

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146631.5 - phase: 0 /pseudo
(359 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
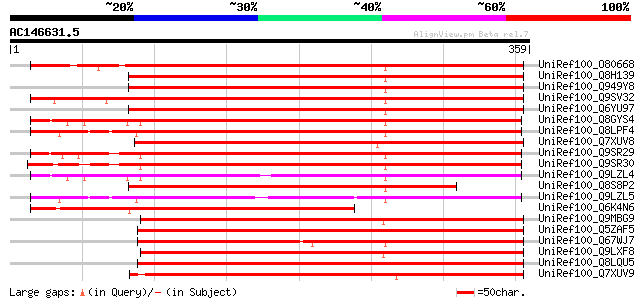
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_O80668 Expressed protein [Arabidopsis thaliana] 472 e-132
UniRef100_Q8H139 Hypothetical protein At2g41190 [Arabidopsis tha... 434 e-120
UniRef100_Q949Y8 Hypothetical protein At2g39130 [Arabidopsis tha... 323 6e-87
UniRef100_Q9SV32 Hypothetical protein F28P10.190 [Arabidopsis th... 313 4e-84
UniRef100_Q6YU97 Putative amino acid transport protein [Oryza sa... 310 3e-83
UniRef100_Q8GYS4 Hypothetical protein At5g02180/T7H20_230 [Arabi... 306 6e-82
UniRef100_Q8LPF4 AT5g02170/T7H20_220 [Arabidopsis thaliana] 291 1e-77
UniRef100_Q7XUV8 OSJNBa0072F16.7 protein [Oryza sativa] 289 7e-77
UniRef100_Q9SR29 F3L24.21 protein [Arabidopsis thaliana] 283 5e-75
UniRef100_Q9SR30 F3L24.20 protein [Arabidopsis thaliana] 282 1e-74
UniRef100_Q9LZL4 Hypothetical protein T7H20_230 [Arabidopsis tha... 278 1e-73
UniRef100_Q8S8P2 Hypothetical protein At2g39130 [Arabidopsis tha... 269 1e-70
UniRef100_Q9LZL5 Hypothetical protein T7H20_220 [Arabidopsis tha... 264 2e-69
UniRef100_Q6K4N6 Amino acid transporter-like [Oryza sativa] 264 2e-69
UniRef100_Q9MBG9 Gb|AAC79623.2 [Arabidopsis thaliana] 260 5e-68
UniRef100_Q5ZAF5 Amino acid transporter-like [Oryza sativa] 254 3e-66
UniRef100_Q67WJ7 Putative amino acid transport protein [Oryza sa... 252 1e-65
UniRef100_Q9LXF8 Hypothetical protein F8M21_130 [Arabidopsis tha... 250 5e-65
UniRef100_Q8LQU5 Amino acid transporter protein-like [Oryza sativa] 236 7e-61
UniRef100_Q7XUV9 OSJNBa0072F16.6 protein [Oryza sativa] 229 1e-58
>UniRef100_O80668 Expressed protein [Arabidopsis thaliana]
Length = 536
Score = 472 bits (1215), Expect = e-132
Identities = 228/350 (65%), Positives = 277/350 (79%), Gaps = 16/350 (4%)
Query: 15 RETTDSYTIATAPNFVSILRGPSSIYSSFSNRSKSDLDIDGKTPCL------SGLDGTTQ 68
RETTDSYTIA +P F S+ P S Y + S+S+LD++ K P L S TQ
Sbjct: 72 RETTDSYTIAASPIFGSLRSNPPSFYRA----SRSNLDVESKAPLLPERHDDSDKASATQ 127
Query: 69 STWWEKRSTQNLVGEMPLG-YGCSYTQTVFNGINVMAGVGLLSTPYTVKQAGWMGLVLML 127
S W K S E+P+G YGCS QT+FN INVMAGVGLLSTPYTVK+AGW +V++L
Sbjct: 128 SAWSHKGS---FAEELPIGGYGCSVIQTIFNAINVMAGVGLLSTPYTVKEAGWASMVILL 184
Query: 128 IFASVCCYTATLMRHCFESREGLTSYPDIGEAAFGRYGRIFVSIILYTELYSYCVEFITL 187
+FA +CCYTATLM+ CFE++ G+ +YPDIGEAAFG+YGRI + ++LYTELYSYCVEFI L
Sbjct: 185 LFAVICCYTATLMKDCFENKTGIITYPDIGEAAFGKYGRILICMLLYTELYSYCVEFIIL 244
Query: 188 EGDNLTGLFPGTSLDIGGLHLDSMHLFGVLTALVILPTVWLKDLRVISYLSVGGIAATIL 247
EGDNLTGLFPGTSLD+ G LDS HLFG+LTAL++LPTVWLKDLR+ISYLS GG+ AT L
Sbjct: 245 EGDNLTGLFPGTSLDLLGFRLDSKHLFGILTALIVLPTVWLKDLRIISYLSAGGVIATAL 304
Query: 248 IIISVFSVGTT--VGFHHTGRVVNWSGIPFAIGVYGFCFAGHSVFPNIYQSMADKKQYTK 305
I +SVF +GTT +GFHHTG+ V W+GIPFAIG+YGFC++GHSVFPNIYQSMADK ++ K
Sbjct: 305 IAVSVFFLGTTGGIGFHHTGQAVKWNGIPFAIGIYGFCYSGHSVFPNIYQSMADKTKFNK 364
Query: 306 ALITCFVLCILIYGSVAVMGFLSFGDDTLSQITLNMPAGAFASKVALWTT 355
A+ITCF++C+L+YG VA+MG+L FG+ TLSQITLNMP F SKVA WTT
Sbjct: 365 AVITCFIICVLLYGGVAIMGYLMFGEATLSQITLNMPQDQFFSKVAQWTT 414
>UniRef100_Q8H139 Hypothetical protein At2g41190 [Arabidopsis thaliana]
Length = 407
Score = 434 bits (1116), Expect = e-120
Identities = 198/276 (71%), Positives = 240/276 (86%), Gaps = 3/276 (1%)
Query: 83 EMPLG-YGCSYTQTVFNGINVMAGVGLLSTPYTVKQAGWMGLVLMLIFASVCCYTATLMR 141
E+P+G YGCS QT+FN INVMAGVGLLSTPYTVK+AGW +V++L+FA +CCYTATLM+
Sbjct: 10 ELPIGGYGCSVIQTIFNAINVMAGVGLLSTPYTVKEAGWASMVILLLFAVICCYTATLMK 69
Query: 142 HCFESREGLTSYPDIGEAAFGRYGRIFVSIILYTELYSYCVEFITLEGDNLTGLFPGTSL 201
CFE++ G+ +YPDIGEAAFG+YGRI + ++LYTELYSYCVEFI LEGDNLTGLFPGTSL
Sbjct: 70 DCFENKTGIITYPDIGEAAFGKYGRILICMLLYTELYSYCVEFIILEGDNLTGLFPGTSL 129
Query: 202 DIGGLHLDSMHLFGVLTALVILPTVWLKDLRVISYLSVGGIAATILIIISVFSVGTT--V 259
D+ G LDS HLFG+LTAL++LPTVWLKDLR+ISYLS GG+ AT LI +SVF +GTT +
Sbjct: 130 DLLGFRLDSKHLFGILTALIVLPTVWLKDLRIISYLSAGGVIATALIAVSVFFLGTTGGI 189
Query: 260 GFHHTGRVVNWSGIPFAIGVYGFCFAGHSVFPNIYQSMADKKQYTKALITCFVLCILIYG 319
GFHHTG+ V W+GIPFAIG+YGFC++GHSVFPNIYQSMADK ++ KA+ITCF++C+L+YG
Sbjct: 190 GFHHTGQAVKWNGIPFAIGIYGFCYSGHSVFPNIYQSMADKTKFNKAVITCFIICVLLYG 249
Query: 320 SVAVMGFLSFGDDTLSQITLNMPAGAFASKVALWTT 355
VA+MG+L FG+ TLSQITLNMP F SKVA WTT
Sbjct: 250 GVAIMGYLMFGEATLSQITLNMPQDQFFSKVAQWTT 285
>UniRef100_Q949Y8 Hypothetical protein At2g39130 [Arabidopsis thaliana]
Length = 550
Score = 323 bits (827), Expect = 6e-87
Identities = 145/275 (52%), Positives = 199/275 (71%), Gaps = 2/275 (0%)
Query: 83 EMPLGYGCSYTQTVFNGINVMAGVGLLSTPYTVKQAGWMGLVLMLIFASVCCYTATLMRH 142
E+P+ SY Q V NG+NV+ GVG+LSTPY K+ GW+GL+++ ++ + YT L+R+
Sbjct: 153 EIPMSRNSSYGQAVLNGLNVLCGVGILSTPYAAKEGGWLGLMILFVYGLLSFYTGILLRY 212
Query: 143 CFESREGLTSYPDIGEAAFGRYGRIFVSIILYTELYSYCVEFITLEGDNLTGLFPGTSLD 202
C +S L +YPDIG+AAFG GRIFVSI+LY ELY+ CVE+I LE DNL+ L+P +L
Sbjct: 213 CLDSESDLETYPDIGQAAFGTTGRIFVSIVLYLELYACCVEYIILEIDNLSSLYPNAALS 272
Query: 203 IGGLHLDSMHLFGVLTALVILPTVWLKDLRVISYLSVGGIAATILIIISVFSVGTT--VG 260
IGG LD+ HLF +LT L +LPTVWL+DL V+SY+S GG+ A++L+++ +F +G VG
Sbjct: 273 IGGFQLDARHLFALLTTLAVLPTVWLRDLSVLSYISAGGVIASVLVVLCLFWIGLVDEVG 332
Query: 261 FHHTGRVVNWSGIPFAIGVYGFCFAGHSVFPNIYQSMADKKQYTKALITCFVLCILIYGS 320
H G +N S +P AIG+YG+C++GH+VFPNIY SMA QY L+TCF +C L+Y
Sbjct: 333 IHSKGTTLNLSTLPVAIGLYGYCYSGHAVFPNIYTSMAKPSQYPAVLLTCFGICTLMYAG 392
Query: 321 VAVMGFLSFGDDTLSQITLNMPAGAFASKVALWTT 355
VAVMG+ FG+ T SQ TLN+P A+K+A+WTT
Sbjct: 393 VAVMGYTMFGESTQSQFTLNLPQDLIATKIAVWTT 427
>UniRef100_Q9SV32 Hypothetical protein F28P10.190 [Arabidopsis thaliana]
Length = 571
Score = 313 bits (803), Expect = 4e-84
Identities = 161/355 (45%), Positives = 225/355 (63%), Gaps = 14/355 (3%)
Query: 15 RETTDSYTIATAPNF-----VSILRGPSSIYSSFSNRSKSDLDIDGKT-PCLSGLDG--- 65
R++ D + +PN S+ R SS SS R + + T P L +
Sbjct: 63 RQSIDMFGSVPSPNLGFLANSSMSRRGSSFMSSTLTRRHTPESLPCVTKPLLEDEEAPKH 122
Query: 66 --TTQSTWWEKRSTQNLVG-EMPLGYGCSYTQTVFNGINVMAGVGLLSTPYTVKQAGWMG 122
+T S K S+ +V +M + S+ Q V NG+NV+ GVG+LSTPY VK+ GW+G
Sbjct: 123 KLSTHSLLPSKPSSSMVVSHDMGISNDSSFGQAVLNGVNVLCGVGILSTPYAVKEGGWLG 182
Query: 123 LVLMLIFASVCCYTATLMRHCFESREGLTSYPDIGEAAFGRYGRIFVSIILYTELYSYCV 182
L+++ F +C YT L+R+C +S + +YPDIG AAFG GRI VS+ILY ELY+ V
Sbjct: 183 LIILFAFGILCFYTGLLLRYCLDSHPDVQTYPDIGHAAFGSTGRILVSVILYMELYAMSV 242
Query: 183 EFITLEGDNLTGLFPGTSLDIGGLHLDSMHLFGVLTALVILPTVWLKDLRVISYLSVGGI 242
E+I LEGDNL+ +FP SL IGG HLD+ LF +LT L +LPTVWL+DL V+SY+S GG+
Sbjct: 243 EYIILEGDNLSSMFPNASLSIGGFHLDAPRLFALLTTLAVLPTVWLRDLSVLSYISAGGV 302
Query: 243 AATILIIISVFSVGTT--VGFHHTGRVVNWSGIPFAIGVYGFCFAGHSVFPNIYQSMADK 300
A++L+++ +F VG VG H G +N + +P ++G+YG+C++GH VFPNIY SMA
Sbjct: 303 IASVLVVLCLFWVGLVDDVGIHSKGTPLNLATLPVSVGLYGYCYSGHGVFPNIYTSMAKP 362
Query: 301 KQYTKALITCFVLCILIYGSVAVMGFLSFGDDTLSQITLNMPAGAFASKVALWTT 355
Q++ L+ F +C L+Y VAVMG+ FG+ T SQ TLN+P ASK+ALWTT
Sbjct: 363 SQFSAVLLASFGICTLMYAGVAVMGYSMFGESTESQFTLNLPQDLVASKIALWTT 417
>UniRef100_Q6YU97 Putative amino acid transport protein [Oryza sativa]
Length = 571
Score = 310 bits (795), Expect = 3e-83
Identities = 143/275 (52%), Positives = 193/275 (70%), Gaps = 2/275 (0%)
Query: 83 EMPLGYGCSYTQTVFNGINVMAGVGLLSTPYTVKQAGWMGLVLMLIFASVCCYTATLMRH 142
E+P CSYTQ V NGINV+ GVG+LSTPY +KQ GW+GLV++ +FA + YT L+R
Sbjct: 176 EVPAYQQCSYTQAVMNGINVLCGVGILSTPYAIKQGGWLGLVILCLFAVLAWYTGVLLRR 235
Query: 143 CFESREGLTSYPDIGEAAFGRYGRIFVSIILYTELYSYCVEFITLEGDNLTGLFPGTSLD 202
C +S+EGL +YPDIG AAFG GRI +SIILY ELY+ C+E++ LE DNL+ LFP L
Sbjct: 236 CLDSKEGLETYPDIGHAAFGTTGRIAISIILYIELYACCIEYLILESDNLSKLFPNAHLT 295
Query: 203 IGGLHLDSMHLFGVLTALVILPTVWLKDLRVISYLSVGGIAATILIIISVFSVGTT--VG 260
IG + L+S F +LT L+++PT WL+DL +SYLS GG+ A+IL+++ + VG VG
Sbjct: 296 IGSMTLNSHVFFAILTTLIVMPTTWLRDLSCLSYLSAGGVIASILVVVCLCWVGVVDHVG 355
Query: 261 FHHTGRVVNWSGIPFAIGVYGFCFAGHSVFPNIYQSMADKKQYTKALITCFVLCILIYGS 320
F + G +N GIP AIG+YG+C++GH VFPNIY S+ ++ Q+ L TC L +++
Sbjct: 356 FENKGTALNLPGIPIAIGLYGYCYSGHGVFPNIYSSLKNRNQFPSILFTCIGLSSILFAG 415
Query: 321 VAVMGFLSFGDDTLSQITLNMPAGAFASKVALWTT 355
AVMG+ FG+ T SQ TLN+P SKVA+WTT
Sbjct: 416 AAVMGYKMFGESTESQFTLNLPENLVVSKVAVWTT 450
>UniRef100_Q8GYS4 Hypothetical protein At5g02180/T7H20_230 [Arabidopsis thaliana]
Length = 550
Score = 306 bits (784), Expect = 6e-82
Identities = 164/354 (46%), Positives = 225/354 (63%), Gaps = 15/354 (4%)
Query: 15 RETTDSYTIATAPNFVSILRGPSS--IYSSFSNRSKSD----LDIDGKTPCLSGLDGTTQ 68
R++ D T T P+ VS + SS + SSF + +S L P LS +
Sbjct: 74 RQSMDLLTGMTPPS-VSFMPQSSSRRLASSFQKKQQSSFCDSLSSSSSKPLLSQPVPDKE 132
Query: 69 STWWEKRSTQNL---VGEMPLGYG--CSYTQTVFNGINVMAGVGLLSTPYTVKQAGWMGL 123
T L V ++PL CS++Q+V NG NV+ G+GL++ PY +K++GW+GL
Sbjct: 133 ETILPVNPQSQLKLSVTDLPLPEPNLCSFSQSVLNGTNVLCGLGLITMPYAIKESGWLGL 192
Query: 124 VLMLIFASVCCYTATLMRHCFESREGLTSYPDIGEAAFGRYGRIFVSIILYTELYSYCVE 183
++L F + CYT LM+ C ES G+ +YPDIG+AAFG GR +SI+LY ELY+ CVE
Sbjct: 193 PILLFFGVITCYTGVLMKRCLESSPGIQTYPDIGQAAFGITGRFIISILLYVELYAACVE 252
Query: 184 FITLEGDNLTGLFPGTSLDI-GGLHLDSMHLFGVLTALVILPTVWLKDLRVISYLSVGGI 242
+I + DNL+GLFP SL I G+ LDS +F +LT L++LPTVWLKDL ++SYLSVGG+
Sbjct: 253 YIIMMSDNLSGLFPNVSLSIASGISLDSPQIFAILTTLLVLPTVWLKDLSLLSYLSVGGV 312
Query: 243 AATILIIISVFSVGTT--VGFHHTGRVVNWSGIPFAIGVYGFCFAGHSVFPNIYQSMADK 300
A+IL+ I +F VG +GFH TGRV + S +P IG++GF ++GHSVFPNIY SM D
Sbjct: 313 LASILLGICLFWVGAVDGIGFHATGRVFDLSNLPVTIGIFGFGYSGHSVFPNIYSSMKDP 372
Query: 301 KQYTKALITCFVLCILIYGSVAVMGFLSFGDDTLSQITLNMPAGAFASKVALWT 354
++ L+ CF C ++Y +VAV G+ FG+ SQ TLNMP F SKVA+WT
Sbjct: 373 SRFPLVLVICFSFCTVLYIAVAVCGYTMFGEAVESQFTLNMPKHFFPSKVAVWT 426
>UniRef100_Q8LPF4 AT5g02170/T7H20_220 [Arabidopsis thaliana]
Length = 526
Score = 291 bits (746), Expect = 1e-77
Identities = 156/351 (44%), Positives = 220/351 (62%), Gaps = 13/351 (3%)
Query: 15 RETTDSYTIATAPNFVSIL-----RGPSSIYSSF-SNRSKSDLDIDGKTPCLSGLDGTTQ 68
R++ D T T P S + R SS++ SF S+ SK L ID K S + + +
Sbjct: 53 RQSMDLLTGVTPPTSTSFVSSFRQRRQSSVFGSFTSSPSKQQLLID-KDEIQSSVVSSIK 111
Query: 69 STWWEKRSTQNLVGEMPL---GYGCSYTQTVFNGINVMAGVGLLSTPYTVKQAGWMGLVL 125
S + ++ G++ C+++Q+V NGINV+ GV LL+ PY VK+ GW+GL +
Sbjct: 112 S-FLASHLQLSVPGDLLTPQENRSCTFSQSVLNGINVLCGVALLTMPYAVKEGGWLGLFI 170
Query: 126 MLIFASVCCYTATLMRHCFESREGLTSYPDIGEAAFGRYGRIFVSIILYTELYSYCVEFI 185
+ F + YT L++ C E+ G+ +YPDIG+AAFG GRI VSI+LY ELY+ CVE+I
Sbjct: 171 LFSFGIITFYTGILLKRCLENSPGIHTYPDIGQAAFGTTGRILVSILLYVELYASCVEYI 230
Query: 186 TLEGDNLTGLFPGTSLDIGGLHLDSMHLFGVLTALVILPTVWLKDLRVISYLSVGGIAAT 245
+ DNL+ +FP TSL I G LDS +F + T L++LPTVWLKDL ++SYLS GG+ ++
Sbjct: 231 IMMSDNLSRMFPNTSLYINGFSLDSTQVFAITTTLIVLPTVWLKDLSLLSYLSAGGVISS 290
Query: 246 ILIIISVFSVGTT--VGFHHTGRVVNWSGIPFAIGVYGFCFAGHSVFPNIYQSMADKKQY 303
IL+ + +F G+ VGFH +G+ ++ + IP AIG+YGF F HSVFPNIY SM + ++
Sbjct: 291 ILLALCLFWAGSVDGVGFHISGQALDITNIPVAIGIYGFGFGSHSVFPNIYSSMKEPSKF 350
Query: 304 TKALITCFVLCILIYGSVAVMGFLSFGDDTLSQITLNMPAGAFASKVALWT 354
L+ F C L Y +VAV GF FGD SQ TLNMP +SK+A+WT
Sbjct: 351 PTVLLISFAFCTLFYIAVAVCGFTMFGDAIQSQFTLNMPPHFTSSKIAVWT 401
>UniRef100_Q7XUV8 OSJNBa0072F16.7 protein [Oryza sativa]
Length = 455
Score = 289 bits (740), Expect = 7e-77
Identities = 131/271 (48%), Positives = 189/271 (69%), Gaps = 2/271 (0%)
Query: 87 GYGCSYTQTVFNGINVMAGVGLLSTPYTVKQAGWMGLVLMLIFASVCCYTATLMRHCFES 146
G G ++ +T FNG+N ++GVGLLS PY + + GW+ LVL+L A VCCYT L+R C +
Sbjct: 65 GDGATFVRTCFNGLNALSGVGLLSIPYALSEGGWLSLVLLLAVAMVCCYTGLLLRRCMAA 124
Query: 147 REGLTSYPDIGEAAFGRYGRIFVSIILYTELYSYCVEFITLEGDNLTGLFPGTSLDIGGL 206
+ YPDIG AFG GR+ VS LY ELY + F+ LEGDNL LFPGTSL +GGL
Sbjct: 125 SPAVRGYPDIGALAFGAKGRLAVSAFLYAELYLVAIGFLILEGDNLDKLFPGTSLAVGGL 184
Query: 207 HLDSMHLFGVLTALVILPTVWLKDLRVISYLSVGGIAATILIIISVF--SVGTTVGFHHT 264
+ LF V+ A+VILPT WL+ L V++Y+S G+ A+++++ V +V VGFH
Sbjct: 185 VVSGKQLFVVVVAVVILPTTWLRSLAVLAYVSASGVLASVVVVFCVLWAAVFDGVGFHGK 244
Query: 265 GRVVNWSGIPFAIGVYGFCFAGHSVFPNIYQSMADKKQYTKALITCFVLCILIYGSVAVM 324
GR++N SG+P A+G+Y FC+ GH++FP + SM +K ++++ L+ CFV C + YGS+A++
Sbjct: 245 GRMLNVSGLPTALGLYTFCYCGHAIFPTLCNSMQEKDKFSRVLVICFVACTVNYGSMAIL 304
Query: 325 GFLSFGDDTLSQITLNMPAGAFASKVALWTT 355
G+L +GDD SQ+TLN+P G +SK+A++TT
Sbjct: 305 GYLMYGDDVKSQVTLNLPEGKISSKLAIYTT 335
>UniRef100_Q9SR29 F3L24.21 protein [Arabidopsis thaliana]
Length = 478
Score = 283 bits (724), Expect = 5e-75
Identities = 152/357 (42%), Positives = 218/357 (60%), Gaps = 24/357 (6%)
Query: 15 RETTDSYTIATAPNFVSILRG-------PSSIYSSFSNR----SKSDLDIDGKTPCLSGL 63
R + D T T P VS ++G SSI S + R + S + K P LS
Sbjct: 51 RHSVDLLTGVTPP-MVSFIQGRSTETSFSSSIASLYKRRPTSIANSFVSSTSKQPLLSEK 109
Query: 64 DGTTQSTWWEKRSTQNLVGEMPLGYG----CSYTQTVFNGINVMAGVGLLSTPYTVKQAG 119
D + S+Q + L YG CS+ Q+V NGINV+ G+ LL+ PY VK+ G
Sbjct: 110 DDVSFL------SSQVGLSNTDLSYGEPNFCSFPQSVLNGINVLCGISLLTMPYAVKEGG 163
Query: 120 WMGLVLMLIFASVCCYTATLMRHCFESREGLTSYPDIGEAAFGRYGRIFVSIILYTELYS 179
W+GL ++L FA + CYT L++ C ES L +YPDIG+AAFG GR+ +SI+LY ELY
Sbjct: 164 WLGLCILLSFAIITCYTGILLKRCLESSSDLRTYPDIGQAAFGFTGRLIISILLYMELYV 223
Query: 180 YCVEFITLEGDNLTGLFPGTSLDIGGLHLDSMHLFGVLTALVILPTVWLKDLRVISYLSV 239
CVE+I + DNL+ +FP +L+I G+ LDS +F + L++LPTVWLKDL ++SYLS
Sbjct: 224 CCVEYIIMMSDNLSRVFPNITLNIVGVSLDSPQIFAISATLIVLPTVWLKDLSLLSYLSA 283
Query: 240 GGIAATILIIISVFSVGTT--VGFHHTGRVVNWSGIPFAIGVYGFCFAGHSVFPNIYQSM 297
GG+ +IL+ + +F VG+ VGFH G+ ++ + +P AIG++GF F+GH+V P+IY SM
Sbjct: 284 GGVFVSILLALCLFWVGSVDGVGFHTGGKALDLANLPVAIGIFGFGFSGHAVLPSIYSSM 343
Query: 298 ADKKQYTKALITCFVLCILIYGSVAVMGFLSFGDDTLSQITLNMPAGAFASKVALWT 354
+ ++ L+ F C+ Y +VA+ G+ FG+ SQ TLNMP ASK+A+WT
Sbjct: 344 KEPSKFPLVLLISFGFCVFFYIAVAICGYSMFGEAIQSQFTLNMPQQYTASKIAVWT 400
>UniRef100_Q9SR30 F3L24.20 protein [Arabidopsis thaliana]
Length = 481
Score = 282 bits (721), Expect = 1e-74
Identities = 148/348 (42%), Positives = 216/348 (61%), Gaps = 22/348 (6%)
Query: 13 YNRETTDSYTIATAPNFVSILRGPSSIYSSFSNRSKSDLDIDGKTPCLSGLDGTTQSTWW 72
+ R + S+T + A + R P+SI +SF++ + K P LS D
Sbjct: 69 HRRSSQTSFTSSVASLYK---RRPTSIANSFASSTS-------KQPLLSEKDDVLFL--- 115
Query: 73 EKRSTQNLVGEMPLGYG----CSYTQTVFNGINVMAGVGLLSTPYTVKQAGWMGLVLMLI 128
S+Q + L YG CS+ Q+V NGINV+ G+ LL+ PY VK+ GW+GL ++L
Sbjct: 116 ---SSQIGLSNTDLSYGEPNFCSFPQSVLNGINVLCGISLLTMPYAVKEGGWLGLCILLS 172
Query: 129 FASVCCYTATLMRHCFESREGLTSYPDIGEAAFGRYGRIFVSIILYTELYSYCVEFITLE 188
FA + CYT L++ C ES L +YPDIG+AAFG GR+ +SI+LY ELY CVE+I +
Sbjct: 173 FAIITCYTGILLKRCLESSSDLRTYPDIGQAAFGFTGRLIISILLYMELYVCCVEYIIMM 232
Query: 189 GDNLTGLFPGTSLDIGGLHLDSMHLFGVLTALVILPTVWLKDLRVISYLSVGGIAATILI 248
DNL+ +FP +L+I G+ LDS +F + L++LPTVWLKDL ++SYLS GG+ +IL+
Sbjct: 233 SDNLSRVFPNITLNIVGVSLDSPQIFAISATLIVLPTVWLKDLSLLSYLSAGGVFVSILL 292
Query: 249 IISVFSVGTT--VGFHHTGRVVNWSGIPFAIGVYGFCFAGHSVFPNIYQSMADKKQYTKA 306
+ +F VG+ VGFH G+ ++ + +P AIG++GF F+GH+V P+IY SM + ++
Sbjct: 293 ALCLFWVGSVDGVGFHTGGKSLDLANLPVAIGIFGFGFSGHAVLPSIYSSMKEPSKFPLV 352
Query: 307 LITCFVLCILIYGSVAVMGFLSFGDDTLSQITLNMPAGAFASKVALWT 354
L+ F C+ Y VA+ G+ FG+ SQ TLNMP ASK+A+WT
Sbjct: 353 LLISFGFCVFFYIVVAICGYSMFGEAIQSQFTLNMPQQYTASKIAVWT 400
>UniRef100_Q9LZL4 Hypothetical protein T7H20_230 [Arabidopsis thaliana]
Length = 543
Score = 278 bits (712), Expect = 1e-73
Identities = 155/354 (43%), Positives = 215/354 (59%), Gaps = 22/354 (6%)
Query: 15 RETTDSYTIATAPNFVSILRGPSS--IYSSFSNRSKSD----LDIDGKTPCLSGLDGTTQ 68
R++ D T T P+ VS + SS + SSF + +S L P LS +
Sbjct: 74 RQSMDLLTGMTPPS-VSFMPQSSSRRLASSFQKKQQSSFCDSLSSSSSKPLLSQPVPDKE 132
Query: 69 STWWEKRSTQNL---VGEMPLGYG--CSYTQTVFNGINVMAGVGLLSTPYTVKQAGWMGL 123
T L V ++PL CS++Q+V NG NV+ G+GL++ PY +K++GW+GL
Sbjct: 133 ETILPVNPQSQLKLSVTDLPLPEPNLCSFSQSVLNGTNVLCGLGLITMPYAIKESGWLGL 192
Query: 124 VLMLIFASVCCYTATLMRHCFESREGLTSYPDIGEAAFGRYGRIFVSIILYTELYSYCVE 183
++L F + CYT LM+ C ES G+ +YPDIG+AAFG ++ CVE
Sbjct: 193 PILLFFGVITCYTGVLMKRCLESSPGIQTYPDIGQAAFGITDSSIRGVVP-------CVE 245
Query: 184 FITLEGDNLTGLFPGTSLDIG-GLHLDSMHLFGVLTALVILPTVWLKDLRVISYLSVGGI 242
+I + DNL+GLFP SL I G+ LDS +F +LT L++LPTVWLKDL ++SYLSVGG+
Sbjct: 246 YIIMMSDNLSGLFPNVSLSIASGISLDSPQIFAILTTLLVLPTVWLKDLSLLSYLSVGGV 305
Query: 243 AATILIIISVFSVGTT--VGFHHTGRVVNWSGIPFAIGVYGFCFAGHSVFPNIYQSMADK 300
A+IL+ I +F VG +GFH TGRV + S +P IG++GF ++GHSVFPNIY SM D
Sbjct: 306 LASILLGICLFWVGAVDGIGFHATGRVFDLSNLPVTIGIFGFGYSGHSVFPNIYSSMKDP 365
Query: 301 KQYTKALITCFVLCILIYGSVAVMGFLSFGDDTLSQITLNMPAGAFASKVALWT 354
++ L+ CF C ++Y +VAV G+ FG+ SQ TLNMP F SKVA+WT
Sbjct: 366 SRFPLVLVICFSFCTVLYIAVAVCGYTMFGEAVESQFTLNMPKHFFPSKVAVWT 419
>UniRef100_Q8S8P2 Hypothetical protein At2g39130 [Arabidopsis thaliana]
Length = 381
Score = 269 bits (687), Expect = 1e-70
Identities = 120/229 (52%), Positives = 166/229 (72%), Gaps = 2/229 (0%)
Query: 83 EMPLGYGCSYTQTVFNGINVMAGVGLLSTPYTVKQAGWMGLVLMLIFASVCCYTATLMRH 142
E+P+ SY Q V NG+NV+ GVG+LSTPY K+ GW+GL+++ ++ + YT L+R+
Sbjct: 153 EIPMSRNSSYGQAVLNGLNVLCGVGILSTPYAAKEGGWLGLMILFVYGLLSFYTGILLRY 212
Query: 143 CFESREGLTSYPDIGEAAFGRYGRIFVSIILYTELYSYCVEFITLEGDNLTGLFPGTSLD 202
C +S L +YPDIG+AAFG GRIFVSI+LY ELY+ CVE+I LE DNL+ L+P +L
Sbjct: 213 CLDSESDLETYPDIGQAAFGTTGRIFVSIVLYLELYACCVEYIILESDNLSSLYPNAALS 272
Query: 203 IGGLHLDSMHLFGVLTALVILPTVWLKDLRVISYLSVGGIAATILIIISVFSVGTT--VG 260
IGG LD+ HLF +LT L +LPTVWL+DL V+SY+S GG+ A++L+++ +F +G VG
Sbjct: 273 IGGFQLDARHLFALLTTLAVLPTVWLRDLSVLSYISAGGVIASVLVVLCLFWIGLVDEVG 332
Query: 261 FHHTGRVVNWSGIPFAIGVYGFCFAGHSVFPNIYQSMADKKQYTKALIT 309
H G +N S +P AIG+YG+C++GH+VFPNIY SMA QY L+T
Sbjct: 333 IHSKGTTLNLSTLPVAIGLYGYCYSGHAVFPNIYTSMAKPSQYPAVLLT 381
>UniRef100_Q9LZL5 Hypothetical protein T7H20_220 [Arabidopsis thaliana]
Length = 516
Score = 264 bits (675), Expect = 2e-69
Identities = 149/351 (42%), Positives = 212/351 (59%), Gaps = 23/351 (6%)
Query: 15 RETTDSYTIATAPNFVSIL-----RGPSSIYSSF-SNRSKSDLDIDGKTPCLSGLDGTTQ 68
R++ D T T P S + R SS++ SF S+ SK L ID K S + + +
Sbjct: 53 RQSMDLLTGVTPPTSTSFVSSFRQRRQSSVFGSFTSSPSKQQLLID-KDEIQSSVVSSIK 111
Query: 69 STWWEKRSTQNLVGEMPL---GYGCSYTQTVFNGINVMAGVGLLSTPYTVKQAGWMGLVL 125
S + ++ G++ C+++Q+V NGINV+ GV LL+ PY VK+ GW+GL +
Sbjct: 112 S-FLASHLQLSVPGDLLTPQENRSCTFSQSVLNGINVLCGVALLTMPYAVKEGGWLGLFI 170
Query: 126 MLIFASVCCYTATLMRHCFESREGLTSYPDIGEAAFGRYGRIFVSIILYTELYSYCVEFI 185
+ F + YT L++ C E+ G+ +YPDIG+AAFG GRI VS + CVE+I
Sbjct: 171 LFSFGIITFYTGILLKRCLENSPGIHTYPDIGQAAFGTTGRILVS--------ASCVEYI 222
Query: 186 TLEGDNLTGLFPGTSLDIGGLHLDSMHLFGVLTALVILPTVWLKDLRVISYLSVGGIAAT 245
+ DNL+ +FP TSL I G LDS +F + T L++LPTVWLKDL ++SYLS G+ ++
Sbjct: 223 IMMSDNLSRMFPNTSLYINGFSLDSTQVFAITTTLIVLPTVWLKDLSLLSYLS--GVISS 280
Query: 246 ILIIISVFSVGTT--VGFHHTGRVVNWSGIPFAIGVYGFCFAGHSVFPNIYQSMADKKQY 303
IL+ + +F G+ VGFH +G+ ++ + IP AIG+YGF F HSVFPNIY SM + ++
Sbjct: 281 ILLALCLFWAGSVDGVGFHISGQALDITNIPVAIGIYGFGFGSHSVFPNIYSSMKEPSKF 340
Query: 304 TKALITCFVLCILIYGSVAVMGFLSFGDDTLSQITLNMPAGAFASKVALWT 354
L+ F C L Y +VAV GF FGD SQ TLNMP +SK+A+WT
Sbjct: 341 PTVLLISFAFCTLFYIAVAVCGFTMFGDAIQSQFTLNMPPHFTSSKIAVWT 391
>UniRef100_Q6K4N6 Amino acid transporter-like [Oryza sativa]
Length = 323
Score = 264 bits (675), Expect = 2e-69
Identities = 132/236 (55%), Positives = 171/236 (71%), Gaps = 14/236 (5%)
Query: 15 RETTDSYTIATAPNFVSILRGPSSIYSSFSNRSKSDLDIDGKTPCLSG-LDGTTQSTWWE 73
RETTD+YTIA +P+F + GPS+ S + +S L D K P LS LDG S
Sbjct: 75 RETTDTYTIAASPSFGYL--GPSTSKYSILDGGRSSLGSDLKLPLLSDKLDGKQDSVKSL 132
Query: 74 KRSTQNLV-----------GEMPLGYGCSYTQTVFNGINVMAGVGLLSTPYTVKQAGWMG 122
+++ + + GE+ + GCS TQTVFNG+NV+AGVGLLSTP+T+ +AGW+G
Sbjct: 133 RKTLGSAIDRKSSLLTQHTGEVYIAQGCSVTQTVFNGVNVLAGVGLLSTPFTIHEAGWVG 192
Query: 123 LVLMLIFASVCCYTATLMRHCFESREGLTSYPDIGEAAFGRYGRIFVSIILYTELYSYCV 182
L ++ +FA VCCYT LM+HCFES++G+++YPDIGEAAFGR GR+ +SIILYTELYSYCV
Sbjct: 193 LAVLAMFAIVCCYTGVLMKHCFESKDGISTYPDIGEAAFGRIGRLLISIILYTELYSYCV 252
Query: 183 EFITLEGDNLTGLFPGTSLDIGGLHLDSMHLFGVLTALVILPTVWLKDLRVISYLS 238
EFI LEGDN+T +F D G+H+D H FGVLTAL++LPTVWL+DLRV+SYLS
Sbjct: 253 EFIILEGDNMTSIFSHIGFDWLGVHIDGKHFFGVLTALIVLPTVWLRDLRVLSYLS 308
>UniRef100_Q9MBG9 Gb|AAC79623.2 [Arabidopsis thaliana]
Length = 429
Score = 260 bits (664), Expect = 5e-68
Identities = 119/267 (44%), Positives = 177/267 (65%), Gaps = 2/267 (0%)
Query: 91 SYTQTVFNGINVMAGVGLLSTPYTVKQAGWMGLVLMLIFASVCCYTATLMRHCFESREGL 150
S+ +T FN +N ++G+G+LS PY++ + GW+ L L+L+ A YT+ L+ C + +
Sbjct: 17 SFFKTCFNALNALSGIGILSVPYSLARGGWLSLSLLLLLAVTAFYTSLLITKCMNADRNI 76
Query: 151 TSYPDIGEAAFGRYGRIFVSIILYTELYSYCVEFITLEGDNLTGLFPGTSLDIGGLHLDS 210
+YPDIGE AFGR GRI VS+ ++ ELY F+ LEGDNL LFPG ++++ GL L+
Sbjct: 77 KTYPDIGERAFGRPGRIIVSVFMHLELYLVTTGFLILEGDNLHNLFPGFTIEMIGLRLNG 136
Query: 211 MHLFGVLTALVILPTVWLKDLRVISYLSVGGIAATILIIISVFSVGT--TVGFHHTGRVV 268
F A VI+PT+W +L V+SY+S+ G+ AT + + S+ +G +GFH G+++
Sbjct: 137 KQAFMATVAFVIMPTLWWDNLSVLSYVSMSGVLATTVTLGSISWIGAFDGIGFHQKGKLI 196
Query: 269 NWSGIPFAIGVYGFCFAGHSVFPNIYQSMADKKQYTKALITCFVLCILIYGSVAVMGFLS 328
NWSGIP A+ +Y FC+ H V P +Y SM K Q+ L+ CF+LC + Y S+AV+G+L
Sbjct: 197 NWSGIPTALSLYAFCYGAHPVLPTLYSSMKSKHQFNNVLLICFILCTIGYTSMAVLGYLM 256
Query: 329 FGDDTLSQITLNMPAGAFASKVALWTT 355
+G TLSQITLN+P +SKVA++TT
Sbjct: 257 YGSQTLSQITLNLPIHKTSSKVAIYTT 283
>UniRef100_Q5ZAF5 Amino acid transporter-like [Oryza sativa]
Length = 631
Score = 254 bits (648), Expect = 3e-66
Identities = 119/268 (44%), Positives = 172/268 (63%), Gaps = 1/268 (0%)
Query: 89 GCSYTQTVFNGINVMAGVGLLSTPYTVKQAGWMGLVLMLIFASVCCYTATLMRHCFESRE 148
G S+++T N N ++G+G+LS PY V Q GW+ L+L ++ +VC YT TL+ C +
Sbjct: 36 GASFSRTCLNLTNAVSGIGVLSMPYAVSQGGWLSLLLFVLVGAVCYYTGTLIERCMRADG 95
Query: 149 GLTSYPDIGEAAFGRYGRIFVSIILYTELYSYCVEFITLEGDNLTGLFPGTSLDIGGLHL 208
+ SYPDIG+ AFG GR V+ +Y ELY + F+ LEGDNL LFPG +++I G L
Sbjct: 96 SIASYPDIGQYAFGATGRRAVAFFMYVELYLVAISFLVLEGDNLDKLFPGATMEILGYQL 155
Query: 209 DSMHLFGVLTALVILPTVWLKDLRVISYLSVGGIAATILIIISVFSVGTT-VGFHHTGRV 267
LF VL A VILPT WLK+L +++Y+S G+ A++ + S+ G GFH
Sbjct: 156 HGKQLFIVLAAAVILPTTWLKNLGMLAYVSAAGLIASVALTASLIWAGVAETGFHRNSNT 215
Query: 268 VNWSGIPFAIGVYGFCFAGHSVFPNIYQSMADKKQYTKALITCFVLCILIYGSVAVMGFL 327
+N +GIP ++G+Y CF GH+VFP IY SM + K ++K L+ VLC L YG AV+G++
Sbjct: 216 LNLAGIPTSLGLYFVCFTGHAVFPTIYSSMKNSKHFSKVLLISSVLCSLNYGLTAVLGYM 275
Query: 328 SFGDDTLSQITLNMPAGAFASKVALWTT 355
+GDD SQ+TLN+P+G +K+A+ T
Sbjct: 276 IYGDDVQSQVTLNLPSGKLYTKIAIVMT 303
>UniRef100_Q67WJ7 Putative amino acid transport protein [Oryza sativa]
Length = 413
Score = 252 bits (644), Expect = 1e-65
Identities = 116/272 (42%), Positives = 175/272 (63%), Gaps = 6/272 (2%)
Query: 89 GCSYTQTVFNGINVMAGVGLLSTPYTVKQAGWMGLVLMLIFASVCCYTATLMRHCFESRE 148
G S+ +T FNG+N ++GVG+LS PY + Q GW+ L + + A++C YT L++ C +S
Sbjct: 10 GTSFLKTCFNGVNALSGVGILSMPYALSQGGWLSLAIFITIAAICFYTGILLQRCIDSSS 69
Query: 149 GLTSYPDIGEAAFGRYGRIFVSIILYTELYSYCVEFITLEGDNLTGLFPGTSLDIGGLHL 208
+ +YPDIGE AFGR GRI V+ +Y ELY ++F+ LEGDNL LFP S H
Sbjct: 70 LVKTYPDIGELAFGRKGRIAVAAFMYLELYLVAIDFLILEGDNLEKLFPNASF-FSSFHR 128
Query: 209 ---DSMHLFGVLTALVILPTVWLKDLRVISYLSVGGIAATILIIISVFSVGTT--VGFHH 263
+ F +L AL++LPT W + L +++Y+S+GG+ A+ +++ SV VG VGF
Sbjct: 129 IAGGTRQGFVLLFALLVLPTTWFRSLDLLAYVSLGGVLASAILVASVLWVGAADGVGFRE 188
Query: 264 TGRVVNWSGIPFAIGVYGFCFAGHSVFPNIYQSMADKKQYTKALITCFVLCILIYGSVAV 323
G V W G+P A+ +Y FCF+GH+VFP IY M +++ + L+ CF++C L YG + V
Sbjct: 189 GGVAVRWGGVPTAMSLYAFCFSGHAVFPMIYTGMRNRRMFPHVLLICFIICTLAYGVMGV 248
Query: 324 MGFLSFGDDTLSQITLNMPAGAFASKVALWTT 355
+G+L +G SQ+TLN+PA +S +A++TT
Sbjct: 249 IGYLMYGGSLRSQVTLNLPARKLSSSIAIYTT 280
>UniRef100_Q9LXF8 Hypothetical protein F8M21_130 [Arabidopsis thaliana]
Length = 423
Score = 250 bits (638), Expect = 5e-65
Identities = 118/268 (44%), Positives = 172/268 (64%), Gaps = 3/268 (1%)
Query: 91 SYTQTVFNGINVMAGVGLLSTPYTVKQAGWMGLVLMLIFASVCCYTATLMRHCFESREGL 150
S+++T F+GIN ++GVG+LS PY + GW+ L+++ A Y A L++ C E L
Sbjct: 34 SFSKTCFHGINALSGVGILSVPYALASGGWLSLIILFTVAITTFYCAILIKRCMEMDPLL 93
Query: 151 TSYPDIGEAAFGRYGRIFVSIILYTELYSYCVEFITLEGDNLTGLFPGTSLDIGGLHLDS 210
SYPDIG AFG GR+ VSI + ELY F+ LEGDNL LF L+ GL
Sbjct: 94 RSYPDIGYKAFGNTGRVIVSIFMNLELYLVATSFLILEGDNLNKLFSNVGLNFMGLEFQG 153
Query: 211 MHLFGVLTALVILPTVWLKDLRVISYLSVGGIAATILIIISVFSVGT--TVGF-HHTGRV 267
+F ++ AL+ILP+VWL ++R++SY+S G+ A+ +I+ S+FSVG VGF ++ V
Sbjct: 154 KQMFIIMVALIILPSVWLDNMRILSYVSASGVFASGVILASIFSVGAFEGVGFKNNDSEV 213
Query: 268 VNWSGIPFAIGVYGFCFAGHSVFPNIYQSMADKKQYTKALITCFVLCILIYGSVAVMGFL 327
+G+ ++ +Y FC+ H VFP +Y SM +K+Q++ +I CF +C IY SVAV+G+L
Sbjct: 214 FRLNGVATSVSLYAFCYCAHPVFPTLYTSMKNKRQFSNVMIICFTICTFIYASVAVLGYL 273
Query: 328 SFGDDTLSQITLNMPAGAFASKVALWTT 355
+G D SQITLN+P +SKVA+WTT
Sbjct: 274 MYGSDVESQITLNLPTDKLSSKVAIWTT 301
>UniRef100_Q8LQU5 Amino acid transporter protein-like [Oryza sativa]
Length = 443
Score = 236 bits (602), Expect = 7e-61
Identities = 112/269 (41%), Positives = 168/269 (61%), Gaps = 2/269 (0%)
Query: 89 GCSYTQTVFNGINVMAGVGLLSTPYTVKQAGWMGLVLMLIFASVCCYTATLMRHCFESRE 148
G S+ ++ N NV++G+G+LS PY + Q GW+ L L + ++C YT L+ C +
Sbjct: 45 GASFGRSCLNLSNVISGIGMLSVPYALSQGGWLSLALFAMVGAICFYTGKLIYRCMRADR 104
Query: 149 GLTSYPDIGEAAFGRYGRIFVSIILYTELYSYCVEFITLEGDNLTGLFPGTSLDIGGLHL 208
+ SYPDIG AFGRYGR + +I+Y ELY + F+ LEGDNL L PGT + I G +
Sbjct: 105 CVRSYPDIGYLAFGRYGRTAIGLIMYVELYLVAISFLILEGDNLDKLLPGTVVKILGYQV 164
Query: 209 DSMHLFGVLTALVILPTVWLKDLRVISYLSVGGIAATILIIISVFSVGTT-VGFHHTG-R 266
LF ++ A VILPT WLK+L +++Y+S G+ +++ + +S+ G GFH G
Sbjct: 165 HGKQLFMLVAAAVILPTTWLKNLSMLAYVSAVGLVSSVALTVSLVWAGVADKGFHMAGSS 224
Query: 267 VVNWSGIPFAIGVYGFCFAGHSVFPNIYQSMADKKQYTKALITCFVLCILIYGSVAVMGF 326
++N SG+P A+ +Y CFAGH VFP +Y SM +K + K L+ VLC L Y AV+G+
Sbjct: 225 ILNLSGLPTALSLYFVCFAGHGVFPTVYSSMRARKDFPKVLLISSVLCSLNYAVTAVLGY 284
Query: 327 LSFGDDTLSQITLNMPAGAFASKVALWTT 355
+G+D +Q+TLN+P G +++A+ TT
Sbjct: 285 KIYGEDVQAQVTLNLPTGKLYTRIAILTT 313
>UniRef100_Q7XUV9 OSJNBa0072F16.6 protein [Oryza sativa]
Length = 397
Score = 229 bits (583), Expect = 1e-58
Identities = 116/277 (41%), Positives = 171/277 (60%), Gaps = 9/277 (3%)
Query: 84 MPLGYGCSYTQTVFNGINVMAGVGLLSTPYTVKQAGWMGLVLMLIFASVCCYTATLMRHC 143
MP+G +T NG+N ++GVGLL+ PY + + GW+ L L+ A+ C YT L+ C
Sbjct: 1 MPVGTA----RTCMNGLNALSGVGLLTVPYALSEGGWVSLALLAAVAAACWYTGILLCRC 56
Query: 144 FESREGLTSYPDIGEAAFGRYGRIFVSIILYTELYSYCVEFITLEGDNLTGLFPGTSLDI 203
++ + + +YPDIGE AFGR GR+ VS Y ELY F+ LEGDNL LFPG + +
Sbjct: 57 MDADDAIRTYPDIGERAFGRTGRLLVSAFTYVELYLVATGFLILEGDNLDKLFPGARVTL 116
Query: 204 GGLHLDSMHLFGVLTALVILPTVWLKDLRVISYLSVGGIAATILIIISVFSVGTTVGFHH 263
G + L LF VL ALV+ PT WL+ L V++Y+S G+ A+++I++SV G
Sbjct: 117 GTVSLAGKRLFVVLVALVVAPTTWLRSLGVLAYVSATGVFASVVIVLSVLWAAAVDGVGF 176
Query: 264 TGR----VVNWSGIPFAIGVYGFCFAGHSVFPNIYQSMADKKQYTKAL-ITCFVLCILIY 318
+GR + +G+P A+G+Y FC+ GH +FP +Y SM K Q+ K T I +
Sbjct: 177 SGRGTTTPLRIAGLPTALGLYIFCYGGHPMFPTLYTSMKRKSQFPKVYPYTHAHNTIDML 236
Query: 319 GSVAVMGFLSFGDDTLSQITLNMPAGAFASKVALWTT 355
++AV+G+L +GD LSQ+TLN+P+ +SKVA++TT
Sbjct: 237 PAMAVLGYLMYGDGVLSQVTLNLPSARLSSKVAIYTT 273
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.325 0.140 0.433
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 591,603,707
Number of Sequences: 2790947
Number of extensions: 24867795
Number of successful extensions: 73594
Number of sequences better than 10.0: 672
Number of HSP's better than 10.0 without gapping: 189
Number of HSP's successfully gapped in prelim test: 483
Number of HSP's that attempted gapping in prelim test: 72739
Number of HSP's gapped (non-prelim): 765
length of query: 359
length of database: 848,049,833
effective HSP length: 128
effective length of query: 231
effective length of database: 490,808,617
effective search space: 113376790527
effective search space used: 113376790527
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 75 (33.5 bits)
Medicago: description of AC146631.5


