

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146584.2 - phase: 0 /pseudo
(1452 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
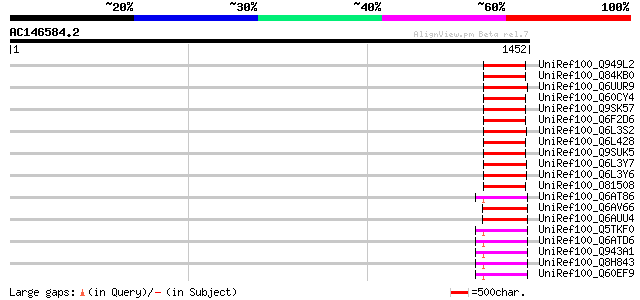
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q949L2 Putative polyprotein [Cicer arietinum] 161 2e-37
UniRef100_Q84KB0 Pol protein [Cucumis melo] 139 8e-31
UniRef100_Q6UUR9 Putative gag-pol protein [Oryza sativa] 120 3e-25
UniRef100_Q60CY4 Putative polyprotein [Solanum tuberosum] 119 5e-25
UniRef100_Q9SK57 Putative retroelement pol polyprotein [Arabidop... 119 5e-25
UniRef100_Q6F2D6 Putative polyprotein [Solanum demissum] 119 7e-25
UniRef100_Q6L3S2 Putative retrotransposon protein [Solanum demis... 119 9e-25
UniRef100_Q6L428 Putative integrase [Solanum demissum] 117 2e-24
UniRef100_Q9SUK5 RETROTRANSPOSON like protein [Arabidopsis thali... 115 8e-24
UniRef100_Q6L3Y7 Putative gag-pol polyprotein [Solanum demissum] 115 1e-23
UniRef100_Q6L3Y6 Putative gag-pol polyprotein [Solanum demissum] 115 1e-23
UniRef100_O81508 T7M24.4 protein [Arabidopsis thaliana] 114 3e-23
UniRef100_Q6AT86 Putative polyprotein [Oryza sativa] 105 1e-20
UniRef100_Q6AV66 Hypothetical protein OSJNBa0035N15.2 [Oryza sat... 104 2e-20
UniRef100_Q6AUU4 Putative pol polyprotein [Oryza sativa] 104 2e-20
UniRef100_Q5TKF0 Hypothetical protein OSJNBa0030I14.16 [Oryza sa... 104 2e-20
UniRef100_Q6ATD6 Putative polyprotein [Oryza sativa] 103 3e-20
UniRef100_Q943A1 Putative polyprotein [Oryza sativa] 103 4e-20
UniRef100_Q8H843 Putative polyprotein [Oryza sativa] 103 4e-20
UniRef100_Q60EF9 Putative polyprotein [Oryza sativa] 103 4e-20
>UniRef100_Q949L2 Putative polyprotein [Cicer arietinum]
Length = 141
Score = 161 bits (407), Expect = 2e-37
Identities = 75/117 (64%), Positives = 95/117 (81%), Gaps = 1/117 (0%)
Query: 1326 KEVDPKFIGPYQILERVGTVAYRVGLPPHLSNLHNVFHVSQLRKYVPDPSHVIQSDDVQV 1385
K++ PKFIGPYQI R+G VAYR+ LPP LSN+H+VFHVSQLRKY+ DPSHVI+ D +Q+
Sbjct: 23 KKLTPKFIGPYQISARIGPVAYRIALPPILSNIHDVFHVSQLRKYLADPSHVIEPDTIQL 82
Query: 1386 RDNLTVETLPVRIDDRKVKTLRGKEIPLVRVVWTGATGESLTWELESKMLESYPELF 1442
+DNL+ E PVRIDD K+K LR K++ LV+V+W TG++ TWELESKM E +PELF
Sbjct: 83 KDNLSFEVPPVRIDDMKIKRLRTKDVSLVKVIWNPITGDA-TWELESKMREQHPELF 138
>UniRef100_Q84KB0 Pol protein [Cucumis melo]
Length = 923
Score = 139 bits (349), Expect = 8e-31
Identities = 63/116 (54%), Positives = 84/116 (72%)
Query: 1327 EVDPKFIGPYQILERVGTVAYRVGLPPHLSNLHNVFHVSQLRKYVPDPSHVIQSDDVQVR 1386
++ P+F+GP++ILER+G VAYR+ LPP LS +H+VFHVS LRKYVPDPSHV+ + +++
Sbjct: 806 KLSPRFVGPFEILERIGPVAYRLALPPSLSTVHDVFHVSMLRKYVPDPSHVVDYEPLEID 865
Query: 1387 DNLTVETLPVRIDDRKVKTLRGKEIPLVRVVWTGATGESLTWELESKMLESYPELF 1442
+NL+ PV + R VKTLR K+IPLV+V+W E TWE E M YPELF
Sbjct: 866 ENLSYVEQPVEVLARGVKTLRNKQIPLVKVLWRNHRVEEATWEREDDMRSRYPELF 921
>UniRef100_Q6UUR9 Putative gag-pol protein [Oryza sativa]
Length = 552
Score = 120 bits (301), Expect = 3e-25
Identities = 55/121 (45%), Positives = 80/121 (65%)
Query: 1327 EVDPKFIGPYQILERVGTVAYRVGLPPHLSNLHNVFHVSQLRKYVPDPSHVIQSDDVQVR 1386
++ P++IGP+ I RVG++AYR+ LP ++ +H+VFHVS LRKY+ DP H I + + V
Sbjct: 430 KLSPRYIGPFVITARVGSLAYRLQLPESMNGVHDVFHVSMLRKYLRDPEHKIDLEPIMVE 489
Query: 1387 DNLTVETLPVRIDDRKVKTLRGKEIPLVRVVWTGATGESLTWELESKMLESYPELFA*GK 1446
+LT+E PVRI D + +R + I V+V+WT + TWELE +M + YPELF G
Sbjct: 490 QDLTIECRPVRIFDWSERVMRRRTIKYVKVLWTNQSEREATWELEEEMKKKYPELFVDGM 549
Query: 1447 F 1447
F
Sbjct: 550 F 550
>UniRef100_Q60CY4 Putative polyprotein [Solanum tuberosum]
Length = 394
Score = 119 bits (299), Expect = 5e-25
Identities = 58/116 (50%), Positives = 77/116 (66%)
Query: 1327 EVDPKFIGPYQILERVGTVAYRVGLPPHLSNLHNVFHVSQLRKYVPDPSHVIQSDDVQVR 1386
++ P++IGP++IL VG VAY + LPP S +H VFHVS LR+YVPD SHV+Q D V++
Sbjct: 273 KLSPRYIGPFEILRTVGEVAYELALPPVFSAIHPVFHVSMLRRYVPDESHVLQYDAVELD 332
Query: 1387 DNLTVETLPVRIDDRKVKTLRGKEIPLVRVVWTGATGESLTWELESKMLESYPELF 1442
D LT PV I R V+ LR + IP+V+V W + E TWE E +M E +P LF
Sbjct: 333 DRLTFVEEPVAILARDVRRLRSRAIPVVKVRWRHCSVEEATWETEQEMREQFPGLF 388
>UniRef100_Q9SK57 Putative retroelement pol polyprotein [Arabidopsis thaliana]
Length = 1611
Score = 119 bits (299), Expect = 5e-25
Identities = 55/117 (47%), Positives = 79/117 (67%)
Query: 1326 KEVDPKFIGPYQILERVGTVAYRVGLPPHLSNLHNVFHVSQLRKYVPDPSHVIQSDDVQV 1385
K++ P+++GPY+++ERVG VAY++ LPP L+ HNVFHVSQLRK + D ++ +
Sbjct: 1466 KKLSPRYVGPYKVIERVGAVAYKLDLPPKLNAFHNVFHVSQLRKCLSDQEESVEDIPPGL 1525
Query: 1386 RDNLTVETLPVRIDDRKVKTLRGKEIPLVRVVWTGATGESLTWELESKMLESYPELF 1442
++N+TVE PVRI DR K RGK L++V+W E TWE E+KM ++PE F
Sbjct: 1526 KENMTVEAWPVRIMDRMTKGTRGKARDLLKVLWNCRGREEYTWETENKMKANFPEWF 1582
>UniRef100_Q6F2D6 Putative polyprotein [Solanum demissum]
Length = 1769
Score = 119 bits (298), Expect = 7e-25
Identities = 58/116 (50%), Positives = 76/116 (65%)
Query: 1327 EVDPKFIGPYQILERVGTVAYRVGLPPHLSNLHNVFHVSQLRKYVPDPSHVIQSDDVQVR 1386
++ P++IGP++IL VG VAY + LPP S +H VFHVS LR+YVPD SHV+Q D V++
Sbjct: 1648 KLSPRYIGPFEILRTVGEVAYELALPPVFSAIHPVFHVSMLRRYVPDESHVLQYDAVELD 1707
Query: 1387 DNLTVETLPVRIDDRKVKTLRGKEIPLVRVVWTGATGESLTWELESKMLESYPELF 1442
D LT PV I R V+ LR + IP+V+V W E TWE E +M E +P LF
Sbjct: 1708 DRLTFVEEPVAILARDVRRLRSRAIPVVKVRWRHRPVEEATWETEQEMREQFPSLF 1763
>UniRef100_Q6L3S2 Putative retrotransposon protein [Solanum demissum]
Length = 1602
Score = 119 bits (297), Expect = 9e-25
Identities = 53/119 (44%), Positives = 84/119 (70%)
Query: 1327 EVDPKFIGPYQILERVGTVAYRVGLPPHLSNLHNVFHVSQLRKYVPDPSHVIQSDDVQVR 1386
++ P++IGPY+I++RVG+VAY + LP L+ +H VFH+S L+K + DPS ++ ++ V+++
Sbjct: 1480 KLSPRYIGPYRIVQRVGSVAYELELPQELAAVHPVFHISMLKKCIGDPSLILPTESVKIK 1539
Query: 1387 DNLTVETLPVRIDDRKVKTLRGKEIPLVRVVWTGATGESLTWELESKMLESYPELFA*G 1445
DNL+ E +PV+I DR+V+ LR K++ V+V+W E TWE E M + YP LF G
Sbjct: 1540 DNLSYEEVPVQILDRQVRRLRTKDVASVKVLWRNQFVEEATWEAEEDMKKRYPHLFESG 1598
>UniRef100_Q6L428 Putative integrase [Solanum demissum]
Length = 1609
Score = 117 bits (294), Expect = 2e-24
Identities = 57/116 (49%), Positives = 75/116 (64%)
Query: 1327 EVDPKFIGPYQILERVGTVAYRVGLPPHLSNLHNVFHVSQLRKYVPDPSHVIQSDDVQVR 1386
++ P++IGP++IL VG VAY + LPP S +H VFHV LR+YVPD SHV+Q D V++
Sbjct: 1347 KLSPRYIGPFEILRTVGEVAYELALPPAFSAIHPVFHVPMLRRYVPDESHVLQYDAVELD 1406
Query: 1387 DNLTVETLPVRIDDRKVKTLRGKEIPLVRVVWTGATGESLTWELESKMLESYPELF 1442
D LT P+ I R VK LR + IP+V+V W E TWE E +M E +P LF
Sbjct: 1407 DRLTFVEEPIAILARDVKRLRSRAIPVVKVHWRHRPVEEATWETEQEMREQFPVLF 1462
>UniRef100_Q9SUK5 RETROTRANSPOSON like protein [Arabidopsis thaliana]
Length = 687
Score = 115 bits (289), Expect = 8e-24
Identities = 52/117 (44%), Positives = 79/117 (67%)
Query: 1326 KEVDPKFIGPYQILERVGTVAYRVGLPPHLSNLHNVFHVSQLRKYVPDPSHVIQSDDVQV 1385
K++ P+++GPY+++ERVG VAY++ LPP L HNVFHVSQLRK + + ++ +
Sbjct: 550 KKLRPRYVGPYKVIERVGAVAYKLDLPPKLDAFHNVFHVSQLRKCLSEQEESMEDVPPGL 609
Query: 1386 RDNLTVETLPVRIDDRKVKTLRGKEIPLVRVVWTGATGESLTWELESKMLESYPELF 1442
++N+TVE PVRI D+ K RGK + L++++W E TWE E+KM ++PE F
Sbjct: 610 KENMTVEAWPVRIMDQMKKGTRGKSMDLLKILWNCGGREEYTWETETKMKANFPEWF 666
>UniRef100_Q6L3Y7 Putative gag-pol polyprotein [Solanum demissum]
Length = 1351
Score = 115 bits (287), Expect = 1e-23
Identities = 51/119 (42%), Positives = 81/119 (67%)
Query: 1327 EVDPKFIGPYQILERVGTVAYRVGLPPHLSNLHNVFHVSQLRKYVPDPSHVIQSDDVQVR 1386
++ P++IGPY+I +R+G VAY + LP L+ +H VFH+S L+K + DPS ++ ++ +++
Sbjct: 1229 KLSPRYIGPYRIAKRIGNVAYELELPQELAAVHPVFHISMLKKCIGDPSLILPTESIKIN 1288
Query: 1387 DNLTVETLPVRIDDRKVKTLRGKEIPLVRVVWTGATGESLTWELESKMLESYPELFA*G 1445
DNL+ E +PV+I DR+V+ LR K++ V+V+W E TWE E M + YP LF G
Sbjct: 1289 DNLSYEEVPVQILDRQVRRLRTKDVASVKVLWRDQFVEEATWEAEEDMKKKYPYLFESG 1347
>UniRef100_Q6L3Y6 Putative gag-pol polyprotein [Solanum demissum]
Length = 1351
Score = 115 bits (287), Expect = 1e-23
Identities = 51/119 (42%), Positives = 81/119 (67%)
Query: 1327 EVDPKFIGPYQILERVGTVAYRVGLPPHLSNLHNVFHVSQLRKYVPDPSHVIQSDDVQVR 1386
++ P++IGPY+I +R+G VAY + LP L+ +H VFH+S L+K + DPS ++ ++ +++
Sbjct: 1229 KLSPRYIGPYRIAKRIGNVAYELELPQELAAVHPVFHISMLKKCIGDPSLILPTESIKIN 1288
Query: 1387 DNLTVETLPVRIDDRKVKTLRGKEIPLVRVVWTGATGESLTWELESKMLESYPELFA*G 1445
DNL+ E +PV+I DR+V+ LR K++ V+V+W E TWE E M + YP LF G
Sbjct: 1289 DNLSYEEVPVQILDRQVRRLRTKDVASVKVLWRDQFVEEATWEAEEDMKKKYPYLFESG 1347
>UniRef100_O81508 T7M24.4 protein [Arabidopsis thaliana]
Length = 973
Score = 114 bits (284), Expect = 3e-23
Identities = 52/117 (44%), Positives = 78/117 (66%)
Query: 1326 KEVDPKFIGPYQILERVGTVAYRVGLPPHLSNLHNVFHVSQLRKYVPDPSHVIQSDDVQV 1385
K++ P+++GPY+++ERVG VAY++ LPP L+ HNVFHVSQLRK + + ++ +
Sbjct: 828 KKLSPRYVGPYKVIERVGAVAYKLDLPPKLNAFHNVFHVSQLRKCLSNQEESVEDVPPGL 887
Query: 1386 RDNLTVETLPVRIDDRKVKTLRGKEIPLVRVVWTGATGESLTWELESKMLESYPELF 1442
++N+TVE PV+I DR K RGK L++V+W E TWE E+KM ++ E F
Sbjct: 888 KENMTVEAWPVQIMDRMTKGTRGKSRDLLKVLWNCGGREQYTWETENKMKANFSEWF 944
>UniRef100_Q6AT86 Putative polyprotein [Oryza sativa]
Length = 1501
Score = 105 bits (261), Expect = 1e-20
Identities = 57/154 (37%), Positives = 89/154 (57%), Gaps = 8/154 (5%)
Query: 1304 SRRRPRVFESHSYDRCRTC-------FEVK-EVDPKFIGPYQILERVGTVAYRVGLPPHL 1355
+RRR FE+ Y R F+ K ++ P+F+GPY+ILER G VAY++ LPP++
Sbjct: 1343 NRRRELTFEAGDYVYLRVTPLRGVHRFQTKGKLAPRFVGPYRILERRGEVAYQLELPPNM 1402
Query: 1356 SNLHNVFHVSQLRKYVPDPSHVIQSDDVQVRDNLTVETLPVRIDDRKVKTLRGKEIPLVR 1415
+H+VFHVSQL+K + P S+ + ++++LT P+RI + + R K I +
Sbjct: 1403 VGIHDVFHVSQLKKCLRVPEEQASSEHIDIQEDLTYVEKPIRILETSERRTRNKVIRFCK 1462
Query: 1416 VVWTGATGESLTWELESKMLESYPELFA*GKFSR 1449
V W+ + E TWE E ++ ++P LFA SR
Sbjct: 1463 VQWSHHSEEEATWEREDELKAAHPHLFANSSESR 1496
>UniRef100_Q6AV66 Hypothetical protein OSJNBa0035N15.2 [Oryza sativa]
Length = 1169
Score = 104 bits (260), Expect = 2e-20
Identities = 52/128 (40%), Positives = 81/128 (62%), Gaps = 1/128 (0%)
Query: 1323 FEVK-EVDPKFIGPYQILERVGTVAYRVGLPPHLSNLHNVFHVSQLRKYVPDPSHVIQSD 1381
F+ K ++ P+F+GP++I+ R G VAY++ LP L N+H+VFHVSQL+K + PS S+
Sbjct: 1037 FQTKGKLAPRFVGPFRIIARRGEVAYQLELPASLGNVHDVFHVSQLKKCLRVPSEQADSE 1096
Query: 1382 DVQVRDNLTVETLPVRIDDRKVKTLRGKEIPLVRVVWTGATGESLTWELESKMLESYPEL 1441
++VR++LT E PV+I D + R + I +V W+ E TWE + ++ +YP+L
Sbjct: 1097 QIEVREDLTYEERPVKILDTMERRTRNRVIRFCKVQWSNHAEEEATWERQDELKAAYPDL 1156
Query: 1442 FA*GKFSR 1449
FA SR
Sbjct: 1157 FASSSESR 1164
>UniRef100_Q6AUU4 Putative pol polyprotein [Oryza sativa]
Length = 922
Score = 104 bits (260), Expect = 2e-20
Identities = 52/128 (40%), Positives = 81/128 (62%), Gaps = 1/128 (0%)
Query: 1323 FEVK-EVDPKFIGPYQILERVGTVAYRVGLPPHLSNLHNVFHVSQLRKYVPDPSHVIQSD 1381
F+ K ++ P+F+GP++I+ R G VAY++ LP L N+H+VFHVSQL+K + PS S+
Sbjct: 790 FQTKGKLAPRFVGPFRIIARRGEVAYQLELPASLGNVHDVFHVSQLKKCLRVPSEQADSE 849
Query: 1382 DVQVRDNLTVETLPVRIDDRKVKTLRGKEIPLVRVVWTGATGESLTWELESKMLESYPEL 1441
++VR++LT E PV+I D + R + I +V W+ E TWE + ++ +YP+L
Sbjct: 850 QIEVREDLTYEERPVKILDTMERRTRNRVIRFCKVQWSNHAEEEATWERQDELKAAYPDL 909
Query: 1442 FA*GKFSR 1449
FA SR
Sbjct: 910 FASSSESR 917
>UniRef100_Q5TKF0 Hypothetical protein OSJNBa0030I14.16 [Oryza sativa]
Length = 1253
Score = 104 bits (259), Expect = 2e-20
Identities = 57/154 (37%), Positives = 88/154 (57%), Gaps = 8/154 (5%)
Query: 1304 SRRRPRVFESHSYDRCRTC-------FEVK-EVDPKFIGPYQILERVGTVAYRVGLPPHL 1355
+RRR VF++ Y R F+ K ++ P+F+GPY+ILER G VAY++ LP ++
Sbjct: 1095 NRRRELVFQAGDYVYLRVTPLRGVHRFQTKGKLAPRFVGPYRILERRGEVAYQLELPSNM 1154
Query: 1356 SNLHNVFHVSQLRKYVPDPSHVIQSDDVQVRDNLTVETLPVRIDDRKVKTLRGKEIPLVR 1415
+HNVFHVSQL+K + P D ++++++LT P RI + + R K I +
Sbjct: 1155 LGIHNVFHVSQLKKCLRVPEEQASPDQIEIQEDLTYVEKPTRILETSERRTRNKVIRFCK 1214
Query: 1416 VVWTGATGESLTWELESKMLESYPELFA*GKFSR 1449
V W+ + E TWE E ++ ++P LFA SR
Sbjct: 1215 VQWSHLSKEEATWEREDELKAAHPHLFASSSQSR 1248
>UniRef100_Q6ATD6 Putative polyprotein [Oryza sativa]
Length = 1524
Score = 103 bits (258), Expect = 3e-20
Identities = 58/154 (37%), Positives = 87/154 (55%), Gaps = 8/154 (5%)
Query: 1304 SRRRPRVFESHSYDRCRTC-------FEVK-EVDPKFIGPYQILERVGTVAYRVGLPPHL 1355
+RRR FE+ Y R F+ K ++ P+F+GPY+ILER G VAY++ LP ++
Sbjct: 1366 NRRRELAFETGDYVYLRVTPLRGVHRFQKKGKLAPRFVGPYRILERRGEVAYQLELPSNM 1425
Query: 1356 SNLHNVFHVSQLRKYVPDPSHVIQSDDVQVRDNLTVETLPVRIDDRKVKTLRGKEIPLVR 1415
+HNVFHVSQL+K + P D ++++++LT P RI + + R K I +
Sbjct: 1426 LGIHNVFHVSQLKKCLRVPEEQASPDHIEIQEDLTYVEKPTRILETNERRTRNKVIRFCK 1485
Query: 1416 VVWTGATGESLTWELESKMLESYPELFA*GKFSR 1449
V W+ T E TWE E ++ ++P LFA SR
Sbjct: 1486 VQWSHHTEEEATWEREDELKATHPHLFASSSESR 1519
>UniRef100_Q943A1 Putative polyprotein [Oryza sativa]
Length = 1524
Score = 103 bits (257), Expect = 4e-20
Identities = 57/154 (37%), Positives = 87/154 (56%), Gaps = 8/154 (5%)
Query: 1304 SRRRPRVFESHSYDRCRTC-------FEVK-EVDPKFIGPYQILERVGTVAYRVGLPPHL 1355
+RRR FE+ Y R F+ K ++ P+F+GPY+ILER G VAY++ LP ++
Sbjct: 1366 NRRRELAFETGDYVYLRVTPLRGVHRFQTKGKLAPRFVGPYRILERRGEVAYQLELPSNM 1425
Query: 1356 SNLHNVFHVSQLRKYVPDPSHVIQSDDVQVRDNLTVETLPVRIDDRKVKTLRGKEIPLVR 1415
+HN+FHVSQL+K + P D ++++++LT P RI + + R K I +
Sbjct: 1426 LGIHNMFHVSQLKKCLRVPEEQASPDHIEIQEDLTYVEKPTRILETSERRTRNKVIRFCK 1485
Query: 1416 VVWTGATGESLTWELESKMLESYPELFA*GKFSR 1449
V W+ T E TWE E ++ ++P LFA SR
Sbjct: 1486 VQWSHHTEEEATWEREDELKATHPHLFASSSESR 1519
>UniRef100_Q8H843 Putative polyprotein [Oryza sativa]
Length = 1796
Score = 103 bits (257), Expect = 4e-20
Identities = 58/154 (37%), Positives = 88/154 (56%), Gaps = 8/154 (5%)
Query: 1304 SRRRPRVFESHSYDRCRTC-------FEVK-EVDPKFIGPYQILERVGTVAYRVGLPPHL 1355
+R+R FE+ Y R F+ K ++ P+F+GPY+ILER G VAY++ LP ++
Sbjct: 1638 NRQRELTFEAGDYVYLRVTPLRGVHRFQTKGKLAPRFVGPYKILERRGEVAYQLELPSNM 1697
Query: 1356 SNLHNVFHVSQLRKYVPDPSHVIQSDDVQVRDNLTVETLPVRIDDRKVKTLRGKEIPLVR 1415
+H+VFHVSQL+K + P S+ + ++++LT PVRI D + R K I R
Sbjct: 1698 IGIHDVFHVSQLKKCLRVPEEQADSEHIDIQEDLTYVEKPVRILDTSERRTRNKVIRFCR 1757
Query: 1416 VVWTGATGESLTWELESKMLESYPELFA*GKFSR 1449
V W+ + E TWE E ++ ++P LFA SR
Sbjct: 1758 VQWSHHSEEEATWEREDELKAAHPHLFASSSESR 1791
>UniRef100_Q60EF9 Putative polyprotein [Oryza sativa]
Length = 1493
Score = 103 bits (257), Expect = 4e-20
Identities = 57/154 (37%), Positives = 87/154 (56%), Gaps = 8/154 (5%)
Query: 1304 SRRRPRVFESHSYDRCRTC-------FEVK-EVDPKFIGPYQILERVGTVAYRVGLPPHL 1355
+RRR FE+ Y R F+ K ++ P+F+GPY+ILER G VAY++ LP ++
Sbjct: 1335 NRRRELAFETGDYVYLRVTPLRGVHRFQTKGKLAPRFVGPYRILERRGEVAYQLELPSNM 1394
Query: 1356 SNLHNVFHVSQLRKYVPDPSHVIQSDDVQVRDNLTVETLPVRIDDRKVKTLRGKEIPLVR 1415
+HNVFHVSQL+K + P D ++++++LT P RI + + R K I +
Sbjct: 1395 LGIHNVFHVSQLKKCLRVPEEQASPDHIEIQEDLTYVEKPTRIPETSERRTRNKVIRFCK 1454
Query: 1416 VVWTGATGESLTWELESKMLESYPELFA*GKFSR 1449
V W+ + E TWE E ++ ++P LFA SR
Sbjct: 1455 VQWSHHSEEEATWEREDELKATHPYLFASSSESR 1488
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.365 0.164 0.591
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,816,814,110
Number of Sequences: 2790947
Number of extensions: 60597693
Number of successful extensions: 303095
Number of sequences better than 10.0: 506
Number of HSP's better than 10.0 without gapping: 418
Number of HSP's successfully gapped in prelim test: 88
Number of HSP's that attempted gapping in prelim test: 301332
Number of HSP's gapped (non-prelim): 851
length of query: 1452
length of database: 848,049,833
effective HSP length: 140
effective length of query: 1312
effective length of database: 457,317,253
effective search space: 600000235936
effective search space used: 600000235936
T: 11
A: 40
X1: 14 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.6 bits)
S2: 81 (35.8 bits)
Medicago: description of AC146584.2


