

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146553.10 - phase: 0
(301 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
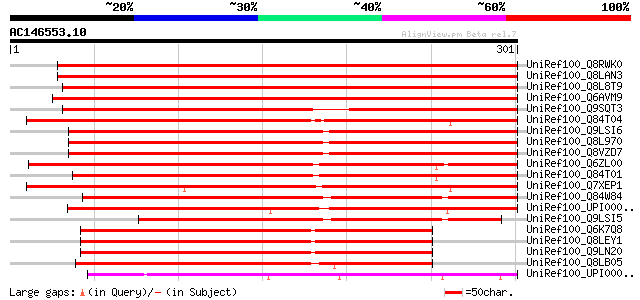
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q8RWK0 Hypothetical protein At5g18900 [Arabidopsis tha... 455 e-127
UniRef100_Q8LAN3 Prolyl 4-hydroxylase alpha subunit-like protein... 454 e-126
UniRef100_Q8L8T9 Prolyl 4-hydroxylase alpha subunit-like protein... 435 e-121
UniRef100_Q6AVM9 Putative prolyl 4-hydroxylase alpha subunit [Or... 397 e-109
UniRef100_Q9SQT3 F24P17.24 protein [Arabidopsis thaliana] 396 e-109
UniRef100_Q84T04 Putative oxidoreductase [Oryza sativa] 354 2e-96
UniRef100_Q9LSI6 Prolyl 4-hydroxylase alpha subunit-like protein... 352 9e-96
UniRef100_Q8L970 Prolyl 4-hydroxylase, putative [Arabidopsis tha... 352 9e-96
UniRef100_Q8VZD7 AT3g28480/MFJ20_16 [Arabidopsis thaliana] 350 2e-95
UniRef100_Q6ZL00 Prolyl 4-hydroxylase alpha-1 subunit-like prote... 342 9e-93
UniRef100_Q84T01 Putative oxidoreductase [Oryza sativa] 335 1e-90
UniRef100_Q7XEP1 Hypothetical protein [Oryza sativa] 333 3e-90
UniRef100_Q84W84 Putative prolyl 4-hydroxylase [Arabidopsis thal... 331 2e-89
UniRef100_UPI00002D6679 UPI00002D6679 UniRef100 entry 272 7e-72
UniRef100_Q9LSI5 Gb|AAF08583.1 [Arabidopsis thaliana] 268 2e-70
UniRef100_Q6K7Q8 Putative prolyl 4-hydroxylase [Oryza sativa] 242 8e-63
UniRef100_Q8LEY1 Putative prolyl 4-hydroxylase, alpha subunit [A... 241 1e-62
UniRef100_Q9LN20 F14O10.12 protein [Arabidopsis thaliana] 240 3e-62
UniRef100_Q8LB05 Putative prolyl 4-hydroxylase, alpha subunit [A... 236 6e-61
UniRef100_UPI00002F63AB UPI00002F63AB UniRef100 entry 228 1e-58
>UniRef100_Q8RWK0 Hypothetical protein At5g18900 [Arabidopsis thaliana]
Length = 298
Score = 455 bits (1170), Expect = e-127
Identities = 212/273 (77%), Positives = 241/273 (87%)
Query: 29 AGTSAIIDPTKVKQVSWKPRAFVYKGFLTDLECDHLISIAKSELKRSAVADNLSGESKLS 88
+ +S ++P+KVKQVS KPRAFVY+GFLT+LECDH++S+AK+ LKRSAVADN SGESK S
Sbjct: 26 SSSSVFVNPSKVKQVSSKPRAFVYEGFLTELECDHMVSLAKASLKRSAVADNDSGESKFS 85
Query: 89 EVRTSSGMFISKNKDAIVSGIEDKISSWTFLPKENGEDIQVLRYEHGQKYDPHYDYFADK 148
EVRTSSG FISK KD IVSGIEDKIS+WTFLPKENGEDIQVLRYEHGQKYD H+DYF DK
Sbjct: 86 EVRTSSGTFISKGKDPIVSGIEDKISTWTFLPKENGEDIQVLRYEHGQKYDAHFDYFHDK 145
Query: 149 VNIARGGHRVATVLMYLTNVTKGGETVFPNAEESPRHKLSETDEDLSECGKKGVAVNPRR 208
VNI RGGHR+AT+LMYL+NVTKGGETVFP+AE R LSE +EDLS+C K+G+AV PR+
Sbjct: 146 VNIVRGGHRMATILMYLSNVTKGGETVFPDAEIPSRRVLSENEEDLSDCAKRGIAVKPRK 205
Query: 209 GDALLFCSLHPNAIPDTLSLHAGCPVIEGEKWSATKWIHVDSFDKTVGAGGDCTDQHESC 268
GDALLF +LHP+AIPD LSLH GCPVIEGEKWSATKWIHVDSFD+ V G+CTD +ESC
Sbjct: 206 GDALLFFNLHPDAIPDPLSLHGGCPVIEGEKWSATKWIHVDSFDRIVTPSGNCTDMNESC 265
Query: 269 ERWAALGECTKNPEYMVGTSGLPGYCRKSCKTC 301
ERWA LGECTKNPEYMVGT+ LPGYCR+SCK C
Sbjct: 266 ERWAVLGECTKNPEYMVGTTELPGYCRRSCKAC 298
>UniRef100_Q8LAN3 Prolyl 4-hydroxylase alpha subunit-like protein [Arabidopsis
thaliana]
Length = 298
Score = 454 bits (1167), Expect = e-126
Identities = 212/273 (77%), Positives = 240/273 (87%)
Query: 29 AGTSAIIDPTKVKQVSWKPRAFVYKGFLTDLECDHLISIAKSELKRSAVADNLSGESKLS 88
+ +S ++P+KVKQVS KPRAFVY+GFLT+LECDH++S+AK+ LKRSAVADN SGESK S
Sbjct: 26 SSSSVFVNPSKVKQVSSKPRAFVYEGFLTELECDHMVSLAKASLKRSAVADNDSGESKFS 85
Query: 89 EVRTSSGMFISKNKDAIVSGIEDKISSWTFLPKENGEDIQVLRYEHGQKYDPHYDYFADK 148
EVRTSSG FISK KD IVSGIEDKIS+WTFLPKENGEDIQVLRYEHGQKYD H+DYF DK
Sbjct: 86 EVRTSSGTFISKGKDPIVSGIEDKISTWTFLPKENGEDIQVLRYEHGQKYDAHFDYFHDK 145
Query: 149 VNIARGGHRVATVLMYLTNVTKGGETVFPNAEESPRHKLSETDEDLSECGKKGVAVNPRR 208
VNI RGGHR+AT+LMYL+NVTKGGETVFP+AE R LSE EDLS+C K+G+AV PR+
Sbjct: 146 VNIVRGGHRMATILMYLSNVTKGGETVFPDAEIPSRRVLSENKEDLSDCAKRGIAVKPRK 205
Query: 209 GDALLFCSLHPNAIPDTLSLHAGCPVIEGEKWSATKWIHVDSFDKTVGAGGDCTDQHESC 268
GDALLF +LHP+AIPD LSLH GCPVIEGEKWSATKWIHVDSFD+ V G+CTD +ESC
Sbjct: 206 GDALLFFNLHPDAIPDPLSLHGGCPVIEGEKWSATKWIHVDSFDRIVTPSGNCTDMNESC 265
Query: 269 ERWAALGECTKNPEYMVGTSGLPGYCRKSCKTC 301
ERWA LGECTKNPEYMVGT+ LPGYCR+SCK C
Sbjct: 266 ERWAVLGECTKNPEYMVGTTELPGYCRRSCKAC 298
>UniRef100_Q8L8T9 Prolyl 4-hydroxylase alpha subunit-like protein [Arabidopsis
thaliana]
Length = 297
Score = 435 bits (1118), Expect = e-121
Identities = 204/270 (75%), Positives = 236/270 (86%)
Query: 32 SAIIDPTKVKQVSWKPRAFVYKGFLTDLECDHLISIAKSELKRSAVADNLSGESKLSEVR 91
S+II+P+KVKQVS KPRAFVY+GFLTDLECDHLIS+AK L+RSAVADN +GES++S+VR
Sbjct: 28 SSIINPSKVKQVSSKPRAFVYEGFLTDLECDHLISLAKENLQRSAVADNDNGESQVSDVR 87
Query: 92 TSSGMFISKNKDAIVSGIEDKISSWTFLPKENGEDIQVLRYEHGQKYDPHYDYFADKVNI 151
TSSG FISK KD IVSGIEDK+S+WTFLPKENGED+QVLRYEHGQKYD H+DYF DKVNI
Sbjct: 88 TSSGTFISKGKDPIVSGIEDKLSTWTFLPKENGEDLQVLRYEHGQKYDAHFDYFHDKVNI 147
Query: 152 ARGGHRVATVLMYLTNVTKGGETVFPNAEESPRHKLSETDEDLSECGKKGVAVNPRRGDA 211
ARGGHR+ATVL+YL+NVTKGGETVFP+A+E R LSE +DLS+C KKG+AV P++G+A
Sbjct: 148 ARGGHRIATVLLYLSNVTKGGETVFPDAQEFSRRSLSENKDDLSDCAKKGIAVKPKKGNA 207
Query: 212 LLFCSLHPNAIPDTLSLHAGCPVIEGEKWSATKWIHVDSFDKTVGAGGDCTDQHESCERW 271
LLF +L +AIPD SLH GCPVIEGEKWSATKWIHVDSFDK + G+CTD +ESCERW
Sbjct: 208 LLFFNLQQDAIPDPFSLHGGCPVIEGEKWSATKWIHVDSFDKILTHDGNCTDVNESCERW 267
Query: 272 AALGECTKNPEYMVGTSGLPGYCRKSCKTC 301
A LGEC KNPEYMVGT +PG CR+SCK C
Sbjct: 268 AVLGECGKNPEYMVGTPEIPGNCRRSCKAC 297
>UniRef100_Q6AVM9 Putative prolyl 4-hydroxylase alpha subunit [Oryza sativa]
Length = 319
Score = 397 bits (1021), Expect = e-109
Identities = 185/276 (67%), Positives = 225/276 (81%)
Query: 26 GSYAGTSAIIDPTKVKQVSWKPRAFVYKGFLTDLECDHLISIAKSELKRSAVADNLSGES 85
GS + ++ P +Q+SWKPR F+Y+ FL+D E +HL+S+A++ELKRSAVADNLSG+S
Sbjct: 44 GSSVYPAPVVYPHHSRQISWKPRVFLYQHFLSDDEANHLVSLARTELKRSAVADNLSGKS 103
Query: 86 KLSEVRTSSGMFISKNKDAIVSGIEDKISSWTFLPKENGEDIQVLRYEHGQKYDPHYDYF 145
+LS+ RTSSG FI K++D IV+GIE+KI++WTFLPKENGEDIQVLRY+HG+KY+ HYDYF
Sbjct: 104 ELSDARTSSGTFIRKSQDPIVAGIEEKIAAWTFLPKENGEDIQVLRYKHGEKYERHYDYF 163
Query: 146 ADKVNIARGGHRVATVLMYLTNVTKGGETVFPNAEESPRHKLSETDEDLSECGKKGVAVN 205
+D VN RGGHR+ATVLMYLT+V +GGETVFP AEE + D LSEC KKGVAV
Sbjct: 164 SDNVNTLRGGHRIATVLMYLTDVAEGGETVFPLAEEFTESGTNNEDSTLSECAKKGVAVK 223
Query: 206 PRRGDALLFCSLHPNAIPDTLSLHAGCPVIEGEKWSATKWIHVDSFDKTVGAGGDCTDQH 265
PR+GDALLF +L P+A D+LSLHAGCPVI+GEKWSATKWI V SFDK G+CTD +
Sbjct: 224 PRKGDALLFFNLSPDASKDSLSLHAGCPVIKGEKWSATKWIRVASFDKVYHTQGNCTDDN 283
Query: 266 ESCERWAALGECTKNPEYMVGTSGLPGYCRKSCKTC 301
ESCE+WAALGEC KNPEYM+GT+ LPGYCRKSC C
Sbjct: 284 ESCEKWAALGECIKNPEYMIGTAALPGYCRKSCNIC 319
>UniRef100_Q9SQT3 F24P17.24 protein [Arabidopsis thaliana]
Length = 278
Score = 396 bits (1018), Expect = e-109
Identities = 191/270 (70%), Positives = 221/270 (81%), Gaps = 21/270 (7%)
Query: 32 SAIIDPTKVKQVSWKPRAFVYKGFLTDLECDHLISIAKSELKRSAVADNLSGESKLSEVR 91
S+II+P+KVKQVS KPRAFVY+GFLTDLECDHLIS+AK L+RSAVADN +GES++S+VR
Sbjct: 30 SSIINPSKVKQVSSKPRAFVYEGFLTDLECDHLISLAKENLQRSAVADNDNGESQVSDVR 89
Query: 92 TSSGMFISKNKDAIVSGIEDKISSWTFLPKENGEDIQVLRYEHGQKYDPHYDYFADKVNI 151
TSSG FISK KD IVSGIEDK+S+WTFLPKENGED+QVLRYEHGQKYD H+DYF DKVNI
Sbjct: 90 TSSGTFISKGKDPIVSGIEDKLSTWTFLPKENGEDLQVLRYEHGQKYDAHFDYFHDKVNI 149
Query: 152 ARGGHRVATVLMYLTNVTKGGETVFPNAEESPRHKLSETDEDLSECGKKGVAVNPRRGDA 211
ARGGHR+ATVL+YL+NVTKGGETVFP+A+ V + P++G+A
Sbjct: 150 ARGGHRIATVLLYLSNVTKGGETVFPDAQ---------------------VCLKPKKGNA 188
Query: 212 LLFCSLHPNAIPDTLSLHAGCPVIEGEKWSATKWIHVDSFDKTVGAGGDCTDQHESCERW 271
LLF +L +AIPD SLH GCPVIEGEKWSATKWIHVDSFDK + G+CTD +ESCERW
Sbjct: 189 LLFFNLQQDAIPDPFSLHGGCPVIEGEKWSATKWIHVDSFDKILTHDGNCTDVNESCERW 248
Query: 272 AALGECTKNPEYMVGTSGLPGYCRKSCKTC 301
A LGEC KNPEYMVGT +PG CR+SCK C
Sbjct: 249 AVLGECGKNPEYMVGTPEIPGNCRRSCKAC 278
>UniRef100_Q84T04 Putative oxidoreductase [Oryza sativa]
Length = 299
Score = 354 bits (908), Expect = 2e-96
Identities = 173/293 (59%), Positives = 215/293 (73%), Gaps = 5/293 (1%)
Query: 11 IIVPLLLICKIHFALGSYAGTSAIIDPTKVKQVSWKPRAFVYKGFLTDLECDHLISIAKS 70
+I LLL+ + G DP +V Q+SW+PRAF+Y GFL+ ECDHL+++AK
Sbjct: 8 VIFLLLLLALVPALSRPDGGGGGFYDPARVTQLSWRPRAFLYSGFLSHDECDHLVNLAKG 67
Query: 71 ELKRSAVADNLSGESKLSEVRTSSGMFISKNKDAIVSGIEDKISSWTFLPKENGEDIQVL 130
+++S VADN SG+S +S+VRTSSG F+SK++D IVSGIE ++++WTFLP+EN E IQ+L
Sbjct: 68 RMEKSMVADNDSGKSIMSQVRTSSGTFLSKHEDDIVSGIEKRVAAWTFLPEENAESIQIL 127
Query: 131 RYEHGQKYDPHYDYFADKVNIARGGHRVATVLMYLTNVTKGGETVFPNAEESPRHKLSET 190
YE GQKYD H+DYF DK N+ RGGHRVATVLMYLT+V KGGETVFPNA + RH L
Sbjct: 128 HYELGQKYDAHFDYFHDKNNLKRGGHRVATVLMYLTDVKKGGETVFPNA--AGRH-LQLK 184
Query: 191 DEDLSECGKKGVAVNPRRGDALLFCSLHPNAIPDTLSLHAGCPVIEGEKWSATKWIHVDS 250
DE S+C + G+AV P++GDALLF SLH NA D SLH CPVIEGEKWSATKWIHV S
Sbjct: 185 DETWSDCARSGLAVKPKKGDALLFFSLHVNATTDPASLHGSCPVIEGEKWSATKWIHVRS 244
Query: 251 FDKTVGAGGD--CTDQHESCERWAALGECTKNPEYMVGTSGLPGYCRKSCKTC 301
FD D C+D++E C RWAA+GEC +NP+YMVGT G+CRKSC C
Sbjct: 245 FDNPPDVSLDLPCSDENERCTRWAAVGECYRNPKYMVGTKDSLGFCRKSCGVC 297
>UniRef100_Q9LSI6 Prolyl 4-hydroxylase alpha subunit-like protein [Arabidopsis
thaliana]
Length = 332
Score = 352 bits (902), Expect = 9e-96
Identities = 162/266 (60%), Positives = 202/266 (75%), Gaps = 3/266 (1%)
Query: 36 DPTKVKQVSWKPRAFVYKGFLTDLECDHLISIAKSELKRSAVADNLSGESKLSEVRTSSG 95
DPT+V Q+SW PR F+Y+GFL+D ECDH I +AK +L++S VADN SGES SEVRTSSG
Sbjct: 68 DPTRVTQLSWTPRVFLYEGFLSDEECDHFIKLAKGKLEKSMVADNDSGESVESEVRTSSG 127
Query: 96 MFISKNKDAIVSGIEDKISSWTFLPKENGEDIQVLRYEHGQKYDPHYDYFADKVNIARGG 155
MF+SK +D IVS +E K+++WTFLP+ENGE +Q+L YE+GQKY+PH+DYF D+ N+ GG
Sbjct: 128 MFLSKRQDDIVSNVEAKLAAWTFLPEENGESMQILHYENGQKYEPHFDYFHDQANLELGG 187
Query: 156 HRVATVLMYLTNVTKGGETVFPNAEESPRHKLSETDEDLSECGKKGVAVNPRRGDALLFC 215
HR+ATVLMYL+NV KGGETVFP + D+ +EC K+G AV PR+GDALLF
Sbjct: 188 HRIATVLMYLSNVEKGGETVFPMWKGKATQL---KDDSWTECAKQGYAVKPRKGDALLFF 244
Query: 216 SLHPNAIPDTLSLHAGCPVIEGEKWSATKWIHVDSFDKTVGAGGDCTDQHESCERWAALG 275
+LHPNA D+ SLH CPV+EGEKWSAT+WIHV SF++ C D++ SCE+WA G
Sbjct: 245 NLHPNATTDSNSLHGSCPVVEGEKWSATRWIHVKSFERAFNKQSGCMDENVSCEKWAKAG 304
Query: 276 ECTKNPEYMVGTSGLPGYCRKSCKTC 301
EC KNP YMVG+ GYCRKSCK C
Sbjct: 305 ECQKNPTYMVGSDKDHGYCRKSCKAC 330
>UniRef100_Q8L970 Prolyl 4-hydroxylase, putative [Arabidopsis thaliana]
Length = 316
Score = 352 bits (902), Expect = 9e-96
Identities = 162/266 (60%), Positives = 202/266 (75%), Gaps = 3/266 (1%)
Query: 36 DPTKVKQVSWKPRAFVYKGFLTDLECDHLISIAKSELKRSAVADNLSGESKLSEVRTSSG 95
DPT+V Q+SW PR F+Y+GFL+D ECDH I +AK +L++S VADN SGES SEVRTSSG
Sbjct: 52 DPTRVTQLSWTPRVFLYEGFLSDEECDHFIKLAKGKLEKSMVADNDSGESVESEVRTSSG 111
Query: 96 MFISKNKDAIVSGIEDKISSWTFLPKENGEDIQVLRYEHGQKYDPHYDYFADKVNIARGG 155
MF+SK +D IVS +E K+++WTFLP+ENGE +Q+L YE+GQKY+PH+DYF D+ N+ GG
Sbjct: 112 MFLSKRQDDIVSNVEAKLAAWTFLPEENGESMQILHYENGQKYEPHFDYFHDQANLELGG 171
Query: 156 HRVATVLMYLTNVTKGGETVFPNAEESPRHKLSETDEDLSECGKKGVAVNPRRGDALLFC 215
HR+ATVLMYL+NV KGGETVFP + D+ +EC K+G AV PR+GDALLF
Sbjct: 172 HRIATVLMYLSNVEKGGETVFPMWKGKATQL---KDDSWTECAKQGYAVKPRKGDALLFF 228
Query: 216 SLHPNAIPDTLSLHAGCPVIEGEKWSATKWIHVDSFDKTVGAGGDCTDQHESCERWAALG 275
+LHPNA D+ SLH CPV+EGEKWSAT+WIHV SF++ C D++ SCE+WA G
Sbjct: 229 NLHPNATTDSNSLHGSCPVVEGEKWSATRWIHVKSFERAFNKQSGCMDENVSCEKWAKAG 288
Query: 276 ECTKNPEYMVGTSGLPGYCRKSCKTC 301
EC KNP YMVG+ GYCRKSCK C
Sbjct: 289 ECQKNPTYMVGSDKDHGYCRKSCKAC 314
>UniRef100_Q8VZD7 AT3g28480/MFJ20_16 [Arabidopsis thaliana]
Length = 316
Score = 350 bits (899), Expect = 2e-95
Identities = 161/266 (60%), Positives = 202/266 (75%), Gaps = 3/266 (1%)
Query: 36 DPTKVKQVSWKPRAFVYKGFLTDLECDHLISIAKSELKRSAVADNLSGESKLSEVRTSSG 95
DPT+V Q+SW PR F+Y+GFL+D ECDH I +AK +L++S VADN SGES SEVRTSSG
Sbjct: 52 DPTRVTQLSWTPRVFLYEGFLSDEECDHFIKLAKGKLEKSMVADNDSGESVESEVRTSSG 111
Query: 96 MFISKNKDAIVSGIEDKISSWTFLPKENGEDIQVLRYEHGQKYDPHYDYFADKVNIARGG 155
MF+SK +D IV+ +E K+++WTFLP+ENGE +Q+L YE+GQKY+PH+DYF D+ N+ GG
Sbjct: 112 MFLSKRQDDIVNNVEAKLAAWTFLPEENGESMQILHYENGQKYEPHFDYFHDQANLELGG 171
Query: 156 HRVATVLMYLTNVTKGGETVFPNAEESPRHKLSETDEDLSECGKKGVAVNPRRGDALLFC 215
HR+ATVLMYL+NV KGGETVFP + D+ +EC K+G AV PR+GDALLF
Sbjct: 172 HRIATVLMYLSNVEKGGETVFPMWKGKATQL---KDDSWTECAKQGYAVKPRKGDALLFF 228
Query: 216 SLHPNAIPDTLSLHAGCPVIEGEKWSATKWIHVDSFDKTVGAGGDCTDQHESCERWAALG 275
+LHPNA D+ SLH CPV+EGEKWSAT+WIHV SF++ C D++ SCE+WA G
Sbjct: 229 NLHPNATTDSNSLHGSCPVVEGEKWSATRWIHVKSFERAFNKQSGCMDENVSCEKWAKAG 288
Query: 276 ECTKNPEYMVGTSGLPGYCRKSCKTC 301
EC KNP YMVG+ GYCRKSCK C
Sbjct: 289 ECQKNPTYMVGSDKDHGYCRKSCKAC 314
>UniRef100_Q6ZL00 Prolyl 4-hydroxylase alpha-1 subunit-like protein [Oryza sativa]
Length = 313
Score = 342 bits (876), Expect = 9e-93
Identities = 167/294 (56%), Positives = 209/294 (70%), Gaps = 9/294 (3%)
Query: 12 IVPLLLICKIHFALGSYAGTSAIIDPTKVKQVSWKPRAFVYKGFLTDLECDHLISIAKSE 71
+VPLLL+ + + + ++V+ VSW+PR FVYKGFL+D ECDHL+ + K +
Sbjct: 23 VVPLLLLGEAGDDGVGAVAAAPPFNASRVRAVSWRPRVFVYKGFLSDDECDHLVKLGKRK 82
Query: 72 LKRSAVADNLSGESKLSEVRTSSGMFISKNKDAIVSGIEDKISSWTFLPKENGEDIQVLR 131
++RS VADN SG+S +SEVRTSSGMF+ K +D +VS IE +I++WTFLP+EN E+IQ+LR
Sbjct: 83 MQRSMVADNKSGKSVMSEVRTSSGMFLDKRQDPVVSRIEKRIAAWTFLPEENAENIQILR 142
Query: 132 YEHGQKYDPHYDYFADKVNIARGGHRVATVLMYLTNVTKGGETVFPNAEESPRHKLSETD 191
YEHGQKY+PH+DYF DKVN A GGHR ATVLMYL+ V KGGETVFPNAE + D
Sbjct: 143 YEHGQKYEPHFDYFHDKVNQALGGHRYATVLMYLSTVEKGGETVFPNAE---GWENQPKD 199
Query: 192 EDLSECGKKGVAVNPRRGDALLFCSLHPNAIPDTLSLHAGCPVIEGEKWSATKWIHVDSF 251
+ SEC +KG+AV P +GD +LF SLH + +PD LSLH CPVIEGEKWSA KWI + S+
Sbjct: 200 DTFSECAQKGLAVKPVKGDTVLFFSLHIDGVPDPLSLHGSCPVIEGEKWSAPKWIRIRSY 259
Query: 252 D----KTVGAGGDCTDQHESCERWAALGECTKNPEYMVGTSGLPGYCRKSCKTC 301
+ V G C+D C +WA GEC KNP YMVG GLPG CRKSC C
Sbjct: 260 EHPPVSKVTEG--CSDNSARCAKWAEAGECEKNPVYMVGAEGLPGNCRKSCGVC 311
>UniRef100_Q84T01 Putative oxidoreductase [Oryza sativa]
Length = 310
Score = 335 bits (858), Expect = 1e-90
Identities = 157/266 (59%), Positives = 196/266 (73%), Gaps = 5/266 (1%)
Query: 38 TKVKQVSWKPRAFVYKGFLTDLECDHLISIAKSELKRSAVADNLSGESKLSEVRTSSGMF 97
+ V +SWKPR F YKGFL+D ECDHL+ + K +LKRS VADN SG+S +SEVRTSSGMF
Sbjct: 46 SSVTIISWKPRIFFYKGFLSDDECDHLVKLGKEKLKRSMVADNESGKSVMSEVRTSSGMF 105
Query: 98 ISKNKDAIVSGIEDKISSWTFLPKENGEDIQVLRYEHGQKYDPHYDYFADKVNIARGGHR 157
+ K +D +VSGIE++I++WT LP+EN E+IQ+LRYE+GQKYDPH+DYF DKVN +GGHR
Sbjct: 106 LDKQQDPVVSGIEERIAAWTLLPQENAENIQILRYENGQKYDPHFDYFQDKVNQLQGGHR 165
Query: 158 VATVLMYLTNVTKGGETVFPNAEESPRHKLSETDEDLSECGKKGVAVNPRRGDALLFCSL 217
ATVL YL+ V KGGETVFPNAE + D+ S+C KKG+AV +GD++LF +L
Sbjct: 166 YATVLTYLSTVEKGGETVFPNAE---GWESQPKDDSFSDCAKKGLAVKAVKGDSVLFFNL 222
Query: 218 HPNAIPDTLSLHAGCPVIEGEKWSATKWIHVDSFD--KTVGAGGDCTDQHESCERWAALG 275
P+ PD LSLH CPVIEGEKWSA KWIHV S+D ++ +C+D E+C WAA G
Sbjct: 223 QPDGTPDPLSLHGSCPVIEGEKWSAPKWIHVRSYDNASSMKQSEECSDLSENCAAWAASG 282
Query: 276 ECTKNPEYMVGTSGLPGYCRKSCKTC 301
EC N YM+GT PG C+KSC C
Sbjct: 283 ECNNNAVYMIGTEDAPGQCQKSCNAC 308
>UniRef100_Q7XEP1 Hypothetical protein [Oryza sativa]
Length = 369
Score = 333 bits (855), Expect = 3e-90
Identities = 168/342 (49%), Positives = 218/342 (63%), Gaps = 54/342 (15%)
Query: 11 IIVPLLLICKIHFALGSYAGTSAIIDPTKVKQVSWKPRAFVYKGFLTDLECDHLISIAKS 70
+ V LL++ + + + DP++V Q+SW+PRAF++KGFLTD EC+HLIS+AK
Sbjct: 16 LAVALLVLAGVASSAAAAGSGRGAFDPSRVVQLSWRPRAFLHKGFLTDAECEHLISLAKD 75
Query: 71 ELKRSAVADNLSGESKLSEVRTSSGMFISKNK---------------------------- 102
+L++S VADN SG+S +SEVRTSSGMF+ K +
Sbjct: 76 KLEKSMVADNESGKSVMSEVRTSSGMFLEKKQVVFFCSLREMKFDQLFPIYISIVCKALN 135
Query: 103 --------------------DAIVSGIEDKISSWTFLPKENGEDIQVLRYEHGQKYDPHY 142
D +V+ IE++I++WTFLP +NGE IQ+L Y++G+KY+PHY
Sbjct: 136 HVVIFAIWTSILSNRPVLKLDEVVARIEERIAAWTFLPPDNGESIQILHYQNGEKYEPHY 195
Query: 143 DYFADKVNIARGGHRVATVLMYLTNVTKGGETVFPNAEESPRHKLSETDEDLSECGKKGV 202
DYF DK N A GGHR+ATVLMYL++V KGGET+FP AE L D+ S+C K G
Sbjct: 196 DYFHDKNNQALGGHRIATVLMYLSDVGKGGETIFPEAEGK---LLQPKDDTWSDCAKNGY 252
Query: 203 AVNPRRGDALLFCSLHPNAIPDTLSLHAGCPVIEGEKWSATKWIHVDSFDKTVGAGGD-- 260
AV P +GDALLF SLHP+A D+ SLH CPVIEG+KWSATKWIHV SFD +V G
Sbjct: 253 AVKPVKGDALLFFSLHPDATTDSDSLHGSCPVIEGQKWSATKWIHVRSFDISVKQGASTD 312
Query: 261 -CTDQHESCERWAALGECTKNPEYMVGTSGLPGYCRKSCKTC 301
C D++ C +WAA+GEC KNP YMVGT+ PG+CRKSC C
Sbjct: 313 GCEDENVLCPQWAAVGECAKNPNYMVGTNEAPGFCRKSCNVC 354
>UniRef100_Q84W84 Putative prolyl 4-hydroxylase [Arabidopsis thaliana]
Length = 253
Score = 331 bits (848), Expect = 2e-89
Identities = 162/260 (62%), Positives = 198/260 (75%), Gaps = 9/260 (3%)
Query: 44 SWKPRAFVYKGFLTDLECDHLISIAKSELKRS-AVADNLSGESKLSEVRTSSGMFISKNK 102
SW PRAF+YKGFL+D ECDHLI +AK +L++S VAD SGES+ SEVRTSSGMF++K +
Sbjct: 1 SWTPRAFLYKGFLSDEECDHLIKLAKGKLEKSMVVADVDSGESEDSEVRTSSGMFLTKRQ 60
Query: 103 DAIVSGIEDKISSWTFLPKENGEDIQVLRYEHGQKYDPHYDYFADKVNIARGGHRVATVL 162
D IV+ +E K+++WTFLP+ENGE +Q+L YE+GQKYDPH+DYF DK + GGHR+ATVL
Sbjct: 61 DDIVANVEAKLAAWTFLPEENGEALQILHYENGQKYDPHFDYFYDKKALELGGHRIATVL 120
Query: 163 MYLTNVTKGGETVFPNAE-ESPRHKLSETDEDLSECGKKGVAVNPRRGDALLFCSLHPNA 221
MYL+NVTKGGETVFPN + ++P+ K D+ S+C K+G AV PR+GDALLF +LH N
Sbjct: 121 MYLSNVTKGGETVFPNWKGKTPQLK----DDSWSKCAKQGYAVKPRKGDALLFFNLHLNG 176
Query: 222 IPDTLSLHAGCPVIEGEKWSATKWIHVDSFDKTVGAGGDCTDQHESCERWAALGECTKNP 281
D SLH CPVIEGEKWSAT+WIHV SF K C D HESC+ WA GEC KNP
Sbjct: 177 TTDPNSLHGSCPVIEGEKWSATRWIHVRSFGKKKLV---CVDDHESCQEWADAGECEKNP 233
Query: 282 EYMVGTSGLPGYCRKSCKTC 301
YMVG+ G+CRKSCK C
Sbjct: 234 MYMVGSETSLGFCRKSCKAC 253
>UniRef100_UPI00002D6679 UPI00002D6679 UniRef100 entry
Length = 345
Score = 272 bits (696), Expect = 7e-72
Identities = 140/273 (51%), Positives = 179/273 (65%), Gaps = 11/273 (4%)
Query: 35 IDPTKVKQVSWKPRAFVYKGFLTDLECDHLISIAKSELKRSAVADNLSGESKLSEVRTSS 94
IDP + ++SWKP A VY+GFLT+ EC+HLI +AK L++S V D +G S SE RTSS
Sbjct: 71 IDPRAIDRISWKPHAEVYRGFLTEEECEHLIELAKPALRKSTVVDVDTGGSVGSEERTSS 130
Query: 95 GMFISKNKDAIVSGIEDKISSWTFLPKENGEDIQVLRYEHGQKYDPHYDYFADKVNIAR- 153
GMF+ + +D +V IE +I++WT +P+ +GE QVLRYE GQ+Y PH+DYF D V+ R
Sbjct: 131 GMFLMRGEDEVVETIERRIAAWTHVPESHGEGFQVLRYERGQEYRPHFDYFHDSVHTERD 190
Query: 154 -GGHRVATVLMYLTNVTKGGETVFPNAEESPRHKLSETDEDLSECGKKGVAVNPRRGDAL 212
GG RV TVLMYLT+V KGGET FP+A+E R D S C + VAV P+ GDAL
Sbjct: 191 KGGQRVGTVLMYLTDVEKGGETTFPSAQEGER-----APSDASACARGKVAVAPKAGDAL 245
Query: 213 LFCSLHPNAIPDTLSLHAGCPVIEGEKWSATKWIHVDSF-DKTVGAG---GDCTDQHESC 268
F SL+ N D LS HAGCPV G K+SATKW+HV + D + GA G C D +++C
Sbjct: 246 YFRSLNHNLSLDVLSAHAGCPVELGVKYSATKWMHVYTIEDGSAGAAMPPGVCKDVNKNC 305
Query: 269 ERWAALGECTKNPEYMVGTSGLPGYCRKSCKTC 301
E WA GEC NP +M+G G C +SC C
Sbjct: 306 EGWAKAGECENNPSFMIGRGRAKGNCMRSCGAC 338
>UniRef100_Q9LSI5 Gb|AAF08583.1 [Arabidopsis thaliana]
Length = 328
Score = 268 bits (684), Expect = 2e-70
Identities = 133/217 (61%), Positives = 161/217 (73%), Gaps = 8/217 (3%)
Query: 77 VADNLSGESKLSEVRTSSGMFISKNKDAIVSGIEDKISSWTFLPKENGEDIQVLRYEHGQ 136
VAD SGES+ SEVRTSSGMF++K +D IV+ +E K+++WTFLP+ENGE +Q+L YE+GQ
Sbjct: 3 VADVDSGESEDSEVRTSSGMFLTKRQDDIVANVEAKLAAWTFLPEENGEALQILHYENGQ 62
Query: 137 KYDPHYDYFADKVNIARGGHRVATVLMYLTNVTKGGETVFPNAE-ESPRHKLSETDEDLS 195
KYDPH+DYF DK + GGHR+ATVLMYL+NVTKGGETVFPN + ++P+ K D+ S
Sbjct: 63 KYDPHFDYFYDKKALELGGHRIATVLMYLSNVTKGGETVFPNWKGKTPQLK----DDSWS 118
Query: 196 ECGKKGVAVNPRRGDALLFCSLHPNAIPDTLSLHAGCPVIEGEKWSATKWIHVDSFDKTV 255
+C K+G AV PR+GDALLF +LH N D SLH CPVIEGEKWSAT+WIHV SF K
Sbjct: 119 KCAKQGYAVKPRKGDALLFFNLHLNGTTDPNSLHGSCPVIEGEKWSATRWIHVRSFGKKK 178
Query: 256 GAGGDCTDQHESCERWAALGECTKNPEYMVGTSGLPG 292
C D HESC+ WA GEC KNP YMVG G
Sbjct: 179 LV---CVDDHESCQEWADAGECEKNPMYMVGVGKKTG 212
>UniRef100_Q6K7Q8 Putative prolyl 4-hydroxylase [Oryza sativa]
Length = 310
Score = 242 bits (618), Expect = 8e-63
Identities = 112/209 (53%), Positives = 154/209 (73%), Gaps = 2/209 (0%)
Query: 43 VSWKPRAFVYKGFLTDLECDHLISIAKSELKRSAVADNLSGESKLSEVRTSSGMFISKNK 102
+SW+PRAFVY FL+ ECD+LI +AK + +S V D+ +G+SK S VRTSSGMF+ + +
Sbjct: 102 ISWEPRAFVYHNFLSKEECDYLIGLAKPHMVKSTVVDSTTGKSKDSRVRTSSGMFLQRGR 161
Query: 103 DAIVSGIEDKISSWTFLPKENGEDIQVLRYEHGQKYDPHYDYFADKVNIARGGHRVATVL 162
D ++ IE +I+ +TF+P E+GE +QVL YE GQKY+PH+DYF D+ N GG R+AT+L
Sbjct: 162 DKVIRAIEKRIADYTFIPMEHGEGLQVLHYEVGQKYEPHFDYFLDEYNTKNGGQRMATLL 221
Query: 163 MYLTNVTKGGETVFPNAEESPRHKLSETDEDLSECGKKGVAVNPRRGDALLFCSLHPNAI 222
MYL++V +GGET+FP+A + +LSEC +KG+AV P+ GDALLF S+ P+A
Sbjct: 222 MYLSDVEEGGETIFPDA--NVNSSSLPWYNELSECARKGLAVKPKMGDALLFWSMKPDAT 279
Query: 223 PDTLSLHAGCPVIEGEKWSATKWIHVDSF 251
D LSLH GCPVI+G KWS+TKW+HV +
Sbjct: 280 LDPLSLHGGCPVIKGNKWSSTKWMHVREY 308
>UniRef100_Q8LEY1 Putative prolyl 4-hydroxylase, alpha subunit [Arabidopsis thaliana]
Length = 287
Score = 241 bits (616), Expect = 1e-62
Identities = 115/209 (55%), Positives = 152/209 (72%), Gaps = 2/209 (0%)
Query: 43 VSWKPRAFVYKGFLTDLECDHLISIAKSELKRSAVADNLSGESKLSEVRTSSGMFISKNK 102
+SW+PRAFVY FL+ EC++LIS+AK + +S V D+ +G+SK S VRTSSG F+ + +
Sbjct: 79 LSWEPRAFVYHNFLSKEECEYLISLAKPHMVKSTVVDSETGKSKDSRVRTSSGTFLRRGR 138
Query: 103 DAIVSGIEDKISSWTFLPKENGEDIQVLRYEHGQKYDPHYDYFADKVNIARGGHRVATVL 162
D I+ IE +I+ +TF+P ++GE +QVL YE GQKY+PHYDYF D+ N GG R+AT+L
Sbjct: 139 DKIIKTIEKRIADYTFIPADHGEGLQVLHYEAGQKYEPHYDYFVDEFNTKNGGQRMATML 198
Query: 163 MYLTNVTKGGETVFPNAEESPRHKLSETDEDLSECGKKGVAVNPRRGDALLFCSLHPNAI 222
MYL++V +GGETVFP A + +LSECGKKG++V PR GDALLF S+ P+A
Sbjct: 199 MYLSDVEEGGETVFPAA--NMNFSSVPWYNELSECGKKGLSVKPRMGDALLFWSMRPDAT 256
Query: 223 PDTLSLHAGCPVIEGEKWSATKWIHVDSF 251
D SLH GCPVI G KWS+TKWIHV +
Sbjct: 257 LDPTSLHGGCPVIRGNKWSSTKWIHVGEY 285
>UniRef100_Q9LN20 F14O10.12 protein [Arabidopsis thaliana]
Length = 287
Score = 240 bits (613), Expect = 3e-62
Identities = 114/209 (54%), Positives = 152/209 (72%), Gaps = 2/209 (0%)
Query: 43 VSWKPRAFVYKGFLTDLECDHLISIAKSELKRSAVADNLSGESKLSEVRTSSGMFISKNK 102
+SW+PRAFVY FL+ EC++LIS+AK + +S V D+ +G+SK S VRTSSG F+ + +
Sbjct: 79 LSWEPRAFVYHNFLSKEECEYLISLAKPHMVKSTVVDSETGKSKDSRVRTSSGTFLRRGR 138
Query: 103 DAIVSGIEDKISSWTFLPKENGEDIQVLRYEHGQKYDPHYDYFADKVNIARGGHRVATVL 162
D I+ IE +I+ +TF+P ++GE +QVL YE GQKY+PHYDYF D+ N GG R+AT+L
Sbjct: 139 DKIIKTIEKRIADYTFIPADHGEGLQVLHYEAGQKYEPHYDYFVDEFNTKNGGQRMATML 198
Query: 163 MYLTNVTKGGETVFPNAEESPRHKLSETDEDLSECGKKGVAVNPRRGDALLFCSLHPNAI 222
MYL++V +GGETVFP A + +LSECGKKG++V PR GDALLF S+ P+A
Sbjct: 199 MYLSDVEEGGETVFPAA--NMNFSSVPWYNELSECGKKGLSVKPRMGDALLFWSMRPDAT 256
Query: 223 PDTLSLHAGCPVIEGEKWSATKWIHVDSF 251
D SLH GCPVI G KWS+TKW+HV +
Sbjct: 257 LDPTSLHGGCPVIRGNKWSSTKWMHVGEY 285
>UniRef100_Q8LB05 Putative prolyl 4-hydroxylase, alpha subunit [Arabidopsis thaliana]
Length = 291
Score = 236 bits (602), Expect = 6e-61
Identities = 116/214 (54%), Positives = 151/214 (70%), Gaps = 6/214 (2%)
Query: 40 VKQVSWKPRAFVYKGFLTDLECDHLISIAKSELKRSAVADNLSGESKLSEVRTSSGMFIS 99
V+ +SW+PRA VY FLT+ EC+HLIS+AK + +S V D +G SK S VRTSSG F+
Sbjct: 80 VEVISWEPRAVVYHNFLTNEECEHLISLAKPSMVKSTVVDEKTGGSKDSRVRTSSGTFLR 139
Query: 100 KNKDAIVSGIEDKISSWTFLPKENGEDIQVLRYEHGQKYDPHYDYFADKVNIARGGHRVA 159
+ D +V IE +IS +TF+P ENGE +QVL Y+ GQKY+PHYDYF D+ N GG R+A
Sbjct: 140 RGHDEVVEVIEKRISDFTFIPVENGEGLQVLHYQVGQKYEPHYDYFLDEFNTKNGGQRIA 199
Query: 160 TVLMYLTNVTKGGETVFPNAEESPRHKLSETD--EDLSECGKKGVAVNPRRGDALLFCSL 217
TVLMYL++V GGETVFP A R +S +LS+CGK+G++V P+ DALLF ++
Sbjct: 200 TVLMYLSDVDDGGETVFPAA----RGNISAVPWWNELSKCGKEGLSVLPKXRDALLFWNM 255
Query: 218 HPNAIPDTLSLHAGCPVIEGEKWSATKWIHVDSF 251
P+A D SLH GCPV++G KWS+TKW HV F
Sbjct: 256 RPDASLDPSSLHGGCPVVKGNKWSSTKWFHVHEF 289
>UniRef100_UPI00002F63AB UPI00002F63AB UniRef100 entry
Length = 287
Score = 228 bits (581), Expect = 1e-58
Identities = 121/279 (43%), Positives = 164/279 (58%), Gaps = 25/279 (8%)
Query: 47 PRAFVYKGFLTDLECDHLISIAKSELKRSAVADNLSGESKLSEVRTSSGMFISKNKDAIV 106
P+A++ + FL++ ECDHL+++AK EL S V + G S S +RTS+GMF+ K +D +V
Sbjct: 8 PKAYLLRNFLSESECDHLMNLAKQELAPSTVVGD-GGTSVSSNIRTSAGMFLRKGQDEVV 66
Query: 107 SGIEDKISSWTFLPKENGEDIQVLRYEHGQKYDPHYDYFADKVNIA--RGGHRVATVLMY 164
IE++I+ + P++NGE IQ+LRY+ GQKYDPHYDYF DKVN A RGG RVATVL+Y
Sbjct: 67 RRIEERIARASGTPEKNGEGIQILRYDKGQKYDPHYDYFHDKVNPAPKRGGQRVATVLIY 126
Query: 165 LTNVTKGGETVFPNAEESPRHKLSETDEDL------SECGKKGVAVNPRRGDALLFCSLH 218
L + +GGET FPNAE + +E + S+C K+G+ V RGDA+LF S+
Sbjct: 127 LVDTEEGGETTFPNAELPEHFEANEENNPFASNVKRSDCAKRGIPVKSVRGDAILFFSMT 186
Query: 219 PNAIPDTLSLHAGCPVIEGEKWSATKWIHVDSFDKTV------------GAGGDCTDQHE 266
+ D+ SLH CPV+ G+KW+A KWI V FD A C D+
Sbjct: 187 SDGALDSGSLHGACPVVAGQKWTAVKWIRVAEFDGNFQHELPMISLTRRTADEPCVDEWS 246
Query: 267 SCERWAALGECTKNPEYMVGTSGL----PGYCRKSCKTC 301
C WA G C +NPE+M C +SC C
Sbjct: 247 ECAAWARDGWCERNPEFMTAPDSARDSESPACARSCGLC 285
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.318 0.135 0.422
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 548,293,172
Number of Sequences: 2790947
Number of extensions: 23021323
Number of successful extensions: 49629
Number of sequences better than 10.0: 313
Number of HSP's better than 10.0 without gapping: 234
Number of HSP's successfully gapped in prelim test: 80
Number of HSP's that attempted gapping in prelim test: 48849
Number of HSP's gapped (non-prelim): 392
length of query: 301
length of database: 848,049,833
effective HSP length: 127
effective length of query: 174
effective length of database: 493,599,564
effective search space: 85886324136
effective search space used: 85886324136
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 74 (33.1 bits)
Medicago: description of AC146553.10


