

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146723.12 + phase: 0
(172 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
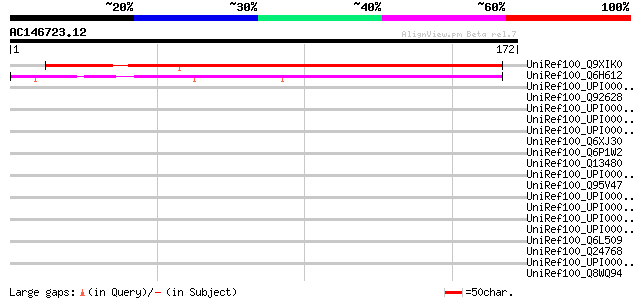
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9XIK0 T10O24.14 [Arabidopsis thaliana] 201 6e-51
UniRef100_Q6H612 Hypothetical protein P0505H05.24 [Oryza sativa] 148 6e-35
UniRef100_UPI000036CBF5 UPI000036CBF5 UniRef100 entry 35 0.79
UniRef100_Q92628 Hypothetical protein KIAA0232 [Homo sapiens] 35 0.79
UniRef100_UPI000031283D UPI000031283D UniRef100 entry 34 1.4
UniRef100_UPI0000293735 UPI0000293735 UniRef100 entry 34 1.8
UniRef100_UPI000036CE99 Grb-2 associated binder protein-1 [Equus... 33 2.3
UniRef100_Q6XJ30 Similar to Drosophila melanogaster Hsc70-4 [Dro... 33 2.3
UniRef100_Q6P1W2 GRB2-associated binding protein 1, isoform a [H... 33 2.3
UniRef100_Q13480 GRB2-associated binding protein 1 [Homo sapiens] 33 2.3
UniRef100_UPI0000325CEF UPI0000325CEF UniRef100 entry 33 3.0
UniRef100_Q95V47 70 kDa heat shock protein [Artemia sanfranciscana] 33 3.9
UniRef100_UPI000032FC7C UPI000032FC7C UniRef100 entry 32 5.1
UniRef100_UPI00002B34CC UPI00002B34CC UniRef100 entry 32 5.1
UniRef100_UPI0000291E36 UPI0000291E36 UniRef100 entry 32 5.1
UniRef100_UPI000026BE20 UPI000026BE20 UniRef100 entry 32 5.1
UniRef100_Q6L509 Putative hsp70 [Oryza sativa] 32 6.7
UniRef100_Q24768 Heat shock protein [Eimeria acervulina] 32 6.7
UniRef100_UPI00003C1736 UPI00003C1736 UniRef100 entry 32 8.8
UniRef100_Q8WQ94 HSC70 protein [Crassostrea gigas] 32 8.8
>UniRef100_Q9XIK0 T10O24.14 [Arabidopsis thaliana]
Length = 179
Score = 201 bits (511), Expect = 6e-51
Identities = 102/157 (64%), Positives = 118/157 (74%), Gaps = 7/157 (4%)
Query: 13 IPSSSSSIPSLCFSQTTTKCIQRFSFPSINRKVSTLPFSSIKRS--ITRAAEYKFPDPIP 70
+P SS C ++T QR KV SS+ R + RAAEYKFPDPIP
Sbjct: 26 LPIISSPAAVSCAIKSTQFFKQR-----CRTKVRDFSLSSLSRRGFVCRAAEYKFPDPIP 80
Query: 71 EFADSETEKFQNHLLNKLSKKDVFEESVEEVVGVCTEIFSTFLHSEYGGPGTLLVDPFID 130
EFA++ETEKF++H+LNKLSK+D+FE+SV+E+VGVCTEIF TFL SEYGGPGTLLV PFID
Sbjct: 81 EFAEAETEKFRDHMLNKLSKRDLFEDSVDEIVGVCTEIFETFLRSEYGGPGTLLVIPFID 140
Query: 131 MADIVNERGLPGGPQAARAAINWAQAHVDKDWNEWTG 167
MAD +NER LPGGPQAARAAI WAQ HVDKDW EWTG
Sbjct: 141 MADTLNERELPGGPQAARAAIKWAQDHVDKDWKEWTG 177
>UniRef100_Q6H612 Hypothetical protein P0505H05.24 [Oryza sativa]
Length = 172
Score = 148 bits (373), Expect = 6e-35
Identities = 86/178 (48%), Positives = 106/178 (59%), Gaps = 19/178 (10%)
Query: 1 MASSTWC-------LVPFLIPSSSSSIPSLCFSQTTTKCIQRFSFPSINRKVSTLPFSSI 53
MA+ W L P L+ S+SS PS S+ + R R + P +
Sbjct: 1 MAARLWAAAVAPATLNPPLLTLSASSSPSS--SRLRRSVLGRL------RSRAPRPADFV 52
Query: 54 KRSITRAA--EYKFPDPIPEFADSETEKFQNHLLNKLSKK--DVFEESVEEVVGVCTEIF 109
R AA +YKFPDPIPEFA ET KF+ H++ +L +K D F E VEE+V VCTEI
Sbjct: 53 CRRAKNAAYDDYKFPDPIPEFAAQETSKFKEHMMWRLEQKKDDYFGEHVEEIVDVCTEIL 112
Query: 110 STFLHSEYGGPGTLLVDPFIDMADIVNERGLPGGPQAARAAINWAQAHVDKDWNEWTG 167
TFL +Y GPGTLLV PF+DM + ERGLPG PQAARAAI WA+ ++DKDW WTG
Sbjct: 113 GTFLEHDYCGPGTLLVHPFLDMKGEIKERGLPGAPQAARAAIAWAEKNIDKDWKAWTG 170
>UniRef100_UPI000036CBF5 UPI000036CBF5 UniRef100 entry
Length = 1401
Score = 35.0 bits (79), Expect = 0.79
Identities = 27/87 (31%), Positives = 41/87 (47%), Gaps = 13/87 (14%)
Query: 62 EYK-----FPDPIPEFAD--SETEKFQNHLLNKLSKKDVFEESVEE----VVGVCTEIFS 110
EYK + +PI E+ S K + N+ D+ E+VEE V G+C I +
Sbjct: 402 EYKEEPLWYTEPIAEYFVPLSRKSKLETTYRNRQDTSDLTSEAVEELSESVHGLC--ISN 459
Query: 111 TFLHSEYGGPGTLLVDPFIDMADIVNE 137
LH Y GT + F++M ++NE
Sbjct: 460 NNLHKTYLAAGTFIDGHFVEMPAVINE 486
>UniRef100_Q92628 Hypothetical protein KIAA0232 [Homo sapiens]
Length = 1402
Score = 35.0 bits (79), Expect = 0.79
Identities = 27/87 (31%), Positives = 41/87 (47%), Gaps = 13/87 (14%)
Query: 62 EYK-----FPDPIPEFAD--SETEKFQNHLLNKLSKKDVFEESVEE----VVGVCTEIFS 110
EYK + +PI E+ S K + N+ D+ E+VEE V G+C I +
Sbjct: 402 EYKEEPLWYTEPIAEYFVPLSRKSKLETTYRNRQDTSDLTSEAVEELSESVHGLC--ISN 459
Query: 111 TFLHSEYGGPGTLLVDPFIDMADIVNE 137
LH Y GT + F++M ++NE
Sbjct: 460 NNLHKTYLAAGTFIDGHFVEMPAVINE 486
>UniRef100_UPI000031283D UPI000031283D UniRef100 entry
Length = 375
Score = 34.3 bits (77), Expect = 1.4
Identities = 25/72 (34%), Positives = 37/72 (50%), Gaps = 12/72 (16%)
Query: 35 RFSFPSINRKVSTLPF--SSIKRSIT-------RAAEYKFPDPIPEFA---DSETEKFQN 82
+FSF +I +++ L F IK SI ++ E+KF I EF D + EK +N
Sbjct: 185 KFSFNTIQKRMRELAFLNKGIKISINDLTQKKIKSTEFKFEGGIAEFVDYLDEKREKLKN 244
Query: 83 HLLNKLSKKDVF 94
N+L KK +F
Sbjct: 245 RNDNELFKKPIF 256
>UniRef100_UPI0000293735 UPI0000293735 UniRef100 entry
Length = 248
Score = 33.9 bits (76), Expect = 1.8
Identities = 28/83 (33%), Positives = 39/83 (46%), Gaps = 7/83 (8%)
Query: 13 IPSSSSSIPSLCFSQTTTKCIQRFSFPSINRKVSTLPFSSIKRSITRAAEYKFPDPIPEF 72
+P SS PS T+ I F F N KV+ + IK ++T E+K P F
Sbjct: 141 LPKSSGYTPSKAALNNFTQGIY-FDFKKFNIKVTLITPGFIKTALTDKNEFKMP-----F 194
Query: 73 ADSETEKFQNHLLNKLSKKDVFE 95
S TE + + N L+KK+ FE
Sbjct: 195 LKS-TEYAADEIFNGLTKKNNFE 216
>UniRef100_UPI000036CE99 Grb-2 associated binder protein-1 [Equus caballus]
Length = 601
Score = 33.5 bits (75), Expect = 2.3
Identities = 23/90 (25%), Positives = 38/90 (41%), Gaps = 3/90 (3%)
Query: 14 PSSSSSIPSLCFSQTTTKCIQRFSF-PSINRKVSTLPFSSIKRSITRAAEYKFPDPIPEF 72
P + SIP ++ CI PS + +ST+ + +++ + Y P P
Sbjct: 356 PVETCSIPRTASDTDSSYCIPTAGMSPSRSNTISTVDLNKLRKDASSQDCYDIPRTFPSD 415
Query: 73 ADSETEKFQNH--LLNKLSKKDVFEESVEE 100
S E F NH + N L+ V E ++E
Sbjct: 416 RSSSLEGFHNHFKVKNVLTVGSVSSEELDE 445
>UniRef100_Q6XJ30 Similar to Drosophila melanogaster Hsc70-4 [Drosophila yakuba]
Length = 84
Score = 33.5 bits (75), Expect = 2.3
Identities = 21/65 (32%), Positives = 30/65 (45%), Gaps = 10/65 (15%)
Query: 86 NKLSKKDVFEESVEEVVGVCTEIFSTFLHSEYGGPGTLLVDPFIDMADIVNERGLPGGPQ 145
N+L+ K+ +E +E+ GVC I + S G PG + P G+PG
Sbjct: 17 NQLADKEEYEHRQKELEGVCNPIITKLYQSAGGAPGGMPGGP----------GGMPGAAG 66
Query: 146 AARAA 150
AA AA
Sbjct: 67 AAGAA 71
>UniRef100_Q6P1W2 GRB2-associated binding protein 1, isoform a [Homo sapiens]
Length = 724
Score = 33.5 bits (75), Expect = 2.3
Identities = 23/90 (25%), Positives = 38/90 (41%), Gaps = 3/90 (3%)
Query: 14 PSSSSSIPSLCFSQTTTKCIQRFSF-PSINRKVSTLPFSSIKRSITRAAEYKFPDPIPEF 72
P + SIP ++ CI PS + +ST+ + +++ + Y P P
Sbjct: 356 PVETCSIPRTASDTDSSYCIPTAGMSPSRSNTISTVDLNKLRKDASSQDCYDIPRAFPSD 415
Query: 73 ADSETEKFQNH--LLNKLSKKDVFEESVEE 100
S E F NH + N L+ V E ++E
Sbjct: 416 RSSSLEGFHNHFKVKNVLTVGSVSSEELDE 445
>UniRef100_Q13480 GRB2-associated binding protein 1 [Homo sapiens]
Length = 694
Score = 33.5 bits (75), Expect = 2.3
Identities = 23/90 (25%), Positives = 38/90 (41%), Gaps = 3/90 (3%)
Query: 14 PSSSSSIPSLCFSQTTTKCIQRFSF-PSINRKVSTLPFSSIKRSITRAAEYKFPDPIPEF 72
P + SIP ++ CI PS + +ST+ + +++ + Y P P
Sbjct: 356 PVETCSIPRTASDTDSSYCIPTAGMSPSRSNTISTVDLNKLRKDASSQDCYDIPRAFPSD 415
Query: 73 ADSETEKFQNH--LLNKLSKKDVFEESVEE 100
S E F NH + N L+ V E ++E
Sbjct: 416 RSSSLEGFHNHFKVKNVLTVGSVSSEELDE 445
>UniRef100_UPI0000325CEF UPI0000325CEF UniRef100 entry
Length = 297
Score = 33.1 bits (74), Expect = 3.0
Identities = 21/82 (25%), Positives = 41/82 (49%), Gaps = 12/82 (14%)
Query: 35 RFSFPSINRKVSTLPF---------SSIKRSITRAAEYKFPDPIPEFA---DSETEKFQN 82
+FSF + +++ L F + + TR++E+K+ I EF D++ EK +N
Sbjct: 95 KFSFSILEKRLRELAFLNKGIKIVLNDFSQKKTRSSEFKYDGGILEFVEFLDNKREKLKN 154
Query: 83 HLLNKLSKKDVFEESVEEVVGV 104
N+L KK ++ E + + +
Sbjct: 155 KNDNELFKKPIYMEGSKNNIDI 176
>UniRef100_Q95V47 70 kDa heat shock protein [Artemia sanfranciscana]
Length = 644
Score = 32.7 bits (73), Expect = 3.9
Identities = 23/72 (31%), Positives = 36/72 (49%), Gaps = 11/72 (15%)
Query: 64 KFPDPIPEFADSET------EKFQNHLLNKLSKKDVFEESVEEVVGVCTEIFSTFLHSEY 117
KF D +PE AD T E + +N+L++K+ +EE +E+ VC I + Y
Sbjct: 557 KFKDKLPE-ADKNTILDKCNETIKWLDVNQLAEKEEYEEKQKEIEKVCNPIITKL----Y 611
Query: 118 GGPGTLLVDPFI 129
G G +L D +
Sbjct: 612 GQAGGMLADSLV 623
>UniRef100_UPI000032FC7C UPI000032FC7C UniRef100 entry
Length = 248
Score = 32.3 bits (72), Expect = 5.1
Identities = 28/83 (33%), Positives = 38/83 (45%), Gaps = 7/83 (8%)
Query: 13 IPSSSSSIPSLCFSQTTTKCIQRFSFPSINRKVSTLPFSSIKRSITRAAEYKFPDPIPEF 72
+P SS PS T+ I F F N KV+ + IK ++T E+K P F
Sbjct: 141 LPKSSGYTPSKAALNNFTQGIY-FDFKKFNVKVTLITPGFIKTALTDKNEFKMP-----F 194
Query: 73 ADSETEKFQNHLLNKLSKKDVFE 95
S TE + + N L KK+ FE
Sbjct: 195 LKS-TEYAADEIYNGLIKKNNFE 216
>UniRef100_UPI00002B34CC UPI00002B34CC UniRef100 entry
Length = 108
Score = 32.3 bits (72), Expect = 5.1
Identities = 25/86 (29%), Positives = 38/86 (44%), Gaps = 3/86 (3%)
Query: 13 IPSSSSSIPSLCFSQTTTKCIQRFSFPSINRKVSTLPFSSIKRSITRAAEYKFPDPIPEF 72
+P SS PS T+ I F F N +V+ + IK ++T E+K P +
Sbjct: 1 LPKSSGYTPSKASLNNFTQGIY-FDFKKFNIRVTLISPGFIKTALTDKNEFKM--PFLKD 57
Query: 73 ADSETEKFQNHLLNKLSKKDVFEESV 98
EK N L+NK S + +F +
Sbjct: 58 TSYAAEKIYNGLVNKKSFEIIFPPQI 83
>UniRef100_UPI0000291E36 UPI0000291E36 UniRef100 entry
Length = 210
Score = 32.3 bits (72), Expect = 5.1
Identities = 28/83 (33%), Positives = 38/83 (45%), Gaps = 7/83 (8%)
Query: 13 IPSSSSSIPSLCFSQTTTKCIQRFSFPSINRKVSTLPFSSIKRSITRAAEYKFPDPIPEF 72
+P SS PS T+ I F F N KV+ + IK ++T E+K P F
Sbjct: 103 LPKSSGYTPSKAALNNFTQGIY-FDFKKFNVKVTLITPGFIKTALTDKNEFKMP-----F 156
Query: 73 ADSETEKFQNHLLNKLSKKDVFE 95
S TE + + N L KK+ FE
Sbjct: 157 LKS-TEYAADEIYNGLIKKNNFE 178
>UniRef100_UPI000026BE20 UPI000026BE20 UniRef100 entry
Length = 248
Score = 32.3 bits (72), Expect = 5.1
Identities = 22/63 (34%), Positives = 33/63 (51%), Gaps = 1/63 (1%)
Query: 30 TKCIQRFSFPSINRKVSTLP-FSSIKRSITRAAEYKFPDPIPEFADSETEKFQNHLLNKL 88
+K +++F N VST+ F I SI + KF I E S++ KF +HL+ L
Sbjct: 186 SKELKKFIESGKNIIVSTIQKFPVISSSINKEKNKKFAVVIDEVHSSQSGKFSDHLVKSL 245
Query: 89 SKK 91
SK+
Sbjct: 246 SKQ 248
>UniRef100_Q6L509 Putative hsp70 [Oryza sativa]
Length = 646
Score = 32.0 bits (71), Expect = 6.7
Identities = 20/61 (32%), Positives = 31/61 (50%), Gaps = 11/61 (18%)
Query: 86 NKLSKKDVFEESVEEVVGVCTEIFSTFLHSEYGGPGTLLVDPFIDMADIVNERGLPGGPQ 145
N+L++ D FE+ ++E+ G+C I + Y GPG DMA ++E GG
Sbjct: 589 NQLAEADEFEDKMKELEGICNPIIAKM----YQGPGA-------DMAGGMDEDAPAGGSG 637
Query: 146 A 146
A
Sbjct: 638 A 638
>UniRef100_Q24768 Heat shock protein [Eimeria acervulina]
Length = 646
Score = 32.0 bits (71), Expect = 6.7
Identities = 20/59 (33%), Positives = 28/59 (46%), Gaps = 4/59 (6%)
Query: 86 NKLSKKDVFEESVEEVVGVCTEIFSTFLHSEYGGPGTLLVDPFIDMADIVNERGLPGGP 144
N+L++K+ FE +EV VCT I + S G G + M D+ G GGP
Sbjct: 586 NQLAEKEEFEAKQKEVEAVCTPIVTKMYQSAAGAQGGMPG----GMPDMSAAAGAAGGP 640
>UniRef100_UPI00003C1736 UPI00003C1736 UniRef100 entry
Length = 1678
Score = 31.6 bits (70), Expect = 8.8
Identities = 34/127 (26%), Positives = 51/127 (39%), Gaps = 17/127 (13%)
Query: 16 SSSSIPSLCFSQTTTKCIQ-------RF-----SFPSINRKVSTLPFSSIKRSITRAAEY 63
+ SS P+ Q T +Q RF S PS+ V T P SI R I +
Sbjct: 617 AGSSAPAARLRQQETSSLQHSSASRTRFLTFHESAPSLRDVVGTFPLFSIARQIHPEPFF 676
Query: 64 KFPDPIP----EFADSETEKFQNHLLNKLSKKDVFEESVEEVVGVCTEIFSTFLHSEYGG 119
P +P E+A FQ+H L L D +++ E V + L+ ++G
Sbjct: 677 SNPSDLPARPREYA-GRMFHFQSHSLPYLRSFDHWDQKSEHVASTQKDKPPKRLYWQFGP 735
Query: 120 PGTLLVD 126
P L++
Sbjct: 736 PPPTLLE 742
>UniRef100_Q8WQ94 HSC70 protein [Crassostrea gigas]
Length = 599
Score = 31.6 bits (70), Expect = 8.8
Identities = 17/58 (29%), Positives = 26/58 (44%)
Query: 86 NKLSKKDVFEESVEEVVGVCTEIFSTFLHSEYGGPGTLLVDPFIDMADIVNERGLPGG 143
N+L+ K+ FE +E+ GVC I + + G PG + + G PGG
Sbjct: 530 NQLADKEEFEHKQKELEGVCNPIITKLYQASGGAPGGGMPGGMPNFGGGAPGGGAPGG 587
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.316 0.132 0.404
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 309,195,333
Number of Sequences: 2790947
Number of extensions: 12479912
Number of successful extensions: 32827
Number of sequences better than 10.0: 25
Number of HSP's better than 10.0 without gapping: 2
Number of HSP's successfully gapped in prelim test: 23
Number of HSP's that attempted gapping in prelim test: 32822
Number of HSP's gapped (non-prelim): 25
length of query: 172
length of database: 848,049,833
effective HSP length: 118
effective length of query: 54
effective length of database: 518,718,087
effective search space: 28010776698
effective search space used: 28010776698
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 70 (31.6 bits)
Medicago: description of AC146723.12


