

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146575.4 - phase: 0
(165 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
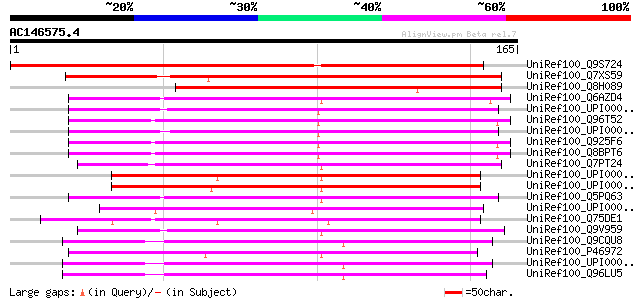
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9S724 Putative mitochondrial inner membrane protease ... 207 8e-53
UniRef100_Q7XS59 OSJNBa0019G23.8 protein [Oryza sativa] 157 7e-38
UniRef100_Q8H089 Putative signal peptidase [Oryza sativa] 136 2e-31
UniRef100_Q6AZD4 Zgc:100888 protein [Brachydanio rerio] 112 3e-24
UniRef100_UPI0000364AEB UPI0000364AEB UniRef100 entry 111 6e-24
UniRef100_Q96T52 Inner mitochondrial membrane peptidase 2 [Homo ... 111 6e-24
UniRef100_UPI0000292EBB UPI0000292EBB UniRef100 entry 110 1e-23
UniRef100_Q925F6 Inner mitochondrial membrane peptidase 2 [Mus m... 110 2e-23
UniRef100_Q8BPT6 Mus musculus 0 day neonate eyeball cDNA, RIKEN ... 109 2e-23
UniRef100_Q7PT24 ENSANGP00000018428 [Anopheles gambiae str. PEST] 106 3e-22
UniRef100_UPI000042BFD7 UPI000042BFD7 UniRef100 entry 105 4e-22
UniRef100_UPI000042CE63 UPI000042CE63 UniRef100 entry 105 6e-22
UniRef100_Q5PQ63 Hypothetical protein [Xenopus laevis] 104 1e-21
UniRef100_UPI000042C6B4 UPI000042C6B4 UniRef100 entry 103 1e-21
UniRef100_Q75DE1 ABR086Wp [Ashbya gossypii] 102 5e-21
UniRef100_Q9V959 CG11110-PA [Drosophila melanogaster] 101 8e-21
UniRef100_Q9CQU8 Mus musculus adult male testis cDNA, RIKEN full... 100 2e-20
UniRef100_P46972 Mitochondrial inner membrane protease subunit 2... 99 5e-20
UniRef100_UPI0000025FFF UPI0000025FFF UniRef100 entry 98 9e-20
UniRef100_Q96LU5 Hypothetical protein FLJ25059 [Homo sapiens] 97 2e-19
>UniRef100_Q9S724 Putative mitochondrial inner membrane protease subunit 2
[Arabidopsis thaliana]
Length = 154
Score = 207 bits (527), Expect = 8e-53
Identities = 91/154 (59%), Positives = 122/154 (79%), Gaps = 2/154 (1%)
Query: 1 MGTRNLVWNVTKKLATIGLITFTVSDRYATVVPVRGASMSPTFNPKTNSFTDDYVFVEKL 60
MG +N++W V KK T +I T+SDR +VVPVRG SMSPTFNP+ NS+ DDYV V+K
Sbjct: 1 MGIQNILWQVAKKSFTGSIIGLTISDRCCSVVPVRGDSMSPTFNPQRNSYLDDYVLVDKF 60
Query: 61 CLDKFKFSHGDIVIFSSPSNFKETHIKRIIALPGEWFVNRHNQDVLKVPEGHCWVEGDNA 120
CL +KF+ GD+V+FSSP++F + +IKRI+ +PGEW + ++DV++VPEGHCWVEGDN
Sbjct: 61 CLKDYKFARGDVVVFSSPTHFGDRYIKRIVGMPGEWISS--SRDVIRVPEGHCWVEGDNK 118
Query: 121 ASSTDSKSYGPVPLGLVRGRVTHVVWPPQRIGAV 154
SS DS+S+GP+PLGL++GRVT V+WPPQRI +
Sbjct: 119 TSSLDSRSFGPIPLGLIQGRVTRVMWPPQRISKI 152
>UniRef100_Q7XS59 OSJNBa0019G23.8 protein [Oryza sativa]
Length = 164
Score = 157 bits (398), Expect = 7e-38
Identities = 73/143 (51%), Positives = 99/143 (69%), Gaps = 5/143 (3%)
Query: 19 LITFTVSDRYATVVPVRGASMSPTFNPKTNSFTDDYVFVEKLCLD-KFKFSHGDIVIFSS 77
L+ TV+DRYA+V+ VRG SM+PT + D V +LCLD ++ S GD+V+F S
Sbjct: 20 LVVVTVNDRYASVITVRGTSMNPTLESQQG----DRALVSRLCLDARYGLSRGDVVVFRS 75
Query: 78 PSNFKETHIKRIIALPGEWFVNRHNQDVLKVPEGHCWVEGDNAASSTDSKSYGPVPLGLV 137
P+ + +KR+IALPG+W Q++ ++P GHCWVEGDN S DS+SYGP+PLGL+
Sbjct: 76 PTEHRSLLVKRLIALPGDWIQVPAAQEIRQIPVGHCWVEGDNPDVSWDSRSYGPIPLGLM 135
Query: 138 RGRVTHVVWPPQRIGAVKNTTPE 160
+GRVTH+VWPP RIG V+ PE
Sbjct: 136 QGRVTHIVWPPNRIGPVERKMPE 158
>UniRef100_Q8H089 Putative signal peptidase [Oryza sativa]
Length = 152
Score = 136 bits (342), Expect = 2e-31
Identities = 62/131 (47%), Positives = 81/131 (61%), Gaps = 25/131 (19%)
Query: 55 VFVEKLCLDKFKFSHGDIVIFSSPSNFKETHIKRIIALPGEWFVNRHNQDVLKVPEGHCW 114
V E+ CL K++FSHGD+V+F PS+ +E +KR+IALPGEW D++K+PEGHCW
Sbjct: 16 VLAERSCLQKYQFSHGDVVLFKCPSDHRELFVKRLIALPGEWMQLPGTPDIIKIPEGHCW 75
Query: 115 VEGDNAASSTDSKSYGP-------------------------VPLGLVRGRVTHVVWPPQ 149
VEGDNAA S DS+S+GP +PLGL++GRV HV+WPP
Sbjct: 76 VEGDNAACSWDSRSFGPEVDGIKDSMGGVRVSSASGMIGPPRIPLGLIKGRVAHVIWPPS 135
Query: 150 RIGAVKNTTPE 160
+IG V PE
Sbjct: 136 KIGRVDTKMPE 146
>UniRef100_Q6AZD4 Zgc:100888 protein [Brachydanio rerio]
Length = 183
Score = 112 bits (280), Expect = 3e-24
Identities = 58/146 (39%), Positives = 86/146 (58%), Gaps = 3/146 (2%)
Query: 20 ITFTVSDRYATVVPVRGASMSPTFNPKTNSFTDDYVFVEKLCLDKFKFSHGDIVIFSSPS 79
+T TV DR A V V GASM P+ NP S + D V + + + + GDIV SP
Sbjct: 24 VTVTVLDRLAYVARVEGASMQPSLNPDGES-SPDVVLLNRWSVRNYHVQRGDIVSVLSPK 82
Query: 80 NFKETHIKRIIALPGEWFVNR-HNQDVLKVPEGHCWVEGDNAASSTDSKSYGPVPLGLVR 138
N ++ IKR+I + G++ + ++VP+GH W+EGD+ S DS ++GPV LGLV
Sbjct: 83 NPQQKIIKRVIGIEGDFIKTLGYKNRYVRVPDGHLWIEGDHHGHSFDSNAFGPVSLGLVH 142
Query: 139 GRVTHVVWPPQRIGAVK-NTTPERLP 163
GR +H++WPP R ++ + P+R P
Sbjct: 143 GRASHIIWPPSRWQRIEPSVPPDRRP 168
>UniRef100_UPI0000364AEB UPI0000364AEB UniRef100 entry
Length = 174
Score = 111 bits (278), Expect = 6e-24
Identities = 54/141 (38%), Positives = 84/141 (59%), Gaps = 3/141 (2%)
Query: 20 ITFTVSDRYATVVPVRGASMSPTFNPKTNSFTDDYVFVEKLCLDKFKFSHGDIVIFSSPS 79
++ TV DR+A V V GASM P+ NP+ D V + + + + GDIV SP
Sbjct: 25 VSLTVFDRFACVARVEGASMQPSLNPEAGP--GDVVLLNRWSVRNHQVQRGDIVSVLSPK 82
Query: 80 NFKETHIKRIIALPGEWFVN-RHNQDVLKVPEGHCWVEGDNAASSTDSKSYGPVPLGLVR 138
N ++ IKR+I L G++ + +++P+GH W+EGD+ S DS S+GPV +GL+
Sbjct: 83 NPQQKIIKRVIGLEGDFIRTLSYKNRYVRIPDGHFWIEGDHHGHSMDSNSFGPVSVGLLH 142
Query: 139 GRVTHVVWPPQRIGAVKNTTP 159
GR +H++WPP+R +K + P
Sbjct: 143 GRASHIIWPPKRWQRIKASLP 163
>UniRef100_Q96T52 Inner mitochondrial membrane peptidase 2 [Homo sapiens]
Length = 175
Score = 111 bits (278), Expect = 6e-24
Identities = 63/146 (43%), Positives = 83/146 (56%), Gaps = 3/146 (2%)
Query: 20 ITFTVSDRYATVVPVRGASMSPTFNPKTNSFTDDYVFVEKLCLDKFKFSHGDIVIFSSPS 79
+ T DR A V V GASM P+ NP S + D V + + F+ GDIV SP
Sbjct: 25 VAVTFLDRVACVARVEGASMQPSLNPG-GSQSSDVVLLNHWKVRNFEVHRGDIVSLVSPK 83
Query: 80 NFKETHIKRIIALPGEWFVN-RHNQDVLKVPEGHCWVEGDNAASSTDSKSYGPVPLGLVR 138
N ++ IKR+IAL G+ H +KVP GH WVEGD+ S DS S+GPV LGL+
Sbjct: 84 NPEQKIIKRVIALEGDIVRTIGHKNRYVKVPRGHIWVEGDHHGHSFDSNSFGPVSLGLLH 143
Query: 139 GRVTHVVWPPQRIGAVKNT-TPERLP 163
TH++WPP+R +++ PERLP
Sbjct: 144 AHATHILWPPERWQKLESVLPPERLP 169
>UniRef100_UPI0000292EBB UPI0000292EBB UniRef100 entry
Length = 173
Score = 110 bits (276), Expect = 1e-23
Identities = 55/141 (39%), Positives = 82/141 (58%), Gaps = 4/141 (2%)
Query: 20 ITFTVSDRYATVVPVRGASMSPTFNPKTNSFTDDYVFVEKLCLDKFKFSHGDIVIFSSPS 79
+T TV DR A V V GASM P+ NP+ D V + + + + GDIV SP
Sbjct: 25 VTLTVFDRVACVARVEGASMQPSLNPEVPG---DVVLLNRWSVRNHQVQRGDIVSVLSPK 81
Query: 80 NFKETHIKRIIALPGEWFVN-RHNQDVLKVPEGHCWVEGDNAASSTDSKSYGPVPLGLVR 138
N ++ IKR+I L G++ + +++PEGH W+EGD+ S DS ++GPV +GL+
Sbjct: 82 NPQQKIIKRVIGLEGDFIRTLSYKNRYVRIPEGHFWIEGDHHGHSLDSNNFGPVSVGLLH 141
Query: 139 GRVTHVVWPPQRIGAVKNTTP 159
GR +H++WPP R +K + P
Sbjct: 142 GRASHIIWPPSRWQRIKASLP 162
>UniRef100_Q925F6 Inner mitochondrial membrane peptidase 2 [Mus musculus]
Length = 175
Score = 110 bits (274), Expect = 2e-23
Identities = 61/146 (41%), Positives = 83/146 (56%), Gaps = 3/146 (2%)
Query: 20 ITFTVSDRYATVVPVRGASMSPTFNPKTNSFTDDYVFVEKLCLDKFKFSHGDIVIFSSPS 79
+ T DR A V V G+SM P+ NP S + D V + + F+ GDIV SP
Sbjct: 25 VAVTFLDRVACVARVEGSSMQPSLNPG-GSQSSDVVLLNHWKVRNFEVQRGDIVSLVSPK 83
Query: 80 NFKETHIKRIIALPGEWFVN-RHNQDVLKVPEGHCWVEGDNAASSTDSKSYGPVPLGLVR 138
N ++ IKR+IAL G+ H ++KVP GH WVEGD+ S DS S+GPV LGL+
Sbjct: 84 NPEQKIIKRVIALEGDIVRTIGHKNGLVKVPRGHMWVEGDHHGHSFDSNSFGPVSLGLLH 143
Query: 139 GRVTHVVWPPQRIGAVKNT-TPERLP 163
TH++WPP+R +++ PER P
Sbjct: 144 AHATHILWPPERWQRLESVLPPERCP 169
>UniRef100_Q8BPT6 Mus musculus 0 day neonate eyeball cDNA, RIKEN full-length enriched
library, clone:E130011P11 product:inner mitochondrial
membrane peptidase 2-like (S. cerevisiae), full insert
sequence [Mus musculus]
Length = 175
Score = 109 bits (273), Expect = 2e-23
Identities = 61/146 (41%), Positives = 83/146 (56%), Gaps = 3/146 (2%)
Query: 20 ITFTVSDRYATVVPVRGASMSPTFNPKTNSFTDDYVFVEKLCLDKFKFSHGDIVIFSSPS 79
+ T DR A V V G+SM P+ NP S + D V + + F+ GDIV SP
Sbjct: 25 VAVTFLDRVACVARVEGSSMQPSLNPG-GSQSSDVVLLNHWKVRNFEVQRGDIVSLVSPK 83
Query: 80 NFKETHIKRIIALPGEWFVN-RHNQDVLKVPEGHCWVEGDNAASSTDSKSYGPVPLGLVR 138
N ++ IKR+IAL G+ H ++KVP GH WVEGD+ S DS S+GPV LGL+
Sbjct: 84 NPEQKIIKRVIALEGDIVRTIGHKNRLVKVPRGHMWVEGDHHGHSFDSNSFGPVSLGLLH 143
Query: 139 GRVTHVVWPPQRIGAVKNT-TPERLP 163
TH++WPP+R +++ PER P
Sbjct: 144 AHATHILWPPERWQRLESVLPPERCP 169
>UniRef100_Q7PT24 ENSANGP00000018428 [Anopheles gambiae str. PEST]
Length = 171
Score = 106 bits (264), Expect = 3e-22
Identities = 55/139 (39%), Positives = 77/139 (54%), Gaps = 3/139 (2%)
Query: 23 TVSDRYATVVPVRGASMSPTFNPKTNSFTDDYVFVEKLCLDKFKFSHGDIVIFSSPSNFK 82
T+ D V V G SM P NP ++ DYVF+ + + GDI+ SP +
Sbjct: 19 TLLDCVGYVARVEGVSMQPALNP--DATVTDYVFLSRWAVRNMDVQRGDIISLISPKDPT 76
Query: 83 ETHIKRIIALPGEWFVNR-HNQDVLKVPEGHCWVEGDNAASSTDSKSYGPVPLGLVRGRV 141
+ IKR++AL G+ + + VPEGHCWVEGD+ +S DS ++GPV LGLV R
Sbjct: 77 QKIIKRVVALQGDVISTLGYKLPYVTVPEGHCWVEGDHTGNSLDSNTFGPVSLGLVTARA 136
Query: 142 THVVWPPQRIGAVKNTTPE 160
T +VWPP R + +T P+
Sbjct: 137 TQIVWPPSRWQQLPSTVPK 155
>UniRef100_UPI000042BFD7 UPI000042BFD7 UniRef100 entry
Length = 162
Score = 105 bits (262), Expect = 4e-22
Identities = 51/124 (41%), Positives = 78/124 (62%), Gaps = 4/124 (3%)
Query: 34 VRGASMSPTFNPKTNSFTDDYVFVEKLCLDKFK-FSHGDIVIFSSPSNFKETHIKRIIAL 92
+ G+SM+PTFNP T++ T D V V+K + K + S GDI++F SP N ++ KR++ +
Sbjct: 33 ITGSSMTPTFNPGTSTMTKDIVLVQKYNIKKPRSLSRGDIIMFRSPENPEKLLTKRVVGI 92
Query: 93 PGEWFVNR---HNQDVLKVPEGHCWVEGDNAASSTDSKSYGPVPLGLVRGRVTHVVWPPQ 149
G+ + + + +K+P H WVEGDN+ S DS +GPV GLV G+V ++WPP
Sbjct: 93 QGDIIRPKSPPYPKSEVKIPRNHFWVEGDNSFHSIDSNKFGPVSQGLVIGKVVTIIWPPS 152
Query: 150 RIGA 153
R G+
Sbjct: 153 RFGS 156
>UniRef100_UPI000042CE63 UPI000042CE63 UniRef100 entry
Length = 162
Score = 105 bits (261), Expect = 6e-22
Identities = 51/124 (41%), Positives = 77/124 (61%), Gaps = 4/124 (3%)
Query: 34 VRGASMSPTFNPKTNSFTDDYVFVEKLCLDK-FKFSHGDIVIFSSPSNFKETHIKRIIAL 92
+ G+SM+PTFNP T++ T D V V+K + K S GDI++F SP N ++ KR++ +
Sbjct: 33 ITGSSMTPTFNPGTSTMTKDIVLVQKYNIKKPGSLSRGDIIMFRSPENPEKLLTKRVVGI 92
Query: 93 PGEWFVNR---HNQDVLKVPEGHCWVEGDNAASSTDSKSYGPVPLGLVRGRVTHVVWPPQ 149
G+ + + + +K+P H WVEGDN+ S DS +GPV GLV G+V ++WPP
Sbjct: 93 QGDIIRPKSPPYPKSEVKIPRNHFWVEGDNSFHSIDSNKFGPVSQGLVIGKVVTIIWPPS 152
Query: 150 RIGA 153
R G+
Sbjct: 153 RFGS 156
>UniRef100_Q5PQ63 Hypothetical protein [Xenopus laevis]
Length = 170
Score = 104 bits (259), Expect = 1e-21
Identities = 56/141 (39%), Positives = 74/141 (51%), Gaps = 2/141 (1%)
Query: 20 ITFTVSDRYATVVPVRGASMSPTFNPKTNSFTDDYVFVEKLCLDKFKFSHGDIVIFSSPS 79
+T T DR A + V G SM P+ NP D V + + + GDIV SP
Sbjct: 22 VTVTFLDRVACIARVEGVSMQPSLNPDARG-ESDIVLLNRWRARNYDVQRGDIVSLVSPK 80
Query: 80 NFKETHIKRIIALPGEWFVNR-HNQDVLKVPEGHCWVEGDNAASSTDSKSYGPVPLGLVR 138
N ++ IKR+IAL G+ H +KVP GH WVEGD+ S DS ++GPV LGL+
Sbjct: 81 NPEQKIIKRVIALEGDIVKTLGHKNRYVKVPRGHVWVEGDHHGHSFDSNAFGPVSLGLLH 140
Query: 139 GRVTHVVWPPQRIGAVKNTTP 159
TH++WPP R +K P
Sbjct: 141 SHATHILWPPNRWQKLKPFLP 161
>UniRef100_UPI000042C6B4 UPI000042C6B4 UniRef100 entry
Length = 187
Score = 103 bits (258), Expect = 1e-21
Identities = 53/129 (41%), Positives = 75/129 (58%), Gaps = 4/129 (3%)
Query: 30 TVVPVRGASMSPTFNPK--TNSFTDDYVFVEKLCLDKFKFSHGDIVIFSSPSNFKETHIK 87
++ V G SM PTFNP TN +D V +E+ K+ GD+V SP N + K
Sbjct: 40 SLATVTGGSMQPTFNPDLATNPLHNDVVLLERWSPAMNKYKRGDVVTLWSPQNPQLLTTK 99
Query: 88 RIIALPGEWF--VNRHNQDVLKVPEGHCWVEGDNAASSTDSKSYGPVPLGLVRGRVTHVV 145
RI+AL G+ + +++P GHCWVEGD+ + DS +YGP+PLGL+ RV+H++
Sbjct: 100 RIVALEGDLVHPLPPSPPTPVRIPPGHCWVEGDSKYQTRDSNTYGPIPLGLITARVSHII 159
Query: 146 WPPQRIGAV 154
WP R G V
Sbjct: 160 WPWARAGEV 168
>UniRef100_Q75DE1 ABR086Wp [Ashbya gossypii]
Length = 168
Score = 102 bits (253), Expect = 5e-21
Identities = 62/153 (40%), Positives = 79/153 (51%), Gaps = 11/153 (7%)
Query: 11 TKKLATIGLITFTVSDRYATVV-------PVRGASMSPTFNPKTNSFTDDYVFVEKLCLD 63
T +LA L+ T Y TV V G SM PT NP + D+V V KL
Sbjct: 3 TSRLAKCSLVAVTWLPVYMTVTHHVVFISKVEGPSMRPTLNPM-DGVASDWVLVWKLGKT 61
Query: 64 KFK-FSHGDIVIFSSPSNFKETHIKRIIALPGEWFVNRHN--QDVLKVPEGHCWVEGDNA 120
+ +HGD+VIF SP N K+ + KRI + R+ + +VP+ H WVEGDN
Sbjct: 62 NIRNLNHGDVVIFRSPMNPKKVYCKRIQGKQYDTVRTRYPYPKSTCEVPKSHIWVEGDNV 121
Query: 121 ASSTDSKSYGPVPLGLVRGRVTHVVWPPQRIGA 153
S DS +GP+ GLV G VT V+WPP R GA
Sbjct: 122 TQSVDSNHFGPISTGLVVGEVTRVIWPPSRWGA 154
>UniRef100_Q9V959 CG11110-PA [Drosophila melanogaster]
Length = 171
Score = 101 bits (251), Expect = 8e-21
Identities = 53/140 (37%), Positives = 76/140 (53%), Gaps = 3/140 (2%)
Query: 23 TVSDRYATVVPVRGASMSPTFNPKTNSFTDDYVFVEKLCLDKFKFSHGDIVIFSSPSNFK 82
T D V V G SM P NP + DYVF+ + + GDI+ SP +
Sbjct: 19 TFLDCVGYVARVDGISMQPALNPVPDE--KDYVFLLRWGTHNSQVERGDIISLISPKDPA 76
Query: 83 ETHIKRIIALPGEWFVNR-HNQDVLKVPEGHCWVEGDNAASSTDSKSYGPVPLGLVRGRV 141
+ IKR++ L G+ + ++++VPEGHCWVEGD+ S DS ++GPV LGL+ R
Sbjct: 77 QKIIKRVVGLQGDVVSTLGYKHEIVRVPEGHCWVEGDHTGHSMDSNTFGPVALGLMSARA 136
Query: 142 THVVWPPQRIGAVKNTTPER 161
+VWPP+R ++N P R
Sbjct: 137 VAIVWPPERWRILENELPRR 156
>UniRef100_Q9CQU8 Mus musculus adult male testis cDNA, RIKEN full-length enriched
library, clone:4930535E08 product:DJ1137O17.1 (SIMILAR
TO PUTATIVE MITOCHONDRIAL INNER MEMBRANE PROTEASE
SUBNUNIT 2) homolog (1500034J20Rik protein) (Mus
musculus adult male cerebellum cDNA, RIKEN full-length
enriched library, clone:1500034J20 product:DJ1137O17.1
(SIMILAR TO PUTATIVE MITOCHONDRIAL INNER MEMBRANE
PROTEASE SUBNUNIT 2) homolog) (Mus musculus adult male
tongue cDNA, RIKEN full-length enriched library,
clone:2310032F17 product:DJ1137O17.1 (SIMILAR TO
PUTATIVE MITOCHONDRIAL INNER MEMBRANE PROTEASE SUBNUNIT
2) homolog) (Mus musculus adult male tongue cDNA, RIKEN
full-length enriched library, clone:2310066K04
product:DJ1137O17.1 (SIMILAR TO PUTATIVE MITOCHONDRIAL
INNER MEMBRANE PROTEASE SUBNUNIT 2) homolog) (Mus
musculus 10 days embryo whole body cDNA, RIKEN
full-length enriched library, clone:2610002D17
product:DJ1137O17.1 (SIMILAR TO PUTATIVE MITOCHONDRIAL
INNER MEMBRANE PROTEASE SUBNUNIT 2) homolog) (Mus
musculus 10 days embryo whole body cDNA, RIKEN
full-length enriched library, clone:2610011N11
product:DJ1137O17.1 (SIMILAR TO PUTATIVE MITOCHONDRIAL
INNER MEMBRANE PROTEASE SUBNUNIT 2) homolog) (Mus
musculus 10 days embryo whole body cDNA, RIKEN
full-length enriched library, clone:2610016I19
product:DJ1137O17.1 (SIMILAR TO PUTATIVE MITOCHONDRIAL
INNER MEMBRANE PROTEASE SUBNUNIT 2) homolog) (Mus
musculus 10 days embryo whole body cDNA, RIKEN
full-length enriched library, clone:2610019J16
product:DJ1137O17.1 (SIMILAR TO PUTATIVE MITOCHONDRIAL
INNER MEMBRANE PROTEASE SUBNUNIT 2) homolog) (Mus
musculus 10 days embryo whole body cDNA, RIKEN
full-length enriched library, clone:2610029L18
product:DJ1137O17.1 (SIMILAR TO PUTATIVE MITOCHONDRIAL
INNER MEMBRANE PROTEASE SUBNUNIT 2) homolog) (Mus
musculus 10 days embryo whole body cDNA, RIKEN
full-length enriched library, clone:2610204A17
product:DJ1137O17.1 (SIMILAR TO PUTATIVE MITOCHONDRIAL
INNER MEMBRANE PROTEASE SUBNUNIT 2) homolog) (Mus
musculus 10 days embryo whole body cDNA, RIKEN
full-length enriched library, clone:2610528O17
product:DJ1137O17.1 (SIMILAR TO PUTATIVE MITOCHONDRIAL
INNER MEMBRANE PROTEASE SUBNUNIT 2) homolog) [Mus
musculus]
Length = 166
Score = 99.8 bits (247), Expect = 2e-20
Identities = 56/144 (38%), Positives = 74/144 (50%), Gaps = 10/144 (6%)
Query: 18 GLITFTVSDRYATVVPVRGASMSPTFNPKTNSFTDDYVFVEKLCLDKFKFSHGDIVIFSS 77
G I + VV G SM PT D VF E L + GDIVI S
Sbjct: 20 GCIAHCAFEYVGGVVMCSGPSMEPTIQ------NSDIVFAENLSRHFYGIQRGDIVIAKS 73
Query: 78 PSNFKETHIKRIIALPGEWFVNRHNQDVLK----VPEGHCWVEGDNAASSTDSKSYGPVP 133
PS+ K KR+I L G+ ++ DV K VP GH W+EGDN +STDS+ YGP+P
Sbjct: 74 PSDPKSNICKRVIGLEGDKILSTSPSDVFKSRSYVPTGHVWLEGDNLQNSTDSRYYGPIP 133
Query: 134 LGLVRGRVTHVVWPPQRIGAVKNT 157
GL+RGR+ +WP G ++++
Sbjct: 134 YGLIRGRIFFKIWPFSDFGFLRDS 157
>UniRef100_P46972 Mitochondrial inner membrane protease subunit 2 [Saccharomyces
cerevisiae]
Length = 177
Score = 98.6 bits (244), Expect = 5e-20
Identities = 47/136 (34%), Positives = 78/136 (56%), Gaps = 3/136 (2%)
Query: 20 ITFTVSDRYATVVPVRGASMSPTFNPKTNSFTDDYVFVEKLCL-DKFKFSHGDIVIFSSP 78
+ T+++ + V+G SM PT NP+T + D+V + K + + S DI++F +P
Sbjct: 23 VLLTINNNVVHIAQVKGTSMQPTLNPQTETLATDWVLLWKFGVKNPSNLSRDDIILFKAP 82
Query: 79 SNFKETHIKRIIALPGEWFVNR--HNQDVLKVPEGHCWVEGDNAASSTDSKSYGPVPLGL 136
+N ++ + KR+ LP + + + + + +P GH WVEGDN S DS ++GP+ GL
Sbjct: 83 TNPRKVYCKRVKGLPFDTIDTKFPYPKPQVNLPRGHIWVEGDNYFHSIDSNTFGPISSGL 142
Query: 137 VRGRVTHVVWPPQRIG 152
V G+ +VWPP R G
Sbjct: 143 VIGKAITIVWPPSRWG 158
>UniRef100_UPI0000025FFF UPI0000025FFF UniRef100 entry
Length = 166
Score = 97.8 bits (242), Expect = 9e-20
Identities = 55/144 (38%), Positives = 73/144 (50%), Gaps = 10/144 (6%)
Query: 18 GLITFTVSDRYATVVPVRGASMSPTFNPKTNSFTDDYVFVEKLCLDKFKFSHGDIVIFSS 77
G I + VV G SM PT D VF E L + GDIVI S
Sbjct: 20 GCIAHCAFEYVGGVVMCSGPSMEPTIQ------NSDIVFAENLSRHFYGIQRGDIVIAKS 73
Query: 78 PSNFKETHIKRIIALPGEWFVNRHNQDVLK----VPEGHCWVEGDNAASSTDSKSYGPVP 133
PS+ K KR+I L G+ ++ DV K VP GH W+EGDN +STDS+ YGP+P
Sbjct: 74 PSDPKSNICKRVIGLEGDKILSTSPSDVFKSRSYVPTGHVWLEGDNLQNSTDSRYYGPIP 133
Query: 134 LGLVRGRVTHVVWPPQRIGAVKNT 157
GL+RG + +WP G ++++
Sbjct: 134 YGLIRGHIFFKIWPFSDFGFLRDS 157
>UniRef100_Q96LU5 Hypothetical protein FLJ25059 [Homo sapiens]
Length = 166
Score = 97.1 bits (240), Expect = 2e-19
Identities = 55/142 (38%), Positives = 70/142 (48%), Gaps = 10/142 (7%)
Query: 18 GLITFTVSDRYATVVPVRGASMSPTFNPKTNSFTDDYVFVEKLCLDKFKFSHGDIVIFSS 77
G I + VV G SM PT D VF E L + GDIVI S
Sbjct: 20 GCIAHCAFEYVGGVVMCSGPSMEPTIQ------NSDIVFAENLSRHFYGIQRGDIVIAKS 73
Query: 78 PSNFKETHIKRIIALPGEWFVNRHNQDVLK----VPEGHCWVEGDNAASSTDSKSYGPVP 133
PS+ K KR+I L G+ + D K VP GH W+EGDN +STDS+ YGP+P
Sbjct: 74 PSDPKSNICKRVIGLEGDKILTTSPSDFFKSHSYVPMGHVWLEGDNLQNSTDSRCYGPIP 133
Query: 134 LGLVRGRVTHVVWPPQRIGAVK 155
GL+RGR+ +WP G ++
Sbjct: 134 YGLIRGRIFFKIWPLSDFGFLR 155
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.319 0.136 0.422
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 305,112,867
Number of Sequences: 2790947
Number of extensions: 12849566
Number of successful extensions: 23420
Number of sequences better than 10.0: 560
Number of HSP's better than 10.0 without gapping: 452
Number of HSP's successfully gapped in prelim test: 109
Number of HSP's that attempted gapping in prelim test: 22536
Number of HSP's gapped (non-prelim): 834
length of query: 165
length of database: 848,049,833
effective HSP length: 117
effective length of query: 48
effective length of database: 521,509,034
effective search space: 25032433632
effective search space used: 25032433632
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 70 (31.6 bits)
Medicago: description of AC146575.4


