

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146569.8 - phase: 0
(166 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
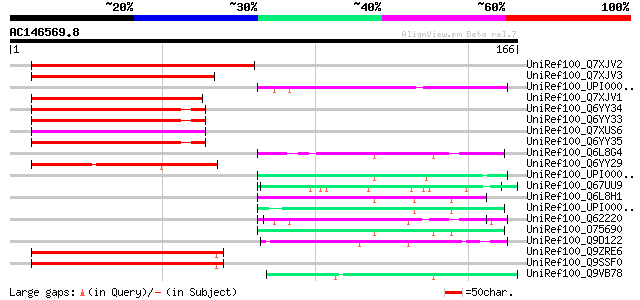
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q7XJV2 OSJNBa0016O02.18 protein [Oryza sativa] 69 4e-11
UniRef100_Q7XJV3 OSJNBa0016O02.17 protein [Oryza sativa] 59 5e-08
UniRef100_UPI00001807DF UPI00001807DF UniRef100 entry 57 1e-07
UniRef100_Q7XJV1 OSJNBa0016O02.19 protein [Oryza sativa] 57 2e-07
UniRef100_Q6YY34 Hypothetical protein OSJNBb0056I22.9 [Oryza sat... 54 2e-06
UniRef100_Q6YY33 Hypothetical protein OSJNBb0056I22.11 [Oryza sa... 52 4e-06
UniRef100_Q7XUS6 OSJNBa0060P14.7 protein [Oryza sativa] 52 4e-06
UniRef100_Q6YY35 Hypothetical protein OSJNBb0056I22.7 [Oryza sat... 49 4e-05
UniRef100_Q6L8G4 Keratin-associated protein 5-11 [Homo sapiens] 49 4e-05
UniRef100_Q6YY29 Hypothetical protein OSJNBb0056I22.16 [Oryza sa... 49 5e-05
UniRef100_UPI000036EF3B UPI000036EF3B UniRef100 entry 49 6e-05
UniRef100_Q67UU9 Putative keratin associated protein [Oryza sativa] 49 6e-05
UniRef100_Q6L8H1 Keratin-associated protein 5-4 [Homo sapiens] 49 6e-05
UniRef100_UPI000021D6F8 UPI000021D6F8 UniRef100 entry 48 8e-05
UniRef100_Q62220 Serine 2 ultra high sulfur protein [Mus musculus] 48 8e-05
UniRef100_O75690 Keratin-associated protein 5-8 [Homo sapiens] 48 1e-04
UniRef100_Q9D122 Mus musculus 18-day embryo whole body cDNA, RIK... 47 1e-04
UniRef100_Q9ZRE6 ATFP4 [Arabidopsis thaliana] 47 1e-04
UniRef100_Q9SSF0 MLP3.5 protein [Arabidopsis thaliana] 47 1e-04
UniRef100_Q9VB78 CG6124-PA [Drosophila melanogaster] 47 1e-04
>UniRef100_Q7XJV2 OSJNBa0016O02.18 protein [Oryza sativa]
Length = 131
Score = 68.9 bits (167), Expect = 4e-11
Identities = 35/73 (47%), Positives = 45/73 (60%)
Query: 8 GVTSVSLEGDDKDRVCVTGDNVDIVCLANQLKKKFNNVTILTTEEVKKKTEAEKKKEKEE 67
GV S+++ GDD+DR+ V GD VD+ CL L+KK +L EEVK K E+KK+ EE
Sbjct: 29 GVISMAITGDDRDRLEVVGDGVDVTCLVTCLRKKVRFADVLQVEEVKDKKPEEEKKKPEE 88
Query: 68 EKKKMIEACHAVL 80
EKK C A L
Sbjct: 89 EKKPPECPCQATL 101
>UniRef100_Q7XJV3 OSJNBa0016O02.17 protein [Oryza sativa]
Length = 122
Score = 58.9 bits (141), Expect = 5e-08
Identities = 28/60 (46%), Positives = 39/60 (64%)
Query: 8 GVTSVSLEGDDKDRVCVTGDNVDIVCLANQLKKKFNNVTILTTEEVKKKTEAEKKKEKEE 67
GV+SV + GD KDR+ V GD VD CL L+KK + ++ EEVK+K EKK E+++
Sbjct: 29 GVSSVEVTGDGKDRLQVVGDGVDAACLVTCLRKKIGHAELVQVEEVKEKKPEEKKPEEKK 88
>UniRef100_UPI00001807DF UPI00001807DF UniRef100 entry
Length = 191
Score = 57.4 bits (137), Expect = 1e-07
Identities = 31/85 (36%), Positives = 38/85 (44%), Gaps = 5/85 (5%)
Query: 82 GSCK--CNNFN-CHGKCNSCTKCESSKCDGTNCVTICSFKCGNSKCDGNKCNGCTPKVTS 138
G CK C + C G C SC C+SS C + C CS CG+S C + C C S
Sbjct: 85 GGCKGGCGSCGGCKGGCGSCGGCQSSCCQSSCCKPCCSSGCGSSCCQSSCCKPCC--CQS 142
Query: 139 PCFQRCPQWCTCSKCCAPYQPYCKP 163
C + C +C K C CKP
Sbjct: 143 SCCKPCCCQSSCCKPCCCQSSCCKP 167
Score = 33.1 bits (74), Expect = 2.7
Identities = 28/98 (28%), Positives = 31/98 (31%), Gaps = 24/98 (24%)
Query: 82 GSCKCNNF-------------NCHGKCNSCTKCES--SKCDGT--NCVTICSFKCGNSKC 124
GSC C C G C SC C+ C G C + K G C
Sbjct: 25 GSCCCKPVCCCVPVCSCSSCGGCKGGCGSCGGCKGGCGSCGGCKGGCGSCGGCKGGCGSC 84
Query: 125 DGNK-----CNGCTPKVTS--PCFQRCPQWCTCSKCCA 155
G K C GC S C C Q C CC+
Sbjct: 85 GGCKGGCGSCGGCKGGCGSCGGCQSSCCQSSCCKPCCS 122
>UniRef100_Q7XJV1 OSJNBa0016O02.19 protein [Oryza sativa]
Length = 150
Score = 56.6 bits (135), Expect = 2e-07
Identities = 28/56 (50%), Positives = 37/56 (66%)
Query: 8 GVTSVSLEGDDKDRVCVTGDNVDIVCLANQLKKKFNNVTILTTEEVKKKTEAEKKK 63
GV+SV + GD KDR+ V GD VD VC+ N+L+KK + I+ EEVK+K KK
Sbjct: 29 GVSSVEVTGDGKDRLQVVGDGVDPVCVVNRLRKKIGHAEIVQVEEVKEKKPDLPKK 84
>UniRef100_Q6YY34 Hypothetical protein OSJNBb0056I22.9 [Oryza sativa]
Length = 117
Score = 53.5 bits (127), Expect = 2e-06
Identities = 29/57 (50%), Positives = 37/57 (64%), Gaps = 3/57 (5%)
Query: 8 GVTSVSLEGDDKDRVCVTGDNVDIVCLANQLKKKFNNVTILTTEEVKKKTEAEKKKE 64
GV SV+L GD KD+V V GD VD + L L+KK + T++T EVKK+ EKK E
Sbjct: 30 GVDSVALAGDGKDQVVVVGDGVDSIKLTTALRKKVGHATLMTVGEVKKE---EKKPE 83
>UniRef100_Q6YY33 Hypothetical protein OSJNBb0056I22.11 [Oryza sativa]
Length = 121
Score = 52.4 bits (124), Expect = 4e-06
Identities = 29/57 (50%), Positives = 37/57 (64%), Gaps = 3/57 (5%)
Query: 8 GVTSVSLEGDDKDRVCVTGDNVDIVCLANQLKKKFNNVTILTTEEVKKKTEAEKKKE 64
GV SV+L GD KD+V V GD VD + L L+KK + T++T EVKK+ EKK E
Sbjct: 30 GVDSVALAGDGKDQVVVVGDGVDSIKLTAALRKKVGHATLVTVGEVKKE---EKKPE 83
>UniRef100_Q7XUS6 OSJNBa0060P14.7 protein [Oryza sativa]
Length = 119
Score = 52.4 bits (124), Expect = 4e-06
Identities = 27/57 (47%), Positives = 34/57 (59%)
Query: 8 GVTSVSLEGDDKDRVCVTGDNVDIVCLANQLKKKFNNVTILTTEEVKKKTEAEKKKE 64
GV S+ + GD KDR+ V GD VD VCL L++K I+ EEVK K EK+ E
Sbjct: 30 GVNSMEVTGDGKDRLQVVGDGVDPVCLVACLRRKIGYAEIVQVEEVKDKKPEEKQPE 86
>UniRef100_Q6YY35 Hypothetical protein OSJNBb0056I22.7 [Oryza sativa]
Length = 119
Score = 49.3 bits (116), Expect = 4e-05
Identities = 27/57 (47%), Positives = 36/57 (62%), Gaps = 3/57 (5%)
Query: 8 GVTSVSLEGDDKDRVCVTGDNVDIVCLANQLKKKFNNVTILTTEEVKKKTEAEKKKE 64
GV SV+L GD KD++ V GD VD + L L+KK + T++T E KK+ EKK E
Sbjct: 30 GVDSVALAGDGKDQLVVVGDGVDSIELTTALRKKVGHATLMTVGEDKKE---EKKPE 83
>UniRef100_Q6L8G4 Keratin-associated protein 5-11 [Homo sapiens]
Length = 156
Score = 49.3 bits (116), Expect = 4e-05
Identities = 31/86 (36%), Positives = 38/86 (44%), Gaps = 13/86 (15%)
Query: 82 GSCKCNNFNCHGKCNSCTKCESSKCDGTNCVTICSFK--CGNSKCDGNKCNGCTPKVT-- 137
GSC C+ NC C C C SS C C + CS C +S C + C C + +
Sbjct: 78 GSCGCSQSNC---CKPC--CSSSGCGSFCCQSSCSKPCCCQSSCCQSSCCKPCCCQSSCC 132
Query: 138 -SPCFQRCPQWCTCSKCCAPYQPYCK 162
S CF+ C C S CC P CK
Sbjct: 133 QSSCFKPC---CCQSSCCVPVCCQCK 155
Score = 35.8 bits (81), Expect = 0.41
Identities = 26/87 (29%), Positives = 32/87 (35%), Gaps = 9/87 (10%)
Query: 84 CKCNNFNCHGKCNSCTKCESSKCDGTNCVTICSFKCGNSKCDGNKCNGCTPKVTSP---- 139
C C C SC+ C S C + S K G C ++ N C P +S
Sbjct: 40 CCCKPVCCCVPACSCSSCGSCGGSKGGCGSCGSSKGGCGSCGCSQSNCCKPCCSSSGCGS 99
Query: 140 --CFQRCPQWCTC-SKCCAPYQPYCKP 163
C C + C C S CC CKP
Sbjct: 100 FCCQSSCSKPCCCQSSCC--QSSCCKP 124
>UniRef100_Q6YY29 Hypothetical protein OSJNBb0056I22.16 [Oryza sativa]
Length = 133
Score = 48.9 bits (115), Expect = 5e-05
Identities = 29/63 (46%), Positives = 39/63 (61%), Gaps = 3/63 (4%)
Query: 8 GVTSVSLEGDDKDRVCVTGDNVDIVCLANQLKKKFNNVTIL--TTEEVKKKTEAEKKKEK 65
GV SV L GDD+ +V V GD VD + L N L++K + L ++ KKK E KKKE+
Sbjct: 29 GVVSVELAGDDRSKVVVVGD-VDSIGLTNALRRKVDGSAELVEVSDASKKKEEEAKKKEE 87
Query: 66 EEE 68
+EE
Sbjct: 88 KEE 90
>UniRef100_UPI000036EF3B UPI000036EF3B UniRef100 entry
Length = 231
Score = 48.5 bits (114), Expect = 6e-05
Identities = 29/101 (28%), Positives = 39/101 (37%), Gaps = 20/101 (19%)
Query: 82 GSCKCNNFNCHGKCNSCTKCESSKCDGTNCVTICSFK-----------CGNSKCDGNKCN 130
GSC C+ +C+ C + C SS C + C CS CG+S C + C
Sbjct: 96 GSCGCSQCSCYKPCCCSSGCGSSCCQSSCCKPCCSQSSCCKPCSCSSGCGSSCCQSSCCK 155
Query: 131 GCTPK--------VTSPCFQRCPQWCTCSKCCAPYQPYCKP 163
C + +S C C Q C CC+ CKP
Sbjct: 156 PCCSQFSCCKPFCCSSGCGSSCCQSSCCKPCCS-QSSCCKP 195
Score = 40.8 bits (94), Expect = 0.013
Identities = 25/90 (27%), Positives = 34/90 (37%), Gaps = 17/90 (18%)
Query: 84 CK--CNNFNCHGKCNSCTKCESSKCDGTNCVTICS-----------FKCGNSKCDGNKCN 130
CK C+ +C C+ + C SS C + C CS CG+S C + C
Sbjct: 125 CKPCCSQSSCCKPCSCSSGCGSSCCQSSCCKPCCSQFSCCKPFCCSSGCGSSCCQSSCCK 184
Query: 131 GCTPKVTS----PCFQRCPQWCTCSKCCAP 156
C + + C C C S CC P
Sbjct: 185 PCCSQSSCCKPCSCSSGCGSSCCQSSCCKP 214
Score = 36.6 bits (83), Expect = 0.24
Identities = 27/83 (32%), Positives = 33/83 (39%), Gaps = 10/83 (12%)
Query: 84 CK--CNNFNCHGKCNSCTKCESSKCDGTNCVTICSFKCGNSKCDGNKCNGCTPKVTSPCF 141
CK C+ F+C + C SS C + C CS S C K C+ S C
Sbjct: 154 CKPCCSQFSCCKPFCCSSGCGSSCCQSSCCKPCCS----QSSC--CKPCSCSSGCGSSCC 207
Query: 142 QR--CPQWCTCSKCCAPYQPYCK 162
Q C C+ S CC P CK
Sbjct: 208 QSSCCKPCCSQSSCCVPICCQCK 230
Score = 33.1 bits (74), Expect = 2.7
Identities = 24/79 (30%), Positives = 27/79 (33%), Gaps = 13/79 (16%)
Query: 90 NCHGKCNSCTKCESSKCDGTNCVTICSFKCGNSK--CDGNKCNGCTPKVTSPCFQRCPQW 147
+C G C C SK C CG SK C C+ C+ C C
Sbjct: 67 SCGGSKGGCGSCGGSKGG-------CG-SCGGSKGGCGSCGCSQCSCYKPCCCSSGCGSS 118
Query: 148 CTCSKCCAP---YQPYCKP 163
C S CC P CKP
Sbjct: 119 CCQSSCCKPCCSQSSCCKP 137
>UniRef100_Q67UU9 Putative keratin associated protein [Oryza sativa]
Length = 426
Score = 48.5 bits (114), Expect = 6e-05
Identities = 34/93 (36%), Positives = 37/93 (39%), Gaps = 11/93 (11%)
Query: 83 SCKCNNFNCHGKCNS--CTKCE-SSKCDGTNCVTICSFKCGNSKCDGNKCNGCTP----- 134
SC C C KC C C SS CD C CS C S C + C+ C P
Sbjct: 101 SCLCYCCKCSPKCKRPRCLNCSCSSCCDEPCCKPNCSACCAGSCCSPDCCSCCKPNCSCC 160
Query: 135 KVTSPCFQRCPQWC-TCSKCCAPYQPYCKPYWT 166
K S C C C +CS CC CKP T
Sbjct: 161 KTPSCCKPNCSCSCPSCSSCCD--TSCCKPSCT 191
Score = 47.8 bits (112), Expect = 1e-04
Identities = 29/91 (31%), Positives = 36/91 (38%), Gaps = 11/91 (12%)
Query: 82 GSCKCNNFNCHGKCNSCTK---CESSKCDGTNCVTICS----FKCGNSKCDGNKCN-GCT 133
GSC N +C CN C C + C +C S FKC + C + C CT
Sbjct: 289 GSCSTNCCSCKPSCNGCCGEQCCRCADCFSCSCPRCSSCFNIFKCSCAGCCSSLCKCPCT 348
Query: 134 PKV---TSPCFQRCPQWCTCSKCCAPYQPYC 161
+ S C +R P C C C QP C
Sbjct: 349 TQCFSCQSSCCKRQPSCCKCQSSCCEGQPSC 379
Score = 42.7 bits (99), Expect = 0.003
Identities = 25/87 (28%), Positives = 35/87 (39%), Gaps = 17/87 (19%)
Query: 84 CKCNNFNCHG------KCNSCTKCESSKCD------GTNCVTI---CSFKCGNSKCDGNK 128
C C + C+G CN C C ++C TNC + C+ CG C
Sbjct: 256 CGCPSCGCNGCGLPSCGCNGCGSCSCAQCKPDCGSCSTNCCSCKPSCNGCCGEQCCRCAD 315
Query: 129 CNGCTPKVTSPCFQRCPQWCTCSKCCA 155
C C+ S CF C+C+ CC+
Sbjct: 316 CFSCSCPRCSSCFNIFK--CSCAGCCS 340
Score = 41.2 bits (95), Expect = 0.010
Identities = 26/95 (27%), Positives = 36/95 (37%), Gaps = 12/95 (12%)
Query: 75 ACHAVLHGSCKCNNFNCHG------KCNSC--TKCESSKCDGTNCVTICSFKCGNSKCDG 126
AC + C C + C+G CN C C + C +C C CG+ C
Sbjct: 237 ACPSCGCNGCGCPSCGCNGCGCPSCGCNGCGLPSCGCNGCGSCSCAQ-CKPDCGS--CST 293
Query: 127 NKCNGCTPKVTSPCFQRCPQWCTCSKCCAPYQPYC 161
N C+ C P C ++C + C C P C
Sbjct: 294 NCCS-CKPSCNGCCGEQCCRCADCFSCSCPRCSSC 327
Score = 41.2 bits (95), Expect = 0.010
Identities = 24/78 (30%), Positives = 31/78 (38%), Gaps = 11/78 (14%)
Query: 84 CKCNNFNCHGKCNSCTKCESSKCDGTNCVTICSFKCGNSKCDGNKCNGCTPKVTSPCFQR 143
C C + C+G C C S C+G C + CG C N C C+ C Q
Sbjct: 236 CACPSCGCNG-CG----CPSCGCNGCGCPSCGCNGCGLPSCGCNGCGSCS------CAQC 284
Query: 144 CPQWCTCSKCCAPYQPYC 161
P +CS C +P C
Sbjct: 285 KPDCGSCSTNCCSCKPSC 302
Score = 38.1 bits (87), Expect = 0.083
Identities = 25/91 (27%), Positives = 35/91 (37%), Gaps = 19/91 (20%)
Query: 76 CHAVLHGSCKCNNFNCHGKC----NSCTKCESSKCDGT------NCVTICSFKCGNSKCD 125
C ++ C F+C C SC KC+SS C+G +C ++ C C
Sbjct: 339 CSSLCKCPCTTQCFSCQSSCCKRQPSCCKCQSSCCEGQPSCCEGHCCSLPKPSCPECSCG 398
Query: 126 GN-KCNGCT-----PKVTSPCFQRCPQWCTC 150
C CT P+ +PC C C C
Sbjct: 399 CVWSCKNCTEGCRCPRCRNPC---CLSGCLC 426
Score = 37.0 bits (84), Expect = 0.19
Identities = 22/79 (27%), Positives = 29/79 (35%), Gaps = 12/79 (15%)
Query: 86 CNNFNCHGKCNSCTKCESSKCDGTNCVTICSFKCGNS--------KCDGNKCNGCTPKVT 137
C NC C SC+ C + C +C F C S C ++CN +P
Sbjct: 165 CCKPNCSCSCPSCSSCCDTSCCKPSCTCFNIFSCFKSLYSCFKIPSCFKSQCNCSSPNC- 223
Query: 138 SPCFQRCPQWCTCSKCCAP 156
C P C+C C P
Sbjct: 224 --CTCTLPS-CSCKGCACP 239
>UniRef100_Q6L8H1 Keratin-associated protein 5-4 [Homo sapiens]
Length = 288
Score = 48.5 bits (114), Expect = 6e-05
Identities = 27/80 (33%), Positives = 33/80 (40%), Gaps = 5/80 (6%)
Query: 82 GSCKCNNFNCHGKCNSCTKCESSKCDGTNCVTICSFK-CGNSKCDGNKCNG--CTPKVTS 138
GSC C+ +C C + C SS C + C CS CG+S C + C C
Sbjct: 192 GSCGCSQCSCCKPCCCSSGCGSSCCQSSCCKPCCSSSGCGSSCCQSSCCKPYCCQSSCCK 251
Query: 139 PCFQR--CPQWCTCSKCCAP 156
PC C C S CC P
Sbjct: 252 PCCSSSGCGSSCCQSSCCNP 271
Score = 42.0 bits (97), Expect = 0.006
Identities = 24/82 (29%), Positives = 32/82 (38%), Gaps = 18/82 (21%)
Query: 84 CK--CNNFNCHGKCNSCTKCESSKCDGTNCVTICSFK-CGNSKCDGNKCNGCTPKVTSPC 140
CK C++ C C + C+ C + C CS CG+S C + CN C
Sbjct: 221 CKPCCSSSGCGSSCCQSSCCKPYCCQSSCCKPCCSSSGCGSSCCQSSCCNPC-------- 272
Query: 141 FQRCPQWCTCSKCCAPYQPYCK 162
C+ S CC P CK
Sbjct: 273 -------CSQSSCCVPVCCQCK 287
Score = 38.1 bits (87), Expect = 0.083
Identities = 28/91 (30%), Positives = 32/91 (34%), Gaps = 19/91 (20%)
Query: 82 GSCKCNNFNC------HGKCNSCTKCESSKCDGTNCVTICSFKCGNSKCDGNKCNGCTPK 135
GSC + C G C SC + S C C + C C S C C P
Sbjct: 172 GSCGGSKGGCGSCGGSKGGCGSCGCSQCSCCKPCCCSSGCGSSCCQSSC-------CKPC 224
Query: 136 VTSPCFQRCPQWCTCSKCCAPY---QPYCKP 163
+S C C S CC PY CKP
Sbjct: 225 CSS---SGCGSSCCQSSCCKPYCCQSSCCKP 252
>UniRef100_UPI000021D6F8 UPI000021D6F8 UniRef100 entry
Length = 216
Score = 48.1 bits (113), Expect = 8e-05
Identities = 28/86 (32%), Positives = 34/86 (38%), Gaps = 9/86 (10%)
Query: 82 GSCKCNNFNCHGKCNSCTKCESSKCDGTNCVTICSFKCGNSKCDGNKCNG--CTPKVTSP 139
GSC C G C + C+ C + C CS CG+S C + C C P
Sbjct: 134 GSCG----GCKGGCCQSSCCKPCCCQSSCCKPCCSSGCGSSCCQSSCCKPCCCQSSCCKP 189
Query: 140 CFQR---CPQWCTCSKCCAPYQPYCK 162
C + C C S CCAP CK
Sbjct: 190 CCCQSSCCKPCCCQSSCCAPVCCQCK 215
Score = 37.7 bits (86), Expect = 0.11
Identities = 29/90 (32%), Positives = 32/90 (35%), Gaps = 11/90 (12%)
Query: 82 GSCK--CNNFN-CHGKCNSCTKCES--SKCDGTNCVTICSFKCGN-SKCDG--NKCNGCT 133
G CK C + C G C SC C+ C G C CG+ C G C GC
Sbjct: 75 GGCKGGCGSCGGCKGGCGSCGGCKGGCGSCGGCGSCGGCKGGCGSCGGCKGGCGSCGGCG 134
Query: 134 PKVTSPCFQRCPQWCTCSKCCAPYQPYCKP 163
C C Q C CC CKP
Sbjct: 135 S--CGGCKGGCCQSSCCKPCCC-QSSCCKP 161
Score = 37.4 bits (85), Expect = 0.14
Identities = 23/74 (31%), Positives = 26/74 (35%), Gaps = 4/74 (5%)
Query: 82 GSCKCNNFNCHGKCNSCTKCESSKCDGTNCVTICSFKCGNSKCDGNKCNGCTPKVTSPCF 141
G CK +C G C SC C+ C C G C G K C PC
Sbjct: 95 GGCKGGCGSCGG-CGSCGGCKGGCGSCGGCKGGCGSCGGCGSCGGCKGGCCQSSCCKPC- 152
Query: 142 QRCPQWCTCSKCCA 155
C Q C CC+
Sbjct: 153 --CCQSSCCKPCCS 164
>UniRef100_Q62220 Serine 2 ultra high sulfur protein [Mus musculus]
Length = 223
Score = 48.1 bits (113), Expect = 8e-05
Identities = 29/92 (31%), Positives = 37/92 (39%), Gaps = 11/92 (11%)
Query: 82 GSCK--CNNFN-CHGKCNSCTKCESSKCDGTNCVTICSFKCGNSKCDGNKC-------NG 131
G CK C + C G C + C+ C + C CS CG+S C + C +
Sbjct: 58 GGCKGGCGSCGGCKGGCCQSSCCKPCCCQSSCCKPCCSSGCGSSCCQSSCCKPCCCQSSC 117
Query: 132 CTPKVTSPCFQRCPQWCTCSKCCAPYQPYCKP 163
C P +S C C Q C CC CKP
Sbjct: 118 CKPCCSSGCGSSCCQSSCCKPCCC-QSSCCKP 148
Score = 45.8 bits (107), Expect = 4e-04
Identities = 27/85 (31%), Positives = 35/85 (40%), Gaps = 8/85 (9%)
Query: 82 GSCKCNNFNCHGKCNSCTKCESSKCDGTNCVTICSFKCGNSKCDGNKCNGCTPKVTSPCF 141
GS C + C C + C+ C + C CS CG+S C + C C + S C
Sbjct: 127 GSSCCQSSCCKPCCCQSSCCKPCCCQSSCCKPCCSSGCGSSCCQSSCCKPCCCQ--SSCC 184
Query: 142 QRCPQWCTCSKCCAP---YQPYCKP 163
+ C C S CC P CKP
Sbjct: 185 KPC---CCQSSCCKPCCCQSSCCKP 206
Score = 44.7 bits (104), Expect = 9e-04
Identities = 24/74 (32%), Positives = 32/74 (42%), Gaps = 6/74 (8%)
Query: 84 CK-CNNFNCHGKCNSCTKCESSKCDGTNCVTICSFKCGNSKCDGNKCNGCTPKVTSPCFQ 142
CK C + C C + C+ C + C CS CG+S C + C C + S C +
Sbjct: 90 CKPCCSSGCGSSCCQSSCCKPCCCQSSCCKPCCSSGCGSSCCQSSCCKPCCCQ--SSCCK 147
Query: 143 RCPQWCTCSKCCAP 156
C C S CC P
Sbjct: 148 PC---CCQSSCCKP 158
Score = 35.0 bits (79), Expect = 0.71
Identities = 31/87 (35%), Positives = 34/87 (38%), Gaps = 21/87 (24%)
Query: 91 CHGKC-NSCTKCESSKCDGTNC-VTICSFK-CGNSK-----CDGNK-----CNGCTPKVT 137
C G C +SC C SS C C V +CS CG K C G K C GC
Sbjct: 6 CSGGCGSSCGGCGSSCCKPVCCCVPVCSCSSCGGCKGGCGSCGGCKGGCGSCGGCKGGCG 65
Query: 138 SP-------CFQRCPQWCTC-SKCCAP 156
S C C + C C S CC P
Sbjct: 66 SCGGCKGGCCQSSCCKPCCCQSSCCKP 92
>UniRef100_O75690 Keratin-associated protein 5-8 [Homo sapiens]
Length = 187
Score = 47.8 bits (112), Expect = 1e-04
Identities = 30/106 (28%), Positives = 40/106 (37%), Gaps = 25/106 (23%)
Query: 82 GSCKCNNFNCHGKCNSCTKCESSKCDGTNCVTICSFK-----------CGNSKCDGNKCN 130
GSC C+ +C+ C + C SS C + C CS CG+S C + C
Sbjct: 81 GSCGCSQCSCYKPCCCSSGCGSSCCQSSCCKPCCSQSSCCKPCSCSSGCGSSCCQSSCCK 140
Query: 131 GCTPKVT------------SPCFQR--CPQWCTCSKCCAPYQPYCK 162
C + + S C Q C C+ S CC P CK
Sbjct: 141 PCCSQSSCCKPCCCSSGCGSSCCQSSCCKPCCSQSSCCVPICCQCK 186
Score = 43.9 bits (102), Expect = 0.002
Identities = 32/110 (29%), Positives = 39/110 (35%), Gaps = 19/110 (17%)
Query: 73 IEACHAVLHGSC---KCNNFNCHGKCNSCTKCESSK-------CDGTNCV--TICSFKCG 120
+ AC GSC K +C G C C SK C +C CS CG
Sbjct: 42 VPACSCSSCGSCGGSKGGRGSCGGSKGDCGSCGGSKGGCGSCGCSQCSCYKPCCCSSGCG 101
Query: 121 NSKCDGNKCNGCTPKVT----SPCFQRCPQWCTCSKCCAP---YQPYCKP 163
+S C + C C + + C C C S CC P CKP
Sbjct: 102 SSCCQSSCCKPCCSQSSCCKPCSCSSGCGSSCCQSSCCKPCCSQSSCCKP 151
>UniRef100_Q9D122 Mus musculus 18-day embryo whole body cDNA, RIKEN full-length
enriched library, clone:1110033F04 product:weakly
similar to KERATIN ASSOCIATED PROTEIN 4.12 [Mus
musculus]
Length = 167
Score = 47.4 bits (111), Expect = 1e-04
Identities = 27/83 (32%), Positives = 38/83 (45%), Gaps = 7/83 (8%)
Query: 83 SCKCNNFNCHGKCNSCTKCESSKCDGTNCVT-ICSFKCGNSKCDGNKC-NGCTPKVTSPC 140
SC C C +C C+ + C + C++ C CG+S C G+ C C SPC
Sbjct: 52 SC-CRPSCCRPQCCQSVCCQPTCCRPSCCISSCCQPSCGSSSCCGSSCCRPCCRPCCSPC 110
Query: 141 FQRCPQWCTCSKCCAPYQPYCKP 163
C + C C CC +P C+P
Sbjct: 111 CSPCCRPC-CRPCC---RPCCRP 129
Score = 36.2 bits (82), Expect = 0.32
Identities = 30/106 (28%), Positives = 39/106 (36%), Gaps = 15/106 (14%)
Query: 72 MIEACHAVLHGSCKCNNFNCHGKCNSCTKCESSKCDGTNCVTIC-------SFKCGNSKC 124
M+ +C +V C C C T C ++ C + CV+ C S C S C
Sbjct: 1 MVSSCGSVCSEK-GCGQGCCQPSCCQTTCCRTTCCRPSCCVSSCCRPSCCVSSCCRPSCC 59
Query: 125 DGNKCNG--CTPKVTSP--CFQRCPQ-WCTCSKCCAP--YQPYCKP 163
C C P P C C Q C S CC +P C+P
Sbjct: 60 RPQCCQSVCCQPTCCRPSCCISSCCQPSCGSSSCCGSSCCRPCCRP 105
>UniRef100_Q9ZRE6 ATFP4 [Arabidopsis thaliana]
Length = 179
Score = 47.4 bits (111), Expect = 1e-04
Identities = 24/66 (36%), Positives = 42/66 (63%), Gaps = 3/66 (4%)
Query: 8 GVTSVSLEGDDKDRVCVTGDNVDIVCLANQLKKKFNNVTILTTEEVKKKTEAEKKKEKE- 66
GV +V ++GD ++++ VTG VD++ L N L+KK +++ +V+ + +KK E+E
Sbjct: 51 GVNAVEIKGDHRNQIEVTGVEVDMIALINTLRKKVAFAELVSVAKVEPPKDGDKKPEEEK 110
Query: 67 --EEKK 70
EEKK
Sbjct: 111 KPEEKK 116
>UniRef100_Q9SSF0 MLP3.5 protein [Arabidopsis thaliana]
Length = 157
Score = 47.4 bits (111), Expect = 1e-04
Identities = 24/66 (36%), Positives = 42/66 (63%), Gaps = 3/66 (4%)
Query: 8 GVTSVSLEGDDKDRVCVTGDNVDIVCLANQLKKKFNNVTILTTEEVKKKTEAEKKKEKE- 66
GV +V ++GD ++++ VTG VD++ L N L+KK +++ +V+ + +KK E+E
Sbjct: 29 GVNAVEIKGDHRNQIEVTGVEVDMIALINTLRKKVAFAELVSVAKVEPPKDGDKKPEEEK 88
Query: 67 --EEKK 70
EEKK
Sbjct: 89 KPEEKK 94
>UniRef100_Q9VB78 CG6124-PA [Drosophila melanogaster]
Length = 979
Score = 47.4 bits (111), Expect = 1e-04
Identities = 30/91 (32%), Positives = 37/91 (39%), Gaps = 10/91 (10%)
Query: 85 KCNNFNCHGKCNSCTKCESSK-----CDGTNCVTICSFKCGNSKCDGNKCNGCTPKVT-- 137
KC N +G C + KCE +K DG NCV IC C N C C T
Sbjct: 199 KCKNGCSNGVCRAPDKCECNKFYEKDIDG-NCVPICENGCVNGNCTAPDVCQCLTGYTKI 257
Query: 138 --SPCFQRCPQWCTCSKCCAPYQPYCKPYWT 166
+ C CP+ C C AP + C +T
Sbjct: 258 GHNVCLAVCPEGCQNGVCVAPNECSCNAGYT 288
Score = 40.4 bits (93), Expect = 0.017
Identities = 31/92 (33%), Positives = 38/92 (40%), Gaps = 17/92 (18%)
Query: 76 CHAVLHGSCKCNNFNCHGKCNSCTKCESSKCDGTNCVTICSFKCGNSKCDG-NKC----- 129
C V G CK GKC SC + SK G +C ICS C N CD KC
Sbjct: 459 CSPVCSGGCKNGFCVAPGKC-SCDE-GYSKETGNSCKPICSKGCENGFCDAPEKCSCNDG 516
Query: 130 ------NGCTPKVTSPC---FQRCPQWCTCSK 152
N C+P + C F P+ C+C +
Sbjct: 517 YEMDGENRCSPVCSGGCKNGFCVAPEKCSCDE 548
Score = 39.3 bits (90), Expect = 0.037
Identities = 31/90 (34%), Positives = 36/90 (39%), Gaps = 6/90 (6%)
Query: 76 CHAVLHGSCKCNNFNCHGKCNSCTKCESSKCDGTNCVTICSFKCGNSKCDG-NKC---NG 131
C V G CK GKC SC + SK G +C ICS C N CD KC +G
Sbjct: 657 CSPVCSGGCKNGFCIAPGKC-SCDE-GYSKETGNSCKPICSKGCENGFCDAPEKCSCNDG 714
Query: 132 CTPKVTSPCFQRCPQWCTCSKCCAPYQPYC 161
+ C C C C AP + C
Sbjct: 715 YEMDSENRCSPVCSGGCKNGFCVAPGKCSC 744
Score = 39.3 bits (90), Expect = 0.037
Identities = 31/90 (34%), Positives = 36/90 (39%), Gaps = 6/90 (6%)
Query: 76 CHAVLHGSCKCNNFNCHGKCNSCTKCESSKCDGTNCVTICSFKCGNSKCDG-NKC---NG 131
C V G CK GKC SC + SK G +C ICS C N CD KC +G
Sbjct: 591 CSPVCSGGCKNGFCVAPGKC-SCDE-GYSKETGNSCKPICSKGCENGFCDAPEKCSCNDG 648
Query: 132 CTPKVTSPCFQRCPQWCTCSKCCAPYQPYC 161
+ C C C C AP + C
Sbjct: 649 YEMDSENRCSPVCSGGCKNGFCIAPGKCSC 678
Score = 39.3 bits (90), Expect = 0.037
Identities = 31/90 (34%), Positives = 36/90 (39%), Gaps = 6/90 (6%)
Query: 76 CHAVLHGSCKCNNFNCHGKCNSCTKCESSKCDGTNCVTICSFKCGNSKCDG-NKC---NG 131
C V G CK GKC SC + SK G +C ICS C N CD KC +G
Sbjct: 393 CSPVCSGGCKNGFCVAPGKC-SCDE-GYSKETGNSCKPICSKGCENGFCDAPEKCSCNDG 450
Query: 132 CTPKVTSPCFQRCPQWCTCSKCCAPYQPYC 161
+ C C C C AP + C
Sbjct: 451 YEMDSENRCSPVCSGGCKNGFCVAPGKCSC 480
Score = 37.4 bits (85), Expect = 0.14
Identities = 30/96 (31%), Positives = 37/96 (38%), Gaps = 25/96 (26%)
Query: 76 CHAVLHGSCKCNNFNCHGKCNSCTKCES----SKCDGTNCVTICSFKCGNSKCDG-NKC- 129
C V G CK +G C + KC SK G +C ICS C N CD KC
Sbjct: 525 CSPVCSGGCK------NGFCVAPEKCSCDEGYSKETGNSCKPICSNGCENGFCDAPEKCS 578
Query: 130 ----------NGCTPKVTSPC---FQRCPQWCTCSK 152
N C+P + C F P C+C +
Sbjct: 579 CNDGYEMDGENRCSPVCSGGCKNGFCVAPGKCSCDE 614
Score = 37.4 bits (85), Expect = 0.14
Identities = 29/94 (30%), Positives = 36/94 (37%), Gaps = 21/94 (22%)
Query: 76 CHAVLHGSCKCNNFNCHGKCNSCTKCESSKCDGT--NCVTICSFKCGNSKCDG-NKC--- 129
C V G CK GKC+ C+ GT +C ICS C N CD KC
Sbjct: 327 CSPVCSGGCKNGFCVAPGKCS----CDEGYIKGTGNSCKPICSKGCENGFCDAPEKCSCN 382
Query: 130 --------NGCTPKVTSPC---FQRCPQWCTCSK 152
N C+P + C F P C+C +
Sbjct: 383 DGYEMDGENRCSPVCSGGCKNGFCVAPGKCSCDE 416
Score = 37.4 bits (85), Expect = 0.14
Identities = 30/92 (32%), Positives = 37/92 (39%), Gaps = 17/92 (18%)
Query: 76 CHAVLHGSCKCNNFNCHGKCNSCTKCESSKCDGTNCVTICSFKCGNSKCDG-NKC----- 129
C V G CK GKC SC + SK G +C ICS C N C+ KC
Sbjct: 723 CSPVCSGGCKNGFCVAPGKC-SCDE-GYSKETGNSCKPICSKGCENGFCEAPEKCSCNDG 780
Query: 130 ------NGCTPKVTSPC---FQRCPQWCTCSK 152
N C+P + C F P C+C +
Sbjct: 781 YEMDGENRCSPVCSGGCKNGFCIAPGKCSCDE 812
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.320 0.134 0.459
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 308,184,545
Number of Sequences: 2790947
Number of extensions: 13655609
Number of successful extensions: 167259
Number of sequences better than 10.0: 1470
Number of HSP's better than 10.0 without gapping: 326
Number of HSP's successfully gapped in prelim test: 1191
Number of HSP's that attempted gapping in prelim test: 157720
Number of HSP's gapped (non-prelim): 6559
length of query: 166
length of database: 848,049,833
effective HSP length: 118
effective length of query: 48
effective length of database: 518,718,087
effective search space: 24898468176
effective search space used: 24898468176
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 70 (31.6 bits)
Medicago: description of AC146569.8


