

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146747.7 - phase: 0 /pseudo
(305 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
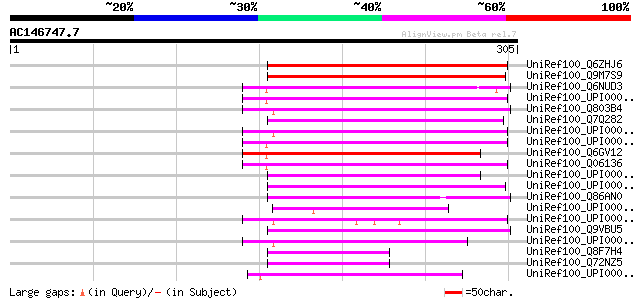
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q6ZHJ6 Putative short-chain dehydrogenase/reductase [O... 205 1e-51
UniRef100_Q9M7S9 Hypothetical protein F24F17.4 [Arabidopsis thal... 199 8e-50
UniRef100_Q6NUD3 MGC81046 protein [Xenopus laevis] 111 2e-23
UniRef100_UPI00003ABCE7 Hypothetical protein [Gallus gallus] 110 4e-23
UniRef100_Q803B4 Zgc:55947 protein [Brachydanio rerio] 108 2e-22
UniRef100_Q7Q282 ENSANGP00000014198 [Anopheles gambiae str. PEST] 108 2e-22
UniRef100_UPI0000363A89 UPI0000363A89 UniRef100 entry 107 3e-22
UniRef100_UPI00001D09A7 UPI00001D09A7 UniRef100 entry 107 5e-22
UniRef100_Q6GV12 Follicular lymphoma variant translocation 1 [Mu... 106 7e-22
UniRef100_Q06136 Follicular variant translocation protein 1 prec... 106 9e-22
UniRef100_UPI00004319F3 UPI00004319F3 UniRef100 entry 99 2e-19
UniRef100_UPI0000221EE3 UPI0000221EE3 UniRef100 entry 89 1e-16
UniRef100_Q86AN0 Similar to Homo sapiens (Human). Follicular var... 86 2e-15
UniRef100_UPI000023DEE0 UPI000023DEE0 UniRef100 entry 75 2e-12
UniRef100_UPI0000318CA3 UPI0000318CA3 UniRef100 entry 75 3e-12
UniRef100_Q9VBU5 CG10425-PA [Drosophila melanogaster] 73 8e-12
UniRef100_UPI000042CB6F UPI000042CB6F UniRef100 entry 72 2e-11
UniRef100_Q8F7H4 3-oxoacyl-(Acyl carrier protein) reductase [Lep... 64 4e-09
UniRef100_Q72NZ5 3-oxoacyl- [Leptospira interrogans] 64 4e-09
UniRef100_UPI00003C1E21 UPI00003C1E21 UniRef100 entry 62 2e-08
>UniRef100_Q6ZHJ6 Putative short-chain dehydrogenase/reductase [Oryza sativa]
Length = 327
Score = 205 bits (521), Expect = 1e-51
Identities = 97/144 (67%), Positives = 118/144 (81%)
Query: 156 ITSWSAYSASKFGLRGLAEALQQEVIGDNIHVSLIFPPDTDTPGLVEENKRKPELTKIIA 215
+ ++AYSASKF LRGL EALQ EVI DNIHVSLIFPPDT+TPG EENKR+PELT IIA
Sbjct: 181 VYGYTAYSASKFALRGLGEALQHEVIADNIHVSLIFPPDTETPGFAEENKRRPELTNIIA 240
Query: 216 ASSGFMKADEVAQKAFDGIRSGSFIISCNLEGIALSLATSGLSPQRSFLMAFVEVIAAGI 275
SSG MKAD+VA+KA DGI+SG FI+ CN EG L++AT+GLSPQ S L AF+E+I AG+
Sbjct: 241 GSSGGMKADDVARKALDGIKSGKFIVPCNFEGAMLAVATAGLSPQSSPLTAFLEIIGAGV 300
Query: 276 MRIAALCFQWNWYGSIEKWHKQRK 299
MR AA+CFQ+NW+ +IE W+ + K
Sbjct: 301 MRFAAICFQFNWFMTIENWYAKNK 324
>UniRef100_Q9M7S9 Hypothetical protein F24F17.4 [Arabidopsis thaliana]
Length = 327
Score = 199 bits (506), Expect = 8e-50
Identities = 94/143 (65%), Positives = 120/143 (83%)
Query: 156 ITSWSAYSASKFGLRGLAEALQQEVIGDNIHVSLIFPPDTDTPGLVEENKRKPELTKIIA 215
+ ++AYSASKFGL+GLA+ALQQEVI D+IHV+LIFPPDT+TPG EE K +PE+T IIA
Sbjct: 182 VYGYAAYSASKFGLQGLAQALQQEVISDDIHVTLIFPPDTNTPGFEEEQKSRPEVTAIIA 241
Query: 216 ASSGFMKADEVAQKAFDGIRSGSFIISCNLEGIALSLATSGLSPQRSFLMAFVEVIAAGI 275
ASSG M+ +EVA+KA DGI++G+F +SCN EG LSLAT+G+SPQRSF +AF+EVI AG
Sbjct: 242 ASSGSMETEEVAKKAMDGIKAGNFTVSCNFEGFLLSLATTGMSPQRSFWLAFLEVITAGP 301
Query: 276 MRIAALCFQWNWYGSIEKWHKQR 298
+R+ AL FQW+WY +IEKW K +
Sbjct: 302 IRLIALFFQWDWYKAIEKWSKTK 324
>UniRef100_Q6NUD3 MGC81046 protein [Xenopus laevis]
Length = 332
Score = 111 bits (278), Expect = 2e-23
Identities = 69/169 (40%), Positives = 97/169 (56%), Gaps = 9/169 (5%)
Query: 141 RRIGKILYLLLLL--CFITSWSAYSASKFGLRGLAEALQQEVIGDNIHVSLIFPPDTDTP 198
RR+G+I+++ + ++AYS +KF LRGLAEALQ EV NI+V++ +PPDTDTP
Sbjct: 163 RRMGRIVFVSSQAGQLGLFGYTAYSPTKFALRGLAEALQMEVKPYNIYVTVAYPPDTDTP 222
Query: 199 GLVEENKRKPELTKIIAASSGFMKADEVAQKAFDGIRSGSFIISCNLEGIALSLATSGLS 258
G EENK KP TK+I+ SS +AD+VA+ G+F S +G LS T G+S
Sbjct: 223 GFAEENKTKPLETKLISESSSLCQADQVARVIVKDAIQGNFNSSVGSDGYMLSSLTCGMS 282
Query: 259 PQRSFLMAFVEVIAAGIMRIAALCFQWNWYGSI------EKWHKQRKCK 301
P + +V+ G+ RI AL F + SI +K Q+ CK
Sbjct: 283 PVTTITEGLQQVVTMGLFRIIAL-FYLGSFDSIVRRCMMQKRDSQKSCK 330
>UniRef100_UPI00003ABCE7 Hypothetical protein [Gallus gallus]
Length = 332
Score = 110 bits (276), Expect = 4e-23
Identities = 63/161 (39%), Positives = 96/161 (59%), Gaps = 2/161 (1%)
Query: 141 RRIGKILYLLLLL--CFITSWSAYSASKFGLRGLAEALQQEVIGDNIHVSLIFPPDTDTP 198
RR+G+I+++ + ++AYS +KF LRGLAEALQ EV NI+V++ +PPDTDTP
Sbjct: 163 RRMGRIVFVSSQAGQLGLFGYTAYSPTKFALRGLAEALQMEVKPYNIYVTVAYPPDTDTP 222
Query: 199 GLVEENKRKPELTKIIAASSGFMKADEVAQKAFDGIRSGSFIISCNLEGIALSLATSGLS 258
G EE+K KP TK+I+ +S +A++VA+ G+F S +G LS+ TSG+S
Sbjct: 223 GFAEESKTKPLETKLISETSSVCQAEQVARVIVKDAIQGNFNSSVGSDGYMLSILTSGMS 282
Query: 259 PQRSFLMAFVEVIAAGIMRIAALCFQWNWYGSIEKWHKQRK 299
P S +V+ GI RI L + ++ + + QR+
Sbjct: 283 PVTSITEGLQQVVCMGIFRIVGLFYLGSFDSIVRRCMMQRE 323
>UniRef100_Q803B4 Zgc:55947 protein [Brachydanio rerio]
Length = 332
Score = 108 bits (270), Expect = 2e-22
Identities = 64/164 (39%), Positives = 96/164 (58%), Gaps = 3/164 (1%)
Query: 141 RRIGKILYLLLLLCFIT--SWSAYSASKFGLRGLAEALQQEVIGDNIHVSLIFPPDTDTP 198
RR+G+I+++ I ++AYS SKF LRGLAEALQ E+ NI+V++ +PPDTDTP
Sbjct: 163 RRMGRIMFVSSQAGQIGLFGYTAYSPSKFALRGLAEALQMEMKPYNIYVTVAYPPDTDTP 222
Query: 199 GLVEENKRKPELTKIIAASSGFMKADEVAQKAFDGIRSGSFIISCNLEGIALSLATSGLS 258
G EENK KP TK+I+ +SG + ++VA+ G+F S +G LS T G+S
Sbjct: 223 GFAEENKTKPLETKLISETSGVSQPEQVAKIVVKDAVQGNFTSSFGPDGYMLSALTCGMS 282
Query: 259 PQRSFLMAFVEVIAAGIMRIAALCFQWNWYGSIEKWHKQR-KCK 301
P S +++ G+ R AL + ++ + + QR +CK
Sbjct: 283 PVTSITEGLQQIVTMGLFRTIALFYLGSFDSIVRRCMIQREQCK 326
>UniRef100_Q7Q282 ENSANGP00000014198 [Anopheles gambiae str. PEST]
Length = 323
Score = 108 bits (269), Expect = 2e-22
Identities = 58/142 (40%), Positives = 78/142 (54%)
Query: 156 ITSWSAYSASKFGLRGLAEALQQEVIGDNIHVSLIFPPDTDTPGLVEENKRKPELTKIIA 215
I + AY+ASKF LRGLAE + E + V+L P DTDTPG EN+ KPE TKII+
Sbjct: 182 IYGYGAYAASKFALRGLAETIAMEARHRGVSVTLALPADTDTPGFENENRTKPEETKIIS 241
Query: 216 ASSGFMKADEVAQKAFDGIRSGSFIISCNLEGIALSLATSGLSPQRSFLMAFVEVIAAGI 275
S G K ++V ++ GSF LE LS+ G++P R L+ FV+ G
Sbjct: 242 GSGGLAKPEDVGKRLVQDALKGSFFSIMGLESWVLSILCVGMAPWRGPLLCFVQFYLLGP 301
Query: 276 MRIAALCFQWNWYGSIEKWHKQ 297
+R+ L QWN+ I+ KQ
Sbjct: 302 LRLIGLVLQWNFQRIIKNCAKQ 323
>UniRef100_UPI0000363A89 UPI0000363A89 UniRef100 entry
Length = 332
Score = 107 bits (268), Expect = 3e-22
Identities = 61/161 (37%), Positives = 95/161 (58%), Gaps = 2/161 (1%)
Query: 141 RRIGKILYLLLLLCFIT--SWSAYSASKFGLRGLAEALQQEVIGDNIHVSLIFPPDTDTP 198
RR+G+I+++ I ++AYS SKF LRGLAE+LQ E+ NI+V++ +PPDTDTP
Sbjct: 163 RRMGRIMFVSSQAGQIGLFGYTAYSPSKFALRGLAESLQMEIKPYNIYVTVAYPPDTDTP 222
Query: 199 GLVEENKRKPELTKIIAASSGFMKADEVAQKAFDGIRSGSFIISCNLEGIALSLATSGLS 258
GL EEN+ KP TK+I+ +SG + D+VA+ G+F S +G L+ T G+S
Sbjct: 223 GLAEENRTKPLETKLISETSGVCQPDQVAKVIVRDAVQGNFNSSVGPDGYMLAAVTCGMS 282
Query: 259 PQRSFLMAFVEVIAAGIMRIAALCFQWNWYGSIEKWHKQRK 299
P S +++ G+ R AL + ++ + + QR+
Sbjct: 283 PVTSITEGLQQIVTMGLFRTIALFYLGSFDSIVRRCMIQRE 323
>UniRef100_UPI00001D09A7 UPI00001D09A7 UniRef100 entry
Length = 332
Score = 107 bits (266), Expect = 5e-22
Identities = 59/161 (36%), Positives = 96/161 (58%), Gaps = 2/161 (1%)
Query: 141 RRIGKILYLLLLL--CFITSWSAYSASKFGLRGLAEALQQEVIGDNIHVSLIFPPDTDTP 198
RR+G+I+++ + ++AYS+SKF +RGLAEALQ EV N++V++ +PPDTDTP
Sbjct: 163 RRVGRIVFVSSQAGQLGLFGFTAYSSSKFAIRGLAEALQMEVKPYNVYVTVAYPPDTDTP 222
Query: 199 GLVEENKRKPELTKIIAASSGFMKADEVAQKAFDGIRSGSFIISCNLEGIALSLATSGLS 258
GL EENK KP T++I+ ++ K ++VA++ G+F S +G LS T G++
Sbjct: 223 GLAEENKTKPLETRLISETTAVCKPEQVAKQIVKDAIQGNFNSSIGSDGYMLSSLTCGMA 282
Query: 259 PQRSFLMAFVEVIAAGIMRIAALCFQWNWYGSIEKWHKQRK 299
P S +V+ G+ R AL + ++ I + Q++
Sbjct: 283 PVTSITEGLQQVVTMGLFRTIALFYLGSFDNIIRRCMVQKR 323
>UniRef100_Q6GV12 Follicular lymphoma variant translocation 1 [Mus musculus]
Length = 332
Score = 106 bits (265), Expect = 7e-22
Identities = 57/145 (39%), Positives = 89/145 (61%), Gaps = 2/145 (1%)
Query: 141 RRIGKILYLLLLL--CFITSWSAYSASKFGLRGLAEALQQEVIGDNIHVSLIFPPDTDTP 198
RR+G+I+++ + ++AYS+SKF +RGLAEALQ EV N++V++ +PPDTDTP
Sbjct: 163 RRVGRIVFVSSQAGQLGLFGFTAYSSSKFAIRGLAEALQMEVKPYNVYVTVAYPPDTDTP 222
Query: 199 GLVEENKRKPELTKIIAASSGFMKADEVAQKAFDGIRSGSFIISCNLEGIALSLATSGLS 258
GL EENK KP T++I+ ++ K ++VA++ G+F S +G LS T G++
Sbjct: 223 GLAEENKTKPLETRLISETTAICKPEQVAKQIVKDAIQGNFNSSIGSDGYMLSSLTCGMA 282
Query: 259 PQRSFLMAFVEVIAAGIMRIAALCF 283
P S +V+ G+ R AL +
Sbjct: 283 PVTSITEGLQQVVTMGLFRTIALFY 307
>UniRef100_Q06136 Follicular variant translocation protein 1 precursor [Homo sapiens]
Length = 332
Score = 106 bits (264), Expect = 9e-22
Identities = 57/161 (35%), Positives = 95/161 (58%), Gaps = 2/161 (1%)
Query: 141 RRIGKILYLLLLL--CFITSWSAYSASKFGLRGLAEALQQEVIGDNIHVSLIFPPDTDTP 198
RR+G+I+++ + ++AYSASKF +RGLAEALQ EV N+++++ +PPDTDTP
Sbjct: 163 RRVGRIVFVSSQAGQLGLFGFTAYSASKFAIRGLAEALQMEVKPYNVYITVAYPPDTDTP 222
Query: 199 GLVEENKRKPELTKIIAASSGFMKADEVAQKAFDGIRSGSFIISCNLEGIALSLATSGLS 258
G EEN+ KP T++I+ ++ K ++VA++ G+F S +G LS T G++
Sbjct: 223 GFAEENRTKPLETRLISETTSVCKPEQVAKQIVKDAIQGNFNSSLGSDGYMLSALTCGMA 282
Query: 259 PQRSFLMAFVEVIAAGIMRIAALCFQWNWYGSIEKWHKQRK 299
P S +V+ G+ R AL + ++ + + QR+
Sbjct: 283 PVTSITEGLQQVVTMGLFRTIALFYLGSFDSIVRRCMMQRE 323
>UniRef100_UPI00004319F3 UPI00004319F3 UniRef100 entry
Length = 293
Score = 98.6 bits (244), Expect = 2e-19
Identities = 50/128 (39%), Positives = 76/128 (59%)
Query: 156 ITSWSAYSASKFGLRGLAEALQQEVIGDNIHVSLIFPPDTDTPGLVEENKRKPELTKIIA 215
I +SAY ++KF LRGLAE++ E+ NI V+L PPDT+TPG E KP TK+I+
Sbjct: 145 IFGYSAYCSTKFALRGLAESISMELAPYNISVTLSLPPDTNTPGFAIEELSKPLETKLIS 204
Query: 216 ASSGFMKADEVAQKAFDGIRSGSFIISCNLEGIALSLATSGLSPQRSFLMAFVEVIAAGI 275
++S + + VA K F +G F + +EG L++ SG+SP S F++ + GI
Sbjct: 205 STSSLVSPELVADKLFKDALAGKFFSTVGMEGFILTVLCSGMSPFNSIRELFIQSMLMGI 264
Query: 276 MRIAALCF 283
R+ + C+
Sbjct: 265 FRLISACY 272
>UniRef100_UPI0000221EE3 UPI0000221EE3 UniRef100 entry
Length = 345
Score = 89.0 bits (219), Expect = 1e-16
Identities = 45/143 (31%), Positives = 79/143 (54%)
Query: 156 ITSWSAYSASKFGLRGLAEALQQEVIGDNIHVSLIFPPDTDTPGLVEENKRKPELTKIIA 215
I +SAYS +KF LRG A+ L E++ ++V++++PP+TDT G E + PE TK+++
Sbjct: 188 IFGYSAYSPTKFALRGFADTLHMELLPYKVNVAVLYPPNTDTEGFKLELETMPEETKLMS 247
Query: 216 ASSGFMKADEVAQKAFDGIRSGSFIISCNLEGIALSLATSGLSPQRSFLMAFVEVIAAGI 275
++G VA+ I G++ + L+G L T+G P++S A + AG+
Sbjct: 248 DAAGLFTPKHVAEAHLKDIADGNYTTTIGLDGWMLGCLTAGACPEKSLFRALTQSALAGV 307
Query: 276 MRIAALCFQWNWYGSIEKWHKQR 298
R L + + G +K +++R
Sbjct: 308 FRAITLVYLGYFNGITKKCYRRR 330
>UniRef100_Q86AN0 Similar to Homo sapiens (Human). Follicular variant translocation
protein 1 [Dictyostelium discoideum]
Length = 323
Score = 85.5 bits (210), Expect = 2e-15
Identities = 49/147 (33%), Positives = 80/147 (54%), Gaps = 4/147 (2%)
Query: 156 ITSWSAYSASKFGLRGLAEALQQEVIGDNIHVSLIFPPDTDTPGLVEENKRKPELTKIIA 215
+ +S Y SKF LRGLAE L+ E+ I S+++PPDTDTPG +EN KPE T I+
Sbjct: 179 VPGYSTYCPSKFALRGLAETLRSELKPYKITFSVVYPPDTDTPGYQQENLTKPEETVAIS 238
Query: 216 ASSGFMKADEVAQKAFDGIRSGSFIISCNLEGIALSLATSGLSPQRSFLMAFVEVIAAGI 275
+ EVA+ GI++G + I+ ++ ++ + GL+P F +F +++ A I
Sbjct: 239 GGGKAVSPLEVAKSIVSGIKNGDYHIAYDVPTKLCAVLSPGLTP---FYFSFFDILLAPI 295
Query: 276 MR-IAALCFQWNWYGSIEKWHKQRKCK 301
R + + N ++ W K+ + K
Sbjct: 296 CRLVGIIAMNQNDAEVLKSWKKRNQLK 322
>UniRef100_UPI000023DEE0 UPI000023DEE0 UniRef100 entry
Length = 346
Score = 75.5 bits (184), Expect = 2e-12
Identities = 41/108 (37%), Positives = 62/108 (56%), Gaps = 2/108 (1%)
Query: 159 WSAYSASKFGLRGLAEALQQEVI--GDNIHVSLIFPPDTDTPGLVEENKRKPELTKIIAA 216
++ Y+ SK+ +RGLAE +QQEV+ N+ V L+ P +PG + EN KPE+TKII
Sbjct: 197 YTPYTPSKYAMRGLAECIQQEVLLYPQNVQVHLLLPGTILSPGNIRENVNKPEITKIIEK 256
Query: 217 SSGFMKADEVAQKAFDGIRSGSFIISCNLEGIALSLATSGLSPQRSFL 264
DEVA + G+ +G I+ N G + ++ G +P+ SFL
Sbjct: 257 DDPKQTPDEVAALSIKGLEAGQHNITVNFLGHLMKWSSFGGTPKNSFL 304
>UniRef100_UPI0000318CA3 UPI0000318CA3 UniRef100 entry
Length = 409
Score = 74.7 bits (182), Expect = 3e-12
Identities = 60/212 (28%), Positives = 92/212 (43%), Gaps = 53/212 (25%)
Query: 141 RRIGKILYLLLLLCFIT--SWSAYSASKFGLRGLAEALQQEVIGDNIHVSLIFPPDTDTP 198
RR+G+I+++ I ++AYS SKF LRGLAE+LQ E+ NI+V++ +PPDTDTP
Sbjct: 190 RRMGRIMFVSSQAGQIGLFGYTAYSPSKFALRGLAESLQMEIKPYNIYVTVAYPPDTDTP 249
Query: 199 GLVEENKRK---------------------PELTKIIAASS-------GFMKADEVAQKA 230
GL EENK K P + +A+ G + A Q
Sbjct: 250 GLAEENKTKVRRVSARRSLRSLLSWFPYGSPLFSSCLASGDQTHLRDVGGLPAGPSGQSD 309
Query: 231 FDG-----------------------IRSGSFIISCNLEGIALSLATSGLSPQRSFLMAF 267
G ++ G+F S +G LS T G+SP S
Sbjct: 310 RSGRSGETQNRLLCEPSARVLISLSDLQQGNFNSSVGPDGYMLSAVTCGMSPVTSITEGL 369
Query: 268 VEVIAAGIMRIAALCFQWNWYGSIEKWHKQRK 299
+++ G+ R AL + ++ + + QR+
Sbjct: 370 QQIVTMGLFRTIALFYLGSFDSIVRRCMIQRE 401
>UniRef100_Q9VBU5 CG10425-PA [Drosophila melanogaster]
Length = 336
Score = 73.2 bits (178), Expect = 8e-12
Identities = 43/146 (29%), Positives = 66/146 (44%)
Query: 156 ITSWSAYSASKFGLRGLAEALQQEVIGDNIHVSLIFPPDTDTPGLVEENKRKPELTKIIA 215
I + YSA+K+ LR +AE + E + V+L P DT+TPG EE K KP TKII+
Sbjct: 183 IYGYGPYSATKYALRAMAETIAMESREHGVSVTLAMPCDTNTPGFEEEEKSKPRETKIIS 242
Query: 216 ASSGFMKADEVAQKAFDGIRSGSFIISCNLEGIALSLATSGLSPQRSFLMAFVEVIAAGI 275
G ++ + +A+ G F + E ++ L P F + I G
Sbjct: 243 GGGGLIEPEVMAKAILKDALKGKFTSTVGAESWLITTLGGALLPWDGFFTNLLHAIVLGP 302
Query: 276 MRIAALCFQWNWYGSIEKWHKQRKCK 301
+RI + + I K ++ K K
Sbjct: 303 LRIVSYALHKYFNSVIRKCAREDKAK 328
>UniRef100_UPI000042CB6F UPI000042CB6F UniRef100 entry
Length = 335
Score = 72.0 bits (175), Expect = 2e-11
Identities = 41/137 (29%), Positives = 75/137 (53%), Gaps = 2/137 (1%)
Query: 141 RRIGKILYLLLLLCFIT--SWSAYSASKFGLRGLAEALQQEVIGDNIHVSLIFPPDTDTP 198
RR GKI+++ L +++ +S+YS +K+ LRGL++AL+ E++ NI + + P P
Sbjct: 167 RRTGKIIFVASFLSYVSFAGYSSYSPAKYALRGLSDALRSEMLLHNIDIHIFLPCGISGP 226
Query: 199 GLVEENKRKPELTKIIAASSGFMKADEVAQKAFDGIRSGSFIISCNLEGIALSLATSGLS 258
G EN+ KP +TK I + + A G++ G + I+ NL + L ++G
Sbjct: 227 GFDAENRTKPAVTKKIEEGDTPITPEVCAAALESGLKKGYYQITDNLVTEPIRLRSNGGV 286
Query: 259 PQRSFLMAFVEVIAAGI 275
P +FL+ + +I + +
Sbjct: 287 PTNNFLLDTLWLIVSSV 303
>UniRef100_Q8F7H4 3-oxoacyl-(Acyl carrier protein) reductase [Leptospira interrogans]
Length = 287
Score = 64.3 bits (155), Expect = 4e-09
Identities = 31/73 (42%), Positives = 44/73 (59%)
Query: 156 ITSWSAYSASKFGLRGLAEALQQEVIGDNIHVSLIFPPDTDTPGLVEENKRKPELTKIIA 215
I + AYSASKF + G A++L+QE++ + V + PP TDTPGL +EN+ KPEL K +
Sbjct: 158 IYGYGAYSASKFAIIGFAQSLRQEMMFHGVRVKVFLPPTTDTPGLEKENRDKPELVKEME 217
Query: 216 ASSGFMKADEVAQ 228
S V +
Sbjct: 218 MGSAINSVHSVGK 230
>UniRef100_Q72NZ5 3-oxoacyl- [Leptospira interrogans]
Length = 287
Score = 64.3 bits (155), Expect = 4e-09
Identities = 31/73 (42%), Positives = 44/73 (59%)
Query: 156 ITSWSAYSASKFGLRGLAEALQQEVIGDNIHVSLIFPPDTDTPGLVEENKRKPELTKIIA 215
I + AYSASKF + G A++L+QE++ + V + PP TDTPGL +EN+ KPEL K +
Sbjct: 158 IYGYGAYSASKFAIIGFAQSLRQEMMFHGVRVKVFLPPTTDTPGLEKENRDKPELVKEME 217
Query: 216 ASSGFMKADEVAQ 228
S V +
Sbjct: 218 MGSAINSVHSVGK 230
>UniRef100_UPI00003C1E21 UPI00003C1E21 UniRef100 entry
Length = 268
Score = 61.6 bits (148), Expect = 2e-08
Identities = 45/132 (34%), Positives = 64/132 (48%), Gaps = 3/132 (2%)
Query: 144 GKILYL--LLLLCFITSWSAYSASKFGLRGLAEALQQEVIGDNIHVSLIFPPDTDTPGLV 201
GKI+ + L L + +S Y+ K +RGLAE+L+ E+I I V FP +PGL
Sbjct: 104 GKIVLVSSTLGLIGLVGYSQYAPMKHAIRGLAESLRSELILYGISVHAYFPGTILSPGLE 163
Query: 202 EENKRKPELTKIIAASSGFMKADEVAQKAFDGIRSGSFIISCNLEGIALSLATSGLSP-Q 260
EENK KP++T I + + A+ G+ G F I+ + A G SP
Sbjct: 164 EENKTKPKITLEIEGQDDGLTPAQCAKGLIKGVERGHFFITTDFNTELFRSAALGASPTN 223
Query: 261 RSFLMAFVEVIA 272
R L V +IA
Sbjct: 224 RGVLDRAVALIA 235
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.337 0.143 0.471
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 459,920,670
Number of Sequences: 2790947
Number of extensions: 17989110
Number of successful extensions: 353727
Number of sequences better than 10.0: 3245
Number of HSP's better than 10.0 without gapping: 1357
Number of HSP's successfully gapped in prelim test: 2001
Number of HSP's that attempted gapping in prelim test: 297307
Number of HSP's gapped (non-prelim): 29374
length of query: 305
length of database: 848,049,833
effective HSP length: 127
effective length of query: 178
effective length of database: 493,599,564
effective search space: 87860722392
effective search space used: 87860722392
T: 11
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.8 bits)
S2: 74 (33.1 bits)
Medicago: description of AC146747.7


