

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146567.9 + phase: 0
(80 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
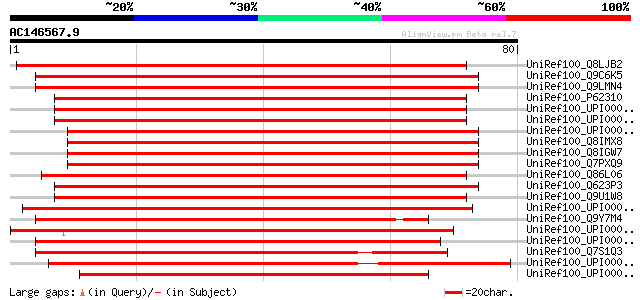
Sequences producing significant alignments: (bits) Value
UniRef100_Q8LJB2 P0505D12.24 protein [Oryza sativa] 134 7e-31
UniRef100_Q9C6K5 Sm-like protein [Arabidopsis thaliana] 127 7e-29
UniRef100_Q9LMN4 F16F4.12 protein [Arabidopsis thaliana] 125 3e-28
UniRef100_P62310 U6 snRNA-associated Sm-like protein LSm3 [Homo ... 114 5e-25
UniRef100_UPI00003AA8E6 UPI00003AA8E6 UniRef100 entry 114 5e-25
UniRef100_UPI00002DA7FE UPI00002DA7FE UniRef100 entry 114 6e-25
UniRef100_UPI0000431571 UPI0000431571 UniRef100 entry 112 2e-24
UniRef100_Q8IMX8 CG31184-PA [Drosophila melanogaster] 109 2e-23
UniRef100_Q8IGW7 RE11655p [Drosophila melanogaster] 109 2e-23
UniRef100_Q7PXQ9 ENSANGP00000011793 [Anopheles gambiae str. PEST] 108 3e-23
UniRef100_Q86L06 Similar to Arabidopsis thaliana (Mouse-ear cres... 107 1e-22
UniRef100_Q623P3 Hypothetical protein CBG01776 [Caenorhabditis b... 100 2e-20
UniRef100_Q9U1W8 Hypothetical protein Y62E10A.12 [Caenorhabditis... 99 3e-20
UniRef100_UPI00003C1491 UPI00003C1491 UniRef100 entry 93 2e-18
UniRef100_Q9Y7M4 Probable U6 snRNA-associated Sm-like protein LS... 91 9e-18
UniRef100_UPI0000235002 UPI0000235002 UniRef100 entry 88 5e-17
UniRef100_UPI000023CAA9 UPI000023CAA9 UniRef100 entry 84 9e-16
UniRef100_Q7S1Q3 Hypothetical protein [Neurospora crassa] 79 2e-14
UniRef100_UPI000021B27C UPI000021B27C UniRef100 entry 75 5e-13
UniRef100_UPI000042E1FB UPI000042E1FB UniRef100 entry 71 6e-12
>UniRef100_Q8LJB2 P0505D12.24 protein [Oryza sativa]
Length = 188
Score = 134 bits (336), Expect = 7e-31
Identities = 68/71 (95%), Positives = 68/71 (95%)
Query: 2 AGTEEESAVKEPLDLIRLSLDERIYVKLRSDRELRGKLHAYDQHLNMILGDVEEIVTTVE 61
A EEE AVKEPLDLIRLSLDERIYVKLRSDRELRGKLHAYDQHLNMILGDVEEIVTTVE
Sbjct: 8 AAAEEEIAVKEPLDLIRLSLDERIYVKLRSDRELRGKLHAYDQHLNMILGDVEEIVTTVE 67
Query: 62 IDDETYEEIVR 72
IDDETYEEIVR
Sbjct: 68 IDDETYEEIVR 78
>UniRef100_Q9C6K5 Sm-like protein [Arabidopsis thaliana]
Length = 98
Score = 127 bits (319), Expect = 7e-29
Identities = 62/70 (88%), Positives = 67/70 (95%)
Query: 5 EEESAVKEPLDLIRLSLDERIYVKLRSDRELRGKLHAYDQHLNMILGDVEEIVTTVEIDD 64
EEE+ V+EPLDLIRLSLDERIYVKLRSDRELRGKLHA+DQHLNMILGDVEE +TTVEIDD
Sbjct: 4 EEEATVREPLDLIRLSLDERIYVKLRSDRELRGKLHAFDQHLNMILGDVEETITTVEIDD 63
Query: 65 ETYEEIVRVS 74
ETYEEIVR +
Sbjct: 64 ETYEEIVRTT 73
>UniRef100_Q9LMN4 F16F4.12 protein [Arabidopsis thaliana]
Length = 97
Score = 125 bits (313), Expect = 3e-28
Identities = 58/70 (82%), Positives = 68/70 (96%)
Query: 5 EEESAVKEPLDLIRLSLDERIYVKLRSDRELRGKLHAYDQHLNMILGDVEEIVTTVEIDD 64
EE++ V+EPLDLIRLS++ERIYVKLRSDRELRGKLHA+DQHLNMILGDVEE++TT+EIDD
Sbjct: 4 EEDATVREPLDLIRLSIEERIYVKLRSDRELRGKLHAFDQHLNMILGDVEEVITTIEIDD 63
Query: 65 ETYEEIVRVS 74
ETYEEIVR +
Sbjct: 64 ETYEEIVRTT 73
>UniRef100_P62310 U6 snRNA-associated Sm-like protein LSm3 [Homo sapiens]
Length = 101
Score = 114 bits (286), Expect = 5e-25
Identities = 54/65 (83%), Positives = 62/65 (95%)
Query: 8 SAVKEPLDLIRLSLDERIYVKLRSDRELRGKLHAYDQHLNMILGDVEEIVTTVEIDDETY 67
+ V+EPLDLIRLSLDERIYVK+R+DRELRG+LHAYDQHLNMILGDVEE VTT+EID+ETY
Sbjct: 11 NTVEEPLDLIRLSLDERIYVKMRNDRELRGRLHAYDQHLNMILGDVEETVTTIEIDEETY 70
Query: 68 EEIVR 72
EEI +
Sbjct: 71 EEIYK 75
>UniRef100_UPI00003AA8E6 UPI00003AA8E6 UniRef100 entry
Length = 102
Score = 114 bits (286), Expect = 5e-25
Identities = 54/65 (83%), Positives = 62/65 (95%)
Query: 8 SAVKEPLDLIRLSLDERIYVKLRSDRELRGKLHAYDQHLNMILGDVEEIVTTVEIDDETY 67
+ V+EPLDLIRLSLDERIYVK+R+DRELRG+LHAYDQHLNMILGDVEE VTT+EID+ETY
Sbjct: 12 NTVEEPLDLIRLSLDERIYVKMRNDRELRGRLHAYDQHLNMILGDVEETVTTIEIDEETY 71
Query: 68 EEIVR 72
EEI +
Sbjct: 72 EEIYK 76
>UniRef100_UPI00002DA7FE UPI00002DA7FE UniRef100 entry
Length = 102
Score = 114 bits (285), Expect = 6e-25
Identities = 54/65 (83%), Positives = 62/65 (95%)
Query: 8 SAVKEPLDLIRLSLDERIYVKLRSDRELRGKLHAYDQHLNMILGDVEEIVTTVEIDDETY 67
+ V+EPLDLIRLSLDERIYVK+R+DRELRG+LHAYDQHLNMILGDVEE VTTVEID+ETY
Sbjct: 12 NTVEEPLDLIRLSLDERIYVKMRNDRELRGRLHAYDQHLNMILGDVEETVTTVEIDEETY 71
Query: 68 EEIVR 72
EE+ +
Sbjct: 72 EELYK 76
>UniRef100_UPI0000431571 UPI0000431571 UniRef100 entry
Length = 93
Score = 112 bits (280), Expect = 2e-24
Identities = 52/65 (80%), Positives = 61/65 (93%)
Query: 10 VKEPLDLIRLSLDERIYVKLRSDRELRGKLHAYDQHLNMILGDVEEIVTTVEIDDETYEE 69
VKEPLDLIRLSLDERIYVK+R++RELRG+LHAYDQHLNM+LG+ EE VTTVEID+ETYEE
Sbjct: 9 VKEPLDLIRLSLDERIYVKMRNERELRGRLHAYDQHLNMVLGEAEETVTTVEIDEETYEE 68
Query: 70 IVRVS 74
+ R +
Sbjct: 69 VYRTT 73
>UniRef100_Q8IMX8 CG31184-PA [Drosophila melanogaster]
Length = 103
Score = 109 bits (273), Expect = 2e-23
Identities = 49/65 (75%), Positives = 61/65 (93%)
Query: 10 VKEPLDLIRLSLDERIYVKLRSDRELRGKLHAYDQHLNMILGDVEEIVTTVEIDDETYEE 69
VKEPLDLIRLSLDE++YVK+R++RELRG+LHA+DQHLNM+LGD EE VTTVEID+ETYEE
Sbjct: 15 VKEPLDLIRLSLDEKVYVKMRNERELRGRLHAFDQHLNMVLGDAEETVTTVEIDEETYEE 74
Query: 70 IVRVS 74
+ + +
Sbjct: 75 VYKTA 79
>UniRef100_Q8IGW7 RE11655p [Drosophila melanogaster]
Length = 103
Score = 109 bits (273), Expect = 2e-23
Identities = 49/65 (75%), Positives = 61/65 (93%)
Query: 10 VKEPLDLIRLSLDERIYVKLRSDRELRGKLHAYDQHLNMILGDVEEIVTTVEIDDETYEE 69
VKEPLDLIRLSLDE++YVK+R++RELRG+LHA+DQHLNM+LGD EE VTTVEID+ETYEE
Sbjct: 15 VKEPLDLIRLSLDEKVYVKMRNERELRGRLHAFDQHLNMVLGDAEETVTTVEIDEETYEE 74
Query: 70 IVRVS 74
+ + +
Sbjct: 75 VYKTA 79
>UniRef100_Q7PXQ9 ENSANGP00000011793 [Anopheles gambiae str. PEST]
Length = 127
Score = 108 bits (271), Expect = 3e-23
Identities = 50/65 (76%), Positives = 60/65 (91%)
Query: 10 VKEPLDLIRLSLDERIYVKLRSDRELRGKLHAYDQHLNMILGDVEEIVTTVEIDDETYEE 69
VKEPLDLIRLSLDE+IYVK+R +RELRG+LHA+DQHLNM+LGD EE VTTVEID+ETYEE
Sbjct: 38 VKEPLDLIRLSLDEKIYVKMRYERELRGRLHAFDQHLNMVLGDAEETVTTVEIDEETYEE 97
Query: 70 IVRVS 74
+ + +
Sbjct: 98 VYKTT 102
>UniRef100_Q86L06 Similar to Arabidopsis thaliana (Mouse-ear cress). Sm-like
protein [Dictyostelium discoideum]
Length = 98
Score = 107 bits (266), Expect = 1e-22
Identities = 52/67 (77%), Positives = 58/67 (85%)
Query: 6 EESAVKEPLDLIRLSLDERIYVKLRSDRELRGKLHAYDQHLNMILGDVEEIVTTVEIDDE 65
EE V+EPLDLIRLSLDERI+VK+R DRELRGKLHAYDQHLNMIL DVEE + VE D+E
Sbjct: 5 EEGTVEEPLDLIRLSLDERIFVKMRQDRELRGKLHAYDQHLNMILSDVEETIKVVEKDEE 64
Query: 66 TYEEIVR 72
T EEI+R
Sbjct: 65 TDEEIIR 71
>UniRef100_Q623P3 Hypothetical protein CBG01776 [Caenorhabditis briggsae]
Length = 102
Score = 99.8 bits (247), Expect = 2e-20
Identities = 44/67 (65%), Positives = 60/67 (88%)
Query: 8 SAVKEPLDLIRLSLDERIYVKLRSDRELRGKLHAYDQHLNMILGDVEEIVTTVEIDDETY 67
+ V+EPLDL+RLSLDE++YVK+R+DRELRG+L A+DQHLNM+L +VEE +TT E+D++T+
Sbjct: 12 ATVEEPLDLLRLSLDEKVYVKMRNDRELRGRLRAFDQHLNMVLSEVEETITTREVDEDTF 71
Query: 68 EEIVRVS 74
EEI R S
Sbjct: 72 EEIYRQS 78
>UniRef100_Q9U1W8 Hypothetical protein Y62E10A.12 [Caenorhabditis elegans]
Length = 102
Score = 99.0 bits (245), Expect = 3e-20
Identities = 43/65 (66%), Positives = 59/65 (90%)
Query: 8 SAVKEPLDLIRLSLDERIYVKLRSDRELRGKLHAYDQHLNMILGDVEEIVTTVEIDDETY 67
+ V+EPLDL+RLSLDER+YVK+R+DRELRG+L A+DQHLNM+L +VEE +TT E+D++T+
Sbjct: 12 ATVEEPLDLLRLSLDERVYVKMRNDRELRGRLRAFDQHLNMVLSEVEETITTREVDEDTF 71
Query: 68 EEIVR 72
EEI +
Sbjct: 72 EEIYK 76
>UniRef100_UPI00003C1491 UPI00003C1491 UniRef100 entry
Length = 110
Score = 92.8 bits (229), Expect = 2e-18
Identities = 44/71 (61%), Positives = 54/71 (75%)
Query: 3 GTEEESAVKEPLDLIRLSLDERIYVKLRSDRELRGKLHAYDQHLNMILGDVEEIVTTVEI 62
G +V EP DLIRLS+ ER+YVKLR DRELRG LHAYD H+N+ILGDVEE + V++
Sbjct: 16 GPSPLGSVTEPFDLIRLSISERVYVKLRGDRELRGTLHAYDGHMNLILGDVEETIYVVDV 75
Query: 63 DDETYEEIVRV 73
+DE+ E VRV
Sbjct: 76 NDESGSETVRV 86
>UniRef100_Q9Y7M4 Probable U6 snRNA-associated Sm-like protein LSm3
[Schizosaccharomyces pombe]
Length = 93
Score = 90.5 bits (223), Expect = 9e-18
Identities = 44/62 (70%), Positives = 51/62 (81%), Gaps = 1/62 (1%)
Query: 5 EEESAVKEPLDLIRLSLDERIYVKLRSDRELRGKLHAYDQHLNMILGDVEEIVTTVEIDD 64
E AV EPLDL+RLSLDE +YVKLR DREL G+LHAYD+HLNM+LGD EEIVT + D+
Sbjct: 2 ESAQAVAEPLDLVRLSLDEIVYVKLRGDRELNGRLHAYDEHLNMVLGDAEEIVTIFD-DE 60
Query: 65 ET 66
ET
Sbjct: 61 ET 62
>UniRef100_UPI0000235002 UPI0000235002 UniRef100 entry
Length = 725
Score = 88.2 bits (217), Expect = 5e-17
Identities = 44/72 (61%), Positives = 54/72 (74%), Gaps = 2/72 (2%)
Query: 1 MAGTEEE--SAVKEPLDLIRLSLDERIYVKLRSDRELRGKLHAYDQHLNMILGDVEEIVT 58
MA TE S+V EPLDL+RLSLDE ++VKLR DREL+G+LHAYD H N++LGDVEE +
Sbjct: 1 MADTEGAGTSSVSEPLDLVRLSLDEIVFVKLRGDRELKGRLHAYDSHCNLVLGDVEETIY 60
Query: 59 TVEIDDETYEEI 70
VE D+ E I
Sbjct: 61 VVEEDENEQEHI 72
>UniRef100_UPI000023CAA9 UPI000023CAA9 UniRef100 entry
Length = 795
Score = 84.0 bits (206), Expect = 9e-16
Identities = 39/64 (60%), Positives = 50/64 (77%)
Query: 5 EEESAVKEPLDLIRLSLDERIYVKLRSDRELRGKLHAYDQHLNMILGDVEEIVTTVEIDD 64
EE V EPLDL+RL L+E ++VKLR DREL+GKLHAYD H N++LG+VEE + TV+ DD
Sbjct: 6 EEPHHVSEPLDLVRLLLNEVVFVKLRGDRELKGKLHAYDSHCNLVLGEVEETIYTVDEDD 65
Query: 65 ETYE 68
+ E
Sbjct: 66 DDDE 69
>UniRef100_Q7S1Q3 Hypothetical protein [Neurospora crassa]
Length = 168
Score = 79.3 bits (194), Expect = 2e-14
Identities = 38/65 (58%), Positives = 49/65 (74%), Gaps = 2/65 (3%)
Query: 5 EEESAVKEPLDLIRLSLDERIYVKLRSDRELRGKLHAYDQHLNMILGDVEEIVTTVEIDD 64
E+ +V EP+DL+RL LDE + VKLR DREL+G+LHAYD H N++LG+VEE T +DD
Sbjct: 6 EDAGSVSEPMDLVRLLLDEVVCVKLRGDRELKGRLHAYDSHCNLVLGEVEE--TIYVVDD 63
Query: 65 ETYEE 69
E EE
Sbjct: 64 EDTEE 68
>UniRef100_UPI000021B27C UPI000021B27C UniRef100 entry
Length = 78
Score = 74.7 bits (182), Expect = 5e-13
Identities = 37/73 (50%), Positives = 51/73 (69%), Gaps = 3/73 (4%)
Query: 7 ESAVKEPLDLIRLSLDERIYVKLRSDRELRGKLHAYDQHLNMILGDVEEIVTTVEIDDET 66
E+ EP L+RL LDE ++VKLR DREL+G+LHAYD H N++LGDVEE T I DE
Sbjct: 6 EAQANEPFGLVRLLLDEVVFVKLRGDRELKGRLHAYDSHCNLVLGDVEE---THYIQDED 62
Query: 67 YEEIVRVSHNPNH 79
E ++V ++ ++
Sbjct: 63 DESELKVRNSDDY 75
>UniRef100_UPI000042E1FB UPI000042E1FB UniRef100 entry
Length = 95
Score = 71.2 bits (173), Expect = 6e-12
Identities = 33/55 (60%), Positives = 43/55 (78%)
Query: 12 EPLDLIRLSLDERIYVKLRSDRELRGKLHAYDQHLNMILGDVEEIVTTVEIDDET 66
EPLDLIRLSLDE+++VK R +REL+G L+AYD H+NM+LG+VEE E +T
Sbjct: 6 EPLDLIRLSLDEQVFVKCRGNRELKGTLYAYDPHMNMVLGNVEETYYEEESKSDT 60
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.313 0.134 0.360
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 126,700,931
Number of Sequences: 2790947
Number of extensions: 4284788
Number of successful extensions: 10482
Number of sequences better than 10.0: 234
Number of HSP's better than 10.0 without gapping: 204
Number of HSP's successfully gapped in prelim test: 30
Number of HSP's that attempted gapping in prelim test: 10277
Number of HSP's gapped (non-prelim): 234
length of query: 80
length of database: 848,049,833
effective HSP length: 56
effective length of query: 24
effective length of database: 691,756,801
effective search space: 16602163224
effective search space used: 16602163224
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 68 (30.8 bits)
Medicago: description of AC146567.9


