

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146550.4 - phase: 0 /pseudo
(322 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
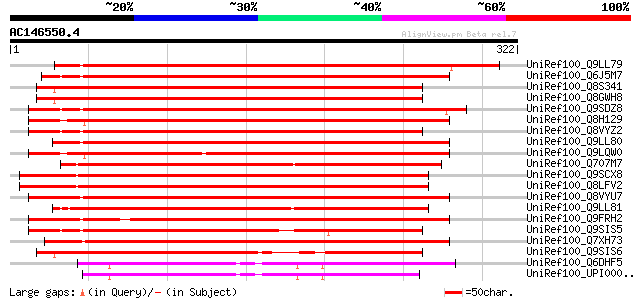
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9LL79 Putative purple acid phosphatase [Phaseolus vul... 347 3e-94
UniRef100_Q6J5M7 Purple acid phosphatase 1 [Solanum tuberosum] 344 2e-93
UniRef100_Q8S341 Purple acid phosphatase [Arabidopsis thaliana] 339 5e-92
UniRef100_Q8GWH8 Putative purple acid phosphatase [Arabidopsis t... 339 5e-92
UniRef100_Q9SDZ8 Putative purple acid phosphatase precursor [Ara... 338 9e-92
UniRef100_Q8H129 Putative purple acid phosphatase [Arabidopsis t... 338 1e-91
UniRef100_Q8VYZ2 Purple acid phosphatase [Arabidopsis thaliana] 333 5e-90
UniRef100_Q9LL80 Putative purple acid phosphatase [Glycine max] 332 6e-90
UniRef100_Q9LQW0 F10B6.10 [Arabidopsis thaliana] 330 3e-89
UniRef100_Q707M7 Putative acid phosphatase [Lupinus luteus] 327 2e-88
UniRef100_Q9SCX8 Acid phosphatase type 5 precursor [Arabidopsis ... 320 3e-86
UniRef100_Q8LFV2 Acid phosphatase type 5 [Arabidopsis thaliana] 318 9e-86
UniRef100_Q8VYU7 At1g25230/F4F7_8 [Arabidopsis thaliana] 315 8e-85
UniRef100_Q9LL81 Putative purple acid phosphatase [Ipomoea batatas] 313 3e-84
UniRef100_Q9FRH2 Hypothetical protein F4F7.38 [Arabidopsis thali... 301 1e-80
UniRef100_Q9SIS5 Putative purple acid phosphatase [Arabidopsis t... 298 1e-79
UniRef100_Q7XH73 Putative purple acid phosphatase [Oryza sativa] 298 2e-79
UniRef100_Q9SIS6 Putative purple acid phosphatase [Arabidopsis t... 255 1e-66
UniRef100_Q6DHF5 Zgc:92339 [Brachydanio rerio] 136 7e-31
UniRef100_UPI0000437F1C UPI0000437F1C UniRef100 entry 129 8e-29
>UniRef100_Q9LL79 Putative purple acid phosphatase [Phaseolus vulgaris]
Length = 331
Score = 347 bits (889), Expect = 3e-94
Identities = 172/297 (57%), Positives = 215/297 (71%), Gaps = 15/297 (5%)
Query: 29 SIAEKLPRFEHHLKPQQQSLNFLVVGDWGRKGNYNQSLVAHQMGIVGDNLNIDFVISTGD 88
+++ L R EH +K SL+ +V+GDWGRKG YNQS V+ QMG VG LNIDFVISTGD
Sbjct: 17 NVSALLQRLEHPVKADG-SLSLVVIGDWGRKGTYNQSEVSAQMGRVGAKLNIDFVISTGD 75
Query: 89 NFYKDGLEGVDDPTFYESFVNIYTAPSLQKIWYSVLGNHDYRGDVDAQLSSILRQKDSRW 148
NFY DGL GVDDP F SF IYTA SLQK WYSVLGNHDYRGDV+AQL++IL++ D RW
Sbjct: 76 NFYDDGLSGVDDPAFELSFSKIYTAKSLQKQWYSVLGNHDYRGDVEAQLNTILQKIDPRW 135
Query: 149 VCLRSFILDGGIVEFFFVDTTPFIEKYFTDPKEHTYDWNGVLPRESYRAELLKNVDLALV 208
+C RSFI+D I EFFFVDTTPF++KYF PK+HTYDW GVLPR+ Y ++LLK++++AL
Sbjct: 136 ICQRSFIVDTEIAEFFFVDTTPFVDKYFLKPKDHTYDWTGVLPRDKYLSKLLKDLEIALK 195
Query: 209 KSKAKWKIVVGHHTIKSVGHHGNTQELEQQLLPILKSNNIDAYINGHDHCLQHIIDNERH 268
S AKWKIVVGHH ++S+GHHG+TQEL + LLPIL++N++D YINGHDHCL+HI
Sbjct: 196 DSTAKWKIVVGHHPVRSIGHHGDTQELIRHLLPILEANDVDMYINGHDHCLEHISSTSSQ 255
Query: 269 GEAISSHGTRK--------------*SFTMTDKDSCLCRSQK*M*MLYSMTFWARFC 311
+ ++S G K FTM DK + R +K + L+ M F A+FC
Sbjct: 256 IQFLTSGGGSKAWKGDHLIKMGKMGQRFTMMDKYLQVWRFKKSIPKLFIMIFLAKFC 312
>UniRef100_Q6J5M7 Purple acid phosphatase 1 [Solanum tuberosum]
Length = 328
Score = 344 bits (883), Expect = 2e-93
Identities = 161/259 (62%), Positives = 206/259 (79%), Gaps = 2/259 (0%)
Query: 21 LRLLSNHSSIAEKLPRFEHHLKPQQQSLNFLVVGDWGRKGNYNQSLVAHQMGIVGDNLNI 80
L LL +++AE L R EH + S++FLVVGDWGR+G +NQS VA QMGI+G+ LNI
Sbjct: 14 LLLLLFPAAMAE-LHRLEHPVNTDG-SISFLVVGDWGRRGTFNQSQVAQQMGIIGEKLNI 71
Query: 81 DFVISTGDNFYKDGLEGVDDPTFYESFVNIYTAPSLQKIWYSVLGNHDYRGDVDAQLSSI 140
DFV+STGDNFY DGL GVDDP F ESF N+YTAPSLQK WY+VLGNHDYRGD AQLS I
Sbjct: 72 DFVVSTGDNFYDDGLTGVDDPAFEESFTNVYTAPSLQKNWYNVLGNHDYRGDALAQLSPI 131
Query: 141 LRQKDSRWVCLRSFILDGGIVEFFFVDTTPFIEKYFTDPKEHTYDWNGVLPRESYRAELL 200
L+QKD+RW+C+RS+I++ + EFFFVDTTPF + YFT PK+HTYDW V+PR+ Y +++L
Sbjct: 132 LKQKDNRWICMRSYIVNTDVAEFFFVDTTPFQDMYFTTPKDHTYDWRNVMPRKDYLSQVL 191
Query: 201 KNVDLALVKSKAKWKIVVGHHTIKSVGHHGNTQELEQQLLPILKSNNIDAYINGHDHCLQ 260
K++D AL +S AKWKIVVGHHTIKS GHHG+++EL +LPIL++NN+D Y+NGHDHCL+
Sbjct: 192 KDLDSALRESSAKWKIVVGHHTIKSAGHHGSSEELGVHILPILQANNVDFYLNGHDHCLE 251
Query: 261 HIIDNERHGEAISSHGTRK 279
HI ++ + ++S G K
Sbjct: 252 HISSSDSPLQFLTSGGGSK 270
>UniRef100_Q8S341 Purple acid phosphatase [Arabidopsis thaliana]
Length = 328
Score = 339 bits (870), Expect = 5e-92
Identities = 160/248 (64%), Positives = 194/248 (77%), Gaps = 3/248 (1%)
Query: 18 SVLLRLLSNH--SSIAEKLPRFEHHLKPQQQ-SLNFLVVGDWGRKGNYNQSLVAHQMGIV 74
SV+L LS + KL R +H +K + SL+FLV+GDWGRKG +NQSLVAHQMG+V
Sbjct: 8 SVILMFLSIFFINGALSKLERLKHPVKKKSDGSLSFLVIGDWGRKGGFNQSLVAHQMGVV 67
Query: 75 GDNLNIDFVISTGDNFYKDGLEGVDDPTFYESFVNIYTAPSLQKIWYSVLGNHDYRGDVD 134
G+ L+IDFVIS GDNFY DGL+GV+DP+F SF +IYT PSLQK WYSVLGNHDYRG+V+
Sbjct: 68 GEKLDIDFVISVGDNFYDDGLKGVNDPSFEASFSHIYTHPSLQKQWYSVLGNHDYRGNVE 127
Query: 135 AQLSSILRQKDSRWVCLRSFILDGGIVEFFFVDTTPFIEKYFTDPKEHTYDWNGVLPRES 194
AQLS +L QKD RW C RSF+L G+V+FFF DT PF+EKYFT+P++HTYDW VLPR
Sbjct: 128 AQLSKVLTQKDWRWFCRRSFVLSSGMVDFFFADTNPFVEKYFTEPEDHTYDWRNVLPRNK 187
Query: 195 YRAELLKNVDLALVKSKAKWKIVVGHHTIKSVGHHGNTQELEQQLLPILKSNNIDAYING 254
Y + LL ++DL + KS+A WK VVGHH IK+ G+HG TQEL QLLPIL+ N +D YING
Sbjct: 188 YISNLLHDLDLEIKKSRATWKFVVGHHGIKTAGNHGVTQELVDQLLPILEENKVDLYING 247
Query: 255 HDHCLQHI 262
HDHCLQHI
Sbjct: 248 HDHCLQHI 255
>UniRef100_Q8GWH8 Putative purple acid phosphatase [Arabidopsis thaliana]
Length = 328
Score = 339 bits (870), Expect = 5e-92
Identities = 160/248 (64%), Positives = 194/248 (77%), Gaps = 3/248 (1%)
Query: 18 SVLLRLLSNH--SSIAEKLPRFEHHLKPQQQ-SLNFLVVGDWGRKGNYNQSLVAHQMGIV 74
SV+L LS + KL R +H +K + SL+FLV+GDWGRKG +NQSLVAHQMG+V
Sbjct: 8 SVILMFLSIFFINGALSKLERLKHPVKKKSDGSLSFLVIGDWGRKGGFNQSLVAHQMGVV 67
Query: 75 GDNLNIDFVISTGDNFYKDGLEGVDDPTFYESFVNIYTAPSLQKIWYSVLGNHDYRGDVD 134
G+ L+IDFVIS GDNFY DGL+GV+DP+F SF +IYT PSLQK WYSVLGNHDYRG+V+
Sbjct: 68 GEKLDIDFVISVGDNFYDDGLKGVNDPSFEASFSHIYTHPSLQKQWYSVLGNHDYRGNVE 127
Query: 135 AQLSSILRQKDSRWVCLRSFILDGGIVEFFFVDTTPFIEKYFTDPKEHTYDWNGVLPRES 194
AQLS +L QKD RW C RSF+L G+V+FFF DT PF+EKYFT+P++HTYDW VLPR
Sbjct: 128 AQLSKVLTQKDWRWFCRRSFVLSSGMVDFFFADTNPFVEKYFTEPEDHTYDWRNVLPRNK 187
Query: 195 YRAELLKNVDLALVKSKAKWKIVVGHHTIKSVGHHGNTQELEQQLLPILKSNNIDAYING 254
Y + LL ++DL + KS+A WK VVGHH IK+ G+HG TQEL QLLPIL+ N +D YING
Sbjct: 188 YISNLLHDLDLEIKKSRATWKFVVGHHGIKTAGNHGVTQELVDQLLPILEENKVDLYING 247
Query: 255 HDHCLQHI 262
HDHCLQHI
Sbjct: 248 HDHCLQHI 255
>UniRef100_Q9SDZ8 Putative purple acid phosphatase precursor [Arabidopsis thaliana]
Length = 314
Score = 338 bits (868), Expect = 9e-92
Identities = 172/291 (59%), Positives = 211/291 (72%), Gaps = 15/291 (5%)
Query: 13 VIFPASVLLRLLSNHSSIAEKLPRFEHHLKPQQQSLNFLVVGDWGRKGNYNQSLVAHQMG 72
+IF L+ +LS +S AE LPRF +P SL+FLVVGDWGR+G+YNQS VA QMG
Sbjct: 12 LIFSIFCLVIILSACNSTAE-LPRFVQPPEPDG-SLSFLVVGDWGRRGSYNQSQVALQMG 69
Query: 73 IVGDNLNIDFVISTGDNFYKDGLEGVDDPTFYESFVNIYTAPSLQKIWYSVLGNHDYRGD 132
+G +LNIDF+ISTGDNFY DG+ D F +SF NIYTA SLQK WY+VLGNHDYRG+
Sbjct: 70 KIGKDLNIDFLISTGDNFYDDGIISPYDSQFQDSFTNIYTATSLQKPWYNVLGNHDYRGN 129
Query: 133 VDAQLSSILRQKDSRWVCLRSFILDGGIVEFFFVDTTPFIEKYFTDPKEHTYDWNGVLPR 192
V AQLS ILR D RW+CLRS++++ IV+ FFVDTTPF+++YF +PK+H YDW GVLPR
Sbjct: 130 VYAQLSPILRDLDCRWICLRSYVVNAEIVDIFFVDTTPFVDRYFDEPKDHVYDWRGVLPR 189
Query: 193 ESYRAELLKNVDLALVKSKAKWKIVVGHHTIKSVGHHGNTQELEQQLLPILKSNNIDAYI 252
Y LL +VD+AL +S AKWKIVVGHHTIKS GHHGNT ELE+QLLPIL++N +D YI
Sbjct: 190 NKYLNSLLTDVDVALQESMAKWKIVVGHHTIKSAGHHGNTIELEKQLLPILEANEVDLYI 249
Query: 253 NGHDHCLQHIIDNERHGEAISSHG-------------TRK*SFTMTDKDSC 290
NGHDHCL+HI + ++S G ++ FTM DKDSC
Sbjct: 250 NGHDHCLEHISGINSGMQFMTSGGGFKAWKGDVNDWNLQEMRFTMMDKDSC 300
>UniRef100_Q8H129 Putative purple acid phosphatase [Arabidopsis thaliana]
Length = 366
Score = 338 bits (867), Expect = 1e-91
Identities = 161/269 (59%), Positives = 207/269 (76%), Gaps = 6/269 (2%)
Query: 13 VIFPASVLLRLLSNHSSIAEKLPRFEHHLKPQQQ--SLNFLVVGDWGRKGNYNQSLVAHQ 70
++F L+ + S+HSS AE L+P + +++FLV+GDWGR+G+YNQS VA Q
Sbjct: 41 LVFYVYNLIIIFSSHSSTAE----LRRLLQPSKTDGTVSFLVIGDWGRRGSYNQSQVALQ 96
Query: 71 MGIVGDNLNIDFVISTGDNFYKDGLEGVDDPTFYESFVNIYTAPSLQKIWYSVLGNHDYR 130
MG +G+ L+IDFVISTGDNFY +GL + DP F +SF NIYTAPSLQK WYSVLGNHDYR
Sbjct: 97 MGEIGEKLDIDFVISTGDNFYDNGLTSLHDPLFQDSFTNIYTAPSLQKPWYSVLGNHDYR 156
Query: 131 GDVDAQLSSILRQKDSRWVCLRSFILDGGIVEFFFVDTTPFIEKYFTDPKEHTYDWNGVL 190
GDV AQLS +LR D+RWVC+RSFI++ IV+ FFVDTTPF++KYF P +H YDW+GVL
Sbjct: 157 GDVRAQLSPMLRALDNRWVCMRSFIVNAEIVDLFFVDTTPFVDKYFIQPNKHVYDWSGVL 216
Query: 191 PRESYRAELLKNVDLALVKSKAKWKIVVGHHTIKSVGHHGNTQELEQQLLPILKSNNIDA 250
PR++Y LLK +D+AL +S AKWKIV+GHHTIKS GHHGNT ELE+ LLPIL++N +D
Sbjct: 217 PRQTYLNNLLKELDVALRESVAKWKIVIGHHTIKSAGHHGNTIELEKHLLPILQANEVDL 276
Query: 251 YINGHDHCLQHIIDNERHGEAISSHGTRK 279
Y+NGHDHCL+HI + + + ++S G K
Sbjct: 277 YVNGHDHCLEHISSVDSNIQFMTSGGGSK 305
>UniRef100_Q8VYZ2 Purple acid phosphatase [Arabidopsis thaliana]
Length = 335
Score = 333 bits (853), Expect = 5e-90
Identities = 162/250 (64%), Positives = 196/250 (77%), Gaps = 2/250 (0%)
Query: 13 VIFPASVLLRLLSNHSSIAEKLPRFEHHLKPQQQSLNFLVVGDWGRKGNYNQSLVAHQMG 72
+IF L+ +LS +S AE LPRF +P SL+FLVVGDWGR+G+YNQS VA QMG
Sbjct: 12 LIFSIFCLVIILSACNSTAE-LPRFVQPPEPDG-SLSFLVVGDWGRRGSYNQSQVALQMG 69
Query: 73 IVGDNLNIDFVISTGDNFYKDGLEGVDDPTFYESFVNIYTAPSLQKIWYSVLGNHDYRGD 132
+G +LNIDF+ISTGDNFY DG+ D F +SF NIYTA SLQK WY+VLGNHDYRG+
Sbjct: 70 KIGKDLNIDFLISTGDNFYDDGIISPYDSQFQDSFTNIYTATSLQKPWYNVLGNHDYRGN 129
Query: 133 VDAQLSSILRQKDSRWVCLRSFILDGGIVEFFFVDTTPFIEKYFTDPKEHTYDWNGVLPR 192
V AQLS ILR D RW+CLRS++++ IV+ FFVDTTPF+++YF +PK+H YDW GVLPR
Sbjct: 130 VYAQLSPILRDLDCRWICLRSYVVNAEIVDIFFVDTTPFVDRYFDEPKDHVYDWRGVLPR 189
Query: 193 ESYRAELLKNVDLALVKSKAKWKIVVGHHTIKSVGHHGNTQELEQQLLPILKSNNIDAYI 252
Y LL +VD+AL +S AKWKIVVGHHTIKS GHHGNT ELE+QLLPIL++N +D YI
Sbjct: 190 NKYLNSLLTDVDVALQESMAKWKIVVGHHTIKSAGHHGNTIELEKQLLPILEANEVDLYI 249
Query: 253 NGHDHCLQHI 262
NGHDHCL+HI
Sbjct: 250 NGHDHCLEHI 259
>UniRef100_Q9LL80 Putative purple acid phosphatase [Glycine max]
Length = 332
Score = 332 bits (852), Expect = 6e-90
Identities = 154/252 (61%), Positives = 195/252 (77%), Gaps = 1/252 (0%)
Query: 28 SSIAEKLPRFEHHLKPQQQSLNFLVVGDWGRKGNYNQSLVAHQMGIVGDNLNIDFVISTG 87
SS KL +H K SL+FLVVGDWGRKG YNQSLVA QMG++G+ L++DFVISTG
Sbjct: 24 SSSKAKLESLQHAPKADG-SLSFLVVGDWGRKGAYNQSLVAFQMGVIGEKLDVDFVISTG 82
Query: 88 DNFYKDGLEGVDDPTFYESFVNIYTAPSLQKIWYSVLGNHDYRGDVDAQLSSILRQKDSR 147
DNFY +GL GV DP+F ESF IYTAPSLQK WY+VLGNHDYRG+ AQ+S +LR +D+R
Sbjct: 83 DNFYDNGLTGVFDPSFEESFTKIYTAPSLQKKWYNVLGNHDYRGNAKAQISHVLRYRDNR 142
Query: 148 WVCLRSFILDGGIVEFFFVDTTPFIEKYFTDPKEHTYDWNGVLPRESYRAELLKNVDLAL 207
WVC RS+ L+ V+FFFVDTTP+++KYF + K H YDW G+LPR+ Y + LLK+VDLAL
Sbjct: 143 WVCFRSYTLNSENVDFFFVDTTPYVDKYFIEDKGHNYDWRGILPRKRYTSNLLKDVDLAL 202
Query: 208 VKSKAKWKIVVGHHTIKSVGHHGNTQELEQQLLPILKSNNIDAYINGHDHCLQHIIDNER 267
+S A WK+V+GHHTIK++GHHG+TQEL LP+LK+NN+D Y+NGHDHCL+HI +
Sbjct: 203 RQSTATWKVVIGHHTIKNIGHHGDTQELLIHFLPLLKANNVDLYMNGHDHCLEHISSLDS 262
Query: 268 HGEAISSHGTRK 279
+ ++S G K
Sbjct: 263 SVQFLTSGGGSK 274
>UniRef100_Q9LQW0 F10B6.10 [Arabidopsis thaliana]
Length = 364
Score = 330 bits (846), Expect = 3e-89
Identities = 159/269 (59%), Positives = 205/269 (76%), Gaps = 8/269 (2%)
Query: 13 VIFPASVLLRLLSNHSSIAEKLPRFEHHLKPQQQ--SLNFLVVGDWGRKGNYNQSLVAHQ 70
++F L+ + S+HSS AE L+P + +++FLV+GDWGR+G+YNQS VA Q
Sbjct: 41 LVFYVYNLIIIFSSHSSTAE----LRRLLQPSKTDGTVSFLVIGDWGRRGSYNQSQVALQ 96
Query: 71 MGIVGDNLNIDFVISTGDNFYKDGLEGVDDPTFYESFVNIYTAPSLQKIWYSVLGNHDYR 130
MG +G+ L+IDFVISTGDNFY +GL + DP F +SF NIYTAPSLQK WYS GNHDYR
Sbjct: 97 MGEIGEKLDIDFVISTGDNFYDNGLTSLHDPLFQDSFTNIYTAPSLQKPWYS--GNHDYR 154
Query: 131 GDVDAQLSSILRQKDSRWVCLRSFILDGGIVEFFFVDTTPFIEKYFTDPKEHTYDWNGVL 190
GDV AQLS +LR D+RWVC+RSFI++ IV+ FFVDTTPF++KYF P +H YDW+GVL
Sbjct: 155 GDVRAQLSPMLRALDNRWVCMRSFIVNAEIVDLFFVDTTPFVDKYFIQPNKHVYDWSGVL 214
Query: 191 PRESYRAELLKNVDLALVKSKAKWKIVVGHHTIKSVGHHGNTQELEQQLLPILKSNNIDA 250
PR++Y LLK +D+AL +S AKWKIV+GHHTIKS GHHGNT ELE+ LLPIL++N +D
Sbjct: 215 PRQTYLNNLLKELDVALRESVAKWKIVIGHHTIKSAGHHGNTIELEKHLLPILQANEVDL 274
Query: 251 YINGHDHCLQHIIDNERHGEAISSHGTRK 279
Y+NGHDHCL+HI + + + ++S G K
Sbjct: 275 YVNGHDHCLEHISSVDSNIQFMTSGGGSK 303
>UniRef100_Q707M7 Putative acid phosphatase [Lupinus luteus]
Length = 330
Score = 327 bits (839), Expect = 2e-88
Identities = 154/242 (63%), Positives = 193/242 (79%), Gaps = 2/242 (0%)
Query: 33 KLPRFEHHLKPQQQSLNFLVVGDWGRKGNYNQSLVAHQMGIVGDNLNIDFVISTGDNFYK 92
+L +F H K SLNFLV+GDWGR+G YNQS +A QMG VG+ L+IDFV+STGDNFY
Sbjct: 28 ELHKFAHSSK-HDGSLNFLVLGDWGRRGAYNQSEIAFQMGKVGEKLDIDFVVSTGDNFYD 86
Query: 93 DGLEGVDDPTFYESFVNIYTAPSLQKIWYSVLGNHDYRGDVDAQLSSILRQKDSRWVCLR 152
+GL D F ESF N+YTA SLQK WYSVLGNHDYRGDV+AQLS L+Q D+RW+CLR
Sbjct: 87 NGLTSDQDTAFEESFTNVYTAKSLQKQWYSVLGNHDYRGDVEAQLSPFLKQIDNRWLCLR 146
Query: 153 SFILDGGIVEFFFVDTTPFIEKYFTDPKEHTYDWNGVLPRESYRAELLKNVDLALVKSKA 212
SFI+D +VE FFVDTTPF+EKYFT+ K H YDW G++P++SY LLK+++LA+ +S A
Sbjct: 147 SFIVDSELVEIFFVDTTPFVEKYFTETK-HKYDWQGIIPQKSYITNLLKDLELAIKESTA 205
Query: 213 KWKIVVGHHTIKSVGHHGNTQELEQQLLPILKSNNIDAYINGHDHCLQHIIDNERHGEAI 272
+WKIVVGHH I+SVGHHG+TQEL +QLLPIL+ N++D Y+NGHDHCL+HI D E + +
Sbjct: 206 QWKIVVGHHAIRSVGHHGDTQELIKQLLPILQENDVDFYMNGHDHCLEHISDTESSIQFL 265
Query: 273 SS 274
+S
Sbjct: 266 TS 267
>UniRef100_Q9SCX8 Acid phosphatase type 5 precursor [Arabidopsis thaliana]
Length = 338
Score = 320 bits (820), Expect = 3e-86
Identities = 146/261 (55%), Positives = 200/261 (75%), Gaps = 2/261 (0%)
Query: 7 QRMLSPVIFPASVLLRLLSNHSSIAE-KLPRFEHHLKPQQQSLNFLVVGDWGRKGNYNQS 65
+R L S+LL + + ++ +L RF K S++F+V+GDWGR+G++NQS
Sbjct: 5 RRSLMSATASLSLLLCIFTTFVVVSNGELQRFIEPAK-SDGSVSFIVIGDWGRRGSFNQS 63
Query: 66 LVAHQMGIVGDNLNIDFVISTGDNFYKDGLEGVDDPTFYESFVNIYTAPSLQKIWYSVLG 125
LVA+QMG +G+ +++DFV+STGDNFY +GL DP F +SF NIYTAPSLQK WYSVLG
Sbjct: 64 LVAYQMGKIGEKIDLDFVVSTGDNFYDNGLFSEHDPNFEQSFSNIYTAPSLQKQWYSVLG 123
Query: 126 NHDYRGDVDAQLSSILRQKDSRWVCLRSFILDGGIVEFFFVDTTPFIEKYFTDPKEHTYD 185
NHDYRGD +AQLSS+LR+ DSRW+CLRSF++D +VE FFVDTTPF+++Y+T+ H+YD
Sbjct: 124 NHDYRGDAEAQLSSVLREIDSRWICLRSFVVDAELVEMFFVDTTPFVKEYYTEADGHSYD 183
Query: 186 WNGVLPRESYRAELLKNVDLALVKSKAKWKIVVGHHTIKSVGHHGNTQELEQQLLPILKS 245
W V R SY LL++++++L SKA+WKIVVGHH ++S+GHHG+T+EL ++LLPILK
Sbjct: 184 WRAVPSRNSYVKALLRDLEVSLKSSKARWKIVVGHHAMRSIGHHGDTKELNEELLPILKE 243
Query: 246 NNIDAYINGHDHCLQHIIDNE 266
N +D Y+NGHDHCLQH+ D +
Sbjct: 244 NGVDLYMNGHDHCLQHMSDED 264
>UniRef100_Q8LFV2 Acid phosphatase type 5 [Arabidopsis thaliana]
Length = 338
Score = 318 bits (816), Expect = 9e-86
Identities = 145/261 (55%), Positives = 201/261 (76%), Gaps = 2/261 (0%)
Query: 7 QRMLSPVIFPASVLLRLLSNHSSIAE-KLPRFEHHLKPQQQSLNFLVVGDWGRKGNYNQS 65
+R L S+LL + + ++ +L RF K S++F+V+GDWGR+G++NQS
Sbjct: 5 RRSLISATASLSLLLCIFTTFVVVSNGELQRFIEPAK-SDGSVSFIVIGDWGRRGSFNQS 63
Query: 66 LVAHQMGIVGDNLNIDFVISTGDNFYKDGLEGVDDPTFYESFVNIYTAPSLQKIWYSVLG 125
LVA+QMG +G+ +++DFV+STGDNFY +GL DP F +SF NIYTAPSL+K WYSVLG
Sbjct: 64 LVAYQMGKIGEKIDLDFVVSTGDNFYDNGLFSEHDPNFEQSFSNIYTAPSLRKQWYSVLG 123
Query: 126 NHDYRGDVDAQLSSILRQKDSRWVCLRSFILDGGIVEFFFVDTTPFIEKYFTDPKEHTYD 185
NHDYRGD +AQLSS+LR+ DSRW+CLRSF++D +VE FFVDTTPF+++Y+T+ H+YD
Sbjct: 124 NHDYRGDAEAQLSSVLREIDSRWICLRSFVVDAELVEMFFVDTTPFVKEYYTEADGHSYD 183
Query: 186 WNGVLPRESYRAELLKNVDLALVKSKAKWKIVVGHHTIKSVGHHGNTQELEQQLLPILKS 245
W V R SY LL++++++L SKA+WKIVVGHH ++S+GHHG+T+EL ++LLPILK
Sbjct: 184 WRAVPSRNSYVKALLRDLEVSLKSSKARWKIVVGHHAMRSIGHHGDTKELNEELLPILKE 243
Query: 246 NNIDAYINGHDHCLQHIIDNE 266
N++D Y+NGHDHCLQH+ D +
Sbjct: 244 NSVDLYMNGHDHCLQHMSDED 264
>UniRef100_Q8VYU7 At1g25230/F4F7_8 [Arabidopsis thaliana]
Length = 339
Score = 315 bits (808), Expect = 8e-85
Identities = 148/267 (55%), Positives = 193/267 (71%), Gaps = 1/267 (0%)
Query: 13 VIFPASVLLRLLSNHSSIAEKLPRFEHHLKPQQQSLNFLVVGDWGRKGNYNQSLVAHQMG 72
++ + LLS + +L +H P S++FLV+GDWGR G YNQS VA QMG
Sbjct: 12 IVMTLLICFLLLSLAPKLEAELATVQHAPNPDG-SISFLVIGDWGRHGLYNQSQVALQMG 70
Query: 73 IVGDNLNIDFVISTGDNFYKDGLEGVDDPTFYESFVNIYTAPSLQKIWYSVLGNHDYRGD 132
+G+ ++I+FV+STGDN Y +G++ +DDP F SF NIYT+PSLQK WY VLGNHDYRGD
Sbjct: 71 RIGEEMDINFVVSTGDNIYDNGMKSIDDPAFQLSFSNIYTSPSLQKPWYLVLGNHDYRGD 130
Query: 133 VDAQLSSILRQKDSRWVCLRSFILDGGIVEFFFVDTTPFIEKYFTDPKEHTYDWNGVLPR 192
V+AQLS ILR DSRW+C+RSFI+D I E FFVDTTPF++ YF P++ TYDW+GV PR
Sbjct: 131 VEAQLSPILRSMDSRWICMRSFIVDAEIAELFFVDTTPFVDAYFLSPQDQTYDWSGVSPR 190
Query: 193 ESYRAELLKNVDLALVKSKAKWKIVVGHHTIKSVGHHGNTQELEQQLLPILKSNNIDAYI 252
+SY +L +++ L +S AKWKIVVGHH IKS HGNT+ELE LLPIL++N +D Y+
Sbjct: 191 KSYLQTILTELEMGLRESSAKWKIVVGHHAIKSASIHGNTKELESLLLPILEANKVDLYM 250
Query: 253 NGHDHCLQHIIDNERHGEAISSHGTRK 279
NGHDHCLQHI ++ + ++S G K
Sbjct: 251 NGHDHCLQHISTSQSPIQFLTSGGGSK 277
>UniRef100_Q9LL81 Putative purple acid phosphatase [Ipomoea batatas]
Length = 312
Score = 313 bits (803), Expect = 3e-84
Identities = 148/239 (61%), Positives = 185/239 (76%), Gaps = 3/239 (1%)
Query: 28 SSIAEKLPRFEHHLKPQQQSLNFLVVGDWGRKGNYNQSLVAHQMGIVGDNLNIDFVISTG 87
S+I E LP F HH SL+FLV+GDWGRKG+YNQS VA QMG +GD L IDFV+STG
Sbjct: 25 SAIVE-LPTF-HHPTKGDGSLSFLVIGDWGRKGDYNQSQVAFQMGEIGDQLAIDFVVSTG 82
Query: 88 DNFYKDGLEGVDDPTFYESFVNIYTAPSLQKIWYSVLGNHDYRGDVDAQLSSILRQKDSR 147
DNFY +GL G D F ESF ++YTA SLQK WYSVLGNHDYRGD +AQLSS LR+ DSR
Sbjct: 83 DNFYDNGLTGEHDDAFTESFTDVYTAESLQKQWYSVLGNHDYRGDAEAQLSSHLRKLDSR 142
Query: 148 WVCLRSFILDGGIVEFFFVDTTPFIEKYFTDPKEHTYDWNGVLPRESYRAELLKNVDLAL 207
W CLRSF+++ V+ FFVDTTPF+E+YF P EH YDW GV P+++Y +L ++ AL
Sbjct: 143 WPCLRSFVVNTETVDLFFVDTTPFVEEYFNSP-EHVYDWRGVFPQQTYTKNVLNGLEYAL 201
Query: 208 VKSKAKWKIVVGHHTIKSVGHHGNTQELEQQLLPILKSNNIDAYINGHDHCLQHIIDNE 266
+KS KW+IV+GHH I+S GHHG+T+EL ++LLPIL++ N+D Y+NGHDH L+HI D+E
Sbjct: 202 MKSTTKWRIVIGHHAIRSAGHHGDTKELVERLLPILRTYNVDLYMNGHDHSLEHISDDE 260
>UniRef100_Q9FRH2 Hypothetical protein F4F7.38 [Arabidopsis thaliana]
Length = 333
Score = 301 bits (772), Expect = 1e-80
Identities = 145/267 (54%), Positives = 188/267 (70%), Gaps = 7/267 (2%)
Query: 13 VIFPASVLLRLLSNHSSIAEKLPRFEHHLKPQQQSLNFLVVGDWGRKGNYNQSLVAHQMG 72
++ + LLS + +L +H P S++FLV+GDWGR G YNQS VA Q
Sbjct: 12 IVMTLLICFLLLSLAPKLEAELATVQHAPNPDG-SISFLVIGDWGRHGLYNQSQVALQ-- 68
Query: 73 IVGDNLNIDFVISTGDNFYKDGLEGVDDPTFYESFVNIYTAPSLQKIWYSVLGNHDYRGD 132
++I+FV+STGDN Y +G++ +DDP F SF NIYT+PSLQK WY VLGNHDYRGD
Sbjct: 69 ----EMDINFVVSTGDNIYDNGMKSIDDPAFQLSFSNIYTSPSLQKPWYLVLGNHDYRGD 124
Query: 133 VDAQLSSILRQKDSRWVCLRSFILDGGIVEFFFVDTTPFIEKYFTDPKEHTYDWNGVLPR 192
V+AQLS ILR DSRW+C+RSFI+D I E FFVDTTPF++ YF P++ TYDW+GV PR
Sbjct: 125 VEAQLSPILRSMDSRWICMRSFIVDAEIAELFFVDTTPFVDAYFLSPQDQTYDWSGVSPR 184
Query: 193 ESYRAELLKNVDLALVKSKAKWKIVVGHHTIKSVGHHGNTQELEQQLLPILKSNNIDAYI 252
+SY +L +++ L +S AKWKIVVGHH IKS HGNT+ELE LLPIL++N +D Y+
Sbjct: 185 KSYLQTILTELEMGLRESSAKWKIVVGHHAIKSASIHGNTKELESLLLPILEANKVDLYM 244
Query: 253 NGHDHCLQHIIDNERHGEAISSHGTRK 279
NGHDHCLQHI ++ + ++S G K
Sbjct: 245 NGHDHCLQHISTSQSPIQFLTSGGGSK 271
>UniRef100_Q9SIS5 Putative purple acid phosphatase [Arabidopsis thaliana]
Length = 351
Score = 298 bits (764), Expect = 1e-79
Identities = 158/275 (57%), Positives = 188/275 (67%), Gaps = 36/275 (13%)
Query: 13 VIFPASVLLRLLSNHSSIAEKLPRFEHHLKPQQQSLNFLVVGDWGRKGNYNQSLVAHQMG 72
+IF L+ +LS +S AE LPRF +P SL+FLVVGDWGR+G+YNQS VA QMG
Sbjct: 12 LIFSIFCLVIILSACNSTAE-LPRFVQPPEPDG-SLSFLVVGDWGRRGSYNQSQVALQMG 69
Query: 73 IVGDNLNIDFVISTGDNFYKDGLEGVDDPTFYESFVNIYTAPSLQKIWYSVLGNHDYRGD 132
+G +LNIDF+ISTGDNFY DG+ D F +SF NIYTA SLQK WY+VLGNHDYRG+
Sbjct: 70 KIGKDLNIDFLISTGDNFYDDGIISPYDSQFQDSFTNIYTATSLQKPWYNVLGNHDYRGN 129
Query: 133 VDAQLSSILRQKDSRWVCLRSFILDGGIVEFFFVDTTPFIEKYFTDPKEHTYDWNGVLPR 192
V AQLS ILR D RW+CLRS++++ IV+ FFVDTTPF +H YDW GVLPR
Sbjct: 130 VYAQLSPILRDLDCRWICLRSYVVNAEIVDIFFVDTTPF---------DHVYDWRGVLPR 180
Query: 193 ESYRAELLK-------------------------NVDLALVKSKAKWKIVVGHHTIKSVG 227
Y LL +VD+AL +S AKWKIVVGHHTIKS G
Sbjct: 181 NKYLNSLLTVTFFFLLYHKLVISSNSLKIRIIKLDVDVALQESMAKWKIVVGHHTIKSAG 240
Query: 228 HHGNTQELEQQLLPILKSNNIDAYINGHDHCLQHI 262
HHGNT ELE+QLLPIL++N +D YINGHDHCL+HI
Sbjct: 241 HHGNTIELEKQLLPILEANEVDLYINGHDHCLEHI 275
>UniRef100_Q7XH73 Putative purple acid phosphatase [Oryza sativa]
Length = 335
Score = 298 bits (762), Expect = 2e-79
Identities = 147/257 (57%), Positives = 183/257 (71%), Gaps = 1/257 (0%)
Query: 23 LLSNHSSIAEKLPRFEHHLKPQQQSLNFLVVGDWGRKGNYNQSLVAHQMGIVGDNLNIDF 82
L S S +L R EH + L LVVGDWGRKG YNQ+ VA QMG V + IDF
Sbjct: 14 LSSCTSPATAELTRHEHPVAAGAP-LRLLVVGDWGRKGGYNQTRVAEQMGKVAEETEIDF 72
Query: 83 VISTGDNFYKDGLEGVDDPTFYESFVNIYTAPSLQKIWYSVLGNHDYRGDVDAQLSSILR 142
V+STGDNF ++GL GVDD F++SF+++YTA SL K WY VLGNHDYRG+V AQ+ LR
Sbjct: 73 VVSTGDNFLENGLAGVDDMAFHDSFMDVYTAQSLHKPWYLVLGNHDYRGNVLAQIDPALR 132
Query: 143 QKDSRWVCLRSFILDGGIVEFFFVDTTPFIEKYFTDPKEHTYDWNGVLPRESYRAELLKN 202
+ DSR++C+RSFI+ GIV+FFFVDTTPF +Y+TDP E YDW GV PR++Y A LL++
Sbjct: 133 KIDSRFICMRSFIVSAGIVDFFFVDTTPFQLQYWTDPGEDHYDWRGVAPRDAYIANLLED 192
Query: 203 VDLALVKSKAKWKIVVGHHTIKSVGHHGNTQELEQQLLPILKSNNIDAYINGHDHCLQHI 262
VD A+ KS A WKI VGHHT++SV HG+TQEL + LLP+LK N +D YINGHDHCL+HI
Sbjct: 193 VDAAMKKSTATWKIAVGHHTMRSVSAHGDTQELLELLLPVLKENGVDFYINGHDHCLEHI 252
Query: 263 IDNERHGEAISSHGTRK 279
+ +S G K
Sbjct: 253 SSRNSPIQYFTSGGGSK 269
>UniRef100_Q9SIS6 Putative purple acid phosphatase [Arabidopsis thaliana]
Length = 304
Score = 255 bits (651), Expect = 1e-66
Identities = 138/248 (55%), Positives = 164/248 (65%), Gaps = 27/248 (10%)
Query: 18 SVLLRLLSNH--SSIAEKLPRFEHHLKPQQQ-SLNFLVVGDWGRKGNYNQSLVAHQMGIV 74
SV+L LS + KL R +H +K + SL+FLV+GDWGRKG +NQSLVAHQMG+V
Sbjct: 8 SVILMFLSIFFINGALSKLERLKHPVKKKSDGSLSFLVIGDWGRKGGFNQSLVAHQMGVV 67
Query: 75 GDNLNIDFVISTGDNFYKDGLEGVDDPTFYESFVNIYTAPSLQKIWYSVLGNHDYRGDVD 134
G+ L+IDFVIS GDNFY DGL+GV+DP+F SF +IYT PSLQK WYSVLGNHDYRG+V+
Sbjct: 68 GEKLDIDFVISVGDNFYDDGLKGVNDPSFEASFSHIYTHPSLQKQWYSVLGNHDYRGNVE 127
Query: 135 AQLSSILRQKDSRWVCLRSFILDGGIVEFFFVDTTPFIEKYFTDPKEHTYDWNGVLPRES 194
AQLS +L QKD RW C RSF+L V F Y G R
Sbjct: 128 AQLSKVLTQKDWRWFCRRSFVLSS--VNMFCCG----------------YRNGGFFLRGH 169
Query: 195 YRAELLKNVDLALVKSKAKWKIVVGHHTIKSVGHHGNTQELEQQLLPILKSNNIDAYING 254
K+ L + KS+A WK VVGHH IK+ G+HG TQEL QLLPIL+ N +D YING
Sbjct: 170 ------KSFYLEIKKSRATWKFVVGHHGIKTAGNHGVTQELVDQLLPILEENKVDLYING 223
Query: 255 HDHCLQHI 262
HDHCLQHI
Sbjct: 224 HDHCLQHI 231
>UniRef100_Q6DHF5 Zgc:92339 [Brachydanio rerio]
Length = 327
Score = 136 bits (343), Expect = 7e-31
Identities = 86/251 (34%), Positives = 136/251 (53%), Gaps = 17/251 (6%)
Query: 44 QQQSLNFLVVGDWGRKGNY-----NQSLVAHQMGIVGDNLNIDFVISTGDNFYKDGLEGV 98
Q SL F+ +GDWG + +Y +++ A ++ V + +DFV+S GDNFY DG++ V
Sbjct: 25 QASSLRFVGIGDWGGRPSYPFYTPHEADTAKELARVAQSSGLDFVLSLGDNFYYDGVKDV 84
Query: 99 DDPTFYESFVNIYTAPSLQKI-WYSVLGNHDYRGDVDAQLSSILRQKDSRWVCLRSFILD 157
DD F S+ +++ P+L I WY + GNHD+RG+V AQ++ R + RW+ +
Sbjct: 85 DDTRFKFSYEQVFSHPALMTIPWYLIAGNHDHRGNVSAQIAYSSRSE--RWIYPDLYYE- 141
Query: 158 GGIVEFFFVDTTPFIEKYFTDPKE---HTYDWNGVLPRESYRA--ELLKNVDLALVKSKA 212
+ F + + TD +TYD + E A + L ++ L +KA
Sbjct: 142 ---LNFKVPHSNTSLTILMTDTVVVCGNTYDGLDPVGPEDLAAANKQLAWIEQRLQSTKA 198
Query: 213 KWKIVVGHHTIKSVGHHGNTQELEQQLLPILKSNNIDAYINGHDHCLQHIIDNERHGEAI 272
+ IVVGH+ I S+GHHG T+ L +L P+LK N+ Y++GHDH LQ I +++ +
Sbjct: 199 DFVIVVGHYPIWSIGHHGPTKCLISKLRPLLKKYNVSLYLSGHDHSLQFIREDDGSSFVV 258
Query: 273 SSHGTRK*SFT 283
S G + S T
Sbjct: 259 SGAGVEEDSST 269
>UniRef100_UPI0000437F1C UPI0000437F1C UniRef100 entry
Length = 264
Score = 129 bits (325), Expect = 8e-29
Identities = 80/225 (35%), Positives = 125/225 (55%), Gaps = 17/225 (7%)
Query: 47 SLNFLVVGDWGRKGNY-----NQSLVAHQMGIVGDNLNIDFVISTGDNFYKDGLEGVDDP 101
SL F+ +GDWG + +Y +++ A ++ V + +DFV+S GDNFY DG++ VDD
Sbjct: 46 SLRFVGIGDWGGRPSYPFYTPHEADTAKELARVAQSSGLDFVLSLGDNFYYDGVKDVDDT 105
Query: 102 TFYESFVNIYTAPSLQKI-WYSVLGNHDYRGDVDAQLSSILRQKDSRWVCLRSFILDGGI 160
F S+ +++ P+L I WY + GNHD+RG+V AQ++ R + RW+ +
Sbjct: 106 RFKFSYEQVFSHPALMTIPWYLIAGNHDHRGNVSAQIAYSSRSE--RWIYPDLYYE---- 159
Query: 161 VEFFFVDTTPFIEKYFTDPKE---HTYDWNGVLPRESYRA--ELLKNVDLALVKSKAKWK 215
+ F + + TD +TYD + E A + L ++ L +KA +
Sbjct: 160 LNFKVPHSNTSLTILMTDTVVVCGNTYDGLDPVGPEDLAAANKQLAWIEQRLQSTKADFV 219
Query: 216 IVVGHHTIKSVGHHGNTQELEQQLLPILKSNNIDAYINGHDHCLQ 260
IVVGH+ I S+GHHG T+ L +L P+LK N+ Y++GHDH LQ
Sbjct: 220 IVVGHYPIWSIGHHGPTKCLISKLRPLLKKYNVSLYLSGHDHSLQ 264
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.327 0.141 0.450
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 567,244,163
Number of Sequences: 2790947
Number of extensions: 24308657
Number of successful extensions: 68796
Number of sequences better than 10.0: 140
Number of HSP's better than 10.0 without gapping: 101
Number of HSP's successfully gapped in prelim test: 39
Number of HSP's that attempted gapping in prelim test: 68540
Number of HSP's gapped (non-prelim): 177
length of query: 322
length of database: 848,049,833
effective HSP length: 127
effective length of query: 195
effective length of database: 493,599,564
effective search space: 96251914980
effective search space used: 96251914980
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 75 (33.5 bits)
Medicago: description of AC146550.4


