

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146632.13 - phase: 0
(126 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
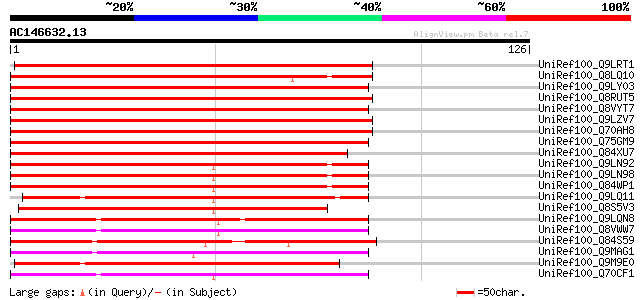
Sequences producing significant alignments: (bits) Value
UniRef100_Q9LRT1 Receptor protein kinase [Arabidopsis thaliana] 145 2e-34
UniRef100_Q8LQ10 Putative receptor protein kinase [Oryza sativa] 135 2e-31
UniRef100_Q9LY03 Hypothetical protein T5P19_20 [Arabidopsis thal... 115 3e-25
UniRef100_Q8RUT5 Putative receptor-like protein kinase [Oryza sa... 115 3e-25
UniRef100_Q8VYT7 Hypothetical protein At3g56370 [Arabidopsis tha... 115 3e-25
UniRef100_Q9LZV7 Hypothetical protein T20L15_160 [Arabidopsis th... 113 1e-24
UniRef100_Q70AH8 Receptor-like kinase with LRR repeats [Triticum... 108 4e-23
UniRef100_Q75GM9 Hypothetical protein OSJNBa0018K15.10 [Oryza sa... 106 1e-22
UniRef100_Q84XU7 Receptor-like protein kinase [Elaeis guineensis... 97 8e-20
UniRef100_Q9LN92 T12C24.1 [Arabidopsis thaliana] 94 7e-19
UniRef100_Q9LN98 F5O11.21 [Arabidopsis thaliana] 94 7e-19
UniRef100_Q84WP1 Putative leucine-rich repeat transmembrane prot... 94 7e-19
UniRef100_Q9LQ11 F16P17.10 protein [Arabidopsis thaliana] 89 2e-17
UniRef100_Q8S5V3 Hypothetical protein OJ1015F07.14 [Oryza sativa] 78 4e-14
UniRef100_Q9LQN8 F24B9.29 protein [Arabidopsis thaliana] 68 5e-11
UniRef100_Q8VWW7 Putative receptor-like protein kinase RLPK1 [Gl... 68 5e-11
UniRef100_Q84S59 Putative serine/threonine kinase protein [Oryza... 67 1e-10
UniRef100_Q9MAG1 F12M16.30 [Arabidopsis thaliana] 67 1e-10
UniRef100_Q9M9E0 T16N11.4 protein [Arabidopsis thaliana] 66 2e-10
UniRef100_Q70CF1 Receptor-like protein kinase [Fagus sylvatica] 65 3e-10
>UniRef100_Q9LRT1 Receptor protein kinase [Arabidopsis thaliana]
Length = 1016
Score = 145 bits (366), Expect = 2e-34
Identities = 66/87 (75%), Positives = 78/87 (88%)
Query: 2 SNRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEYGEDNVLILNDHVR 61
+NRFQ+ALGYVAPEL CQ LRVNEKCDVYGFGV+ILE+VTG+RPVEYGED+ +IL+DHVR
Sbjct: 889 NNRFQNALGYVAPELECQNLRVNEKCDVYGFGVLILELVTGRRPVEYGEDSFVILSDHVR 948
Query: 62 VLLEHGNALECVDPSLMSEYPEDEVLP 88
V+LE GN LEC+DP + +Y EDEVLP
Sbjct: 949 VMLEQGNVLECIDPVMEEQYSEDEVLP 975
>UniRef100_Q8LQ10 Putative receptor protein kinase [Oryza sativa]
Length = 1013
Score = 135 bits (341), Expect = 2e-31
Identities = 65/91 (71%), Positives = 79/91 (86%), Gaps = 4/91 (4%)
Query: 1 MSNRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEYGEDNVLILNDHV 60
MS+RFQ +GYVAPELACQ LR+NEKCD+YGFGV+ILE+VTG+R VEYG+D+V+IL D V
Sbjct: 881 MSSRFQGGMGYVAPELACQSLRINEKCDIYGFGVLILELVTGRRAVEYGDDDVVILIDQV 940
Query: 61 RVLLEHG---NALECVDPSLMSEYPEDEVLP 88
RVLL+HG N LECVDPS+ E+PE+EVLP
Sbjct: 941 RVLLDHGGGSNVLECVDPSI-GEFPEEEVLP 970
>UniRef100_Q9LY03 Hypothetical protein T5P19_20 [Arabidopsis thaliana]
Length = 964
Score = 115 bits (287), Expect = 3e-25
Identities = 50/87 (57%), Positives = 70/87 (79%)
Query: 1 MSNRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEYGEDNVLILNDHV 60
+S++ QSALGY+APE AC+ +++ EKCDVYGFGV++LE+VTGK+PVEY ED+V++L D V
Sbjct: 835 LSSKIQSALGYMAPEFACRTVKITEKCDVYGFGVLVLEVVTGKKPVEYMEDDVVVLCDMV 894
Query: 61 RVLLEHGNALECVDPSLMSEYPEDEVL 87
R LE G A EC+DP L ++P +E +
Sbjct: 895 REALEDGRADECIDPRLQGKFPVEEAV 921
>UniRef100_Q8RUT5 Putative receptor-like protein kinase [Oryza sativa]
Length = 947
Score = 115 bits (287), Expect = 3e-25
Identities = 50/88 (56%), Positives = 70/88 (78%)
Query: 1 MSNRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEYGEDNVLILNDHV 60
+S++ QSALGY+APE AC+ +++ EKCDVYGFGV++LE++TG+RPVEY ED+V++L D V
Sbjct: 819 LSSKIQSALGYMAPEFACKTVKITEKCDVYGFGVLVLEVLTGRRPVEYLEDDVVVLCDLV 878
Query: 61 RVLLEHGNALECVDPSLMSEYPEDEVLP 88
R LE G +C+DP L E+P +E LP
Sbjct: 879 RSALEEGRLEDCMDPRLCGEFPMEEALP 906
>UniRef100_Q8VYT7 Hypothetical protein At3g56370 [Arabidopsis thaliana]
Length = 964
Score = 115 bits (287), Expect = 3e-25
Identities = 50/87 (57%), Positives = 70/87 (79%)
Query: 1 MSNRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEYGEDNVLILNDHV 60
+S++ QSALGY+APE AC+ +++ EKCDVYGFGV++LE+VTGK+PVEY ED+V++L D V
Sbjct: 835 LSSKIQSALGYMAPEFACRTVKITEKCDVYGFGVLVLEVVTGKKPVEYMEDDVVVLCDMV 894
Query: 61 RVLLEHGNALECVDPSLMSEYPEDEVL 87
R LE G A EC+DP L ++P +E +
Sbjct: 895 REALEDGRADECIDPRLQGKFPVEEAV 921
>UniRef100_Q9LZV7 Hypothetical protein T20L15_160 [Arabidopsis thaliana]
Length = 967
Score = 113 bits (282), Expect = 1e-24
Identities = 48/88 (54%), Positives = 67/88 (75%)
Query: 1 MSNRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEYGEDNVLILNDHV 60
+S + QSALGY APE AC+ +++ ++CDVYGFG+++LE+VTGKRPVEY ED+V++L + V
Sbjct: 843 LSGKVQSALGYTAPEFACRTVKITDRCDVYGFGILVLEVVTGKRPVEYAEDDVVVLCETV 902
Query: 61 RVLLEHGNALECVDPSLMSEYPEDEVLP 88
R LE G ECVDP L +P +E +P
Sbjct: 903 REGLEEGRVEECVDPRLRGNFPAEEAIP 930
>UniRef100_Q70AH8 Receptor-like kinase with LRR repeats [Triticum aestivum]
Length = 168
Score = 108 bits (269), Expect = 4e-23
Identities = 46/88 (52%), Positives = 67/88 (75%)
Query: 1 MSNRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEYGEDNVLILNDHV 60
+S++ QSALGY+APE C+ +++ EKCDVYGFGV++LE++TG+ PVEY ED+V++L D V
Sbjct: 40 LSSKVQSALGYMAPEFTCRTVKITEKCDVYGFGVLVLEVMTGRTPVEYMEDDVIVLCDVV 99
Query: 61 RVLLEHGNALECVDPSLMSEYPEDEVLP 88
R L+ G ECVD L ++P +E +P
Sbjct: 100 RAALDEGKVEECVDEKLCGKFPLEEAVP 127
>UniRef100_Q75GM9 Hypothetical protein OSJNBa0018K15.10 [Oryza sativa]
Length = 917
Score = 106 bits (264), Expect = 1e-22
Identities = 47/87 (54%), Positives = 66/87 (75%)
Query: 1 MSNRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEYGEDNVLILNDHV 60
+S++ QSALGY+APE C+ + V EKCDVYGFGV++LEI+TG+RPVEY ED+V++L D V
Sbjct: 789 LSSKIQSALGYMAPEFTCRTVNVTEKCDVYGFGVIVLEILTGRRPVEYLEDDVVVLCDVV 848
Query: 61 RVLLEHGNALECVDPSLMSEYPEDEVL 87
R L+ G +C+DP L E+ +E +
Sbjct: 849 RAALDDGRVEDCMDPRLSGEFSMEEAM 875
>UniRef100_Q84XU7 Receptor-like protein kinase [Elaeis guineensis var. tenera]
Length = 938
Score = 97.1 bits (240), Expect = 8e-20
Identities = 46/82 (56%), Positives = 60/82 (73%)
Query: 1 MSNRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEYGEDNVLILNDHV 60
+S++ QSALGY+APE AC+ +++ EKCDVY FGV++LEI KR +EY ED+V++L D V
Sbjct: 817 LSSKIQSALGYMAPEFACRTVKITEKCDVYAFGVLVLEIQARKRLLEYMEDDVVVLCDMV 876
Query: 61 RVLLEHGNALECVDPSLMSEYP 82
R LE G ECVD LM E P
Sbjct: 877 RGALEEGKVAECVDGRLMWEVP 898
>UniRef100_Q9LN92 T12C24.1 [Arabidopsis thaliana]
Length = 221
Score = 94.0 bits (232), Expect = 7e-19
Identities = 47/88 (53%), Positives = 68/88 (76%), Gaps = 2/88 (2%)
Query: 1 MSNRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEY-GEDNVLILNDH 59
++ +F +A+GY+APELA Q LR +EKCDVY +GV++LE+VTG++PVE E+ VLIL D+
Sbjct: 99 LTKKFHNAVGYIAPELAQQSLRASEKCDVYSYGVVLLELVTGRKPVESPSENQVLILRDY 158
Query: 60 VRVLLEHGNALECVDPSLMSEYPEDEVL 87
VR LLE G+A +C D L E+ E+E++
Sbjct: 159 VRDLLETGSASDCFDRRL-REFEENELI 185
>UniRef100_Q9LN98 F5O11.21 [Arabidopsis thaliana]
Length = 893
Score = 94.0 bits (232), Expect = 7e-19
Identities = 47/88 (53%), Positives = 68/88 (76%), Gaps = 2/88 (2%)
Query: 1 MSNRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEY-GEDNVLILNDH 59
++ +F +A+GY+APELA Q LR +EKCDVY +GV++LE+VTG++PVE E+ VLIL D+
Sbjct: 771 LTKKFHNAVGYIAPELAQQSLRASEKCDVYSYGVVLLELVTGRKPVESPSENQVLILRDY 830
Query: 60 VRVLLEHGNALECVDPSLMSEYPEDEVL 87
VR LLE G+A +C D L E+ E+E++
Sbjct: 831 VRDLLETGSASDCFDRRL-REFEENELI 857
>UniRef100_Q84WP1 Putative leucine-rich repeat transmembrane protein kinase
[Arabidopsis thaliana]
Length = 882
Score = 94.0 bits (232), Expect = 7e-19
Identities = 47/88 (53%), Positives = 68/88 (76%), Gaps = 2/88 (2%)
Query: 1 MSNRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEY-GEDNVLILNDH 59
++ +F +A+GY+APELA Q LR +EKCDVY +GV++LE+VTG++PVE E+ VLIL D+
Sbjct: 760 LTKKFHNAVGYIAPELAQQSLRASEKCDVYSYGVVLLELVTGRKPVESPSENQVLILRDY 819
Query: 60 VRVLLEHGNALECVDPSLMSEYPEDEVL 87
VR LLE G+A +C D L E+ E+E++
Sbjct: 820 VRDLLETGSASDCFDRRL-REFEENELI 846
>UniRef100_Q9LQ11 F16P17.10 protein [Arabidopsis thaliana]
Length = 890
Score = 89.0 bits (219), Expect = 2e-17
Identities = 46/85 (54%), Positives = 66/85 (77%), Gaps = 3/85 (3%)
Query: 4 RFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEY-GEDNVLILNDHVRV 62
+F +A+GY+APELA Q LRV++KCDVY +GV++LE+VTG++PVE E+ V+IL DHVR
Sbjct: 772 KFHNAVGYIAPELA-QSLRVSDKCDVYSYGVVLLELVTGRKPVESPSENEVVILRDHVRN 830
Query: 63 LLEHGNALECVDPSLMSEYPEDEVL 87
LLE G+A +C D L + E+E++
Sbjct: 831 LLETGSASDCFDRRLRG-FEENELI 854
>UniRef100_Q8S5V3 Hypothetical protein OJ1015F07.14 [Oryza sativa]
Length = 891
Score = 78.2 bits (191), Expect = 4e-14
Identities = 37/76 (48%), Positives = 54/76 (70%), Gaps = 1/76 (1%)
Query: 3 NRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEY-GEDNVLILNDHVR 61
+R +A+GY+APELA LR ++K DV+ FGV++LEIVTG++PVE G ++L D+VR
Sbjct: 773 SRLHAAIGYIAPELASPSLRYSDKSDVFSFGVVLLEIVTGRKPVESPGVATAVVLRDYVR 832
Query: 62 VLLEHGNALECVDPSL 77
+LE G +C D S+
Sbjct: 833 AILEDGTVSDCFDRSM 848
>UniRef100_Q9LQN8 F24B9.29 protein [Arabidopsis thaliana]
Length = 554
Score = 67.8 bits (164), Expect = 5e-11
Identities = 41/89 (46%), Positives = 55/89 (61%), Gaps = 4/89 (4%)
Query: 1 MSNRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEYG--EDNVLILND 58
+S R +GY+APE A + + EK DVY FGV+ LEIV+GK + ED V +L D
Sbjct: 377 ISTRIAGTIGYMAPEYAMRGY-LTEKADVYSFGVVALEIVSGKSNTNFRPTEDFVYLL-D 434
Query: 59 HVRVLLEHGNALECVDPSLMSEYPEDEVL 87
VL E G+ LE VDP+L S+Y E+E +
Sbjct: 435 WAYVLQERGSLLELVDPTLASDYSEEEAM 463
>UniRef100_Q8VWW7 Putative receptor-like protein kinase RLPK1 [Glycine max]
Length = 225
Score = 67.8 bits (164), Expect = 5e-11
Identities = 39/88 (44%), Positives = 53/88 (59%), Gaps = 2/88 (2%)
Query: 1 MSNRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEYG-EDNVLILNDH 59
+S R +GY+APE A + + +K DVY FGV+ LEIV+GK Y ++ + L D
Sbjct: 30 ISTRIAGTIGYMAPEYAMRGY-LTDKADVYSFGVVALEIVSGKSNANYRPKEEFVYLLDW 88
Query: 60 VRVLLEHGNALECVDPSLMSEYPEDEVL 87
VL E GN LE VDPSL S+Y +E +
Sbjct: 89 AYVLQEQGNLLELVDPSLGSKYSSEEAM 116
>UniRef100_Q84S59 Putative serine/threonine kinase protein [Oryza sativa]
Length = 643
Score = 66.6 bits (161), Expect = 1e-10
Identities = 39/93 (41%), Positives = 58/93 (61%), Gaps = 8/93 (8%)
Query: 1 MSNRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPV-EYGEDNVLILNDH 59
+++R GY++PE A + + + K DVY FGV++LEI+TG+R YG D+V+ D
Sbjct: 504 ITHRIAGTYGYMSPEYAMRG-QYSMKLDVYSFGVLVLEIITGRRNFGSYGSDHVV---DL 559
Query: 60 VRVLLEH---GNALECVDPSLMSEYPEDEVLPC 89
+ V EH A+E +DPSL + YP D+VL C
Sbjct: 560 IYVTWEHWTSDKAIELIDPSLGNHYPVDKVLKC 592
>UniRef100_Q9MAG1 F12M16.30 [Arabidopsis thaliana]
Length = 854
Score = 66.6 bits (161), Expect = 1e-10
Identities = 38/88 (43%), Positives = 51/88 (57%), Gaps = 2/88 (2%)
Query: 1 MSNRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGK-RPVEYGEDNVLILNDH 59
+S R GY+APE A + + +K DVY FG++ LEIV G+ +E ++N L D
Sbjct: 683 ISTRIAGTFGYMAPEYAMRG-HLTDKADVYSFGIVALEIVHGRSNKIERSKNNTFYLIDW 741
Query: 60 VRVLLEHGNALECVDPSLMSEYPEDEVL 87
V VL E N LE VDP L SEY +E +
Sbjct: 742 VEVLREKNNLLELVDPRLGSEYNREEAM 769
>UniRef100_Q9M9E0 T16N11.4 protein [Arabidopsis thaliana]
Length = 656
Score = 66.2 bits (160), Expect = 2e-10
Identities = 34/79 (43%), Positives = 48/79 (60%), Gaps = 1/79 (1%)
Query: 2 SNRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEYGEDNVLILNDHVR 61
+ R LGY+APELA E DVY FGV++LE+V+G+RP+EY E+ ++L D VR
Sbjct: 518 TTRVVGTLGYLAPELA-SASAPTEASDVYSFGVVVLEVVSGRRPIEYAEEEDMVLVDWVR 576
Query: 62 VLLEHGNALECVDPSLMSE 80
L G ++ D + SE
Sbjct: 577 DLYGGGRVVDAADERVRSE 595
>UniRef100_Q70CF1 Receptor-like protein kinase [Fagus sylvatica]
Length = 217
Score = 65.1 bits (157), Expect = 3e-10
Identities = 37/88 (42%), Positives = 51/88 (57%), Gaps = 2/88 (2%)
Query: 1 MSNRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEY-GEDNVLILNDH 59
+S R +GY+APE A + + +K DVY FG++ LEIV+GK Y ++ + L D
Sbjct: 33 ISTRIAGTIGYMAPEYAMRGY-LTDKADVYSFGIVALEIVSGKSNTNYRPKEEFVYLLDW 91
Query: 60 VRVLLEHGNALECVDPSLMSEYPEDEVL 87
VL E GN LE VDP L S Y +E +
Sbjct: 92 AYVLQEQGNLLELVDPRLGSSYSSEEAM 119
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.321 0.141 0.423
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 155,044,780
Number of Sequences: 2790947
Number of extensions: 5353039
Number of successful extensions: 19566
Number of sequences better than 10.0: 2609
Number of HSP's better than 10.0 without gapping: 1736
Number of HSP's successfully gapped in prelim test: 873
Number of HSP's that attempted gapping in prelim test: 17422
Number of HSP's gapped (non-prelim): 2661
length of query: 126
length of database: 848,049,833
effective HSP length: 102
effective length of query: 24
effective length of database: 563,373,239
effective search space: 13520957736
effective search space used: 13520957736
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 67 (30.4 bits)
Medicago: description of AC146632.13


