

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146566.2 + phase: 2 /partial
(358 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
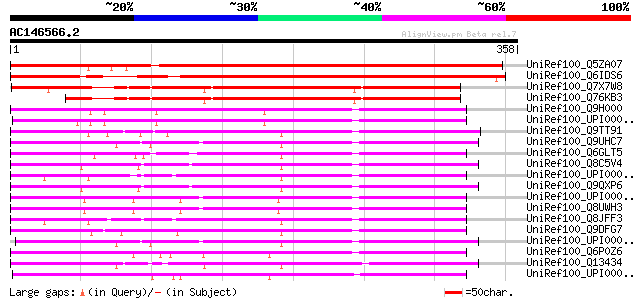
Sequences producing significant alignments: (bits) Value
UniRef100_Q5ZA07 Putative makorin RING finger protein [Oryza sat... 473 e-132
UniRef100_Q6IDS6 Hypothetical protein At3g08505 [Arabidopsis tha... 459 e-128
UniRef100_Q7X7W8 Makorin ring-zinc-finger protein [Pisum sativum] 313 4e-84
UniRef100_Q76KB3 Makorin ring-zinc-finger protein [Pisum sativum] 272 9e-72
UniRef100_Q9H000 Makorin 2 [Homo sapiens] 249 6e-65
UniRef100_UPI000036B3BD UPI000036B3BD UniRef100 entry 241 2e-62
UniRef100_Q9TT91 Makorin 1 [Macropus eugenii] 241 2e-62
UniRef100_Q9UHC7 Makorin 1 [Homo sapiens] 239 1e-61
UniRef100_Q6GLT5 MGC84269 protein [Xenopus laevis] 238 2e-61
UniRef100_Q8C5V4 Mus musculus adult male testis cDNA, RIKEN full... 238 2e-61
UniRef100_UPI000027CA99 UPI000027CA99 UniRef100 entry 238 2e-61
UniRef100_Q9QXP6 Makorin 1 [Mus musculus] 238 2e-61
UniRef100_UPI00003AA829 UPI00003AA829 UniRef100 entry 237 4e-61
UniRef100_Q8UWH3 Makorin-2 [Gallus gallus] 237 4e-61
UniRef100_Q8JFF3 Gene encoding protein featuring ring-finger [Se... 236 6e-61
UniRef100_Q9DFG7 Makorin RING zinc-finger protein 1 [Brachydanio... 235 2e-60
UniRef100_UPI000036DFF0 UPI000036DFF0 UniRef100 entry 234 3e-60
UniRef100_Q6P0Z6 Mkrn2 protein [Brachydanio rerio] 231 3e-59
UniRef100_Q13434 Makorin 4 [Homo sapiens] 228 3e-58
UniRef100_UPI000001360B UPI000001360B UniRef100 entry 226 1e-57
>UniRef100_Q5ZA07 Putative makorin RING finger protein [Oryza sativa]
Length = 368
Score = 473 bits (1216), Expect = e-132
Identities = 228/364 (62%), Positives = 267/364 (72%), Gaps = 21/364 (5%)
Query: 1 VLCKFFAHGACLKGEHCEFSHDWKAPPNNICTFYQKGVCAYGSRCRYDHVKASR------ 54
VLCKFF HGACLKGE+CEFSHDW PNN+CTFYQKG C+YGSRCRYDHVK SR
Sbjct: 6 VLCKFFMHGACLKGEYCEFSHDWNDQPNNVCTFYQKGSCSYGSRCRYDHVKVSRNPTVAP 65
Query: 55 -AQSSTPSSSITEHQPL-------VSESAVLGNTR--VTSNGVATAAEFSLFSTPFVLPS 104
SST + + + QPL V A N R ++ + +A + ++ F
Sbjct: 66 PPSSSTTTRASSSLQPLSFGRPHHVGYQADSSNPRQQISMDVLAHSGSKPVWRNDF---- 121
Query: 105 EQAWNQESAQLDFLREDDVVQSVITSPSELPICSFAAAGNCPRGEQCPHVHGDLCPSCGR 164
Q + +D+ V SP++LPICSFAA GNCP GE+CP +HGDLC +CG+
Sbjct: 122 -QHESVLEDGIDWSISPTVQNQTTLSPADLPICSFAAGGNCPYGEECPQMHGDLCTTCGK 180
Query: 165 QCLHPFRPEEREEHMMSCRNKQKHLEALKRSQEIECSVCLERVLSKPTAAERKFGLLSEC 224
CLHP+RP+EREEH C K LE+LKRSQEIECSVCL+RVLSKPTAAERKFGLLSEC
Sbjct: 181 MCLHPYRPDEREEHTKLCEKNHKRLESLKRSQEIECSVCLDRVLSKPTAAERKFGLLSEC 240
Query: 225 DHPFCVSCIRNWRSSNPTLGMDVNSTLRACPICRKLSYFVVPSVIWYATSEEKMEIIDTY 284
DHPFC+SCIRNWR+++PT GMDVNS LRACPICRKLSY+V+PSV+WY + EEK+EIID Y
Sbjct: 241 DHPFCISCIRNWRNNSPTSGMDVNSALRACPICRKLSYYVIPSVLWYFSKEEKLEIIDNY 300
Query: 285 KAKLKSIDCKHFDFGEGNCPFGTSCFYKHAYRDGRLEEVALRHLGAADGDTIIAKDIRFL 344
KAKLKSIDCK+FDFG G CPFG+SCFYKHAYRDGRLEEV LRHL A DG T+IAK+IR
Sbjct: 301 KAKLKSIDCKYFDFGTGTCPFGSSCFYKHAYRDGRLEEVILRHLDADDGSTVIAKNIRLS 360
Query: 345 STLS 348
LS
Sbjct: 361 DFLS 364
>UniRef100_Q6IDS6 Hypothetical protein At3g08505 [Arabidopsis thaliana]
Length = 323
Score = 459 bits (1181), Expect = e-128
Identities = 220/353 (62%), Positives = 254/353 (71%), Gaps = 37/353 (10%)
Query: 1 VLCKFFAHGACLKGEHCEFSHDWKAPPNNICTFYQKGVCAYGSRCRYDHVKASRAQSSTP 60
+LCKFF HG+CLKGE+CEFSHD K PPNN+CTFYQK +C YGSRCRYDHV RA S+ P
Sbjct: 5 ILCKFFVHGSCLKGENCEFSHDSKDPPNNVCTFYQKRICLYGSRCRYDHV---RAASNLP 61
Query: 61 SSSITEHQPLVSESAVLGNTRVTSNGVATAAEFSLFSTPFVLPSEQAWNQESAQLDFLRE 120
SS +E + + S+ +TP +Q N +
Sbjct: 62 LSSDSE-----------------------SLDRSISTTPSRHLQQQGDNNDG-------- 90
Query: 121 DDVVQSVITSPSELPICSFAAAGNCPRGEQCPHVHGDLCPSCGRQCLHPFRPEEREEHMM 180
D P E PICSFAAAG+CPRG QCPH+HGDLC +CG++CLHPFRPEEREEH
Sbjct: 91 DKSSNVYCIHPREYPICSFAAAGDCPRGNQCPHMHGDLCNTCGKKCLHPFRPEEREEHTK 150
Query: 181 SCRNKQKHLEALKRSQEIECSVCLERVLSKPTAAERKFGLLSECDHPFCVSCIRNWRSSN 240
C KQKH+EALK+SQ+IECSVCL+R+LSK T ERKFGLL+ECDHPFC+ CIRNWRSS
Sbjct: 151 ECEKKQKHIEALKQSQDIECSVCLDRILSKATPGERKFGLLTECDHPFCIQCIRNWRSSA 210
Query: 241 PTLGMDVNSTLRACPICRKLSYFVVPSVIWYATSEEKMEIIDTYKAKLKSIDCKHFDFGE 300
P GMDVNSTLRACPICRKLSYFVVPSV+WY++ EEK EIID YKAKL+SIDCKHF+FG
Sbjct: 211 PVSGMDVNSTLRACPICRKLSYFVVPSVVWYSSPEEKKEIIDIYKAKLRSIDCKHFNFGN 270
Query: 301 GNCPFGTSCFYKHAYRDGRLEEVALRHLGAADGDTIIAKDIR---FLSTLSWF 350
GNCPFG SCFYKHAY DG LEEV LRHLG+ +G+T+I IR FL L F
Sbjct: 271 GNCPFGASCFYKHAYSDGHLEEVVLRHLGSQEGETVITDSIRLSEFLGGLQIF 323
>UniRef100_Q7X7W8 Makorin ring-zinc-finger protein [Pisum sativum]
Length = 461
Score = 313 bits (803), Expect = 4e-84
Identities = 165/334 (49%), Positives = 207/334 (61%), Gaps = 36/334 (10%)
Query: 2 LCKFFAHGACLKGEHCEFSHDWKAP--------PNNICTFYQKGVCAYGSRCRYDHVKAS 53
+CKF+A G CLKG+ C+FSH K IC++YQKG CAY SRCRY HVKAS
Sbjct: 5 VCKFYARGICLKGDQCDFSHQRKDTHQRKDNPVDKQICSYYQKGSCAYDSRCRYKHVKAS 64
Query: 54 RAQSSTPSSSITEHQPLVSESAVLGNTRVTSNGVATAAEFSLFSTPFVLPSEQAWNQESA 113
+A SS ++++ N VT +GV A ++ T V P S
Sbjct: 65 QASSS---------------ASLIPNCAVT-HGVKLAPSWAPKVTR-VPPPPCKHGVRSF 107
Query: 114 QLDFLREDDVVQSVITSPSELPI-----CSFAAAGNCPRGEQCPHVHGDLCPSCGRQCLH 168
Q ++ DV +S T + + C FAAA NCP G C HVHG+ C C + CLH
Sbjct: 108 QHNYQDSSDVGESSSTGSTSARLHGHLFCKFAAA-NCPFGHGCSHVHGNQCLYCRKYCLH 166
Query: 169 PFRPEEREEHMMSCRNKQKHLEALKRSQEIECSVCLERVLSKPTAAERKFGLLSECDHPF 228
P E+E H+ +C K+K++ ALK S+EIEC+VCLE VLSKP +ERKFGLL ECDH F
Sbjct: 167 PSDRREKENHLKTCDKKEKYVLALKNSEEIECNVCLEHVLSKPKPSERKFGLLPECDHAF 226
Query: 229 CVSCIRNWRSSNPT----LGMDVNSTLRACPICRKLSYFVVPSVIWYATSEEKMEIIDTY 284
C+SCIRNWR+S P+ G ++N T+R CP+CR+LSYFV+PS IWY T EEK EIID Y
Sbjct: 227 CLSCIRNWRNSAPSSEFETGNNIN-TVRTCPVCRQLSYFVIPSGIWYTTKEEKQEIIDNY 285
Query: 285 KAKLKSIDCKHFDFGEGNCPFGTSCFYKHAYRDG 318
KA + IDCKHF+ G GNCPFG SCFYKH + G
Sbjct: 286 KANCRLIDCKHFESGNGNCPFGASCFYKHTVKPG 319
>UniRef100_Q76KB3 Makorin ring-zinc-finger protein [Pisum sativum]
Length = 411
Score = 272 bits (696), Expect = 9e-72
Identities = 145/288 (50%), Positives = 181/288 (62%), Gaps = 28/288 (9%)
Query: 40 AYGSRCRYDHVKASRAQSSTPSSSITEHQPLVSESAVLGNTRVTSNGVATAAEFSLFSTP 99
AYGSRCRY HVKAS+A SS ++++ N VT +GV A ++ T
Sbjct: 1 AYGSRCRYKHVKASQASSS---------------ASLIPNCAVT-HGVKLAPSWAPKVTR 44
Query: 100 FVLPSEQAWNQESAQLDFLREDDVVQSVITSPSELPI-----CSFAAAGNCPRGEQCPHV 154
V P S Q ++ DV +S T + + C FAAA NCP G C HV
Sbjct: 45 -VPPPPCKHGVRSFQHNYQDSSDVGESSSTGSTSARLHGHLFCKFAAA-NCPFGHGCSHV 102
Query: 155 HGDLCPSCGRQCLHPFRPEEREEHMMSCRNKQKHLEALKRSQEIECSVCLERVLSKPTAA 214
HG+ C C + CLHP E+E H+ +C K+K++ ALK S+EIEC+VCLE VLSKP +
Sbjct: 103 HGNQCLYCRKYCLHPSDRREKENHLKTCDKKEKYVLALKNSEEIECNVCLEHVLSKPKPS 162
Query: 215 ERKFGLLSECDHPFCVSCIRNWRSSNPT----LGMDVNSTLRACPICRKLSYFVVPSVIW 270
ERKFGLL ECDH FC+SCIRNWR+S P+ G ++N T+R CP+CR+LSYFV+PS IW
Sbjct: 163 ERKFGLLPECDHAFCLSCIRNWRNSAPSSEFETGNNIN-TVRTCPVCRQLSYFVIPSGIW 221
Query: 271 YATSEEKMEIIDTYKAKLKSIDCKHFDFGEGNCPFGTSCFYKHAYRDG 318
Y T EEK EIID YKA + IDCKHF+ G GNCPFG SCFYKH + G
Sbjct: 222 YTTKEEKQEIIDNYKANCRLIDCKHFESGNGNCPFGASCFYKHTVKPG 269
>UniRef100_Q9H000 Makorin 2 [Homo sapiens]
Length = 416
Score = 249 bits (637), Expect = 6e-65
Identities = 132/355 (37%), Positives = 196/355 (55%), Gaps = 37/355 (10%)
Query: 1 VLCKFFAHGACLKGEHCEFSHDW-KAPPNNICTFYQKGVCAYGSRCRYDHVKASRA---- 55
+ C++F HG C +G C FSHD + P+ IC +YQKG CAYG+RCRYDH + S A
Sbjct: 6 ITCRYFMHGVCREGSQCLFSHDLANSKPSTICKYYQKGYCAYGTRCRYDHTRPSAAAGGA 65
Query: 56 -------------QSSTPSSSIT-------EHQPLVSESAVLGNTRVTSNGVATA-AEFS 94
S P S +T H+P E L +G+A + S
Sbjct: 66 VGTMAHSVPSPAFHSPHPPSEVTASIVKTNSHEPGKREKRTLVLRDRNLSGMAERKTQPS 125
Query: 95 LFSTPFVL--PSEQAWNQESAQLDFLRED-DVVQSVITSPSELPICSFAAAGNCPRGEQC 151
+ S P P + + LD +R D V++ + +E +C +AAAG C G+ C
Sbjct: 126 MVSNPGSCSDPQPSPEMKPHSYLDAIRSGLDDVEASSSYSNEQQLCPYAAAGECRFGDAC 185
Query: 152 PHVHGDLCPSCGRQCLHPFRPEEREEH----MMSCRNKQKHLEALKRSQEIECSVCLERV 207
++HG++C C Q LHPF PE+R+ H M++ ++ + A + SQ+ CS+C+E +
Sbjct: 186 VYLHGEVCEICRLQVLHPFDPEQRKAHEKICMLTFEHEMEKAFAFQASQDKVCSICMEVI 245
Query: 208 LSKPTAAERKFGLLSECDHPFCVSCIRNWRSSNPTLGMDVNSTLRACPICRKLSYFVVPS 267
L K +A+ER+FG+LS C+H +C+SCIR WR + N +++CP CR +S FV+PS
Sbjct: 246 LEKASASERRFGILSNCNHTYCLSCIRQWRCAKQF----ENPIIKSCPECRVISEFVIPS 301
Query: 268 VIWYATSEEKMEIIDTYKAKLKSIDCKHFDFGEGNCPFGTSCFYKHAYRDGRLEE 322
V W +K E+I+ +K + CK+F+ G+G CPFG+ C Y+HAY DGRL E
Sbjct: 302 VYWVEDQNKKNELIEAFKQGMGKKACKYFEQGKGTCPFGSKCLYRHAYPDGRLAE 356
>UniRef100_UPI000036B3BD UPI000036B3BD UniRef100 entry
Length = 418
Score = 241 bits (615), Expect = 2e-62
Identities = 132/362 (36%), Positives = 195/362 (53%), Gaps = 46/362 (12%)
Query: 3 CKFFAHGACLKGEHCEFSHDW-KAPPNNICTFYQKGVCAYGSRCR---------YDHVKA 52
C++F HG C +G C FSHD + P+ IC +YQKG CAYG+RCR YDH +
Sbjct: 1 CRYFMHGVCREGSQCLFSHDLANSKPSTICKYYQKGYCAYGTRCRQGLCNRLQLYDHTRP 60
Query: 53 SRA-----------------QSSTPSSSIT-------EHQPLVSESAVLGNTRVTSNGVA 88
S A S P S +T H+P E L +G+A
Sbjct: 61 SAAAGGAVGTMAHSVPSPAFHSPHPPSEVTASIVKTNSHEPGKREKRTLVLRDRNLSGMA 120
Query: 89 TA-AEFSLFSTPFVL--PSEQAWNQESAQLDFLRED-DVVQSVITSPSELPICSFAAAGN 144
+ S+ S P P + + LD +R D V++ + +E +C +AAAG
Sbjct: 121 EGKTQPSMASNPGSCSDPQPSPEMKPHSYLDAIRSGLDDVEASSSYSNEQQLCPYAAAGE 180
Query: 145 CPRGEQCPHVHGDLCPSCGRQCLHPFRPEEREEH----MMSCRNKQKHLEALKRSQEIEC 200
C G+ C ++HG++C C Q LHPF PE+R+ H M++ ++ + A + SQ+ C
Sbjct: 181 CRFGDACVYLHGEVCEICRLQVLHPFDPEQRKAHEKICMLTFEHEMEKAFAFQASQDKVC 240
Query: 201 SVCLERVLSKPTAAERKFGLLSECDHPFCVSCIRNWRSSNPTLGMDVNSTLRACPICRKL 260
S+C+E +L K +A+ER+FG+LS C+H +C+SCIR WR + N +++CP CR +
Sbjct: 241 SICMEVILEKASASERRFGILSNCNHTYCLSCIRQWRCAKQF----ENPIIKSCPECRVI 296
Query: 261 SYFVVPSVIWYATSEEKMEIIDTYKAKLKSIDCKHFDFGEGNCPFGTSCFYKHAYRDGRL 320
S FV+PSV W +K E+I+ +K + CK+F+ G+G CPFG+ C Y+HAY DGRL
Sbjct: 297 SEFVIPSVYWVEDQNKKNELIEAFKQGMGKKACKYFEQGKGTCPFGSKCLYRHAYPDGRL 356
Query: 321 EE 322
E
Sbjct: 357 AE 358
>UniRef100_Q9TT91 Makorin 1 [Macropus eugenii]
Length = 478
Score = 241 bits (615), Expect = 2e-62
Identities = 130/357 (36%), Positives = 195/357 (54%), Gaps = 31/357 (8%)
Query: 1 VLCKFFAHGACLKGEHCEFSHDWKAPPNN-ICTFYQKGVCAYGSRCRYDHVKASR----- 54
V C++F HG C KG +C +SHD + +C +YQ+G CAYG RCRY+H K +
Sbjct: 55 VTCRYFMHGVCKKGNNCRYSHDLSTSQSAMVCRYYQRGCCAYGDRCRYEHTKPLKREEVT 114
Query: 55 -----AQSSTPSSSITEH--QPLVSESAVLGNTRVTSNGVATAA--EFSLFSTPFVLPSE 105
A+S P+SS +PL S + V SN A A E + + FV P +
Sbjct: 115 AANLAAKSDLPASSSLPALVEPLAEVSTGEAES-VNSNFAAAGAGGEDWVNAIEFV-PGQ 172
Query: 106 QAWNQ------ESAQLDFLREDDVVQSVITSPSELPICSFAAAGNCPRGEQCPHVHGDLC 159
+ E+ + E+++ + + +C +AA G C GE C ++HGD C
Sbjct: 173 PYCGRAAPSCTEAPLQGMVIEEELEKQQTNVEMKKQLCPYAAVGECRYGENCVYLHGDAC 232
Query: 160 PSCGRQCLHPFRPEEREEHMMSC-RNKQKHLE---ALKRSQEIECSVCLERVLSKPTAAE 215
CG Q LHP +R +H+ SC +K +E A++RS+++ C +C+E V K +E
Sbjct: 233 DMCGLQVLHPVDAAQRSQHIKSCIEAHEKDMELSFAVQRSKDMVCGICMEVVYEKANPSE 292
Query: 216 RKFGLLSECDHPFCVSCIRNWRSSNPTLGMDVNSTLRACPICRKLSYFVVPSVIWYATSE 275
R+FG+LS C+H +C+ CIR WRS+ + +++CP CR S FV+PS W E
Sbjct: 293 RRFGILSNCNHTYCLKCIRKWRSAKQF----ESKIIKSCPECRITSNFVIPSEYWVEEKE 348
Query: 276 EKMEIIDTYKAKLKSIDCKHFDFGEGNCPFGTSCFYKHAYRDGRLEEVALRHLGAAD 332
EK ++I YK + + C++FD G G+CPFG +CFYKHAY DGR EE + +G ++
Sbjct: 349 EKQKLIQKYKEAMSNKPCRYFDEGRGSCPFGGNCFYKHAYPDGRREEPQRQKVGTSN 405
>UniRef100_Q9UHC7 Makorin 1 [Homo sapiens]
Length = 482
Score = 239 bits (609), Expect = 1e-61
Identities = 124/357 (34%), Positives = 196/357 (54%), Gaps = 33/357 (9%)
Query: 1 VLCKFFAHGACLKGEHCEFSHDWKAPPNNI-CTFYQKGVCAYGSRCRYDHVKASRAQSST 59
V C++F HG C +G++C +SHD P ++ C ++Q+G C YG RCRY+H K + + +T
Sbjct: 59 VTCRYFMHGVCKEGDNCRYSHDLSDSPYSVVCKYFQRGYCIYGDRCRYEHSKPLKQEEAT 118
Query: 60 PSSSITEHQPLVSES--AVLGNTRVTSNGVATAAEFSLFST------------------P 99
+ T+ S S +++G + G A + S F+T P
Sbjct: 119 ATELTTKSSLAASSSLSSIVGPLVEMNTGEAESRN-SNFATVGAGSEDWVNAIEFVPGQP 177
Query: 100 FVLPSEQAWNQESAQLDFLRED-DVVQSVITSPSELPICSFAAAGNCPRGEQCPHVHGDL 158
+ + + + Q +E+ + Q+ + + +L C +AA G C GE C ++HGD
Sbjct: 178 YCGRTAPSCTEAPLQGSVTKEESEKEQTAVETKKQL--CPYAAVGECRYGENCVYLHGDS 235
Query: 159 CPSCGRQCLHPFRPEEREEHMMSC-RNKQKHLE---ALKRSQEIECSVCLERVLSKPTAA 214
C CG Q LHP +R +H+ SC +K +E A++RS+++ C +C+E V K +
Sbjct: 236 CDMCGLQLLHPMDAAQRSQHIKSCIEAHEKDMELSFAVQRSKDMVCGICMEVVYEKANPS 295
Query: 215 ERKFGLLSECDHPFCVSCIRNWRSSNPTLGMDVNSTLRACPICRKLSYFVVPSVIWYATS 274
ER+FG+LS C+H +C+ CIR WRS+ + +++CP CR S FV+PS W
Sbjct: 296 ERRFGILSNCNHTYCLKCIRKWRSAKQF----ESKIIKSCPECRITSNFVIPSEYWVEEK 351
Query: 275 EEKMEIIDTYKAKLKSIDCKHFDFGEGNCPFGTSCFYKHAYRDGRLEEVALRHLGAA 331
EEK ++I YK + + C++FD G G+CPFG +CFYKHAY DGR EE + +G +
Sbjct: 352 EEKQKLILKYKEAMSNKACRYFDEGRGSCPFGGNCFYKHAYPDGRREEPQRQKVGTS 408
>UniRef100_Q6GLT5 MGC84269 protein [Xenopus laevis]
Length = 408
Score = 238 bits (607), Expect = 2e-61
Identities = 128/341 (37%), Positives = 190/341 (55%), Gaps = 32/341 (9%)
Query: 1 VLCKFFAHGACLKGEHCEFSHDWKAPPNN-ICTFYQKGVCAYGSRCRYDHVKASRAQSS- 58
V C++F HG C +G +C +SHD + IC ++Q+G CAYG RCRY+H K + +
Sbjct: 38 VTCRYFIHGVCKEGINCRYSHDLATSRSAMICRYFQRGCCAYGDRCRYEHNKPLQEDPTG 97
Query: 59 ----TPSSSITEHQPLV-SESAVLGNTRVTSNGV-----ATAAEF---SLFSTPFVLPSE 105
PS S+ E + S++A L + + S G A EF L+S +
Sbjct: 98 DTCTAPSESLPEPSGNINSKAAELAASELASGGPRAQDWVNAVEFVPGQLYSGR----AP 153
Query: 106 QAWNQESAQLDFLREDDVVQSVITSPSELPICSFAAAGNCPRGEQCPHVHGDLCPSCGRQ 165
+A+ Q + + D RE+ + + +C +AA G C GE C ++HGD C CG Q
Sbjct: 154 EAYTQGTVKPDEGREEPADPEL-----KKQLCPYAAMGECRYGENCVYLHGDPCDMCGLQ 208
Query: 166 CLHPFRPEEREEHMMSC-RNKQKHLE---ALKRSQEIECSVCLERVLSKPTAAERKFGLL 221
LHP +R +H+ SC +K +E A++RS++I C +C+E V K +ER+FG+L
Sbjct: 209 VLHPVDTCQRSQHIKSCIEAHEKDMELSFAVQRSKDIVCGICMEVVYEKTNPSERRFGIL 268
Query: 222 SECDHPFCVSCIRNWRSSNPTLGMDVNSTLRACPICRKLSYFVVPSVIWYATSEEKMEII 281
S C H +C+ CIR WRS+ + +++CP CR S F++PS W EEK ++I
Sbjct: 269 SNCSHSYCLKCIRKWRSAKQF----ESKIIKSCPECRITSNFIIPSEYWVEEKEEKHKLI 324
Query: 282 DTYKAKLKSIDCKHFDFGEGNCPFGTSCFYKHAYRDGRLEE 322
YK + S C++FD G G CPFG +CFY+HAY DGR+EE
Sbjct: 325 HKYKEAMSSKSCRYFDEGRGTCPFGGNCFYRHAYPDGRIEE 365
>UniRef100_Q8C5V4 Mus musculus adult male testis cDNA, RIKEN full-length enriched
library, clone:4932442G21 product:makorin, ring finger
protein, 1, full insert sequence [Mus musculus]
Length = 481
Score = 238 bits (607), Expect = 2e-61
Identities = 126/356 (35%), Positives = 191/356 (53%), Gaps = 31/356 (8%)
Query: 1 VLCKFFAHGACLKGEHCEFSHDWKAPPNNI-CTFYQKGVCAYGSRCRYDH---------- 49
V C++F HG C +G++C +SHD P + C ++Q+G C YG RCRY+H
Sbjct: 59 VTCRYFMHGVCKEGDNCRYSHDLSDSPYGVVCKYFQRGYCVYGDRCRYEHSKPLKQEEVT 118
Query: 50 -----VKASRAQSSTPSSSITEHQPLVSESAVLGNTRVTSNGVAT-----AAEFSLFSTP 99
K S A SS+ SS + + S A N + G + A EF + P
Sbjct: 119 ATDLSAKPSLAASSSLSSGVGSLAEMNSGEAESRNPSFPTVGAGSEDWVNAIEF-VPGQP 177
Query: 100 FVLPSEQAWNQESAQLDFLREDDVVQSVITSPSELPICSFAAAGNCPRGEQCPHVHGDLC 159
+ + + + Q +E+ + T ++ +C +AA G C GE C ++HGD C
Sbjct: 178 YCGRTAPSCTEVPPQGSVTKEESEKEPT-TVETKKQLCPYAAVGECRYGENCVYLHGDSC 236
Query: 160 PSCGRQCLHPFRPEEREEHMMSC-RNKQKHLE---ALKRSQEIECSVCLERVLSKPTAAE 215
CG Q LHP +R +H+ SC +K +E A++RS+++ C +C+E V K +E
Sbjct: 237 DMCGLQVLHPVDAAQRSQHIKSCIEAHEKDMELSFAVQRSKDMVCGICMEVVYEKANPSE 296
Query: 216 RKFGLLSECDHPFCVSCIRNWRSSNPTLGMDVNSTLRACPICRKLSYFVVPSVIWYATSE 275
R+FG+LS C+H +C+ CIR WRS+ + +++CP CR S FV+PS W E
Sbjct: 297 RRFGILSNCNHTYCLKCIRKWRSAKQF----ESKIIKSCPECRITSNFVIPSENWVEEKE 352
Query: 276 EKMEIIDTYKAKLKSIDCKHFDFGEGNCPFGTSCFYKHAYRDGRLEEVALRHLGAA 331
EK ++I YK + + C++FD G G+CPFG +CFYKHAY DGR EE + +G +
Sbjct: 353 EKQKLIQKYKEAMSNKACRYFDEGRGSCPFGGNCFYKHAYPDGRREEPQRQKVGTS 408
>UniRef100_UPI000027CA99 UPI000027CA99 UniRef100 entry
Length = 372
Score = 238 bits (606), Expect = 2e-61
Identities = 128/334 (38%), Positives = 184/334 (54%), Gaps = 23/334 (6%)
Query: 1 VLCKFFAHGACLKGEHCEFSHDW--KAPPNNICTFYQKGVCAYGSRCRYDHVKASR---- 54
V C++F HG C +G++C +SHD P IC F+QKG C +G RCR+DH K ++
Sbjct: 22 VTCRYFMHGLCKEGDNCRYSHDLTNSKPAAMICKFFQKGNCVFGERCRFDHCKPTKNEEF 81
Query: 55 --AQSSTPSSSITEHQPLVSESAVLGNTRVTSNGVATAAEFSLFSTPFVLPSEQAWNQES 112
Q PSS P S+ A T+ +N AAEF + P+ +E + S
Sbjct: 82 SSPQMLPPSSPSPSTDPESSQPAPRPKTQDWAN----AAEF-VPGQPYCGRAESVKVEIS 136
Query: 113 AQLDFLREDDVVQSVITSPSELPICSFAAAGNCPRGEQCPHVHGDLCPSCGRQCLHPFRP 172
L + E D +V +C +AA G C G C ++HGD+C CG Q LHP
Sbjct: 137 IPL--IEELDCDAAVDKEALRKQLCPYAAVGECRYGINCAYLHGDVCDMCGLQVLHPTDN 194
Query: 173 EEREEHMMSC-RNKQKHLE---ALKRSQEIECSVCLERVLSKPTAAERKFGLLSECDHPF 228
+R +H +C +K +E A++RS+++ C VC+E V K +ER+FG+LS C+H +
Sbjct: 195 SQRSQHTKACIEAHEKDMEISFAIQRSKDMMCGVCMEVVFEKANPSERRFGILSNCNHCY 254
Query: 229 CVSCIRNWRSSNPTLGMDVNSTLRACPICRKLSYFVVPSVIWYATSEEKMEIIDTYKAKL 288
C+ CIR WRS+ + +++CP CR S FV+PS W E+K ++I YK +
Sbjct: 255 CLKCIRKWRSAKQF----ESKIIKSCPECRITSNFVIPSEYWVEDKEDKQKLIQKYKDGM 310
Query: 289 KSIDCKHFDFGEGNCPFGTSCFYKHAYRDGRLEE 322
C++FD G G CPFG +CFYKHA+ DGRLEE
Sbjct: 311 GRKPCRYFDEGRGICPFGANCFYKHAFPDGRLEE 344
>UniRef100_Q9QXP6 Makorin 1 [Mus musculus]
Length = 481
Score = 238 bits (606), Expect = 2e-61
Identities = 125/356 (35%), Positives = 191/356 (53%), Gaps = 31/356 (8%)
Query: 1 VLCKFFAHGACLKGEHCEFSHDWKAPPNNI-CTFYQKGVCAYGSRCRYDH---------- 49
V C++F HG C +G++C +SHD P + C ++Q+G C YG RCRY+H
Sbjct: 59 VTCRYFMHGVCKEGDNCRYSHDLSDSPYGVVCKYFQRGYCVYGDRCRYEHSKPLKQEEVT 118
Query: 50 -----VKASRAQSSTPSSSITEHQPLVSESAVLGNTRVTSNGVAT-----AAEFSLFSTP 99
K S A SS+ SS + + S A N + G + A EF + P
Sbjct: 119 ATDLSAKPSLAASSSLSSGVGSLAEMNSGEAESRNPSFPTVGAGSEDWVNAIEF-VPGQP 177
Query: 100 FVLPSEQAWNQESAQLDFLREDDVVQSVITSPSELPICSFAAAGNCPRGEQCPHVHGDLC 159
+ + + + Q +E+ + T ++ +C +AA G C GE C ++HGD C
Sbjct: 178 YCGRTAPSCTEVPPQGSVTKEESEKEPT-TVETKKQLCPYAAVGECRYGENCVYLHGDSC 236
Query: 160 PSCGRQCLHPFRPEEREEHMMSC-RNKQKHLE---ALKRSQEIECSVCLERVLSKPTAAE 215
CG Q LHP +R +H+ SC +K +E A++R++++ C +C+E V K +E
Sbjct: 237 DMCGLQVLHPVDAAQRSQHIKSCIEAHEKDMELSFAVQRTKDMVCGICMEVVYEKANPSE 296
Query: 216 RKFGLLSECDHPFCVSCIRNWRSSNPTLGMDVNSTLRACPICRKLSYFVVPSVIWYATSE 275
R+FG+LS C+H +C+ CIR WRS+ + +++CP CR S FV+PS W E
Sbjct: 297 RRFGILSNCNHTYCLKCIRKWRSAKQF----ESKIIKSCPECRITSNFVIPSEYWVEEKE 352
Query: 276 EKMEIIDTYKAKLKSIDCKHFDFGEGNCPFGTSCFYKHAYRDGRLEEVALRHLGAA 331
EK ++I YK + + C++FD G G+CPFG +CFYKHAY DGR EE + +G +
Sbjct: 353 EKQKLIQKYKEAMSNKACRYFDEGRGSCPFGGNCFYKHAYPDGRREEPQRQKVGTS 408
>UniRef100_UPI00003AA829 UPI00003AA829 UniRef100 entry
Length = 417
Score = 237 bits (604), Expect = 4e-61
Identities = 127/356 (35%), Positives = 190/356 (52%), Gaps = 40/356 (11%)
Query: 1 VLCKFFAHGACLKGEHCEFSHDWKAPPNN-ICTFYQKGVCAYGSRCRYDHVK-------A 52
V C+++ G C +G C FSHD + ++ +C +YQKG CAYG+RCRYDH++ A
Sbjct: 6 VTCRYYVQGVCREGSKCLFSHDLSSSKSSTVCKYYQKGQCAYGARCRYDHIRLPASGGAA 65
Query: 53 SRAQSSTPSSSITEHQPLVSESAVLGNTRVTSNG----------------VATAAEFSLF 96
+ A S+++ +P SA +++ +G + E +
Sbjct: 66 APAPPPMASAALLSPRPAPEHSAPAAKSKLRESGKREKKTLVLRDRNLCGLNEEKEKASA 125
Query: 97 STPFVLPSEQAWNQESAQLDFLR------EDDVVQSVITSPSELPICSFAAAGNCPRGEQ 150
+ V S+++ N E +L ED V S +L C +AAAG C G++
Sbjct: 126 TNDVVCCSDKSDNVEMKPHSYLEAICSGLEDPVAGSSFGDGEQL--CPYAAAGACHFGDR 183
Query: 151 CPHVHGDLCPSCGRQCLHPFRPEEREEHMMSCRNKQKH----LEALKRSQEIECSVCLER 206
C ++HGD+C CG Q LHPF E+R+ H M C +H A + SQ+ CS+C+E
Sbjct: 184 CLYLHGDVCEICGLQVLHPFDQEQRKAHEMMCMATFEHEMEKAFAFQASQDKVCSICMEV 243
Query: 207 VLSKPTAAERKFGLLSECDHPFCVSCIRNWRSSNPTLGMDVNSTLRACPICRKLSYFVVP 266
V KP+A+ER+FG+LS C+H +C+SCIR WR + N +++CP CR +S FV+P
Sbjct: 244 VYEKPSASERRFGILSNCNHTYCLSCIRQWRCAKQF----ENPIIKSCPECRVISEFVIP 299
Query: 267 SVIWYATSEEKMEIIDTYKAKLKSIDCKHFDFGEGNCPFGTSCFYKHAYRDGRLEE 322
S W E+K E+I+ +K + CK+F+ G+G CPFG C Y HAY DG E
Sbjct: 300 SAYWVEDQEKKNELIEAFKQGVGKKPCKYFEQGKGTCPFGGKCLYFHAYPDGTRAE 355
>UniRef100_Q8UWH3 Makorin-2 [Gallus gallus]
Length = 417
Score = 237 bits (604), Expect = 4e-61
Identities = 127/356 (35%), Positives = 190/356 (52%), Gaps = 40/356 (11%)
Query: 1 VLCKFFAHGACLKGEHCEFSHDWKAPPNN-ICTFYQKGVCAYGSRCRYDHVK-------A 52
V C+++ G C +G C FSHD + ++ +C +YQKG CAYG+RCRYDH++ A
Sbjct: 6 VTCRYYVQGVCREGSKCLFSHDLSSSKSSTVCKYYQKGQCAYGARCRYDHIRLPASGGAA 65
Query: 53 SRAQSSTPSSSITEHQPLVSESAVLGNTRVTSNG----------------VATAAEFSLF 96
+ A S+++ +P SA +++ +G + E +
Sbjct: 66 APAPPPKASAALLSPRPAPEHSAPAAKSKLRESGKREKKTLVLRDRNLCGLNEEKEKAST 125
Query: 97 STPFVLPSEQAWNQESAQLDFLR------EDDVVQSVITSPSELPICSFAAAGNCPRGEQ 150
+ V S+++ N E +L ED V S +L C +AAAG C G++
Sbjct: 126 TNNVVCCSDKSDNVEMKPHSYLEAICSGLEDPVAGSSFGDGEQL--CPYAAAGACHFGDR 183
Query: 151 CPHVHGDLCPSCGRQCLHPFRPEEREEHMMSCRNKQKH----LEALKRSQEIECSVCLER 206
C ++HGD+C CG Q LHPF E+R+ H M C +H A + SQ+ CS+C+E
Sbjct: 184 CLYLHGDVCEICGLQVLHPFDQEQRKAHEMMCMATFEHEMEKAFAFQASQDKVCSICMEV 243
Query: 207 VLSKPTAAERKFGLLSECDHPFCVSCIRNWRSSNPTLGMDVNSTLRACPICRKLSYFVVP 266
V KP+A+ER+FG+LS C+H +C+SCIR WR + N +++CP CR +S FV+P
Sbjct: 244 VYEKPSASERRFGILSNCNHTYCLSCIRQWRCAKQF----ENPIIKSCPECRVISEFVIP 299
Query: 267 SVIWYATSEEKMEIIDTYKAKLKSIDCKHFDFGEGNCPFGTSCFYKHAYRDGRLEE 322
S W E+K E+I+ +K + CK+F+ G+G CPFG C Y HAY DG E
Sbjct: 300 SAYWVEDQEKKNELIEAFKQGVGKKPCKYFEQGKGTCPFGGKCLYLHAYPDGTRAE 355
>UniRef100_Q8JFF3 Gene encoding protein featuring ring-finger [Seriola
quinqueradiata]
Length = 435
Score = 236 bits (603), Expect = 6e-61
Identities = 128/334 (38%), Positives = 185/334 (55%), Gaps = 21/334 (6%)
Query: 1 VLCKFFAHGACLKGEHCEFSHDW--KAPPNNICTFYQKGVCAYGSRCRYDHVKASR---- 54
V C++F HG C +G++C +SHD P IC F+QKG C +G RCR++H K ++
Sbjct: 22 VTCRYFMHGLCKEGDNCRYSHDLTNSKPAAMICKFFQKGNCVFGDRCRFEHCKPAKNEEL 81
Query: 55 -AQSSTPSSSITEHQPLVSESAVLGNTRVT-SNGVATAAEFSLFSTPFVLPSEQAWNQES 112
A P S + P S+ G T V + AAEF + P+ +EQA + S
Sbjct: 82 PAPQMLPLPSASLAGP--SDPEPSGPTPVPGAQDWVNAAEF-VPGQPYCGRAEQAKVESS 138
Query: 113 AQLDFLREDDVVQSVITSPSELPICSFAAAGNCPRGEQCPHVHGDLCPSCGRQCLHPFRP 172
L + E D + +C +AA G C G C ++HGD+C CG Q LHP
Sbjct: 139 VPL--IEEFDSYPAPDNKQLRKQLCPYAAVGECRYGINCAYLHGDVCYMCGLQVLHPTDN 196
Query: 173 EEREEHMMSC-RNKQKHLE---ALKRSQEIECSVCLERVLSKPTAAERKFGLLSECDHPF 228
+R EH +C +K +E A++RS+++ C VC+E V K +ER+FG+LS C H +
Sbjct: 197 NQRSEHTKACIEAHEKDMEISFAIQRSKDMMCGVCMEVVFEKANPSERRFGILSNCSHCY 256
Query: 229 CVSCIRNWRSSNPTLGMDVNSTLRACPICRKLSYFVVPSVIWYATSEEKMEIIDTYKAKL 288
C+ CIR WRS+ + +++CP CR S FV+PS W ++K ++I YK +
Sbjct: 257 CLKCIRKWRSAKQF----ESKIIKSCPECRITSNFVIPSEYWVEDKDDKQKLIQKYKDGM 312
Query: 289 KSIDCKHFDFGEGNCPFGTSCFYKHAYRDGRLEE 322
S C++FD G G CPFG++CFYKHA+ DGRLEE
Sbjct: 313 GSKPCRYFDEGRGTCPFGSNCFYKHAFPDGRLEE 346
>UniRef100_Q9DFG7 Makorin RING zinc-finger protein 1 [Brachydanio rerio]
Length = 442
Score = 235 bits (599), Expect = 2e-60
Identities = 124/338 (36%), Positives = 181/338 (52%), Gaps = 21/338 (6%)
Query: 1 VLCKFFAHGACLKGEHCEFSHDWKAPPNN-ICTFYQKGVCAYGSRCRYDHVKASRAQ--- 56
V C++F HG C +GE+C +SHD + IC F+QKG CA+G RCRY+H K S+
Sbjct: 25 VTCRYFMHGLCKEGENCRYSHDLSSCKQTMICKFFQKGCCAFGDRCRYEHTKPSKQDEVP 84
Query: 57 SSTPSSSITEHQPLVSESAVL-----GNTRVTSNGVATAAEFSLFSTPFVLPSEQAWNQE 111
SS PS +T PL + G T + A + + FV +
Sbjct: 85 SSKPSMPLTA-APLAGTPEPVSDGPGGTTGAQEKPQGSGAVDWVNAAEFVPGQPYCGRAD 143
Query: 112 SAQLDF---LREDDVVQSVITSPSELPICSFAAAGNCPRGEQCPHVHGDLCPSCGRQCLH 168
+ L E++ + + +C +AA G C G C ++HGD+C CG Q LH
Sbjct: 144 PVLCEGPGPLIEEEYEKEQANKEMKKQLCPYAAVGECRYGLNCAYLHGDVCDMCGLQVLH 203
Query: 169 PFRPEEREEHMMSC-RNKQKHLE---ALKRSQEIECSVCLERVLSKPTAAERKFGLLSEC 224
P +R +H+ +C +K +E A++RS+++ C VC+E V K +ER+FG+LS C
Sbjct: 204 PSDTSQRSQHIRACIEAHEKDMEISFAIQRSKDMMCGVCMEVVFEKTNPSERRFGILSNC 263
Query: 225 DHPFCVSCIRNWRSSNPTLGMDVNSTLRACPICRKLSYFVVPSVIWYATSEEKMEIIDTY 284
H +C+ CIR WRS+ + +++CP CR S V+PS W EEK ++I Y
Sbjct: 264 CHCYCLKCIRKWRSAKQF----ESKIIKSCPECRITSNLVIPSEYWVEDKEEKQQLIQKY 319
Query: 285 KAKLKSIDCKHFDFGEGNCPFGTSCFYKHAYRDGRLEE 322
K + + C++FD G G CPFG +CFYKHA+ DGRLEE
Sbjct: 320 KDGMGTKPCRYFDEGRGTCPFGANCFYKHAFPDGRLEE 357
>UniRef100_UPI000036DFF0 UPI000036DFF0 UniRef100 entry
Length = 421
Score = 234 bits (597), Expect = 3e-60
Identities = 122/353 (34%), Positives = 193/353 (54%), Gaps = 33/353 (9%)
Query: 5 FFAHGACLKGEHCEFSHDWKAPPNNI-CTFYQKGVCAYGSRCRYDHVKASRAQSSTPSSS 63
+F HG C +G++C +SHD P ++ C ++Q+G C YG RCRY+H K + + +T +
Sbjct: 2 YFMHGVCKEGDNCRYSHDLSDSPYSVVCKYFQRGYCIYGDRCRYEHSKPLKQEEATATEL 61
Query: 64 ITEHQPLVSES--AVLGNTRVTSNGVATAAEFSLFST------------------PFVLP 103
T+ S S +++G + G A + S F+T P+
Sbjct: 62 TTKSSLAASSSLSSIVGPLVEMNTGEAESRN-SNFATVGAGSEDWVNAIEFVPGQPYCGR 120
Query: 104 SEQAWNQESAQLDFLRED-DVVQSVITSPSELPICSFAAAGNCPRGEQCPHVHGDLCPSC 162
+ + + Q +E+ + Q+ + + +L C +AA G C GE C ++HGD C C
Sbjct: 121 TAPSCTEAPLQGSVTKEESEKEQTAVETKKQL--CPYAAVGECRYGENCVYIHGDSCDMC 178
Query: 163 GRQCLHPFRPEEREEHMMSC-RNKQKHLE---ALKRSQEIECSVCLERVLSKPTAAERKF 218
G Q LHP +R +H+ SC +K +E A++RS+++ C +C+E V K +ER+F
Sbjct: 179 GLQVLHPMDAAQRSQHIKSCIEAHEKDMELSFAVQRSKDMVCGICMEVVYEKANPSERRF 238
Query: 219 GLLSECDHPFCVSCIRNWRSSNPTLGMDVNSTLRACPICRKLSYFVVPSVIWYATSEEKM 278
G+LS C+H +C+ CIR WRS+ + +++CP CR S FV+PS W EEK
Sbjct: 239 GILSNCNHTYCLKCIRKWRSAKQF----ESKIIKSCPECRITSNFVIPSEYWVEEKEEKQ 294
Query: 279 EIIDTYKAKLKSIDCKHFDFGEGNCPFGTSCFYKHAYRDGRLEEVALRHLGAA 331
++I YK + + C++FD G G+CPFG +CFYKHAY DGR EE + +G +
Sbjct: 295 KLILKYKEAMSNKACRYFDEGRGSCPFGGNCFYKHAYPDGRREEPQRQKVGTS 347
>UniRef100_Q6P0Z6 Mkrn2 protein [Brachydanio rerio]
Length = 414
Score = 231 bits (588), Expect = 3e-59
Identities = 127/354 (35%), Positives = 175/354 (48%), Gaps = 36/354 (10%)
Query: 1 VLCKFFAHGACLKGEHCEFSHDWK-APPNNICTFYQKGVCAYGSRCRYDHVKASRAQSST 59
V C++F HG C +G C FSHD + P+ IC +YQ+G CAYG RCRYDH+K S
Sbjct: 6 VTCRYFLHGVCREGSRCLFSHDLTTSKPSTICKYYQRGACAYGDRCRYDHIKPPGRGSGA 65
Query: 60 PSSSITEHQPLVSESAV-LGNTRVTSNG-----VATAAEFSLFSTPFVLPSE-----QAW 108
P+ SA G TS V S P V +E + W
Sbjct: 66 PADHSNRSSSSAGASAPGPGPPAHTSKHLKKPLVLRDKALCSDSRPRVFSAESSELNECW 125
Query: 109 NQES-------AQLDFLRED-DVVQSVITSPSELP--------ICSFAAAGNCPRGEQCP 152
Q + L+ +R D + + P IC F AAG C GE CP
Sbjct: 126 EQRDDGAQKPHSYLEAIRSGLDASAAAAAAAGTFPELQQTSPQICPFLAAGQCQYGESCP 185
Query: 153 HVHGDLCPSCGRQCLHPFRPEEREEHMMSC----RNKQKHLEALKRSQEIECSVCLERVL 208
++HG++C C + LHP PE+R H C + A+++SQ+ C +CL+ V
Sbjct: 186 YLHGEMCEICRQHVLHPHDPEQRAAHEKKCMVAFEMDMERAFAVQQSQDKVCKICLDVVY 245
Query: 209 SKPTAAERKFGLLSECDHPFCVSCIRNWRSSNPTLGMDVNSTLRACPICRKLSYFVVPSV 268
K + +ER+FG+LS C H +C++CIR WR N ++CP CR +S FV+PS+
Sbjct: 246 EKSSPSERRFGILSSCAHTYCLNCIRQWRCVEQL----HNQIRKSCPECRVVSEFVIPSI 301
Query: 269 IWYATSEEKMEIIDTYKAKLKSIDCKHFDFGEGNCPFGTSCFYKHAYRDGRLEE 322
W E+K +I+ +K+ + CK+FD G G CPFG CFY HAY DGR E
Sbjct: 302 YWVEDQEQKNLLIEEFKSGVSKKACKYFDQGRGTCPFGGKCFYMHAYADGRRAE 355
>UniRef100_Q13434 Makorin 4 [Homo sapiens]
Length = 485
Score = 228 bits (580), Expect = 3e-58
Identities = 124/360 (34%), Positives = 186/360 (51%), Gaps = 39/360 (10%)
Query: 1 VLCKFFAHGACLKGEHCEFSHDWKAPPNNI-CTFYQKGVCAYGSRCRYDHVKASRAQSST 59
V C++F +G C +G++C +SHD + C ++Q+G C YG RCR +H K + + +T
Sbjct: 94 VTCRYFKYGICKEGDNCRYSHDLSDRLCGVVCKYFQRGCCVYGDRCRCEHSKPLKQEEAT 153
Query: 60 PSSSITEHQPLVSES--AVLGNTRVTSNGVATAAEFSLFSTPFVLPSEQAWNQESAQLDF 117
+ T+ S S +++G V N + S F+T V+ + W + ++F
Sbjct: 154 ATELTTKSSLAASSSLSSIVGPL-VEMNTNEAESRNSNFAT--VVAGSEDW---ANAIEF 207
Query: 118 LREDDVVQSVITSPSELPI----------------------CSFAAAGNCPRGEQCPHVH 155
+ + S +E P+ C +AA G C GE C ++H
Sbjct: 208 VPGQPYCGRTVPSCTEAPLQGSVTKEESEEEQTAVETKKQLCPYAAVGQCRYGENCVYLH 267
Query: 156 GDLCPSCGRQCLHPFRPEEREEHMMSC-RNKQKHLE---ALKRSQEIECSVCLERVLSKP 211
GDLC CG Q LHP +R +H+ +C +K +E A++RS++ C +C+E V K
Sbjct: 268 GDLCDMCGLQVLHPMDAAQRSQHIQACIEAHEKDMEFSFAVQRSKDKVCGICMEVVYEKA 327
Query: 212 TAAERKFGLLSECDHPFCVSCIRNWRSSNPTLGMDVNSTLRACPICRKLSYFVVPSVIWY 271
E +FG+LS C+H FC+ CIR WRS+ V S CP CR S FV+PS W
Sbjct: 328 NPNEHRFGILSNCNHTFCLKCIRKWRSAKEFESRIVKS----CPQCRITSNFVIPSEYWV 383
Query: 272 ATSEEKMEIIDTYKAKLKSIDCKHFDFGEGNCPFGTSCFYKHAYRDGRLEEVALRHLGAA 331
EEK ++I YK + + CK+FD G G+CPFG +CFYKH Y DGR EE + +G +
Sbjct: 384 EEKEEKQKLIQKYKEAMSNKACKYFDEGRGSCPFGENCFYKHMYPDGRREEPQRQQVGTS 443
>UniRef100_UPI000001360B UPI000001360B UniRef100 entry
Length = 402
Score = 226 bits (575), Expect = 1e-57
Identities = 125/342 (36%), Positives = 178/342 (51%), Gaps = 26/342 (7%)
Query: 3 CKFFAHGACLKGEHCEFSHD-WKAPPNNICTFYQKGVCAYGSRCRYDHVK-ASRAQSSTP 60
C++F HG C +G HC+FSHD + P+ IC FYQ+G CAYG RCRYDHVK +SR +
Sbjct: 1 CRYFLHGVCREGNHCQFSHDPSSSKPSTICKFYQRGTCAYGERCRYDHVKLSSRGGGAFD 60
Query: 61 SSSITEHQPLVSESAVLGNTRVTSNGVATAAEFSLFSTP---FVLPSEQAWNQESAQ--- 114
+ + + S T V + S+ +P F P+E A
Sbjct: 61 MAGVGGARDGASTRGAAKKTFVHQERASVLLLPSVLLSPENMFRAPAESFGADVMAPAPH 120
Query: 115 --LDFLR-------EDDVVQSVITSPSELP-ICSFAAAGNCPRGEQCPHVHGDLCPSCGR 164
+D +R +D ++ +LP +C +AA G+C E C ++HGD C CG
Sbjct: 121 TYVDAIRTGLSSSSQDHTPPTMAGVNQDLPRLCPYAAVGHCYYEENCIYLHGDKCEVCGL 180
Query: 165 QCLHPFRPEEREEHMMSC----RNKQKHLEALKRSQEIECSVCLERVLSKPTAAERKFGL 220
Q L P PE+R H C + A++ SQE CS+C+E V+ K ++R+FG+
Sbjct: 181 QVLDPHNPEQRSMHEKMCLLAFEADMEKAFAVQLSQEKVCSICMEVVVQKMNPSDRRFGI 240
Query: 221 LSECDHPFCVSCIRNWRSSNPTLGMDVNSTLRACPICRKLSYFVVPSVIWYATSEEKMEI 280
LS C H FC++CIR WR + N +++CP CR S FV+PSV W E+K +
Sbjct: 241 LSSCCHVFCLACIRKWRCTRNFS----NKIIKSCPECRVASEFVIPSVYWVENQEDKDHL 296
Query: 281 IDTYKAKLKSIDCKHFDFGEGNCPFGTSCFYKHAYRDGRLEE 322
I+ +K+ + CK+FD G G+CPFG C Y HA DG E
Sbjct: 297 IELFKSGVGKKPCKYFDQGRGSCPFGGKCLYLHALPDGSRAE 338
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.322 0.135 0.435
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 621,756,598
Number of Sequences: 2790947
Number of extensions: 24984855
Number of successful extensions: 66132
Number of sequences better than 10.0: 1051
Number of HSP's better than 10.0 without gapping: 244
Number of HSP's successfully gapped in prelim test: 807
Number of HSP's that attempted gapping in prelim test: 63343
Number of HSP's gapped (non-prelim): 2624
length of query: 358
length of database: 848,049,833
effective HSP length: 128
effective length of query: 230
effective length of database: 490,808,617
effective search space: 112885981910
effective search space used: 112885981910
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 75 (33.5 bits)
Medicago: description of AC146566.2


