

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146751.11 - phase: 0
(399 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
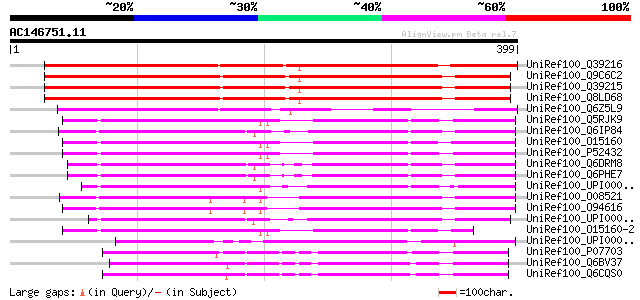
UniRef100_Q39216 RNA polymerase subunit [Arabidopsis thaliana] 475 e-133
UniRef100_Q9C6C2 RNA polymerase subunit; 10595-12672 [Arabidopsi... 431 e-119
UniRef100_Q39215 RNA polymerase subunit [Arabidopsis thaliana] 429 e-119
UniRef100_Q8LD68 RNA polymerase subunit [Arabidopsis thaliana] 429 e-119
UniRef100_Q6Z5L9 Putative RNA polymerase subunit [Oryza sativa] 292 1e-77
UniRef100_Q5RJK9 Polr1c protein [Rattus norvegicus] 266 6e-70
UniRef100_Q6IP84 MGC78824 protein [Xenopus laevis] 266 8e-70
UniRef100_O15160 DNA-directed RNA polymerase I 40 kDa polypeptid... 266 1e-69
UniRef100_P52432 DNA-directed RNA polymerase I 40 kDa polypeptid... 265 2e-69
UniRef100_Q6DRM8 RNA polymerase I 140 kDa subunit [Brachydanio r... 259 7e-68
UniRef100_Q6PHE7 RNA polymerase 1-1 [Brachydanio rerio] 259 9e-68
UniRef100_UPI00003ACD10 UPI00003ACD10 UniRef100 entry 257 5e-67
UniRef100_O08521 Homologous to 40kD subunit of RNA-polymerase I ... 236 8e-61
UniRef100_O94616 DNA-directed RNA polymerases I and III 40 kDa p... 236 1e-60
UniRef100_UPI000042E7AF UPI000042E7AF UniRef100 entry 228 3e-58
UniRef100_O15160-2 Splice isoform 2 of O15160 [Homo sapiens] 224 4e-57
UniRef100_UPI0000366163 UPI0000366163 UniRef100 entry 218 3e-55
UniRef100_P07703 DNA-directed RNA polymerases I and III 40 kDa p... 216 7e-55
UniRef100_Q6BV37 Similar to sp|P07703 Saccharomyces cerevisiae Y... 214 3e-54
UniRef100_Q6CQS0 Kluyveromyces lactis strain NRRL Y-1140 chromos... 214 4e-54
>UniRef100_Q39216 RNA polymerase subunit [Arabidopsis thaliana]
Length = 385
Score = 475 bits (1223), Expect = e-133
Identities = 242/375 (64%), Positives = 295/375 (78%), Gaps = 15/375 (4%)
Query: 28 IMNLEDVPSTLPTHLELQKTRVFCNLDAPQHTDTIQYSGAYAALGVDNSVRLDNFYQNFK 87
I +L DVP+ LP HLELQ+TRV C D+ H I +SGAY+++GVDNSVRL+NF ++FK
Sbjct: 23 IFDLPDVPTGLPPHLELQRTRVVCKKDSNIHPTAITFSGAYSSMGVDNSVRLENFSEDFK 82
Query: 88 VEVKRITDEEMEFDMIGIDPAVANAFRRILIAEVPTMAIERVYIANNTSLVQDEVLSHRL 147
V+V +T+ +M FDMIG+ +ANAFRRIL+AE+P+MAIE+VY+ANNTS++QDEVL+HRL
Sbjct: 83 VDVISLTETDMVFDMIGVHAGIANAFRRILLAELPSMAIEKVYVANNTSVIQDEVLAHRL 142
Query: 148 GLIPIDADPKLFEYPDDAGGENNEKNTIVFKQHVRCEKGQPRLTVKSDTLKWLPNGSELI 207
GLIPI ADP+LFEY + + NEKNTIVFK HV+C KG PR V + LKWLPNGSELI
Sbjct: 143 GLIPIAADPRLFEYLSE-NDQPNEKNTIVFKLHVKCLKGDPRRKVLTSELKWLPNGSELI 201
Query: 208 AEGTKSAADPAPKTFTTFN--QKSLPKFSKDP-APYNLDVIVAKLGPGQEIELEAHAVKG 264
E S PKT+T+FN Q S P+F+++P P D+++AKLGPGQEIELEAHAVKG
Sbjct: 202 KESGGSTT--TPKTYTSFNHSQDSFPEFAENPIRPTLKDILIAKLGPGQEIELEAHAVKG 259
Query: 265 IGKTHAKWSPVATAWYRMLPEVVLTKDVKDELAEELVSKCPAKVFDIEDIGRGRKKAVVK 324
IGKTHAKWSPVATAWYRMLPEVVL K+ + + AEELV CP KVFDIED+G+GRK+A V
Sbjct: 260 IGKTHAKWSPVATAWYRMLPEVVLLKEFEGKHAEELVKVCPKKVFDIEDMGQGRKRATVA 319
Query: 325 NARACTLCRECIREADEGEEGGEGEKWTDRVSLRRVKDHFIFTIESTGAIPPDVLFTEAV 384
R C+LCRECIR +G +W D+V LRRVK+HFIFTIESTG+ PP+VLF EAV
Sbjct: 320 RPRDCSLCRECIR---------DGVEWEDQVDLRRVKNHFIFTIESTGSQPPEVLFNEAV 370
Query: 385 KILEDKCERVITELS 399
KILEDKCERVI+ELS
Sbjct: 371 KILEDKCERVISELS 385
>UniRef100_Q9C6C2 RNA polymerase subunit; 10595-12672 [Arabidopsis thaliana]
Length = 375
Score = 431 bits (1109), Expect = e-119
Identities = 225/371 (60%), Positives = 278/371 (74%), Gaps = 17/371 (4%)
Query: 28 IMNLEDVPSTLPTHLELQKTRVFCNLDAPQHTDTIQYSGAY-AALGVDNSVRLDNFYQNF 86
I +L DVP+ LP HL+ Q+TRV +AP HT + YSG Y ++ D++V+L NFY NF
Sbjct: 15 IDDLPDVPAGLPPHLKAQQTRVVSKNNAPAHTASAIYSGTYVSSTEEDDNVKLGNFYDNF 74
Query: 87 KVEVKRITDEEMEFDMIGIDPAVANAFRRILIAEVPTMAIERVYIANNTSLVQDEVLSHR 146
KV+V +T +MEFDMIGID A ANAFRRILIAEVP+MAIE+V IA NTS++ DEVL+HR
Sbjct: 75 KVDVVSLTKTDMEFDMIGIDAAFANAFRRILIAEVPSMAIEKVLIAYNTSVIIDEVLAHR 134
Query: 147 LGLIPIDADPKLFEYPDDAGGENNEKNTIVFKQHVRCEKGQPRLTVKSDTLKWLPNGSEL 206
+GLIPI ADP+LFEY + + NEKNTIVFK HV+C K +PRL V + LKWLPNGSEL
Sbjct: 135 MGLIPIAADPRLFEYLSEHD-QANEKNTIVFKLHVKCPKNRPRLKVLTSDLKWLPNGSEL 193
Query: 207 IAEGTKSAADPAPKTFTTFN--QKSLPKFSKDP-APYNLDVIVAKLGPGQEIELEAHAVK 263
+ E + P KT+T+F+ Q SLP+F+ +P P +LD+++AKL PGQEIELEAHAVK
Sbjct: 194 LRESENKTSKP--KTYTSFSCSQDSLPEFANNPITPCDLDILIAKLAPGQEIELEAHAVK 251
Query: 264 GIGKTHAKWSPVATAWYRMLPEVVLTKDVKDELAEELVSKCPAKVFDIEDIGRGRKKAVV 323
GIGKTHAKWSPV TAWYRM PEVVL +V+DELAE LV+ CP VFDIED+G+G+K+A V
Sbjct: 252 GIGKTHAKWSPVGTAWYRMHPEVVLRGEVEDELAERLVNVCPQNVFDIEDMGKGKKRATV 311
Query: 324 KNARACTLCRECIREADEGEEGGEGEKWTDRVSLRRVKDHFIFTIESTGAIPPDVLFTEA 383
R CTLC+EC+R+ D D V L VK+HFIF IESTG++PP+VLFTEA
Sbjct: 312 AQPRKCTLCKECVRDDD----------LVDHVDLGSVKNHFIFNIESTGSLPPEVLFTEA 361
Query: 384 VKILEDKCERV 394
VKILE KCE +
Sbjct: 362 VKILEAKCEAI 372
>UniRef100_Q39215 RNA polymerase subunit [Arabidopsis thaliana]
Length = 375
Score = 429 bits (1104), Expect = e-119
Identities = 224/371 (60%), Positives = 277/371 (74%), Gaps = 17/371 (4%)
Query: 28 IMNLEDVPSTLPTHLELQKTRVFCNLDAPQHTDTIQYSGAY-AALGVDNSVRLDNFYQNF 86
I +L DVP+ LP HL+ Q+TRV +AP HT + YSG Y ++ D++V+L NFY NF
Sbjct: 15 IDDLPDVPAGLPPHLKAQQTRVVSKNNAPAHTASAIYSGTYVSSTEEDDNVKLGNFYDNF 74
Query: 87 KVEVKRITDEEMEFDMIGIDPAVANAFRRILIAEVPTMAIERVYIANNTSLVQDEVLSHR 146
KV+V +T +MEFDMIGID A ANAFRRILIAEVP+MAIE+V IA NTS++ DEVL+HR
Sbjct: 75 KVDVVSLTKTDMEFDMIGIDAAFANAFRRILIAEVPSMAIEKVLIAYNTSVIIDEVLAHR 134
Query: 147 LGLIPIDADPKLFEYPDDAGGENNEKNTIVFKQHVRCEKGQPRLTVKSDTLKWLPNGSEL 206
+GLIPI ADP+LFEY + + NEKN IVFK HV+C K +PRL V + LKWLPNGSEL
Sbjct: 135 MGLIPIAADPRLFEYLSEHD-QANEKNNIVFKLHVKCPKNRPRLKVLTSDLKWLPNGSEL 193
Query: 207 IAEGTKSAADPAPKTFTTFN--QKSLPKFSKDP-APYNLDVIVAKLGPGQEIELEAHAVK 263
+ E + P KT+T+F+ Q SLP+F+ +P P +LD+++AKL PGQEIELEAHAVK
Sbjct: 194 LRESENKTSKP--KTYTSFSCSQDSLPEFANNPITPCDLDILIAKLAPGQEIELEAHAVK 251
Query: 264 GIGKTHAKWSPVATAWYRMLPEVVLTKDVKDELAEELVSKCPAKVFDIEDIGRGRKKAVV 323
GIGKTHAKWSPV TAWYRM PEVVL +V+DELAE LV+ CP VFDIED+G+G+K+A V
Sbjct: 252 GIGKTHAKWSPVGTAWYRMHPEVVLRGEVEDELAERLVNVCPQNVFDIEDMGKGKKRATV 311
Query: 324 KNARACTLCRECIREADEGEEGGEGEKWTDRVSLRRVKDHFIFTIESTGAIPPDVLFTEA 383
R CTLC+EC+R+ D D V L VK+HFIF IESTG++PP+VLFTEA
Sbjct: 312 AQPRKCTLCKECVRDDD----------LVDHVDLGSVKNHFIFNIESTGSLPPEVLFTEA 361
Query: 384 VKILEDKCERV 394
VKILE KCE +
Sbjct: 362 VKILEAKCEAI 372
>UniRef100_Q8LD68 RNA polymerase subunit [Arabidopsis thaliana]
Length = 375
Score = 429 bits (1103), Expect = e-119
Identities = 224/371 (60%), Positives = 278/371 (74%), Gaps = 17/371 (4%)
Query: 28 IMNLEDVPSTLPTHLELQKTRVFCNLDAPQHTDTIQYSGAY-AALGVDNSVRLDNFYQNF 86
I +L DVP+ LP HL+ Q+TRV +AP HT + YSG Y ++ D++V+L NFY NF
Sbjct: 15 IDDLPDVPAGLPPHLKAQQTRVVSKNNAPAHTASAIYSGTYVSSTEEDDNVKLGNFYDNF 74
Query: 87 KVEVKRITDEEMEFDMIGIDPAVANAFRRILIAEVPTMAIERVYIANNTSLVQDEVLSHR 146
KV+V +T +MEFDMIGID A ANAFRRILIAEVP+MAIE+V+IA NTS++ DEVL+HR
Sbjct: 75 KVDVVSLTKTDMEFDMIGIDAAFANAFRRILIAEVPSMAIEKVFIAYNTSVIIDEVLAHR 134
Query: 147 LGLIPIDADPKLFEYPDDAGGENNEKNTIVFKQHVRCEKGQPRLTVKSDTLKWLPNGSEL 206
+GLIPI ADP+LFEY + + NEKNTIVFK HV+C K +PRL V + LKWLPNGSEL
Sbjct: 135 MGLIPIAADPRLFEYLSEHD-QANEKNTIVFKLHVKCPKNRPRLKVLTSDLKWLPNGSEL 193
Query: 207 IAEGTKSAADPAPKTFTTFN--QKSLPKFSKDP-APYNLDVIVAKLGPGQEIELEAHAVK 263
+ E + KT+T+F+ Q SLP+F+ +P P +LD+++AKL PGQEIELEAHAVK
Sbjct: 194 LRESENKTSKL--KTYTSFSCSQDSLPEFANNPITPCDLDILIAKLAPGQEIELEAHAVK 251
Query: 264 GIGKTHAKWSPVATAWYRMLPEVVLTKDVKDELAEELVSKCPAKVFDIEDIGRGRKKAVV 323
GIGKTHAKWSPV TAWYRM PEVVL +V+DELAE LV+ CP VFDIED+G+G+K+A V
Sbjct: 252 GIGKTHAKWSPVGTAWYRMHPEVVLRGEVEDELAERLVNVCPQNVFDIEDMGKGKKRATV 311
Query: 324 KNARACTLCRECIREADEGEEGGEGEKWTDRVSLRRVKDHFIFTIESTGAIPPDVLFTEA 383
R CTLC+EC+R+ D D V L VK+HFIF IESTG++PP+VLFTEA
Sbjct: 312 AQPRKCTLCKECVRDDD----------LVDHVDLGSVKNHFIFNIESTGSLPPEVLFTEA 361
Query: 384 VKILEDKCERV 394
VKILE KCE +
Sbjct: 362 VKILEAKCEAI 372
>UniRef100_Q6Z5L9 Putative RNA polymerase subunit [Oryza sativa]
Length = 300
Score = 292 bits (747), Expect = 1e-77
Identities = 167/367 (45%), Positives = 213/367 (57%), Gaps = 93/367 (25%)
Query: 38 LPTHLELQKTRVFCNLDAPQHTDTIQYSGAYAALGVDNSVRLDNFYQNFKVEVKRITDEE 97
LP HLE+Q+TRV C DAP +T+ QY+GAY+A+G+DNSV + F +NFKVE+ R+T+++
Sbjct: 22 LPEHLEVQRTRVVCKADAPVNTEGFQYAGAYSAMGIDNSVSAEKFCKNFKVEISRLTEDD 81
Query: 98 MEFDMIGIDPAVANAFRRILIAEVPTMAIERVYIANNTSLVQDEVLSHRLGLIPIDADPK 157
MEFDMIGID ++ANAFRRILIAE+PTMAIE+V + +NTS++ DEVLSHRLGLIP+DADP+
Sbjct: 82 MEFDMIGIDASIANAFRRILIAELPTMAIEKVLMVDNTSVIADEVLSHRLGLIPLDADPR 141
Query: 158 LFEYPDDAGGENNEKNTIVFKQHVRCEKGQPRLTVKSDTLKWLPNGSELIAEGTKSAADP 217
FEY + NE+NTIV+K HV C+KG PRLTVKS L+WLP GS L A P
Sbjct: 142 HFEYMSE-NDVPNERNTIVYKLHVSCKKGSPRLTVKSGDLEWLPEGSRL------PLASP 194
Query: 218 AP-----KTFTTFNQKSLPKFSKDPAPYNLDVIVAKLGPGQEIELEAHAVKGIGKTHAKW 272
A KT+T+F+Q K D+ +A+LGPGQ
Sbjct: 195 AQSRYKQKTYTSFSQSQKDILEKPLGVKFKDITIARLGPGQ------------------- 235
Query: 273 SPVATAWYRMLPEVVLTKDVKDELAEELVSKCPAKVFDIEDIGRGRKKAVVKNARACTLC 332
VV K++K + AE+LV KCP VFDIED+G
Sbjct: 236 -------------VVFRKEIKGDNAEKLVKKCPVNVFDIEDLGN---------------- 266
Query: 333 RECIREADEGEEGGEGEKWTDRVSLRRVKDHFIFTIESTGAIPPDVLFTEAVKILEDKCE 392
TIESTG +PP+ LFTEAV+ILE+KCE
Sbjct: 267 ---------------------------------VTIESTGGLPPEALFTEAVRILEEKCE 293
Query: 393 RVITELS 399
RVI+ LS
Sbjct: 294 RVISGLS 300
>UniRef100_Q5RJK9 Polr1c protein [Rattus norvegicus]
Length = 346
Score = 266 bits (681), Expect = 6e-70
Identities = 157/372 (42%), Positives = 217/372 (58%), Gaps = 53/372 (14%)
Query: 42 LELQKTRVFCNLDAPQHTDTIQYSGAYAALGVDNSVRLDNFYQNFKVEVKRITDEEMEFD 101
+E +TRV ++ T + G Y+ G D++ D F +NF+V+V + + +EFD
Sbjct: 7 VEEMRTRVVLGEFGVRNVHTTDFPGNYS--GYDDAWDQDRFEKNFRVDVVHMDESTLEFD 64
Query: 102 MIGIDPAVANAFRRILIAEVPTMAIERVYIANNTSLVQDEVLSHRLGLIPIDADPKLFEY 161
M+GID A+ANAFRRIL+AEVPTMA+E+V + NNTS+VQDE+L+HRLGLIPI ADP+LFEY
Sbjct: 65 MVGIDAAIANAFRRILLAEVPTMAVEKVLVYNNTSIVQDEILAHRLGLIPILADPRLFEY 124
Query: 162 PDDAGGENNEKNTIVFKQHVRCEKGQPRLTVKSDT-------------LKWLP--NGSEL 206
+ E E +T+ F+ VRC + SD + W+P N +++
Sbjct: 125 RNPGEEEGTEIDTLQFRLQVRCTRNPSAAKDSSDPNELYVNHKVYTRHMTWVPLGNQADV 184
Query: 207 IAEGTKSAADPAPKTFTTFNQKSLPKFSKDPAPYNLDVIVAKLGPGQEIELEAHAVKGIG 266
EGT P + D+++A+L PGQEI+L H VKGIG
Sbjct: 185 FPEGTVR-------------------------PVHDDILIAQLRPGQEIDLMMHCVKGIG 219
Query: 267 KTHAKWSPVATAWYRMLPEVVLTKDVKDELAEELVSKCPAKVFDIEDIGRGRKKAVVKNA 326
K HAK+SPVATA YR+LP++ L + V+ E AEEL V ++++ +G+K A V NA
Sbjct: 220 KDHAKFSPVATASYRLLPDITLLEPVEGEAAEELSQCFSPGVIAVQEV-QGKKVARVANA 278
Query: 327 RACTLCRECIREADEGEEGGEGEKWTDRVSLRRVKDHFIFTIESTGAIPPDVLFTEAVKI 386
R T RE R EK V L RV+DH+IF++ESTG +PPDVL +EA+KI
Sbjct: 279 RLDTFSREIFRH----------EKLKKAVRLARVRDHYIFSVESTGVLPPDVLVSEAIKI 328
Query: 387 LEDKCERVITEL 398
L KC R + EL
Sbjct: 329 LMGKCRRFLDEL 340
>UniRef100_Q6IP84 MGC78824 protein [Xenopus laevis]
Length = 347
Score = 266 bits (680), Expect = 8e-70
Identities = 149/374 (39%), Positives = 225/374 (59%), Gaps = 50/374 (13%)
Query: 39 PTHLELQKTRVFCNLDAPQHTDTIQYSGAYAALGVDNSVRLDNFYQNFKVEVKRITDEEM 98
P ++L + RV ++ T + G Y +G D++ F +NF++++ ++ D+ +
Sbjct: 4 PGSVQLIRERVVLGEFGLRNVHTTDFPGNY--MGYDDTWNQGRFEKNFRIDIVKMEDDIL 61
Query: 99 EFDMIGIDPAVANAFRRILIAEVPTMAIERVYIANNTSLVQDEVLSHRLGLIPIDADPKL 158
EFDM+GID ++ANAFRRIL++EVPT+A+E+VY+ NNTS++QDE+L+HRLGLIPI ADP+L
Sbjct: 62 EFDMVGIDASIANAFRRILLSEVPTVAVEKVYVYNNTSIIQDEILAHRLGLIPIRADPRL 121
Query: 159 FEYPDDAGGENNEKNTIVFKQHVRCEKGQPRLT--------------VKSDTLKWLPNGS 204
FEY E E +T+ F+ +V+C + PR + V + +KWLP G+
Sbjct: 122 FEYRSADDAEGTEIDTLQFELNVKCTR-NPRASKDCSDPNELYLNHKVYTGHMKWLPQGN 180
Query: 205 ELIAEGTKSAADPAPKTFTTFNQKSLPKFSKDPAPYNLDVIVAKLGPGQEIELEAHAVKG 264
++ D P+T P P + D+++A+L PGQEI + H VKG
Sbjct: 181 QV---------DLFPET-------------DPPRPVHKDILIAQLRPGQEINVIMHCVKG 218
Query: 265 IGKTHAKWSPVATAWYRMLPEVVLTKDVKDELAEELVSKCPAKVFDIEDIGRGRKKAVVK 324
IGK HAK+SPVATA YR+LPEV L ++ ELA++L V ++++ G++ A V+
Sbjct: 219 IGKDHAKFSPVATASYRLLPEVTLLSSIEGELADKLQGCFSPGVISVDEV-NGKRVARVE 277
Query: 325 NARACTLCRECIREADEGEEGGEGEKWTDRVSLRRVKDHFIFTIESTGAIPPDVLFTEAV 384
NAR T RE R D + V + RV+DH+IF++ESTG +PPD+L +EA+
Sbjct: 278 NARMDTCSREIFRHDD----------LKNLVRMARVRDHYIFSVESTGILPPDILVSEAI 327
Query: 385 KILEDKCERVITEL 398
K+L KC R + EL
Sbjct: 328 KVLMGKCRRFLDEL 341
>UniRef100_O15160 DNA-directed RNA polymerase I 40 kDa polypeptide [Homo sapiens]
Length = 346
Score = 266 bits (679), Expect = 1e-69
Identities = 155/372 (41%), Positives = 217/372 (57%), Gaps = 53/372 (14%)
Query: 42 LELQKTRVFCNLDAPQHTDTIQYSGAYAALGVDNSVRLDNFYQNFKVEVKRITDEEMEFD 101
+E ++RV ++ T + G Y+ G D++ D F +NF+V+V + + +EFD
Sbjct: 7 VEEMRSRVVLGEFGVRNVHTTDFPGNYS--GYDDAWDQDRFEKNFRVDVVHMDENSLEFD 64
Query: 102 MIGIDPAVANAFRRILIAEVPTMAIERVYIANNTSLVQDEVLSHRLGLIPIDADPKLFEY 161
M+GID A+ANAFRRIL+AEVPTMA+E+V + NNTS+VQDE+L+HRLGLIPI ADP+LFEY
Sbjct: 65 MVGIDAAIANAFRRILLAEVPTMAVEKVLVYNNTSIVQDEILAHRLGLIPIHADPRLFEY 124
Query: 162 PDDAGGENNEKNTIVFKQHVRCEKGQPRLTVKSDT-------------LKWLP--NGSEL 206
+ E E +T+ F+ VRC + SD + W+P N ++L
Sbjct: 125 RNQGDEEGTEIDTLQFRLQVRCTRNPHAAKDSSDPNELYVNHKVYTRHMTWIPLGNQADL 184
Query: 207 IAEGTKSAADPAPKTFTTFNQKSLPKFSKDPAPYNLDVIVAKLGPGQEIELEAHAVKGIG 266
EGT P + D+++A+L PGQEI+L H VKGIG
Sbjct: 185 FPEGTIR-------------------------PVHDDILIAQLRPGQEIDLLMHCVKGIG 219
Query: 267 KTHAKWSPVATAWYRMLPEVVLTKDVKDELAEELVSKCPAKVFDIEDIGRGRKKAVVKNA 326
K HAK+SPVATA YR+LP++ L + V+ E AEEL V +++++ +G+K A V N
Sbjct: 220 KDHAKFSPVATASYRLLPDITLLEPVEGEAAEELSRCFSPGVIEVQEV-QGKKVARVANP 278
Query: 327 RACTLCRECIREADEGEEGGEGEKWTDRVSLRRVKDHFIFTIESTGAIPPDVLFTEAVKI 386
R T RE R EK V L RV+DH+IF++ESTG +PPDVL +EA+K+
Sbjct: 279 RLDTFSREIFR----------NEKLKKVVRLARVRDHYIFSVESTGVLPPDVLVSEAIKV 328
Query: 387 LEDKCERVITEL 398
L KC R + EL
Sbjct: 329 LMGKCRRFLDEL 340
>UniRef100_P52432 DNA-directed RNA polymerase I 40 kDa polypeptide [Mus musculus]
Length = 346
Score = 265 bits (676), Expect = 2e-69
Identities = 157/372 (42%), Positives = 217/372 (58%), Gaps = 53/372 (14%)
Query: 42 LELQKTRVFCNLDAPQHTDTIQYSGAYAALGVDNSVRLDNFYQNFKVEVKRITDEEMEFD 101
+E +TRV ++ T + G YA G D++ + F +NF+V V ++ ++ +EFD
Sbjct: 7 VEEMRTRVVLGEFGVRNVHTTDFPGNYA--GYDDAWDQNRFEKNFRVGVVQMDEDTLEFD 64
Query: 102 MIGIDPAVANAFRRILIAEVPTMAIERVYIANNTSLVQDEVLSHRLGLIPIDADPKLFEY 161
M+GID A+ANAFRRIL+A VPTMA+E+V + NNTS+VQDE+L+HRLGLIPI ADP+LFEY
Sbjct: 65 MVGIDAAIANAFRRILLARVPTMAVEKVLVYNNTSIVQDEILAHRLGLIPILADPRLFEY 124
Query: 162 PDDAGGENNEKNTIVFKQHVRCEKGQPRLTVKSDT-------------LKWLP--NGSEL 206
+ E E +T+ F+ VRC + SD + W+P N +++
Sbjct: 125 RNQGEEEGTEIDTLQFRLQVRCTRNPNAAKDSSDPNELYVNHKVYTRHMTWVPLGNQADV 184
Query: 207 IAEGTKSAADPAPKTFTTFNQKSLPKFSKDPAPYNLDVIVAKLGPGQEIELEAHAVKGIG 266
EGT P + D+++A+L PGQEI+L H VKGIG
Sbjct: 185 FPEGTIR-------------------------PVHDDILIAQLRPGQEIDLMMHCVKGIG 219
Query: 267 KTHAKWSPVATAWYRMLPEVVLTKDVKDELAEELVSKCPAKVFDIEDIGRGRKKAVVKNA 326
K HAK+SPVATA YR+LP + L + V+ E AEEL V ++E++ +G+K A V NA
Sbjct: 220 KDHAKFSPVATASYRLLPAITLLEPVEGEAAEELSQCFSPGVIEVEEV-QGKKVARVANA 278
Query: 327 RACTLCRECIREADEGEEGGEGEKWTDRVSLRRVKDHFIFTIESTGAIPPDVLFTEAVKI 386
R T RE R EK V L RV+DH+IF++ESTG +PPDVL +EA+KI
Sbjct: 279 RLDTFSREIFRH----------EKLKKAVRLARVRDHYIFSVESTGVLPPDVLVSEAIKI 328
Query: 387 LEDKCERVITEL 398
L KC R + EL
Sbjct: 329 LMGKCRRFLDEL 340
>UniRef100_Q6DRM8 RNA polymerase I 140 kDa subunit [Brachydanio rerio]
Length = 343
Score = 259 bits (663), Expect = 7e-68
Identities = 153/367 (41%), Positives = 215/367 (57%), Gaps = 51/367 (13%)
Query: 46 KTRVFCNLDAPQHTDTIQYSGAYAALGVDNSVRLDNFYQNFKVEVKRITDEEMEFDMIGI 105
+ RV ++ T Y G Y G D++ + F +NF++E+ + +EFDMIGI
Sbjct: 8 RDRVILGEFGVKNVHTTDYPGNYP--GYDDTWDFEKFKKNFRIEIISSDENTLEFDMIGI 65
Query: 106 DPAVANAFRRILIAEVPTMAIERVYIANNTSLVQDEVLSHRLGLIPIDADPKLFEYPDDA 165
D A+ANAFRRIL+AEVPTMAIE+V+I NNTS++QDE+L+HRLGL+PI ADP+LFEY +
Sbjct: 66 DAAIANAFRRILLAEVPTMAIEKVFIYNNTSIIQDEILAHRLGLVPIKADPRLFEYRNSE 125
Query: 166 GGENNEKNTIVFKQHVRCEKGQPRLT--------------VKSDTLKWLPNGSELIAEGT 211
E E +TI + V+C + PR + S +KWLP G++
Sbjct: 126 DQEGTEIDTIQLQLKVKCTRN-PRAPKDSSDPKELYLNHMIYSGDIKWLPIGNQ------ 178
Query: 212 KSAADPAPKTFTTFNQKSLPKFSKDPAPYNLDVIVAKLGPGQEIELEAHAVKGIGKTHAK 271
AD F + P + D+++A++ PGQE+++ H VKGIGK HAK
Sbjct: 179 ---AD-------VFADAKI-------GPVHGDILLAQVRPGQELDIVMHCVKGIGKNHAK 221
Query: 272 WSPVATAWYRMLPEVVLTKDVKDELAEELVSKCPAKVFDIEDIGRGRKKAVVKNARACTL 331
+SPVATA YR+LPE+ L + ++ E AE L V ++E++ GRK A V N+R T
Sbjct: 222 FSPVATASYRLLPEITLLQTIEGEQAERLKRCFSPGVIELENV-NGRKVAKVVNSRLDTC 280
Query: 332 CRECIREADEGEEGGEGEKWTDRVSLRRVKDHFIFTIESTGAIPPDVLFTEAVKILEDKC 391
RE +R D V L RV+DHFIF++ESTG +PPDVL TEA+ +L KC
Sbjct: 281 SREVLRHDD----------LKSLVKLGRVRDHFIFSVESTGILPPDVLVTEAINVLMSKC 330
Query: 392 ERVITEL 398
++ +TEL
Sbjct: 331 QKFLTEL 337
>UniRef100_Q6PHE7 RNA polymerase 1-1 [Brachydanio rerio]
Length = 347
Score = 259 bits (662), Expect = 9e-68
Identities = 153/367 (41%), Positives = 215/367 (57%), Gaps = 51/367 (13%)
Query: 46 KTRVFCNLDAPQHTDTIQYSGAYAALGVDNSVRLDNFYQNFKVEVKRITDEEMEFDMIGI 105
+ RV ++ T Y G Y G D++ + F +NF++E+ + +EFDMIGI
Sbjct: 12 RDRVILGEFGVKNVHTTDYPGNYP--GYDDTWDFEKFKKNFRIEIISSDENTLEFDMIGI 69
Query: 106 DPAVANAFRRILIAEVPTMAIERVYIANNTSLVQDEVLSHRLGLIPIDADPKLFEYPDDA 165
D A+ANAFRRIL+AEVPTMAIE+V+I NNTS++QDE+L+HRLGL+PI ADP+LFEY +
Sbjct: 70 DAAIANAFRRILLAEVPTMAIEKVFIYNNTSIIQDEILAHRLGLVPIKADPRLFEYRNSE 129
Query: 166 GGENNEKNTIVFKQHVRCEKGQPRLT--------------VKSDTLKWLPNGSELIAEGT 211
E E +TI + V+C + PR + S +KWLP G++
Sbjct: 130 DQEGTEIDTIQLQLKVKCTRN-PRAPKDSSDPKELYLNHMIYSGDIKWLPIGNQ------ 182
Query: 212 KSAADPAPKTFTTFNQKSLPKFSKDPAPYNLDVIVAKLGPGQEIELEAHAVKGIGKTHAK 271
AD F + P + D+++A++ PGQE+++ H VKGIGK HAK
Sbjct: 183 ---AD-------VFADAKI-------GPVHGDILLAQVRPGQELDIVMHCVKGIGKDHAK 225
Query: 272 WSPVATAWYRMLPEVVLTKDVKDELAEELVSKCPAKVFDIEDIGRGRKKAVVKNARACTL 331
+SPVATA YR+LPE+ L + ++ E AE L V ++E++ GRK A V N+R T
Sbjct: 226 FSPVATASYRLLPEITLLQTIEGEQAERLKRCFSPGVIELENV-NGRKVAKVVNSRLDTC 284
Query: 332 CRECIREADEGEEGGEGEKWTDRVSLRRVKDHFIFTIESTGAIPPDVLFTEAVKILEDKC 391
RE +R D V L RV+DHFIF++ESTG +PPDVL TEA+ +L KC
Sbjct: 285 SREVLRHDD----------LKSLVKLGRVRDHFIFSVESTGILPPDVLVTEAINVLMSKC 334
Query: 392 ERVITEL 398
++ +TEL
Sbjct: 335 QKFLTEL 341
>UniRef100_UPI00003ACD10 UPI00003ACD10 UniRef100 entry
Length = 346
Score = 257 bits (656), Expect = 5e-67
Identities = 153/355 (43%), Positives = 211/355 (59%), Gaps = 49/355 (13%)
Query: 57 QHTDTIQYSGAYAALGVDNSVRLDNFYQNFKVEVKRITDEEMEFDMIGIDPAVANAFRRI 116
++ T + G Y G D++ F + F+V+V R D +EFDM+GID A+ANAFRRI
Sbjct: 22 RNVHTTDFPGNYP--GYDDAWDQRRFEEAFRVDVVREEDGVLEFDMVGIDAAIANAFRRI 79
Query: 117 LIAEVPTMAIERVYIANNTSLVQDEVLSHRLGLIPIDADPKLFEYPDDAGGENNEKNTIV 176
L+AEVPTMA+E+V++ NNTS+VQDE+L+HRLGLIPI ADP+LFEY + E E +T+
Sbjct: 80 LLAEVPTMAVEKVFVYNNTSIVQDEILAHRLGLIPIRADPRLFEYRNQGDEEGTEIDTLQ 139
Query: 177 FKQHVRCEKGQPRLTVKSDT-------------LKWLPNGSELIAEGTKSAADPAPKTFT 223
F+ ++C + SD + W+P G++ D P
Sbjct: 140 FQLKIKCSRNPQAAKESSDPNELYFNHKVYSKHMTWVPLGNQ---------TDLFP---- 186
Query: 224 TFNQKSLPKFSKDPAPYNLDVIVAKLGPGQEIELEAHAVKGIGKTHAKWSPVATAWYRML 283
D P + D+++A L PGQEI++ H VKGIGK HAK+SPVATA YR+L
Sbjct: 187 ----------DADFRPVHDDILIALLRPGQEIDVLMHCVKGIGKDHAKFSPVATASYRLL 236
Query: 284 PEVVLTKDVKDELAEELVSKCPAKVFDIEDIGRGRKKAVVKNARACTLCRECIREADEGE 343
P++ L + V+DE AE L V +I++I +G+K A V NAR T RE R
Sbjct: 237 PDITLLQPVEDEAAETLQKCFSPGVIEIQNI-KGKKVARVANARLDTFSREVFRH----- 290
Query: 344 EGGEGEKWTDRVSLRRVKDHFIFTIESTGAIPPDVLFTEAVKILEDKCERVITEL 398
EG K + V L RV++H+IF++ESTG +PPDVL TEA+KIL KC+R + EL
Sbjct: 291 ---EGLK--NLVRLARVRNHYIFSVESTGILPPDVLVTEAIKILMGKCQRFLNEL 340
>UniRef100_O08521 Homologous to 40kD subunit of RNA-polymerase I and III [Cricetulus
griseus]
Length = 347
Score = 236 bits (602), Expect = 8e-61
Identities = 144/377 (38%), Positives = 211/377 (55%), Gaps = 54/377 (14%)
Query: 40 THLELQKTRVFCNLDAPQHTDTIQYSGAYAALGVDNSVRLDNFYQNFKVEVKRITDEEME 99
T + +T + D ++ + G Y DN LD F +N KV + + E M
Sbjct: 1 TRPDRSRTEISVLSDRVTDVGSVDFPGYY--FDEDNIWDLDKFKKNLKVSITSLDQETMV 58
Query: 100 FDMIGIDPAVANAFRRILIAEVPTMAIERVYIANNTSLVQDEVLSHRLGLIPIDADP--- 156
F++ GID ++ANAFRRILIAE+PT+A E VYI NNTS++QDEVLSHR+GL+PI ADP
Sbjct: 59 FEISGIDASIANAFRRILIAEIPTLAFEFVYIINNTSIIQDEVLSHRIGLVPISADPDMF 118
Query: 157 KLFEYP-DDAGGENNEKNTIVFKQHVRC---------EKGQPRLTVKSDT----LKWLPN 202
K F++P + + +T+VF + +C EK RL V S+ L W P
Sbjct: 119 KWFQHPLPGQEATHTDYDTVVFSLNKKCEFNKNAATDEKDPKRLYVNSEVYSGDLIWKPQ 178
Query: 203 GSELIAEGTKSAADPAPKTFTTFNQKSLPKFSKDP-APYNLDVIVAKLGPGQEIELEAHA 261
G + +F+ +P N D++VAKL PGQEI+LEAHA
Sbjct: 179 G------------------------RQEERFADNPIRVVNPDIVVAKLRPGQEIDLEAHA 214
Query: 262 VKGIGKTHAKWSPVATAWYRMLPEVVLTKDVKDELAEELVSKCPAKVFDIEDIGRGRKKA 321
+ GIG+ HAK+SPVATA YR+LP + + ++ E A + P V ++E+ G+K+A
Sbjct: 215 ILGIGQDHAKFSPVATASYRLLPTIHILSPIEGEDAVKFQKCFPKGVIELEEGPDGKKQA 274
Query: 322 VVKNARACTLCRECIREADEGEEGGEGEKWTDRVSLRRVKDHFIFTIESTGAIPPDVLFT 381
V + R T+ REC+R + + D+V L RV+DH++F++ESTG + PDVLF
Sbjct: 275 RVADVRKDTVSRECLRHPE----------FADKVQLGRVRDHYLFSVESTGIMKPDVLFI 324
Query: 382 EAVKILEDKCERVITEL 398
+++ +L+ KC V + L
Sbjct: 325 KSIAVLKSKCLAVKSSL 341
>UniRef100_O94616 DNA-directed RNA polymerases I and III 40 kDa polypeptide
[Schizosaccharomyces pombe]
Length = 348
Score = 236 bits (601), Expect = 1e-60
Identities = 143/375 (38%), Positives = 211/375 (56%), Gaps = 54/375 (14%)
Query: 42 LELQKTRVFCNLDAPQHTDTIQYSGAYAALGVDNSVRLDNFYQNFKVEVKRITDEEMEFD 101
++ +T + D ++ + G Y DN LD F +N KV + + E M F+
Sbjct: 4 VDRSRTEISVLSDRVTDVGSVDFPGYY--FDEDNIWDLDKFKKNLKVSITSLDQETMVFE 61
Query: 102 MIGIDPAVANAFRRILIAEVPTMAIERVYIANNTSLVQDEVLSHRLGLIPIDADP---KL 158
+ GID ++ANAFRRILIAE+PT+A E VYI NNTS++QDEVLSHR+GL+PI ADP K
Sbjct: 62 ISGIDASIANAFRRILIAEIPTLAFEFVYIINNTSIIQDEVLSHRIGLVPISADPDMFKW 121
Query: 159 FEYP-DDAGGENNEKNTIVFKQHVRC---------EKGQPRLTVKSDT----LKWLPNGS 204
F++P + + +T+VF + +C EK RL V S+ L W P G
Sbjct: 122 FQHPLPGQEATHTDYDTVVFSLNKKCEFNKNAATDEKDPKRLYVNSEVYSGDLIWKPQG- 180
Query: 205 ELIAEGTKSAADPAPKTFTTFNQKSLPKFSKDP-APYNLDVIVAKLGPGQEIELEAHAVK 263
+ +F+ +P N D++VAKL PGQEI+LEAHA+
Sbjct: 181 -----------------------RQEERFADNPIRVVNPDIVVAKLRPGQEIDLEAHAIL 217
Query: 264 GIGKTHAKWSPVATAWYRMLPEVVLTKDVKDELAEELVSKCPAKVFDIEDIGRGRKKAVV 323
GIG+ HAK+SPVATA YR+LP + + ++ E A + P V ++E+ G+K+A V
Sbjct: 218 GIGQDHAKFSPVATASYRLLPTIHILSPIEGEDAVKFQKCFPKGVIELEEGPDGKKQARV 277
Query: 324 KNARACTLCRECIREADEGEEGGEGEKWTDRVSLRRVKDHFIFTIESTGAIPPDVLFTEA 383
+ R T+ REC+R + + D+V L RV+DH++F++ESTG + PDVLF ++
Sbjct: 278 ADVRKDTVSRECLRHPE----------FADKVQLGRVRDHYLFSVESTGIMKPDVLFIKS 327
Query: 384 VKILEDKCERVITEL 398
+ +L+ KC V + L
Sbjct: 328 IAVLKSKCLAVKSSL 342
>UniRef100_UPI000042E7AF UPI000042E7AF UniRef100 entry
Length = 350
Score = 228 bits (580), Expect = 3e-58
Identities = 133/346 (38%), Positives = 190/346 (54%), Gaps = 50/346 (14%)
Query: 63 QYSGAYAALGVDNSVRLDNFYQNFKVEVKRITDEEMEFDMIGIDPAVANAFRRILIAEVP 122
++ G Y G D+S L F Q+ V+R+T +EFD++G+D ++ANA RR LIAEVP
Sbjct: 26 EFPGHYP--GEDHSWNLQKFKQDLVASVQRLTPSTIEFDLVGVDASIANALRRTLIAEVP 83
Query: 123 TMAIERVYIANNTSLVQDEVLSHRLGLIPIDADPKLFEYPDDAGGENNEKNTIVFKQHVR 182
T+AIE VY+ NNTS++QDEVL HR+GL+P+ DP+ Y A + +E +T+VF VR
Sbjct: 84 TVAIEEVYVWNNTSIMQDEVLCHRVGLVPLKIDPRSLRYRPSANSQPHEHDTVVFDLRVR 143
Query: 183 CEKGQPRL--------------TVKSDTLKWLPNGSELIAEGTKSAADPAPKTFTTFNQK 228
C++ +P + V S LKW+P+G + K F
Sbjct: 144 CDR-RPNVDRSETDPKKMYFDSNVYSGMLKWVPSGDQ-------------KKKFK----- 184
Query: 229 SLPKFSKDPAPYNLDVIVAKLGPGQEIELEAHAVKGIGKTHAKWSPVATAWYRMLPEVVL 288
K P P N D+++ KL PGQ+++L A KG+G HAK+SPVATA YR+LP ++L
Sbjct: 185 -----GKAPRPVNDDILLVKLRPGQQVDLHCFARKGVGADHAKFSPVATASYRLLPHIIL 239
Query: 289 TKDVKDELAEELVSKCPAKVFDIEDIGRGRKKAVVKNARACTLCRECIREADEGEEGGEG 348
+ E ++ P V +IE G + VVKN R T+ RE +R +
Sbjct: 240 RGPIPKEHQQKFKGCFPPGVIEIEKDENGEDQCVVKNPRKDTVSREVLRHPE-------- 291
Query: 349 EKWTDRVSLRRVKDHFIFTIESTGAIPPDVLFTEAVKILEDKCERV 394
+ D V L R++DHFIF +ES G P+ L EA+KIL K + V
Sbjct: 292 --FQDLVQLARIRDHFIFNVESAGQYSPEELVPEAIKILLGKIKDV 335
>UniRef100_O15160-2 Splice isoform 2 of O15160 [Homo sapiens]
Length = 342
Score = 224 bits (570), Expect = 4e-57
Identities = 136/339 (40%), Positives = 191/339 (56%), Gaps = 53/339 (15%)
Query: 42 LELQKTRVFCNLDAPQHTDTIQYSGAYAALGVDNSVRLDNFYQNFKVEVKRITDEEMEFD 101
+E ++RV ++ T + G Y+ G D++ D F +NF+V+V + + +EFD
Sbjct: 7 VEEMRSRVVLGEFGVRNVHTTDFPGNYS--GYDDAWDQDRFEKNFRVDVVHMDENSLEFD 64
Query: 102 MIGIDPAVANAFRRILIAEVPTMAIERVYIANNTSLVQDEVLSHRLGLIPIDADPKLFEY 161
M+GID A+ANAFRRIL+AEVPTMA+E+V + NNTS+VQDE+L+HRLGLIPI ADP+LFEY
Sbjct: 65 MVGIDAAIANAFRRILLAEVPTMAVEKVLVYNNTSIVQDEILAHRLGLIPIHADPRLFEY 124
Query: 162 PDDAGGENNEKNTIVFKQHVRCEKGQPRLTVKSDT-------------LKWLP--NGSEL 206
+ E E +T+ F+ VRC + SD + W+P N ++L
Sbjct: 125 RNQGDEEGTEIDTLQFRLQVRCTRNPHAAKDSSDPNELYVNHKVYTRHMTWIPLGNQADL 184
Query: 207 IAEGTKSAADPAPKTFTTFNQKSLPKFSKDPAPYNLDVIVAKLGPGQEIELEAHAVKGIG 266
EGT P + D+++A+L PGQEI+L H VKGIG
Sbjct: 185 FPEGTIR-------------------------PVHDDILIAQLRPGQEIDLLMHCVKGIG 219
Query: 267 KTHAKWSPVATAWYRMLPEVVLTKDVKDELAEELVSKCPAKVFDIEDIGRGRKKAVVKNA 326
K HAK+SPVATA YR+LP++ L + V+ E AEEL V +++++ +G+K A V N
Sbjct: 220 KDHAKFSPVATASYRLLPDITLLEPVEGEAAEELSRCFSPGVIEVQEV-QGKKVARVANP 278
Query: 327 RACTLCRECIREADEGEEGGEGEKWTDRVSLRRVKDHFI 365
R T RE R EK V L RV+DH+I
Sbjct: 279 RLDTFSREIFR----------NEKLKKVVRLARVRDHYI 307
>UniRef100_UPI0000366163 UPI0000366163 UniRef100 entry
Length = 295
Score = 218 bits (554), Expect = 3e-55
Identities = 131/324 (40%), Positives = 188/324 (57%), Gaps = 38/324 (11%)
Query: 84 QNFKVEVKRITDEEMEFDMIGIDPAVANAFRRILIAEVPTMAIERVYIANNTSLVQDEVL 143
QNF++++ + + +MEFDM+G+D A+ NAFRRIL+AEVPTMAIE+V+I NNTS+VQDEVL
Sbjct: 1 QNFQIDLMHLDESQMEFDMMGVDAAITNAFRRILLAEVPTMAIEKVFIYNNTSIVQDEVL 60
Query: 144 SHRLGLIPIDADPKLFEYPDDAGGENNEKNTIVFKQHVRCEKGQPRLTVKSDTLKWLPNG 203
+HRLGLIPI ADP+LFEY N+ K TI+ C P + +
Sbjct: 61 AHRLGLIPIKADPRLFEY------RNSGKMTII----KMCVCVSPSI---------IQPF 101
Query: 204 SELIAEGTKSAADPAPKTFTTFNQKSLPKFSK-DPAPYNLDVIVAKLGPGQEIELEAHAV 262
S ++ G ++ + + T Q + F+ P + D+++A+L PGQE+++ H +
Sbjct: 102 SHFVSSGDEATENEGSEIDTIQLQLKINVFANCSIGPVHGDILIAQLRPGQELDIIMHCI 161
Query: 263 KGIGKTHAKWSPVATAWYRMLPEVVLTKDVKDELAEELVSKCPAKVFDIEDIGRGRKKAV 322
KGIGK HAK+SPVATA YR+LPE+ L + V+ E AE L V D+ED
Sbjct: 162 KGIGKDHAKFSPVATASYRLLPEITLLEPVEGEKAERLQRCFFRGVIDLED--------- 212
Query: 323 VKNARACTLCRECIREADEGEEGGEG--------EKWTDRVSLRRVKDHFIFTIESTGAI 374
N R L + + +AD E E E+ R + + + T+E+TG +
Sbjct: 213 -SNGRNAFLPKAVVGDADPSTETSETSCRSFVLVEQLLTRTNGTLIDEIQPVTVETTGIL 271
Query: 375 PPDVLFTEAVKILEDKCERVITEL 398
PPDVL TEA+K+L KC+R + EL
Sbjct: 272 PPDVLVTEAIKVLMTKCQRFLNEL 295
>UniRef100_P07703 DNA-directed RNA polymerases I and III 40 kDa polypeptide
[Saccharomyces cerevisiae]
Length = 335
Score = 216 bits (551), Expect = 7e-55
Identities = 128/324 (39%), Positives = 194/324 (59%), Gaps = 30/324 (9%)
Query: 74 DNSVRLDNFYQNFKVEVKRITDEEMEFDMIGIDPAVANAFRRILIAEVPTMAIERVYIAN 133
+N ++ F ++F+V + + E FD+I ID ++ANAFRRI+I+EVP++A E VY N
Sbjct: 28 ENEWNVEKFKKDFEVNISSLDAREANFDLINIDTSIANAFRRIMISEVPSVAAEYVYFFN 87
Query: 134 NTSLVQDEVLSHRLGLIPIDADPKLFEY-----PDDAGGENNEKNTIVFKQHVRCEKGQP 188
NTS++QDEVL+HR+GL+P+ DP + + PDD + ++NTIV +V+C + P
Sbjct: 88 NTSVIQDEVLAHRIGLVPLKVDPDMLTWVDSNLPDDE--KFTDENTIVLSLNVKCTR-NP 144
Query: 189 RLTVKSDTLKWLPNGSELIAEGTKSAADPAPKTFTTFNQKSLPKFSKDPAPYNLDVIVAK 248
S K L N + + A K +P + TTF P DP D+++AK
Sbjct: 145 DAPKGSTDPKELYNNAHVYARDLK--FEPQGRQSTTF--ADCPVVPADP-----DILLAK 195
Query: 249 LGPGQEIELEAHAVKGIGKTHAKWSPVATAWYRMLPEVVLTKDVKDELAEELVSKCPAKV 308
L PGQEI L+AH + GIG HAK+SPV+TA YR+LP++ + + +K E A P V
Sbjct: 196 LRPGQEISLKAHCILGIGGDHAKFSPVSTASYRLLPQINILQPIKGESARRFQKCFPPGV 255
Query: 309 FDIEDIGRGRKKAVVKNARACTLCRECIREADEGEEGGEGEKWTDRVSLRRVKDHFIFTI 368
I++ G +A VK+AR T+ RE +R E++ D+V L RV++HFIF +
Sbjct: 256 IGIDE---GSDEAYVKDARKDTVSREVLRY----------EEFADKVKLGRVRNHFIFNV 302
Query: 369 ESTGAIPPDVLFTEAVKILEDKCE 392
ES GA+ P+ +F ++V+IL++K E
Sbjct: 303 ESAGAMTPEEIFFKSVRILKNKAE 326
>UniRef100_Q6BV37 Similar to sp|P07703 Saccharomyces cerevisiae YPR110c RPC40
[Debaryomyces hansenii]
Length = 335
Score = 214 bits (546), Expect = 3e-54
Identities = 124/317 (39%), Positives = 189/317 (59%), Gaps = 27/317 (8%)
Query: 79 LDNFYQNFKVEVKRITDEEMEFDMIGIDPAVANAFRRILIAEVPTMAIERVYIANNTSLV 138
++ F +F++ + +++ FD+I ID ++ANAFRRI+I+EVP++A E VY+ NNTS++
Sbjct: 34 IEKFKDSFEINISHLSERSSTFDLIHIDTSIANAFRRIMISEVPSVAAETVYVFNNTSVI 93
Query: 139 QDEVLSHRLGLIPIDADPKLFEYPDDAGGENN---EKNTIVFKQHVRCEKGQPRLTVKSD 195
QDEVL+HR+GLIP+ DP + D A E + ++NTIV V C K P S
Sbjct: 94 QDEVLAHRIGLIPLKVDPDTLTWIDTAQDEKDRFTDENTIVMSLDVSCTK-NPHPPQGST 152
Query: 196 TLKWLPNGSELIAEGTKSAADPAPKTFTTFNQKSLPKFSKDPAPYNLDVIVAKLGPGQEI 255
K L S + A+ K +P + F K+ P + DP D+++AKL P QEI
Sbjct: 153 DPKELYRNSHVYAKDLK--FEPQGRQLEIF--KNNPVVAADP-----DILLAKLRPNQEI 203
Query: 256 ELEAHAVKGIGKTHAKWSPVATAWYRMLPEVVLTKDVKDELAEELVSKCPAKVFDIEDIG 315
L AH + G+G HAK+SPVATA YR++P + +T+ +K E + P+ V DI G
Sbjct: 204 SLRAHCILGVGSDHAKFSPVATASYRLMPVITITEPIKGESGKRFQKCFPSGVIDINKNG 263
Query: 316 RGRKKAVVKNARACTLCRECIREADEGEEGGEGEKWTDRVSLRRVKDHFIFTIESTGAIP 375
+AVVK++R T+ RE +R E++ +V L R +DHFIF +ESTGA+
Sbjct: 264 ----EAVVKDSRKDTVSREVLRH----------EEFNGKVKLGRKRDHFIFNVESTGAMS 309
Query: 376 PDVLFTEAVKILEDKCE 392
P+ +F ++V++L++K E
Sbjct: 310 PEEIFFKSVRVLKNKAE 326
>UniRef100_Q6CQS0 Kluyveromyces lactis strain NRRL Y-1140 chromosome D of strain NRRL
Y- 1140 of Kluyveromyces lactis [Kluyveromyces lactis]
Length = 334
Score = 214 bits (544), Expect = 4e-54
Identities = 122/322 (37%), Positives = 195/322 (59%), Gaps = 27/322 (8%)
Query: 74 DNSVRLDNFYQNFKVEVKRITDEEMEFDMIGIDPAVANAFRRILIAEVPTMAIERVYIAN 133
DNS L+ F + F V+++ + + E+ FD++ +D ++ANAFRRI+I+EVP++A E VY N
Sbjct: 28 DNSWDLEKFKETFDVQIESLDEREVNFDLVHVDTSIANAFRRIMISEVPSVAPEYVYFFN 87
Query: 134 NTSLVQDEVLSHRLGLIPIDADPKLFEYPDDAGGEN---NEKNTIVFKQHVRCEKGQPRL 190
NTS++QDEVL+HR+GL+P+ DP + + D + E+ ++NTIV +V+C + P
Sbjct: 88 NTSVLQDEVLAHRIGLVPLKVDPDMLSWVDTSLPESERFTDENTIVMSLNVKCTR-NPDA 146
Query: 191 TVKSDTLKWLPNGSELIAEGTKSAADPAPKTFTTFNQKSLPKFSKDPAPYNLDVIVAKLG 250
+ K L + + A K +P K F P + DP D+++AKL
Sbjct: 147 PKDCEDPKVLYRNAHIYARDLK--FEPQGKQVEQF--AKCPVVAADP-----DILLAKLR 197
Query: 251 PGQEIELEAHAVKGIGKTHAKWSPVATAWYRMLPEVVLTKDVKDELAEELVSKCPAKVFD 310
PGQEI L AH + GIG HAK+SPV+TA YR+LP + +T+ + + A++ V P V D
Sbjct: 198 PGQEISLRAHCILGIGADHAKFSPVSTASYRLLPHIQITQPITGDDAKKFVKCFPKGVAD 257
Query: 311 IEDIGRGRKKAVVKNARACTLCRECIREADEGEEGGEGEKWTDRVSLRRVKDHFIFTIES 370
+ + G +A +K+ R T+ RE +R +++ +V L RV+DHFIF +ES
Sbjct: 258 VNEKG----EAYIKDPRKDTVSREVLRH----------DEFQGKVKLGRVRDHFIFNVES 303
Query: 371 TGAIPPDVLFTEAVKILEDKCE 392
+GA+ P+ +F ++V+IL++K E
Sbjct: 304 SGAMTPEEIFFKSVRILKNKAE 325
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.315 0.134 0.389
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 697,189,893
Number of Sequences: 2790947
Number of extensions: 30605038
Number of successful extensions: 69136
Number of sequences better than 10.0: 155
Number of HSP's better than 10.0 without gapping: 141
Number of HSP's successfully gapped in prelim test: 14
Number of HSP's that attempted gapping in prelim test: 68682
Number of HSP's gapped (non-prelim): 250
length of query: 399
length of database: 848,049,833
effective HSP length: 129
effective length of query: 270
effective length of database: 488,017,670
effective search space: 131764770900
effective search space used: 131764770900
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.5 bits)
S2: 76 (33.9 bits)
Medicago: description of AC146751.11


