

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146574.11 + phase: 0 /pseudo
(738 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
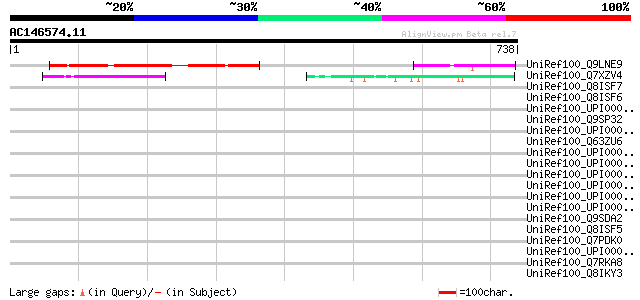
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9LNE9 T21E18.2 protein [Arabidopsis thaliana] 239 3e-61
UniRef100_Q7XZV4 Aleurone layer morphogenesis protein [Oryza sat... 107 2e-21
UniRef100_Q8ISF7 2MDa_1 protein [Caenorhabditis elegans] 41 0.15
UniRef100_Q8ISF6 2MDa_2 protein [Caenorhabditis elegans] 41 0.15
UniRef100_UPI0000499259 UPI0000499259 UniRef100 entry 40 0.33
UniRef100_Q9SP32 Endoribonuclease Dicer homolog [Arabidopsis tha... 39 0.43
UniRef100_UPI000046CED4 UPI000046CED4 UniRef100 entry 39 0.56
UniRef100_Q63ZU6 Hypothetical protein [Xenopus laevis] 39 0.56
UniRef100_UPI000046AC95 UPI000046AC95 UniRef100 entry 39 0.73
UniRef100_UPI000042ED83 UPI000042ED83 UniRef100 entry 39 0.73
UniRef100_UPI000042D7EE UPI000042D7EE UniRef100 entry 39 0.73
UniRef100_UPI0000311BB8 UPI0000311BB8 UniRef100 entry 39 0.73
UniRef100_UPI0000075E61 UPI0000075E61 UniRef100 entry 38 0.95
UniRef100_UPI000046AE99 UPI000046AE99 UniRef100 entry 38 0.95
UniRef100_Q9SDA2 Hypothetical protein F13G24.20 [Arabidopsis tha... 38 0.95
UniRef100_Q8ISF5 1MDa_1 protein [Caenorhabditis elegans] 38 0.95
UniRef100_Q7PDK0 Erythrocyte membrane protein PFEMP3 [Plasmodium... 38 0.95
UniRef100_UPI0000260F9A UPI0000260F9A UniRef100 entry 37 2.1
UniRef100_Q7RKA8 Hypothetical protein [Plasmodium yoelii yoelii] 37 2.1
UniRef100_Q8IKY3 Hypothetical protein [Plasmodium falciparum] 37 2.1
>UniRef100_Q9LNE9 T21E18.2 protein [Arabidopsis thaliana]
Length = 1628
Score = 239 bits (609), Expect = 3e-61
Identities = 133/307 (43%), Positives = 187/307 (60%), Gaps = 34/307 (11%)
Query: 58 SDVCPTADAVRAFVEHLVDPMLPAKASIRDAPSISQQQKVAKQVHSVVLLYNYYHRKRHP 117
+D CPT DA+RA +E LVDP+LP+K + D PS S ++ VAKQVH+VVLLYNYYHRK +P
Sbjct: 1015 TDSCPTEDAIRALLESLVDPLLPSKPT-DDLPSTSIRESVAKQVHAVVLLYNYYHRKDNP 1073
Query: 118 ELEFVAFKDFCKLIVGLRPALLIYMKFTQKPDQTDLVDVEQQLSLTEKAIASSCDICTCL 177
LE ++F+ F L ++PALL ++K + V Q L EK I +C + L
Sbjct: 1074 HLECLSFESFRSLATVMKPALLQHLK--------EDGGVSGQTVLLEKVIVDACSLSMSL 1125
Query: 178 DASKNVPNIEGWPVSKVAILLVDSKMENCLLRFCSTTDGVWSVIEKDVGSSDQISEDMNE 237
DAS ++ + P+ +VA+LLVDS+ ++C L+ S T GVWS++EK + E+ E
Sbjct: 1126 DASSDLFILNKCPIRRVAVLLVDSEKKSCYLQHSSITQGVWSLLEKPIEKEKAARENQKE 1185
Query: 238 LKHTYQKRRVIQNPTKNGLNVDDDEFLQVGYSAVKEATGVNSNDIMLLESYTVYSQRKEK 297
+ F +V ++ VKEATGVN DI++LE + V S +EK
Sbjct: 1186 ----------------------EGVFQKVAFAVVKEATGVNHKDIVILERHLVCSLSEEK 1223
Query: 298 TASRFYIMKC-SQSTADGSIQVPIKDLIESFRGPLLKKSSSSWTITSVVEYFHVLPYSEI 356
TA RFYIMKC SQ G + P+++++ +GPL +KS S WT+ S+VEYFHVLPY+ +
Sbjct: 1224 TAVRFYIMKCTSQDKFSG--ENPVEEVLSCMQGPLFEKSFSDWTMNSIVEYFHVLPYATL 1281
Query: 357 ISDWISR 363
I DW SR
Sbjct: 1282 IEDWFSR 1288
Score = 109 bits (272), Expect = 3e-22
Identities = 69/180 (38%), Positives = 97/180 (53%), Gaps = 36/180 (20%)
Query: 589 NANSSNKNLEKIQTFIASKGTILSQTALNALIRKRNALALQQRAIEDEMAVCNMKIHRWL 648
+A++SN NLE++QT + SK T LS+TAL L+ KR+ L QQR IEDE+A C+ + +
Sbjct: 1445 SAHNSNPNLEELQTSLLSKATSLSETALKVLLCKRDKLTRQQRNIEDEIAKCD----KCI 1500
Query: 649 AGEEDDFELKLESVIEGCNGTWLR--------------------------------KLDG 676
+ D+EL+LE+V+E CN T+ R +LD
Sbjct: 1501 QNIKGDWELQLETVLECCNETYPRRNLQESLDKSACQSNKRLKLSETLPSTKSLCQRLDD 1560
Query: 677 ICHENNWILPTYSVSLSDGEFHATVRVKGVDFEYSCEGNTCPFPREARDSAAAQMLTKFR 736
IC NNW+LP Y V+ SDG + A VR+ G + G EAR+SAAA +LTK +
Sbjct: 1561 ICLMNNWVLPNYRVAPSDGGYEAEVRITGNHVACTIHGEEKSDAEEARESAAACLLTKLQ 1620
>UniRef100_Q7XZV4 Aleurone layer morphogenesis protein [Oryza sativa]
Length = 454
Score = 107 bits (266), Expect = 2e-21
Identities = 68/181 (37%), Positives = 101/181 (55%), Gaps = 6/181 (3%)
Query: 49 VDSKGKMGRSDVCPTADAVRAFVEHLVDPMLPAKASIRDAPSISQQQKVAKQVHSVVLLY 108
V+ +G + V DAV V++LV P L + R P Q VA+QVH V+LY
Sbjct: 3 VERRGGGEVNGVVEMEDAVGILVDYLVRPAL--RKGSRMTPE--NQADVARQVHMAVILY 58
Query: 109 NYYHRKRHPELEFVAFKDFCKLI-VGLRPALLIYMKFTQKPDQTDLVDVEQ-QLSLTEKA 166
NYYHRK+ P+L F F K + L +LL Y + +++ E LS+T+KA
Sbjct: 59 NYYHRKQFPQLAFADAMRFFKCASLTLGDSLLAYSNMVHQHEKSSGSPGEGVNLSVTDKA 118
Query: 167 IASSCDICTCLDASKNVPNIEGWPVSKVAILLVDSKMENCLLRFCSTTDGVWSVIEKDVG 226
+ +C I LDA+++ P++ WP+SKVA+LL+DS + CLL S V S++EK++
Sbjct: 119 VVDACGIAEALDANQDSPDMAMWPISKVAVLLLDSTRKRCLLESGSVGKNVRSLLEKEID 178
Query: 227 S 227
S
Sbjct: 179 S 179
Score = 62.0 bits (149), Expect = 6e-08
Identities = 93/376 (24%), Positives = 144/376 (37%), Gaps = 88/376 (23%)
Query: 432 SNSLQDSKLAEKQFPKHEVTESHVSSKGLYIDLDNKPGSDTKVALNQKEK-NGCGITKRC 490
S +L DS LA + H S G PG +++ K + CGI +
Sbjct: 82 SLTLGDSLLAYSN-----MVHQHEKSSG-------SPGEGVNLSVTDKAVVDACGIAEAL 129
Query: 491 DSVKED-----WDMDVDKSLVLPSKNKECQ-------KHTANTLHISEDQKIENPSVQHH 538
D+ ++ W + L+L S K C K+ + L D +I P V++
Sbjct: 130 DANQDSPDMAMWPISKVAVLLLDSTRKRCLLESGSVGKNVRSLLEKEIDSEI--PRVKNA 187
Query: 539 SNECTRPSKAEKAVSKRKHITE-----GGIKDQSAFDKICAGTTFENDSV---------- 583
S K VS ++ E G D A K+ + EN S+
Sbjct: 188 LPPVVDVSTM-KFVSCSVNVKETAAANAGFVDMEAGVKMNSNKRRENHSLDLNISHDMAM 246
Query: 584 EKCILNANSS--------NKNLEKIQTFIASKGTILSQTALNALIRKRNALALQQRAIED 635
EK ++ N+ +K + Q A L + + ++KR+ L +QR IED
Sbjct: 247 EKALVIKNTELEWRSAEVSKKSDFHQGGGAKDNKDLKFASFKSYLKKRDDLHRKQRMIED 306
Query: 636 EMAVCNMKIHRWLAGEE-------------------------------DDFEL----KLE 660
E +M I AG E D +E + +
Sbjct: 307 ETVQFDMDIQSVFAGGEWTPEAMSLLEKYGILVDSLDMVEVNGSSYSGDGYETLTIERKK 366
Query: 661 SVIEGCNGTWLRKLDGICHENNWILPTYSV--SLSDGEFHATVRVKGVDFEYSCEGNTCP 718
+E ++LD +C ENNWILP Y V SL+DG + A V + ++F G+
Sbjct: 367 LTVERLLRNKCQELDEVCRENNWILPRYKVMPSLTDGMYVANVDIACLEFSQMTFGDPKT 426
Query: 719 FPREARDSAAAQMLTK 734
PR+AR+SAAA +L +
Sbjct: 427 NPRDARESAAANLLAE 442
>UniRef100_Q8ISF7 2MDa_1 protein [Caenorhabditis elegans]
Length = 18534
Score = 40.8 bits (94), Expect = 0.15
Identities = 62/266 (23%), Positives = 106/266 (39%), Gaps = 44/266 (16%)
Query: 432 SNSLQDSKLAEKQFPKHEVTESHVSSKGLYIDLDNKPGS--------------------- 470
S+SL +S L K+ K EV +S +S ID+ N+ +
Sbjct: 11180 SDSLAESTLRSKKTKKGEVEKSELS-----IDMKNQDKTTLATTLLEDDLAKTTSAEESE 11234
Query: 471 -DTKVALNQKEKNGCGITKRCDSVKEDWDMDVDKSLVLPSKNKECQKHTANTL------- 522
+ VAL KEK + ++ S + + V+P KN++ K + +
Sbjct: 11235 AEHLVALQNKEKTSLAMRRKRVSFDSSTKSESIED-VIPDKNRDSDKMSITGIKKKMSQK 11293
Query: 523 -HISEDQKIENPSVQHHSN--ECTRPSKAEKAVSKRKHITEGGIKDQSAFDKICAGTTFE 579
+E QK E+P V+ S+ E T SK +K+ + R TE + D++ DK T +
Sbjct: 11294 SESAEAQKNESPEVKEISSFEEKTLKSK-KKSKADRNQGTEANLGDKT-IDKDYLSVTDK 11351
Query: 580 NDSVEKCILNANSSNK----NLEKIQTFIASKGTILSQTALNALIRKRNALALQQRAIED 635
N S+EK + + N K+ T A + L A + ++ D
Sbjct: 11352 NQSLEKSEASGQAEKSIKAPNKSKVTTSFADESLTSELDRLMADEEMAEMMFAEEEKAAD 11411
Query: 636 EMAVCNMKIHRWLAGEEDDFELKLES 661
+ V N + +E+ E+ L+S
Sbjct: 11412 LLNVMNKNKGLNKSEQEESQEISLKS 11437
Score = 38.1 bits (87), Expect = 0.95
Identities = 43/202 (21%), Positives = 84/202 (41%), Gaps = 19/202 (9%)
Query: 442 EKQFPKHEVTESHVSSKGLYIDLDNKPGSDTKVALNQKEKNGCGITKRCDSVKEDWDMDV 501
EKQ ++ E+ K +D NK + K+A + + K + +E +D
Sbjct: 9441 EKQAQINKAAEADAVKKQNELDEQNKLEATKKLAAEKLKLEEQSAAKSKQAAEEQAKLDA 9500
Query: 502 DKSLVLPSKNKECQKHTANTLHISEDQKIENPSVQHHSNECTRPSKAEKAVSKRKHITEG 561
+K K +K T + +D+K S SNE +K + K+ ++
Sbjct: 9501 Q------TKAKAAEKQTG----LEKDEKSNKDS---GSNETVEEKPKKKVLKKKTEKSDS 9547
Query: 562 GIKDQSAFDKICAGTTFENDSVEKCILNANSSNKNL---EKIQTFIASKGTILSQTAL-- 616
I +S K A + ++S + + +A S K +K++ I +K + ++ L
Sbjct: 9548 SISQKSDTSKTVAESAGSSESETQKVADATSKQKETDKKQKLEAEITAKKSADEKSKLET 9607
Query: 617 -NALIRKRNALALQQRAIEDEM 637
+ LI+ A +Q+ ED++
Sbjct: 9608 ESKLIKAAEDAAKKQKEKEDKL 9629
Score = 35.8 bits (81), Expect = 4.7
Identities = 44/212 (20%), Positives = 89/212 (41%), Gaps = 42/212 (19%)
Query: 436 QDSKLAEKQFPKHEVTESHVSSKGLYIDLDNKPGSDTKVALNQKEKNGCGITKRCDSVKE 495
++ +LAEKQ + E +++ L ++ K ++T QKE+ ++ +
Sbjct: 9915 KEKELAEKQKLESEAATKKAAAEKLKLEEQKKKDAETASIEKQKEQ------EKLAQEQS 9968
Query: 496 DWDMDVDKSLVLPSKNKECQKHTANTLHISEDQKIENPSVQHHSNECTRPSKAE----KA 551
++D KS +E QK+E+ + + E + S E K
Sbjct: 9969 KLEVDAKKS--------------------AEKQKLESETKSKKTEEAPKESVDEKPKKKV 10008
Query: 552 VSKRKHITEGGIKDQSAFDKICAGTTFENDSVEKCILNANSSNKNLE-----KIQTFIAS 606
+ K+ ++ I +S K A + ++DS + + A+ ++K E K+++ IA+
Sbjct: 10009 LKKKTEKSDSSISQKSDTAKTVAESAGQSDSETQKVSEADKAHKQKESDEKQKLESEIAA 10068
Query: 607 KGTILSQTALNALIRKRNALALQQRAIEDEMA 638
K + ++ K A ++ IEDE A
Sbjct: 10069 KKSAEQKS-------KLETEAKTKKVIEDESA 10093
>UniRef100_Q8ISF6 2MDa_2 protein [Caenorhabditis elegans]
Length = 18519
Score = 40.8 bits (94), Expect = 0.15
Identities = 62/266 (23%), Positives = 106/266 (39%), Gaps = 44/266 (16%)
Query: 432 SNSLQDSKLAEKQFPKHEVTESHVSSKGLYIDLDNKPGS--------------------- 470
S+SL +S L K+ K EV +S +S ID+ N+ +
Sbjct: 11180 SDSLAESTLRSKKTKKGEVEKSELS-----IDMKNQDKTTLATTLLEDDLAKTTSAEESE 11234
Query: 471 -DTKVALNQKEKNGCGITKRCDSVKEDWDMDVDKSLVLPSKNKECQKHTANTL------- 522
+ VAL KEK + ++ S + + V+P KN++ K + +
Sbjct: 11235 AEHLVALQNKEKTSLAMRRKRVSFDSSTKSESIED-VIPDKNRDSDKMSITGIKKKMSQK 11293
Query: 523 -HISEDQKIENPSVQHHSN--ECTRPSKAEKAVSKRKHITEGGIKDQSAFDKICAGTTFE 579
+E QK E+P V+ S+ E T SK +K+ + R TE + D++ DK T +
Sbjct: 11294 SESAEAQKNESPEVKEISSFEEKTLKSK-KKSKADRNQGTEANLGDKT-IDKDYLSVTDK 11351
Query: 580 NDSVEKCILNANSSNK----NLEKIQTFIASKGTILSQTALNALIRKRNALALQQRAIED 635
N S+EK + + N K+ T A + L A + ++ D
Sbjct: 11352 NQSLEKSEASGQAEKSIKAPNKSKVTTSFADESLTSELDRLMADEEMAEMMFAEEEKAAD 11411
Query: 636 EMAVCNMKIHRWLAGEEDDFELKLES 661
+ V N + +E+ E+ L+S
Sbjct: 11412 LLNVMNKNKGLNKSEQEESQEISLKS 11437
Score = 38.1 bits (87), Expect = 0.95
Identities = 43/202 (21%), Positives = 84/202 (41%), Gaps = 19/202 (9%)
Query: 442 EKQFPKHEVTESHVSSKGLYIDLDNKPGSDTKVALNQKEKNGCGITKRCDSVKEDWDMDV 501
EKQ ++ E+ K +D NK + K+A + + K + +E +D
Sbjct: 9441 EKQAQINKAAEADAVKKQNELDEQNKLEATKKLAAEKLKLEEQSAAKSKQAAEEQAKLDA 9500
Query: 502 DKSLVLPSKNKECQKHTANTLHISEDQKIENPSVQHHSNECTRPSKAEKAVSKRKHITEG 561
+K K +K T + +D+K S SNE +K + K+ ++
Sbjct: 9501 Q------TKAKAAEKQTG----LEKDEKSNKDS---GSNETVEEKPKKKVLKKKTEKSDS 9547
Query: 562 GIKDQSAFDKICAGTTFENDSVEKCILNANSSNKNL---EKIQTFIASKGTILSQTAL-- 616
I +S K A + ++S + + +A S K +K++ I +K + ++ L
Sbjct: 9548 SISQKSDTSKTVAESAGSSESETQKVADATSKQKETDKKQKLEAEITAKKSADEKSKLET 9607
Query: 617 -NALIRKRNALALQQRAIEDEM 637
+ LI+ A +Q+ ED++
Sbjct: 9608 ESKLIKAAEDAAKKQKEKEDKL 9629
Score = 35.8 bits (81), Expect = 4.7
Identities = 44/212 (20%), Positives = 89/212 (41%), Gaps = 42/212 (19%)
Query: 436 QDSKLAEKQFPKHEVTESHVSSKGLYIDLDNKPGSDTKVALNQKEKNGCGITKRCDSVKE 495
++ +LAEKQ + E +++ L ++ K ++T QKE+ ++ +
Sbjct: 9915 KEKELAEKQKLESEAATKKAAAEKLKLEEQKKKDAETASIEKQKEQ------EKLAQEQS 9968
Query: 496 DWDMDVDKSLVLPSKNKECQKHTANTLHISEDQKIENPSVQHHSNECTRPSKAE----KA 551
++D KS +E QK+E+ + + E + S E K
Sbjct: 9969 KLEVDAKKS--------------------AEKQKLESETKSKKTEEAPKESVDEKPKKKV 10008
Query: 552 VSKRKHITEGGIKDQSAFDKICAGTTFENDSVEKCILNANSSNKNLE-----KIQTFIAS 606
+ K+ ++ I +S K A + ++DS + + A+ ++K E K+++ IA+
Sbjct: 10009 LKKKTEKSDSSISQKSDTAKTVAESAGQSDSETQKVSEADKAHKQKESDEKQKLESEIAA 10068
Query: 607 KGTILSQTALNALIRKRNALALQQRAIEDEMA 638
K + ++ K A ++ IEDE A
Sbjct: 10069 KKSAEQKS-------KLETEAKTKKVIEDESA 10093
>UniRef100_UPI0000499259 UPI0000499259 UniRef100 entry
Length = 1598
Score = 39.7 bits (91), Expect = 0.33
Identities = 47/202 (23%), Positives = 81/202 (39%), Gaps = 24/202 (11%)
Query: 435 LQDSKLAEKQFPKHEVTESHVSSKGLYIDLDNKPGSDTKVAL---NQKEKNGCGITKRCD 491
+ D K E Q KH++ + ++ + N+ + KV L N+K +N K D
Sbjct: 945 INDKKDLEIQLSKHQIENQKIQNQNDELKKINQTKENEKVLLNEINEKLQNKINELKEKD 1004
Query: 492 SVKEDWDMDVDKSLVLPSKNKECQKHTANTLHISEDQKIENPSVQHHSNECTRPSK-AEK 550
++ED + + + SK L +E+ KIEN +Q NE K E
Sbjct: 1005 KIQEDENNKLQSEITNYSKTITSLTEKIE-LCKTENTKIEN-KIQQKENEIEEIKKEKEI 1062
Query: 551 AVSKRKHITEGGIKDQSAFDKICAGTTFENDSVEKCILNAN------SSNKNLEKIQTFI 604
A+ + H + IKD FEN E+ I+ N ++N+ +++ I
Sbjct: 1063 ALEELNHEIKKKIKD------------FENQIKEQEIIQNNQIITIKEKDQNIYELKQHI 1110
Query: 605 ASKGTILSQTALNALIRKRNAL 626
I+ +T L + N +
Sbjct: 1111 EKMKKIIEETPLKEYEEENNKI 1132
>UniRef100_Q9SP32 Endoribonuclease Dicer homolog [Arabidopsis thaliana]
Length = 1909
Score = 39.3 bits (90), Expect = 0.43
Identities = 45/152 (29%), Positives = 67/152 (43%), Gaps = 15/152 (9%)
Query: 592 SSNKNLEKIQTFIASKGTILSQTALNALIRK---RNALALQQRAIEDEMAVCNMK-IHRW 647
S + N ++ FI ++Q + +K RNALA + E E+A K I+
Sbjct: 1754 SRSGNTATVEVFIDGVQVGVAQNPQKKMAQKLAARNALAALK---EKEIAESKEKHINNG 1810
Query: 648 LAGEEDDFELKLESVIEGCNGTWLRKLDGICHENNWILPTYSVSLSDGEFHAT-----VR 702
AGE D E + + G + L+ IC NW +P+Y G HA VR
Sbjct: 1811 NAGE-DQGENENGNKKNGHQPFTRQTLNDICLRKNWPMPSYRCVKEGGPAHAKRFTFGVR 1869
Query: 703 VKGVDFEYS--CEGNTCPFPREARDSAAAQML 732
V D ++ C G P ++A+DSAA +L
Sbjct: 1870 VNTSDRGWTDECIGEPMPSVKKAKDSAAVLLL 1901
>UniRef100_UPI000046CED4 UPI000046CED4 UniRef100 entry
Length = 2078
Score = 38.9 bits (89), Expect = 0.56
Identities = 38/146 (26%), Positives = 55/146 (37%), Gaps = 18/146 (12%)
Query: 465 DNKPGSDTKVALNQKEKNGCGITKRCDSVKEDWDMDVDKSLVLPSKNKECQKHTANTLHI 524
+N K NQKE I KR D V + D + L+L E K+ N
Sbjct: 1298 ENNEKQKIKKIKNQKEVEP--IEKRRDIVNGNTSADANSDLILDKYGSEFNKYEGNNTTF 1355
Query: 525 SEDQKIENPSVQHHSNECTRPSKAEKAVSKRKHITEGGIK-DQSAFDKICAGTT--FEND 581
++ S ++SN K+ + E IK S DK G EN+
Sbjct: 1356 ENVKENFRKSATNNSN-------------KKLYTKESNIKASNSEIDKETCGNVEQIENE 1402
Query: 582 SVEKCILNANSSNKNLEKIQTFIASK 607
+ +KCI + N N + +Q I K
Sbjct: 1403 NRDKCISKQDIKNSNNQSLQNNILVK 1428
>UniRef100_Q63ZU6 Hypothetical protein [Xenopus laevis]
Length = 1489
Score = 38.9 bits (89), Expect = 0.56
Identities = 57/228 (25%), Positives = 102/228 (44%), Gaps = 36/228 (15%)
Query: 439 KLAEKQFPKHEVTESHVSSKGLYI-DLDNKPGSDTKVALNQKEKNGCGITKRCD--SVKE 495
+L K ++ E+ VS K + +LDN K+ ++ ++ D +++
Sbjct: 1082 ELENKNHALNDGNEALVSDKEKMLSELDNAKQELLKITMDNEDLQAALEKMDADLKELQK 1141
Query: 496 DWDMDVDKSLVLPSKNKECQKHTANTLHISEDQKIENPSVQHHSNECTRPSK--AEKAVS 553
DM V + L +N+E Q + N E +++ +E K AE+ ++
Sbjct: 1142 SKDMLVAQCEDLQYQNQELQNYQKNLTE-------EKITLEREKDEIIEMLKNTAEEMLN 1194
Query: 554 KRKHITEGGIKDQSAFDKICAGTTFENDS-VEKCI-LNAN-----SSNKNLEKIQTFIAS 606
K+KH+TE +G E +S VEK + L +N S NL K T I +
Sbjct: 1195 KQKHLTEE-----------ISGLKMEKESVVEKHLELESNLSALISERDNLLKATTEIKT 1243
Query: 607 --KGTILSQTALNALI----RKRNALALQQRAIEDEMAVCNMKIHRWL 648
+G +L Q LN LI +++ LAL++ A E+E+ ++++ L
Sbjct: 1244 EREGLMLKQNELNTLIENLHQEKERLALERNAKEEELIAVISQLNQLL 1291
>UniRef100_UPI000046AC95 UPI000046AC95 UniRef100 entry
Length = 1207
Score = 38.5 bits (88), Expect = 0.73
Identities = 34/144 (23%), Positives = 57/144 (38%), Gaps = 12/144 (8%)
Query: 460 LYIDLDNKPGSDTKVALNQKEKNGCGITKRCDSVKEDWDMDVDKSLVLPSKNKECQKHTA 519
LY L+N G++ LN + N +V +++ + S KN ++
Sbjct: 353 LYNYLNNNNGNNKNGLLNNHQHNNRNGHSSLSNVNNNYNSSSNVSNSKTKKNNYSNQNIL 412
Query: 520 NTLHISEDQKIENPSVQHHSNECTRPSKAEKAVSKRKHITEGGIKDQSAFDKICAGTTFE 579
N L++S D N ++ E AEK V K K+ + KD C+ +T
Sbjct: 413 NKLNVSIDYNSFNDYNNNYIQE------AEKVVEKYKNFNKMNSKDSDNMYSACSYSTGA 466
Query: 580 NDSVEKCILNANSSNKNLEKIQTF 603
D N ++ N N+ I+ F
Sbjct: 467 TD------FNNHNGNNNINTIKHF 484
>UniRef100_UPI000042ED83 UPI000042ED83 UniRef100 entry
Length = 705
Score = 38.5 bits (88), Expect = 0.73
Identities = 51/187 (27%), Positives = 82/187 (43%), Gaps = 27/187 (14%)
Query: 436 QDSKLAEKQFPKHEVTESHVSSKGLYIDLDNKPGSDTKVALNQKEKNGCGITKRCDSVKE 495
Q S LAE PK ++ E+ V++ G D+ TK+ALN + G KR D VK
Sbjct: 16 QSSYLAESLKPKPDI-ENKVTTNGDSNDI-------TKIALNTNKPGDDGSLKRDDRVKN 67
Query: 496 DWDMDVDKSLVLPSKNKECQKHTANTLHISEDQKI-ENPSVQHHSNECTRPSKAEKAVSK 554
+ L + + K T + S+ K+ +NP + + A+ VSK
Sbjct: 68 LINR-------LTATGRHEAKRNPTTTNASDAYKLFQNPD----KLKALKRMFAQPKVSK 116
Query: 555 R---KHITEGGIKDQSAFD-KICAGTTFENDSVEKCILNANSSNKNLEKIQTFIASKGTI 610
R KH+ E K + + K F++ S +K + + N NLE+ F+A K
Sbjct: 117 RRVKKHVKEKFQKKHTEKNTKTSTEPFFKDKSTDKYLTSNNIKKHNLEQ---FVAPKNED 173
Query: 611 LSQTALN 617
+++ A N
Sbjct: 174 IARLAHN 180
>UniRef100_UPI000042D7EE UPI000042D7EE UniRef100 entry
Length = 455
Score = 38.5 bits (88), Expect = 0.73
Identities = 51/187 (27%), Positives = 82/187 (43%), Gaps = 27/187 (14%)
Query: 436 QDSKLAEKQFPKHEVTESHVSSKGLYIDLDNKPGSDTKVALNQKEKNGCGITKRCDSVKE 495
Q S LAE PK ++ E+ V++ G D+ TK+ALN + G KR D VK
Sbjct: 16 QSSYLAESLKPKPDI-ENKVTTNGDSNDI-------TKIALNTNKPGDDGSLKRDDRVKN 67
Query: 496 DWDMDVDKSLVLPSKNKECQKHTANTLHISEDQKI-ENPSVQHHSNECTRPSKAEKAVSK 554
+ L + + K T + S+ K+ +NP + + A+ VSK
Sbjct: 68 LINR-------LTATGRHEAKRNPTTTNASDAYKLFQNPD----KLKALKRMFAQPKVSK 116
Query: 555 R---KHITEGGIKDQSAFD-KICAGTTFENDSVEKCILNANSSNKNLEKIQTFIASKGTI 610
R KH+ E K + + K F++ S +K + + N NLE+ F+A K
Sbjct: 117 RRVKKHVKEKFQKKHTEKNTKTSTEPFFKDKSTDKYLTSNNIKKHNLEQ---FVAPKNED 173
Query: 611 LSQTALN 617
+++ A N
Sbjct: 174 IARLAHN 180
>UniRef100_UPI0000311BB8 UPI0000311BB8 UniRef100 entry
Length = 336
Score = 38.5 bits (88), Expect = 0.73
Identities = 38/141 (26%), Positives = 64/141 (44%), Gaps = 14/141 (9%)
Query: 220 VIEKDVGSSDQISEDMNELKHTYQKRRVIQNPTKNGLNVDDDEFLQVGYSAVKEATGVNS 279
VIEK + + + ED K+ Q+ +N K LN+D+D L + S VK+
Sbjct: 39 VIEKIL--KELLQEDDLLPKNQQQEIDFNENNNKENLNIDEDTALVLIKSDVKKEV---- 92
Query: 280 NDIMLLESYTVYSQRKEKTASRFYIMKCSQSTADGSIQVPIKDLIESFRGPLLKKSSSSW 339
N+I L + +T +Q KEK S + ++ +K+L S LK++ +
Sbjct: 93 NEISLNKRFTNNNQYKEKANSNI--------SLPNFVEKKVKNLRRSINSEKLKENINDI 144
Query: 340 TITSVVEYFHVLPYSEIISDW 360
I ++ E + E+IS W
Sbjct: 145 QINTIKETELIKCRIELISHW 165
>UniRef100_UPI0000075E61 UPI0000075E61 UniRef100 entry
Length = 2083
Score = 38.1 bits (87), Expect = 0.95
Identities = 43/202 (21%), Positives = 84/202 (41%), Gaps = 19/202 (9%)
Query: 442 EKQFPKHEVTESHVSSKGLYIDLDNKPGSDTKVALNQKEKNGCGITKRCDSVKEDWDMDV 501
EKQ ++ E+ K +D NK + K+A + + K + +E +D
Sbjct: 901 EKQAQINKAAEADAVKKQNELDEQNKLEATKKLAAEKLKLEEQSAAKSKQAAEEQAKLDA 960
Query: 502 DKSLVLPSKNKECQKHTANTLHISEDQKIENPSVQHHSNECTRPSKAEKAVSKRKHITEG 561
+K K +K T + +D+K S SNE +K + K+ ++
Sbjct: 961 Q------TKAKAAEKQTG----LEKDEKSNKDS---GSNETVEEKPKKKVLKKKTEKSDS 1007
Query: 562 GIKDQSAFDKICAGTTFENDSVEKCILNANSSNKNL---EKIQTFIASKGTILSQTAL-- 616
I +S K A + ++S + + +A S K +K++ I +K + ++ L
Sbjct: 1008 SISQKSDTSKTVAESAGSSESETQKVADATSKQKETDKKQKLEAEITAKKSADEKSKLET 1067
Query: 617 -NALIRKRNALALQQRAIEDEM 637
+ LI+ A +Q+ ED++
Sbjct: 1068 ESKLIKAAEDAAKKQKEKEDKL 1089
Score = 35.8 bits (81), Expect = 4.7
Identities = 44/212 (20%), Positives = 89/212 (41%), Gaps = 42/212 (19%)
Query: 436 QDSKLAEKQFPKHEVTESHVSSKGLYIDLDNKPGSDTKVALNQKEKNGCGITKRCDSVKE 495
++ +LAEKQ + E +++ L ++ K ++T QKE+ ++ +
Sbjct: 1375 KEKELAEKQKLESEAATKKAAAEKLKLEEQKKKDAETASIEKQKEQ------EKLAQEQS 1428
Query: 496 DWDMDVDKSLVLPSKNKECQKHTANTLHISEDQKIENPSVQHHSNECTRPSKAE----KA 551
++D KS +E QK+E+ + + E + S E K
Sbjct: 1429 KLEVDAKKS--------------------AEKQKLESETKSKKTEEAPKESVDEKPKKKV 1468
Query: 552 VSKRKHITEGGIKDQSAFDKICAGTTFENDSVEKCILNANSSNKNLE-----KIQTFIAS 606
+ K+ ++ I +S K A + ++DS + + A+ ++K E K+++ IA+
Sbjct: 1469 LKKKTEKSDSSISQKSDTAKTVAESAGQSDSETQKVSEADKAHKQKESDEKQKLESEIAA 1528
Query: 607 KGTILSQTALNALIRKRNALALQQRAIEDEMA 638
K + ++ K A ++ IEDE A
Sbjct: 1529 KKSAEQKS-------KLETEAKTKKVIEDESA 1553
>UniRef100_UPI000046AE99 UPI000046AE99 UniRef100 entry
Length = 725
Score = 38.1 bits (87), Expect = 0.95
Identities = 54/231 (23%), Positives = 92/231 (39%), Gaps = 32/231 (13%)
Query: 439 KLAEKQFPKHEVTESHVSSKGLYIDLDNKPGSDTKVALNQKEKNGCGITKRCDSVKEDWD 498
+L EK K + +S ++S D +N+ +V+L + + +K + K
Sbjct: 415 QLLEKAENKRDENDSLLNSSDDENDQNNQ-SDQNEVSLVETTEISKLSSKEVEKAKLQLK 473
Query: 499 MDVDKSLVLPSKNKECQKHTANTLHIS---EDQKIENPSVQHHSNECTRPSKAEKAVSKR 555
D+D +L K +T N +++ + +IE P V+ N T K EK +++
Sbjct: 474 EDMDLGNML---RKNAVLNTNNIINLDFELNETEIEEPDVKDVENNNTEEKKEEKTNNEK 530
Query: 556 KHITEGGIKDQSAFDKI------------CAGTTFENDSVEKCILNANSSNKNLEKIQTF 603
I E I D DKI FEN+ E I N N + N E + F
Sbjct: 531 DKIKEKKINDLVDIDKINDEDILKQYGEKITVYNFENNIYENYI-NINDTINNRENEELF 589
Query: 604 IASKGTILSQTALNA---------LIRKRNALALQQRAIEDEMAVCNMKIH 645
S ++ LN + R++N + ++ IE + NM +H
Sbjct: 590 ELSDSEKITGEELNVNEWCNYNTLIEREQNIIKKEKEIIEKKK---NMPLH 637
>UniRef100_Q9SDA2 Hypothetical protein F13G24.20 [Arabidopsis thaliana]
Length = 561
Score = 38.1 bits (87), Expect = 0.95
Identities = 43/166 (25%), Positives = 73/166 (43%), Gaps = 17/166 (10%)
Query: 455 VSSKGLYIDLDNKPGSDTKVALNQKEKNGCGITKRCDSVKEDWDMDVDKSLVLPSKNKEC 514
V +K I + P D L KEK RCD V E D++V K S+ E
Sbjct: 180 VVNKASRISQNKSPKEDLSKNLKNKEKGKIVEPVRCDDVLEKTDLEVKK----VSRISEN 235
Query: 515 QKHTANTLHISEDQKIENP-----SVQHHSNECTRPSK-AEKAVSKRKHITEGGIKDQSA 568
+ +TL E KI+ P ++ S + + S+ +E SK + + K+++
Sbjct: 236 KSSKEDTLKNKEKAKIDEPVRCDDVLEKTSLDAQKVSRISENKNSKEERLKNLKNKEKTN 295
Query: 569 FDKICAGTTFENDSVEKCILNANSSNKNLEKIQTFIASKGTILSQT 614
D+ +D+VEK + SS +EK + +++K +S+T
Sbjct: 296 IDE----PVRPDDAVEKTLYVVESS---VEKKKKKMSTKSVKISET 334
>UniRef100_Q8ISF5 1MDa_1 protein [Caenorhabditis elegans]
Length = 10578
Score = 38.1 bits (87), Expect = 0.95
Identities = 43/202 (21%), Positives = 84/202 (41%), Gaps = 19/202 (9%)
Query: 442 EKQFPKHEVTESHVSSKGLYIDLDNKPGSDTKVALNQKEKNGCGITKRCDSVKEDWDMDV 501
EKQ ++ E+ K +D NK + K+A + + K + +E +D
Sbjct: 9421 EKQAQINKAAEADAVKKQNELDEQNKLEATKKLAAEKLKLEEQSAAKSKQAAEEQAKLDA 9480
Query: 502 DKSLVLPSKNKECQKHTANTLHISEDQKIENPSVQHHSNECTRPSKAEKAVSKRKHITEG 561
+K K +K T + +D+K S SNE +K + K+ ++
Sbjct: 9481 Q------TKAKAAEKQTG----LEKDEKSNKDS---GSNETVEEKPKKKVLKKKTEKSDS 9527
Query: 562 GIKDQSAFDKICAGTTFENDSVEKCILNANSSNKNL---EKIQTFIASKGTILSQTAL-- 616
I +S K A + ++S + + +A S K +K++ I +K + ++ L
Sbjct: 9528 SISQKSDTSKTVAESAGSSESETQKVADATSKQKETDKKQKLEAEITAKKSADEKSKLET 9587
Query: 617 -NALIRKRNALALQQRAIEDEM 637
+ LI+ A +Q+ ED++
Sbjct: 9588 ESKLIKAAEDAAKKQKEKEDKL 9609
Score = 35.8 bits (81), Expect = 4.7
Identities = 44/212 (20%), Positives = 89/212 (41%), Gaps = 42/212 (19%)
Query: 436 QDSKLAEKQFPKHEVTESHVSSKGLYIDLDNKPGSDTKVALNQKEKNGCGITKRCDSVKE 495
++ +LAEKQ + E +++ L ++ K ++T QKE+ ++ +
Sbjct: 9895 KEKELAEKQKLESEAATKKAAAEKLKLEEQKKKDAETASIEKQKEQ------EKLAQEQS 9948
Query: 496 DWDMDVDKSLVLPSKNKECQKHTANTLHISEDQKIENPSVQHHSNECTRPSKAE----KA 551
++D KS +E QK+E+ + + E + S E K
Sbjct: 9949 KLEVDAKKS--------------------AEKQKLESETKSKKTEEAPKESVDEKPKKKV 9988
Query: 552 VSKRKHITEGGIKDQSAFDKICAGTTFENDSVEKCILNANSSNKNLE-----KIQTFIAS 606
+ K+ ++ I +S K A + ++DS + + A+ ++K E K+++ IA+
Sbjct: 9989 LKKKTEKSDSSISQKSDTAKTVAESAGQSDSETQKVSEADKAHKQKESDEKQKLESEIAA 10048
Query: 607 KGTILSQTALNALIRKRNALALQQRAIEDEMA 638
K + ++ K A ++ IEDE A
Sbjct: 10049 KKSAEQKS-------KLETEAKTKKVIEDESA 10073
>UniRef100_Q7PDK0 Erythrocyte membrane protein PFEMP3 [Plasmodium yoelii yoelii]
Length = 1444
Score = 38.1 bits (87), Expect = 0.95
Identities = 33/144 (22%), Positives = 57/144 (38%), Gaps = 12/144 (8%)
Query: 460 LYIDLDNKPGSDTKVALNQKEKNGCGITKRCDSVKEDWDMDVDKSLVLPSKNKECQKHTA 519
LY L+N ++ V LN + N + +++ + + KN ++
Sbjct: 707 LYNYLNNNNANNKNVLLNNHQHNNRNGHSGLSNTNNNYNSSSNVLISKTKKNNYSNQNIL 766
Query: 520 NTLHISEDQKIENPSVQHHSNECTRPSKAEKAVSKRKHITEGGIKDQSAFDKICAGTTFE 579
N L++S D N ++ E AEK V K K+ + KD C+ +T
Sbjct: 767 NKLNVSIDYNSFNDYNNNYIQE------AEKVVEKYKNFNKMNSKDSDNMYSACSYSTGA 820
Query: 580 NDSVEKCILNANSSNKNLEKIQTF 603
D LN ++ N N+ I+ F
Sbjct: 821 TD------LNNHNGNNNINTIKHF 838
>UniRef100_UPI0000260F9A UPI0000260F9A UniRef100 entry
Length = 374
Score = 37.0 bits (84), Expect = 2.1
Identities = 38/171 (22%), Positives = 71/171 (41%), Gaps = 14/171 (8%)
Query: 571 KICAGTTFENDSVEKCILNANSSNKNLEKIQTFIASKGTILSQTALNALIRKRNALALQQ 630
K + F N + E N+ K+ ++++ F S T + Q+ +N L R + QQ
Sbjct: 191 KAFTDSLFRNQAAENVTRQTNA--KSTQEVERFFTSLDTAVEQSNVNRLTAIRQFNSGQQ 248
Query: 631 RAIEDEMAVCNMKIHRWLAGEEDDFELKLESVIEGCNGTWLRKLDGICHENNWILPTYS- 689
I N + R D F ++ + I N W R + I + N + +++
Sbjct: 249 NVISQFNK--NQEFAR------DTFNTQMLASIRAANANWFRSVATIDNGNQMLANSFAA 300
Query: 690 ---VSLSDGEFHATVRVKGVDFEYSCEGNTCPFPREARDSAAAQMLTKFRS 737
++L D E++ + + D + E R+A+ +A AQ + RS
Sbjct: 301 QSELALRDKEYNNLWQRRRDDAFFVFEAAERQLDRQAQAAALAQQESLVRS 351
>UniRef100_Q7RKA8 Hypothetical protein [Plasmodium yoelii yoelii]
Length = 1668
Score = 37.0 bits (84), Expect = 2.1
Identities = 44/178 (24%), Positives = 72/178 (39%), Gaps = 28/178 (15%)
Query: 433 NSLQDSKLAEKQFPKHEVTESHVSSKGLYIDLDNKPGSDTKVALNQKEKNGCGITKRCD- 491
N+LQ EK +E++E H ++ ID K + N K G K CD
Sbjct: 639 NNLQTKYSDEKY--NNELSEEHDTTH--LIDKKRKFNDINEHKNNSKNGKYNGTIKACDV 694
Query: 492 ---SVKEDWDMDVDKSLVLPSKNKECQKHTANTLH---ISEDQKIENPSVQHHSNECTRP 545
+K+ D + + S E Q +T +H I+ D+K +N + +NE +
Sbjct: 695 NENKIKKCHDEKNPEDKKISSNKNETQINTKKEIHEKGITRDKK-KNTEKKKTNNEIIK- 752
Query: 546 SKAEKAVSKRKHITEGGIKDQSAFDKICA--GTTFENDSVEKCILNANSSNKNLEKIQ 601
TEG + + + KI T F+N K I N ++ ++EKI+
Sbjct: 753 -------------TEGTVLNNKKYKKITIKNDTNFDNKEENKIIFNEHTKENDMEKIK 797
>UniRef100_Q8IKY3 Hypothetical protein [Plasmodium falciparum]
Length = 1029
Score = 37.0 bits (84), Expect = 2.1
Identities = 36/190 (18%), Positives = 83/190 (42%), Gaps = 14/190 (7%)
Query: 442 EKQFPKHEVTESHVSSKGLYIDLDNKPGSDTKVALNQKEKNGCGITKRCDSVKEDWDM-- 499
+K+ PK+ + +S+++ DNK S K+ N + D + +D
Sbjct: 584 DKRNPKNNINKSNINYN------DNKYKSTHKMNNNYNNDDDDNNNDYNDDAYDYYDHYH 637
Query: 500 ---DVDKSLVLPSKNKECQKHTANTLHISE--DQKIENPSVQHHSNECTRPSKAEKAVSK 554
D + L + N++ + N + D++ ++ ++ H +E +K K ++K
Sbjct: 638 KCDDTNNKLQINKFNRKEETTIINVCSEKKPYDERPKSKNIMHKHDEQDEHNKNVKIMNK 697
Query: 555 RKHITEGGIKDQSAFDKICAGTTFENDSVEKCILNANSSNKNLEKIQTFIASKGTILSQT 614
+K+ T ++ +S +++ G N++ N N+S+ N + +K +S+
Sbjct: 698 KKYCTNNSVQGKSGSERV-VGYRSANNNNNNNNKNNNNSSSNNNISSNNMCNKNMNISRK 756
Query: 615 ALNALIRKRN 624
+LN + +N
Sbjct: 757 SLNVSSKIKN 766
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.324 0.136 0.409
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,177,407,042
Number of Sequences: 2790947
Number of extensions: 48356605
Number of successful extensions: 188502
Number of sequences better than 10.0: 44
Number of HSP's better than 10.0 without gapping: 2
Number of HSP's successfully gapped in prelim test: 42
Number of HSP's that attempted gapping in prelim test: 188447
Number of HSP's gapped (non-prelim): 85
length of query: 738
length of database: 848,049,833
effective HSP length: 135
effective length of query: 603
effective length of database: 471,271,988
effective search space: 284177008764
effective search space used: 284177008764
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.5 bits)
S2: 79 (35.0 bits)
Medicago: description of AC146574.11


