

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146747.6 + phase: 0
(498 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
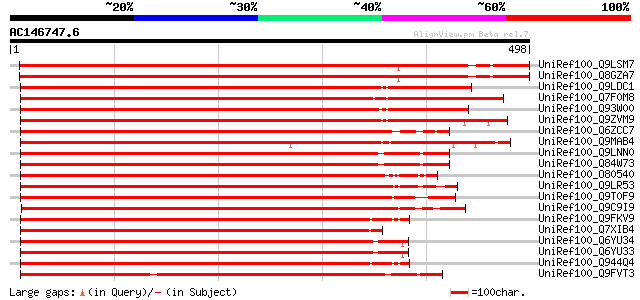
Sequences producing significant alignments: (bits) Value
UniRef100_Q9LSM7 Cyclin-dependent protein kinase-like [Arabidops... 714 0.0
UniRef100_Q8GZA7 Putative cyclin-dependent protein kinase [Arabi... 712 0.0
UniRef100_Q9LDC1 CRK1 protein [Beta vulgaris subsp. vulgaris] 684 0.0
UniRef100_Q7F0M8 Putative CRK1 protein [Oryza sativa] 677 0.0
UniRef100_Q93W00 Putative CRK1 protein [Oryza sativa] 676 0.0
UniRef100_Q9ZVM9 Hypothetical protein T22H22.5 [Arabidopsis thal... 660 0.0
UniRef100_Q6ZCC7 Putative CRK1 protein [Oryza sativa] 592 e-167
UniRef100_Q9MAB4 Putative cyclin-dependent protein kinase [Arabi... 590 e-167
UniRef100_Q9LNN0 F8L10.9 protein [Arabidopsis thaliana] 587 e-166
UniRef100_Q84W73 Putative cell division-related protein [Arabido... 585 e-165
UniRef100_O80540 F14J9.26 protein [Arabidopsis thaliana] 556 e-157
UniRef100_Q9LR53 F21B7.34 [Arabidopsis thaliana] 540 e-152
UniRef100_Q9T0F9 Hypothetical protein T5L19.140 [Arabidopsis tha... 538 e-151
UniRef100_Q9C9I9 Hypothetical protein F26A9.10 [Arabidopsis thal... 528 e-148
UniRef100_Q9FKV9 Cyclin-dependent protein kinase-like protein [A... 525 e-147
UniRef100_Q7XIB4 Putative cyclin-dependent kinase CDC2C [Oryza s... 525 e-147
UniRef100_Q6YU34 Putative CRK1 protein [Oryza sativa] 525 e-147
UniRef100_Q6YU33 Putative CRK1 protein [Oryza sativa] 525 e-147
UniRef100_Q944Q4 AT5g44290/K9L2_5 [Arabidopsis thaliana] 524 e-147
UniRef100_Q9FVT3 CRK1 protein, putative [Arabidopsis thaliana] 523 e-147
>UniRef100_Q9LSM7 Cyclin-dependent protein kinase-like [Arabidopsis thaliana]
Length = 580
Score = 714 bits (1844), Expect = 0.0
Identities = 353/493 (71%), Positives = 415/493 (83%), Gaps = 13/493 (2%)
Query: 10 QGWPSWLMAVAGEAIGDWTPRRANSFEKLAKIGQGTYSNVYKAKDLVTGKIVALKKVRFD 69
+GWP WL+A G++I D TPRRA ++EKL KIGQGTYSNVYKAKDL++GKIVALKKVRFD
Sbjct: 89 EGWPPWLIAACGDSIKDLTPRRATTYEKLEKIGQGTYSNVYKAKDLLSGKIVALKKVRFD 148
Query: 70 NLEPESVKFMAREILVLRKLDHPNVVKLEGLVTSRMSCSLYLVFEYMEHDLAGLSAGQGV 129
NLE ESVKFMAREILVLR+L+HPNV+KL+GLVTSR+SCSLYLVFEYMEHDL+GL+A QG+
Sbjct: 149 NLEAESVKFMAREILVLRRLNHPNVIKLQGLVTSRVSCSLYLVFEYMEHDLSGLAATQGL 208
Query: 130 KFTEPQVKCFMKQLLSGLEHCHSRGVLHRDIKGSNLLIDNEGILKIADFGLATFYNPNKK 189
KF PQVKCFMKQLLSGLEHCHSRGVLHRDIKGSNLLIDN+GILKIADFGLATFY+P +K
Sbjct: 209 KFDLPQVKCFMKQLLSGLEHCHSRGVLHRDIKGSNLLIDNDGILKIADFGLATFYDPKQK 268
Query: 190 QSMTSRVVTLWYRPPELLLGATFYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIF 249
Q+MTSRVVTLWYRPPELLLGAT YG G+DLWSAGCI+AELLAGKP+MPGRTEVEQLHKIF
Sbjct: 269 QTMTSRVVTLWYRPPELLLGATSYGTGVDLWSAGCIMAELLAGKPVMPGRTEVEQLHKIF 328
Query: 250 KLCGSPAEEYWRKHKLPNATIFKPQQPYKRCISETFKDFPPSSLPLIDSLLAIDPDRRGT 309
KLCGSP++ YW+K++LPNAT+FKPQ PYKRC++E F F PSS+ L+++LL IDP RGT
Sbjct: 329 KLCGSPSDSYWKKYRLPNATLFKPQHPYKRCVAEAFNGFTPSSVHLVETLLTIDPADRGT 388
Query: 310 ASAALNHEFFTTEPYACEPSSLPKYPPSKELDVKMRDEEARRQKALNGKANAVDGAKRVR 369
+++ALN EFFTTEP C+PSSLPKYPPSKEL+VK+RDEE RRQK L GK + +DGA+R+R
Sbjct: 389 STSALNSEFFTTEPLPCDPSSLPKYPPSKELNVKLRDEELRRQKGLAGKGSGIDGARRIR 448
Query: 370 AR--ERGRAIPAPEANAEIQTNLDRWRVVTHANAKSKSEKFPPPHQDGAVGYPQDESSKG 427
R GRAIPAPEANAE Q NLDRWR ++ N KSKSEKFPPPHQDGAVGYP ++ SK
Sbjct: 449 YRGDRTGRAIPAPEANAESQANLDRWRSISQTNGKSKSEKFPPPHQDGAVGYPLEDLSKK 508
Query: 428 PVSFGA-SDTSFSSGTFNVKPSGPTRSHDGTGLHKGTKTKKEESQMASSWKFMRPFK-PS 485
FGA ++TSF S +S +GT + K + K+ ++ ASS K++ K P
Sbjct: 509 TSVFGAKTETSFGL-------SRSLKSGEGTSMRK--ISNKDGARGASSRKYIWGLKPPP 559
Query: 486 TIGLSMDLLFRSK 498
+GLSMDLLFRS+
Sbjct: 560 ALGLSMDLLFRSR 572
>UniRef100_Q8GZA7 Putative cyclin-dependent protein kinase [Arabidopsis thaliana]
Length = 580
Score = 712 bits (1839), Expect = 0.0
Identities = 352/493 (71%), Positives = 414/493 (83%), Gaps = 13/493 (2%)
Query: 10 QGWPSWLMAVAGEAIGDWTPRRANSFEKLAKIGQGTYSNVYKAKDLVTGKIVALKKVRFD 69
+GWP WL+A G++I D TPRRA ++EKL KIGQGTYSNVYKAKDL++GKIVALKKVRFD
Sbjct: 89 EGWPPWLIAACGDSIKDLTPRRATTYEKLEKIGQGTYSNVYKAKDLLSGKIVALKKVRFD 148
Query: 70 NLEPESVKFMAREILVLRKLDHPNVVKLEGLVTSRMSCSLYLVFEYMEHDLAGLSAGQGV 129
NLE ESVKFMAREILVLR+L+HPNV+KL+GLVTSR+SCSLYLVFEYMEHDL+GL+A QG+
Sbjct: 149 NLEAESVKFMAREILVLRRLNHPNVIKLQGLVTSRVSCSLYLVFEYMEHDLSGLAATQGL 208
Query: 130 KFTEPQVKCFMKQLLSGLEHCHSRGVLHRDIKGSNLLIDNEGILKIADFGLATFYNPNKK 189
KF PQVKCFMKQLLSGLEHCHSRGVLHRDIKGSNLLIDN+GILKIADFGLATFY+P +K
Sbjct: 209 KFDLPQVKCFMKQLLSGLEHCHSRGVLHRDIKGSNLLIDNDGILKIADFGLATFYDPKQK 268
Query: 190 QSMTSRVVTLWYRPPELLLGATFYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIF 249
Q+MTSRVVTLWYRPPELLLGAT YG G+DLWSAGCI+AELLAGKP+MPGRTEVEQLHKIF
Sbjct: 269 QTMTSRVVTLWYRPPELLLGATSYGTGVDLWSAGCIMAELLAGKPVMPGRTEVEQLHKIF 328
Query: 250 KLCGSPAEEYWRKHKLPNATIFKPQQPYKRCISETFKDFPPSSLPLIDSLLAIDPDRRGT 309
KLCGSP++ YW+K++LPNAT+FKPQ PYKRC++E F F PSS+ L+++LL IDP RGT
Sbjct: 329 KLCGSPSDSYWKKYRLPNATLFKPQHPYKRCVAEAFNGFTPSSVHLVETLLTIDPADRGT 388
Query: 310 ASAALNHEFFTTEPYACEPSSLPKYPPSKELDVKMRDEEARRQKALNGKANAVDGAKRVR 369
+++ALN EFFTTEP C+PSSLPKYPPSKEL+VK+RDEE RRQK L GK + +DGA+R+R
Sbjct: 389 STSALNSEFFTTEPLPCDPSSLPKYPPSKELNVKLRDEELRRQKGLAGKGSGIDGARRIR 448
Query: 370 AR--ERGRAIPAPEANAEIQTNLDRWRVVTHANAKSKSEKFPPPHQDGAVGYPQDESSKG 427
R GRAIPAPEANAE Q NLDRWR ++ N KSKSEKFPPPHQDGAVGYP ++ SK
Sbjct: 449 YRGDRTGRAIPAPEANAESQANLDRWRSISQTNGKSKSEKFPPPHQDGAVGYPLEDLSKK 508
Query: 428 PVSFGA-SDTSFSSGTFNVKPSGPTRSHDGTGLHKGTKTKKEESQMASSWKFMRPFK-PS 485
FGA ++TSF S +S +G + K + K+ ++ ASS K++ K P
Sbjct: 509 TSVFGAKTETSFGL-------SRSLKSGEGASMRK--ISNKDGARGASSRKYIWGLKPPP 559
Query: 486 TIGLSMDLLFRSK 498
+GLSMDLLFRS+
Sbjct: 560 ALGLSMDLLFRSR 572
>UniRef100_Q9LDC1 CRK1 protein [Beta vulgaris subsp. vulgaris]
Length = 599
Score = 684 bits (1764), Expect = 0.0
Identities = 335/433 (77%), Positives = 377/433 (86%), Gaps = 2/433 (0%)
Query: 11 GWPSWLMAVAGEAIGDWTPRRANSFEKLAKIGQGTYSNVYKAKDLVTGKIVALKKVRFDN 70
GWPSWL AVAGEAI W PRRA++FEK+ KIGQGTYSNVYKA+D +TGKIVALKKVRFDN
Sbjct: 117 GWPSWLSAVAGEAIDGWVPRRADTFEKIDKIGQGTYSNVYKARDSLTGKIVALKKVRFDN 176
Query: 71 LEPESVKFMAREILVLRKLDHPNVVKLEGLVTSRMSCSLYLVFEYMEHDLAGLSAGQGVK 130
LEPESVKFMAREIL+LR+LDHPNVVKLEGLVTSRMSCSLYLVFEYMEHDLAGL+A +K
Sbjct: 177 LEPESVKFMAREILILRRLDHPNVVKLEGLVTSRMSCSLYLVFEYMEHDLAGLAASPDIK 236
Query: 131 FTEPQVKCFMKQLLSGLEHCHSRGVLHRDIKGSNLLIDNEGILKIADFGLATFYNPNKKQ 190
FTEPQVKC+M QL+SGLEHCH+RGVLHRDIKGSNLL+DN GILKIADFGLATF++PNKK
Sbjct: 237 FTEPQVKCYMHQLISGLEHCHNRGVLHRDIKGSNLLLDNGGILKIADFGLATFFDPNKKH 296
Query: 191 SMTSRVVTLWYRPPELLLGATFYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFK 250
MTSRVVTLWYR PELLLGAT YGVGIDL SAGCILAELLAG+PIMPGRTEVEQLHKI+K
Sbjct: 297 PMTSRVVTLWYRAPELLLGATDYGVGIDLRSAGCILAELLAGRPIMPGRTEVEQLHKIYK 356
Query: 251 LCGSPAEEYWRKHKLPNATIFKPQQPYKRCISETFKDFPPSSLPLIDSLLAIDPDRRGTA 310
LCGSP++EYW+K KLPNATIFKP++PYKRCI ETF+DFPPS+L LIDSLLAIDP R TA
Sbjct: 357 LCGSPSDEYWKKSKLPNATIFKPREPYKRCIRETFRDFPPSALSLIDSLLAIDPAERKTA 416
Query: 311 SAALNHEFFTTEPYACEPSSLPKYPPSKELDVKMRDEEARRQKALNGKANAVDGAKRVRA 370
+ ALN +FF+TEP AC+PS+LPKYPPSKE+D K RD+EARR +A + KA D K+ R
Sbjct: 417 TDALNSDFFSTEPLACDPSTLPKYPPSKEMDAKRRDDEARRLRAAS-KAQG-DATKKTRT 474
Query: 371 RERGRAIPAPEANAEIQTNLDRWRVVTHANAKSKSEKFPPPHQDGAVGYPQDESSKGPVS 430
R+R RA+PAPEANAE+Q NLDR R++THANAKSKSEKFPPPHQDG +GYP S S
Sbjct: 475 RDRPRAMPAPEANAELQANLDRRRIITHANAKSKSEKFPPPHQDGGLGYPLGASQHIDPS 534
Query: 431 FGASDTSFSSGTF 443
D +SS +F
Sbjct: 535 NIPPDIPYSSTSF 547
>UniRef100_Q7F0M8 Putative CRK1 protein [Oryza sativa]
Length = 573
Score = 677 bits (1746), Expect = 0.0
Identities = 336/465 (72%), Positives = 389/465 (83%), Gaps = 3/465 (0%)
Query: 11 GWPSWLMAVAGEAIGDWTPRRANSFEKLAKIGQGTYSNVYKAKDLVTGKIVALKKVRFDN 70
GWP+WL AVAG+AI WTPRRA+SFEK+ KIGQGTYSNVYKA+D V+GKIVALKKVRFDN
Sbjct: 102 GWPAWLSAVAGDAIDGWTPRRADSFEKIDKIGQGTYSNVYKARDSVSGKIVALKKVRFDN 161
Query: 71 LEPESVKFMAREILVLRKLDHPNVVKLEGLVTSRMSCSLYLVFEYMEHDLAGLSAGQGVK 130
LEPESV+FMAREIL+LR+LDHPNV+KL+GLVTSRMSCSLYLVF+YM HDLAGL+A +K
Sbjct: 162 LEPESVRFMAREILILRRLDHPNVIKLDGLVTSRMSCSLYLVFDYMVHDLAGLAASPEIK 221
Query: 131 FTEPQVKCFMKQLLSGLEHCHSRGVLHRDIKGSNLLIDNEGILKIADFGLATFYNPNKKQ 190
FT PQVKC++ QLLSGLEHCH RGVLHRDIKGSNLL+DN G+LKI DFGLA+F++PN KQ
Sbjct: 222 FTLPQVKCYVHQLLSGLEHCHDRGVLHRDIKGSNLLLDNNGVLKIGDFGLASFFDPNHKQ 281
Query: 191 SMTSRVVTLWYRPPELLLGATFYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFK 250
MTSRVVTLWYRPPELLLGAT YGVG+DLWSAGCILAELLAGKPIMPGRTEVEQLHKIFK
Sbjct: 282 PMTSRVVTLWYRPPELLLGATDYGVGVDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFK 341
Query: 251 LCGSPAEEYWRKHKLPNATIFKPQQPYKRCISETFKDFPPSSLPLIDSLLAIDPDRRGTA 310
LCGSP EEYW+K KLP+ATIFKPQQPYKR I++TFKDFP S+L LI++LLAIDP R TA
Sbjct: 342 LCGSPTEEYWKKSKLPHATIFKPQQPYKRRIADTFKDFPQSALRLIETLLAIDPADRLTA 401
Query: 311 SAALNHEFFTTEPYACEPSSLPKYPPSKELDVKMRDEEARRQKALNGKANAVDGAKRVRA 370
++AL EFF TEP+AC+PSSLP+YPPSKE+D K RDEEA R +A G+ N +GA++ R
Sbjct: 402 TSALESEFFKTEPHACDPSSLPQYPPSKEMDAKRRDEEA-RLRAAGGRVNG-EGARKTRT 459
Query: 371 RERGRAIPAPEANAEIQTNLDRWRVVTHANAKSKSEKFPPPHQDGAVGYPQDESSKGPVS 430
RER RA+PAPEANAE+Q N+D+ R++THANAKSKSEKFPPPHQDGA+GYP S+ +
Sbjct: 460 RERPRAVPAPEANAELQANIDKRRLITHANAKSKSEKFPPPHQDGALGYPLGCSNHMEPA 519
Query: 431 FGASDTSFSSGTFNV-KPSGPTRSHDGTGLHKGTKTKKEESQMAS 474
F D S S F K S PT S G + +K +S +S
Sbjct: 520 FEPPDPSSFSTVFPYEKGSVPTWSGPLADPSSGNQKRKHKSGRSS 564
>UniRef100_Q93W00 Putative CRK1 protein [Oryza sativa]
Length = 558
Score = 676 bits (1743), Expect = 0.0
Identities = 333/432 (77%), Positives = 373/432 (86%), Gaps = 4/432 (0%)
Query: 11 GWPSWLMAVAGEAIGDWTPRRANSFEKLAKIGQGTYSNVYKAKDLVTGKIVALKKVRFDN 70
GWP WL+AVAGEA+ WTPRRA++FEKL KIG GTYSNVY+A+D V+G+IVALKKVRFDN
Sbjct: 75 GWPPWLVAVAGEALRGWTPRRADTFEKLNKIGSGTYSNVYRARDTVSGRIVALKKVRFDN 134
Query: 71 LEPESVKFMAREILVLRKLDHPNVVKLEGLVTSRMSCSLYLVFEYMEHDLAGLSAGQGVK 130
LEPESVKFMAREIL+LRKLDHPNV+KLEGLVTSRMSCSLYLVFEYMEHDLAGL+A VK
Sbjct: 135 LEPESVKFMAREILILRKLDHPNVIKLEGLVTSRMSCSLYLVFEYMEHDLAGLAASPDVK 194
Query: 131 FTEPQVKCFMKQLLSGLEHCHSRGVLHRDIKGSNLLIDNEGILKIADFGLATFYNPNKKQ 190
FT PQ+KC+++QLLSGLEHCH+ VLHRDIKGSNLL+DN GILKIADFGLATF++P K+
Sbjct: 195 FTLPQIKCYVQQLLSGLEHCHNNNVLHRDIKGSNLLLDNNGILKIADFGLATFFDPRHKR 254
Query: 191 SMTSRVVTLWYRPPELLLGATFYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFK 250
MTSRVVTLWYRPPELLLGAT YGVG+DLWSAGCILAELL GKPIMPGRTEVEQLHKIFK
Sbjct: 255 PMTSRVVTLWYRPPELLLGATDYGVGVDLWSAGCILAELLHGKPIMPGRTEVEQLHKIFK 314
Query: 251 LCGSPAEEYWRKHKLPNATIFKPQQPYKRCISETFKDFPPSSLPLIDSLLAIDPDRRGTA 310
LCGSP+EEYW+K KLP+ATIFKPQQPYKRCI E FKDFPPSSLPL+++LLAIDP R TA
Sbjct: 315 LCGSPSEEYWKKSKLPHATIFKPQQPYKRCIREAFKDFPPSSLPLVETLLAIDPAERQTA 374
Query: 311 SAALNHEFFTTEPYACEPSSLPKYPPSKELDVKMRDEEARRQKALNGKANAVDGAKRVRA 370
++AL EFF TEPYAC+PSSLP YPPSKE+D KMRDEEARR +A KA +G KR R
Sbjct: 375 TSALQSEFFATEPYACDPSSLPTYPPSKEMDAKMRDEEARRLRAA-AKAKG-EGVKRTRT 432
Query: 371 RERG-RAIPAPEANAEIQTNLD-RWRVVTHANAKSKSEKFPPPHQDGAVGYPQDESSKGP 428
R+R RA PAPEANAE+Q NLD R R++THANAKSKSEKFPPPHQDGA+G P S
Sbjct: 433 RDRSQRAGPAPEANAELQANLDQRRRMITHANAKSKSEKFPPPHQDGAMGNPLGSSRHME 492
Query: 429 VSFGASDTSFSS 440
+ D SFS+
Sbjct: 493 PMYEHQDASFST 504
>UniRef100_Q9ZVM9 Hypothetical protein T22H22.5 [Arabidopsis thaliana]
Length = 572
Score = 660 bits (1702), Expect = 0.0
Identities = 334/476 (70%), Positives = 392/476 (82%), Gaps = 11/476 (2%)
Query: 11 GWPSWLMAVAGEAIGDWTPRRANSFEKLAKIGQGTYSNVYKAKDLVTGKIVALKKVRFDN 70
GWPSWL GEA+ W PR+A++FEK+ KIGQGTYSNVYKAKD++TGKIVALKKVRFDN
Sbjct: 94 GWPSWLSDACGEALNGWVPRKADTFEKIDKIGQGTYSNVYKAKDMLTGKIVALKKVRFDN 153
Query: 71 LEPESVKFMAREILVLRKLDHPNVVKLEGLVTSRMSCSLYLVFEYMEHDLAGLSAGQGVK 130
LEPESVKFMAREILVLR+LDHPNVVKLEGLVTSRMSCSLYLVF+YM+HDLAGL++ VK
Sbjct: 154 LEPESVKFMAREILVLRRLDHPNVVKLEGLVTSRMSCSLYLVFQYMDHDLAGLASSPVVK 213
Query: 131 FTEPQVKCFMKQLLSGLEHCHSRGVLHRDIKGSNLLIDNEGILKIADFGLATFYNPNKKQ 190
F+E +VKC M+QL+SGLEHCHSRGVLHRDIKGSNLLID+ G+LKIADFGLAT ++PN K+
Sbjct: 214 FSESEVKCLMRQLISGLEHCHSRGVLHRDIKGSNLLIDDGGVLKIADFGLATIFDPNHKR 273
Query: 191 SMTSRVVTLWYRPPELLLGATFYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFK 250
MTSRVVTLWYR PELLLGAT YGVGIDLWSAGCILAELLAG+PIMPGRTEVEQLHKI+K
Sbjct: 274 PMTSRVVTLWYRAPELLLGATDYGVGIDLWSAGCILAELLAGRPIMPGRTEVEQLHKIYK 333
Query: 251 LCGSPAEEYWRKHKLPNATIFKPQQPYKRCISETFKDFPPSSLPLIDSLLAIDPDRRGTA 310
LCGSP+E+YW+K K + I+KP++PYKR I ETFKDFPPSSLPLID+LL+I+P+ R TA
Sbjct: 334 LCGSPSEDYWKKGKFTHGAIYKPREPYKRSIRETFKDFPPSSLPLIDALLSIEPEDRQTA 393
Query: 311 SAALNHEFFTTEPYACEPSSLPKYPPSKELDVKMRDEEARRQKALNGKANAVDGAKRVRA 370
SAAL EFFT+EPYACEP+ LPKYPPSKE+D K RDEE RRQ+A + KA DGA++ R
Sbjct: 394 SAALKSEFFTSEPYACEPADLPKYPPSKEIDAKRRDEETRRQRAAS-KAQG-DGARKNRH 451
Query: 371 RER-GRAIPAPEANAEIQTNLDRWRVVTHANAKSKSEKFPPPHQD-GAVGYPQDESSKGP 428
R+R RA+PAPEANAE+Q+N+DR R++THANAKSKSEKFPPPHQD GA+G P S
Sbjct: 452 RDRSNRALPAPEANAELQSNVDRRRLITHANAKSKSEKFPPPHQDGGAMGVPLGASQHID 511
Query: 429 VSFGASD--TSFSSGTFNV-KPSGPTRSHDGTG----LHKGTKTKKEESQMASSWK 477
+F D SF+S +FN K PT+ +G G KK+++ +S K
Sbjct: 512 PTFIPRDMVPSFTSSSFNFSKDEPPTQVQTWSGPLGHPITGVSRKKKDNTKSSKGK 567
>UniRef100_Q6ZCC7 Putative CRK1 protein [Oryza sativa]
Length = 748
Score = 592 bits (1525), Expect = e-167
Identities = 286/412 (69%), Positives = 335/412 (80%), Gaps = 14/412 (3%)
Query: 11 GWPSWLMAVAGEAIGDWTPRRANSFEKLAKIGQGTYSNVYKAKDLVTGKIVALKKVRFDN 70
GWP WL VA EA+ W PR+A SFEKL KIGQGTYS+VYKA+DL +GKIVALKKVRF N
Sbjct: 159 GWPRWLTEVAAEAVRGWQPRKAESFEKLDKIGQGTYSSVYKARDLESGKIVALKKVRFAN 218
Query: 71 LEPESVKFMAREILVLRKLDHPNVVKLEGLVTSRMSCSLYLVFEYMEHDLAGLSAGQGVK 130
++PESV+FMAREI +LR+LDHPNV+KLEGLVTSRMS SLYLVFEYMEHDLAGL+A G+K
Sbjct: 219 MDPESVRFMAREIHILRRLDHPNVIKLEGLVTSRMSSSLYLVFEYMEHDLAGLAATPGIK 278
Query: 131 FTEPQVKCFMKQLLSGLEHCHSRGVLHRDIKGSNLLIDNEGILKIADFGLATFYNPNKKQ 190
FTEPQVKC+M+QLLSGLEHCH+RGVLHRDIKG+NLLIDN G+LKIADFGLATF+NPN+KQ
Sbjct: 279 FTEPQVKCYMQQLLSGLEHCHNRGVLHRDIKGANLLIDNNGVLKIADFGLATFFNPNQKQ 338
Query: 191 SMTSRVVTLWYRPPELLLGATFYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFK 250
+TSRVVTLWYRPPELLLGAT YG +DLWSAGCILAELL+GKPIMPGRTEVEQLHKIFK
Sbjct: 339 HLTSRVVTLWYRPPELLLGATNYGAAVDLWSAGCILAELLSGKPIMPGRTEVEQLHKIFK 398
Query: 251 LCGSPAEEYWRKHKLPNATIFKPQQPYKRCISETFKDFPPSSLPLIDSLLAIDPDRRGTA 310
LCGSP+E++W KL ATIFKPQ PY+RC+++ +KDFPP +L L+D LLA++P RGTA
Sbjct: 399 LCGSPSEDFWANLKLSRATIFKPQHPYRRCVNDVYKDFPPPALALLDCLLAVEPQNRGTA 458
Query: 311 SAALNHEFFTTEPYACEPSSLPKYPPSKELDVKMRDEEARRQKALNGKANAVDGAKRVRA 370
++AL EFFTT+PYAC+PSSLPKYPPSKE D K+RDEEARRQ+A K + + +R
Sbjct: 459 ASALGSEFFTTKPYACDPSSLPKYPPSKEYDAKLRDEEARRQRAAAVKGHESEAGRR--- 515
Query: 371 RERGRAIPAPEANAEIQTNLDRWRVVTHANAKSKSEKFPPPHQDGAVGYPQD 422
+ +PAP N E+Q N KS S KF P +D G+P D
Sbjct: 516 ----KQLPAPNGNNELQQRR------VQLNPKSSSNKF-IPKEDAVTGFPID 556
>UniRef100_Q9MAB4 Putative cyclin-dependent protein kinase [Arabidopsis thaliana]
Length = 593
Score = 590 bits (1521), Expect = e-167
Identities = 306/481 (63%), Positives = 363/481 (74%), Gaps = 15/481 (3%)
Query: 11 GWPSWLMAVAGEAIGDWTPRRANSFEKLAKIGQGTYSNVYKAKDLVTGKIVALKKVRFDN 70
GWPSWL V GEA+ W PR+A+SFEK+ KIG GTYSNVYKAKD +TG IVALKKVR D
Sbjct: 114 GWPSWLSEVCGEALSGWLPRKADSFEKIDKIGSGTYSNVYKAKDSLTGNIVALKKVRCDV 173
Query: 71 LEPESVKFMAREILVLRKLDHPNVVKLEGLVTSRMSCSLYLVFEYMEHDLAGLSAGQGVK 130
E ES+KFMAREIL+LR+LDHPNV+KLEGLVTSRMS SLYLVF YM+HDLAGL+A +K
Sbjct: 174 NERESLKFMAREILILRRLDHPNVIKLEGLVTSRMSSSLYLVFRYMDHDLAGLAASPEIK 233
Query: 131 FTEPQVKCFMKQLLSGLEHCHSRGVLHRDIKGSNLLIDNEGILKIADFGLATFYNPNKKQ 190
FTE QVKC+MKQLLSGLEHCH+RGVLHRDIKGSNLLID+ G+L+I DFGLATF++ +K+Q
Sbjct: 234 FTEQQVKCYMKQLLSGLEHCHNRGVLHRDIKGSNLLIDDGGVLRIGDFGLATFFDASKRQ 293
Query: 191 SMTSRVVTLWYRPPELLLGATFYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFK 250
MT+RVVTLWYR PELL G Y VG+DLWSAGCILAELLAG+ IMPGR EVEQLH+I+K
Sbjct: 294 EMTNRVVTLWYRSPELLHGVVEYSVGVDLWSAGCILAELLAGRAIMPGRNEVEQLHRIYK 353
Query: 251 LCGSPAEEYWRKHKLPNA---TIFKPQQPYKRCISETFKDFPPSSLPLIDSLLAIDPDRR 307
LCGSP+EEYW+K +LP+ KP YKR I E +KDF P +L L+D+LLA+DP R
Sbjct: 354 LCGSPSEEYWKKIRLPSTHKHAHHKPLPQYKRRIREVYKDFSPEALSLLDTLLALDPAER 413
Query: 308 GTASAALNHEFFTTEPYACEPSSLPKYPPSKELDVKMRDEEARRQKALNGKANAVDGAKR 367
TA+ L +FFTTEP AC+PS LPKYPPSKE+D K RDEE RRQ+ KA G +R
Sbjct: 414 QTATDVLMSDFFTTEPLACQPSDLPKYPPSKEIDAKRRDEEYRRQREAR-KAQGESG-RR 471
Query: 368 VRARERG-RAIPAPEANAEIQTNLDRWRVVTHANAKSKSEKFPPPHQDGAVGYPQDES-- 424
+R RER RA+PAPEANAE Q+N+DR R++THANAKSKSEKFPPPHQDG++G+ S
Sbjct: 472 MRPRERAPRAMPAPEANAENQSNIDRMRMITHANAKSKSEKFPPPHQDGSLGFQVGSSRR 531
Query: 425 ---SKGPVSFGASDTSFSSGTFNV--KPSGPTRSHDGTGLHKGTKTKKEESQMASSWKFM 479
S+ P S + +S+S F P P + D T + K E +MAS K
Sbjct: 532 LDPSEIPYSTNSFTSSYSKEPFQTWSGPLAPIGAPDSTTRRRNDINK--ERRMASKVKGK 589
Query: 480 R 480
R
Sbjct: 590 R 590
>UniRef100_Q9LNN0 F8L10.9 protein [Arabidopsis thaliana]
Length = 694
Score = 587 bits (1513), Expect = e-166
Identities = 284/412 (68%), Positives = 340/412 (81%), Gaps = 6/412 (1%)
Query: 11 GWPSWLMAVAGEAIGDWTPRRANSFEKLAKIGQGTYSNVYKAKDLVTGKIVALKKVRFDN 70
GWP WL +VAGEAI W PRRA+SFEKL KIGQGTYSNVY+A+DL KIVALKKVRFDN
Sbjct: 110 GWPPWLASVAGEAIRGWVPRRADSFEKLDKIGQGTYSNVYRARDLDQKKIVALKKVRFDN 169
Query: 71 LEPESVKFMAREILVLRKLDHPNVVKLEGLVTSRMSCSLYLVFEYMEHDLAGLSAGQGVK 130
LEPESV+FMAREI +LR+LDHPN++KLEGLVTSRMSCSLYLVFEYMEHDLAGL++ +K
Sbjct: 170 LEPESVRFMAREIQILRRLDHPNIIKLEGLVTSRMSCSLYLVFEYMEHDLAGLASHPAIK 229
Query: 131 FTEPQVKCFMKQLLSGLEHCHSRGVLHRDIKGSNLLIDNEGILKIADFGLATFYNPNKKQ 190
F+E QVKC+++QLL GL+HCHSRGVLHRDIKGSNLLIDN G+LKIADFGLA+F++P + Q
Sbjct: 230 FSESQVKCYLQQLLHGLDHCHSRGVLHRDIKGSNLLIDNSGVLKIADFGLASFFDPRQTQ 289
Query: 191 SMTSRVVTLWYRPPELLLGATFYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFK 250
+TSRVVTLWYRPPELLLGAT YG +DLWSAGCILAEL AGKPIMPGRTEVEQLHKIFK
Sbjct: 290 PLTSRVVTLWYRPPELLLGATRYGAAVDLWSAGCILAELYAGKPIMPGRTEVEQLHKIFK 349
Query: 251 LCGSPAEEYWRKHKLPNATIFKPQQPYKRCISETFKDFPPSSLPLIDSLLAIDPDRRGTA 310
LCGSP E+YW K +LP+ATIFKP QPYKR + ETFK+FP +L L+++LL+++PD RGTA
Sbjct: 350 LCGSPTEDYWVKSRLPHATIFKPTQPYKRLVGETFKEFPQPALALLETLLSVNPDDRGTA 409
Query: 311 SAALNHEFFTTEPYACEPSSLPKYPPSKELDVKMRDEEARRQKALNGKANAVDGAKRVRA 370
+AAL EFF+T P C+PSSLPKYPPSKELD +MRDEE+RRQ N + R
Sbjct: 410 TAALKSEFFSTRPLPCDPSSLPKYPPSKELDARMRDEESRRQVG----GNRDQRHQERRG 465
Query: 371 RERGRAIPAPEANAEIQTNLDRWRVVTHANAKSKSEKFPPPHQDGAVGYPQD 422
+ RAIPAP+ANAE+ ++ + + + + +S+SEKF P ++ A G+P D
Sbjct: 466 TKESRAIPAPDANAELVASMQKRQ--SQSTNRSRSEKFNPHPEEVASGFPID 515
>UniRef100_Q84W73 Putative cell division-related protein [Arabidopsis thaliana]
Length = 694
Score = 585 bits (1508), Expect = e-165
Identities = 283/412 (68%), Positives = 340/412 (81%), Gaps = 6/412 (1%)
Query: 11 GWPSWLMAVAGEAIGDWTPRRANSFEKLAKIGQGTYSNVYKAKDLVTGKIVALKKVRFDN 70
GWP WL +VAGEAI W PRRA+SFEKL KIGQGT+SNVY+A+DL KIVALKKVRFDN
Sbjct: 110 GWPPWLASVAGEAIRGWVPRRADSFEKLDKIGQGTHSNVYRARDLDQKKIVALKKVRFDN 169
Query: 71 LEPESVKFMAREILVLRKLDHPNVVKLEGLVTSRMSCSLYLVFEYMEHDLAGLSAGQGVK 130
LEPESV+FMAREI +LR+LDHPN++KLEGLVTSRMSCSLYLVFEYMEHDLAGL++ +K
Sbjct: 170 LEPESVRFMAREIQILRRLDHPNIIKLEGLVTSRMSCSLYLVFEYMEHDLAGLASHPAIK 229
Query: 131 FTEPQVKCFMKQLLSGLEHCHSRGVLHRDIKGSNLLIDNEGILKIADFGLATFYNPNKKQ 190
F+E QVKC+++QLL GL+HCHSRGVLHRDIKGSNLLIDN G+LKIADFGLA+F++P + Q
Sbjct: 230 FSESQVKCYLQQLLHGLDHCHSRGVLHRDIKGSNLLIDNSGVLKIADFGLASFFDPRQTQ 289
Query: 191 SMTSRVVTLWYRPPELLLGATFYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFK 250
+TSRVVTLWYRPPELLLGAT YG +DLWSAGCILAEL AGKPIMPGRTEVEQLHKIFK
Sbjct: 290 PLTSRVVTLWYRPPELLLGATRYGAAVDLWSAGCILAELYAGKPIMPGRTEVEQLHKIFK 349
Query: 251 LCGSPAEEYWRKHKLPNATIFKPQQPYKRCISETFKDFPPSSLPLIDSLLAIDPDRRGTA 310
LCGSP E+YW K +LP+ATIFKP QPYKR + ETFK+FP +L L+++LL+++PD RGTA
Sbjct: 350 LCGSPTEDYWVKSRLPHATIFKPTQPYKRLVGETFKEFPQPALALLETLLSVNPDDRGTA 409
Query: 311 SAALNHEFFTTEPYACEPSSLPKYPPSKELDVKMRDEEARRQKALNGKANAVDGAKRVRA 370
+AAL EFF+T P C+PSSLPKYPPSKELD +MRDEE+RRQ N + R
Sbjct: 410 TAALKSEFFSTRPLPCDPSSLPKYPPSKELDARMRDEESRRQVG----GNRDQRHQERRG 465
Query: 371 RERGRAIPAPEANAEIQTNLDRWRVVTHANAKSKSEKFPPPHQDGAVGYPQD 422
+ RAIPAP+ANAE+ ++ + + + + +S+SEKF P ++ A G+P D
Sbjct: 466 TKESRAIPAPDANAELVASMQKRQ--SQSTNRSRSEKFNPHPEEVASGFPID 515
>UniRef100_O80540 F14J9.26 protein [Arabidopsis thaliana]
Length = 967
Score = 556 bits (1433), Expect = e-157
Identities = 271/400 (67%), Positives = 333/400 (82%), Gaps = 7/400 (1%)
Query: 11 GWPSWLMAVAGEAIGDWTPRRANSFEKLAKIGQGTYSNVYKAKDLVTGKIVALKKVRFDN 70
GWPSWL +VAGEAI W PR+A+SFEKL KIGQGTYS+VYKA+DL T ++VALKKVRF N
Sbjct: 139 GWPSWLASVAGEAINGWIPRKADSFEKLEKIGQGTYSSVYKARDLETNQLVALKKVRFAN 198
Query: 71 LEPESVKFMAREILVLRKLDHPNVVKLEGLVTSRMSCSLYLVFEYMEHDLAGLSAGQGVK 130
++P+SV+FMAREI++LR+LDHPNV+KLEGL+TSR+S S+YL+FEYMEHDLAGL++ G+
Sbjct: 199 MDPDSVRFMAREIIILRRLDHPNVMKLEGLITSRVSGSMYLIFEYMEHDLAGLASTPGIN 258
Query: 131 FTEPQVKCFMKQLLSGLEHCHSRGVLHRDIKGSNLLIDNEGILKIADFGLATFYNPNKKQ 190
F+E Q+KC+MKQLL GLEHCHSRGVLHRDIKGSNLL+D+ LKI DFGLA FY ++KQ
Sbjct: 259 FSEAQIKCYMKQLLHGLEHCHSRGVLHRDIKGSNLLLDHNNNLKIGDFGLANFYQGHQKQ 318
Query: 191 SMTSRVVTLWYRPPELLLGATFYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFK 250
+TSRVVTLWYRPPELLLG+T YGV +DLWS GCILAEL GKPIMPGRTEVEQLHKIFK
Sbjct: 319 PLTSRVVTLWYRPPELLLGSTDYGVTVDLWSTGCILAELFTGKPIMPGRTEVEQLHKIFK 378
Query: 251 LCGSPAEEYWRKHKLPNATIFKPQQPYKRCISETFKDFPPSSLPLIDSLLAIDPDRRGTA 310
LCGSP+EEYW+ KLP+ATIFKPQQPYKRC++ETFK P S+L L++ LLA++PD RGT
Sbjct: 379 LCGSPSEEYWKISKLPHATIFKPQQPYKRCVAETFKSLPSSALALVEVLLAVEPDARGTT 438
Query: 311 SAALNHEFFTTEPYACEPSSLPKYPPSKELDVKMRDEEARRQKALNGKANAVDGAKRVRA 370
++AL EFFTT P A +PSSLPKY P KE+DVK ++EEA+R+K + K N +K+V +
Sbjct: 439 ASALESEFFTTSPLASDPSSLPKYQPRKEIDVKAQEEEAKRKKDTSSKQN---DSKQV-S 494
Query: 371 RERGRAIPAPEANAEIQTNLDRWRVVTHANAKSKSEKFPP 410
RE +A+PAP++NAE T++ + R H N S S+KF P
Sbjct: 495 RE-SKAVPAPDSNAESLTSIQK-RQGQH-NQVSNSDKFNP 531
>UniRef100_Q9LR53 F21B7.34 [Arabidopsis thaliana]
Length = 740
Score = 540 bits (1392), Expect = e-152
Identities = 264/419 (63%), Positives = 330/419 (78%), Gaps = 14/419 (3%)
Query: 11 GWPSWLMAVAGEAIGDWTPRRANSFEKLAKIGQGTYSNVYKAKDLVTGKIVALKKVRFDN 70
GWPSWL++VAGE++ DW PRRAN+FEKL KIGQGTYS+VY+A+DL+ KIVALKKVRFD
Sbjct: 189 GWPSWLVSVAGESLVDWAPRRANTFEKLEKIGQGTYSSVYRARDLLHNKIVALKKVRFDL 248
Query: 71 LEPESVKFMAREILVLRKLDHPNVVKLEGLVTSRMSCSLYLVFEYMEHDLAGLSAGQGVK 130
+ ESVKFMAREI+V+R+LDHPNV+KLEGL+T+ +S SLYLVFEYM+HDL GLS+ GVK
Sbjct: 249 NDMESVKFMAREIIVMRRLDHPNVLKLEGLITAPVSSSLYLVFEYMDHDLLGLSSLPGVK 308
Query: 131 FTEPQVKCFMKQLLSGLEHCHSRGVLHRDIKGSNLLIDNEGILKIADFGLATFYNPNKKQ 190
FTEPQVKC+M+QLLSGLEHCHSRGVLHRDIKGSNLLID++G+LKIADFGLATF++P K
Sbjct: 309 FTEPQVKCYMRQLLSGLEHCHSRGVLHRDIKGSNLLIDSKGVLKIADFGLATFFDPAKSV 368
Query: 191 SMTSRVVTLWYRPPELLLGATFYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFK 250
S+TS VVTLWYRPPELLLGA+ YGVG+DLWS GCIL EL AGKPI+PG+TEVEQLHKIFK
Sbjct: 369 SLTSHVVTLWYRPPELLLGASHYGVGVDLWSTGCILGELYAGKPILPGKTEVEQLHKIFK 428
Query: 251 LCGSPAEEYWRKHKLPNATIFKPQQPYKRCISETFKDFPPSSLPLIDSLLAIDPDRRGTA 310
LCGSP E YWRK KLP++ FK PY+R +SE FKDFP S L L+++LL+IDPD R +A
Sbjct: 429 LCGSPTENYWRKQKLPSSAGFKTAIPYRRKVSEMFKDFPASVLSLLETLLSIDPDHRSSA 488
Query: 311 SAALNHEFFTTEPYACEPSSLPKYPPSKELDVKMRDEEARRQKALNGKANAVDGAKRVRA 370
AL E+F T+P+AC+PS+LPKYPPSKE+D KMRDE R+Q K D R+ +
Sbjct: 489 DRALESEYFKTKPFACDPSNLPKYPPSKEIDAKMRDEAKRQQPMRAEKQERQDSMTRI-S 547
Query: 371 RERGRAIPAPEANAEIQTNLDRWRVVTHANAKSKSEKFPPPHQDGAVGYPQDESSKGPV 429
ER + +P +AN + +++ + + +S+++ F + ++ + GPV
Sbjct: 548 HER-KFVPPVKANNSLSMTMEK----QYKDLRSRNDSFK--------SFKEERTPHGPV 593
>UniRef100_Q9T0F9 Hypothetical protein T5L19.140 [Arabidopsis thaliana]
Length = 649
Score = 538 bits (1386), Expect = e-151
Identities = 267/417 (64%), Positives = 319/417 (76%), Gaps = 13/417 (3%)
Query: 11 GWPSWLMAVAGEAIGDWTPRRANSFEKLAKIGQGTYSNVYKAKDLVTGKIVALKKVRFDN 70
GWPSWL +VAGEAI W PRRA SFEKL KIGQGTYS+VY+A+DL TGK+VA+KKVRF N
Sbjct: 132 GWPSWLTSVAGEAIKGWVPRRAESFEKLDKIGQGTYSSVYRARDLETGKMVAMKKVRFVN 191
Query: 71 LEPESVKFMAREILVLRKLDHPNVVKLEGLVTSRMSCSLYLVFEYMEHDLAGLSAGQGVK 130
++PESV+FMAREI +LRKLDHPNV+KLE LVTS++S SLYLVFEYMEHDL+GL+ GVK
Sbjct: 192 MDPESVRFMAREINILRKLDHPNVMKLECLVTSKLSGSLYLVFEYMEHDLSGLALRPGVK 251
Query: 131 FTEPQVKCFMKQLLSGLEHCHSRGVLHRDIKGSNLLIDNEGILKIADFGLATFYNPNKKQ 190
FTE Q+KC+MKQLLSGLEHCHSRG+LHRDIKG NLL++N+G+LKI DFGLA Y+P + Q
Sbjct: 252 FTESQIKCYMKQLLSGLEHCHSRGILHRDIKGPNLLVNNDGVLKIGDFGLANIYHPEQDQ 311
Query: 191 SMTSRVVTLWYRPPELLLGATFYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFK 250
+TSRVVTLWYR PELLLGAT YG GIDLWS GCIL EL GKPIMPGRTEVEQ+HKIFK
Sbjct: 312 PLTSRVVTLWYRAPELLLGATEYGPGIDLWSVGCILTELFLGKPIMPGRTEVEQMHKIFK 371
Query: 251 LCGSPAEEYWRKHKLPNATIFKPQQPYKRCISETFKDFPPSSLPLIDSLLAIDPDRRGTA 310
CGSP+++YW+K KLP AT FKPQQPYKR + ETFK+ PPS+L L+D LL+++P +RGTA
Sbjct: 372 FCGSPSDDYWQKTKLPLATSFKPQQPYKRVLLETFKNLPPSALALVDKLLSLEPAKRGTA 431
Query: 311 SAALNHEFFTTEPYACEPSSLPKYPPSKELDVKMRDEEARRQKALNGKANAVDGAKRVRA 370
S+ L+ +FFT EP C SSLPKYPPSKELD K+RDEEARR+K+ K + +R +
Sbjct: 432 SSTLSSKFFTMEPLPCNVSSLPKYPPSKELDAKVRDEEARRKKSETVKGRGPESVRR-GS 490
Query: 371 RERGRAIPAPEANAEIQTNLDRWRVVTHANAKSKSEKFPPPHQDGAVGYPQDESSKG 427
R+ PE A Q+ + + K P +D G D KG
Sbjct: 491 RDFKSTATTPEFVASGQSK------------DTITTKRFNPQEDSRTGLRGDRDQKG 535
>UniRef100_Q9C9I9 Hypothetical protein F26A9.10 [Arabidopsis thaliana]
Length = 655
Score = 528 bits (1361), Expect = e-148
Identities = 267/426 (62%), Positives = 322/426 (74%), Gaps = 13/426 (3%)
Query: 12 WPSWLMAVAGEAIGDWTPRRANSFEKLAKIGQGTYSNVYKAKDLVTGKIVALKKVRFDNL 71
WPSWL +VAGEAI W PR A SFEKL KIGQGTYS+VYKA+DL TGKIVA+KKVRF N+
Sbjct: 124 WPSWLASVAGEAIKGWVPRCAESFEKLDKIGQGTYSSVYKARDLETGKIVAMKKVRFVNM 183
Query: 72 EPESVKFMAREILVLRKLDHPNVVKLEGLVTSRMSCSLYLVFEYMEHDLAGLSAGQGVKF 131
+PESV+FMAREIL+LRKLDHPNV+KLEGLVTSR+S SLYLVFEYMEHDLAGL+A G+KF
Sbjct: 184 DPESVRFMAREILILRKLDHPNVMKLEGLVTSRLSGSLYLVFEYMEHDLAGLAATPGIKF 243
Query: 132 TEPQVKCFMKQLLSGLEHCHSRGVLHRDIKGSNLLIDNEGILKIADFGLATFYNPNKKQS 191
+EPQ+KC+M+QL GLEHCH RG+LHRDIKGSNLLI+NEG+LKI DFGLA FY +
Sbjct: 244 SEPQIKCYMQQLFRGLEHCHRRGILHRDIKGSNLLINNEGVLKIGDFGLANFYRGDGDLQ 303
Query: 192 MTSRVVTLWYRPPELLLGATFYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFKL 251
+TSRVVTLWYR PELLLGAT YG IDLWSAGCIL EL AGKPIMPGRTEVEQ+HKIFKL
Sbjct: 304 LTSRVVTLWYRAPELLLGATEYGPAIDLWSAGCILTELFAGKPIMPGRTEVEQMHKIFKL 363
Query: 252 CGSPAEEYWRKHKLPNATIFKPQQPYKRCISETFKDFPPSSLPLIDSLLAIDPDRRGTAS 311
CGSP+E+YWR+ LP AT FKP PYK ++ETF FP S+L LI+ LLAI+P++RG+A+
Sbjct: 364 CGSPSEDYWRRATLPLATSFKPSHPYKPVLAETFNHFPSSALMLINKLLAIEPEKRGSAA 423
Query: 312 AALNHEFFTTEPYACEPSSLPKYPPSKELDVKMRDEEARRQKALNGKANAVDGAKRVRAR 371
+ L EFFTTEP PS+LP+YPPSKELD K+R+EEAR+ +A K + R R +
Sbjct: 424 STLRSEFFTTEPLPANPSNLPRYPPSKELDAKLRNEEARKLRAEGNKRRGGETVTRGRPK 483
Query: 372 ERGRAIPAPEANAEIQTNLDRWRVVTHANAKSKSEKFPPPHQDGAVGYPQDESSKGPVSF 431
+ + PE A Q+ VT + K K++ ++G G+ + +G
Sbjct: 484 DL-KTAQTPEFMAAGQSK------VTCISHKFKTD------EEGGTGFRIEPPRRGIQQN 530
Query: 432 GASDTS 437
G + S
Sbjct: 531 GKAHAS 536
>UniRef100_Q9FKV9 Cyclin-dependent protein kinase-like protein [Arabidopsis thaliana]
Length = 644
Score = 525 bits (1352), Expect = e-147
Identities = 253/373 (67%), Positives = 310/373 (82%), Gaps = 3/373 (0%)
Query: 11 GWPSWLMAVAGEAIGDWTPRRANSFEKLAKIGQGTYSNVYKAKDLVTGKIVALKKVRFDN 70
GWP+WL++VAGEA+ +WTPRRA++FEKL KIGQGTYS+VYKA+DL KIVALK+VRFD
Sbjct: 113 GWPAWLVSVAGEALVNWTPRRASTFEKLEKIGQGTYSSVYKARDLTNNKIVALKRVRFDL 172
Query: 71 LEPESVKFMAREILVLRKLDHPNVVKLEGLVTSRMSCSLYLVFEYMEHDLAGLSAGQGVK 130
+ ESVKFMAREI+V+R+LDHPNV+KLEGL+T+ +S SLYLVFEYM+HDL GL++ G+K
Sbjct: 173 SDLESVKFMAREIIVMRRLDHPNVLKLEGLITASVSSSLYLVFEYMDHDLVGLASIPGIK 232
Query: 131 FTEPQVKCFMKQLLSGLEHCHSRGVLHRDIKGSNLLIDNEGILKIADFGLATFYNPNKKQ 190
F+EPQVKC+M+QLLSGL HCHSRGVLHRDIKGSNLLID+ G+LKIADFGLATF++P
Sbjct: 233 FSEPQVKCYMQQLLSGLHHCHSRGVLHRDIKGSNLLIDSNGVLKIADFGLATFFDPQNCV 292
Query: 191 SMTSRVVTLWYRPPELLLGATFYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFK 250
+TSRVVTLWYRPPELLLGA YGVG+DLWS GCIL EL +GKPI+ G+TEVEQLHKIFK
Sbjct: 293 PLTSRVVTLWYRPPELLLGACHYGVGVDLWSTGCILGELYSGKPILAGKTEVEQLHKIFK 352
Query: 251 LCGSPAEEYWRKHKLPNATIFKPQQPYKRCISETFKDFPPSSLPLIDSLLAIDPDRRGTA 310
LCGSP E+YWRK KLP + F+P PY R ++E FKD P + L L+++LL+IDPDRRG+A
Sbjct: 353 LCGSPTEDYWRKLKLPPSAAFRPALPYGRRVAEMFKDLPTNVLSLLEALLSIDPDRRGSA 412
Query: 311 SAALNHEFFTTEPYACEPSSLPKYPPSKELDVKMRDEEARRQKALNGKANAVDGAKRVRA 370
+ AL E+F TEP+AC+PSSLPKYPPSKE+D K+RD +A+RQ+ K D R R+
Sbjct: 413 ARALESEYFRTEPFACDPSSLPKYPPSKEIDAKIRD-DAKRQRPTQEKHERQDSQTR-RS 470
Query: 371 RERGRAIPAPEAN 383
ER + IP +AN
Sbjct: 471 HER-KLIPPVKAN 482
>UniRef100_Q7XIB4 Putative cyclin-dependent kinase CDC2C [Oryza sativa]
Length = 707
Score = 525 bits (1352), Expect = e-147
Identities = 242/347 (69%), Positives = 299/347 (85%), Gaps = 1/347 (0%)
Query: 11 GWPSWLMAVAGEAIGDWTPRRANSFEKLAKIGQGTYSNVYKAKDLVTGKIVALKKVRFDN 70
GWP WL AVAGEAI W P +A+SFEKL K+GQGTYS+V++A++L TGKIVALKKVRFDN
Sbjct: 105 GWPPWLSAVAGEAIQGWIPLKADSFEKLEKVGQGTYSSVFRARELDTGKIVALKKVRFDN 164
Query: 71 LEPESVKFMAREILVLRKLDHPNVVKLEGLVTSRMSCSLYLVFEYMEHDLAGLSAGQGVK 130
EPESV+FMAREI +LR+LDHPNV+KLEGL+TSR+SCSLYLVFEYMEHDLAGLS+ +K
Sbjct: 165 FEPESVRFMAREIQILRRLDHPNVMKLEGLITSRLSCSLYLVFEYMEHDLAGLSSSPDIK 224
Query: 131 FTEPQVKCFMKQLLSGLEHCHSRGVLHRDIKGSNLLIDNEGILKIADFGLATFYNPNKKQ 190
F+E QVKC+M QLLSGLEHCHSR ++HRDIKG+NLL++NEG+LKIADFGLA +++PNK
Sbjct: 225 FSEAQVKCYMNQLLSGLEHCHSRRIVHRDIKGANLLVNNEGVLKIADFGLANYFDPNKNH 284
Query: 191 SMTSRVVTLWYRPPELLLGATFYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFK 250
+TSRVVTLWYRPPELLLG+T Y +DLWSAGC+ AE+ GKPI+ GRTEVEQLHKIFK
Sbjct: 285 PLTSRVVTLWYRPPELLLGSTHYDAAVDLWSAGCVFAEMFRGKPILQGRTEVEQLHKIFK 344
Query: 251 LCGSPAEEYWRKHKLPNATIFKPQQPYKRCISETFKDFPPSSLPLIDSLLAIDPDRRGTA 310
LCGSPA+EYW+K KLP+ATIFKP PY+ + + FK+ P ++L L+++LL+++P +RGTA
Sbjct: 345 LCGSPADEYWKKSKLPHATIFKPHCPYQSTLQDVFKEMPANALRLLETLLSVEPYKRGTA 404
Query: 311 SAALNHEFFTTEPYACEPSSLPKYPPSKELDVKMRDEEARRQKALNG 357
SAAL EFF T+PYAC+PSSLPKY P+KE+D K+R E++ R+KA G
Sbjct: 405 SAALTSEFFKTKPYACDPSSLPKYAPNKEMDAKLR-EDSHRRKASRG 450
>UniRef100_Q6YU34 Putative CRK1 protein [Oryza sativa]
Length = 729
Score = 525 bits (1352), Expect = e-147
Identities = 257/379 (67%), Positives = 312/379 (81%), Gaps = 11/379 (2%)
Query: 11 GWPSWLMAVAGEAIGDWTPRRANSFEKLAKIGQGTYSNVYKAKDLVTGKIVALKKVRFDN 70
GWPSWL +VAGE + W PRRA++FE+L KIGQGTYSNVYKA+DL TGK+VALK+VRF N
Sbjct: 134 GWPSWLTSVAGEVVQGWLPRRADTFERLDKIGQGTYSNVYKARDLETGKVVALKRVRFVN 193
Query: 71 LEPESVKFMAREILVLRKLD-HPNVVKLEGLVTSRMSCSLYLVFEYMEHDLAGLSAGQGV 129
++PESV+FMAREI VLR+LD HPNVV+LEG+VTSR+S SLYLVFEYM+HDLAGL+A G+
Sbjct: 194 MDPESVRFMAREIHVLRRLDGHPNVVRLEGIVTSRLSHSLYLVFEYMDHDLAGLAATPGL 253
Query: 130 KFTEPQVKCFMKQLLSGLEHCHSRGVLHRDIKGSNLLIDNEGILKIADFGLATFYNPNKK 189
+FTEPQVKC M Q+L+GL HCH RGVLHRDIKG+NLLI +G LKIADFGLATF++ +
Sbjct: 254 RFTEPQVKCLMAQILAGLRHCHDRGVLHRDIKGANLLIGGDGALKIADFGLATFFDAARP 313
Query: 190 QSMTSRVVTLWYRPPELLLGATFYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIF 249
Q +TSRVVTLWYRPPELLLGAT YGV +DLWS GCILAELLAGKPI+PG+TE+EQLHKIF
Sbjct: 314 QPLTSRVVTLWYRPPELLLGATEYGVAVDLWSTGCILAELLAGKPILPGQTEIEQLHKIF 373
Query: 250 KLCGSPAEEYWRKHKLPNATIFKPQQPYKRCISETFKDFPPSSLPLIDSLLAIDPDRRGT 309
KLCGSP+EEYW K KLP+ T+FKPQ+PY+R I+ETF+DF P +L L+D+LLAI+P RGT
Sbjct: 374 KLCGSPSEEYWAKAKLPDVTLFKPQRPYRRKIAETFRDFSPPALDLLDTLLAIEPSDRGT 433
Query: 310 ASAALNHEFFTTEPYACEPSSLPKYPPSKELDVKMRDEEARRQKALNGKANAVDGAKRVR 369
A+AAL+ +FF ++P AC+P+SLPK PPSKE D K+R +EA A+ A A G K
Sbjct: 434 AAAALDSDFFRSKPLACDPASLPKLPPSKEYDAKLRGKEA----AMRQNATAAIGGKGSM 489
Query: 370 ARERGR------AIPAPEA 382
+ + GR A PA +A
Sbjct: 490 SVKPGRNEQSKAAAPAQDA 508
>UniRef100_Q6YU33 Putative CRK1 protein [Oryza sativa]
Length = 725
Score = 525 bits (1352), Expect = e-147
Identities = 257/379 (67%), Positives = 312/379 (81%), Gaps = 11/379 (2%)
Query: 11 GWPSWLMAVAGEAIGDWTPRRANSFEKLAKIGQGTYSNVYKAKDLVTGKIVALKKVRFDN 70
GWPSWL +VAGE + W PRRA++FE+L KIGQGTYSNVYKA+DL TGK+VALK+VRF N
Sbjct: 134 GWPSWLTSVAGEVVQGWLPRRADTFERLDKIGQGTYSNVYKARDLETGKVVALKRVRFVN 193
Query: 71 LEPESVKFMAREILVLRKLD-HPNVVKLEGLVTSRMSCSLYLVFEYMEHDLAGLSAGQGV 129
++PESV+FMAREI VLR+LD HPNVV+LEG+VTSR+S SLYLVFEYM+HDLAGL+A G+
Sbjct: 194 MDPESVRFMAREIHVLRRLDGHPNVVRLEGIVTSRLSHSLYLVFEYMDHDLAGLAATPGL 253
Query: 130 KFTEPQVKCFMKQLLSGLEHCHSRGVLHRDIKGSNLLIDNEGILKIADFGLATFYNPNKK 189
+FTEPQVKC M Q+L+GL HCH RGVLHRDIKG+NLLI +G LKIADFGLATF++ +
Sbjct: 254 RFTEPQVKCLMAQILAGLRHCHDRGVLHRDIKGANLLIGGDGALKIADFGLATFFDAARP 313
Query: 190 QSMTSRVVTLWYRPPELLLGATFYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIF 249
Q +TSRVVTLWYRPPELLLGAT YGV +DLWS GCILAELLAGKPI+PG+TE+EQLHKIF
Sbjct: 314 QPLTSRVVTLWYRPPELLLGATEYGVAVDLWSTGCILAELLAGKPILPGQTEIEQLHKIF 373
Query: 250 KLCGSPAEEYWRKHKLPNATIFKPQQPYKRCISETFKDFPPSSLPLIDSLLAIDPDRRGT 309
KLCGSP+EEYW K KLP+ T+FKPQ+PY+R I+ETF+DF P +L L+D+LLAI+P RGT
Sbjct: 374 KLCGSPSEEYWAKAKLPDVTLFKPQRPYRRKIAETFRDFSPPALDLLDTLLAIEPSDRGT 433
Query: 310 ASAALNHEFFTTEPYACEPSSLPKYPPSKELDVKMRDEEARRQKALNGKANAVDGAKRVR 369
A+AAL+ +FF ++P AC+P+SLPK PPSKE D K+R +EA A+ A A G K
Sbjct: 434 AAAALDSDFFRSKPLACDPASLPKLPPSKEYDAKLRGKEA----AMRQNATAAIGGKGSM 489
Query: 370 ARERGR------AIPAPEA 382
+ + GR A PA +A
Sbjct: 490 SVKPGRNEQSKAAAPAQDA 508
>UniRef100_Q944Q4 AT5g44290/K9L2_5 [Arabidopsis thaliana]
Length = 644
Score = 524 bits (1350), Expect = e-147
Identities = 253/373 (67%), Positives = 309/373 (82%), Gaps = 3/373 (0%)
Query: 11 GWPSWLMAVAGEAIGDWTPRRANSFEKLAKIGQGTYSNVYKAKDLVTGKIVALKKVRFDN 70
GWP+WL++VAGEA+ +WTPRRA++FEKL KIGQGTYS+VYKA+DL KIVALK+VRFD
Sbjct: 113 GWPAWLVSVAGEALVNWTPRRASTFEKLEKIGQGTYSSVYKARDLTNNKIVALKRVRFDL 172
Query: 71 LEPESVKFMAREILVLRKLDHPNVVKLEGLVTSRMSCSLYLVFEYMEHDLAGLSAGQGVK 130
+ ESVKFMAREI+V+R+LDHPNV+KLEGL+T+ +S SLYLVFEYM+HDL GL++ G+K
Sbjct: 173 SDLESVKFMAREIIVMRRLDHPNVLKLEGLITASVSSSLYLVFEYMDHDLVGLASIPGIK 232
Query: 131 FTEPQVKCFMKQLLSGLEHCHSRGVLHRDIKGSNLLIDNEGILKIADFGLATFYNPNKKQ 190
F+EPQVKC+M+QLLSGL HCHSRGVLHRDIKGSNLLID+ G+LKIADFGLATF++P
Sbjct: 233 FSEPQVKCYMQQLLSGLHHCHSRGVLHRDIKGSNLLIDSNGVLKIADFGLATFFDPQNCV 292
Query: 191 SMTSRVVTLWYRPPELLLGATFYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFK 250
+TSRVVTLWYRPPELLLGA YGVG+DLWS GCIL EL +GKPI+ G+TEVEQLHKIFK
Sbjct: 293 PLTSRVVTLWYRPPELLLGACHYGVGVDLWSTGCILGELYSGKPILAGKTEVEQLHKIFK 352
Query: 251 LCGSPAEEYWRKHKLPNATIFKPQQPYKRCISETFKDFPPSSLPLIDSLLAIDPDRRGTA 310
LCGSP E YWRK KLP + F+P PY R ++E FKD P + L L+++LL+IDPDRRG+A
Sbjct: 353 LCGSPTENYWRKLKLPPSAAFRPALPYGRRVAEMFKDLPTNVLSLLEALLSIDPDRRGSA 412
Query: 311 SAALNHEFFTTEPYACEPSSLPKYPPSKELDVKMRDEEARRQKALNGKANAVDGAKRVRA 370
+ AL E+F TEP+AC+PSSLPKYPPSKE+D K+RD +A+RQ+ K D R R+
Sbjct: 413 ARALESEYFRTEPFACDPSSLPKYPPSKEIDAKIRD-DAKRQRPTQEKHERQDSQTR-RS 470
Query: 371 RERGRAIPAPEAN 383
ER + IP +AN
Sbjct: 471 HER-KLIPPVKAN 482
>UniRef100_Q9FVT3 CRK1 protein, putative [Arabidopsis thaliana]
Length = 686
Score = 523 bits (1348), Expect = e-147
Identities = 254/405 (62%), Positives = 317/405 (77%), Gaps = 13/405 (3%)
Query: 11 GWPSWLMAVAGEAIGDWTPRRANSFEKLAKIGQGTYSNVYKAKDLVTGKIVALKKVRFDN 70
GWPSWL++VAGEAI W PR A+SFEKL IGQGTYS+VY+A+DL T +IVALKKVRF N
Sbjct: 122 GWPSWLVSVAGEAINGWIPRSADSFEKLEMIGQGTYSSVYRARDLETNQIVALKKVRFAN 181
Query: 71 LEPESVKFMAREILVLRKLDHPNVVKLEGLVTSRMSCSLYLVFEYMEHDLAGLSAGQGVK 130
++PESV+FMAREI++LR+L+HPNV+KLEGL+ S+ S S+YL+FEYM+HDLAGL++ G+K
Sbjct: 182 MDPESVRFMAREIIILRRLNHPNVMKLEGLIISKASGSMYLIFEYMDHDLAGLASTPGIK 241
Query: 131 FTEPQVKCFMKQLLSGLEHCHSRGVLHRDIKGSNLLIDNEGILKIADFGLATFYNPNKKQ 190
F++ Q QLL GLEHCHS GVLHRDIK SNLL+D LKI DFGL+ FY +KQ
Sbjct: 242 FSQAQ------QLLLGLEHCHSCGVLHRDIKCSNLLLDRNNNLKIGDFGLSNFYRGQRKQ 295
Query: 191 SMTSRVVTLWYRPPELLLGATFYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFK 250
+TSRVVTLWYRPPELLLG+T YGV +DLWS GCILAEL GKP++PGRTEVEQ+HKIFK
Sbjct: 296 PLTSRVVTLWYRPPELLLGSTDYGVTVDLWSTGCILAELFTGKPLLPGRTEVEQMHKIFK 355
Query: 251 LCGSPAEEYWRKHKLPNATIFKPQQPYKRCISETFKDFPPSSLPLIDSLLAIDPDRRGTA 310
LCGSP+EEYWR+ +L +ATIFKPQ PYKRC+++TFKD P S+L L++ LLA++PD RGTA
Sbjct: 356 LCGSPSEEYWRRSRLRHATIFKPQHPYKRCVADTFKDLPSSALALLEVLLAVEPDARGTA 415
Query: 311 SAALNHEFFTTEPYACEPSSLPKYPPSKELDVKMRDEEARRQKALNGKANAVDGAKRVRA 370
S+AL EFFTT+P+ EPSSLP+Y P KE D K+R+EEARR+K + K N ++ R
Sbjct: 416 SSALQSEFFTTKPFPSEPSSLPRYQPRKEFDAKLREEEARRRKGSSSKQN-----EQKRL 470
Query: 371 RERGRAIPAPEANAEIQTNLDRWRVVTHANAKSKSEKFPPPHQDG 415
+A+PAP ANAE+ ++ + + N S SEKF P G
Sbjct: 471 ARESKAVPAPSANAELLASIQ--KRLGETNRTSISEKFNPEGDSG 513
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.317 0.134 0.403
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 878,778,236
Number of Sequences: 2790947
Number of extensions: 38603202
Number of successful extensions: 138912
Number of sequences better than 10.0: 19021
Number of HSP's better than 10.0 without gapping: 14325
Number of HSP's successfully gapped in prelim test: 4696
Number of HSP's that attempted gapping in prelim test: 94557
Number of HSP's gapped (non-prelim): 23044
length of query: 498
length of database: 848,049,833
effective HSP length: 132
effective length of query: 366
effective length of database: 479,644,829
effective search space: 175550007414
effective search space used: 175550007414
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 77 (34.3 bits)
Medicago: description of AC146747.6


