

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146567.10 + phase: 0
(393 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
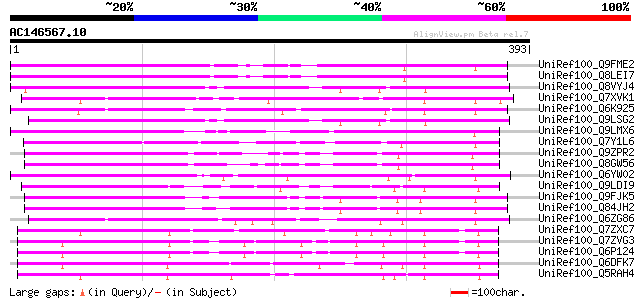
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9FME2 Similarity to RNA-binding protein [Arabidopsis ... 253 5e-66
UniRef100_Q8LEI7 Ras-GTPase-activating protein SH3-domain bindin... 253 9e-66
UniRef100_Q8VYJ4 AT3g25150/MJL12_9 [Arabidopsis thaliana] 244 2e-63
UniRef100_Q7XVK1 OSJNBa0069D17.2 protein [Oryza sativa] 244 4e-63
UniRef100_Q6K925 Putative Ras-GTPase-activating protein binding ... 239 7e-62
UniRef100_Q9LSG2 RNA-binding protein-like [Arabidopsis thaliana] 236 6e-61
UniRef100_Q9LMX6 F21F23.16 protein [Arabidopsis thaliana] 226 7e-58
UniRef100_Q7Y1L6 Putative GAP SH3 binding protein [Oryza sativa] 220 6e-56
UniRef100_Q9ZPR2 Hypothetical protein At2g03640 [Arabidopsis tha... 213 7e-54
UniRef100_Q8GW56 Hypothetical protein At2g03640/F19B11.9 [Arabid... 213 1e-53
UniRef100_Q6YW02 Putative Ras-GTPase activating protein SH3 doma... 205 2e-51
UniRef100_Q9LDI9 Putative RNA-binding protein; 63745-61607 [Arab... 195 2e-48
UniRef100_Q9FJK5 RNA-binding protein-like [Arabidopsis thaliana] 157 4e-37
UniRef100_Q84JH2 Putative NTF2-containing RNA-binding protein [A... 156 8e-37
UniRef100_Q6ZG86 Putative Ras-GTPase activating protein SH3 doma... 155 1e-36
UniRef100_Q7ZXC7 G3bp-prov protein [Xenopus laevis] 124 4e-27
UniRef100_Q7ZVG3 Ras-GTPase-activating protein SH3-domain-bindin... 120 5e-26
UniRef100_Q6P124 Ras-GTPase-activating protein SH3-domain-bindin... 120 7e-26
UniRef100_Q6DFK7 MGC81268 protein [Xenopus laevis] 119 1e-25
UniRef100_Q5RAH4 Hypothetical protein DKFZp459E0328 [Pongo pygma... 119 2e-25
>UniRef100_Q9FME2 Similarity to RNA-binding protein [Arabidopsis thaliana]
Length = 460
Score = 253 bits (647), Expect = 5e-66
Identities = 161/386 (41%), Positives = 222/386 (56%), Gaps = 37/386 (9%)
Query: 1 MAVPEAVQTPTPQMVGNAFVEQYYSILHRDPDQVHRFYHDSSVMSRPEEDGTMTTVTTTA 60
MA EA +P ++VG AFVEQYY ILH+ P VHRFY DSS ++RP+ G +TTVTT
Sbjct: 1 MAQQEASPSPGAEVVGRAFVEQYYHILHQSPGLVHRFYQDSSFLTRPDVTGAVTTVTTMQ 60
Query: 61 EIDKKIQSLEYTSFRVEVLSADAQPSYNNGVMVVVTGCLTGTDNIKRKFAQSFFLAPQDK 120
I+ KI SL+Y + E+ +ADAQ S+ GV+V+VTG LTG DN+++KF+QSFFLAPQDK
Sbjct: 61 AINDKILSLKYEDYTAEIETADAQESHERGVIVLVTGRLTGNDNVRKKFSQSFFLAPQDK 120
Query: 121 GFYVLNDVFRYVDAYK-SIDIESVPANDADESAPSEAIITPEPEPV-HVPEVIPPTQTVI 178
G++VLNDVFR+++ + + SVP N +A I PE V H PEV P
Sbjct: 121 GYFVLNDVFRFLEEKEVTAQARSVPINGTTRDV--QAPIEPERVVVSHEPEVEPE----- 173
Query: 179 PTAQTVIPPTQTVIADTETIISKEVSLPLENGKLSVTENVIPVNHVKESSHHVKEPEQPT 238
P A I + + EV P + + V +V P+ EP
Sbjct: 174 PVAS---------IEEEDLDNVAEVYDPSDKDE-GVVVDVEPI-----------EPPTQI 212
Query: 239 SIEKVASNTQEDTPKKSFASIVNALKDNSAP-FHLRASPAKPAVHPPRVHSVPAPEAPTP 297
S ++ S Q D PK S+ASI+ +K + AP H+ + +PA ++ + PA A P
Sbjct: 213 SHNEILSVPQGDAPKHSYASILKQMKSSPAPTTHVARNKPRPAPVNQKLTAPPAEPAARP 272
Query: 298 ----NMDIPLEKNNENAGRAHAIFVANLPMSATVEQLDRAFKKFGPIKRDGIQVRSNK-- 351
+ ++P + + H+I+V NLP +T QL+ FK FG IK +GIQVRSNK
Sbjct: 273 EASAHENVPNSSHVDVEDDGHSIYVRNLPFDSTPTQLEEVFKNFGAIKHEGIQVRSNKQQ 332
Query: 352 GSCFGFVEFESAASMQSALEVWSISI 377
G CFGFVEFE+++ QSALE ++I
Sbjct: 333 GFCFGFVEFETSSGKQSALEASPVTI 358
>UniRef100_Q8LEI7 Ras-GTPase-activating protein SH3-domain binding protein-like
[Arabidopsis thaliana]
Length = 459
Score = 253 bits (645), Expect = 9e-66
Identities = 161/385 (41%), Positives = 221/385 (56%), Gaps = 36/385 (9%)
Query: 1 MAVPEAVQTPTPQMVGNAFVEQYYSILHRDPDQVHRFYHDSSVMSRPEEDGTMTTVTTTA 60
MA EA +P ++VG AFVEQYY ILH+ P VHRFY DSS ++RP+ G +TTVTT
Sbjct: 1 MAQQEASPSPGAEVVGRAFVEQYYHILHQSPGLVHRFYQDSSFLTRPDVTGAVTTVTTMQ 60
Query: 61 EIDKKIQSLEYTSFRVEVLSADAQPSYNNGVMVVVTGCLTGTDNIKRKFAQSFFLAPQDK 120
I+ KI SL+Y + E+ +ADAQ S+ GV+V VTG LTG DN+++KF+QSFFLAPQDK
Sbjct: 61 AINDKILSLKYEDYTAEIETADAQESHERGVIVPVTGRLTGNDNVRKKFSQSFFLAPQDK 120
Query: 121 GFYVLNDVFRYVDAYK-SIDIESVPANDADESAPSEAIITPEPEPV-HVPEVIPPTQTVI 178
G++VLNDVFR+++ + + SVP N +A I PE V H PEV P
Sbjct: 121 GYFVLNDVFRFLEEKEVTAQARSVPINGTTRDV--QAPIEPERVVVSHEPEVEPE----- 173
Query: 179 PTAQTVIPPTQTVIADTETIISKEVSLPLENGKLSVTENVIPVNHVKESSHHVKEPEQPT 238
P A I + + EV P + + V +V P+ EP
Sbjct: 174 PVAS---------IEEEDLDNVAEVYDPSDKDE-GVVVDVEPI-----------EPPTQI 212
Query: 239 SIEKVASNTQEDTPKKSFASIVNALKDNSAP-FHLRASPAKPAVHPPRVHSVPAPEAPTP 297
S ++ S Q D PK S+ASI+ +K + AP H+ + +PA ++ + PA A P
Sbjct: 213 SHNEILSVPQGDAPKHSYASILKQMKSSPAPTTHVARNKPRPAPVNQKLTAPPAEPAARP 272
Query: 298 ----NMDIPLEKNNENAGRAHAIFVANLPMSATVEQLDRAFKKFGPIKRDGIQVRSNK-G 352
+ ++P + + H+I+V NLP +T QL+ FK FG IK +GIQVRSNK G
Sbjct: 273 EASAHENVPNSSHVDVEDDGHSIYVRNLPFDSTPTQLEEVFKNFGAIKHEGIQVRSNKQG 332
Query: 353 SCFGFVEFESAASMQSALEVWSISI 377
CFGFVEFE+++ QSALE ++I
Sbjct: 333 FCFGFVEFETSSGKQSALEASPVTI 357
>UniRef100_Q8VYJ4 AT3g25150/MJL12_9 [Arabidopsis thaliana]
Length = 488
Score = 244 bits (624), Expect = 2e-63
Identities = 148/394 (37%), Positives = 214/394 (53%), Gaps = 34/394 (8%)
Query: 1 MAVPEAVQTP----TPQMVGNAFVEQYYSILHRDPDQVHRFYHDSSVMSRPEEDGTMTTV 56
MA+ A Q P TP MVGNAFV QYY ILH+ P+ VHRFY + S + RPEE+G M+
Sbjct: 1 MAMLGAQQVPAAACTPDMVGNAFVPQYYHILHQSPEHVHRFYQEISKLGRPEENGLMSIT 60
Query: 57 TTTAEIDKKIQSLEYTSFRVEVLSADAQPSYNNGVMVVVTGCLTGTDNIKRKFAQSFFLA 116
+T IDKKI +L Y E+ + D Q S+ G +V+VTG LTG D+++R F+Q+FFLA
Sbjct: 61 STLQAIDKKIMALGYGVISAEIATVDTQESHGGGYIVLVTGYLTGKDSVRRTFSQTFFLA 120
Query: 117 PQDKGFYVLNDVFRYVDAYKSIDIESVPANDADESAPSEAIITPEPEPVHVPEVIPPTQT 176
PQ+ G++VLND+FR++D + +P N+ AP + + IP
Sbjct: 121 PQETGYFVLNDMFRFIDEGTVVHGNQIPVNNV--QAPVNTY-----QDTAAAKEIPDDFV 173
Query: 177 VIPTAQTVIPPTQTVIADTETIISKEVSLPLENGKLSVTENVIPVNHVKESSHHVKEPEQ 236
Q QT + ++V P E+ ++S E + V E+
Sbjct: 174 QEKYVQENHAVKQTEVLSKSINEPEKVFTPSEDEQVSAAEEALVTETVNEA--------- 224
Query: 237 PTSIEKVASNTQE--DTPKKSFASIVNALKDNSAPFHLRASPAK--------PAVHPPRV 286
P ++KV + + PK+S+ASIV +K+N+AP +P K A+H P
Sbjct: 225 PIEVQKVGESDSRTGEIPKRSYASIVKVMKENAAPMSASRTPTKVEPKKQEDQAIHIPL- 283
Query: 287 HSVPAPEAPTPNMDIPLEKNNENAGRA--HAIFVANLPMSATVEQLDRAFKKFGPIKRDG 344
P E ++ + +NN+ RA +I++ LP+ AT L+ F+KFG I+ +G
Sbjct: 284 -PTPLSEKSDSGANVAVNENNQENERALGPSIYLKGLPLDATPALLENEFQKFGLIRTNG 342
Query: 345 IQVRSNKGSCFGFVEFESAASMQSALEVWSISIN 378
IQVRS KG CFGFVEFESA+SMQSA+E + +N
Sbjct: 343 IQVRSQKGFCFGFVEFESASSMQSAIEASPVMLN 376
>UniRef100_Q7XVK1 OSJNBa0069D17.2 protein [Oryza sativa]
Length = 486
Score = 244 bits (622), Expect = 4e-63
Identities = 160/391 (40%), Positives = 215/391 (54%), Gaps = 34/391 (8%)
Query: 10 PTPQMVGNAFVEQYYSILHRDPDQVHRFYHDSSVMSRPEEDGT--MTTVTTTAEIDKKIQ 67
P+ Q+VGNAFV QYY+ILH+ PD VHRFY D S + RP M TVTT I+ KI
Sbjct: 18 PSAQVVGNAFVHQYYNILHQSPDLVHRFYQDGSRIGRPASPAAAEMDTVTTMEAINAKIV 77
Query: 68 SLEYTSFRVEVLSADAQPSYNNGVMVVVTGCLTGTDNIKRKFAQSFFLAPQDKGFYVLND 127
S++ R E+ + DAQ S GV V+VTG LTG+D+++R+F+QSFFLAPQ+KG++VLND
Sbjct: 78 SMDIV--RAEIKAVDAQESLGGGVTVLVTGHLTGSDDVRREFSQSFFLAPQEKGYFVLND 135
Query: 128 VFRYVDAYKSIDIESVPANDADESAPSEAIITPEPEPVHVPEVIPPTQTVIPTAQTVIPP 187
+ RYV ++E P + + S P + +P+ V P T +P Q
Sbjct: 136 ILRYVGGEGDQEVE--PEPELELSFPP----SQQPDSVPAPSA---NGTSVPREQEAFSQ 186
Query: 188 TQTVIAD------TETIISKEVSLPLENGKLSVTENVIPVNHVKESSHHVKEPEQPTSIE 241
+ +AD + +EV P N + V E + E ++V
Sbjct: 187 PEQHVADPAPNAQEADLNGEEVYNPPNNTEGPVVEETPIPEVIDEVPNNVAVAMPTPPAP 246
Query: 242 KVASNTQEDTPKKSFASIVNALKDNSAPFHLRASPAKPAVHPPRVHSVPAPEAPTPNMDI 301
A QE+ PKKS+ASIV +K+ P + A P++PA PP+ AP P D
Sbjct: 247 APAPVPQEEAPKKSYASIVKVMKE--IPPQISAIPSRPA--PPKQEKQVAPAPVAPVADA 302
Query: 302 PLEKNNENAGR-------AHAIFVANLPMSATVEQLDRAFKKFGPIKRDGIQVRSNK--G 352
P N + AHAI+V NLP+SAT EQL+ AFKKFG IK DGIQVRS+K G
Sbjct: 303 PTFSPNPESSNIQEAEVDAHAIYVRNLPLSATPEQLEEAFKKFGAIKPDGIQVRSHKIQG 362
Query: 353 SCFGFVEFESAASMQSAL--EVWSISINSCY 381
C+GFVEFE +S+QSA+ +IS CY
Sbjct: 363 FCYGFVEFEDPSSVQSAIAGSPVTISDRQCY 393
>UniRef100_Q6K925 Putative Ras-GTPase-activating protein binding protein 1 [Oryza
sativa]
Length = 480
Score = 239 bits (611), Expect = 7e-62
Identities = 159/395 (40%), Positives = 216/395 (54%), Gaps = 32/395 (8%)
Query: 1 MAVPEAVQTPTPQMVGNAFVEQYYSILHRDPDQVHRFYHDSSVMSRPEED--GTMTTVTT 58
MA P+A +P+ Q+VGNAFV+QYY ILH+ PD V+RFY D+S + RP D G M +VTT
Sbjct: 1 MAAPQASPSPSAQVVGNAFVQQYYQILHQSPDLVYRFYQDASRLGRPPADRYGDMVSVTT 60
Query: 59 TAEIDKKIQSLEYTSFRVEVLSADAQPSYNNGVMVVVTGCLTGTDNIKRKFAQSFFLAPQ 118
I++KI +++ + R E+ + D+Q S GV V+VTG LT D + R+F+QSFFLAPQ
Sbjct: 61 MEAINEKIMAMDMS--RAEIKTVDSQESLGGGVTVLVTGHLTVRDGVCREFSQSFFLAPQ 118
Query: 119 DKGFYVLNDVFRYVDAYKSIDIESVPANDADESAPSEAIITPEPEPVHVPEVIPPTQTVI 178
+KG++VLND+FRYV D + A A E P + P P+ P Q
Sbjct: 119 EKGYFVLNDMFRYVG-----DGPTPAAAAAAEVQPEADAVAP---PLANGTATAPLQPAA 170
Query: 179 PTAQTVIPPTQTVIADTETIISKEVSL---PLENGKLSVTENVIPVNHVKESSHHVKEPE 235
P + V+ + +E + PLE + E V E + V
Sbjct: 171 PDYDAMPHEEPDVVENVAVPPEEEEEVYNPPLEEVEGGAVEE---EQSVPEVINEVPNNV 227
Query: 236 QPTSIEKVASNTQEDTPKKSFASIVNALKDNSAPFHLRASPAKPAVHP----PRVHSVPA 291
P A + E+ PKKS+ASIV +K+ P + A+ PA P P S PA
Sbjct: 228 VPVVAPADAPVSHEEAPKKSYASIVKVMKEAPVPAPIPATRPAPAARPAPPKPEKQS-PA 286
Query: 292 PEAPTPNMD-IPLEKNNENAGR------AHAIFVANLPMSATVEQLDRAFKKFGPIKRDG 344
P AP P D P N E++ AHAI+V +LP++AT QL+ FKKFG IK DG
Sbjct: 287 PPAPAPVADATPFSSNAESSNTHEPEVDAHAIYVRSLPLNATTTQLEDEFKKFGTIKPDG 346
Query: 345 IQVRSNK--GSCFGFVEFESAASMQSALEVWSISI 377
IQVRS+K G C+GFVEFE A ++QSA+E + I
Sbjct: 347 IQVRSHKIQGFCYGFVEFEEATAVQSAIEASPVMI 381
>UniRef100_Q9LSG2 RNA-binding protein-like [Arabidopsis thaliana]
Length = 473
Score = 236 bits (603), Expect = 6e-61
Identities = 140/376 (37%), Positives = 205/376 (54%), Gaps = 30/376 (7%)
Query: 15 VGNAFVEQYYSILHRDPDQVHRFYHDSSVMSRPEEDGTMTTVTTTAEIDKKIQSLEYTSF 74
VGNAFV QYY ILH+ P+ VHRFY + S + RPEE+G M+ +T IDKKI +L Y
Sbjct: 4 VGNAFVPQYYHILHQSPEHVHRFYQEISKLGRPEENGLMSITSTLQAIDKKIMALGYGVI 63
Query: 75 RVEVLSADAQPSYNNGVMVVVTGCLTGTDNIKRKFAQSFFLAPQDKGFYVLNDVFRYVDA 134
E+ + D Q S+ G +V+VTG LTG D+++R F+Q+FFLAPQ+ G++VLND+FR++D
Sbjct: 64 SAEIATVDTQESHGGGYIVLVTGYLTGKDSVRRTFSQTFFLAPQETGYFVLNDMFRFIDE 123
Query: 135 YKSIDIESVPANDADESAPSEAIITPEPEPVHVPEVIPPTQTVIPTAQTVIPPTQTVIAD 194
+ +P N+ AP + + IP Q QT +
Sbjct: 124 GTVVHGNQIPVNNV--QAPVNTY-----QDTAAAKEIPDDFVQEKYVQENHAVKQTEVLS 176
Query: 195 TETIISKEVSLPLENGKLSVTENVIPVNHVKESSHHVKEPEQPTSIEKVASNTQE--DTP 252
++V P E+ ++S E + V E+ P ++KV + + P
Sbjct: 177 KSINEPEKVFTPSEDEQVSAAEEALVTETVNEA---------PIEVQKVGESDSRTGEIP 227
Query: 253 KKSFASIVNALKDNSAPFHLRASPAK--------PAVHPPRVHSVPAPEAPTPNMDIPLE 304
K+S+ASIV +K+N+AP +P K A+H P P E ++ +
Sbjct: 228 KRSYASIVKVMKENAAPMSASRTPTKVEPKKQEDQAIHIPL--PTPLSEKSDSGANVAVN 285
Query: 305 KNNENAGRA--HAIFVANLPMSATVEQLDRAFKKFGPIKRDGIQVRSNKGSCFGFVEFES 362
+NN+ RA +I++ LP+ AT L+ F+KFG I+ +GIQVRS KG CFGFVEFES
Sbjct: 286 ENNQENERALGPSIYLKGLPLDATPALLENEFQKFGLIRTNGIQVRSQKGFCFGFVEFES 345
Query: 363 AASMQSALEVWSISIN 378
A+SMQSA+E + +N
Sbjct: 346 ASSMQSAIEASPVMLN 361
>UniRef100_Q9LMX6 F21F23.16 protein [Arabidopsis thaliana]
Length = 428
Score = 226 bits (577), Expect = 7e-58
Identities = 144/374 (38%), Positives = 200/374 (52%), Gaps = 41/374 (10%)
Query: 1 MAVPEAVQTPTPQMVGNAFVEQYYSILHRDPDQVHRFYHDSSVMSRPEEDGTMTTVTTTA 60
MA+ P +GN+FVE+YY++L++ P QVH+FY D SV+ RP DG M +V +
Sbjct: 1 MALESNAPVVDPNTIGNSFVEKYYNLLYKSPSQVHQFYLDDSVLGRPGSDGEMVSVKSLK 60
Query: 61 EIDKKIQSLEYTSFRVEVLSADAQPSYNNGVMVVVTGCLTGTDNIKRKFAQSFFLAPQDK 120
I+++I S +Y ++++L+AD+Q SY NGV+ +VTG LT + + +F+QSFFL P +
Sbjct: 61 AINEQIMSFDYEISKIQILTADSQASYMNGVVTLVTGLLTVKEGQRMRFSQSFFLVPLNG 120
Query: 121 GFYVLNDVFRYVDAYKSIDIESVPANDADESAPSEAIITPEPEPVHVPEVIPPTQTVIPT 180
++VLNDVFRYV ADE EA E E V +P+V+ PT+ V
Sbjct: 121 SYFVLNDVFRYV---------------ADEIVEPEA-NKKEVEEV-IPQVVQPTEQVDEV 163
Query: 181 AQTVIPPTQTVIADTETIISKEVSLPLENGKLSVTENVIPVNHVKESSHHVKEPEQPTSI 240
A+ V PTQ A TEN + ++ H K E
Sbjct: 164 AEPVTIPTQQPEAK------------------QTTENTVKKPERAVANGHPKTQEDNVVN 205
Query: 241 EKVASNTQEDTPKKSFASIVNALKDNSAPFHLRASPAKPAVHP-PRVHSVPAPEAPTPNM 299
+K + D PKKSFA IV L N A F+ +ASPAKP P + + +AP P
Sbjct: 206 DK---SNGVDAPKKSFAHIVQDLAQNGATFNAKASPAKPKSKPVTKPSAARESKAPAPVS 262
Query: 300 DIPLEKNNENAGRAHAIFVANLPMSATVEQLDRAFKKFGPIKRDGIQVRSN--KGSCFGF 357
+ + + IFVANL M AT EQL+ FK FG I +DGIQVRS KG+CFGF
Sbjct: 263 EHSSAATIDQQAEGYTIFVANLLMDATPEQLNETFKGFGAITKDGIQVRSYRLKGNCFGF 322
Query: 358 VEFESAASMQSALE 371
V F SA +++ L+
Sbjct: 323 VTFASAEAVKLVLQ 336
>UniRef100_Q7Y1L6 Putative GAP SH3 binding protein [Oryza sativa]
Length = 488
Score = 220 bits (560), Expect = 6e-56
Identities = 138/374 (36%), Positives = 207/374 (54%), Gaps = 40/374 (10%)
Query: 11 TPQMVGNAFVEQYYSILHRDPDQVHRFYHDSSVMSRPEEDGTMTTVTTTAEIDKKIQSLE 70
+PQ++ AFV+QYY ILH PDQV++FY D+S++ RP+ +G M V+TTA+I+K I S++
Sbjct: 13 SPQVISGAFVQQYYHILHETPDQVYKFYQDASIVGRPDSNGVMKYVSTTADINKIILSMD 72
Query: 71 YTSFRVEVLSADAQPSYNNGVMVVVTGCLTGTDNIKRKFAQSFFLAPQDKGFY-VLNDVF 129
++++ E+ +ADAQ S+ +GV++VVTG LT ++ I R+F QSFFLAPQ+ G Y VLND+F
Sbjct: 73 FSNYLTEIETADAQLSHQDGVLIVVTGSLT-SEGICRRFTQSFFLAPQESGGYVVLNDIF 131
Query: 130 RYVDAYKSIDIESVPANDADESAPSEAIITPEPEPVHVPEVIPPTQTVIPTAQTVIPPTQ 189
R++ + I V + + + + + + +P P + P +
Sbjct: 132 RFIVERPPVAISQV-SQENENNQNTATLPETDPNPAGDGMISEPV------------AVE 178
Query: 190 TVIADTETIISKEVSLPLENGKLSVTENVIPVNHVKESSHHVKEPEQPTSIEKVASNTQE 249
+A+ E S +EN + E PV KE + P Q+
Sbjct: 179 NNVAEGEVTNSTVDGTSIENNATAAVEP--PVQMTKEEPRKISVAAPPPP-------AQK 229
Query: 250 DTPKKSFASIVNALKDNSAPFHLRASPAKPAVHPPRVHSVPAPEAP----------TPNM 299
D KKS+ASIV +K+ S ++ PA V V +V A E P TPN
Sbjct: 230 DVTKKSYASIVKVMKEVSLTPVVKPKPAPKHV----VKTVEASEKPSVKSSQTVEITPND 285
Query: 300 DIPLEKNNENAGRAHAIFVANLPMSATVEQLDRAFKKFGPIKRDGIQVRSNK--GSCFGF 357
+ E N N + +++FV +LP + TV+ ++ FKKFG IK GIQVR+NK CFGF
Sbjct: 286 NNDAENNTSNDEQGYSVFVKSLPHNVTVQTVEEEFKKFGAIKPGGIQVRNNKIDRFCFGF 345
Query: 358 VEFESAASMQSALE 371
+EFES SMQ+A+E
Sbjct: 346 IEFESQQSMQAAIE 359
>UniRef100_Q9ZPR2 Hypothetical protein At2g03640 [Arabidopsis thaliana]
Length = 423
Score = 213 bits (542), Expect = 7e-54
Identities = 137/365 (37%), Positives = 202/365 (54%), Gaps = 46/365 (12%)
Query: 12 PQMVGNAFVEQYYSILHRDPDQVHRFYHDSSVMSRPEEDGTMTTVTTTAEIDKKIQSLEY 71
PQ VGN FV++YY+ L+ +VH+FY + S++SRP DG + T+ + I+ +I S++Y
Sbjct: 12 PQFVGNGFVQEYYNHLYDSTSEVHKFYLEDSMISRPGLDGEIVTIKSLKGINDQIMSIDY 71
Query: 72 TSFRVEVLSADAQPSYNNGVMVVVTGCLTGTDNIKRKFAQSFFLAPQDKGFYVLNDVFRY 131
S R+E+L+AD+Q + NGV+ +VTG + G D +RKF+QSFFL ++ ++VLND FRY
Sbjct: 72 KSSRIEILTADSQSTLKNGVVTLVTGLVIGNDGGRRKFSQSFFLVSRNGSYFVLNDTFRY 131
Query: 132 V-DAYKSIDIESVPANDADESAPSEAIITPEPEPVHVPEVIPPTQTVIPTAQTVIPPTQT 190
V D + +E + +ES + AI T EP V VI PT+
Sbjct: 132 VSDEF----VEPEATKEVEESQSTNAI-TAEPANESVEAVIVPTEA-------------- 172
Query: 191 VIADTETIISKEVSLPLENGKLSVTENVIPVNHVKESSHHVKEPEQPTSIEKVASNTQED 250
+T ++K S + NG V E + V E+S K E + QE+
Sbjct: 173 -----KTTVTKPASA-IPNGHAKVPEEKV----VNENSSLPKAAE---------AKLQEE 213
Query: 251 TPKKSFASIVNALKDNSAPFHLRASPAKPAVHPPRVHSVPAPE--APTPNMDIPLEKNNE 308
PKKSFA IV +L ++ ++ASP K P V APE AP+P ++ +
Sbjct: 214 VPKKSFALIVQSLAQSAGTLQVKASPVK---RKPVEKPVAAPERKAPSPIRKQASAESIK 270
Query: 309 NAGRAHAIFVANLPMSATVEQLDRAFKKFGPIKRDGIQVRS--NKGSCFGFVEFESAASM 366
+ +IFVANLPM AT+EQL FK FG I++DGIQVRS K +C GFV FE+ ++
Sbjct: 271 PQAQGSSIFVANLPMDATIEQLYETFKSFGAIRKDGIQVRSYPEKKNCIGFVAFENGEAV 330
Query: 367 QSALE 371
++ +
Sbjct: 331 KNVFQ 335
>UniRef100_Q8GW56 Hypothetical protein At2g03640/F19B11.9 [Arabidopsis thaliana]
Length = 422
Score = 213 bits (541), Expect = 1e-53
Identities = 136/365 (37%), Positives = 202/365 (55%), Gaps = 47/365 (12%)
Query: 12 PQMVGNAFVEQYYSILHRDPDQVHRFYHDSSVMSRPEEDGTMTTVTTTAEIDKKIQSLEY 71
PQ VGN FV++YY+ L+ +VH+FY + S++SRP DG + T+ + I+ +I S++Y
Sbjct: 12 PQFVGNGFVQEYYNHLYDSTSEVHKFYLEDSMISRPGLDGEIVTIKSLKGINDQIMSIDY 71
Query: 72 TSFRVEVLSADAQPSYNNGVMVVVTGCLTGTDNIKRKFAQSFFLAPQDKGFYVLNDVFRY 131
S R+E+L+AD+Q + NGV+ +VTG + G D +RKF+QSFFL ++ ++VLND FRY
Sbjct: 72 KSSRIEILTADSQSTLKNGVVTLVTGLVIGNDGGRRKFSQSFFLVSRNGSYFVLNDTFRY 131
Query: 132 V-DAYKSIDIESVPANDADESAPSEAIITPEPEPVHVPEVIPPTQTVIPTAQTVIPPTQT 190
V D + +E + +ES + AI P E V + VI PT
Sbjct: 132 VSDEF----VEPEATKEVEESQSTNAITEPANESV----------------EAVIVPT-- 169
Query: 191 VIADTETIISKEVSLPLENGKLSVTENVIPVNHVKESSHHVKEPEQPTSIEKVASNTQED 250
+ +T ++K S + NG V E + V E+S K E + QE+
Sbjct: 170 ---EAKTTVTKPAS-AIPNGHAKVPEEKV----VNENSSLPKAAE---------AKLQEE 212
Query: 251 TPKKSFASIVNALKDNSAPFHLRASPAKPAVHPPRVHSVPAPE--APTPNMDIPLEKNNE 308
PKKSFA IV +L ++ ++ASP K P V APE AP+P ++ +
Sbjct: 213 VPKKSFALIVQSLAQSAGTLQVKASPVK---RKPVEKPVAAPERKAPSPIRKQASAESIK 269
Query: 309 NAGRAHAIFVANLPMSATVEQLDRAFKKFGPIKRDGIQVRS--NKGSCFGFVEFESAASM 366
+ +IFVANLPM AT+EQL FK FG I++DGIQVRS K +C GFV FE+ ++
Sbjct: 270 PQAQGSSIFVANLPMDATIEQLYETFKSFGAIRKDGIQVRSYPEKKNCIGFVAFENGEAV 329
Query: 367 QSALE 371
++ +
Sbjct: 330 KNVFQ 334
>UniRef100_Q6YW02 Putative Ras-GTPase activating protein SH3 domain-binding protein 2
[Oryza sativa]
Length = 569
Score = 205 bits (522), Expect = 2e-51
Identities = 138/395 (34%), Positives = 215/395 (53%), Gaps = 27/395 (6%)
Query: 1 MAVPEAVQTPTPQMVGNAFVEQYYSILHRDPDQVHRFYHDSSVMSRPEEDGTMTTVTTTA 60
M V E+V +PQM+GNAFV+QYY++LH P QV +FYHDSS + RP+ +GTMT+VTT
Sbjct: 3 MQVGESVAPLSPQMIGNAFVQQYYNVLHSSPGQVCKFYHDSSTLGRPDSNGTMTSVTTLT 62
Query: 61 EIDKKIQSLEYTSFRVEVLSADAQPSYNNGVMVVVTGCLTGTDNIKRKFAQSFFLAPQDK 120
I+ + S +++S +++ + DAQ S N GV ++VTG + ++ +F+QSFFLAPQ+
Sbjct: 63 AINDEFLSTDFSSCLIKLENVDAQLSLNGGVHILVTGSIGHNGTMRHRFSQSFFLAPQES 122
Query: 121 -GFYVLNDVFRYVDAYKSIDIESVPANDADESAPSEAIITP--EPEPVHVPEVIPPTQTV 177
G++VLND+ RY +++ E+ ND +P E ++T + P HV + T +V
Sbjct: 123 GGYFVLNDMLRYDSLQETLLTET---ND----SPQERLLTEINDSLPNHVDD---NTHSV 172
Query: 178 IPTAQTVIPPTQTVIADTETIISKEVSLPLEN---GKLSVTENVIPVNHVKESSHHVKEP 234
T++ AD E ++ V+ +EN S ENV+ + S +
Sbjct: 173 TFTSEPETSGNVNETADLELPSAENVNDNVENLPANDSSPEENVLVEACTEVVSSCAENI 232
Query: 235 EQPTSIEKVASNTQEDTPKKSFASIVNALKDNSAPFHLRASPAKPAVHPPR------VHS 288
++TQ+D K+S+AS+V K+ + + KP P +
Sbjct: 233 PAAAPAPAPRASTQKDVTKQSYASVVKVTKEGTPTPPVAKPKPKPKPKPTAKVTDNVEKA 292
Query: 289 VPAPEAPTPNMDI--PLEKNNENAGRAHAIFVANLPMSATVEQLDRAFKKFGPIKRDGIQ 346
V +P PT D P +K N + ++++V +LP T + ++ F+KFG I+ GIQ
Sbjct: 293 VSSPVKPTNAADTTSPNDK-NVLVEQGYSVYVKHLPYECTTKDVEEKFRKFGAIRPGGIQ 351
Query: 347 VRSNK--GSCFGFVEFESAASMQSALEVWSISINS 379
VR + G CFGFVEFES SM +A+E +SI S
Sbjct: 352 VRHRQPDGFCFGFVEFESRQSMLAAIEASPVSIGS 386
>UniRef100_Q9LDI9 Putative RNA-binding protein; 63745-61607 [Arabidopsis thaliana]
Length = 427
Score = 195 bits (495), Expect = 2e-48
Identities = 132/375 (35%), Positives = 203/375 (53%), Gaps = 54/375 (14%)
Query: 10 PTPQMVGNAFVEQYYSILHRDPDQVHRFYHDSSVMSRPEEDGTMTTVTTTAEIDKKIQSL 69
P+ Q + FV QYY +L + P + R Y D+SV+SRP+ GTM + T+ I+K I S
Sbjct: 8 PSAQDIAAEFVRQYYHVLGQLPHEARRLYVDASVVSRPDVTGTMMSFTSVEAINKHILSC 67
Query: 70 EYTSFRVEVLSADAQPSYNNGVMVVVTGCLTGTDNIKRKFAQSFFLAPQDKGFYVLNDVF 129
++ + + EVLS D+Q S +G+ ++V G +TG DN +RKF+Q F+LA Q+ VLND+
Sbjct: 68 DFENTKFEVLSVDSQNSLEDGIFIMVIGFMTGKDNQRRKFSQMFYLARQNT-LVVLNDML 126
Query: 130 RYVDAYKSIDIESVPANDADESAPSEAIITPEPEPVHVPEVIPPTQTVIPTAQTVIPPTQ 189
RYV D ++S+ +E P V E++ P + +T + Q
Sbjct: 127 RYV--------------DQEDSSTTETPCEP------VTEIVRPADGLKKAEKTEL--KQ 164
Query: 190 TVIADTETIISKEV---SLPLENGKLSVTENVIPVNHVKESSHHVKEPEQPTSIEKVASN 246
+A E ++ V + PL+NGK+ +E + + V EP+ +
Sbjct: 165 KNVASVEKSVNAAVEKNAAPLDNGKMKQSEKAV-------ITQKVTEPD---------AA 208
Query: 247 TQEDTPKKSFASIVNALKDNSAPFHLRASPAKPAVHPPRVHSVP----APEAPTPNMDIP 302
Q D K+SFA IV ++ N+APF ++ SP + V P+ P AP+ P +
Sbjct: 209 PQPDGAKRSFADIVGSMAKNAAPFQVK-SPVQAPVQKPKYVGQPRAAAAPQKPA-YVSKS 266
Query: 303 LEKNNENAGR--AHAIFVANLPMSATVEQLDRAFKKFGPIKRDGIQVRSNKGS----CFG 356
++KN++ +IFVANLP++A QL FK FGPIK +GIQVRS++G+ CFG
Sbjct: 267 IKKNDQKVIEVPGTSIFVANLPLNAMPPQLFELFKDFGPIKENGIQVRSSRGNANPVCFG 326
Query: 357 FVEFESAASMQSALE 371
F+ FE+ AS+QS L+
Sbjct: 327 FISFETVASVQSVLQ 341
>UniRef100_Q9FJK5 RNA-binding protein-like [Arabidopsis thaliana]
Length = 461
Score = 157 bits (398), Expect = 4e-37
Identities = 124/404 (30%), Positives = 191/404 (46%), Gaps = 67/404 (16%)
Query: 12 PQMVGNAFVEQYYSILHRDPDQVHRFYHDSSVMSRPEEDGTMTTVTTTAEIDKKIQSLEY 71
P VG+AFV QYY I P+ + RFY + S + R +DG M +T I ++++ L Y
Sbjct: 13 PLTVGSAFVNQYYYIFCNMPEHLPRFYQEISRVGRVGQDGVMRDFSTFQGISEELKRLTY 72
Query: 72 TSFR-VEVLSADAQPSYNNGVMVVVTGCLTGTDNIKRKFAQSFFLAPQDKGFYVLNDVFR 130
E+ S D Q S+N G ++ VTG T + +RKF Q+FFLAPQ+KGF+VLND+ R
Sbjct: 73 GDCNSAEITSYDTQESHNGGFLLFVTGYFTLNERSRRKFTQTFFLAPQEKGFFVLNDILR 132
Query: 131 YVDAYKSIDIESVPANDADESAPSEAIITPEPEPVHVPEVIPPTQTVIPTAQTVIPPTQT 190
+V+ +DA ++ PE V I T I A T + ++
Sbjct: 133 FVN------------DDAKDN-------VPETIDGEVVSGINSTTPTIINAPTGMKGSEQ 173
Query: 191 VIADTETIISKEVSLPLENGKLSVTENVIPVNHVKESSHHVKEPEQPTSIEKVASNTQE- 249
+ + KEVS PL+N + +NV+ V E ++ V E + ++VA ++Q+
Sbjct: 174 AACVSVNPVCKEVSKPLDNE--NAKDNVL----VPEIANEVARTE--ITCKEVADDSQKN 225
Query: 250 --------DTPKKSFASIVNALKDN-SAPFHLRASPAKPAVHPPRVHSVPAP-------- 292
D PKKS+AS++ KD P SP K + + H P+
Sbjct: 226 YDPDDGLADAPKKSYASVLKVTKDKFGVPAVSLPSPKK--IPKDQEHQAPSDPSTGQILK 283
Query: 293 --------------EAPTPNMDIPLEKNNEN---AGRAHAIFVANLPMSATVEQLDRAFK 335
E+ T + + +N N +I+V +LP +A ++ L+ FK
Sbjct: 284 DQGQQASSDPSQVIESDTVSESVDASENGHNQEAVAEGTSIYVRHLPFNANIDMLEAEFK 343
Query: 336 KFGPIKRDGIQVRSNK--GSCFGFVEFESAASMQSALEVWSISI 377
+FG I GIQV + + G +GFVEFE A + A+E + I
Sbjct: 344 QFGAITNGGIQVINQRGLGYPYGFVEFEEADAAHRAIEASPVKI 387
>UniRef100_Q84JH2 Putative NTF2-containing RNA-binding protein [Arabidopsis thaliana]
Length = 458
Score = 156 bits (395), Expect = 8e-37
Identities = 121/404 (29%), Positives = 190/404 (46%), Gaps = 70/404 (17%)
Query: 12 PQMVGNAFVEQYYSILHRDPDQVHRFYHDSSVMSRPEEDGTMTTVTTTAEIDKKIQSLEY 71
P VG+AFV QYY I P+ + RFY + S + R +DG M +T I ++++ L Y
Sbjct: 13 PLTVGSAFVNQYYYIFCNMPEHLPRFYQEISRVGRVGQDGVMRDFSTFQGISEELKRLTY 72
Query: 72 TSFR-VEVLSADAQPSYNNGVMVVVTGCLTGTDNIKRKFAQSFFLAPQDKGFYVLNDVFR 130
E+ S D Q S+N G ++ VTG T + +RKF Q+FFLAPQ+KGF+VLND+ R
Sbjct: 73 GDCNSAEITSYDTQESHNGGFLLFVTGYFTLNERSRRKFTQTFFLAPQEKGFFVLNDILR 132
Query: 131 YVDAYKSIDIESVPANDADESAPSEAIITPEPEPVHVPEVIPPTQTVIPTAQTVIPPTQT 190
+V+ +DA ++ P EV+ + PT + ++
Sbjct: 133 FVN------------DDAKDNVPETI----------DGEVVSGINSTTPTIINGMKGSEQ 170
Query: 191 VIADTETIISKEVSLPLENGKLSVTENVIPVNHVKESSHHVKEPEQPTSIEKVASNTQE- 249
+ + KEVS PL+N + +NV+ V E ++ V E + ++VA ++Q+
Sbjct: 171 AACVSVNPVCKEVSKPLDNE--NAKDNVL----VPEIANEVARTE--ITCKEVADDSQKN 222
Query: 250 --------DTPKKSFASIVNALKDN-SAPFHLRASPAKPAVHPPRVHSVPAP-------- 292
D PKKS+AS++ KD P SP K + + H P+
Sbjct: 223 YDPDDGLADAPKKSYASVLKVTKDKFGVPAVSLPSPKK--IPKDQEHQAPSDPSTGQILK 280
Query: 293 --------------EAPTPNMDIPLEKNNEN---AGRAHAIFVANLPMSATVEQLDRAFK 335
E+ T + + +N N +I+V +LP +A ++ L+ FK
Sbjct: 281 DQGQQASSDPSQVIESDTVSESVDASENGHNQEAVAEGTSIYVRHLPFNANIDMLEAEFK 340
Query: 336 KFGPIKRDGIQVRSNK--GSCFGFVEFESAASMQSALEVWSISI 377
+FG I GIQV + + G +GFVEFE A + A+E + I
Sbjct: 341 QFGAITNGGIQVINQRGLGYPYGFVEFEEADAAHRAIEASPVKI 384
>UniRef100_Q6ZG86 Putative Ras-GTPase activating protein SH3 domain-binding protein 2
[Oryza sativa]
Length = 511
Score = 155 bits (393), Expect = 1e-36
Identities = 117/383 (30%), Positives = 191/383 (49%), Gaps = 38/383 (9%)
Query: 15 VGNAFVEQYYSILHRDPDQVHRFYHDSSVMSRPEE-DGTMTTVTTTAEIDKKIQSLEYTS 73
VG F+ YY++L + PD VH+FY+D+S M R ++ GT TT +T +I I SL +T
Sbjct: 12 VGTYFLRNYYNLLQQSPDVVHQFYNDASTMVRVDDLAGTNTTASTMMDIHSLIMSLNFT- 70
Query: 74 FRVEVLSADAQPSYNNGVMVVVTGCL-TGTDNIKRKFAQSFFLAPQDKGFYVLNDVFRYV 132
++E+ +A+ S+ +GV+V+V+G + T + +RKF Q FFLAPQ+KG++VLND F +V
Sbjct: 71 -QIEIKTANFLNSWGDGVLVMVSGLVQTKEYSHQRKFIQMFFLAPQEKGYFVLNDYFHFV 129
Query: 133 DAYKSIDIESVPANDADESAPSEAIITPEPEPVHVPE---VIPPTQTVIPTAQ--TVIPP 187
D + + ++ + + S +++ EPE +H E +P T + T P
Sbjct: 130 DEEQVQPAPVIAQDNFETNMASNSVV--EPEYIHEEENQSAVPITSEESDAVENYTYSEP 187
Query: 188 TQTVIADTET----IISKEVSLPLENGKLSVTENVIPVNHVKESSHHVKEPEQPTSIEKV 243
Q V++ ++ + +E NG E PV HV+EP
Sbjct: 188 PQQVVSQSDNWGDEPLPEEPISSFTNGMAMAPEE--PVQSPPVPPPHVEEP--------- 236
Query: 244 ASNTQEDTPKKSFASIVNALKDNSAPFHLRASPAKP----AVHPPRVHSVPAPEAPT--P 297
+ KK++ASI+ K + +P +P + HSV T P
Sbjct: 237 ----VGEPVKKTYASILRTAKAPLVFPVAQPAPTRPHQATETNQAAQHSVMTSSVATEKP 292
Query: 298 NMDIPLEKNNENAGRAHAIFVANLPMSATVEQLDRAFKKFGPIKRDGIQVRSNK--GSCF 355
D+ E ++ + +++V N+P S + L+ FKKFG + DG+ +RS K G +
Sbjct: 293 KTDVYGEFAVQDDEESKSVYVGNVPSSVSEADLENEFKKFGRLIPDGVAIRSRKETGGYY 352
Query: 356 GFVEFESAASMQSALEVWSISIN 378
FVEFE + + +AL+ I IN
Sbjct: 353 AFVEFEELSGVHNALKASPIEIN 375
>UniRef100_Q7ZXC7 G3bp-prov protein [Xenopus laevis]
Length = 470
Score = 124 bits (312), Expect = 4e-27
Identities = 113/406 (27%), Positives = 172/406 (41%), Gaps = 52/406 (12%)
Query: 7 VQTPTPQMVGNAFVEQYYSILHRDPDQVHRFYHDSSVMSRPEEDGT---MTTVTTTAEID 63
++ P+P +VG FV QYY++L++ PD +HRFY SS D V +I
Sbjct: 3 MEKPSPLLVGREFVRQYYTLLNQAPDFLHRFYGKSSSYVHGGLDSNGKPADAVYGQTDIH 62
Query: 64 KKIQSLEYTSFRVEVLSADAQPSYNNGVMVVVTGCLTGTDNIKRKFAQSFFLAPQD---K 120
KK+ SL + R ++ DA + N+GV+V V G L+ R+F Q+F LAP+
Sbjct: 63 KKVMSLNFKDCRTKIRHVDAHATLNDGVVVQVMGELSNNRQPMRRFMQTFLLAPEGSVAN 122
Query: 121 GFYVLNDVFRYVDAYKSIDIESVPANDADESAPSEAIITPEPEPVHVPEVIPPTQTVIPT 180
FYV ND+FRY D D ++ P ++DE A PE V E T P
Sbjct: 123 KFYVHNDIFRYQDEVFG-DSDTEPPEESDEEAEEPEERQQSPELVATDEA---TYYEQPG 178
Query: 181 AQTVIPPTQTVIADTETIISKEVSLP---LENGKLSVTENVIPVNHVKESSHHVKEPEQP 237
+ + P + I + E E+ +E +E P +E+ + K P P
Sbjct: 179 SNDLDEPLEDQIVEAEPEQEPELEQEPEMVEEKCEPTSEEPAPEEEEEETVNIEKSPSPP 238
Query: 238 TSIEKVASNTQEDTPKKSFASIVN-------ALKDNSAPFHL----------------RA 274
+ A QED S+AS+ + A+ + P H+ +
Sbjct: 239 PA--DPAPAVQEDPRTFSWASVTSKNLPPSGAVPVSGIPPHVVKVQTSQPRPDTKSEAQT 296
Query: 275 SPAKPAVHPPRVH----SVPAPEAPTPNMDIPLEKNNENAGR---AHAIFVANLPMSATV 327
+ +P RV +VP P P P + E R +H +FV NLP
Sbjct: 297 TTQRPPQRDQRVREQRTTVPPPRGPRPLREGEPESETRRINRYPDSHQLFVGNLPHDVDK 356
Query: 328 EQLDRAFKKFGPIKRDGIQVRSNKGS---CFGFVEFESAASMQSAL 370
+L F+ +G + +++R N G FGFV F+ A +Q L
Sbjct: 357 TELKEFFQTYGNV----VELRINSGGKLPNFGFVVFDDAEPVQKIL 398
>UniRef100_Q7ZVG3 Ras-GTPase-activating protein SH3-domain-binding protein
[Brachydanio rerio]
Length = 477
Score = 120 bits (302), Expect = 5e-26
Identities = 115/412 (27%), Positives = 180/412 (42%), Gaps = 64/412 (15%)
Query: 7 VQTPTPQMVGNAFVEQYYSILHRDPDQVHRFY--HDSSVMSRPEEDGTMT-TVTTTAEID 63
++ P+ Q+VG FV QYY++L++ PD +HRFY + S V + +G V +EI
Sbjct: 3 MEKPSAQLVGREFVRQYYTLLNQAPDYLHRFYGKNSSYVHGGLDNNGKPAEAVYGQSEIH 62
Query: 64 KKIQSLEYTSFRVEVLSADAQPSYNNGVMVVVTGCLTGTDNIKRKFAQSFFLAPQD---K 120
KK+ +L + ++ DA + N GV+V V G L+ RKF Q+F LAP+
Sbjct: 63 KKVMALSFRDCHTKIRHVDAHATLNEGVVVQVLGGLSNNMQPMRKFMQTFVLAPEGTVAN 122
Query: 121 GFYVLNDVFRYVDAYKSIDIESVPANDADESAPSEAIITPEPEPVHVPEVIPPTQT---- 176
FYV ND+FRY D D +S P +++E E E VH PEV+ +
Sbjct: 123 KFYVHNDIFRYQDEVFG-DSDSEPPEESEED-------VEELERVHSPEVVQEESSGYYE 174
Query: 177 VIPTAQTVIPPTQTVIADTETIISKEVSLPLENGKLSVTENVI--PVNHVKESSHHVKEP 234
P + + P + V E EV + E + + I P HV+E + + P
Sbjct: 175 QTPCVEPEV-PQEEVSVTPEPQPEPEVEVEPEPAAVELKAEPISQPEVHVEEKTQ--RSP 231
Query: 235 EQPTSIEKVASNTQEDTPKKSFASIVN-------ALKDNSAPFH-LRASPAKPAV----- 281
PT + A ED S+AS+ + + P H +R A+P V
Sbjct: 232 PSPTPAD-TAPTMPEDNRPSSWASVTSKNLPPGGVVPATGVPPHVVRVPSAQPRVEVKTE 290
Query: 282 ------------HPPRVHSVPAPEAPTPNMDIPLE-KNNENAGR-------AHAIFVANL 321
P P+P TP + E ++ E+ R +H +FV N+
Sbjct: 291 TQTTAQRPQRDQRPRDQRPGPSPAHRTPRPGVVREGESGESEVRRTVRYPDSHQLFVGNV 350
Query: 322 PMSATVEQLDRAFKKFGPIKRDGIQVRSNKGS---CFGFVEFESAASMQSAL 370
P +L F+++G + +++R N G FGFV F+ + +Q L
Sbjct: 351 PHDVDKNELKEFFEQYGTV----LELRINSGGKLPNFGFVVFDDSEPVQKIL 398
>UniRef100_Q6P124 Ras-GTPase-activating protein SH3-domain-binding protein
[Brachydanio rerio]
Length = 477
Score = 120 bits (301), Expect = 7e-26
Identities = 115/412 (27%), Positives = 180/412 (42%), Gaps = 64/412 (15%)
Query: 7 VQTPTPQMVGNAFVEQYYSILHRDPDQVHRFY--HDSSVMSRPEEDG-TMTTVTTTAEID 63
++ P+ Q+VG FV QYY++L++ PD +HRFY + S V + +G V +EI
Sbjct: 3 MEKPSAQLVGREFVRQYYTLLNQAPDYLHRFYGKNSSYVHGGLDNNGKPAEAVYGQSEIH 62
Query: 64 KKIQSLEYTSFRVEVLSADAQPSYNNGVMVVVTGCLTGTDNIKRKFAQSFFLAPQD---K 120
KK+ +L + ++ DA + N GV+V V G L+ RKF Q+F LAP+
Sbjct: 63 KKVMALSFRDCHTKIRHVDAHATLNEGVVVQVLGELSNNMQPMRKFMQTFVLAPEGTVAN 122
Query: 121 GFYVLNDVFRYVDAYKSIDIESVPANDADESAPSEAIITPEPEPVHVPEVIPPTQT---- 176
FYV ND+FRY D D +S P +++E E E VH PEV+ +
Sbjct: 123 KFYVHNDIFRYQDEVFG-DSDSEPPEESEED-------VEELERVHSPEVVQEESSGYYE 174
Query: 177 VIPTAQTVIPPTQTVIADTETIISKEVSLPLENGKLSVTENVI--PVNHVKESSHHVKEP 234
P + + P + V E EV + E + + I P HV+E + + P
Sbjct: 175 QTPCVEPEV-PQEEVSVTPEPQPEPEVEVEPEPAAVELKAEPISQPEVHVEEKTQ--RSP 231
Query: 235 EQPTSIEKVASNTQEDTPKKSFASIVN-------ALKDNSAPFH-LRASPAKPAV----- 281
PT + A ED S+AS+ + + P H +R A+P V
Sbjct: 232 PSPTPAD-TAPTMPEDNRPSSWASVTSKNLPPGGVVPATGVPPHVVRVPSAQPRVEVKTE 290
Query: 282 ------------HPPRVHSVPAPEAPTPNMDIPLE-KNNENAGR-------AHAIFVANL 321
P P+P TP + E ++ E+ R +H +FV N+
Sbjct: 291 TQTTAQRPQRDQRPRDQRPGPSPAHRTPRPGVVREGESGESEVRRTVRYPDSHQLFVGNV 350
Query: 322 PMSATVEQLDRAFKKFGPIKRDGIQVRSNKGS---CFGFVEFESAASMQSAL 370
P +L F+++G + +++R N G FGFV F+ + +Q L
Sbjct: 351 PHDVDKNELKEFFEQYGTV----LELRINSGGKLPNFGFVVFDDSEPVQKIL 398
>UniRef100_Q6DFK7 MGC81268 protein [Xenopus laevis]
Length = 483
Score = 119 bits (299), Expect = 1e-25
Identities = 112/397 (28%), Positives = 164/397 (41%), Gaps = 42/397 (10%)
Query: 7 VQTPTPQMVGNAFVEQYYSILHRDPDQVHRFY--HDSSVMSRPEEDGT-MTTVTTTAEID 63
++ P+P +VG FV QYY++L++ PD +HRFY + S V + +G V AEI
Sbjct: 3 MEKPSPLLVGREFVRQYYTLLNKAPDFLHRFYGRNSSYVHGGLDANGKPQEAVYGQAEIH 62
Query: 64 KKIQSLEYTSFRVEVLSADAQPSYNNGVMVVVTGCLTGTDNIKRKFAQSFFLAPQ---DK 120
KK+ SL+++ R ++ DA + ++GV+V V G L+ RKF Q+F LAP+
Sbjct: 63 KKVMSLQFSECRTKIRHVDAHATLSDGVVVQVMGELSNNGQPMRKFMQTFVLAPEGSVPN 122
Query: 121 GFYVLNDVFRYVDAYKSIDIESVPANDADESAPSEAIITPEPEPVHVPEVIPPTQTVIPT 180
FYV ND+FRY D + +E + P PEPV P
Sbjct: 123 KFYVHNDIFRYEDEVFGDSEAEMDEESEEEVEEEQEERQPSPEPVQ-ENASNTYYEPHPV 181
Query: 181 AQTVIPPTQTVIADTETIISKEVSLPLENGKLSVTENVIPVNHVKESSHHVKE----PEQ 236
+ V P + I + E E E+ K E V K + E P++
Sbjct: 182 SNGVEEPLEEPITEPEP--EPEPETKNEDIKPETEEKVQEELEEKSPTPPPAEPTVLPQE 239
Query: 237 PTSIEKVASNTQEDTPKKSFASIVNALKDNSAPFHLRASPAKPA---------VHPPRVH 287
P AS T ++ P + + P H+ +PA A VH PRV
Sbjct: 240 PPKAFSWASVTSKNLPPS------GTVPASGIPPHVIKAPASQARVDVKPEAPVHSPRVR 293
Query: 288 SVPAPEAPT--PNMDIPLEKNNENAGR----------AHAIFVANLPMSATVEQLDRAFK 335
E PT P +NE A +H +FV NLP +L F
Sbjct: 294 EQRPRERPTFQPRGPRAGRSDNEQAESDNRRMYRYPDSHQLFVGNLPHDIDESELKEFFM 353
Query: 336 KFGPIKRDGIQVRSNKGSC--FGFVEFESAASMQSAL 370
+G + I + G FGFV F+ + +Q L
Sbjct: 354 SYGNVMELRINTKGVGGKLPNFGFVVFDDSEPVQRIL 390
>UniRef100_Q5RAH4 Hypothetical protein DKFZp459E0328 [Pongo pygmaeus]
Length = 482
Score = 119 bits (297), Expect = 2e-25
Identities = 108/394 (27%), Positives = 167/394 (41%), Gaps = 37/394 (9%)
Query: 7 VQTPTPQMVGNAFVEQYYSILHRDPDQVHRFYHDSSVMSRPEEDGT---MTTVTTTAEID 63
++ P+P +VG FV QYY++L++ P+ +HRFY +S D + V +I
Sbjct: 3 MEKPSPLLVGREFVRQYYTLLNKAPEYLHRFYGRNSSYVHGGVDASGKPQEAVYGQNDIH 62
Query: 64 KKIQSLEYTSFRVEVLSADAQPSYNNGVMVVVTGCLTGTDNIKRKFAQSFFLAPQ---DK 120
K+ SL ++ ++ DA + ++GV+V V G L+ + +RKF Q+F LAP+
Sbjct: 63 HKVLSLNFSECHTKIRHVDAHATLSDGVVVQVMGLLSNSGQPERKFMQTFVLAPEGSVPN 122
Query: 121 GFYVLNDVFRYVDAYKSIDIESVPANDADESAPSEAIITPEPEPVH------VPEVIPPT 174
FYV ND+FRY D + DE + P PEPV E P T
Sbjct: 123 KFYVHNDMFRYEDEVFGDSEPELDEESEDEVEEEQEERQPSPEPVQENANSGYYEAHPVT 182
Query: 175 QTV-IPTAQTVIPPTQTVIADTETIISKEVSLPLENGKLSVTENVIPVNHVKESSHHVKE 233
+ P ++ P ++T+T +E+ +E L E + + + V
Sbjct: 183 NGIEEPLEESSHEPEPEPESETKT---EELKPQVEEKNL---EELEEKSTTPPPAEPVSL 236
Query: 234 PEQPTSIEKVASNTQEDTPKKSFASIVNALKDNSAPFHLRASPAKPAV--HPPRVHSVP- 290
P++P AS T ++ P S AP AKP V PPRV
Sbjct: 237 PQEPPKAFSWASVTSKNLPPSGTVSSSGIPPHVKAPVSQPRVEAKPEVQSQPPRVREQRP 296
Query: 291 ------APEAPTPNMDIPLEKNNENAGR------AHAIFVANLPMSATVEQLDRAFKKFG 338
P P P+ +E+N+ + R +H +FV NLP +L F FG
Sbjct: 297 RERPGFPPRGPRPDRG-DMEQNDSDNRRIIRYPDSHQLFVGNLPHDIDENELKEFFMSFG 355
Query: 339 PIKRDGIQVRSNKGSC--FGFVEFESAASMQSAL 370
+ I + G FGFV F+ + +Q L
Sbjct: 356 NVVELRINTKGVGGKLPNFGFVVFDDSEPVQRIL 389
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.313 0.129 0.371
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 683,834,998
Number of Sequences: 2790947
Number of extensions: 30076549
Number of successful extensions: 102008
Number of sequences better than 10.0: 1442
Number of HSP's better than 10.0 without gapping: 208
Number of HSP's successfully gapped in prelim test: 1264
Number of HSP's that attempted gapping in prelim test: 98045
Number of HSP's gapped (non-prelim): 3471
length of query: 393
length of database: 848,049,833
effective HSP length: 129
effective length of query: 264
effective length of database: 488,017,670
effective search space: 128836664880
effective search space used: 128836664880
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 76 (33.9 bits)
Medicago: description of AC146567.10


