

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146564.3 - phase: 0 /pseudo
(281 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
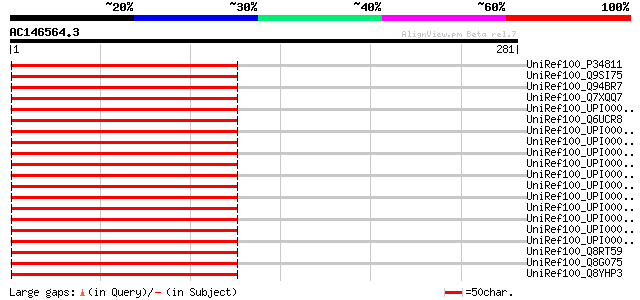
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_P34811 Elongation factor G, chloroplast precursor [Gly... 186 8e-46
UniRef100_Q9SI75 F23N19.11 [Arabidopsis thaliana] 180 3e-44
UniRef100_Q94BR7 Hypothetical protein At1g62750 [Arabidopsis tha... 180 3e-44
UniRef100_Q7XQQ7 OSJNBa0091D06.15 protein [Oryza sativa] 176 6e-43
UniRef100_UPI00002A8CC6 UPI00002A8CC6 UniRef100 entry 164 3e-39
UniRef100_Q6UCR8 Predicted translation elongation factor G [uncu... 164 3e-39
UniRef100_UPI000032D345 UPI000032D345 UniRef100 entry 162 7e-39
UniRef100_UPI000034262E UPI000034262E UniRef100 entry 162 1e-38
UniRef100_UPI00003294D3 UPI00003294D3 UniRef100 entry 161 2e-38
UniRef100_UPI00002C1B69 UPI00002C1B69 UniRef100 entry 160 3e-38
UniRef100_UPI00002AFF51 UPI00002AFF51 UniRef100 entry 160 3e-38
UniRef100_UPI00002D90C0 UPI00002D90C0 UniRef100 entry 159 6e-38
UniRef100_UPI00002C5359 UPI00002C5359 UniRef100 entry 159 8e-38
UniRef100_UPI000029848F UPI000029848F UniRef100 entry 159 8e-38
UniRef100_UPI00003413B4 UPI00003413B4 UniRef100 entry 158 1e-37
UniRef100_UPI00002C653B UPI00002C653B UniRef100 entry 158 1e-37
UniRef100_UPI000029D235 UPI000029D235 UniRef100 entry 158 1e-37
UniRef100_Q8RT59 Elongation factor EfG [Bartonella bacilliformis] 158 2e-37
UniRef100_Q8G075 Elongation factor G [Brucella suis] 158 2e-37
UniRef100_Q8YHP3 Elongation factor G [Brucella melitensis] 158 2e-37
>UniRef100_P34811 Elongation factor G, chloroplast precursor [Glycine max]
Length = 788
Score = 186 bits (471), Expect = 8e-46
Identities = 91/126 (72%), Positives = 108/126 (85%), Gaps = 1/126 (0%)
Query: 2 DIVTLVGLKDTIAGETLCDPKGPFVLDQMDFPVHVIKIAIEPKTKADMEKMEAGLIKLAE 61
DI+ L GLKDTI GETLCDP P VL++MDFP VIK+AIEPKTKAD++KM GLIKLA+
Sbjct: 465 DIIALAGLKDTITGETLCDPDNPIVLERMDFPDPVIKVAIEPKTKADVDKMATGLIKLAQ 524
Query: 62 EDPSFRVSQDKE-DRTIIQGTGELHLETIVDRLKREFKVEANVGAPQANYRESISEVTEV 120
EDPSF S+D+E ++T+I+G GELHLE IVDRLKREFKVEANVGAPQ NYRESIS+++EV
Sbjct: 525 EDPSFHFSRDEEINQTVIEGMGELHLEIIVDRLKREFKVEANVGAPQVNYRESISKISEV 584
Query: 121 RYVFNK 126
+YV K
Sbjct: 585 KYVHKK 590
Score = 37.4 bits (85), Expect = 0.44
Identities = 16/28 (57%), Positives = 22/28 (78%)
Query: 253 LTEGRASYSMQFDRFDTVSRNIQNELTT 280
+T+GRASY+MQ FD V ++IQN+L T
Sbjct: 754 MTKGRASYTMQLAMFDVVPQHIQNQLAT 781
>UniRef100_Q9SI75 F23N19.11 [Arabidopsis thaliana]
Length = 783
Score = 180 bits (457), Expect = 3e-44
Identities = 89/126 (70%), Positives = 106/126 (83%), Gaps = 1/126 (0%)
Query: 2 DIVTLVGLKDTIAGETLCDPKGPFVLDQMDFPVHVIKIAIEPKTKADMEKMEAGLIKLAE 61
DI+ L GLKDTI GETL DP+ P VL++MDFP VIK+AIEPKTKAD++KM GLIKLA+
Sbjct: 460 DIIALAGLKDTITGETLSDPENPVVLERMDFPDPVIKVAIEPKTKADIDKMATGLIKLAQ 519
Query: 62 EDPSFRVSQDKE-DRTIIQGTGELHLETIVDRLKREFKVEANVGAPQANYRESISEVTEV 120
EDPSF S+D+E ++T+I+G GELHLE IVDRLKREFKVEANVGAPQ NYRESIS++ EV
Sbjct: 520 EDPSFHFSRDEEMNQTVIEGMGELHLEIIVDRLKREFKVEANVGAPQVNYRESISKIAEV 579
Query: 121 RYVFNK 126
+Y K
Sbjct: 580 KYTHKK 585
Score = 37.4 bits (85), Expect = 0.44
Identities = 15/28 (53%), Positives = 24/28 (85%)
Query: 253 LTEGRASYSMQFDRFDTVSRNIQNELTT 280
+T+GRASY+MQ +FD V ++IQN+L++
Sbjct: 749 MTKGRASYTMQLAKFDVVPQHIQNQLSS 776
>UniRef100_Q94BR7 Hypothetical protein At1g62750 [Arabidopsis thaliana]
Length = 783
Score = 180 bits (457), Expect = 3e-44
Identities = 89/126 (70%), Positives = 106/126 (83%), Gaps = 1/126 (0%)
Query: 2 DIVTLVGLKDTIAGETLCDPKGPFVLDQMDFPVHVIKIAIEPKTKADMEKMEAGLIKLAE 61
DI+ L GLKDTI GETL DP+ P VL++MDFP VIK+AIEPKTKAD++KM GLIKLA+
Sbjct: 460 DIIALAGLKDTITGETLSDPENPVVLERMDFPDPVIKVAIEPKTKADIDKMATGLIKLAQ 519
Query: 62 EDPSFRVSQDKE-DRTIIQGTGELHLETIVDRLKREFKVEANVGAPQANYRESISEVTEV 120
EDPSF S+D+E ++T+I+G GELHLE IVDRLKREFKVEANVGAPQ NYRESIS++ EV
Sbjct: 520 EDPSFHFSRDEEMNQTVIEGMGELHLEIIVDRLKREFKVEANVGAPQVNYRESISKIAEV 579
Query: 121 RYVFNK 126
+Y K
Sbjct: 580 KYTHKK 585
Score = 37.4 bits (85), Expect = 0.44
Identities = 15/28 (53%), Positives = 24/28 (85%)
Query: 253 LTEGRASYSMQFDRFDTVSRNIQNELTT 280
+T+GRASY+MQ +FD V ++IQN+L++
Sbjct: 749 MTKGRASYTMQLAKFDVVPQHIQNQLSS 776
>UniRef100_Q7XQQ7 OSJNBa0091D06.15 protein [Oryza sativa]
Length = 749
Score = 176 bits (446), Expect = 6e-43
Identities = 88/126 (69%), Positives = 106/126 (83%), Gaps = 1/126 (0%)
Query: 2 DIVTLVGLKDTIAGETLCDPKGPFVLDQMDFPVHVIKIAIEPKTKADMEKMEAGLIKLAE 61
DIV L GLKDTI GETL DP P VL++M+FP VIK+AIEPKTKAD +KM GLIKLA+
Sbjct: 424 DIVALAGLKDTITGETLSDPDKPVVLERMEFPDPVIKVAIEPKTKADADKMATGLIKLAQ 483
Query: 62 EDPSFRVSQDKE-DRTIIQGTGELHLETIVDRLKREFKVEANVGAPQANYRESISEVTEV 120
EDPSF S+D+E ++T+I+G GELHL+ IVDRLKREF+VEANVGAPQ NYRESIS+++EV
Sbjct: 484 EDPSFHFSRDEETNQTVIEGMGELHLDIIVDRLKREFRVEANVGAPQVNYRESISKISEV 543
Query: 121 RYVFNK 126
+YV K
Sbjct: 544 QYVHKK 549
Score = 38.1 bits (87), Expect = 0.26
Identities = 16/27 (59%), Positives = 23/27 (84%)
Query: 253 LTEGRASYSMQFDRFDTVSRNIQNELT 279
+T+GRASY+MQ +FD V ++IQNEL+
Sbjct: 713 MTKGRASYTMQLAKFDVVPQHIQNELS 739
>UniRef100_UPI00002A8CC6 UPI00002A8CC6 UniRef100 entry
Length = 691
Score = 164 bits (414), Expect = 3e-39
Identities = 82/126 (65%), Positives = 98/126 (77%), Gaps = 1/126 (0%)
Query: 2 DIVTLVGLKDTIAGETLCDPKGPFVLDQMDFPVHVIKIAIEPKTKADMEKMEAGLIKLAE 61
DIV + GLKDTI GETLCD + P +L+ MDFP VI+IA+EPKTKAD EKM L +LA+
Sbjct: 373 DIVAIAGLKDTITGETLCDSQKPIILESMDFPDPVIEIAVEPKTKADQEKMGEALARLAK 432
Query: 62 EDPSFRVSQDKED-RTIIQGTGELHLETIVDRLKREFKVEANVGAPQANYRESISEVTEV 120
EDPSFRV+ D E +T+I+G GELHL+ IVDR+KREFKVEANVGAPQ YRE+IS+ EV
Sbjct: 433 EDPSFRVTSDDESGQTVIKGMGELHLDIIVDRMKREFKVEANVGAPQVAYRETISKEVEV 492
Query: 121 RYVFNK 126
Y K
Sbjct: 493 TYTHKK 498
Score = 34.7 bits (78), Expect = 2.8
Identities = 13/26 (50%), Positives = 21/26 (80%)
Query: 253 LTEGRASYSMQFDRFDTVSRNIQNEL 278
+T+GRA YSM FD ++ V +N+Q+E+
Sbjct: 661 MTQGRAQYSMFFDHYEQVPQNVQDEV 686
>UniRef100_Q6UCR8 Predicted translation elongation factor G [uncultured marine alpha
proteobacterium HOT2C01]
Length = 691
Score = 164 bits (414), Expect = 3e-39
Identities = 82/126 (65%), Positives = 98/126 (77%), Gaps = 1/126 (0%)
Query: 2 DIVTLVGLKDTIAGETLCDPKGPFVLDQMDFPVHVIKIAIEPKTKADMEKMEAGLIKLAE 61
DIV + GLKDTI GETLCD + P +L+ MDFP VI+IA+EPKTKAD EKM L +LA+
Sbjct: 373 DIVAIAGLKDTITGETLCDSQKPIILESMDFPDPVIEIAVEPKTKADQEKMGEALARLAK 432
Query: 62 EDPSFRVSQDKED-RTIIQGTGELHLETIVDRLKREFKVEANVGAPQANYRESISEVTEV 120
EDPSFRV+ D E +T+I+G GELHL+ IVDR+KREFKVEANVGAPQ YRE+IS+ EV
Sbjct: 433 EDPSFRVTSDDESGQTVIKGMGELHLDIIVDRMKREFKVEANVGAPQVAYRETISKEVEV 492
Query: 121 RYVFNK 126
Y K
Sbjct: 493 TYTHKK 498
Score = 34.7 bits (78), Expect = 2.8
Identities = 13/26 (50%), Positives = 21/26 (80%)
Query: 253 LTEGRASYSMQFDRFDTVSRNIQNEL 278
+T+GRA YSM FD ++ V +N+Q+E+
Sbjct: 661 MTQGRAQYSMFFDHYEQVPQNVQDEV 686
>UniRef100_UPI000032D345 UPI000032D345 UniRef100 entry
Length = 264
Score = 162 bits (411), Expect = 7e-39
Identities = 81/126 (64%), Positives = 98/126 (77%), Gaps = 1/126 (0%)
Query: 2 DIVTLVGLKDTIAGETLCDPKGPFVLDQMDFPVHVIKIAIEPKTKADMEKMEAGLIKLAE 61
DIV LVGLKD GETLCDP P +L++MDFP VI++A+EPKTK D EKM L +LA+
Sbjct: 57 DIVALVGLKDVTTGETLCDPNKPIILEKMDFPDPVIEVAVEPKTKVDHEKMGQALARLAQ 116
Query: 62 EDPSFRVSQDKED-RTIIQGTGELHLETIVDRLKREFKVEANVGAPQANYRESISEVTEV 120
EDPSFRV+ D+E +TII+G GELHLE +VDR+KREFKVEANVGAPQ YRE+IS+ +V
Sbjct: 117 EDPSFRVNTDEESGQTIIKGMGELHLEILVDRMKREFKVEANVGAPQVAYRETISKTYDV 176
Query: 121 RYVFNK 126
Y K
Sbjct: 177 DYTHKK 182
>UniRef100_UPI000034262E UPI000034262E UniRef100 entry
Length = 276
Score = 162 bits (409), Expect = 1e-38
Identities = 81/126 (64%), Positives = 98/126 (77%), Gaps = 1/126 (0%)
Query: 2 DIVTLVGLKDTIAGETLCDPKGPFVLDQMDFPVHVIKIAIEPKTKADMEKMEAGLIKLAE 61
DIV LVGLKD GETLCDP P +L++MDFP VI++A+EPKTK D EKM L +LA+
Sbjct: 41 DIVALVGLKDVTTGETLCDPNKPIILEKMDFPDPVIEVAVEPKTKVDHEKMGQALARLAQ 100
Query: 62 EDPSFRVSQDKED-RTIIQGTGELHLETIVDRLKREFKVEANVGAPQANYRESISEVTEV 120
EDPSFRV+ D+E +TII+G GELHLE +VDR+KREFKVEANVGAPQ YRE+IS+ +V
Sbjct: 101 EDPSFRVNTDEESGQTIIKGMGELHLEILVDRMKREFKVEANVGAPQVAYRETISKSYDV 160
Query: 121 RYVFNK 126
Y K
Sbjct: 161 DYTHKK 166
>UniRef100_UPI00003294D3 UPI00003294D3 UniRef100 entry
Length = 291
Score = 161 bits (408), Expect = 2e-38
Identities = 83/126 (65%), Positives = 97/126 (76%), Gaps = 1/126 (0%)
Query: 2 DIVTLVGLKDTIAGETLCDPKGPFVLDQMDFPVHVIKIAIEPKTKADMEKMEAGLIKLAE 61
DIV LVGLKD GETLCDP P VL++MDFP VI++A+EPKTKAD EKM L +LA+
Sbjct: 42 DIVALVGLKDVTTGETLCDPNKPIVLERMDFPDPVIEVAVEPKTKADHEKMGQALGRLAQ 101
Query: 62 EDPSFRVSQDKED-RTIIQGTGELHLETIVDRLKREFKVEANVGAPQANYRESISEVTEV 120
EDPSFRV+ D E +TII+G GELHLE +VDR+KREFKVEANVGAPQ YRE+IS +V
Sbjct: 102 EDPSFRVTTDDESGQTIIKGMGELHLEILVDRMKREFKVEANVGAPQVAYRETISSSFDV 161
Query: 121 RYVFNK 126
Y K
Sbjct: 162 DYTHKK 167
>UniRef100_UPI00002C1B69 UPI00002C1B69 UniRef100 entry
Length = 303
Score = 160 bits (406), Expect = 3e-38
Identities = 82/126 (65%), Positives = 98/126 (77%), Gaps = 1/126 (0%)
Query: 2 DIVTLVGLKDTIAGETLCDPKGPFVLDQMDFPVHVIKIAIEPKTKADMEKMEAGLIKLAE 61
DIV LVGLKD GETLCD P VL++MDFP VI++A+EPKTK D EKM L +LA+
Sbjct: 26 DIVALVGLKDVTTGETLCDINKPIVLEKMDFPDPVIEVAVEPKTKVDHEKMGTALSRLAQ 85
Query: 62 EDPSFRVSQDKED-RTIIQGTGELHLETIVDRLKREFKVEANVGAPQANYRESISEVTEV 120
EDPSFRV+ D E +TII+G GELHLE IVDR+KREFKVEANVGAPQ YRE+IS++++V
Sbjct: 86 EDPSFRVTTDHESGQTIIKGMGELHLEIIVDRMKREFKVEANVGAPQVAYRETISKLSDV 145
Query: 121 RYVFNK 126
Y K
Sbjct: 146 DYTHKK 151
>UniRef100_UPI00002AFF51 UPI00002AFF51 UniRef100 entry
Length = 270
Score = 160 bits (405), Expect = 3e-38
Identities = 82/126 (65%), Positives = 97/126 (76%), Gaps = 1/126 (0%)
Query: 2 DIVTLVGLKDTIAGETLCDPKGPFVLDQMDFPVHVIKIAIEPKTKADMEKMEAGLIKLAE 61
DIV LVGLKD GETLC P+ P VL++MDFP VI++A+EPKTKAD EKM L +LA+
Sbjct: 62 DIVALVGLKDVTTGETLCAPEKPIVLERMDFPDPVIEVAVEPKTKADHEKMGTALARLAQ 121
Query: 62 EDPSFRVSQDKED-RTIIQGTGELHLETIVDRLKREFKVEANVGAPQANYRESISEVTEV 120
EDPSFRV+ D E +TII+G GELHLE +VDR+KREFKVEANVGAPQ YRE+IS +V
Sbjct: 122 EDPSFRVTTDHESGQTIIKGMGELHLEILVDRMKREFKVEANVGAPQVAYRETISSSFDV 181
Query: 121 RYVFNK 126
Y K
Sbjct: 182 DYTHKK 187
>UniRef100_UPI00002D90C0 UPI00002D90C0 UniRef100 entry
Length = 451
Score = 159 bits (403), Expect = 6e-38
Identities = 82/126 (65%), Positives = 96/126 (76%), Gaps = 1/126 (0%)
Query: 2 DIVTLVGLKDTIAGETLCDPKGPFVLDQMDFPVHVIKIAIEPKTKADMEKMEAGLIKLAE 61
DIV LVGLKD GETLC P P VL++MDFP VI++A+EPKTKAD EKM L +LA+
Sbjct: 203 DIVALVGLKDVTTGETLCSPNEPIVLEKMDFPDPVIEVAVEPKTKADHEKMGTALGRLAQ 262
Query: 62 EDPSFRVSQDKED-RTIIQGTGELHLETIVDRLKREFKVEANVGAPQANYRESISEVTEV 120
EDPSFRV+ D E +TII+G GELHLE +VDR+KREFKVEANVGAPQ YRE+IS +V
Sbjct: 263 EDPSFRVTTDHESGQTIIKGMGELHLEILVDRMKREFKVEANVGAPQVAYRETISSSYDV 322
Query: 121 RYVFNK 126
Y K
Sbjct: 323 DYTHKK 328
>UniRef100_UPI00002C5359 UPI00002C5359 UniRef100 entry
Length = 556
Score = 159 bits (402), Expect = 8e-38
Identities = 82/126 (65%), Positives = 96/126 (76%), Gaps = 1/126 (0%)
Query: 2 DIVTLVGLKDTIAGETLCDPKGPFVLDQMDFPVHVIKIAIEPKTKADMEKMEAGLIKLAE 61
DIV LVGLKD GETLC P P VL++MDFP VI++A+EPKTKAD EKM L +LA+
Sbjct: 244 DIVALVGLKDVTTGETLCSPTEPIVLEKMDFPDPVIEVAVEPKTKADHEKMGTALGRLAQ 303
Query: 62 EDPSFRVSQDKED-RTIIQGTGELHLETIVDRLKREFKVEANVGAPQANYRESISEVTEV 120
EDPSFRV+ D E +TII+G GELHLE +VDR+KREFKVEANVGAPQ YRE+IS +V
Sbjct: 304 EDPSFRVTTDHESGQTIIKGMGELHLEILVDRMKREFKVEANVGAPQVAYRETISSSYDV 363
Query: 121 RYVFNK 126
Y K
Sbjct: 364 DYTHKK 369
>UniRef100_UPI000029848F UPI000029848F UniRef100 entry
Length = 303
Score = 159 bits (402), Expect = 8e-38
Identities = 82/126 (65%), Positives = 95/126 (75%), Gaps = 1/126 (0%)
Query: 2 DIVTLVGLKDTIAGETLCDPKGPFVLDQMDFPVHVIKIAIEPKTKADMEKMEAGLIKLAE 61
DIV + GLKDTI GETL D P VL+ MDFP VI+IA+EPKTKAD EKM L +LA+
Sbjct: 160 DIVAIAGLKDTITGETLSDSNKPIVLESMDFPDPVIEIAVEPKTKADQEKMGEALARLAK 219
Query: 62 EDPSFRVSQDKED-RTIIQGTGELHLETIVDRLKREFKVEANVGAPQANYRESISEVTEV 120
EDPSFRVS D E +T+I+G GELHL+ IVDR+KREF VEANVGAPQ YRE+IS+ EV
Sbjct: 220 EDPSFRVSSDDESGQTVIKGMGELHLDIIVDRMKREFNVEANVGAPQVAYRETISKEVEV 279
Query: 121 RYVFNK 126
Y K
Sbjct: 280 TYTHKK 285
>UniRef100_UPI00003413B4 UPI00003413B4 UniRef100 entry
Length = 692
Score = 158 bits (400), Expect = 1e-37
Identities = 81/126 (64%), Positives = 96/126 (75%), Gaps = 1/126 (0%)
Query: 2 DIVTLVGLKDTIAGETLCDPKGPFVLDQMDFPVHVIKIAIEPKTKADMEKMEAGLIKLAE 61
DIV LVGLKD GETLCD P +L++MDFP VI++A+EPKTK D EKM L +LA+
Sbjct: 374 DIVALVGLKDVTTGETLCDNNDPIILEKMDFPDPVIEVAVEPKTKVDHEKMGTALGRLAQ 433
Query: 62 EDPSFRVSQDKED-RTIIQGTGELHLETIVDRLKREFKVEANVGAPQANYRESISEVTEV 120
EDPSFRV+ D E +TII+G GELHLE IVDR+KREFKVEANVGAPQ YRE+IS ++V
Sbjct: 434 EDPSFRVTTDHESGQTIIKGMGELHLEIIVDRMKREFKVEANVGAPQVAYRETISRDSDV 493
Query: 121 RYVFNK 126
Y K
Sbjct: 494 DYTHKK 499
Score = 35.4 bits (80), Expect = 1.7
Identities = 14/26 (53%), Positives = 20/26 (76%)
Query: 253 LTEGRASYSMQFDRFDTVSRNIQNEL 278
LT+GRA+YSMQFD ++ V N+ E+
Sbjct: 662 LTQGRANYSMQFDHYEQVPSNVAEEV 687
>UniRef100_UPI00002C653B UPI00002C653B UniRef100 entry
Length = 412
Score = 158 bits (400), Expect = 1e-37
Identities = 81/126 (64%), Positives = 97/126 (76%), Gaps = 1/126 (0%)
Query: 2 DIVTLVGLKDTIAGETLCDPKGPFVLDQMDFPVHVIKIAIEPKTKADMEKMEAGLIKLAE 61
DIV L GLK TI G TLCD K P +L+ M+FP VI+IA+EPKTKAD EKM L +LA+
Sbjct: 134 DIVALAGLKYTITGHTLCDEKKPILLEPMEFPDPVIEIAVEPKTKADQEKMGEALARLAK 193
Query: 62 EDPSFRVSQDKED-RTIIQGTGELHLETIVDRLKREFKVEANVGAPQANYRESISEVTEV 120
EDPSFRV+ D+E +TII+G GELHL+ IVDR+KREFKVEAN+GAPQ YRE+I TEV
Sbjct: 194 EDPSFRVTSDEESGQTIIKGMGELHLDIIVDRMKREFKVEANIGAPQVAYRETILGSTEV 253
Query: 121 RYVFNK 126
Y+ K
Sbjct: 254 TYIHKK 259
>UniRef100_UPI000029D235 UPI000029D235 UniRef100 entry
Length = 692
Score = 158 bits (400), Expect = 1e-37
Identities = 82/126 (65%), Positives = 96/126 (76%), Gaps = 1/126 (0%)
Query: 2 DIVTLVGLKDTIAGETLCDPKGPFVLDQMDFPVHVIKIAIEPKTKADMEKMEAGLIKLAE 61
DIV LVGLKD GETLCD P VL++MDFP VI++A+EPKTKAD EKM L +LA+
Sbjct: 374 DIVALVGLKDVTTGETLCDVNKPIVLERMDFPDPVIEVAVEPKTKADHEKMGQALGRLAQ 433
Query: 62 EDPSFRVSQDKED-RTIIQGTGELHLETIVDRLKREFKVEANVGAPQANYRESISEVTEV 120
EDPSFRV+ D E +TII+G GELHLE +VDR+KREFKVEANVGAPQ YRE+IS +V
Sbjct: 434 EDPSFRVTTDHESGQTIIKGMGELHLEILVDRMKREFKVEANVGAPQVAYRETISAAFDV 493
Query: 121 RYVFNK 126
Y K
Sbjct: 494 DYTHKK 499
Score = 33.1 bits (74), Expect = 8.3
Identities = 12/26 (46%), Positives = 21/26 (80%)
Query: 253 LTEGRASYSMQFDRFDTVSRNIQNEL 278
L++GRA+Y+MQFD ++ V N+ +E+
Sbjct: 662 LSQGRANYTMQFDHYEQVPSNVADEV 687
>UniRef100_Q8RT59 Elongation factor EfG [Bartonella bacilliformis]
Length = 694
Score = 158 bits (399), Expect = 2e-37
Identities = 80/126 (63%), Positives = 96/126 (75%), Gaps = 1/126 (0%)
Query: 2 DIVTLVGLKDTIAGETLCDPKGPFVLDQMDFPVHVIKIAIEPKTKADMEKMEAGLIKLAE 61
DIV L GLK+T G+TLCDP P +L++M+FP VI+IAIEPKTKAD EKM L +LA
Sbjct: 377 DIVALAGLKETTTGDTLCDPLKPVILERMEFPEPVIEIAIEPKTKADQEKMGIALNRLAA 436
Query: 62 EDPSFRVSQDKED-RTIIQGTGELHLETIVDRLKREFKVEANVGAPQANYRESISEVTEV 120
EDPSFRV D+E +TII G GELHL+ IVDR++REFKVEANVG PQ YRESI++V E+
Sbjct: 437 EDPSFRVKSDEESGQTIIAGMGELHLDIIVDRMRREFKVEANVGQPQVAYRESITKVAEI 496
Query: 121 RYVFNK 126
Y K
Sbjct: 497 DYTHKK 502
>UniRef100_Q8G075 Elongation factor G [Brucella suis]
Length = 694
Score = 158 bits (399), Expect = 2e-37
Identities = 80/126 (63%), Positives = 95/126 (74%), Gaps = 1/126 (0%)
Query: 2 DIVTLVGLKDTIAGETLCDPKGPFVLDQMDFPVHVIKIAIEPKTKADMEKMEAGLIKLAE 61
DIV L GLK+T G+TLCDP P +L++M+FP VI+IAIEPKTKAD EKM L +LA
Sbjct: 377 DIVALAGLKETTTGDTLCDPLKPVILERMEFPDPVIEIAIEPKTKADQEKMGIALNRLAA 436
Query: 62 EDPSFRVSQDKED-RTIIQGTGELHLETIVDRLKREFKVEANVGAPQANYRESISEVTEV 120
EDPSFRV D+E +TII G GELHL+ +VDR+KREFKVEANVGAPQ YRESI+ E+
Sbjct: 437 EDPSFRVKSDEESGQTIIAGMGELHLDILVDRMKREFKVEANVGAPQVAYRESITRAAEI 496
Query: 121 RYVFNK 126
Y K
Sbjct: 497 DYTHKK 502
>UniRef100_Q8YHP3 Elongation factor G [Brucella melitensis]
Length = 694
Score = 158 bits (399), Expect = 2e-37
Identities = 80/126 (63%), Positives = 95/126 (74%), Gaps = 1/126 (0%)
Query: 2 DIVTLVGLKDTIAGETLCDPKGPFVLDQMDFPVHVIKIAIEPKTKADMEKMEAGLIKLAE 61
DIV L GLK+T G+TLCDP P +L++M+FP VI+IAIEPKTKAD EKM L +LA
Sbjct: 377 DIVALAGLKETTTGDTLCDPLKPVILERMEFPDPVIEIAIEPKTKADQEKMGIALNRLAA 436
Query: 62 EDPSFRVSQDKED-RTIIQGTGELHLETIVDRLKREFKVEANVGAPQANYRESISEVTEV 120
EDPSFRV D+E +TII G GELHL+ +VDR+KREFKVEANVGAPQ YRESI+ E+
Sbjct: 437 EDPSFRVKSDEESGQTIIAGMGELHLDILVDRMKREFKVEANVGAPQVAYRESITRAAEI 496
Query: 121 RYVFNK 126
Y K
Sbjct: 497 DYTHKK 502
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.334 0.145 0.477
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 429,193,374
Number of Sequences: 2790947
Number of extensions: 16328143
Number of successful extensions: 59364
Number of sequences better than 10.0: 2010
Number of HSP's better than 10.0 without gapping: 1317
Number of HSP's successfully gapped in prelim test: 693
Number of HSP's that attempted gapping in prelim test: 56250
Number of HSP's gapped (non-prelim): 2532
length of query: 281
length of database: 848,049,833
effective HSP length: 126
effective length of query: 155
effective length of database: 496,390,511
effective search space: 76940529205
effective search space used: 76940529205
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.6 bits)
S2: 74 (33.1 bits)
Medicago: description of AC146564.3


