

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
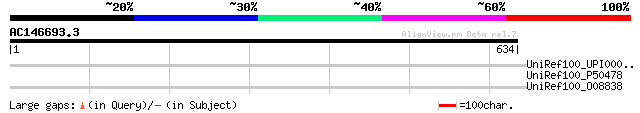
Query= AC146693.3 - phase: 0 /pseudo
(634 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_UPI00003ABBC4 UPI00003ABBC4 UniRef100 entry 36 3.0
UniRef100_P50478 Amphiphysin [Gallus gallus] 36 3.0
UniRef100_O08838 Amphiphysin [Rattus norvegicus] 35 8.8
>UniRef100_UPI00003ABBC4 UPI00003ABBC4 UniRef100 entry
Length = 682
Score = 36.2 bits (82), Expect = 3.0
Identities = 21/55 (38%), Positives = 27/55 (48%), Gaps = 7/55 (12%)
Query: 318 PNRISERLSPNSLPRLILSPHTPKCLRPKSPKWPNKWPPLLKH*ESFPVNPKQTP 372
P +S SP + P +L+P +P RPKSP K PP+ P PK TP
Sbjct: 264 PEEVSPLPSPTASPNHMLAPASPAPARPKSPTQLRKGPPV-------PPLPKLTP 311
>UniRef100_P50478 Amphiphysin [Gallus gallus]
Length = 682
Score = 36.2 bits (82), Expect = 3.0
Identities = 21/55 (38%), Positives = 27/55 (48%), Gaps = 7/55 (12%)
Query: 318 PNRISERLSPNSLPRLILSPHTPKCLRPKSPKWPNKWPPLLKH*ESFPVNPKQTP 372
P +S SP + P +L+P +P RPKSP K PP+ P PK TP
Sbjct: 264 PEEVSPLPSPTASPNHMLAPASPAPARPKSPTQLRKGPPV-------PPLPKLTP 311
>UniRef100_O08838 Amphiphysin [Rattus norvegicus]
Length = 683
Score = 34.7 bits (78), Expect = 8.8
Identities = 20/55 (36%), Positives = 26/55 (46%), Gaps = 7/55 (12%)
Query: 318 PNRISERLSPNSLPRLILSPHTPKCLRPKSPKWPNKWPPLLKH*ESFPVNPKQTP 372
P S SP + P L+P +P +RP+SP K PP+ P PK TP
Sbjct: 264 PEEASPLPSPTASPNHTLAPASPAPVRPRSPSQTRKGPPV-------PPLPKVTP 311
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.364 0.164 0.631
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 851,754,565
Number of Sequences: 2790947
Number of extensions: 29630132
Number of successful extensions: 136764
Number of sequences better than 10.0: 3
Number of HSP's better than 10.0 without gapping: 0
Number of HSP's successfully gapped in prelim test: 3
Number of HSP's that attempted gapping in prelim test: 136764
Number of HSP's gapped (non-prelim): 3
length of query: 634
length of database: 848,049,833
effective HSP length: 134
effective length of query: 500
effective length of database: 474,062,935
effective search space: 237031467500
effective search space used: 237031467500
T: 11
A: 40
X1: 14 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.5 bits)
S2: 78 (34.7 bits)
Medicago: description of AC146693.3


