sbjct: 951 MEILLLAVKEAQSWQEAQVELHSERHGSLSIDSRIRVRDVSQDTTFSGFRCFIDGSWKAS 1010
Query: 1574 LHRVGFG-ACIRDANGNHVISRTECFTPLLDVEMGEAIGLLHAMRWAKDLNLVNMDFETD 1632
G G C+ + + L + E LL AM+ + N+ F TD
Sbjct: 1011 DQFSGTGWFCLSSLGESPTMGAANVRRSLSPLHT-EMEALLWAMKCMIGADNQNVAFFTD 1069
Query: 1633 SKVVVENIYKGDGVSDFMAIIHDCRHLLMTDLANSDVKFIRRQANSAAHNLAREALNHAS 1692
+V+ + F + + + + N + I R AN A LAR+
Sbjct: 1070 CSDLVKMVSSPTEWPAFSVYLEELQS-DREEFTNFSLSLISRSANVKADKLARKI----- 1123
Query: 1693 FHYHLNIPHCIHTLINN 1709
+PH + T +NN
Sbjct: 1124 ----RTVPHHV-TYVNN 1135
>UniRef100_Q9SJ38 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1229
Score = 653 bits (1685), Expect = 0.0
Identities = 392/1200 (32%), Positives = 613/1200 (50%), Gaps = 32/1200 (2%)
Query: 492 NDILSSEEKKGRSERAPWLIRGFRQAVLDAGLFDLHMTGYAFTWFKSLGTARAVEEKLDR 551
N+IL + EK+G R FR + GL+DL +G F+W + + V ++LDR
Sbjct: 36 NEILDNSEKRGGPPRDQGSFIDFRSFISKNGLWDLKYSGNPFSW-RGMRYDWFVRQRLDR 94
Query: 552 ALVNSDWCNMFPNAGVECLSTTSSDHYPLWLTCKPVTAVNRNPHRFKFENAWLAEPDFKQ 611
A+ N+ W FP+ E L SDH PL + R +F+F+N
Sbjct: 95 AMSNNSWLESFPSGRSEYLRFEGSDHRPLVVFVDEARVKRRG--QFRFDNRLRDNDVVNA 152
Query: 612 QVQQRWKLYPEEGITQKLSYCAEDLTDWSRNNN-NFRRDISKVQKKIEKLRTHVTAANVS 670
+Q+ W + + K++ C ++ +W+R N N I K QK +E+ T N +
Sbjct: 153 LIQETWTNAGDASVLTKMNQCRREIINWTRLQNLNSAELIEKTQKALEEALT-ADPPNPT 211
Query: 671 YFNSLKNKLDKLLVQDDLFWKQRAKTFWYKEGDLNTKFFHAAATSRRKVNRIEHLENSNG 730
+L L+ ++ FWKQR++ W GD NT +FHA +RR NR+ +E+ NG
Sbjct: 212 TIGALTATLEHAYKLEEQFWKQRSRVLWLHSGDRNTGYFHAVTRNRRTQNRLTVMEDING 271
Query: 731 TVCRKEDELQNIARNYFSHLFTKLSSTRADVISK-VATSISDDDNCFLTAAFTLEEFKTA 789
+E ++ I YF +FT S V+ + + +S DN FLT EE K A
Sbjct: 272 VAQHEEHQISQIISGYFQQIFTSESDGDFSVVDEAIEPMVSQGDNDFLTRIPNDEEVKDA 331
Query: 790 AFSMQADKCPGPDGFNPGFYQNFWNMCGSEIHKAGCEWLENGVFPPHLNSTNIALIPKGD 849
FS+ A K PGPDGF GFY ++W++ +++ + + + FP +N T+I LIPK
Sbjct: 332 VFSINASKAPGPDGFTAGFYHSYWHIISTDVGREIRLFFTSKNFPRRMNETHIRLIPKDL 391
Query: 850 AQTTMKDWRPIALCNVVYKMVAKVLANRLKNVLDKCISTSQSAFVPGRSILDNALVAIEL 909
+ D+RPIALCN+ YK+VAK++ R++ +L K IS +QSAFVPGR I DN L+ E+
Sbjct: 392 GPRKVADYRPIALCNIFYKIVAKIMTKRMQLILPKLISENQSAFVPGRVISDNVLITHEV 451
Query: 910 IHYMKAKTKGSQGDVALKLDISKAYDRLDWEYLRDIMIQMRFSEKWIRWIMLCVETVDYT 969
+H+++ + +A+K D+SKAYDR++W++L+ ++ + F WI W++ CV +V Y+
Sbjct: 452 LHFLRTSSAKKHCSMAVKTDMSKAYDRVEWDFLKKVLQRFGFHSIWIDWVLECVTSVSYS 511
Query: 970 VLVNGVQVGPLIPGRGIRQGDPLSPYLFILCAEGLSALIRDAERRGVITGTRICTSAPAI 1029
L+NG G ++P RG+RQGDPLSP LFILC E LS L A+R + G R+ + P +
Sbjct: 512 FLINGTPQGKVVPTRGLRQGDPLSPCLFILCTEVLSGLCTRAQRLRQLPGVRVSINGPRV 571
Query: 1030 SHLLFADDCFLFFRASEQEASVMKNILTTYEAASGQAINLQKSEMYCSRNTPTECQNRIA 1089
+HLLFADD F ++ + + + IL+ Y ASGQ+IN KS + S TP + ++
Sbjct: 572 NHLLFADDTMFFSKSDPESCNKLSEILSRYGKASGQSINFHKSSVTFSSKTPRSVKGQVK 631
Query: 1090 TTLGVKQVLGTGKYLGLPSMIGRSKKATFKFVKDRIWNRINSWSSRCLSQAGREILIKSV 1149
L +++ GTGKYLGLP GR K+ F + D+I + +SW+SR LSQAG+++++K+V
Sbjct: 632 RILKIRKEGGTGKYLGLPEHFGRRKRDIFGAIIDKIRQKSHSWASRFLSQAGKQVMLKAV 691
Query: 1150 LQSIPSYVMSIFLLPSTFIGEIEKMLNAFWWGHNSANSRGMHWLSWERLSVPKTFGGMGF 1209
L S+P Y MS F LPS +I+ +L FWW + R W++W +L+ PK GG+GF
Sbjct: 692 LASMPLYSMSCFKLPSALCRKIQSLLTRFWW-DTKPDVRKTSWVAWSKLTNPKNAGGLGF 750
Query: 1210 KSLRAFNLAMIGKQAWKLISNPDSLITKLLKAKYFPHSDYFSASIGHNPSYVWRSLWSVR 1269
+ + N +++ K W+L+++P+SL++++L KY S + + PS+ WRS+ + R
Sbjct: 751 RDIERCNDSLLAKLGWRLLNSPESLLSRILLGKYCHSSSFMECKLPSQPSHGWRSIIAGR 810
Query: 1270 DYLKHGLKWSIGTGETISVWNQPWLKDPLCLQPVTEVQIMWDALTVGHLFKPNTKEWNEN 1329
+ LK GL W I GE +S+WN PWL L P+ L V L NT +W+ N
Sbjct: 811 EILKEGLGWLITNGEKVSIWNDPWLSISKPLVPIGPALREHQDLRVSALINQNTLQWDWN 870
Query: 1330 FIRYVFNAETSNQILQTPLLQSVQVDKATWRFEKNGLYSVRSAYREIINRNDVLLQHRVP 1389
I + N I Q P S VDK W K+G Y+ RS Y + + Q +
Sbjct: 871 KIAVIL-PNYENLIKQLPAPSSRGVDKLAWLPVKSGQYTSRSGYGIASVASIPIPQTQFN 929
Query: 1390 GKWNIIWNLKLPPKIKNFLWRVCRNCLPTRMRLISKGVQCPPTCAICNNHEEDGKHLFFA 1449
+ N +W L+ PKIK+ +W+ LP ++L+ + + C C E HLFF
Sbjct: 930 WQSN-LWKLQTLPKIKHLMWKAAMEALPVGIQLVRRHISPSAACHRC-GAPESTTHLFFH 987
Query: 1450 CSKSVGCWQRLGFWPSIQQIWSHNAFCADIIFSLLQQLDIAHQQIFAVTLWSLWRHRNNK 1509
C + W+ P + + D + L + + + + + L+ W
Sbjct: 988 CEFAAQVWE---LAPLQETTVPPGSSMLDALSLLKKAIILPPTGVTSAALFP-W------ 1037
Query: 1510 VWNNIVETTEEIGDRAVAFLN--SWKAAQETRIRSSPANPHFDISKWSKPSVGRFKCNVD 1567
I ++G A L+ +W++AQ R N IS+ G F C VD
Sbjct: 1038 ----ICGIYGKLGTMTRAILDALAWQSAQ--RCLPKTRNVVHPISQLPVLRSGYF-CFVD 1090
Query: 1568 AAFSASLHRVGFGACIRDANG--NHVISRTECFTPLLDVEMGEAIGLLHAMRWAKDLNLV 1625
AA+ A G G + A + + L EA + A+ A L
Sbjct: 1091 AAWIAQSSLAGSGWVFQSATALEKETATYSAGCRRLPSALSAEAWAIKSALLHALQLGRT 1150
Query: 1626 NMDFETDSKVVVENIYKGDGVSDFMAIIHDCRHLLMTDLANSDVKFIRRQANSAAHNLAR 1685
++ +DSK VV+ + +++ ++ + R L + FI R AN+ A A+
Sbjct: 1151 DLMVLSDSKSVVDALTSNISINEIYGLLMEIR-ALRVSFHSLCFNFISRSANAIADATAK 1209
>UniRef100_Q8H0A5 Putative reverse transcriptase [Oryza sativa]
Length = 1557
Score = 650 bits (1676), Expect = 0.0
Identities = 378/1169 (32%), Positives = 578/1169 (49%), Gaps = 88/1169 (7%)
Query: 348 PGPPRKMSFIVWNCRGLGSPSTVPTLKYLVRTYKPEGIFLSETMAALNKIEELKYILSFD 407
P P M+ + WNCRG+G+ +TV L LV+ + +FL ET ++KI+ L+ L
Sbjct: 428 PAPSGAMNCLSWNCRGIGNATTVQELCALVKEASSQFVFLCETRQPVDKIKRLRNRLGLR 487
Query: 408 SCFTVDRIGRGGGLAFLWKNSANCTITNFSQNHIDVEV----DDLLIGKWRLTGFYGMPE 463
V G GGLA W S + I N ++ ID + +D L W T YG P
Sbjct: 488 GFAGVSSDGMSGGLALFWHESVHVEILNKNERFIDAYIRLSPNDPL---WHATFVYGEPR 544
Query: 464 NGRRKESWAFLKNLARTSSLPWCVIGDFNDILSSEEKKGRSERAPWLIRGFRQAVLDAGL 523
R + W+ L ++ ++S+L W VIGDFN+ L E R R+ ++ FR + L
Sbjct: 545 VENRHKMWSLLNSIKQSSNLSWLVIGDFNEALWQFEHFSRKPRSDAQMQAFRDVLDTCEL 604
Query: 524 FDLHMTGYAFTWFKSLGTARAVEEKLDRALVNSDWCNMFPNAGVECLSTTSSDHYPLWLT 583
DL ++G T+ G V+ +LDRAL + +W ++F V+ L + S+H P+ L
Sbjct: 605 HDLGISGLPHTYDNRRGGWNNVKVRLDRALADDEWRDLFSQYQVQHLVSPCSNHCPIVL- 663
Query: 584 CKPVTAVNRNPHRFKFENAWLAEPDFKQQVQQRWKLYPEEGITQKLSYCAEDLTDWSRNN 643
KL D + +
Sbjct: 664 --------------------------------------------KLHGSVPDQSWSKKKY 679
Query: 644 NNFRRDISKVQKKIEKLRTHVTAANVSYFNSLKNKLDKLLVQDDLFWKQRAKTFWYKEGD 703
N R++ K +KK+ L T + + + +++LL ++++ W QR++ W KEGD
Sbjct: 680 KNIGRELEKTRKKLSNLLEAGT--DKAAIRQTSDYMNELLYREEMLWLQRSRVNWLKEGD 737
Query: 704 LNTKFFHAAATSRRKVNRIEHLENSNGTVCRKEDELQNIARNYFSHLFTKLSSTRADVIS 763
NTKFFH R K N+I L ++ GT R + ++ +A YF +FT + ++
Sbjct: 738 RNTKFFHNRVAWRAKKNKITKLRDAAGTFHRSKMVMEQMATQYFKDMFTADPNLDHSPVT 797
Query: 764 K-VATSISDDDNCFLTAAFTLEEFKTAAFSMQADKCPGPDGFNPGFYQNFWNMCGSEIHK 822
+ + ++++ N L + F+ +E A F + K PG DGF FYQ W ++I +
Sbjct: 798 RLIQPKVTEEMNNRLCSEFSEDEIANALFQIGPLKAPGTDGFLARFYQRNWGCIKNDIVR 857
Query: 823 AGCEWLENGVFPPHLNSTNIALIPKGDAQTTMKDWRPIALCNVVYKMVAKVLANRLKNVL 882
A E+ +G+ +N T I LIPK + ++D+RPI+LCNV+YK+++K L NRL+ L
Sbjct: 858 AVKEFFHSGIMLEGVNDTAIVLIPKIEQPRDLRDFRPISLCNVIYKVLSKCLVNRLRPFL 917
Query: 883 DKCISTSQSAFVPGRSILDNALVAIELIHYMKAKTKGSQGDVALKLDISKAYDRLDWEYL 942
D+ IS +QSAFVP R I DNAL+A E H+++ A KLD+SKAYDR+DW +L
Sbjct: 918 DELISKNQSAFVPRRMITDNALLAFECFHFIQRNKNPRSAACAYKLDLSKAYDRVDWRFL 977
Query: 943 RDIMIQMRFSEKWIRWIMLCVETVDYTVLVNGVQVGPLIPGRGIRQGDPLSPYLFILCAE 1002
M + FS W+ W+M C+ TV +G P+LF+ A+
Sbjct: 978 EQSMYKWGFSHCWVSWVMTCISTV-------------------CDKGTRCDPFLFLFVAD 1018
Query: 1003 GLSALIRDAERRGVITGTRICTSAPAISHLLFADDCFLFFRASEQEASVMKNILTTYEAA 1062
GLS L+ + +G I+ R+C AP ISHLLFADD LFF+A ++ +K + +Y +
Sbjct: 1019 GLSLLLEEKVEQGAISPIRVCHQAPGISHLLFADDTLLFFKADLSQSQAIKEVFGSYATS 1078
Query: 1063 SGQAINLQKSEMYCSRNTPTECQNRIATTLGVKQVLGTGKYLGLPSMIGRSKKATFKFVK 1122
+GQ IN K + + P ++ I L + KYLGLP+ GR K F+ ++
Sbjct: 1079 TGQLINPTKCSILFGNSLPIASRDAITNCLQIASTEFEDKYLGLPTPGGRMHKGRFQSLR 1138
Query: 1123 DRIWNRINSWSSRCLSQAGREILIKSVLQSIPSYVMSIFLLPSTFIGEIEKMLNAFWWGH 1182
+RIW +I W LS G+E+LIK+V+Q+IP YVM IF LP + ++ + FWWG
Sbjct: 1139 ERIWKKILQWGENYLSSGGKEVLIKAVIQAIPVYVMGIFKLPESVCEDLTSLTRNFWWGA 1198
Query: 1183 NSANSRGMHWLSWERLSVPKTFGGMGFKSLRAFNLAMIGKQAWKLISNPDSLITKLLKAK 1242
R HW +W+ L+ K+ GG+GFK +R FN A++ +QAW+LI NPDSL ++LKAK
Sbjct: 1199 EK-GKRKTHWKAWKSLTKSKSLGGLGFKDIRLFNQALLARQAWRLIDNPDSLCARVLKAK 1257
Query: 1243 YFPHSDYFSASIGHNPSYVWRSLWSVRDYLKHGLKWSIGTGETISVWNQPWLKDPLCLQP 1302
Y+P+ S G N S W+++ + +K G+ W IG G ++ VW PWL L +P
Sbjct: 1258 YYPNGSIVDTSFGGNASPGWQAIEHGLELVKKGIIWRIGNGRSVRVWQDPWLPRDLSRRP 1317
Query: 1303 VT---EVQIMWDALTVGHLFKPNTKEWNENFIRYVFNAETSNQILQTPLLQSVQVDKATW 1359
+T +I W V L N W+ N I +F IL+ + D W
Sbjct: 1318 ITPKNNCRIKW----VADLMLDNGM-WDANKINQIFLPVDVEIILKLRTSSRDEEDFIAW 1372
Query: 1360 RFEKNGLYSVRSAYREIIN-----RNDVLLQHRVPGKWNIIWNLKLPPKIKNFLWRVCRN 1414
+K G +SVR+AYR N + + W ++W +P K+K F WR N
Sbjct: 1373 HPDKLGNFSVRTAYRLAENWAKEEASSSSSDVNIRKAWELLWKCNVPSKVKIFTWRATSN 1432
Query: 1415 CLPTRMRLISKGVQCPPTCAICNNHEEDGKHLFFACSKSVGCWQRLGFWPSIQQIWSHNA 1474
CLPT + ++ TC IC +ED H C ++ W + + +
Sbjct: 1433 CLPTWDNKKKRNLEISDTCVICGMEKEDTMHALCRCPQAKHLWLAMKESNDLSLRMDDHL 1492
Query: 1475 FCADIIFSLLQQLDIAHQQIFAVTLWSLW 1503
+F+ L L Q +F + LW +W
Sbjct: 1493 LGPSWLFNRLALLPDHEQPMFLMVLWRIW 1521
>UniRef100_Q9SLE9 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1319
Score = 635 bits (1638), Expect = e-180
Identities = 401/1343 (29%), Positives = 667/1343 (48%), Gaps = 78/1343 (5%)
Query: 404 LSFDSCFTVDRIGRGGGLAFLWKNSANCTITNFSQNHIDVEVDDLLIGK--WRLTGFYGM 461
L +D T++ +G+ GGLA WKN +N +D++V GK W ++ YG
Sbjct: 14 LGYDYFHTIEPVGKSGGLAIFWKNHLEIDFLFEDKNLLDLKVSQ---GKKSWFVSCVYGN 70
Query: 462 PENGRRKESWAFLKNLARTSSLPWCVIGDFNDILSSEEKKGRSERAPWLIRGFRQAVLDA 521
P R L ++ + WC+IGDFNDILS++ K G R + F+ +L+
Sbjct: 71 PVLHLRYLLLDKLSSIGVQRNSAWCMIGDFNDILSNDGKLGGPSRLISSFQPFKNMLLNC 130
Query: 522 GLFDLHMTGYAFTWFKSLGTARAVEEKLDRALVNSDWCNMFPNAGVECLSTTSSDHYPLW 581
+ + +G +FTW + + ++ KLDR NS+W MF N+ L S H P+
Sbjct: 131 DMHQMGSSGNSFTWGGTRND-QWIQCKLDRCFGNSEWFTMFSNSHQWFLEKLGSHHRPVL 189
Query: 582 LTCKPVTAVNRNPHRFKFENAWLAEPDFKQQVQQRW---KLYPEEGITQKLSYCAEDLTD 638
+ V R +F ++ + +P W + ++ C + ++
Sbjct: 190 VNFVNDQEVFRG--QFCYDKRFAEDPQCAASTLSSWIGNGISDVSSSMLRMVKCRKAISG 247
Query: 639 WSRNNN-NFRRDISKVQKKI--EKLRTHVTAANVSYFNSLKNKLDKLLVQDDLFWKQRAK 695
W +N++ N + I +++ ++ EK + + + +S ++ +L +++ FW+ ++K
Sbjct: 248 WKKNSDFNAQNRILRLRSELDEEKSKQYPCWSRISV---IQTQLGVAFREEESFWRLKSK 304
Query: 696 TFWYKEGDLNTKFFHAAATSRRKVNRIEHLENSNGTVCRKEDELQNIARNYFSHLF-TKL 754
W GD N+KFF A + R N + L + NG E NIA YF +LF +
Sbjct: 305 DKWLFGGDRNSKFFQAMVKANRTKNSLRFLVDENGNEHTLNREKGNIASVYFENLFMSSY 364
Query: 755 SSTRADVISKVATSISDDDNCFLTAAFTLEEFKTAAFSMQADKCPGPDGFNPGFYQNFWN 814
+ + T +S++ N LT A T E +A FS+ + P
Sbjct: 365 PANSQSALDGFKTRVSEEMNQELTQAVTELEIHSAVFSINVESAPEK-----------LE 413
Query: 815 MCGSEIHKAGCEWLENGVFPPHLNSTNIALIPKGDAQTTMKDWRPIALCNVVYKMVAKVL 874
C + + E GV P N T++ LIPK M D RPI+LC+V+YK+++K+L
Sbjct: 414 CCQGSDYIEILGFFETGVLPQEWNHTHLYLIPKFTNPQRMSDIRPISLCSVLYKIISKIL 473
Query: 875 ANRLKNVLDKCISTSQSAFVPGRSILDNALVAIELIHYMKAKTKGSQGDVALKLDISKAY 934
+ +LK L +S SQSAF R I DN L+A E++H ++ K S+ + K D+SKAY
Sbjct: 474 SFKLKKHLPSIVSPSQSAFFAERLISDNILIAHEIVHSLRTNDKISKEFMVFKTDMSKAY 533
Query: 935 DRLDWEYLRDIMIQMRFSEKWIRWIMLCVETVDYTVLVNGVQVGPLIPGRGIRQGDPLSP 994
DR++W +L++I++ + F++KWI WIM CV +V Y+VL+NG G + P RGIRQGDP+SP
Sbjct: 534 DRVEWSFLQEILVALGFNDKWISWIMGCVTSVTYSVLINGQHFGHITPERGIRQGDPISP 593
Query: 995 YLFILCAEGLSALIRDAERRGVITGTRICTSAPAISHLLFADDCFLFFRASEQEASVMKN 1054
+LF+LC E L +++ AE ++G + S P+++HLLF DD L RA++ + M
Sbjct: 594 FLFVLCTEALIHILQQAENSKKVSGIQFNGSGPSVNHLLFVDDTQLVCRATKSDCEQMML 653
Query: 1055 ILTTYEAASGQAINLQKSEMYCSRNTPTECQNRIATTLGVKQVLGTGKYLGLPSMIGRSK 1114
L+ Y SGQ IN++KS + + + I G+ GTGKYLGLP + SK
Sbjct: 654 CLSQYGHISGQLINVEKSSITFGVKVDEDTKRWIKNRSGIHLEGGTGKYLGLPENLSGSK 713
Query: 1115 KATFKFVKDRIWNRINSWSSRCLSQAGREILIKSVLQSIPSYVMSIFLLPSTFIGEIEKM 1174
+ F ++K+++ + ++ W + LSQ G+EIL+KS+ ++P Y+M+ F LP ++ +
Sbjct: 714 QDLFGYIKEKLQSHLSGWYDKTLSQGGKEILLKSIALALPVYIMTCFRLPKGLCTKLTSV 773
Query: 1175 LNAFWWGHNSANSRGMHWLSWERLSVPKTFGGMGFKSLRAFNLAMIGKQAWKLISNPDSL 1234
+ FWW ++ S +HW+ ++L++PK+ GG GFK L+ FN A++ KQAW+L S+ S+
Sbjct: 774 MMDFWW-NSMEFSNKIHWIGGKKLTLPKSLGGFGFKDLQCFNQALLAKQAWRLFSDSKSI 832
Query: 1235 ITKLLKAKYFPHSDYFSASIGHNPSYVWRSLWSVRDYLKHGLKWSIGTGETISVWNQPWL 1294
++++ K++YF ++D+ +A G PSY WRS+ R+ L GLK IG GE +VW WL
Sbjct: 833 VSQIFKSRYFMNTDFLNARQGTRPSYTWRSILYGRELLNGGLKRLIGNGEQTNVWIDKWL 892
Query: 1295 KDPLCLQPVTEVQIMWDALTVGHLFKPNTKEWNENFIRYVFNAETSNQIL-QTPLLQSVQ 1353
D +P+ +M + V HL P T+ WN + +F+ + I+ Q PL+ S
Sbjct: 893 FDGHSRRPMNLHSLMNIHMKVSHLIDPLTRNWNLKKLTELFHEKDVQLIMHQRPLISS-- 950
Query: 1354 VDKATWRFEKNGLYSVRSAY----REIINR--NDVLLQHRVPGKWNIIWNLKLPPKIKNF 1407
D W NGLY+V+S Y RE + + V ++ +W+L+ PKIK F
Sbjct: 951 EDSYCWAGTNNGLYTVKSGYERSSRETFKNLFKEADVYPSVNPLFDKVWSLETVPKIKVF 1010
Query: 1408 LWRVCRNCLPTRMRLISKGVQCPPTCAICNNHEEDGKHLFFACSKSVGCWQRLGFWPSIQ 1467
+W+ + L RL S+G++ C C E HL F C P +
Sbjct: 1011 MWKALKGALAVEDRLRSRGIRTADGCLFCKEEIETINHLLFQC-------------PFAR 1057
Query: 1468 QIWSHNAFCADI------IFSLLQQLDIAHQQIFAV----------TLWSLWRHRNNKVW 1511
Q+W+ + A IFS + + I + Q F + LW +W++RN ++
Sbjct: 1058 QVWALSLIQAPATGFGTSIFSNINHV-IQNSQNFGIPRHMRTVSPWLLWEIWKNRNKTLF 1116
Query: 1512 NNIVETTEEIGDRAVAFLNSWKAAQETRIRSSPANPHFDISKWSKPSVGRFKCNVDAAFS 1571
T+ EI +A N W AQE + H KW+ P G KCN+ A+S
Sbjct: 1117 QGTGLTSSEIVAKAYEECNLWINAQEKSSGGVSPSEH----KWNPPPAGELKCNIGVAWS 1172
Query: 1572 ASLHRVGFGACIRDANGNHVISRTECFTPLLDVEMGEAIGLLHAMRWAKDLNLVNMDFET 1631
G +RD+ G ++ ++ + + + A+ + + F
Sbjct: 1173 RQKQLAGVSWVLRDSMGQVLLHSRRSYSQVYSLFDAKIKSWDWALESMDHFHFDKVTFAA 1232
Query: 1632 DSKVVVENIYKGDGVSDFMAIIHDCRHLLMT-DLANSDVKFIRRQANSAAHNLAREALNH 1690
S +++ ++K + M I H L T D+++ + Q N+ A +A+ +
Sbjct: 1233 TSHDIIKALHKPNEWP--MLIGHIAEFLSFTKDISDWFMMMESTQCNNGAAEIAKSVITG 1290
Query: 1691 ASFHYHL--NIPHCIHTLINNEK 1711
+ ++ + PH + +L E+
Sbjct: 1291 MRWQSYVACSFPHWLRSLFEEER 1313
>UniRef100_Q9SN65 Hypothetical protein F25I24.40 [Arabidopsis thaliana]
Length = 1294
Score = 627 bits (1618), Expect = e-178
Identities = 343/933 (36%), Positives = 517/933 (54%), Gaps = 16/933 (1%)
Query: 362 RGLGSPSTVPTLKYLVRTYKPEGIFLSETMAALNKIEELKYILSFDSCFTVDRIGRGGGL 421
+G+G P T L L + +K + +FL ET+ I L +L F + T G GGL
Sbjct: 370 KGIGVPLTQSQLSNLCKVFKFDVLFLIETLNKCEVISNLASVLGFPNVITQPPQGHSGGL 429
Query: 422 AFLWKNSANCTITNFSQNHIDVEVDDLLIGKWRLTGFYGMPENGRRKESWAFLKNLARTS 481
A LWK+S + HIDV + I + L+ YG P R W +NL++T
Sbjct: 430 ALLWKDSVRLSNLYQDDRHIDVHISINNINFY-LSRVYGHPCQSERHSLWTHFENLSKTR 488
Query: 482 SLPWCVIGDFNDILSSEEKKGRSERAPWLIRGFRQAVLDAGLFDLHMTGYAFTWFKSLGT 541
+ PW +IGDFN+ILS+ EK G +R W RGFR V L D+ G F+W +
Sbjct: 489 NDPWILIGDFNEILSNNEKIGGPQRDEWTFRGFRNMVSTCDLKDIRSIGDRFSWVGERHS 548
Query: 542 ARAVEEKLDRALVNSDWCNMFPNAGVECLSTTSSDHYPLWLTCKPVTAVNRNPHRFKFEN 601
V+ LDRA +NS+ +FP A +E L T SDH PL+L+ + P F+F+
Sbjct: 549 -HTVKCCLDRAFINSEGAFLFPFAELEFLEFTGSDHKPLFLSLEKTETRKMRP--FRFDK 605
Query: 602 AWLAEPDFKQQVQQRWKLY---PEEGITQKLSYCAEDLTDWSRNNN-NFRRDISKVQKKI 657
L P FK V+ W + + ++ C + + +N N R I+++Q +
Sbjct: 606 RLLEVPHFKTYVKAGWNKAINGQRKHLPDQVRTCRQAMAKLKHKSNLNSRIRINQLQAAL 665
Query: 658 EKLRTHVTAANVSYFNSLKNKLDKLLVQDDLFWKQRAKTFWYKEGDLNTKFFHAAATSRR 717
+K + V + ++ +L ++ +W+Q+++ W KEGD NT+FFHA +R
Sbjct: 666 DKAMSSVNRTERRTISHIQRELTVAYRDEERYWQQKSRNQWMKEGDRNTEFFHACTKTRF 725
Query: 718 KVNRIEHLENSNGTVCRKEDELQNIARNYFSHLFTKLSSTRADV----ISKVATSISDDD 773
VNR+ +++ G + R + E+ A+ +F+ ++ + + + T +DD
Sbjct: 726 SVNRLVTIKDEEGMIYRGDKEIGVHAQEFFTKVYESNGRPVSIIDFAGFKPIVTEQINDD 785
Query: 774 NCFLTAAFTLEEFKTAAFSMQADKCPGPDGFNPGFYQNFWNMCGSEIHKAGCEWLENGVF 833
LT + E A + DK PGPDG FY++ W + G ++ K +
Sbjct: 786 ---LTKDLSDLEIYNAICHIGDDKAPGPDGLTARFYKSCWEIVGPDVIKEVKIFFRTSYM 842
Query: 834 PPHLNSTNIALIPKGDAQTTMKDWRPIALCNVVYKMVAKVLANRLKNVLDKCISTSQSAF 893
+N TNI +IPK T+ D+RPIALCNV+YK+++K L RLK LD +S SQ+AF
Sbjct: 843 KQSINHTNICMIPKITNPETLSDYRPIALCNVLYKIISKCLVERLKGHLDAIVSDSQAAF 902
Query: 894 VPGRSILDNALVAIELIHYMKAKTKGSQGDVALKLDISKAYDRLDWEYLRDIMIQMRFSE 953
+PGR + DN ++A E++H +K + + SQ +A+K D+SKAYDR++W +L M FSE
Sbjct: 903 IPGRLVNDNVMIAHEMMHSLKTRKRVSQSYMAVKTDVSKAYDRVEWNFLETTMRLFGFSE 962
Query: 954 KWIRWIMLCVETVDYTVLVNGVQVGPLIPGRGIRQGDPLSPYLFILCAEGLSALIRDAER 1013
WI+WIM V++V+Y+VLVNG+ G + P RGIRQGDPLSPYLFILCA+ L+ LI++
Sbjct: 963 TWIKWIMGAVKSVNYSVLVNGIPHGTIQPQRGIRQGDPLSPYLFILCADILNHLIKNRVA 1022
Query: 1014 RGVITGTRICTSAPAISHLLFADDCFLFFRASEQEASVMKNILTTYEAASGQAINLQKSE 1073
G I G RI P ++HL FADD F +++ + +K++ YE SGQ IN+ KS
Sbjct: 1023 EGDIRGIRIGNGVPGVTHLQFADDSLFFCQSNVRNCQALKDVFDVYEYYSGQKINMSKSM 1082
Query: 1074 MYCSRNTPTECQNRIATTLGVKQVLGTGKYLGLPSMIGRSKKATFKFVKDRIWNRINSWS 1133
+ QNR+ LG++ G GKYLGLP GR K+ F ++ +R+ R +SWS
Sbjct: 1083 ITFGSRVHGTTQNRLKNILGIQSHGGGGKYLGLPEQFGRKKRDMFNYIIERVKKRTSSWS 1142
Query: 1134 SRCLSQAGREILIKSVLQSIPSYVMSIFLLPSTFIGEIEKMLNAFWWGHNSANSRGMHWL 1193
++ LS AG+EI++KSV S+P Y MS F LP + EIE +L FWW N A R + W+
Sbjct: 1143 AKYLSPAGKEIMLKSVAMSMPVYAMSCFKLPLNIVSEIEALLMNFWWEKN-AKKREIPWI 1201
Query: 1194 SWERLSVPKTFGGMGFKSLRAFNLAMIGKQAWKLISNPDSLITKLLKAKYFPHSDYFSAS 1253
+W+RL K GG+GF+ L FN A++ KQ W++I+NP+SL +++KA+YF A
Sbjct: 1202 AWKRLQYSKKEGGLGFRDLAKFNDALLAKQVWRMINNPNSLFARIMKARYFREDSILDAK 1261
Query: 1254 IGHNPSYVWRSLWSVRDYLKHGLKWSIGTGETI 1286
SY W S+ + D +K G ++ +G G+T+
Sbjct: 1262 RQRYQSYGWTSMLAGLDVIKKGSRFIVGDGKTV 1294
>UniRef100_O22220 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1094
Score = 562 bits (1449), Expect = e-158
Identities = 325/988 (32%), Positives = 532/988 (52%), Gaps = 34/988 (3%)
Query: 628 KLSYCAEDLTDWSRNNN-NFRRDISKVQKKIEKLRTHVTAANVSYFNSLKNKLDKLLVQD 686
+L C + ++ W R N+ N + I+K++++++K ++ T + + + L++ L ++
Sbjct: 3 RLVECRKAISQWKRENDFNAKSRITKLRRELDKEKS-ATFPSWTQISLLQDVLGDAYREE 61
Query: 687 DLFWKQRAKTFWYKEGDLNTKFFHAAATSRRKVNRIEHLENSNGTVCRKEDELQNIARNY 746
+ FW+ +++ W GD N+KFF A + R N + L + NG E IA +
Sbjct: 62 EDFWRLKSRDKWMVGGDKNSKFFQATVKANRVSNSLRFLVDENGNEQTVNREKGKIAVTF 121
Query: 747 FSHLFTKLSSTRAD-VISKVATSISDDDNCFLTAAFTLEEFKTAAFSMQADKCPGPDGFN 805
F LF+ + D V+ +++D N LT +E A FS+ A+ PGPDGF
Sbjct: 122 FEDLFSSSYPSSMDSVLEGFNKRVTEDMNQDLTKKVNEQEIYKAVFSINAESAPGPDGFT 181
Query: 806 PGFYQNFWNMCGSEIHKAGCEWLENGVFPPHLNSTNIALIPKGDAQTTMKDWRPIALCNV 865
F+Q W + ++I + + G+ P N T++ LIPK M D RPI+LC+V
Sbjct: 182 ALFFQRQWPLVKNQIISDIELFFQTGILPEDWNHTHLCLIPKITKPARMADIRPISLCSV 241
Query: 866 VYKMVAKVLANRLKNVLDKCISTSQSAFVPGRSILDNALVAIELIHYMKAKTKGSQGDVA 925
+YK+++K+L+ RLK L +S +QSAFV R + DN ++A E++H ++ K S+ +
Sbjct: 242 MYKIISKILSARLKKYLPVIVSPTQSAFVAERLVSDNIILAHEIVHNLRTNEKISKDFMV 301
Query: 926 LKLDISKAYDRLDWEYLRDIMIQMRFSEKWIRWIMLCVETVDYTVLVNGVQVGPLIPGRG 985
K D+SKAYDR++W +L+ I++ + F+ WI W+M CV +V Y+VL+NG G + P RG
Sbjct: 302 FKTDMSKAYDRVEWPFLKGILLALGFNSTWINWMMACVSSVSYSVLINGQPFGHITPHRG 361
Query: 986 IRQGDPLSPYLFILCAEGLSALIRDAERRGVITGTRICTSAPAISHLLFADDCFLFFRAS 1045
+RQGDPLSP+LF+LC E L ++ AE+ G I+G + + P+++HLLFADD L +AS
Sbjct: 362 LRQGDPLSPFLFVLCTEALIHILNQAEKIGKISGIQFNGTGPSVNHLLFADDTLLICKAS 421
Query: 1046 EQEASVMKNILTTYEAASGQAINLQKSEMYCSRNTPTECQNRIATTLGVKQVLGTGKYLG 1105
+ E + + + L+ Y SGQ IN +KS + E + I G++ GTGKYLG
Sbjct: 422 QLECAEIMHCLSQYGHISGQMINSEKSAITFGAKVNEETKQWIMNRSGIQTEGGTGKYLG 481
Query: 1106 LPSMIGRSKKATFKFVKDRIWNRINSWSSRCLSQAGREILIKSVLQSIPSYVMSIFLLPS 1165
LP SK+ F F+K+++ +R++ W ++ LSQ G++IL+KS+ + P Y M+ F L
Sbjct: 482 LPECFQGSKQVLFGFIKEKLQSRLSGWYAKTLSQGGKDILLKSIAMAFPVYAMTCFRLSK 541
Query: 1166 TFIGEIEKMLNAFWWGHNSANSRGMHWLSWERLSVPKTFGGMGFKSLRAFNLAMIGKQAW 1225
T ++ ++ FWW ++ + + +HW+ ++L +PK GG GFK L+ FN A++ KQA
Sbjct: 542 TLCTKLTSVMMDFWW-NSVQDKKKIHWIGAQKLMLPKFLGGFGFKDLQCFNQALLAKQAS 600
Query: 1226 KLISNPDSLITKLLKAKYFPHSDYFSASIGHNPSYVWRSLWSVRDYLKHGLKWSIGTGET 1285
+L ++ DSL++++LK++Y+ +SD+ SA+ G PSY W+S+ R+ L GLK IG GE
Sbjct: 601 RLHTDSDSLLSQILKSRYYMNSDFLSATKGTRPSYAWQSILYGRELLVSGLKKIIGNGEN 660
Query: 1286 ISVWNQPWLKDPLCLQPVTEVQIMWD-ALTVGHLFKPNTKEWNENFIRYVFNAETSNQIL 1344
VW W+ D +P +QIM D L V L P ++ WN N +R +F + I
Sbjct: 661 TYVWMDNWIFDDKPRRP-ESLQIMVDIQLKVSQLIDPFSRNWNLNMLRDLFPWKEIQIIC 719
Query: 1345 QTPLLQSVQVDKATWRFEKNGLYSVRSAY----REIINR--NDVLLQHRVPGKWNIIWNL 1398
Q + S Q D W +GLY+V+S Y R++ + + Q + + IWNL
Sbjct: 720 QQRPMASRQ-DSFCWFGTNHGLYTVKSEYDLCSRQVHKQMFKEAEEQPSLNPLFGKIWNL 778
Query: 1399 KLPPKIKNFLWRVCRNCLPTRMRLISKGVQCPPTCAICNNHEEDGKHLFFACSKSVGCWQ 1458
PKIK FLW+V + + RL ++GV C++C E H+ F C + W
Sbjct: 779 NSAPKIKVFLWKVLKGAVAVEDRLRTRGVLIEDGCSMCPEKNETLNHILFQCPLARQVW- 837
Query: 1459 RLGFWPSIQQIWSHNAFCADIIFSLLQ---------QLDIAHQQIFAVTLWSLWRHRNNK 1509
++ + S N D IF+ + +L + + +W LW++RN +
Sbjct: 838 ------ALTPMQSPNHGFGDSIFTNVNHVIGNCHNTELSPHLRYVSPWIIWILWKNRNKR 891
Query: 1510 VWNNIVETTEEIGDRAVAFLNSWKAAQETRIRSSPANPHFDISKWSKPSVGRFKCNVDAA 1569
++ I + I +A+ W A E P D++ W P + KCN+ A
Sbjct: 892 LFEGIGSVSLSIVGKALEDCKEWLKAHELICSKEPTK---DLT-WIPPLMNELKCNIGIA 947
Query: 1570 FSASLHRVGFGACIRDANGNHVISRTEC 1597
+S G +R+ G V+ + C
Sbjct: 948 WSKKHQMAGVSWVVRNWKG-RVLLHSRC 974
>UniRef100_Q8S6P1 Putative reverse transcriptase [Oryza sativa]
Length = 1509
Score = 560 bits (1442), Expect = e-157
Identities = 318/887 (35%), Positives = 454/887 (50%), Gaps = 74/887 (8%)
Query: 760 DVISKVATSISDDDNCFLTAAFTLEEFKTAAFSMQADKCPGPDGFNPGFYQNFWNMCGSE 819
+++ V +S N L A FT EE K A ++ K PGPDG GFY+ W++ G +
Sbjct: 633 NLLDVVDRKVSGAMNESLRAEFTREEVKEALDAIGDLKAPGPDGMPAGFYKACWDVVGEK 692
Query: 820 IHKAGCEWLENGVFPPHLNSTNIALIPKGDAQTTMKDWRPIALCNVVYKMVAKVLANRLK 879
+ E L G P N T I LIPK VLANRLK
Sbjct: 693 VTVEVLEVLRGGAIPEGWNDTTIVLIPK-------------------------VLANRLK 727
Query: 880 NVLDKCISTSQSAFVPGRSILDNALVAIELIHYMKAKTKGSQGDVALKLDISKAYDRLDW 939
+L IS +QSAFVPGR I DN L+A E+ HYM+ K G G A KLD+SKAYDR++W
Sbjct: 728 KILPDVISPAQSAFVPGRLISDNILIAYEMTHYMRNKRSGQVGYAAFKLDMSKAYDRVEW 787
Query: 940 EYLRDIMIQMRFSEKWIRWIMLCVETVDYTVLVNGVQVGPLIPGRGIRQGDPLSPYLFIL 999
+L D+M+++ F W+ IM CV TV Y + VNG P RG+RQGDPLSPYLF+L
Sbjct: 788 SFLHDMMLKLGFHTDWVNLIMKCVSTVTYRIRVNGELSESFSPERGLRQGDPLSPYLFLL 847
Query: 1000 CAEGLSALIRDAERRGVITGTRICTSAPAISHLLFADDCFLFFRASEQEASVMKNILTTY 1059
CAEG SAL+ E G + G RIC AP++SHLLFADD + RA+ EA ++ IL Y
Sbjct: 848 CAEGFSALLSKTEEEGRLHGIRICQGAPSVSHLLFADDSLILCRANGGEAQQLQTILQIY 907
Query: 1060 EAASGQAINLQKSEMYCSRNTPTECQNRIATTLGVKQVLGTGKYLGLPSMIGRSKKATFK 1119
E SGQ IN KS + S NT + + + L +++ KYLGLP +GRS+ F
Sbjct: 908 EECSGQVINKDKSAVMFSPNTSSLEKGAVMAALNMQRETTNEKYLGLPVFVGRSRTKIFS 967
Query: 1120 FVKDRIWNRINSWSSRCLSQAGREILIKSVLQSIPSYVMSIFLLPSTFIGEIEKMLNAFW 1179
++K+RIW RI W + LS+AG+EILIK+V Q IP++ M F L +I KM+ +W
Sbjct: 968 YLKERIWQRIQGWKEKLLSRAGKEILIKAVAQVIPTFAMGCFELTKDLCDQISKMIAKYW 1027
Query: 1180 WGHNSANSRGMHWLSWERLSVPKTFGGMGFKSLRAFNLAMIGKQAWKLISNPDSLITKLL 1239
W + +++ MHWLSW +L++PK GG+GF+ + FNLAM+ KQ W+LI +PDSL +++L
Sbjct: 1028 WSNQEKDNK-MHWLSWNKLTLPKNMGGLGFRDIYIFNLAMLAKQGWRLIQDPDSLCSRVL 1086
Query: 1240 KAKYFPHSDYFSASIGHNPSYVWRSLWSVRDYLKHGLKWSIGTGETISVWNQPWLKDPLC 1299
+AKYFP D F N SY WRS+ L++G+ W +G G I++W PW+
Sbjct: 1087 RAKYFPLGDCFRPKQTSNVSYTWRSIQKGLRVLQNGMIWRVGDGSKINIWADPWIPRGWS 1146
Query: 1300 LQPVTEVQIMWDALTVGHLFKPNTKEWNENFIRYVFNAETSNQILQTPLLQSVQVDKATW 1359
+P+T V L P T W+E+ + F E I P+ ++ D W
Sbjct: 1147 RKPMTPRGANL-VTKVEELIDPYTGTWDEDLLSQTFWEEDVAAIKSIPVHVEME-DVLAW 1204
Query: 1360 RFEKNGLYSVRSAYREIINRNDVLLQHRVPGK----------WNIIWNLKLPPKIKNFLW 1409
F+ G ++V+SAY+ ++ PG W +W L +P KIK+FLW
Sbjct: 1205 HFDARGCFTVKSAYKVQREMERRASRNGCPGVSNWESGDDDFWKKLWKLGVPGKIKHFLW 1264
Query: 1410 RVCRNCLPTRMRLISKGVQCPPTCAICNNHEEDGKHLFFACSKSVGCWQRLGFWPSIQQI 1469
R+C N L R L +G+ C +C + ED HLFF C WQ L ++ +
Sbjct: 1265 RMCHNTLALRANLHHRGMDVDTRCVMCGRYNEDAGHLFFKCKPVKKVWQALNL-EELRSM 1323
Query: 1470 WSHNAFCADIIFSLLQQLDIAHQQIFAVTLWSLWRHRNNKVWNNIVETTEEIGDRAVAFL 1529
+++ S+ + + V LW W+ RN E + G+ AV
Sbjct: 1324 LEQQTSGKNVLQSIYCRPENERTSAI-VCLWQWWKERNE-------EKSPRTGECAV--- 1372
Query: 1530 NSWKAAQETRIRSSPANPHFDISKWSKPSVGRFKCNVDAAFSASLHRVGFGACIRDANGN 1589
W +P + K N D A+S+++ + G+G IRD G
Sbjct: 1373 ------------------------WRRPPLNFVKINTDGAYSSNMKQGGWGFVIRDQTGA 1408
Query: 1590 HVISRTECFTPLLDVEMGEAIGLLHAMRWAKDLNLVNMDFETDSKVV 1636
+ + L D E + A++ A + + ++ ETDS ++
Sbjct: 1409 VLQAGAGPAAYLQDAFHAEVVACAAAIKTASERGMSRIELETDSMML 1455
Score = 107 bits (268), Expect = 2e-21
Identities = 80/237 (33%), Positives = 117/237 (48%), Gaps = 9/237 (3%)
Query: 459 YGMPENGRRKESWAFLKNLARTSSLPWCVIGDFNDILSSEEKKGRSERAPWLIRGFRQAV 518
YG + + +W ++ L + PW + GDFN+IL S EK+ +A + FR A+
Sbjct: 397 YGDAHSETKHRTWTTMRGLIDNPTTPWLMAGDFNEILFSHEKQCGRMKAQSAMDEFRHAL 456
Query: 519 LDAGLFDLHMTGYAFTWFK-SLGTARAVEEKLDRALVNSDWCNMFPNAGVECLSTTSSDH 577
D GL DL G AFTW S + E LDRA+ N +W MFP A V SDH
Sbjct: 457 TDCGLDDLGFEGDAFTWRNHSHSQEGYIREWLDRAVANPEWRAMFPAARVINGDPRHSDH 516
Query: 578 YP--LWLTCKPVTAVNRNPHR-FKFENAWLAEPDFKQQVQQRWKLYPE-EG--ITQKLSY 631
P + L K RN H F+FE AWL E FK+ V++ W + +G + L+
Sbjct: 517 RPVIIELEGKNKGVRGRNGHNDFRFEAAWLEEEKFKEVVKEAWDVSAGLQGLLVHASLAG 576
Query: 632 CAEDLTDWSRN-NNNFRRDISKVQKKIEKLRTH-VTAANVSYFNSLKNKLDKLLVQD 686
A L+ WS N + + + KV+K++E R ++ V L+ +L+KL Q+
Sbjct: 577 VAAGLSSWSSNVLGDLEKRLKKVKKELETCRRQPISRDQVVREEVLRYRLEKLEQQN 633
>UniRef100_Q9M1F2 Hypothetical protein F9K21.130 [Arabidopsis thaliana]
Length = 851
Score = 535 bits (1379), Expect = e-150
Identities = 313/843 (37%), Positives = 462/843 (54%), Gaps = 18/843 (2%)
Query: 743 ARNYFSHLFTKLSSTRADV-ISKVATSISDDDNCFLTAAFTLEEFKTAAFSMQADKCPGP 801
A+ +F+ +FT + + + +S+++ N LT F E A + DK PGP
Sbjct: 10 AQKFFTDIFTTNGIQVSPIDFADFPSSVTNIINSELTQDFRDSEIFEAICQIGDDKAPGP 69
Query: 802 DGFNPGFYQNFWNMCGSEIHKAGCEWLENGVFPPHLNSTNIALIPKGDAQTTMKDWRPIA 861
DG FY+ W++ G+++ K + E+ +N TNI +IPK T+ D+RPIA
Sbjct: 70 DGLTARFYKQCWDIVGNDVIKEVKLFFESSHMKTSVNHTNICMIPKIQNPQTLSDYRPIA 129
Query: 862 LCNVVYKMVAKVLANRLKNVLDKCISTSQSAFVPGRSILDNALVAIELIHYMKAKTKGSQ 921
LCNV+YK+++K + NRLK L+ +S SQ+AF+PGR I DN ++A E++H +K + + S+
Sbjct: 130 LCNVLYKVISKCMVNRLKAHLNSIVSDSQAAFIPGRIINDNVMIAHEIMHSLKVRKRVSK 189
Query: 922 GDVALKLDISKAYDRLDWEYLRDIMIQMRFSEKWIRWIMLCVETVDYTVLVNGVQVGPLI 981
+A+K D+SKAYDR++W++L M F +KWI WIM V++V Y+VL+NG G +
Sbjct: 190 TYMAVKTDVSKAYDRVEWDFLETTMRLFGFCDKWIGWIMAAVKSVHYSVLINGSPHGYIS 249
Query: 982 PGRGIRQGDPLSPYLFILCAEGLSALIRDAERRGVITGTRICTSAPAISHLLFADDCFLF 1041
P RGIRQGDPLSPYLFILC + LS LI+ G I G RI APAI+HL FADD F
Sbjct: 250 PTRGIRQGDPLSPYLFILCGDILSHLIKVKASSGDIRGVRIGNGAPAITHLQFADDSLFF 309
Query: 1042 FRASEQEASVMKNILTTYEAASGQAINLQKSEMYCSRNTPTECQNRIATTLGVKQVLGTG 1101
+A+ + +K++ YE SGQ IN+QKS + Q R+ T L + G G
Sbjct: 310 CQANVRNCQALKDVFDVYEYYSGQKINVQKSLITFGSRVYGSTQTRLKTLLNIPNQGGGG 369
Query: 1102 KYLGLPSMIGRSKKATFKFVKDRIWNRINSWSSRCLSQAGREILIKSVLQSIPSYVMSIF 1161
KYLGLP GR KK F ++ DR+ R SWS++ LS AG+EIL+KSV ++P Y MS F
Sbjct: 370 KYLGLPEQFGRKKKEMFNYIIDRVKERTASWSAKFLSPAGKEILLKSVALAMPVYAMSCF 429
Query: 1162 LLPSTFIGEIEKMLNAFWWGHNSANSRGMHWLSWERLSVPKTFGGMGFKSLRAFNLAMIG 1221
LP + EIE +L FWW ++N RG+ W++W+RL K GG+GF+ L FN A++
Sbjct: 430 KLPQGIVSEIESLLMNFWW-EKASNKRGIPWVAWKRLQYSKKEGGLGFRDLAKFNDALLA 488
Query: 1222 KQAWKLISNPDSLITKLLKAKYFPHSDYFSASIGHNPSYVWRSLWSVRDYLKHGLKWSIG 1281
KQAW++I P+SL +++KA+YF + A SY W SL S L+ G ++ IG
Sbjct: 489 KQAWRIIQYPNSLFARVMKARYFKDNSIIDAKTRSQQSYGWSSLLSGIALLRKGTRYVIG 548
Query: 1282 TGETISVWNQPWLKDPLCLQPVTEVQIMWDALTVGHLF--KPNTKEWNENFIRYVFNAET 1339
G+TI + + +T+ Q + L++ +LF + +++ W+ ++ +
Sbjct: 549 DGKTIRLGIDNVVDSHPPRPLLTDEQ--HNGLSLDNLFQHRGHSRCWDNAKLQTFVDQSD 606
Query: 1340 SNQILQTPLLQSVQVDKATWRFEKNGLYSVRSAYREIIN--RNDVLLQHRVPGKWNI--- 1394
+ I + L + D+ W + G Y+VRS Y + N + + G ++
Sbjct: 607 HDYIKRIYLSTRSKTDRLIWSYNSTGDYTVRSGYWLSTHDPSNTIPTMAKPHGSVDLKTK 666
Query: 1395 IWNLKLPPKIKNFLWRVCRNCLPTRMRLISKGVQCPPTCAICNNHEEDGKHLFFACSKSV 1454
IWNL + PK+K+FLWR+ LPT RL ++G++ P C C E H F C +
Sbjct: 667 IWNLPIMPKLKHFLWRILSKALPTTDRLTTRGMRIDPGCPRCRRENESINHALFTCPFAT 726
Query: 1455 GCWQRLGFWPSIQQIWSHNAFCADI--IFSLLQQLDIAHQQ--IFAVTLWSLWRHRNNKV 1510
W RL P + N +I I LLQ I Q I LW +W+ RNN V
Sbjct: 727 MAW-RLSDTPLYRSSILSNNIEDNISNILLLLQNTTITDSQKLIPFWLLWRIWKARNNVV 785
Query: 1511 WNNIVETTEEIGDRAVAFLNSWKAAQETR--IRSSPANPHFDISKWSKPSVGRFKCNVDA 1568
+NN+ E+ RA A N W A +T+ R + W KP + KCN DA
Sbjct: 786 FNNLRESPSITVVRAKAETNEWLNATQTQGPRRLPKRTTAAGNTTWVKPQMPYIKCNFDA 845
Query: 1569 AFS 1571
+F+
Sbjct: 846 SFN 848
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.333 0.142 0.482
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,838,494,874
Number of Sequences: 2790947
Number of extensions: 118130988
Number of successful extensions: 413845
Number of sequences better than 10.0: 1192
Number of HSP's better than 10.0 without gapping: 668
Number of HSP's successfully gapped in prelim test: 524
Number of HSP's that attempted gapping in prelim test: 409576
Number of HSP's gapped (non-prelim): 2215
length of query: 1711
length of database: 848,049,833
effective HSP length: 141
effective length of query: 1570
effective length of database: 454,526,306
effective search space: 713606300420
effective search space used: 713606300420
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.5 bits)
S2: 82 (36.2 bits)
Medicago: description of AC146755.8


