

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146750.10 + phase: 0
(221 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
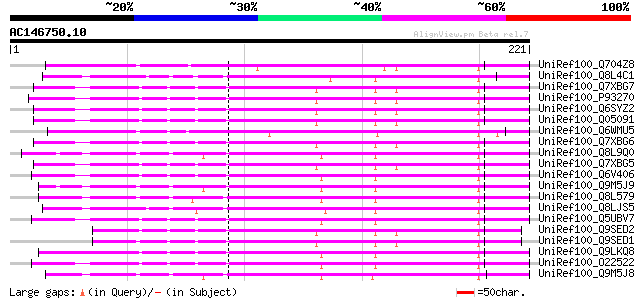
UniRef100_Q704Z8 Putative polygalacturonase-inhibiting protein p... 147 3e-34
UniRef100_Q8L4C1 Polygalacturonase inhibitor protein [Brassica n... 137 3e-31
UniRef100_Q7XBG7 Polygalacturonase-inhibiting protein [Pyrus pyr... 136 4e-31
UniRef100_P93270 Polygalacturonase-inhibiting protein [Malus dom... 136 4e-31
UniRef100_Q6SYZ2 Polygalacturonase-inhibiting protein [Eucalyptu... 134 2e-30
UniRef100_Q05091 Polygalacturonase inhibitor precursor [Pyrus co... 134 2e-30
UniRef100_Q6WMU5 Polygalacturonase-inhibiting protein precursor ... 133 3e-30
UniRef100_Q7XBG6 Polygalacturonase-inhibiting protein [Pyrus hyb... 133 4e-30
UniRef100_Q8L9Q0 Polygalacturonase inhibiting protein 1 [Arabido... 132 8e-30
UniRef100_Q7XBG5 Polygalacturonase-inhibiting protein [Pyrus com... 131 1e-29
UniRef100_Q6V406 Polygalacturonase inhibiting protein [Prunus pe... 131 1e-29
UniRef100_Q9M5J9 Polygalacturonase inhibitor 1 precursor [Arabid... 130 2e-29
UniRef100_Q8L579 Polygalacturonase inhibitor protein [Brassica n... 129 4e-29
UniRef100_Q8LJS5 Polygalacturonase inhibitory protein [Brassica ... 128 9e-29
UniRef100_Q5UBV7 Polygalacturonase inhibiting protein [Prunus mume] 127 2e-28
UniRef100_Q9SED2 Polygalacturonase-inhibiting protein [Eucalyptu... 127 3e-28
UniRef100_Q9SED1 Polygalacturonase-inhibiting protein [Eucalyptu... 127 3e-28
UniRef100_Q9LKQ8 Polygalacturonase inhibiting protein [Prunus ma... 126 4e-28
UniRef100_O22522 Polygalacturonase inhibiting protein [Prunus ar... 126 4e-28
UniRef100_Q9M5J8 Polygalacturonase inhibitor 2 precursor [Arabid... 126 5e-28
>UniRef100_Q704Z8 Putative polygalacturonase-inhibiting protein precursor [Rubus
idaeus]
Length = 331
Score = 147 bits (370), Expect = 3e-34
Identities = 81/187 (43%), Positives = 107/187 (56%), Gaps = 2/187 (1%)
Query: 16 LFLSSIYFSTSNADDVEFSCNANDKATLLKIRDHFGGPNGRLSDWDNGTDCCSDWSFVGC 75
LF ++ FST + CN DK L +I+ F P LS W + DCC+DW V C
Sbjct: 6 LFSLTLLFSTILTPALSELCNPKDKKVLFEIKTAFNNPY-ILSSWKSDADCCTDWYCVEC 64
Query: 76 GKPLPGRITVVTISRGWGLSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSKLQRLQK 135
P RI +TI L+G +PA+ GDLPYL L L ++P +TGPI S +KL+ L+
Sbjct: 65 D-PTTHRINSLTIFTDNNLTGQIPAQVGDLPYLETLELRKLPHLTGPIQPSIAKLKHLKM 123
Query: 136 LDLGSNSLSGSIPTFLGQLKGLQEFDLSNNKLTGVIPASFGSLPSLSQFNVSFNQLCGAI 195
L L N LSGS+P F+ QLK L +L+ NK TG IP+S LP+L ++ NQL G I
Sbjct: 124 LRLSWNGLSGSVPDFISQLKNLTFLELNFNKFTGSIPSSLSQLPNLGALHLDRNQLTGQI 183
Query: 196 PTGLSKF 202
P+ KF
Sbjct: 184 PSSFGKF 190
Score = 85.1 bits (209), Expect = 1e-15
Identities = 60/157 (38%), Positives = 78/157 (49%), Gaps = 34/157 (21%)
Query: 94 LSGTLPAEFGD----LPYLNFLSLAEMPKVTGPIPNSFSKLQRLQKLDLGSNSLSGSIPT 149
L+G +P+ FG +P L FLS ++ TG IP SF+ + ++DL N L G
Sbjct: 179 LTGQIPSSFGKFVGTVPAL-FLSHNQL---TGKIPTSFANMN-FDQIDLSRNKLEGDASV 233
Query: 150 FLGQLKGLQ---------EFDLS--------------NNKLTGVIPASFGSLPSLSQFNV 186
G K Q EFDLS +N +TG IPA L L FNV
Sbjct: 234 IFGLNKTTQIVDLSRNMLEFDLSKVVFSTSLRAVDLNHNSITGSIPAQLTQLDDLVLFNV 293
Query: 187 SFNQLCGAIPTG--LSKFAKSSFDHNKCLCGAPLAAC 221
S+N+LCG IP G L +S+ HN+CLCGAPL +C
Sbjct: 294 SYNRLCGKIPVGGKLQSLDTTSYFHNRCLCGAPLPSC 330
>UniRef100_Q8L4C1 Polygalacturonase inhibitor protein [Brassica napus]
Length = 342
Score = 137 bits (344), Expect = 3e-31
Identities = 81/193 (41%), Positives = 115/193 (58%), Gaps = 8/193 (4%)
Query: 15 LLFLSSIYFSTSNADDVEFSCNANDKATLLKIRDHFGGPNGRLSDWDNGTDCCSDWSFVG 74
LL L ++ F+TS + D+ C+ +DK TLLKI+ P +S WD DCC+ W V
Sbjct: 8 LLLLFALLFTTSLSKDL---CHKDDKNTLLKIKKAMNNPYTIIS-WDPKDDCCT-WVSVE 62
Query: 75 CGKPLPGRITVVTISRGWGLSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSKLQRLQ 134
CG R+T + IS +S +P E GDLPYL +L+L ++P +TG IP + +KL+ L+
Sbjct: 63 CGNA--NRVTSLDISDD-DVSAQIPPEVGDLPYLQYLTLRKLPNLTGEIPPTIAKLKYLK 119
Query: 135 KLDLGSNSLSGSIPTFLGQLKGLQEFDLSNNKLTGVIPASFGSLPSLSQFNVSFNQLCGA 194
L L NSL+G +P FL QLK L+ +LS NKL+G IP S LP L +S N+L G
Sbjct: 120 SLWLSWNSLTGPVPEFLSQLKNLEYINLSFNKLSGSIPGSLSLLPKLDFLELSRNKLTGP 179
Query: 195 IPTGLSKFAKSSF 207
IP F ++++
Sbjct: 180 IPESFGSFKRAAY 192
Score = 87.0 bits (214), Expect = 3e-16
Identities = 63/177 (35%), Positives = 81/177 (45%), Gaps = 51/177 (28%)
Query: 94 LSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSKLQRLQK-LDLGSNSLSGSIPTFLG 152
LSG++P LP L+FL L+ K+TGPIP SF +R + L N LSGSIP LG
Sbjct: 152 LSGSIPGSLSLLPKLDFLELSRN-KLTGPIPESFGSFKRAAYGIYLSHNQLSGSIPKSLG 210
Query: 153 QL----------------------------------------------KGLQEFDLSNNK 166
+ K + DL++N
Sbjct: 211 NVDFNTIDLSRNKLEGDASMLFGAKKTTWHIDLSRNMFQFDISKVKVAKTVNFLDLNHNS 270
Query: 167 LTGVIPASFGSLPSLSQFNVSFNQLCGAIPTG--LSKFAKSSFDHNKCLCGAPLAAC 221
LTG IP + L L FNVS+N+LCG IP G L +F ++ HNKCLC APL +C
Sbjct: 271 LTGSIPDQWTQL-DLQTFNVSYNRLCGRIPQGGDLQRFDVYAYLHNKCLCDAPLPSC 326
>UniRef100_Q7XBG7 Polygalacturonase-inhibiting protein [Pyrus pyrifolia]
Length = 330
Score = 136 bits (342), Expect = 4e-31
Identities = 85/192 (44%), Positives = 110/192 (57%), Gaps = 10/192 (5%)
Query: 11 LHLTLLFLSSIYFSTSNADDVEFSCNANDKATLLKIRDHFGGPNGRLSDWDNGTDCCSDW 70
L LTLLF S + + S+ CN +DK LL+I+ FG P L+ W + TDCC DW
Sbjct: 9 LSLTLLFSSVLNPALSDL------CNPDDKKVLLQIKKAFGDPYV-LASWKSDTDCC-DW 60
Query: 71 SFVGCGKPLPGRITVVTISRGWGLSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSKL 130
V C RI +TI G +SG +PA GDLPYL L + P +TGPI + +KL
Sbjct: 61 YCVTCDSTT-NRINSLTIFAGQ-VSGQIPALVGDLPYLETLEFHKQPNLTGPIQPAIAKL 118
Query: 131 QRLQKLDLGSNSLSGSIPTFLGQLKGLQEFDLSNNKLTGVIPASFGSLPSLSQFNVSFNQ 190
+ L+ L L +LSGS+P FL QLK L DLS N LTG IP+S LP+LS ++ N+
Sbjct: 119 KGLKSLRLSWTNLSGSVPDFLSQLKNLTFLDLSFNNLTGAIPSSLSELPNLSALHLDRNK 178
Query: 191 LCGAIPTGLSKF 202
L G IP +F
Sbjct: 179 LTGHIPKSFGQF 190
Score = 82.0 bits (201), Expect = 1e-14
Identities = 60/177 (33%), Positives = 83/177 (45%), Gaps = 51/177 (28%)
Query: 94 LSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSK-LQRLQKLDLGSNSLSGSIPTFLG 152
L+G +P+ +LP L+ L L + K+TG IP SF + + + L L N LSG+IPT
Sbjct: 155 LTGAIPSSLSELPNLSALHL-DRNKLTGHIPKSFGQFIGNVPDLYLSHNQLSGNIPTSFA 213
Query: 153 QL--------------------------------KGLQEFDLS--------------NNK 166
Q+ + L EF+LS +NK
Sbjct: 214 QMDFTSIDLSRNKLEGDASVIFGLNKTTQIVDLSRNLLEFNLSKVEFPTSLTSLDINHNK 273
Query: 167 LTGVIPASFGSLPSLSQFNVSFNQLCGAIPTG--LSKFAKSSFDHNKCLCGAPLAAC 221
+ G IP F L + NVS+N+LCG IP G L F + S+ HN+CLCGAPL +C
Sbjct: 274 IYGSIPVEFTQL-NFQFLNVSYNRLCGQIPVGGKLQSFDEYSYFHNRCLCGAPLPSC 329
>UniRef100_P93270 Polygalacturonase-inhibiting protein [Malus domestica]
Length = 330
Score = 136 bits (342), Expect = 4e-31
Identities = 85/194 (43%), Positives = 112/194 (56%), Gaps = 10/194 (5%)
Query: 9 LLLHLTLLFLSSIYFSTSNADDVEFSCNANDKATLLKIRDHFGGPNGRLSDWDNGTDCCS 68
+ L LTLLF S + + S+ CN +DK LL+I+ FG P L+ W + TDCC
Sbjct: 7 IFLSLTLLFSSVLKPALSDL------CNPDDKKVLLQIKKAFGDPYV-LTSWKSDTDCC- 58
Query: 69 DWSFVGCGKPLPGRITVVTISRGWGLSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFS 128
DW V C RI +TI G +SG +PA GDLPYL L + P +TGPI + +
Sbjct: 59 DWYCVTCDSTT-NRINSLTIFAGQ-VSGQIPALVGDLPYLETLEFHKQPNLTGPIQPAIA 116
Query: 129 KLQRLQKLDLGSNSLSGSIPTFLGQLKGLQEFDLSNNKLTGVIPASFGSLPSLSQFNVSF 188
KL+ L+ L L +LSGS+P FL QLK L DLS N LTG IP+S LP+L+ ++
Sbjct: 117 KLKGLKFLRLSWTNLSGSVPDFLSQLKNLTFLDLSFNNLTGAIPSSLSQLPNLNALHLDR 176
Query: 189 NQLCGAIPTGLSKF 202
N+L G IP L +F
Sbjct: 177 NKLTGHIPKSLGQF 190
Score = 80.9 bits (198), Expect = 2e-14
Identities = 60/177 (33%), Positives = 81/177 (44%), Gaps = 51/177 (28%)
Query: 94 LSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSK-LQRLQKLDLGSNSLSGSIPTFLG 152
L+G +P+ LP LN L L + K+TG IP S + + + L L N LSG+IPT
Sbjct: 155 LTGAIPSSLSQLPNLNALHL-DRNKLTGHIPKSLGQFIGNVPDLYLSHNQLSGNIPTSFA 213
Query: 153 QL--------------------------------KGLQEFDLS--------------NNK 166
Q+ + L EF+LS +NK
Sbjct: 214 QMDFTSIDLSRNKLEGDASVIFGLNKTTQIVDLSRNLLEFNLSKVEFPTSLTSLDINHNK 273
Query: 167 LTGVIPASFGSLPSLSQFNVSFNQLCGAIPTG--LSKFAKSSFDHNKCLCGAPLAAC 221
+ G IP F L + NVS+N+LCG IP G L F + S+ HN+CLCGAPL +C
Sbjct: 274 IYGSIPVEFTQL-NFQFLNVSYNRLCGQIPVGGKLQSFDEYSYFHNRCLCGAPLPSC 329
>UniRef100_Q6SYZ2 Polygalacturonase-inhibiting protein [Eucalyptus grandis]
Length = 331
Score = 134 bits (337), Expect = 2e-30
Identities = 84/192 (43%), Positives = 108/192 (55%), Gaps = 10/192 (5%)
Query: 11 LHLTLLFLSSIYFSTSNADDVEFSCNANDKATLLKIRDHFGGPNGRLSDWDNGTDCCSDW 70
L LTLLF S + + S+ CN +DK LL+I+ FG P L+ W + TDCC DW
Sbjct: 9 LSLTLLFSSVLNPALSDL------CNPDDKKVLLQIKKAFGDPYV-LASWKSDTDCC-DW 60
Query: 71 SFVGCGKPLPGRITVVTISRGWGLSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSKL 130
V C RI +TI G +SG +PA GDLPYL L + P +TGPI + +KL
Sbjct: 61 YCVTCDSTT-NRINSLTIFAGQ-VSGQIPALVGDLPYLETLEFHKQPNLTGPIQPAIAKL 118
Query: 131 QRLQKLDLGSNSLSGSIPTFLGQLKGLQEFDLSNNKLTGVIPASFGSLPSLSQFNVSFNQ 190
+ L+ L L +LSGS+P FL QLK L DLS N LTG IP+S LP+L + N+
Sbjct: 119 KGLKSLRLSWTNLSGSVPDFLSQLKNLTFLDLSFNNLTGAIPSSLSELPNLGALRLDRNK 178
Query: 191 LCGAIPTGLSKF 202
L G IP +F
Sbjct: 179 LTGHIPISFGQF 190
Score = 80.1 bits (196), Expect = 4e-14
Identities = 60/177 (33%), Positives = 81/177 (44%), Gaps = 51/177 (28%)
Query: 94 LSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSK-LQRLQKLDLGSNSLSGSIPTFLG 152
L+G +P+ +LP L L L + K+TG IP SF + + + L L N LSG+IPT
Sbjct: 155 LTGAIPSSLSELPNLGALRL-DRNKLTGHIPISFGQFIGNVPDLYLSHNQLSGNIPTSFA 213
Query: 153 QL--------------------------------KGLQEFDLS--------------NNK 166
Q+ + L EF+LS +NK
Sbjct: 214 QMDFTSIDLSRNKLEGDASVIFGLNKTTQIVDLSRNLLEFNLSKVEFPTSLTSLDINHNK 273
Query: 167 LTGVIPASFGSLPSLSQFNVSFNQLCGAIPTG--LSKFAKSSFDHNKCLCGAPLAAC 221
+ G IP F L + NVS+N+LCG IP G L F + S+ HN+CLCGAPL C
Sbjct: 274 IYGSIPVEFTQL-NFQFLNVSYNRLCGQIPVGGKLQSFDEYSYFHNRCLCGAPLGPC 329
>UniRef100_Q05091 Polygalacturonase inhibitor precursor [Pyrus communis]
Length = 330
Score = 134 bits (337), Expect = 2e-30
Identities = 84/192 (43%), Positives = 108/192 (55%), Gaps = 10/192 (5%)
Query: 11 LHLTLLFLSSIYFSTSNADDVEFSCNANDKATLLKIRDHFGGPNGRLSDWDNGTDCCSDW 70
L LTLLF S + + S+ CN +DK LL+I+ FG P L+ W + TDCC DW
Sbjct: 9 LSLTLLFSSVLNPALSDL------CNPDDKKVLLQIKKAFGDPYV-LASWKSDTDCC-DW 60
Query: 71 SFVGCGKPLPGRITVVTISRGWGLSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSKL 130
V C RI +TI G +SG +PA GDLPYL L + P +TGPI + +KL
Sbjct: 61 YCVTCDSTT-NRINSLTIFAGQ-VSGQIPALVGDLPYLETLEFHKQPNLTGPIQPAIAKL 118
Query: 131 QRLQKLDLGSNSLSGSIPTFLGQLKGLQEFDLSNNKLTGVIPASFGSLPSLSQFNVSFNQ 190
+ L+ L L +LSGS+P FL QLK L DLS N LTG IP+S LP+L + N+
Sbjct: 119 KGLKSLRLSWTNLSGSVPDFLSQLKNLTFLDLSFNNLTGAIPSSLSELPNLGALRLDRNK 178
Query: 191 LCGAIPTGLSKF 202
L G IP +F
Sbjct: 179 LTGHIPISFGQF 190
Score = 80.5 bits (197), Expect = 3e-14
Identities = 60/177 (33%), Positives = 82/177 (45%), Gaps = 51/177 (28%)
Query: 94 LSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSK-LQRLQKLDLGSNSLSGSIPTFLG 152
L+G +P+ +LP L L L + K+TG IP SF + + + L L N LSG+IPT
Sbjct: 155 LTGAIPSSLSELPNLGALRL-DRNKLTGHIPISFGQFIGNVPDLYLSHNQLSGNIPTSFA 213
Query: 153 QL--------------------------------KGLQEFDLS--------------NNK 166
Q+ + L EF+LS +NK
Sbjct: 214 QMDFTSIDLSRNKLEGDASVIFGLNKTTQIVDLSRNLLEFNLSKVEFPTSLTSLDINHNK 273
Query: 167 LTGVIPASFGSLPSLSQFNVSFNQLCGAIPTG--LSKFAKSSFDHNKCLCGAPLAAC 221
+ G IP F L + NVS+N+LCG IP G L F + S+ HN+CLCGAPL +C
Sbjct: 274 IYGSIPVEFTQL-NFQFLNVSYNRLCGQIPVGGKLQSFDEYSYFHNRCLCGAPLPSC 329
>UniRef100_Q6WMU5 Polygalacturonase-inhibiting protein precursor [Gossypium
barbadense]
Length = 330
Score = 133 bits (335), Expect = 3e-30
Identities = 78/200 (39%), Positives = 107/200 (53%), Gaps = 8/200 (4%)
Query: 17 FLSSIYFSTSNADDVEFSCNANDKATLLKIRDHFGGPNGRLSDWDNGTDCCSDWSFVGCG 76
FLS ++ + + V CNA DK LLKI+ G P L+ WD TDCC DW + C
Sbjct: 7 FLSFLFITIFISPSVSDHCNAQDKKVLLKIKKALGNPY-LLASWDPKTDCC-DWYCLEC- 63
Query: 77 KPLPGRITVVTISRGWGLSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSKLQRLQKL 136
P R+ +T+ L+G +P E GDLPYL L +P + G I + +KL+ L+ L
Sbjct: 64 HPNTHRVVSLTLFSDDRLTGQIPPEVGDLPYLETLLFRHLPNLNGTIQPAIAKLKNLKML 123
Query: 137 DLGSNSLSGSIPTFLGQLKGLQEFDLSNNKLTGVIPASFGSLPSLSQFNVSFNQLCGAIP 196
L +LSG +P FL QLK L DLS N L+G IP+S +LP+L ++ N+L G IP
Sbjct: 124 RLSWTNLSGPVPNFLSQLKNLTYLDLSFNNLSGSIPSSLSTLPNLEDLHLDRNKLTGTIP 183
Query: 197 TGLSKFAKSS-----FDHNK 211
F + + HNK
Sbjct: 184 ESFGMFPRKNLYLFILSHNK 203
Score = 84.3 bits (207), Expect = 2e-15
Identities = 56/154 (36%), Positives = 73/154 (47%), Gaps = 28/154 (18%)
Query: 94 LSGTLPAEFGDLPYLN-FLSLAEMPKVTGPIPNSFSKLQRLQKLDLGSNSLSGSIPTFLG 152
L+GT+P FG P N +L + K++G IP S + + +DL N L G G
Sbjct: 178 LTGTIPESFGMFPRKNLYLFILSHNKLSGTIPASLANMD-FNTIDLSRNLLEGDPSVLFG 236
Query: 153 QLK-----------------------GLQEFDLSNNKLTGVIPASFGSLPSLSQFNVSFN 189
K L DL++NK+TG IPA L L NVS+N
Sbjct: 237 PNKTTFEIDLSRNMFQFGLSKVQFPKSLARLDLNHNKITGSIPAGLTDL-ELQFMNVSYN 295
Query: 190 QLCGAIPTG--LSKFAKSSFDHNKCLCGAPLAAC 221
+LCG IP G L F S++ HN+CLCGAPL C
Sbjct: 296 RLCGQIPVGGRLQSFDYSTYFHNRCLCGAPLDTC 329
>UniRef100_Q7XBG6 Polygalacturonase-inhibiting protein [Pyrus hybrid cultivar]
Length = 330
Score = 133 bits (334), Expect = 4e-30
Identities = 84/192 (43%), Positives = 108/192 (55%), Gaps = 10/192 (5%)
Query: 11 LHLTLLFLSSIYFSTSNADDVEFSCNANDKATLLKIRDHFGGPNGRLSDWDNGTDCCSDW 70
L LTLLF S + + S+ CN +DK LL+I+ FG P L+ W + TDCC DW
Sbjct: 9 LSLTLLFSSVLNPALSDL------CNPDDKKVLLQIKKAFGDPYV-LASWKSDTDCC-DW 60
Query: 71 SFVGCGKPLPGRITVVTISRGWGLSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSKL 130
V C RI +TI G +SG +PA GDLPYL L + P +TGPI + +KL
Sbjct: 61 YCVTCDSTT-NRINSLTIFAGQ-VSGQIPALVGDLPYLETLEFHKQPNLTGPIQPAIAKL 118
Query: 131 QRLQKLDLGSNSLSGSIPTFLGQLKGLQEFDLSNNKLTGVIPASFGSLPSLSQFNVSFNQ 190
+ L+ L L +LSGS+P FL QLK L DLS N LTG IP+S LP+L + N+
Sbjct: 119 KGLKFLRLSWTNLSGSVPDFLSQLKNLTFLDLSFNNLTGAIPSSLSELPNLDALRLDRNK 178
Query: 191 LCGAIPTGLSKF 202
L G IP +F
Sbjct: 179 LTGHIPISFGQF 190
Score = 80.9 bits (198), Expect = 2e-14
Identities = 60/177 (33%), Positives = 83/177 (45%), Gaps = 51/177 (28%)
Query: 94 LSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSK-LQRLQKLDLGSNSLSGSIPTFLG 152
L+G +P+ +LP L+ L L + K+TG IP SF + + + L L N LSG+IPT
Sbjct: 155 LTGAIPSSLSELPNLDALRL-DRNKLTGHIPISFGQFIGNVPDLCLSHNQLSGNIPTSFA 213
Query: 153 QL--------------------------------KGLQEFDLS--------------NNK 166
Q+ + L EF+LS +NK
Sbjct: 214 QMDFGSIDLSRNKLEGDASVIFVLNKTTQIVDLSRNLLEFNLSKVEFPTSLTSLDINHNK 273
Query: 167 LTGVIPASFGSLPSLSQFNVSFNQLCGAIPTG--LSKFAKSSFDHNKCLCGAPLAAC 221
+ G IP F L + NVS+N+LCG IP G L F + S+ HN+CLCGAPL +C
Sbjct: 274 IYGSIPVEFTQL-NFQFLNVSYNRLCGQIPVGGKLQSFDEYSYFHNRCLCGAPLPSC 329
>UniRef100_Q8L9Q0 Polygalacturonase inhibiting protein 1 [Arabidopsis thaliana]
Length = 332
Score = 132 bits (331), Expect = 8e-30
Identities = 82/198 (41%), Positives = 114/198 (57%), Gaps = 8/198 (4%)
Query: 6 NKFLLLHLTLLFLSSIYFSTSNADDVEFSCNANDKATLLKIRDHFGGPNGRLSDWDNGTD 65
+K L TLLFL + + +T + D+ CN NDK TLLKI+ P L+ WD TD
Sbjct: 2 DKTATLLSTLLFLFT-FLTTCLSKDL---CNQNDKNTLLKIKKSLNNPY-HLASWDPQTD 56
Query: 66 CCSDWSFVGCGKPLPG-RITVVTISRGWGLSGTLPAEFGDLPYLNFLSLAEMPKVTGPIP 124
CCS W + CG R+T +TI G +SG +PAE GDLPYL L ++ +TG I
Sbjct: 57 CCS-WYCLECGDATVNHRVTALTIFSGQ-ISGQIPAEVGDLPYLETLVFRKLSNLTGTIQ 114
Query: 125 NSFSKLQRLQKLDLGSNSLSGSIPTFLGQLKGLQEFDLSNNKLTGVIPASFGSLPSLSQF 184
+ +KL+ L+ L L +L+G IP F+ QLK L+ +LS N L+G IP+S +LP +
Sbjct: 115 PTIAKLKNLRMLRLSWTNLTGPIPDFISQLKNLEFLELSFNDLSGSIPSSLSTLPKILAL 174
Query: 185 NVSFNQLCGAIPTGLSKF 202
+S N+L G+IP F
Sbjct: 175 ELSRNKLTGSIPESFGSF 192
Score = 82.0 bits (201), Expect = 1e-14
Identities = 62/177 (35%), Positives = 77/177 (43%), Gaps = 51/177 (28%)
Query: 94 LSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSKLQ-RLQKLDLGSNSLSGSIPTFLG 152
LSG++P+ LP + L L+ K+TG IP SF + L L N LSG IP LG
Sbjct: 157 LSGSIPSSLSTLPKILALELSRN-KLTGSIPESFGSFPGTVPDLRLSHNQLSGPIPKSLG 215
Query: 153 QL----------------------------------------------KGLQEFDLSNNK 166
+ K L DL++N
Sbjct: 216 NIDFNRIDLSRNKLQGDASMLFGSNKTTWSIDLSRNMFQFDISKVDIPKTLGILDLNHNG 275
Query: 167 LTGVIPASFGSLPSLSQFNVSFNQLCGAIPTG--LSKFAKSSFDHNKCLCGAPLAAC 221
+TG IP + P L FNVS+N+LCG IPTG L F S+ HNKCLCGAPL C
Sbjct: 276 ITGNIPVQWTEAP-LQFFNVSYNKLCGHIPTGGKLQTFDSYSYFHNKCLCGAPLEIC 331
>UniRef100_Q7XBG5 Polygalacturonase-inhibiting protein [Pyrus communis]
Length = 330
Score = 131 bits (330), Expect = 1e-29
Identities = 83/192 (43%), Positives = 108/192 (56%), Gaps = 10/192 (5%)
Query: 11 LHLTLLFLSSIYFSTSNADDVEFSCNANDKATLLKIRDHFGGPNGRLSDWDNGTDCCSDW 70
L LTLLF S + + S+ CN +DK LL+I+ FG P L+ W + TDCC DW
Sbjct: 9 LSLTLLFSSVLNPALSDL------CNPDDKKVLLQIKKAFGDPYV-LASWKSDTDCC-DW 60
Query: 71 SFVGCGKPLPGRITVVTISRGWGLSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSKL 130
V C RI +TI G +SG +PA GDLPYL L + P +TGPI + + L
Sbjct: 61 YCVTCDSTT-NRINSLTIFAGQ-VSGQIPALVGDLPYLETLEFHKQPNLTGPIQPAIANL 118
Query: 131 QRLQKLDLGSNSLSGSIPTFLGQLKGLQEFDLSNNKLTGVIPASFGSLPSLSQFNVSFNQ 190
+ L+ L L +LSGS+P FL QLK L DLS N LTG IP+S LP+L ++ N+
Sbjct: 119 KGLKFLRLSWTNLSGSVPDFLSQLKNLTFLDLSFNNLTGAIPSSLSELPNLGALHLDRNK 178
Query: 191 LCGAIPTGLSKF 202
L G IP +F
Sbjct: 179 LTGHIPISFGQF 190
Score = 80.5 bits (197), Expect = 3e-14
Identities = 60/177 (33%), Positives = 82/177 (45%), Gaps = 51/177 (28%)
Query: 94 LSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSK-LQRLQKLDLGSNSLSGSIPTFLG 152
L+G +P+ +LP L L L + K+TG IP SF + + + L L N LSG+IPT
Sbjct: 155 LTGAIPSSLSELPNLGALHL-DRNKLTGHIPISFGQFIGNVPDLYLSHNQLSGNIPTSFA 213
Query: 153 QL--------------------------------KGLQEFDLS--------------NNK 166
Q+ + L EF+LS +NK
Sbjct: 214 QMDFGSIDLSRNKLEGDASVIFGLNKTTQIVDLSRNLLEFNLSKVEFPTSLTSLDINHNK 273
Query: 167 LTGVIPASFGSLPSLSQFNVSFNQLCGAIPTG--LSKFAKSSFDHNKCLCGAPLAAC 221
+ G IP F L + NVS+N+LCG IP G L F + S+ HN+CLCGAPL +C
Sbjct: 274 IYGSIPVEFTQL-NFQFLNVSYNRLCGQIPVGGKLQSFDEYSYFHNRCLCGAPLPSC 329
>UniRef100_Q6V406 Polygalacturonase inhibiting protein [Prunus persica]
Length = 330
Score = 131 bits (330), Expect = 1e-29
Identities = 82/193 (42%), Positives = 109/193 (55%), Gaps = 10/193 (5%)
Query: 10 LLHLTLLFLSSIYFSTSNADDVEFSCNANDKATLLKIRDHFGGPNGRLSDWDNGTDCCSD 69
LL LTLLF + + + S CN DK LL+I+ F P L+ W TDCC D
Sbjct: 8 LLCLTLLFSTILNQALSEL------CNPEDKKVLLQIKKAFNDPYV-LTSWKPETDCC-D 59
Query: 70 WSFVGCGKPLPGRITVVTISRGWGLSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSK 129
W V C RI +TI G +SG +P + GDLPYL L + P +TGPI S +K
Sbjct: 60 WYCVTCDSTT-NRINSLTIFSGQ-VSGQIPTQVGDLPYLETLEFHKQPNLTGPIQPSIAK 117
Query: 130 LQRLQKLDLGSNSLSGSIPTFLGQLKGLQEFDLSNNKLTGVIPASFGSLPSLSQFNVSFN 189
L+RL++L L ++SGS+P FL QLK L DLS + LTG IP+S LP+L+ ++ N
Sbjct: 118 LKRLKELRLSWTNISGSVPDFLSQLKNLTFLDLSFSNLTGSIPSSLSQLPNLNALHLDRN 177
Query: 190 QLCGAIPTGLSKF 202
+L G IP +F
Sbjct: 178 KLTGHIPKSFGEF 190
Score = 87.0 bits (214), Expect = 3e-16
Identities = 61/177 (34%), Positives = 82/177 (45%), Gaps = 51/177 (28%)
Query: 94 LSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSKLQ-RLQKLDLGSNSLSGSIPTFLG 152
L+G++P+ LP LN L L + K+TG IP SF + + +L L N LSG+IPT L
Sbjct: 155 LTGSIPSSLSQLPNLNALHL-DRNKLTGHIPKSFGEFHGSVPELYLSHNQLSGNIPTSLA 213
Query: 153 QL----------------------------------------------KGLQEFDLSNNK 166
+L K L DL++NK
Sbjct: 214 KLDFNRIDFSRNKLEGDASMIFGLNKTTQIVDLSRNLLEFNLSKVEFSKSLISLDLNHNK 273
Query: 167 LTGVIPASFGSLPSLSQFNVSFNQLCGAIPTG--LSKFAKSSFDHNKCLCGAPLAAC 221
+TG IP + L NVS+N+LCG IP G L F S++ HN+CLCGAPL +C
Sbjct: 274 ITGGIPVGLTQV-DLQFLNVSYNRLCGQIPVGGKLQSFDSSTYFHNRCLCGAPLPSC 329
>UniRef100_Q9M5J9 Polygalacturonase inhibitor 1 precursor [Arabidopsis thaliana]
Length = 330
Score = 130 bits (328), Expect = 2e-29
Identities = 80/191 (41%), Positives = 111/191 (57%), Gaps = 8/191 (4%)
Query: 13 LTLLFLSSIYFSTSNADDVEFSCNANDKATLLKIRDHFGGPNGRLSDWDNGTDCCSDWSF 72
L LLFL + + +T + D+ CN NDK TLLKI+ P L+ WD TDCCS W
Sbjct: 7 LCLLFLFT-FLTTCLSKDL---CNQNDKNTLLKIKKSLNNPY-HLASWDPQTDCCS-WYC 60
Query: 73 VGCGKPLPG-RITVVTISRGWGLSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSKLQ 131
+ CG R+T +TI G +SG +PAE GDLPYL L ++ +TG I + +KL+
Sbjct: 61 LECGDATVNHRVTALTIFSGQ-ISGQIPAEVGDLPYLETLVFRKLSNLTGTIQPTIAKLK 119
Query: 132 RLQKLDLGSNSLSGSIPTFLGQLKGLQEFDLSNNKLTGVIPASFGSLPSLSQFNVSFNQL 191
L+ L L +L+G IP F+ QLK L+ +LS N L+G IP+S +LP + +S N+L
Sbjct: 120 NLRMLRLSWTNLTGPIPDFISQLKNLEFLELSFNDLSGSIPSSLSTLPKILALELSRNKL 179
Query: 192 CGAIPTGLSKF 202
G+IP F
Sbjct: 180 TGSIPESFGSF 190
Score = 82.0 bits (201), Expect = 1e-14
Identities = 62/177 (35%), Positives = 77/177 (43%), Gaps = 51/177 (28%)
Query: 94 LSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSKLQ-RLQKLDLGSNSLSGSIPTFLG 152
LSG++P+ LP + L L+ K+TG IP SF + L L N LSG IP LG
Sbjct: 155 LSGSIPSSLSTLPKILALELSRN-KLTGSIPESFGSFPGTVPDLRLSHNQLSGPIPKSLG 213
Query: 153 QL----------------------------------------------KGLQEFDLSNNK 166
+ K L DL++N
Sbjct: 214 NIDFNRIDLSRNKLQGDASMLFGSNKTTWSIDLSRNMFQFDISKVDIPKTLGILDLNHNG 273
Query: 167 LTGVIPASFGSLPSLSQFNVSFNQLCGAIPTG--LSKFAKSSFDHNKCLCGAPLAAC 221
+TG IP + P L FNVS+N+LCG IPTG L F S+ HNKCLCGAPL C
Sbjct: 274 ITGNIPVQWTEAP-LQFFNVSYNKLCGHIPTGGKLQTFDSYSYFHNKCLCGAPLEIC 329
>UniRef100_Q8L579 Polygalacturonase inhibitor protein [Brassica napus]
Length = 331
Score = 129 bits (325), Expect = 4e-29
Identities = 78/191 (40%), Positives = 110/191 (56%), Gaps = 7/191 (3%)
Query: 13 LTLLFLSSIYFSTSNADDVEFSCNANDKATLLKIRDHFGGPNGRLSDWDNGTDCCSDWSF 72
L L F + +TS + ++ CN +DK TLLKI+ P L+ WD TDCCS W
Sbjct: 7 LLLFFFLFTFLTTSFSKNL---CNQDDKTTLLKIKKALNNPY-HLASWDPRTDCCS-WYC 61
Query: 73 VGCG-KPLPGRITVVTISRGWGLSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSKLQ 131
+ CG + R+T +TI G +SG +P E GDL YL L ++ +TG IP + +KL+
Sbjct: 62 LECGDATVNHRVTALTIFSG-QISGQIPPEVGDLSYLQTLVFRKLTNLTGQIPRTIAKLK 120
Query: 132 RLQKLDLGSNSLSGSIPTFLGQLKGLQEFDLSNNKLTGVIPASFGSLPSLSQFNVSFNQL 191
L+ L L +L+G +P FL +LK LQ DLS N L+G IP+S LP+L ++S N+L
Sbjct: 121 YLRSLRLSWTNLTGPVPGFLSELKNLQWVDLSFNDLSGSIPSSLSLLPNLVSLDLSRNKL 180
Query: 192 CGAIPTGLSKF 202
G+IP F
Sbjct: 181 TGSIPESFGSF 191
Score = 83.6 bits (205), Expect = 3e-15
Identities = 64/177 (36%), Positives = 80/177 (45%), Gaps = 51/177 (28%)
Query: 94 LSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSKLQ-RLQKLDLGSNSLSGSIPTFLG 152
LSG++P+ LP L L L+ K+TG IP SF ++ L L N LSG IP LG
Sbjct: 156 LSGSIPSSLSLLPNLVSLDLSRN-KLTGSIPESFGSFPAKVPDLYLSHNQLSGYIPKTLG 214
Query: 153 QL----------------------------------------------KGLQEFDLSNNK 166
L K L DL++N
Sbjct: 215 NLDFSKIDFSRNKLGGDASMLFRANKTTWYIDLSRNMLQFDLSRVVIPKTLGILDLNHNG 274
Query: 167 LTGVIPASFGSLPSLSQFNVSFNQLCGAIPTG--LSKFAKSSFDHNKCLCGAPLAAC 221
+TG IP + P L FNVS+N+LCG IPTG L +F S+ HNKCLCGAPL +C
Sbjct: 275 ITGNIPVQWTEAP-LQFFNVSYNRLCGHIPTGGTLQEFDTYSYFHNKCLCGAPLDSC 330
>UniRef100_Q8LJS5 Polygalacturonase inhibitory protein [Brassica napus]
Length = 331
Score = 128 bits (322), Expect = 9e-29
Identities = 79/189 (41%), Positives = 110/189 (57%), Gaps = 7/189 (3%)
Query: 15 LLFLSSIYFSTSNADDVEFSCNANDKATLLKIRDHFGGPNGRLSDWDNGTDCCSDWSFVG 74
LL L ++ +TS + D+ C+ +DK TLLKI+ P +S WD DCC+ W V
Sbjct: 9 LLLLFALLLTTSLSKDL---CHKDDKNTLLKIKKAMNDPYTIIS-WDPKDDCCT-WYAVE 63
Query: 75 CGKP-LPGRITVVTISRGWGLSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSKLQRL 133
CG + R+T + IS +S +P E GDLPYL +L ++P +TG IP + +KL+ L
Sbjct: 64 CGNASINHRVTSLDISND-DVSTQIPPEVGDLPYLEYLIFHKLPNLTGEIPPTITKLKYL 122
Query: 134 QKLDLGSNSLSGSIPTFLGQLKGLQEFDLSNNKLTGVIPASFGSLPSLSQFNVSFNQLCG 193
+ L L N+LSG +P FL QLK L+ +LS NKL+G IP S LP L +S N+L G
Sbjct: 123 RYLWLSWNNLSGPVPEFLSQLKNLEYINLSFNKLSGSIPGSLSLLPKLEFLELSRNKLTG 182
Query: 194 AIPTGLSKF 202
+IP F
Sbjct: 183 SIPESFGSF 191
Score = 84.7 bits (208), Expect = 2e-15
Identities = 63/177 (35%), Positives = 78/177 (43%), Gaps = 51/177 (28%)
Query: 94 LSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSKLQRL-QKLDLGSNSLSGSIPTFLG 152
LSG++P LP L FL L+ K+TG IP SF + + L L N LSGSIP LG
Sbjct: 156 LSGSIPGSLSLLPKLEFLELSRN-KLTGSIPESFGSFKGVVYALYLSHNQLSGSIPKSLG 214
Query: 153 QL----------------------------------------------KGLQEFDLSNNK 166
L K + DL++N
Sbjct: 215 NLDINQIDLSRNKLEGDASMLFGAKKTTQHIDLSRNMFQFNISKVKVAKTVNFLDLNHNS 274
Query: 167 LTGVIPASFGSLPSLSQFNVSFNQLCGAIPTG--LSKFAKSSFDHNKCLCGAPLAAC 221
LTG IP + L L FNVS+N+LCG IP G L +F + HNKCLC APL +C
Sbjct: 275 LTGSIPVQWTQL-DLQTFNVSYNRLCGRIPQGGDLQRFDAYEYLHNKCLCDAPLQSC 330
>UniRef100_Q5UBV7 Polygalacturonase inhibiting protein [Prunus mume]
Length = 330
Score = 127 bits (320), Expect = 2e-28
Identities = 82/193 (42%), Positives = 107/193 (54%), Gaps = 10/193 (5%)
Query: 10 LLHLTLLFLSSIYFSTSNADDVEFSCNANDKATLLKIRDHFGGPNGRLSDWDNGTDCCSD 69
LL LTLLF + + + S CN DK LL+I+ F P L+ W TDCC D
Sbjct: 8 LLCLTLLFSAILNPALSEL------CNQEDKKVLLQIKKAFNDPYV-LTSWKPETDCC-D 59
Query: 70 WSFVGCGKPLPGRITVVTISRGWGLSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSK 129
W V C RI +TI G +SG +PA+ GDLPYL L + P +TGPI S K
Sbjct: 60 WYCVTCDSTT-NRINSLTIFAGQ-VSGQIPAQVGDLPYLETLEFHKQPNLTGPIQPSIVK 117
Query: 130 LQRLQKLDLGSNSLSGSIPTFLGQLKGLQEFDLSNNKLTGVIPASFGSLPSLSQFNVSFN 189
L+ L+ L L ++SGS+P FL QLK L DLS + LTG IP+S LP+L+ ++ N
Sbjct: 118 LKSLKFLRLSWTNISGSVPDFLSQLKNLTFLDLSFSNLTGSIPSSLSQLPNLNALHLDRN 177
Query: 190 QLCGAIPTGLSKF 202
+L G IP +F
Sbjct: 178 KLTGHIPKSFGEF 190
Score = 87.0 bits (214), Expect = 3e-16
Identities = 61/177 (34%), Positives = 82/177 (45%), Gaps = 51/177 (28%)
Query: 94 LSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSKLQ-RLQKLDLGSNSLSGSIPTFLG 152
L+G++P+ LP LN L L + K+TG IP SF + + +L L N LSG+IPT L
Sbjct: 155 LTGSIPSSLSQLPNLNALHL-DRNKLTGHIPKSFGEFHGSVPELYLSHNQLSGNIPTSLA 213
Query: 153 QL----------------------------------------------KGLQEFDLSNNK 166
+L K L DL++NK
Sbjct: 214 KLDFNRIDFSRNKLEGDASMIFGLNKTTQIVDLSRNLLEFNLSKVEFSKSLISLDLNHNK 273
Query: 167 LTGVIPASFGSLPSLSQFNVSFNQLCGAIPTG--LSKFAKSSFDHNKCLCGAPLAAC 221
+TG IP + L NVS+N+LCG IP G L F S++ HN+CLCGAPL +C
Sbjct: 274 ITGGIPVGLTQV-DLQFLNVSYNRLCGQIPVGGKLQSFDSSTYFHNRCLCGAPLPSC 329
>UniRef100_Q9SED2 Polygalacturonase-inhibiting protein [Eucalyptus saligna]
Length = 303
Score = 127 bits (318), Expect = 3e-28
Identities = 75/167 (44%), Positives = 98/167 (57%), Gaps = 4/167 (2%)
Query: 36 NANDKATLLKIRDHFGGPNGRLSDWDNGTDCCSDWSFVGCGKPLPGRITVVTISRGWGLS 95
N +DK LL+I+ FG P L+ W + TDCC DW V C RI +TI G +S
Sbjct: 3 NPDDKKVLLQIKKAFGDPYV-LASWKSDTDCC-DWYCVTCDSTT-NRINSLTIFAGQ-VS 58
Query: 96 GTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSKLQRLQKLDLGSNSLSGSIPTFLGQLK 155
G +PA GDLPYL L + P +TGPI + +KL+ L+ L L +LSGS+P FL QLK
Sbjct: 59 GQIPALVGDLPYLETLEFHKQPNLTGPIQPAIAKLKGLKFLRLSWTNLSGSVPDFLSQLK 118
Query: 156 GLQEFDLSNNKLTGVIPASFGSLPSLSQFNVSFNQLCGAIPTGLSKF 202
L DLS N LTG IP+S LP+L+ ++ N+L G IP +F
Sbjct: 119 NLTFLDLSFNNLTGAIPSSLSQLPNLNALHLDRNKLTGHIPKSFGQF 165
Score = 80.1 bits (196), Expect = 4e-14
Identities = 60/173 (34%), Positives = 80/173 (45%), Gaps = 50/173 (28%)
Query: 94 LSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSK-LQRLQKLDLGSNSLSGSIPTFLG 152
L+G +P+ LP LN L L + K+TG IP SF + + + L L N LSG+IPT
Sbjct: 130 LTGAIPSSLSQLPNLNALHL-DRNKLTGHIPKSFGQFIGNVPDLYLSHNQLSGNIPTSFA 188
Query: 153 QL-------------------------------KGLQEFDLS--------------NNKL 167
Q+ + L EF+LS +NK+
Sbjct: 189 QMDFGKHRLSRNKLEDASVIFGLNKTTQIVDLSRNLLEFNLSKVEFPTSLTSLDVNHNKI 248
Query: 168 TGVIPASFGSLPSLSQFNVSFNQLCGAIPTG--LSKFAKSSFDHNKCLCGAPL 218
G IP F L + NVS+N+LCG IP G L F + S+ HN+CLCGAPL
Sbjct: 249 YGSIPVEFTQL-NFQFLNVSYNRLCGQIPVGGKLQSFNEYSYFHNRCLCGAPL 300
>UniRef100_Q9SED1 Polygalacturonase-inhibiting protein [Eucalyptus nitens]
Length = 303
Score = 127 bits (318), Expect = 3e-28
Identities = 75/167 (44%), Positives = 98/167 (57%), Gaps = 4/167 (2%)
Query: 36 NANDKATLLKIRDHFGGPNGRLSDWDNGTDCCSDWSFVGCGKPLPGRITVVTISRGWGLS 95
N +DK LL+I+ FG P L+ W + TDCC DW V C RI +TI G +S
Sbjct: 3 NPDDKKVLLQIKKAFGDPY-ILASWKSDTDCC-DWYCVTCDSTT-NRINSLTIFAGQ-VS 58
Query: 96 GTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSKLQRLQKLDLGSNSLSGSIPTFLGQLK 155
G +PA GDLPYL L + P +TGPI + +KL+ L+ L L +LSGS+P FL QLK
Sbjct: 59 GEIPALVGDLPYLETLEFHKQPNLTGPIQPAIAKLKGLKFLRLSWTNLSGSVPDFLSQLK 118
Query: 156 GLQEFDLSNNKLTGVIPASFGSLPSLSQFNVSFNQLCGAIPTGLSKF 202
L DLS N LTG IP+S LP+L+ ++ N+L G IP +F
Sbjct: 119 NLTFLDLSFNNLTGAIPSSLSQLPNLNALHLDRNKLTGHIPKSFGQF 165
Score = 80.1 bits (196), Expect = 4e-14
Identities = 60/173 (34%), Positives = 80/173 (45%), Gaps = 50/173 (28%)
Query: 94 LSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSK-LQRLQKLDLGSNSLSGSIPTFLG 152
L+G +P+ LP LN L L + K+TG IP SF + + + L L N LSG+IPT
Sbjct: 130 LTGAIPSSLSQLPNLNALHL-DRNKLTGHIPKSFGQFIGNVPDLYLSHNQLSGNIPTSFA 188
Query: 153 QL-------------------------------KGLQEFDLS--------------NNKL 167
Q+ + L EF+LS +NK+
Sbjct: 189 QMDFGKHRLSRNKLGDASVIFGLNKTTQIVDLSRNLLEFNLSKVEFPTSLTSLDVNHNKI 248
Query: 168 TGVIPASFGSLPSLSQFNVSFNQLCGAIPTG--LSKFAKSSFDHNKCLCGAPL 218
G IP F L + NVS+N+LCG IP G L F + S+ HN+CLCGAPL
Sbjct: 249 YGSIPVEFTQL-NFQFLNVSYNRLCGQIPVGGKLQSFNEYSYFHNRCLCGAPL 300
>UniRef100_Q9LKQ8 Polygalacturonase inhibiting protein [Prunus mahaleb]
Length = 330
Score = 126 bits (317), Expect = 4e-28
Identities = 77/190 (40%), Positives = 104/190 (54%), Gaps = 4/190 (2%)
Query: 13 LTLLFLSSIYFSTSNADDVEFSCNANDKATLLKIRDHFGGPNGRLSDWDNGTDCCSDWSF 72
L+ L ++ FST + CN DK LL+I+ F P L+ W TDCC DW
Sbjct: 5 LSTLLCLTLLFSTILNPALSELCNPEDKKVLLQIKKAFNDPYV-LTSWKPETDCC-DWYC 62
Query: 73 VGCGKPLPGRITVVTISRGWGLSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSKLQR 132
V C RI +TI G +S +P + GDLPYL L + P +TGPI S +KL+
Sbjct: 63 VTCDSTT-NRINSLTIFAGQ-VSAQIPTQVGDLPYLETLEFHKQPNLTGPIQPSIAKLKS 120
Query: 133 LQKLDLGSNSLSGSIPTFLGQLKGLQEFDLSNNKLTGVIPASFGSLPSLSQFNVSFNQLC 192
L++L L ++SGS+P FL QLK L DLS + LTG IP+S LP+L+ + N+L
Sbjct: 121 LKELRLSWTNISGSVPDFLSQLKNLTFLDLSFSNLTGSIPSSLSQLPNLNALRLDRNKLT 180
Query: 193 GAIPTGLSKF 202
G IP +F
Sbjct: 181 GHIPKSFGEF 190
Score = 88.2 bits (217), Expect = 1e-16
Identities = 62/177 (35%), Positives = 81/177 (45%), Gaps = 51/177 (28%)
Query: 94 LSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSKLQ-RLQKLDLGSNSLSGSIPTFLG 152
L+G++P+ LP LN L L + K+TG IP SF + + L L N LSG+IPT L
Sbjct: 155 LTGSIPSSLSQLPNLNALRL-DRNKLTGHIPKSFGEFHGSVPDLYLSHNQLSGTIPTSLA 213
Query: 153 QL----------------------------------------------KGLQEFDLSNNK 166
+L K L DL++NK
Sbjct: 214 KLNFSTIDFSRNKLEGDASMIFGLNKTTQIVDLSRNLLEFNLSKVEFSKSLTSLDLNHNK 273
Query: 167 LTGVIPASFGSLPSLSQFNVSFNQLCGAIPTG--LSKFAKSSFDHNKCLCGAPLAAC 221
+TG IP L L NVS+N+LCG IP G L F S++ HN+CLCGAPL +C
Sbjct: 274 ITGGIPVGLTQL-DLQFLNVSYNRLCGQIPVGGKLQSFDSSTYFHNRCLCGAPLPSC 329
>UniRef100_O22522 Polygalacturonase inhibiting protein [Prunus armeniaca]
Length = 330
Score = 126 bits (317), Expect = 4e-28
Identities = 81/193 (41%), Positives = 106/193 (53%), Gaps = 10/193 (5%)
Query: 10 LLHLTLLFLSSIYFSTSNADDVEFSCNANDKATLLKIRDHFGGPNGRLSDWDNGTDCCSD 69
LL LTLLF + + + S CN DK LL+I+ F P L+ W TDCC D
Sbjct: 8 LLCLTLLFSTILNPALSEL------CNPEDKKVLLQIKKAFNDPYV-LTSWKPETDCC-D 59
Query: 70 WSFVGCGKPLPGRITVVTISRGWGLSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSK 129
W V C RI +TI G +SG +P + GDLPYL L + P +TGPI S +K
Sbjct: 60 WYCVTCDSTT-NRINSLTIFAGQ-VSGQIPTQVGDLPYLETLEFHKQPNLTGPIQPSIAK 117
Query: 130 LQRLQKLDLGSNSLSGSIPTFLGQLKGLQEFDLSNNKLTGVIPASFGSLPSLSQFNVSFN 189
L+ L++L L ++SGS+P FL QLK L DLS + LTG IP+ LP+L+ V N
Sbjct: 118 LKLLKELRLSWTNISGSVPDFLSQLKNLTFLDLSFSNLTGSIPSWLSQLPNLNALRVDRN 177
Query: 190 QLCGAIPTGLSKF 202
+L G IP +F
Sbjct: 178 KLTGHIPKSFGEF 190
Score = 84.7 bits (208), Expect = 2e-15
Identities = 60/177 (33%), Positives = 81/177 (44%), Gaps = 51/177 (28%)
Query: 94 LSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSKLQ-RLQKLDLGSNSLSGSIPTFLG 152
L+G++P+ LP LN L + + K+TG IP SF + + L L N LSG+IPT L
Sbjct: 155 LTGSIPSWLSQLPNLNALRV-DRNKLTGHIPKSFGEFDGSVPDLYLSHNQLSGTIPTSLA 213
Query: 153 QL----------------------------------------------KGLQEFDLSNNK 166
+L K L DL++NK
Sbjct: 214 KLNFSTIDFSRNKLEGDASMIFGLNKTTQIVDLSRNLLEINLSNVEFSKSLTSLDLNHNK 273
Query: 167 LTGVIPASFGSLPSLSQFNVSFNQLCGAIPTG--LSKFAKSSFDHNKCLCGAPLAAC 221
+TG IP + L NVS+N+LCG IP G L F S++ HN+CLCGAPL +C
Sbjct: 274 ITGGIPVGLTQV-DLQFLNVSYNRLCGQIPVGGKLQSFDSSTYFHNRCLCGAPLPSC 329
>UniRef100_Q9M5J8 Polygalacturonase inhibitor 2 precursor [Arabidopsis thaliana]
Length = 330
Score = 126 bits (316), Expect = 5e-28
Identities = 78/189 (41%), Positives = 106/189 (55%), Gaps = 7/189 (3%)
Query: 16 LFLSSIYFSTSNADDVEFSCNANDKATLLKIRDHFGGPNGRLSDWDNGTDCCSDWSFVGC 75
L LS++ +TS A D+ C+ +DK TLLKI+ P L+ WD TDCCS W + C
Sbjct: 9 LLLSTLLLTTSLAKDL---CHKDDKTTLLKIKKSLNNPY-HLASWDPKTDCCS-WYCLEC 63
Query: 76 GKPLPG-RITVVTISRGWGLSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSKLQRLQ 134
G R+T + I G +SG +P E GDLPYL L ++ +TG I + +KL+ L
Sbjct: 64 GDATVNHRVTSLIIQDG-EISGQIPPEVGDLPYLTSLIFRKLTNLTGHIQPTIAKLKNLT 122
Query: 135 KLDLGSNSLSGSIPTFLGQLKGLQEFDLSNNKLTGVIPASFGSLPSLSQFNVSFNQLCGA 194
L L +L+G +P FL QLK L+ DLS N L+G IP+S SL L +S N+L G
Sbjct: 123 FLRLSWTNLTGPVPEFLSQLKNLEYIDLSFNDLSGSIPSSLSSLRKLEYLELSRNKLTGP 182
Query: 195 IPTGLSKFA 203
IP F+
Sbjct: 183 IPESFGTFS 191
Score = 84.7 bits (208), Expect = 2e-15
Identities = 60/177 (33%), Positives = 79/177 (43%), Gaps = 51/177 (28%)
Query: 94 LSGTLPAEFGDLPYLNFLSLAEMPKVTGPIPNSFSKLQ-RLQKLDLGSNSLSGSIPTFLG 152
LSG++P+ L L +L L+ K+TGPIP SF ++ L L N LSG+IP LG
Sbjct: 155 LSGSIPSSLSSLRKLEYLELSRN-KLTGPIPESFGTFSGKVPSLFLSHNQLSGTIPKSLG 213
Query: 153 Q----------------------------------------------LKGLQEFDLSNNK 166
K L D+++N
Sbjct: 214 NPDFYRIDLSRNKLQGDASILFGAKKTTWIVDISRNMFQFDLSKVKLAKTLNNLDMNHNG 273
Query: 167 LTGVIPASFGSLPSLSQFNVSFNQLCGAIPTG--LSKFAKSSFDHNKCLCGAPLAAC 221
+TG IPA + S NVS+N+LCG IP G + +F SF HNKCLCGAPL +C
Sbjct: 274 ITGSIPAEW-SKAYFQLLNVSYNRLCGRIPKGEYIQRFDSYSFFHNKCLCGAPLPSC 329
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.320 0.138 0.429
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 398,495,372
Number of Sequences: 2790947
Number of extensions: 17731712
Number of successful extensions: 68804
Number of sequences better than 10.0: 3749
Number of HSP's better than 10.0 without gapping: 2446
Number of HSP's successfully gapped in prelim test: 1346
Number of HSP's that attempted gapping in prelim test: 38597
Number of HSP's gapped (non-prelim): 18242
length of query: 221
length of database: 848,049,833
effective HSP length: 123
effective length of query: 98
effective length of database: 504,763,352
effective search space: 49466808496
effective search space used: 49466808496
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 72 (32.3 bits)
Medicago: description of AC146750.10


