

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
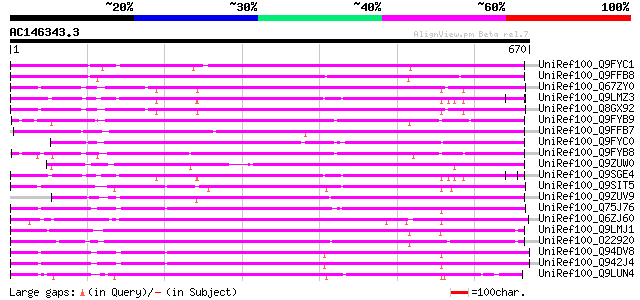
Query= AC146343.3 - phase: 0
(670 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9FYC1 Hypothetical protein F28D10_90 [Arabidopsis tha... 432 e-119
UniRef100_Q9FFB8 Na+/H+ antiporter-like protein [Arabidopsis tha... 412 e-113
UniRef100_Q67ZY0 Hypothetical protein At1g08140 [Arabidopsis tha... 356 1e-96
UniRef100_Q9LMZ3 T6D22.24 [Arabidopsis thaliana] 353 7e-96
UniRef100_Q8GX92 Hypothetical protein At1g08140/T6D22_24 [Arabid... 353 9e-96
UniRef100_Q9FYB9 Hypothetical protein F28D10_110 [Arabidopsis th... 348 2e-94
UniRef100_Q9FFB7 Na+/H+ antiporter-like protein [Arabidopsis tha... 348 2e-94
UniRef100_Q9FYC0 Hypothetical protein F28D10_100 [Arabidopsis th... 348 3e-94
UniRef100_Q9FYB8 Hypothetical protein F28D10_120 [Arabidopsis th... 345 2e-93
UniRef100_Q9ZUW0 Hypothetical protein At2g28180 [Arabidopsis tha... 344 4e-93
UniRef100_Q9SGE4 T23G18.2 [Arabidopsis thaliana] 341 5e-92
UniRef100_Q9SIT5 Putative Na/H antiporter [Arabidopsis thaliana] 333 1e-89
UniRef100_Q9ZUV9 Hypothetical protein At2g28170 [Arabidopsis tha... 332 3e-89
UniRef100_Q75J76 Putative cation/hydrogen exchanger [Oryza sativa] 301 3e-80
UniRef100_Q6ZJ60 Putative cation/hydrogen exchanger [Oryza sativa] 301 4e-80
UniRef100_Q9LMJ1 F10K1.31 protein [Arabidopsis thaliana] 301 5e-80
UniRef100_O22920 Putative Na/H antiporter [Arabidopsis thaliana] 298 3e-79
UniRef100_Q94DV8 Na+/H+ antiporter-like protein [Oryza sativa] 295 3e-78
UniRef100_Q942J4 B1148D12.14 protein [Oryza sativa] 295 3e-78
UniRef100_Q9LUN4 Na+/H+ exchangeing protein-like [Arabidopsis th... 259 2e-67
>UniRef100_Q9FYC1 Hypothetical protein F28D10_90 [Arabidopsis thaliana]
Length = 817
Score = 432 bits (1111), Expect = e-119
Identities = 260/686 (37%), Positives = 389/686 (55%), Gaps = 31/686 (4%)
Query: 1 MDFTMITRTGRKAWTIAFCSFVIPMFFGLVVCYRFQEFWKLEMGNFEAKNLPVIVIGQSG 60
MD ++I TGRKA I S ++ + ++ + + G +I I
Sbjct: 137 MDLSLIRSTGRKAVAIGLSSVLLSITVCALIFFLILRDVGTKKGEPVMSFFEIIFIYLIQ 196
Query: 61 CY--FAVIASLLSDLEILNSELGRLALSTAMVMDSFNSIVTGIGTAFISSIKTDSHDNGD 118
C F VI +LL +L + NSELGRLA+S+A++ D SI++ + F+ +K D G
Sbjct: 197 CLSSFPVIGNLLFELRLQNSELGRLAMSSAVISDFSTSILSAV-LVFLKELKDDKSRLGS 255
Query: 119 --------GKGTLKAFLNVFYYLCFMVVTPLVLRPILKWFVKKTPEGRPMKKVYMYIVFI 170
G +K V ++CF + + RP++ + +K+TP GRP+KK Y+Y + I
Sbjct: 256 VFIGDVIVGNRPMKRAGTVVLFVCFAIY---IFRPLMFFIIKRTPSGRPVKKFYIYAIII 312
Query: 171 IALAVGMLGLLTKQSVLGGICIVGLIVPEGPPLGTEMIKQLELFCSWFLFPIFVTSCAMK 230
+ +L KQS+ G I+GL VP GPPLG+ ++++ E P FV + A +
Sbjct: 313 LVFGSAILADWCKQSIFIGPFILGLAVPHGPPLGSAILQKFESVVFGTFLPFFVATSAEE 372
Query: 231 IDLSVY-----VKSDYIYVWLGIIVAVHLFKMLVTIGICWYCNMPMADGLCLALMLSCKG 285
ID S+ +KS I V + IV K +T + MP D + L+L++S KG
Sbjct: 373 IDTSILQSWIDLKSIVILVSVSFIV-----KFALTTLPAFLYGMPAKDCIALSLIMSFKG 427
Query: 286 VVDFCNNVFLHDSMLLSSEALSVLSINVLVIGTLARIGVKYLYDPSRKYAGYQKRNILSL 345
+ +F + + + +VLS+ +L+ + +K +YDPSR YAGY+KRN+L +
Sbjct: 428 IFEFGAYGYAYQRGTIRPVTFTVLSLYILLNSAVIPPLLKRIYDPSRMYAGYEKRNMLHM 487
Query: 346 KPNSELKIVSCILKPSHIIPIKNVLDICSPTSSNPLVIHILHLLELVGRSSPVFISHRLQ 405
KPNSEL+I+SCI K I P+ N+L+ P+ NP+ ++LHL+ELVG+++PV ISHRLQ
Sbjct: 488 KPNSELRILSCIYKTDDIRPMINLLEATCPSRENPVATYVLHLMELVGQANPVLISHRLQ 547
Query: 406 ERVGSSSHTFSEAVIVTFDLFEHDNAGTASVSTYTAISPVRFMHDDICYLALDKLASIII 465
R + SE V+V+F+ F +D G+ VSTYTA+S + MH DIC LAL+ S+II
Sbjct: 548 TRKSENMSYNSENVVVSFEQFHNDFFGSVFVSTYTALSVPKMMHGDICMLALNNTTSLII 607
Query: 466 LPFHLRWSEDGSVESADE-TTRSLNTKVLERAPCSVAILVNRGHSSPFNHNE-----NSK 519
LPFH WS DGS +D R LN VL+ +PCSV I V R + E +S
Sbjct: 608 LPFHQTWSADGSAIVSDSLMIRQLNKSVLDLSPCSVGIFVYRSSNGRRTIKETAANFSSY 667
Query: 520 QIAMIFLGGSDDREALCLAKRTIKEDTYHLVVYHLVSTIKN-DEFTSWDVMLDDELLKGV 578
Q+ M+FLGG DDREAL LAKR ++ + V L+S+ + ++ T WD MLD ELL+ V
Sbjct: 668 QVCMLFLGGKDDREALSLAKRMARDSRITITVVSLISSEQRANQATDWDRMLDLELLRDV 727
Query: 579 KGVYGSVDNVTYEKVEVENTSDTTEFISDIAIQHDFIIVGRRNGIKSPQTQALASWTEYP 638
K + ++ + + V + + T++ + IA ++D IVGR G KS T+ L W+E+
Sbjct: 728 KSNVLAGADIVFSEEVVNDANQTSQLLKSIANEYDLFIVGREKGRKSVFTEGLEEWSEFE 787
Query: 639 ELGVLGDLLASPDTNTKASILVVQQQ 664
ELG++GDLL S D N +AS+LV+QQQ
Sbjct: 788 ELGIIGDLLTSQDLNCQASVLVIQQQ 813
>UniRef100_Q9FFB8 Na+/H+ antiporter-like protein [Arabidopsis thaliana]
Length = 822
Score = 412 bits (1060), Expect = e-113
Identities = 248/685 (36%), Positives = 383/685 (55%), Gaps = 25/685 (3%)
Query: 1 MDFTMITRTGRKAWTIAFCSFVIPMFFGLVVCYRFQEFWKLEMGNFEAKNLPVIVIGQSG 60
MD +I TGRKA TI S ++ V+ + + + +L +VI
Sbjct: 138 MDTGLIRTTGRKAITIGLSSVLLSTLVCSVIFFGNLRDVGTKNSDHTLNSLEYVVIYSIQ 197
Query: 61 CY--FAVIASLLSDLEILNSELGRLALSTAMVMDSFNSIVTGIGTAFISSIKTD-----S 113
C F V+ +LL +L + NSELGRLA+S+A++ D SI+ + F+ +K + S
Sbjct: 198 CLSSFPVVGNLLFELRLQNSELGRLAISSAVISDFSTSILASV-LIFMKELKDEQTRLGS 256
Query: 114 HDNGDGKGTLKAFLNVFYYLCFMVVTPLVLRPILKWFVKKTPEGRPMKKVYMYIVFIIAL 173
GD + + + F+ + V RP++ + +K+TP GRP+K +Y+ + ++
Sbjct: 257 VFIGDVIAGNRPLMRAGIVVLFVCIAIYVFRPLMFYIIKQTPSGRPVKAIYLSTIIVMVS 316
Query: 174 AVGMLGLLTKQSVLGGICIVGLIVPEGPPLGTEMIKQLELFCSWFLFPIFVTSCAMKIDL 233
+L KQS+ G I+GL VP GPPLG+ +I++ E P F+ S + +ID+
Sbjct: 317 GSAILANWCKQSIFMGPFILGLAVPHGPPLGSAIIQKYESAIFGTFLPFFIASSSTEIDI 376
Query: 234 SVYVKSDYIYVWLGIIVAVHLFKMLVTIGICWYCNMPMADGLCLALMLSCKGVVDFCNNV 293
S + + + I+V + K + T + MPM D L+L++S KG+ +
Sbjct: 377 SALFGWEGLNGIILIMVTSFVVKFIFTTVPALFYGMPMEDCFALSLIMSFKGIFELGAYA 436
Query: 294 FLHDSMLLSSEALSVLSINVLVIGTLARIGVKYLYDPSRKYAGYQKRNILSLKPNSELKI 353
+ + E +V + + + + ++YLYDPSR YAGY+KRN+ LKPNSEL+I
Sbjct: 437 LAYQRGSVRPETFTVACLYITLNSAIIPPILRYLYDPSRMYAGYEKRNMQHLKPNSELRI 496
Query: 354 VSCILKPSHIIPIKNVLDICSPTSSNPLVIHILHLLELVGRSSPVFISHRLQERVGSSSH 413
+SCI + I P+ N+L+ P+ +P+ ++LHL+ELVG+++P+FISH+LQ R +
Sbjct: 497 LSCIYRTDDISPMINLLEAICPSRESPVATYVLHLMELVGQANPIFISHKLQTR-RTEET 555
Query: 414 TFSEAVIVTFDLFEHDNAGTASVSTYTAISPVRFMHDDICYLALDKLASIIILPFHLRWS 473
++S V+V+F+ F D G+ VSTYTA+S MH DIC LAL+ S+I+LPFH WS
Sbjct: 556 SYSNNVLVSFEKFRKDFYGSVFVSTYTALSMPDTMHGDICMLALNNTTSLILLPFHQTWS 615
Query: 474 EDGS-VESADETTRSLNTKVLERAPCSVAILVNRGHSSPFN------------HNENSKQ 520
DGS + S + R+LN VL+ APCSV + V R S N N +S
Sbjct: 616 ADGSALISNNNMIRNLNKSVLDVAPCSVGVFVYRSSSGRKNISSGRKTINGTVPNLSSYN 675
Query: 521 IAMIFLGGSDDREALCLAKRTIKEDTYHLVVYHLVST-IKNDEFTSWDVMLDDELLKGVK 579
I MIFLGG DDREA+ LA R ++ ++ + L++T K E T WD MLDDELL+ VK
Sbjct: 676 ICMIFLGGKDDREAVTLATRMARDPRINITIVRLITTDEKARENTVWDKMLDDELLRDVK 735
Query: 580 GVYGSVDNVTYEKVEVENTSDTTEFISDIAIQHDFIIVGRRNGIKSPQTQALASWTEYPE 639
++ ++ Y + +E+ ++T+ + + D IVGR NG S T+ L W+E+ E
Sbjct: 736 S--NTLVDIFYSEKAIEDAAETSSLLRSMVSDFDMFIVGRGNGRTSVFTEGLEEWSEFKE 793
Query: 640 LGVLGDLLASPDTNTKASILVVQQQ 664
LG++GDLL S D N +AS+LV+QQQ
Sbjct: 794 LGIIGDLLTSQDFNCQASVLVIQQQ 818
>UniRef100_Q67ZY0 Hypothetical protein At1g08140 [Arabidopsis thaliana]
Length = 818
Score = 356 bits (914), Expect = 1e-96
Identities = 234/686 (34%), Positives = 368/686 (53%), Gaps = 44/686 (6%)
Query: 1 MDFTMITRTGRKAWTIAFCSFVIPMFFGLVVCYRFQEFWKLEMGNFEAKNLPVIVIGQSG 60
MD RTG+++ I F + +IP+ G + +R++E + + +I+ QS
Sbjct: 143 MDAESPFRTGKRSSVIGFITVIIPLICGSLT-FRYRE--RRGDSSILRMEYRLIIFLQSI 199
Query: 61 CYFAVIASLLSDLEILNSELGRLALSTAMVMDSFNSIVTGIGTAFISSIKTDSHDNGDGK 120
F I +LL DL+I +SE GR+ALS AMV D + G F ++I +
Sbjct: 200 SAFTSIDTLLKDLQIKHSEFGRIALSGAMVTD-----MLAFGVTFFNAIYYEK------- 247
Query: 121 GTLKAFLNVFYYLCFMVVTPLVLRPILKWFVKKTPEGRPMKKVYMYIVFIIALAVGMLGL 180
L F+ + F+VV V+RP + W +K+TPEGRP+K Y+Y +F IA A
Sbjct: 248 --LYGFMQTVGFCLFVVVMICVVRPAMYWVIKQTPEGRPVKDFYLYSIFGIAFAC--FTF 303
Query: 181 LTKQSVL---GGICIVGLIVPEGPPLGTEMIKQLELFCSWFLFPIFVTSCAMKIDLSVYV 237
K L G + GL VP G PLGT +I++ E F + P+F + M++DL
Sbjct: 304 FNKVIHLFGPAGSFVFGLTVPNGYPLGTTLIQKFESFNLGSILPLFGSLTMMQVDLLRLF 363
Query: 238 KSD--------YIYVWLGIIVAVHLFKMLVTIGICWYCNMPMADGLCLALMLSCKGVVDF 289
K IY + I+ V+ K +VT + MP+ D LAL+LS KG+ +
Sbjct: 364 KESGDLIRMEGQIYEVISFILLVNTTKFVVTTITAYAFKMPLRDSFALALVLSNKGIFEL 423
Query: 290 CNNVFLHDSMLLSSEALSVLSINVLVIGTLARIGVKYLYDPSRKYAGYQKRNILSLKPNS 349
+ + L+ E ++L+ L+ + ++ ++DP++++ Y+KRN+ LK +
Sbjct: 424 AYYTYAVELKLIRPEVFTILAAYTLLNSIFIPMLLELVHDPTKRFRCYRKRNLGILKDGA 483
Query: 350 ELKIVSCILKPSHIIPIKNVLDICSPTSSNPLVIHILHLLELVGRSSPVFISHRLQERVG 409
L+ + C+ +P HI + ++L+ SP+ +P+ +ILHL+ELVG+++P+FISH+LQ+
Sbjct: 484 ALQCIMCVYRPDHITSMTDLLETFSPSQDSPMACNILHLVELVGQANPMFISHQLQKPEP 543
Query: 410 SSSHTFSEAVIVTFDLFEHDNAGTASVSTYTAISPVRFMHDDICYLALDKLASIIILPFH 469
S+ + S+ VI++F F+ S+ +T++S + MH+DIC+LAL + S+I+LPFH
Sbjct: 544 GST-SLSDNVIISFRGFQRQFFEYTSLDIFTSVSVSQHMHEDICWLALSRSLSLIVLPFH 602
Query: 470 LRWSEDGS-VESADETTRSLNTKVLERAPCSVAILVNRGHSSPFNHNENSKQIAMIFLGG 528
WS D S V S D+ R LN VL RAPCSV I V R + ++ +I +IF GG
Sbjct: 603 RTWSVDRSTVISNDDNLRMLNVNVLRRAPCSVGIFVYRKPIVESHMAKSHSKICLIFNGG 662
Query: 529 SDDREALCLAKR-TIKEDTYHLVVYHLV---STIKNDEFTSWDVMLDDELLKGVKGVYGS 584
DDREAL + R + E L + + S + NDE W+ L + V + GS
Sbjct: 663 KDDREALAITNRMRLTEKRTRLTIIRFIPKSSEMDNDE---WEQQQSINLKESVTSIVGS 719
Query: 585 -----VDNVTYEKVEVENTSDTTEFISDIAIQHDFIIVGRRNGIKSPQTQALASWTEYPE 639
VTY V + S+T+ + +A +D IVG +GI + T ++ WTE+ E
Sbjct: 720 NIKENDAKVTYIDKAVSDGSETSRILRAMANDYDLFIVGSGSGIGTEATSGISEWTEFNE 779
Query: 640 LGVLGDLLASPDTNTKASILVVQQQV 665
LG +GDLLAS + + AS+LVVQ+QV
Sbjct: 780 LGPIGDLLASHEYPSSASVLVVQKQV 805
>UniRef100_Q9LMZ3 T6D22.24 [Arabidopsis thaliana]
Length = 2621
Score = 353 bits (907), Expect = 7e-96
Identities = 233/686 (33%), Positives = 368/686 (52%), Gaps = 44/686 (6%)
Query: 1 MDFTMITRTGRKAWTIAFCSFVIPMFFGLVVCYRFQEFWKLEMGNFEAKNLPVIVIGQSG 60
MD + RT +++ I F + +IP+ G + +R++E + + +I+ QS
Sbjct: 1130 MDAELPFRTEKRSSVIGFITVIIPLICGSLT-FRYRE--RRGDSSILRMEYRLIIFLQSI 1186
Query: 61 CYFAVIASLLSDLEILNSELGRLALSTAMVMDSFNSIVTGIGTAFISSIKTDSHDNGDGK 120
F I +LL DL+I +SE GR+ALS AMV D + G F ++I +
Sbjct: 1187 SAFTSIDTLLKDLQIKHSEFGRIALSGAMVTD-----MLAFGVTFFNAIYYEK------- 1234
Query: 121 GTLKAFLNVFYYLCFMVVTPLVLRPILKWFVKKTPEGRPMKKVYMYIVFIIALAVGMLGL 180
L F+ + F+VV V+RP + W +K+TPEGRP+K Y+Y +F IA A
Sbjct: 1235 --LYGFMQTVGFCLFVVVMICVVRPAMYWVIKQTPEGRPVKDFYLYSIFGIAFAC--FTF 1290
Query: 181 LTKQSVL---GGICIVGLIVPEGPPLGTEMIKQLELFCSWFLFPIFVTSCAMKIDLSVYV 237
K L G + GL VP G PLGT +I++ E F + P+F + M++DL
Sbjct: 1291 FNKVIHLFGPAGSFVFGLTVPNGYPLGTTLIQKFESFNLGSILPLFGSLTMMQVDLLRLF 1350
Query: 238 KSD--------YIYVWLGIIVAVHLFKMLVTIGICWYCNMPMADGLCLALMLSCKGVVDF 289
K IY + I+ V+ K +VT + MP+ D LAL+LS KG+ +
Sbjct: 1351 KESGDLIRMEGQIYEVISFILLVNTTKFVVTTITAYAFKMPLRDSFALALVLSNKGIFEL 1410
Query: 290 CNNVFLHDSMLLSSEALSVLSINVLVIGTLARIGVKYLYDPSRKYAGYQKRNILSLKPNS 349
+ + L+ E ++L+ L+ + ++ ++DP++++ Y+KRN+ LK +
Sbjct: 1411 AYYTYAVELKLIRPEVFTILAAYTLLNSIFIPMLLELVHDPTKRFRCYRKRNLGILKDGA 1470
Query: 350 ELKIVSCILKPSHIIPIKNVLDICSPTSSNPLVIHILHLLELVGRSSPVFISHRLQERVG 409
L+ + C+ +P HI + ++L+ SP+ +P+ +ILHL+ELVG+++P+FISH+LQ+
Sbjct: 1471 ALQCLMCVYRPDHITSMTDLLETFSPSQDSPMACNILHLVELVGQANPMFISHQLQKPEP 1530
Query: 410 SSSHTFSEAVIVTFDLFEHDNAGTASVSTYTAISPVRFMHDDICYLALDKLASIIILPFH 469
S+ + S+ VI++F F+ S+ +T++S + MH+DIC+LAL + S+I+LPFH
Sbjct: 1531 GST-SLSDNVIISFRGFQRQFFEYTSLDIFTSVSVSQHMHEDICWLALSRSLSLIVLPFH 1589
Query: 470 LRWSEDGS-VESADETTRSLNTKVLERAPCSVAILVNRGHSSPFNHNENSKQIAMIFLGG 528
WS D S V S D+ R LN VL RAPCSV I V R + ++ +I +IF GG
Sbjct: 1590 RTWSVDRSTVISNDDNLRMLNVNVLRRAPCSVGIFVYRKPIVESHMAKSHSKICLIFNGG 1649
Query: 529 SDDREALCLAKR-TIKEDTYHLVVYHLV---STIKNDEFTSWDVMLDDELLKGVKGVYGS 584
DDREAL + R + E L + + S + NDE W+ L + V + GS
Sbjct: 1650 KDDREALAITNRMRLTEKRTRLTIIRFIPKSSEMDNDE---WEQQQSINLKESVTSIVGS 1706
Query: 585 -----VDNVTYEKVEVENTSDTTEFISDIAIQHDFIIVGRRNGIKSPQTQALASWTEYPE 639
VTY V + S+T+ + +A +D IVG +GI + T ++ WTE+ E
Sbjct: 1707 NIKENDAKVTYIDKAVSDGSETSRILRAMANDYDLFIVGSGSGIGTEATSGISEWTEFNE 1766
Query: 640 LGVLGDLLASPDTNTKASILVVQQQV 665
LG +GDLLAS + + AS+LVVQ+QV
Sbjct: 1767 LGPIGDLLASHEYPSSASVLVVQKQV 1792
Score = 341 bits (874), Expect = 5e-92
Identities = 215/653 (32%), Positives = 348/653 (52%), Gaps = 31/653 (4%)
Query: 1 MDFTMITRTGRKAWTIAFCSFVIPMFFGLVVCYRFQEFWKLEMGNFEAKNLPVIVIGQSG 60
MD M+ +TG K + ++P+ +V + +E + E + I+ QS
Sbjct: 356 MDVGMVRKTGTKVIVTGIATVILPIIAANMVFGKLRETGGKYLTGMEYRT---ILFMQSI 412
Query: 61 CYFAVIASLLSDLEILNSELGRLALSTAMVMDSFNSIVTGIGTAFISSIKTDSHDNGDGK 120
F I+ LL DL I +SE GR+ +STAMV D TG G + + D +
Sbjct: 413 SAFTGISRLLRDLRINHSEFGRIVISTAMVADG-----TGFGVNLFALVAWM-----DWR 462
Query: 121 GTLKAFLNVFYYLCFMVVTPLVLRPILKWFVKKTPEGRPMKKVYMYIVFIIALAVGMLGL 180
+ + + Y+ FMV V+RP + W +K+TP+ RP+K+ ++YI+ I+A
Sbjct: 463 VSALQGVGIIGYVIFMV---WVVRPAMFWVIKRTPQERPVKECFIYIILILAFGGYYFLK 519
Query: 181 LTKQSVLGGICIVGLIVPEGPPLGTEMIKQLELFCSWFLFPIFVTSCAMKIDLSVYVKS- 239
G ++GL VP GPPLG++++++ E F + L P+F+ ++ID
Sbjct: 520 EIHMFPAVGPFLLGLCVPHGPPLGSQLVEKFESFNTGILLPLFLFFSMLQIDGPWLANQI 579
Query: 240 -------DYIYVWLGIIVAVHLFKMLVTIGICWYCNMPMADGLCLALMLSCKGVVDFCNN 292
+Y L II+ V + K++ ++ MP+ D +AL+LS KG+V+ C
Sbjct: 580 GQLRHFDGQLYEALTIIIVVFVAKIIFSMIPALLAKMPLTDSFVMALILSNKGIVELCYF 639
Query: 293 VFLHDSMLLSSEALSVLSINVLVIGTLARIGVKYLYDPSRKYAGYQKRNILSLKPNSELK 352
++ +S +L ++ ++++ +LV T++ + + YLYD S+++ +QKRN++SLK SELK
Sbjct: 640 LYGVESNVLHVKSFTIMATMILVSSTISPVLIHYLYDSSKRFISFQKRNLMSLKLGSELK 699
Query: 353 IVSCILKPSHIIPIKNVLDICSPTSSNPLVIHILHLLELVGRSSPVFISHRLQERVGSSS 412
+ CI K HI + N+L P + + +++HL+ELVG +PVFISH++Q + +
Sbjct: 700 FLVCIHKADHISGMINLLAQSFPLHESTISCYVIHLVELVGLDNPVFISHQMQ-KAEPGN 758
Query: 413 HTFSEAVIVTFDLFEHDNAGTASVSTYTAISPVRFMHDDICYLALDKLASIIILPFHLRW 472
++S V++ FD F+H + S+ +T IS R+MH +I LALDK AS ++LPFH+ W
Sbjct: 759 RSYSNNVLIAFDNFKH-YWKSISLELFTCISNPRYMHQEIYSLALDKQASFLMLPFHIIW 817
Query: 473 SED-GSVESADETTRSLNTKVLERAPCSVAILVNRGHSSPFNHNENSKQIAMIFLGGSDD 531
S D +V S D R+ N VL +APCSV I V+R + S ++ IF+GG DD
Sbjct: 818 SLDQTTVVSDDVMRRNANLNVLRQAPCSVGIFVHRQKLLSAQKSSPSFEVCAIFVGGKDD 877
Query: 532 REALCLAKRTIKEDTYHLVVYHLVSTIKNDEFTSWDVMLDD----ELLKGVKGVYGSVDN 587
REAL L ++ ++ +L V L+ + T WD MLD E+L+ G
Sbjct: 878 REALALGRQMMRNPNVNLTVLKLIPAKMDGMTTGWDQMLDSAEVKEVLRNNNNTVGQHSF 937
Query: 588 VTYEKVEVENTSDTTEFISDIAIQHDFIIVGRRNGIKSPQTQALASWTEYPEL 640
V Y + V + SDT+ + IA D +VGR G+ + AL+ WTE+ EL
Sbjct: 938 VEYVEETVNDGSDTSTLLLSIANSFDLFVVGRSAGVGTDVVSALSEWTEFDEL 990
Score = 303 bits (775), Expect = 1e-80
Identities = 207/679 (30%), Positives = 352/679 (51%), Gaps = 47/679 (6%)
Query: 2 DFTMITRTGRKAWTIAFCSFVIPMFFGLVVCYRFQEFWKLEMGNFEAKNLPVIVIGQSGC 61
D ++ ++G K+ I S +IP G ++ ++ L M E V+ S
Sbjct: 1973 DVGIMKKSGTKSVVIGITSMIIPWQIGKLLYSSREKSSILTMTEME---YTVMTFTMSMT 2029
Query: 62 YFAVIASLLSDLEILNSELGRLALSTAMVMDSFNSIVTGIGTAFISSIKTDSHDNGDGKG 121
F + LL+DL+I++++ G++A S MV D +T +A++S +T G
Sbjct: 2030 PFTCVNMLLTDLKIVHTDFGQIAQSAGMVTDLLAFFLTV--SAYVSRDETQGVKMG---- 2083
Query: 122 TLKAFLNVFYYLCFMVVTPLVLRPILKWFVKKTPEGRPMKKVYMYIVFIIA-LAVGMLGL 180
AF+ F ++ ++R + W ++ TPEG P+K VY+YI ++A L+
Sbjct: 2084 --LAFMAFFIFV-------YLVRQFMLWVIRHTPEGAPVKNVYLYIGLLLAYLSYLYWSR 2134
Query: 181 LTKQSVLGGICIVGLIVPEGPPLGTEMIKQLELFCSWFLFPIFVTSCAMKIDLSVYVKS- 239
LG + GL VP GPPLG+ I++ + F P+F + +K+D S K
Sbjct: 2135 FLFFGPLGAFAL-GLAVPNGPPLGSVFIQKFDSFNEGIFLPLFGSLSMIKLDWSFLRKEF 2193
Query: 240 -------DYIYVWLGIIVAVHLFKMLVTIGICWYCNMPMADGLCLALMLSCKGVVDFCNN 292
++Y + V++ K + +P+ D + L +++ K +
Sbjct: 2194 GNGRHLHGHMYECFSFLPIVYIAKFATSFLAALATKIPLRDSIILGVIMGTKSSFELGYV 2253
Query: 293 VFLHDSMLLSSEALSVLSINVLVIGTLARIGVKYLYDPSRKYAGYQKRNILSLKPNSELK 352
+ + +S E LS+L + +LV L + + +LYD S+++ Y +RN LK E++
Sbjct: 2254 LTAFEKDRISLEVLSLLGVYILVNSLLTPMAIHFLYDRSKRFVCYGRRN---LKEKPEMQ 2310
Query: 353 IVSCILKPSHIIPIKNVLDICSPTSSNPLVIHILHLLELVGRSSPVFISHRLQERVGSSS 412
+ CI KP +I + ++L SP+ +P+ +LHL+EL+G+++P FISH+LQ + S
Sbjct: 2311 TLVCINKPDNITSMISLLRATSPSKDSPMECCVLHLIELLGQATPTFISHQLQ-KPKPGS 2369
Query: 413 HTFSEAVIVTFDLFEHDNAGTASVSTYTAISPVRFMHDDICYLALDKLASIIILPFHLRW 472
++SE VI +F LF+ +AS++ +T+++ + MH+ IC+ AL + +++I+L FH W
Sbjct: 2370 RSYSENVISSFQLFQEVYWDSASINMFTSLTSAKEMHEQICWFALSQGSNLILLSFHRTW 2429
Query: 473 SEDGSVE-SADETTRSLNTKVLERAPCSVAILVNRGHSSPFNHNENSKQIAMIFLGGSDD 531
+G+V S D+T RSLN VL+RAPCSV I V R E+ ++ +I++GG+DD
Sbjct: 2430 EPNGNVIISDDQTLRSLNLNVLKRAPCSVGIFVYRKPIWQTKALESPCRVCLIYVGGNDD 2489
Query: 532 REALCLAKRTIKEDTYHLVVYHLVSTIKNDEFT------SWDVMLDDELLKGVKGVYGSV 585
+EAL LA L V L+ T DE + D+ ++ G
Sbjct: 2490 KEALALADHMRGNQQVILTVLRLIPTSYADESSLRIHSQMVDMNRHEDQRPG-------- 2541
Query: 586 DNVTYEKVEVENTSDTTEFISDIAIQHDFIIVGRRNGIKSPQTQALASWTEYPELGVLGD 645
D T V + ++T++ + ++ +D IVGRR+G+ + T+ L W E+ ELGV+GD
Sbjct: 2542 DKSTIIDWTVGDGTETSKILHSVSYDYDLFIVGRRSGVGTTVTRGLGDWMEFEELGVIGD 2601
Query: 646 LLASPDTNTKASILVVQQQ 664
LLAS ++AS+LVVQQQ
Sbjct: 2602 LLASEYFPSRASVLVVQQQ 2620
>UniRef100_Q8GX92 Hypothetical protein At1g08140/T6D22_24 [Arabidopsis thaliana]
Length = 818
Score = 353 bits (906), Expect = 9e-96
Identities = 233/686 (33%), Positives = 367/686 (52%), Gaps = 44/686 (6%)
Query: 1 MDFTMITRTGRKAWTIAFCSFVIPMFFGLVVCYRFQEFWKLEMGNFEAKNLPVIVIGQSG 60
MD RT +++ I F + +IP+ G + +R++E + + +I+ QS
Sbjct: 143 MDAESPFRTEKRSSVIGFITVIIPLICGSLT-FRYRE--RRGDSSILRMEYRLIIFLQSI 199
Query: 61 CYFAVIASLLSDLEILNSELGRLALSTAMVMDSFNSIVTGIGTAFISSIKTDSHDNGDGK 120
F I +LL DL+I +SE GR+ALS AMV D + G F ++I +
Sbjct: 200 SAFTSIDTLLKDLQIKHSEFGRIALSGAMVTD-----MLAFGVTFFNAIYYEK------- 247
Query: 121 GTLKAFLNVFYYLCFMVVTPLVLRPILKWFVKKTPEGRPMKKVYMYIVFIIALAVGMLGL 180
L F+ + F+VV V+RP + W +K+TPEGRP+K Y+Y +F IA A
Sbjct: 248 --LYGFMQTVGFCLFVVVMICVVRPAMYWVIKQTPEGRPVKDFYLYSIFGIAFAC--FTF 303
Query: 181 LTKQSVL---GGICIVGLIVPEGPPLGTEMIKQLELFCSWFLFPIFVTSCAMKIDLSVYV 237
K L G + GL VP G PLGT +I++ E F + P+F + M++DL
Sbjct: 304 FNKVIHLFGPAGSFVFGLTVPNGYPLGTTLIQKFESFNLGSILPLFGSLTMMQVDLLRLF 363
Query: 238 KSD--------YIYVWLGIIVAVHLFKMLVTIGICWYCNMPMADGLCLALMLSCKGVVDF 289
K IY + I+ V+ K +VT + MP+ D LAL+LS KG+ +
Sbjct: 364 KESGDLIRMEGQIYEVISFILLVNTTKFVVTTITAYAFKMPLRDSFALALVLSNKGIFEL 423
Query: 290 CNNVFLHDSMLLSSEALSVLSINVLVIGTLARIGVKYLYDPSRKYAGYQKRNILSLKPNS 349
+ + L+ E ++L+ L+ + ++ ++DP++++ Y+KRN+ LK +
Sbjct: 424 AYYTYAVELKLIRPEVFTILAAYTLLNSIFIPMLLELVHDPTKRFRCYRKRNLGILKDGA 483
Query: 350 ELKIVSCILKPSHIIPIKNVLDICSPTSSNPLVIHILHLLELVGRSSPVFISHRLQERVG 409
L+ + C+ +P HI + ++L+ SP+ +P+ +ILHL+ELVG+++P+FISH+LQ+
Sbjct: 484 ALQCIMCVYRPDHITSMTDLLETFSPSQDSPMACNILHLVELVGQANPMFISHQLQKPEP 543
Query: 410 SSSHTFSEAVIVTFDLFEHDNAGTASVSTYTAISPVRFMHDDICYLALDKLASIIILPFH 469
S+ + S+ VI++F F+ S+ +T++S + MH+DIC+LAL + S+I+LPFH
Sbjct: 544 GST-SLSDNVIISFRGFQRQFFEYTSLDIFTSVSVSQHMHEDICWLALSRSLSLIVLPFH 602
Query: 470 LRWSEDGS-VESADETTRSLNTKVLERAPCSVAILVNRGHSSPFNHNENSKQIAMIFLGG 528
WS D S V S D+ R LN VL RAPCSV I V R + ++ +I +IF GG
Sbjct: 603 RTWSVDRSTVISNDDNLRMLNVNVLRRAPCSVGIFVYRKPIVESHMAKSHSKICLIFNGG 662
Query: 529 SDDREALCLAKR-TIKEDTYHLVVYHLV---STIKNDEFTSWDVMLDDELLKGVKGVYGS 584
DDREAL + R + E L + + S + NDE W+ L + V + GS
Sbjct: 663 KDDREALAITNRMRLTEKRTRLTIIRFIPKSSEMDNDE---WEQQQSINLKESVTSIVGS 719
Query: 585 -----VDNVTYEKVEVENTSDTTEFISDIAIQHDFIIVGRRNGIKSPQTQALASWTEYPE 639
VTY V + S+T+ + +A +D IVG +GI + T ++ WTE+ E
Sbjct: 720 NIKENDAKVTYIDKAVSDGSETSRILRAMANDYDLFIVGSGSGIGTEATSGISEWTEFNE 779
Query: 640 LGVLGDLLASPDTNTKASILVVQQQV 665
LG +GDLLAS + + AS+LVVQ+QV
Sbjct: 780 LGPIGDLLASHEYPSSASVLVVQKQV 805
>UniRef100_Q9FYB9 Hypothetical protein F28D10_110 [Arabidopsis thaliana]
Length = 671
Score = 348 bits (894), Expect = 2e-94
Identities = 218/683 (31%), Positives = 366/683 (52%), Gaps = 40/683 (5%)
Query: 3 FTMITRTGRKAWTIAFCSFVIPMFFGLVVCYRFQEFWKLEMGNFEAKNLP---------- 52
F M RT R+ +AF S +P+ G+V F + L N N+
Sbjct: 4 FLMTVRTSRR---VAFHSGKLPVVIGIVSF--FAPLFSLSFLNLFTDNIDPHYMSLDKAL 58
Query: 53 ----VIVIGQSGCYFAVIASLLSDLEILNSELGRLALSTAMVMDSFNSIVTGIGTAFISS 108
VIVI QS +L +L+I+NSELGRLALS + + D + T +
Sbjct: 59 AERTVIVITQSQILLPSTTYILLELKIINSELGRLALSASAINDMLGIFAMIVATTQATY 118
Query: 109 IKTDSHDNGDGKGTLKAFLNVFYYLCFMVVTPLVLRPILKWFVKKTPEGRPMKKVYMYIV 168
I SH A+ ++ + F ++ V +P+++W + +TPE +P++ +Y++ V
Sbjct: 119 IHV-SH--------AIAYRDLVAVIIFFLIVFFVFKPMVQWIIDRTPEDKPVEDIYIHAV 169
Query: 169 FIIALAVGMLGLLTKQSVLGGICIVGLIVPEGPPLGTEMIKQLELFCSWFLFPIFVTSCA 228
+ A A + + G I+G+I+PEGPPLG+ + + E PI +T A
Sbjct: 170 ILTAFASAAYFVFFNMKYVLGPLIIGIIIPEGPPLGSALEAKFERLTMNVFLPISITFSA 229
Query: 229 MKID-LSVYVKSDYIYVWLGIIVAVHLFKMLVTIGICWYCNMPMADGLCLALMLSCKGVV 287
M+ D L + + IY + + + + + K++ + +C Y +P ++ L ++L+LS K V
Sbjct: 230 MRCDGLRILSQFTDIYFNIFLTLLILVIKLVACLTLCLYYKLPRSESLAVSLILSYKSFV 289
Query: 288 DFCNNVFLHDSMLLSSEALSVLSINVLVIGTLARIGVKYLYDPSRKYAGYQKRNILSLKP 347
+F + + +S + L + L+ + + V+ +YDP RKY YQKR+IL L+
Sbjct: 290 EFVLYEAVLEEKFISQATYAFLILYSLLSAGIVPMVVRSMYDPKRKYVNYQKRDILHLEA 349
Query: 348 NSELKIVSCILKPSHIIPIKNVLDI-CSPTSSNPLVIHILHLLELVGRSSPVFISH-RLQ 405
NS L+I++C+ KP ++ L + SP P+ + +LHL++LVG+ +P+ +SH +
Sbjct: 350 NSGLRILTCLHKPENVSETIAFLQLFSSPIHDFPIAVTVLHLVKLVGQINPIIVSHDKKL 409
Query: 406 ERVGSSSHTFSEAVIVTFDLFEHDNAGTASVSTYTAISPVRFMHDDICYLALDKLASIII 465
+R+ +S+ + + F F ++ + +V+T+TA S MH+DIC LALD+ S+I+
Sbjct: 410 KRLHKNSYIHTANL--AFRQFMQESLESVTVTTFTAFSHENLMHEDICTLALDRTTSMIV 467
Query: 466 LPFHLRWSEDGSVESADETTRSLNTKVLERAPCSVAILVNRGHSSPFNH----NENSKQI 521
+P +W+ DG ES D R LN +L+RAPCS+ ILV+RG S ++ N + +
Sbjct: 468 VPSGRKWTVDGMFESDDLAARQLNQSLLDRAPCSIGILVDRGQFSRKSYVTSKNRYNIDV 527
Query: 522 AMIFLGGSDDREALCLAKRTIKEDTYHLVVYHLVSTIKNDEFTSWDVMLDDELLKGVKGV 581
++F+GG DDREAL L KR + V L+ ++ + WD +LD+E LK +K
Sbjct: 528 GVLFIGGKDDREALSLVKRMKYNPRVRVTVIRLI--FDHEIESEWDYILDNEGLKDLKST 585
Query: 582 YGSVDNVTYEKVEVENTSDTTEFISDIAIQHDFIIVGRRNGIKSPQTQALASWTEYPELG 641
+ D + E++ V + + + + +A ++D ++VGR + + S L W E PELG
Sbjct: 586 ESNEDILYTERI-VTSVVEVVKAVQLLAEEYDLMVVGRDHDMTSQDLSGLTEWVELPELG 644
Query: 642 VLGDLLASPDTNTKASILVVQQQ 664
V+GDLLA+ D N+K S+LVVQQQ
Sbjct: 645 VIGDLLAARDLNSKVSVLVVQQQ 667
>UniRef100_Q9FFB7 Na+/H+ antiporter-like protein [Arabidopsis thaliana]
Length = 800
Score = 348 bits (894), Expect = 2e-94
Identities = 224/671 (33%), Positives = 354/671 (52%), Gaps = 23/671 (3%)
Query: 6 ITRTGRKAWTIAFCSFVIPMFFGLVVC-YRFQEFWKLEMG-NFEAKNLPVIVIGQSGCYF 63
+ + G++ IA SF + M GL +R + L M VIV Q+
Sbjct: 137 VYQIGKRPIVIAVSSFFVTMISGLAFRNFRLDKVDPLYMPLRLAPTERSVIVSIQAVTLL 196
Query: 64 AVIASLLSDLEILNSELGRLALSTAMVMDSFNSIVTGIGTAFISSIKTDSHDNGDGKGTL 123
VI L+ +L++ NSELGR+A+STA V D F +T + +++ + + S +
Sbjct: 197 PVITHLVYELKMSNSELGRIAISTAAVSD-FLGFLTLVCISYVGTYRYVSPGIANR---- 251
Query: 124 KAFLNVFYYLCFMVVTPLVLRPILKWFVKKTPEGRPMKKVYMYIVFIIALAVGMLGLLTK 183
++ + ++V + +P+ + V TPEG+P+ KVY+Y+ + A+A + +
Sbjct: 252 ----DIVALIILVLVILFIFKPMAQRIVDMTPEGKPVPKVYLYVTILTAIAASIYLSVFN 307
Query: 184 QSVLGGICIVGLIVPEGPPLGTEMIKQLELFCSWFLFPIFVTSCAMKIDL--SVYVKSDY 241
Q + G +VGL +P+GPPLG+ + + E + FPI + AMK D+ ++Y D
Sbjct: 308 QMYILGALLVGLAIPDGPPLGSALEARFESLVTNIFFPISIAVMAMKADVVRALYSFDDI 367
Query: 242 IY--VWLGIIVAVHLFKMLVTIGICWYCNMPMADGLCLALMLSCKGVVDFCNNVFLHDSM 299
+ + LG+ V V V I +C +P + + +A +++ KG VD C
Sbjct: 368 SFNILLLGLTVVVKWTASFVPCLI--FCELPTRESVIIATIMNYKGFVDLCFFDVALRRR 425
Query: 300 LLSSEALSVLSINVLVIGTLARIGVKYLYDPSRKYAGYQKRNILSLKPNSELKIVSCILK 359
LS +V+ I VL+ + +K LYDP RKY GY KR+I+ LK NS+LKI++C+ K
Sbjct: 426 NLSRATYTVMIIYVLLNAGILPTIIKALYDPKRKYIGYVKRDIMHLKTNSDLKILTCLHK 485
Query: 360 PSHIIPIKNVLDICSPTSSNP------LVIHILHLLELVGRSSPVFISHRLQERVGSSSH 413
P +I ++L++ S +N + + LHL++L GR+ P+ I H + + +
Sbjct: 486 PDNISGAISLLELLSSPLNNDNKDRGVIAVTALHLVKLAGRTFPILIPHDKRSKARLLQN 545
Query: 414 TFSEAVIVTFDLFEHDNAGTASVSTYTAISPVRFMHDDICYLALDKLASIIILPFHLRWS 473
++ + +++ F F+ +N + +VS++TA S M DIC LALD L S+II+P +WS
Sbjct: 546 SYIQTMMLAFTEFQQENWESTTVSSFTAYSHENLMDQDICNLALDHLTSMIIVPSGRKWS 605
Query: 474 EDGSVESADETTRSLNTKVLERAPCSVAILVNRGHSSPFNHNENSKQIAMIFLGGSDDRE 533
DG ES D R +N +L+ APCSV IL RG++ + + +IF+GG DDRE
Sbjct: 606 PDGEYESDDIMIRRVNESLLDLAPCSVGILNYRGYNKGKKKTNSIINVGVIFIGGKDDRE 665
Query: 534 ALCLAKRTIKEDTYHLVVYHLVSTIKNDEFTSWDVMLDDELLKGVKGVYGSVDNVTYEKV 593
AL LAK + L V +S + D+ +WD ++DDE+L +K Y +N Y +
Sbjct: 666 ALSLAKWMGQNSRVCLTVIRFLSGQELDKSKNWDYLVDDEVLNDLKATYSLANNFNYMEK 725
Query: 594 EVENTSDTTEFISDIAIQHDFIIVGRRNGIKSPQTQALASWTEYPELGVLGDLLASPDTN 653
V + +A HD +IVGR + S LA W E PELGV+GDLLAS D
Sbjct: 726 VVNGGPAVATTVRLVAEDHDLMIVGRDHEDYSLDLTGLAQWMELPELGVIGDLLASKDLR 785
Query: 654 TKASILVVQQQ 664
+ S+LVVQQQ
Sbjct: 786 ARVSVLVVQQQ 796
>UniRef100_Q9FYC0 Hypothetical protein F28D10_100 [Arabidopsis thaliana]
Length = 705
Score = 348 bits (893), Expect = 3e-94
Identities = 205/614 (33%), Positives = 340/614 (54%), Gaps = 24/614 (3%)
Query: 53 VIVIGQSGCYFAVIASLLSDLEILNSELGRLALSTAMVMDSFNSIVTGIGTAFISSIKTD 112
V++ QS + LS+L+ILNSELGRL LS +++ D F S V+ I + + K
Sbjct: 111 VVISSQSSILLPTVVHFLSELKILNSELGRLVLSASLINDIFASTVS-IFAYLVGTYKNI 169
Query: 113 SHDNGDGKGTLKAFLNVFYYLCFMVVTPLVLRPILKWFVKKTPEGRPMKKVYMYIVFIIA 172
S + A+ ++ + ++V VLRP+++W V++TPEG+P+ VY++ V +
Sbjct: 170 S--------PMTAYRDLIAVIILILVAFCVLRPVVEWIVERTPEGKPVADVYVHAVVLSV 221
Query: 173 LAVGMLGLLTKQSVLGGICIVGLIVPEGPPLGTEMIKQLELFCSWFLFPIFVTSCAMKID 232
+A L G ++G+I+PEGPP+G+ + + E L PI +T M+ D
Sbjct: 222 IASAAYSSFFNMKYLLGPFLLGIIIPEGPPIGSALEAKYEALTMNVLIPISITFSTMRCD 281
Query: 233 -LSVYVKSDYIYVWLGIIVAVHLFKMLVTIGICWYCNMPMADGLCLALMLSCKGVVDFCN 291
+ + + D I+ + ++ KM + C YC +P + + +L+L K +
Sbjct: 282 VMKIVYQYDDIWYNIFLMTFTGFLKMATGMVPCLYCKIPFKEAIAASLLLCSKSFSEIFL 341
Query: 292 NVFLHDSMLLSSEALSVLSINVLVIGTLARIGVKYLYDPSRKYAGYQKRNILSLKPNSEL 351
+D +S + L L+ + + LYDP RKY GYQK+NI++LKP+S+L
Sbjct: 342 YESTYDDSYISQATYTFLITCALINSGIIPTALAGLYDPKRKYVGYQKKNIMNLKPDSDL 401
Query: 352 KIVSCILKPSHIIPIKNVLDICSPTSSNPLVIHILHLLELVGRSSPVFISHRLQ-ERVGS 410
+I++CI +P +I + L T +V+ +LHL++LVG++ PV ISH Q RV +
Sbjct: 402 RILTCIHRPENISAAISFLQFLPST----IVVTVLHLVKLVGKTVPVLISHNKQINRVVT 457
Query: 411 SSHTFSEAVIVTFDLFEHDNAGTASVSTYTAISPVRFMHDDICYLALDKLASIIILPFHL 470
+S+ + + F E + +++ +TAI+ MHD+IC +AL++ SIII+P
Sbjct: 458 NSYIHTANL--AFSQLE-----SVTMTMFTAITHENLMHDEICKVALEQATSIIIVPSGR 510
Query: 471 RWSEDGSVESADETTRSLNTKVLERAPCSVAILVNRGHSSPFNHNENSKQIAMIFLGGSD 530
+W+ DG+ ES DE R LN +L+ A CS+ ILV+RG S + + + +IF+GG D
Sbjct: 511 KWTVDGAFESEDEAIRRLNESLLKSASCSIGILVDRGQLSLKGTRKFNIDVGVIFIGGKD 570
Query: 531 DREALCLAKRTIKEDTYHLVVYHLVSTIKNDEFTSWDVMLDDELLKGVKGVYGSVDNVTY 590
DREAL L K+ + + V L+S + E T+WD +LD E+L+ +K + +++ Y
Sbjct: 571 DREALSLVKKMKQNPRVKITVIRLISD-RETESTNWDYILDHEVLEDLKDT-EATNSIAY 628
Query: 591 EKVEVENTSDTTEFISDIAIQHDFIIVGRRNGIKSPQTQALASWTEYPELGVLGDLLASP 650
+ V + + ++ +D ++VGR +G+ SP L W E PELGV+GDLLAS
Sbjct: 629 TERIVTGGPEVATTVRSLSEDYDLMVVGRDHGMASPDFDGLMEWVELPELGVIGDLLASR 688
Query: 651 DTNTKASILVVQQQ 664
+ +++ S+LVVQQQ
Sbjct: 689 ELDSRVSVLVVQQQ 702
>UniRef100_Q9FYB8 Hypothetical protein F28D10_120 [Arabidopsis thaliana]
Length = 731
Score = 345 bits (885), Expect = 2e-93
Identities = 222/689 (32%), Positives = 371/689 (53%), Gaps = 52/689 (7%)
Query: 3 FTMITRTGRKAWTIAFCSFVIPMFFGLVVCYR------FQEFWKLEMGNFEAKNLPV--- 53
F M RT R+ +AF S +P+ G+V + FQ F+ N + +P+
Sbjct: 64 FLMTVRTSRR---VAFHSGKLPVVIGIVSFFAPLFGLGFQNFFS---DNIDPHYMPLTKA 117
Query: 54 ------IVIGQSGCYFAVIASLLSDLEILNSELGRLALSTAMVMDSFNSIVTGIGTAFIS 107
IVI QS +L +L+I+NSELGRLALS ++ D + GI + ++
Sbjct: 118 LGERTAIVITQSSILLPSTTYILLELKIINSELGRLALSACVIND-----ILGIFSMIVA 172
Query: 108 SIKTD----SHDNGDGKGTLKAFLNVFYYLCFMVVTPLVLRPILKWFVKKTPEGRPMKKV 163
SI+ SH A+ + + F +V LV +P+++W + +TPE +P++ +
Sbjct: 173 SIQATYIHVSHAT--------AYRDTVAVIIFFLVVFLVFKPMVQWVIDRTPEDKPVEDM 224
Query: 164 YMYIVFIIALAVGMLGLLTKQSVLGGICIVGLIVPEGPPLGTEMIKQLELFCSWFLFPIF 223
Y++ V I ALA + + G ++G+I+PEGPPLG+ + + E PI
Sbjct: 225 YIHAVIITALASAAYFVFFNMKYILGPLMIGIIIPEGPPLGSALEAKFERLTMNVFLPIS 284
Query: 224 VTSCAMKIDLSVYVKSDYIYVWLGIIVA--VHLFKMLVTIGICWYCNMPMADGLCLALML 281
+T AM+ D + S + ++ I + + + K++ + C Y +P+++ L ++ +L
Sbjct: 285 ITFSAMRCD-GARILSQFNDIFFNIFLTFLILVIKLVACLAPCLYYKLPLSESLAVSFIL 343
Query: 282 SCKGVVDFCNNVFLHDSMLLSSEALSVLSINVLVIGTLARIGVKYLYDPSRKYAGYQKRN 341
S K DF + D +S S L + L+ + ++ +YDP RKY YQKR+
Sbjct: 344 SYKSFADFVLYEAVLDDTYISQATYSFLILYSLLNAGIVPTVLRRMYDPRRKYVNYQKRD 403
Query: 342 ILSLKPNSELKIVSCILKPSHIIPIKNVLDICS-PTSSNPLVIHILHLLELVGRSSPVFI 400
IL L+ NS+L+I++C+ KP ++ L + S P P+ + +LHL++LVG+ +P+ +
Sbjct: 404 ILHLERNSDLRILTCLHKPENVSETIAFLQLLSSPNLDFPIAVTVLHLVKLVGQINPIIV 463
Query: 401 SH-RLQERVGSSSHTFSEAVIVTFDLFEHDNAGTASVSTYTAISPVRFMHDDICYLALDK 459
SH + +R+ S+ + + F F ++ + +V+T+TA S MH+DIC LALDK
Sbjct: 464 SHDKKLKRLNKDSYIHTANL--AFRQFVLESLESVTVTTFTAFSHENLMHEDICTLALDK 521
Query: 460 LASIIILPFHLRWSEDGSVESADETTRSLNTKVLERAPCSVAILVNRGHSSPFNHNENSK 519
S+I++P +W+ DG ES + R LN +L+RAPCS+ ILV+RG S + + K
Sbjct: 522 TTSMIVVPSGRKWTVDGLFESDNTAIRHLNQSLLDRAPCSIGILVDRGQFSRKSIVTSKK 581
Query: 520 Q----IAMIFLGGSDDREALCLAKRTIKEDTYHLVVYHLVSTIKNDEFTSWDVMLDDELL 575
+ + ++F+GG DDREAL L KR + V LV ++ + WD +LD+E L
Sbjct: 582 RYIIDVGVLFIGGKDDREALSLVKRMKNNPRIRVTVIRLV--FDHEIESDWDYILDNEGL 639
Query: 576 KGVKGVYGSVDNVTYEKVEVENTSDTTEFISDIAIQHDFIIVGRRNGIKSPQTQALASWT 635
K +K + D + Y + V ++ + + + +A ++D ++VGR + + S L W
Sbjct: 640 KDLKSTEDNKD-IDYIERIVTSSVEVVKAVQLLAEEYDLMVVGRDHDMTSQDLSGLMEWV 698
Query: 636 EYPELGVLGDLLASPDTNTKASILVVQQQ 664
E PELGV+GDLLA+ D ++K S+LVVQQQ
Sbjct: 699 ELPELGVIGDLLAARDLSSKVSVLVVQQQ 727
>UniRef100_Q9ZUW0 Hypothetical protein At2g28180 [Arabidopsis thaliana]
Length = 822
Score = 344 bits (883), Expect = 4e-93
Identities = 215/637 (33%), Positives = 346/637 (53%), Gaps = 57/637 (8%)
Query: 48 AKNLP-------VIVIGQSGCYFAVIASLLSDLEILNSELGRLALSTAMVMDSFNSIVTG 100
AKN P V+++ +S F+ IA LL DL + +S +GR+ALS+A+V D ++
Sbjct: 216 AKNRPLTFQEYDVMLLMESITSFSGIARLLRDLGMNHSSIGRVALSSALVSDIVGLLL-- 273
Query: 101 IGTAFISSIKTDSHDNGDGKGTLKAFLNVFYYLCFMVVTPLVLRPILKWFVKKTPEGRPM 160
I+++ S DG L F+V+ V+RPI+ +K+ EGRP+
Sbjct: 274 ----LIANVSRSSATLADGLAILTEIT------LFLVIAFAVVRPIMFKIIKRKGEGRPI 323
Query: 161 KKVYMYIVFIIALAVGMLGLLTKQSVLGGICIVGLIVPEGPPLGTEMIKQLELFCSWFLF 220
+ Y++ V ++ M Q G +GL +P GPP+G+ ++++LE F +
Sbjct: 324 EDKYIHGVLVLVCLSCMYWEDLSQFPPLGAFFLGLAIPNGPPIGSALVERLESFNFGIIL 383
Query: 221 PIFVTSCAMKID-------LSVYVKSDYIYVWLGIIVAVHLFKMLVTIGICWYCNMPMAD 273
P+F+T+ ++ D L+ + D + +++ + L K+ V++ + + MP+ D
Sbjct: 384 PLFLTAVMLRTDTTAWKGALTFFSGDDKKFAVASLVLLIFLLKLSVSVIVPYLYKMPLRD 443
Query: 274 GLCLALMLSCKGVVDFCNNVFLHDSMLLSSEALSVLSINVLVIGTLARIGVKYLYDPSRK 333
+ LAL++S K VLSI ++ L + + +LYDPS++
Sbjct: 444 SIILALIMSHK-----------------------VLSI--VLNSLLIPMAIGFLYDPSKQ 478
Query: 334 YAGYQKRNILSLKPNSELKIVSCILKPSHIIPIKNVLDICSPTSSNPLVIHILHLLELVG 393
+ YQKRN+ S+K ELK + CI +P HI + N+L+ + +PL ++LHL+EL G
Sbjct: 479 FICYQKRNLASMKNMGELKTLVCIHRPDHISSMINLLEASYQSEDSPLTCYVLHLVELRG 538
Query: 394 RSSPVFISHRLQERVGSSSHTFSEAVIVTFDLFEHDNAGTASVSTYTAISPVRFMHDDIC 453
+ P ISH++Q+ + + +SE VI++F+ F + S+ T+T I+ M DDIC
Sbjct: 539 QDVPTLISHKVQKLGVGAGNKYSENVILSFEHFHRSVCSSISIDTFTCIANANHMQDDIC 598
Query: 454 YLALDKLASIIILPFHLRWSED-GSVESADETTRSLNTKVLERAPCSVAILVNRGHSSPF 512
+LALDK ++IILPFH WS D S+ S E R LN VL++APCSV IL+ R +
Sbjct: 599 WLALDKAVTLIILPFHRTWSLDRTSIVSDVEAIRFLNVNVLKQAPCSVGILIERHLVNKK 658
Query: 513 NHNENSKQIAMIFLGGSDDREALCLAKRTIKEDTYHLVVYHLVSTIKNDEFTSWDVMLDD 572
S ++ +IF+GG DDREAL AKR +++ L V L+++ K+ + T WD MLD
Sbjct: 659 QEPHESLKVCVIFVGGKDDREALAFAKRMARQENVTLTVLRLLASGKSKDATGWDQMLDT 718
Query: 573 ----ELLKGVK-GVYGSVDNVTYEKVEVENTSDTTEFISDIAIQHDFIIVGRRNGIKSPQ 627
EL+K G+ + Y + E+ + +DT+ + +A +D +VGR G
Sbjct: 719 VELRELIKSNNAGMVKEETSTIYLEQEILDGADTSMLLRSMAFDYDLFVVGRTCGENHEA 778
Query: 628 TQALASWTEYPELGVLGDLLASPDTNTKASILVVQQQ 664
T+ + +W E+ ELGV+GD LASPD +K S+LVVQQQ
Sbjct: 779 TKGIENWCEFEELGVIGDFLASPDFPSKTSVLVVQQQ 815
>UniRef100_Q9SGE4 T23G18.2 [Arabidopsis thaliana]
Length = 2658
Score = 341 bits (874), Expect = 5e-92
Identities = 226/676 (33%), Positives = 359/676 (52%), Gaps = 44/676 (6%)
Query: 1 MDFTMITRTGRKAWTIAFCSFVIPMFFGLVVCYRFQEFWKLEMGNFEAKNLPVIVIGQSG 60
MD + RT +++ I F + +IP+ G + +R++E + + +I+ QS
Sbjct: 1139 MDAELPFRTEKRSSVIGFITVIIPLICGSLT-FRYRE--RRGDSSILRMEYRLIIFLQSI 1195
Query: 61 CYFAVIASLLSDLEILNSELGRLALSTAMVMDSFNSIVTGIGTAFISSIKTDSHDNGDGK 120
F I +LL DL+I +SE GR+ALS AMV D + G F ++I +
Sbjct: 1196 SAFTSIDTLLKDLQIKHSEFGRIALSGAMVTD-----MLAFGVTFFNAIYYEK------- 1243
Query: 121 GTLKAFLNVFYYLCFMVVTPLVLRPILKWFVKKTPEGRPMKKVYMYIVFIIALAVGMLGL 180
L F+ + F+VV V+RP + W +K+TPEGRP+K Y+Y +F IA A
Sbjct: 1244 --LYGFMQTVGFCLFVVVMICVVRPAMYWVIKQTPEGRPVKDFYLYSIFGIAFAC--FTF 1299
Query: 181 LTKQSVL---GGICIVGLIVPEGPPLGTEMIKQLELFCSWFLFPIFVTSCAMKIDLSVYV 237
K L G + GL VP G PLGT +I++ E F + P+F + M++DL
Sbjct: 1300 FNKVIHLFGPAGSFVFGLTVPNGYPLGTTLIQKFESFNLGSILPLFGSLTMMQVDLLRLF 1359
Query: 238 KSD--------YIYVWLGIIVAVHLFKMLVTIGICWYCNMPMADGLCLALMLSCKGVVDF 289
K IY + I+ V+ K +VT + MP+ D LAL+LS KG+ +
Sbjct: 1360 KESGDLIRMEGQIYEVISFILLVNTTKFVVTTITAYAFKMPLRDSFALALVLSNKGIFEL 1419
Query: 290 CNNVFLHDSMLLSSEALSVLSINVLVIGTLARIGVKYLYDPSRKYAGYQKRNILSLKPNS 349
+ + L+ E ++L+ L+ + ++ ++DP++++ Y+KRN+ LK +
Sbjct: 1420 AYYTYAVELKLIRPEVFTILAAYTLLNSIFIPMLLELVHDPTKRFRCYRKRNLGILKDGA 1479
Query: 350 ELKIVSCILKPSHIIPIKNVLDICSPTSSNPLVIHILHLLELVGRSSPVFISHRLQERVG 409
L+ + C+ +P HI + ++L+ SP+ +P+ +ILHL+ELVG+++P+FISH+LQ+
Sbjct: 1480 ALQCLMCVYRPDHITSMTDLLETFSPSQDSPMACNILHLVELVGQANPMFISHQLQKPEP 1539
Query: 410 SSSHTFSEAVIVTFDLFEHDNAGTASVSTYTAISPVRFMHDDICYLALDKLASIIILPFH 469
S+ + S+ VI++F F+ S+ +T++S + MH+DIC+LAL + S+I+LPFH
Sbjct: 1540 GST-SLSDNVIISFRGFQRQFFEYTSLDIFTSVSVSQHMHEDICWLALSRSLSLIVLPFH 1598
Query: 470 LRWSEDGS-VESADETTRSLNTKVLERAPCSVAILVNRGHSSPFNHNENSKQIAMIFLGG 528
WS D S V S D+ R LN VL RAPCSV I V R + ++ +I +IF GG
Sbjct: 1599 RTWSVDRSTVISNDDNLRMLNVNVLRRAPCSVGIFVYRKPIVESHMAKSHSKICLIFNGG 1658
Query: 529 SDDREALCLAKR-TIKEDTYHLVVYHLV---STIKNDEFTSWDVMLDDELLKGVKGVYGS 584
DDREAL + R + E L + + S + NDE W+ L + V + GS
Sbjct: 1659 KDDREALAITNRMRLTEKRTRLTIIRFIPKSSEMDNDE---WEQQQSINLKESVTSIVGS 1715
Query: 585 -----VDNVTYEKVEVENTSDTTEFISDIAIQHDFIIVGRRNGIKSPQTQALASWTEYPE 639
VTY V + S+T+ + +A +D IVG +GI + T ++ WTE+ E
Sbjct: 1716 NIKENDAKVTYIDKAVSDGSETSRILRAMANDYDLFIVGSGSGIGTEATSGISEWTEFNE 1775
Query: 640 LGVLGDLLASPDTNTK 655
LG +GDLLAS + +K
Sbjct: 1776 LGPIGDLLASHEYPSK 1791
Score = 341 bits (874), Expect = 5e-92
Identities = 215/653 (32%), Positives = 348/653 (52%), Gaps = 31/653 (4%)
Query: 1 MDFTMITRTGRKAWTIAFCSFVIPMFFGLVVCYRFQEFWKLEMGNFEAKNLPVIVIGQSG 60
MD M+ +TG K + ++P+ +V + +E + E + I+ QS
Sbjct: 365 MDVGMVRKTGTKVIVTGIATVILPIIAANMVFGKLRETGGKYLTGMEYRT---ILFMQSI 421
Query: 61 CYFAVIASLLSDLEILNSELGRLALSTAMVMDSFNSIVTGIGTAFISSIKTDSHDNGDGK 120
F I+ LL DL I +SE GR+ +STAMV D TG G + + D +
Sbjct: 422 SAFTGISRLLRDLRINHSEFGRIVISTAMVADG-----TGFGVNLFALVAWM-----DWR 471
Query: 121 GTLKAFLNVFYYLCFMVVTPLVLRPILKWFVKKTPEGRPMKKVYMYIVFIIALAVGMLGL 180
+ + + Y+ FMV V+RP + W +K+TP+ RP+K+ ++YI+ I+A
Sbjct: 472 VSALQGVGIIGYVIFMV---WVVRPAMFWVIKRTPQERPVKECFIYIILILAFGGYYFLK 528
Query: 181 LTKQSVLGGICIVGLIVPEGPPLGTEMIKQLELFCSWFLFPIFVTSCAMKIDLSVYVKS- 239
G ++GL VP GPPLG++++++ E F + L P+F+ ++ID
Sbjct: 529 EIHMFPAVGPFLLGLCVPHGPPLGSQLVEKFESFNTGILLPLFLFFSMLQIDGPWLANQI 588
Query: 240 -------DYIYVWLGIIVAVHLFKMLVTIGICWYCNMPMADGLCLALMLSCKGVVDFCNN 292
+Y L II+ V + K++ ++ MP+ D +AL+LS KG+V+ C
Sbjct: 589 GQLRHFDGQLYEALTIIIVVFVAKIIFSMIPALLAKMPLTDSFVMALILSNKGIVELCYF 648
Query: 293 VFLHDSMLLSSEALSVLSINVLVIGTLARIGVKYLYDPSRKYAGYQKRNILSLKPNSELK 352
++ +S +L ++ ++++ +LV T++ + + YLYD S+++ +QKRN++SLK SELK
Sbjct: 649 LYGVESNVLHVKSFTIMATMILVSSTISPVLIHYLYDSSKRFISFQKRNLMSLKLGSELK 708
Query: 353 IVSCILKPSHIIPIKNVLDICSPTSSNPLVIHILHLLELVGRSSPVFISHRLQERVGSSS 412
+ CI K HI + N+L P + + +++HL+ELVG +PVFISH++Q + +
Sbjct: 709 FLVCIHKADHISGMINLLAQSFPLHESTISCYVIHLVELVGLDNPVFISHQMQ-KAEPGN 767
Query: 413 HTFSEAVIVTFDLFEHDNAGTASVSTYTAISPVRFMHDDICYLALDKLASIIILPFHLRW 472
++S V++ FD F+H + S+ +T IS R+MH +I LALDK AS ++LPFH+ W
Sbjct: 768 RSYSNNVLIAFDNFKH-YWKSISLELFTCISNPRYMHQEIYSLALDKQASFLMLPFHIIW 826
Query: 473 SED-GSVESADETTRSLNTKVLERAPCSVAILVNRGHSSPFNHNENSKQIAMIFLGGSDD 531
S D +V S D R+ N VL +APCSV I V+R + S ++ IF+GG DD
Sbjct: 827 SLDQTTVVSDDVMRRNANLNVLRQAPCSVGIFVHRQKLLSAQKSSPSFEVCAIFVGGKDD 886
Query: 532 REALCLAKRTIKEDTYHLVVYHLVSTIKNDEFTSWDVMLDD----ELLKGVKGVYGSVDN 587
REAL L ++ ++ +L V L+ + T WD MLD E+L+ G
Sbjct: 887 REALALGRQMMRNPNVNLTVLKLIPAKMDGMTTGWDQMLDSAEVKEVLRNNNNTVGQHSF 946
Query: 588 VTYEKVEVENTSDTTEFISDIAIQHDFIIVGRRNGIKSPQTQALASWTEYPEL 640
V Y + V + SDT+ + IA D +VGR G+ + AL+ WTE+ EL
Sbjct: 947 VEYVEETVNDGSDTSTLLLSIANSFDLFVVGRSAGVGTDVVSALSEWTEFDEL 999
Score = 303 bits (775), Expect = 1e-80
Identities = 207/679 (30%), Positives = 352/679 (51%), Gaps = 47/679 (6%)
Query: 2 DFTMITRTGRKAWTIAFCSFVIPMFFGLVVCYRFQEFWKLEMGNFEAKNLPVIVIGQSGC 61
D ++ ++G K+ I S +IP G ++ ++ L M E V+ S
Sbjct: 2010 DVGIMKKSGTKSVVIGITSMIIPWQIGKLLYSSREKSSILTMTEME---YTVMTFTMSMT 2066
Query: 62 YFAVIASLLSDLEILNSELGRLALSTAMVMDSFNSIVTGIGTAFISSIKTDSHDNGDGKG 121
F + LL+DL+I++++ G++A S MV D +T +A++S +T G
Sbjct: 2067 PFTCVNMLLTDLKIVHTDFGQIAQSAGMVTDLLAFFLTV--SAYVSRDETQGVKMG---- 2120
Query: 122 TLKAFLNVFYYLCFMVVTPLVLRPILKWFVKKTPEGRPMKKVYMYIVFIIA-LAVGMLGL 180
AF+ F ++ ++R + W ++ TPEG P+K VY+YI ++A L+
Sbjct: 2121 --LAFMAFFIFV-------YLVRQFMLWVIRHTPEGAPVKNVYLYIGLLLAYLSYLYWSR 2171
Query: 181 LTKQSVLGGICIVGLIVPEGPPLGTEMIKQLELFCSWFLFPIFVTSCAMKIDLSVYVKS- 239
LG + GL VP GPPLG+ I++ + F P+F + +K+D S K
Sbjct: 2172 FLFFGPLGAFAL-GLAVPNGPPLGSVFIQKFDSFNEGIFLPLFGSLSMIKLDWSFLRKEF 2230
Query: 240 -------DYIYVWLGIIVAVHLFKMLVTIGICWYCNMPMADGLCLALMLSCKGVVDFCNN 292
++Y + V++ K + +P+ D + L +++ K +
Sbjct: 2231 GNGRHLHGHMYECFSFLPIVYIAKFATSFLAALATKIPLRDSIILGVIMGTKSSFELGYV 2290
Query: 293 VFLHDSMLLSSEALSVLSINVLVIGTLARIGVKYLYDPSRKYAGYQKRNILSLKPNSELK 352
+ + +S E LS+L + +LV L + + +LYD S+++ Y +RN LK E++
Sbjct: 2291 LTAFEKDRISLEVLSLLGVYILVNSLLTPMAIHFLYDRSKRFVCYGRRN---LKEKPEMQ 2347
Query: 353 IVSCILKPSHIIPIKNVLDICSPTSSNPLVIHILHLLELVGRSSPVFISHRLQERVGSSS 412
+ CI KP +I + ++L SP+ +P+ +LHL+EL+G+++P FISH+LQ + S
Sbjct: 2348 TLVCINKPDNITSMISLLRATSPSKDSPMECCVLHLIELLGQATPTFISHQLQ-KPKPGS 2406
Query: 413 HTFSEAVIVTFDLFEHDNAGTASVSTYTAISPVRFMHDDICYLALDKLASIIILPFHLRW 472
++SE VI +F LF+ +AS++ +T+++ + MH+ IC+ AL + +++I+L FH W
Sbjct: 2407 RSYSENVISSFQLFQEVYWDSASINMFTSLTSAKEMHEQICWFALSQGSNLILLSFHRTW 2466
Query: 473 SEDGSVE-SADETTRSLNTKVLERAPCSVAILVNRGHSSPFNHNENSKQIAMIFLGGSDD 531
+G+V S D+T RSLN VL+RAPCSV I V R E+ ++ +I++GG+DD
Sbjct: 2467 EPNGNVIISDDQTLRSLNLNVLKRAPCSVGIFVYRKPIWQTKALESPCRVCLIYVGGNDD 2526
Query: 532 REALCLAKRTIKEDTYHLVVYHLVSTIKNDEFT------SWDVMLDDELLKGVKGVYGSV 585
+EAL LA L V L+ T DE + D+ ++ G
Sbjct: 2527 KEALALADHMRGNQQVILTVLRLIPTSYADESSLRIHSQMVDMNRHEDQRPG-------- 2578
Query: 586 DNVTYEKVEVENTSDTTEFISDIAIQHDFIIVGRRNGIKSPQTQALASWTEYPELGVLGD 645
D T V + ++T++ + ++ +D IVGRR+G+ + T+ L W E+ ELGV+GD
Sbjct: 2579 DKSTIIDWTVGDGTETSKILHSVSYDYDLFIVGRRSGVGTTVTRGLGDWMEFEELGVIGD 2638
Query: 646 LLASPDTNTKASILVVQQQ 664
LLAS ++AS+LVVQQQ
Sbjct: 2639 LLASEYFPSRASVLVVQQQ 2657
>UniRef100_Q9SIT5 Putative Na/H antiporter [Arabidopsis thaliana]
Length = 821
Score = 333 bits (853), Expect = 1e-89
Identities = 214/693 (30%), Positives = 354/693 (50%), Gaps = 49/693 (7%)
Query: 1 MDFTMITRTGRKAWTIAFCSFVIPMFFGLVVCYRFQEFWKLEMGNFEAKNLPVIVIGQSG 60
MD ++ +TG++A TIA V+P G + + E + + + + S
Sbjct: 119 MDIMVVRKTGKRALTIAIGGMVLPFLIGAAFSFSMH---RSEDHLGQGTYILFLGVALSV 175
Query: 61 CYFAVIASLLSDLEILNSELGRLALSTAMVMDSFNSIVTGIGTAFISSIKTDSHDNGDGK 120
F V+A +L++L+++N+E+GR+++S A+V D F I+ + A S KT
Sbjct: 176 TAFPVLARILAELKLINTEIGRISMSAALVNDMFAWILLALAIALAESDKT--------- 226
Query: 121 GTLKAFLNVFYYLC---FMVVTPLVLRPILKWFVKKTPEGRPMKKVYMYIVFIIALAVGM 177
+F +++ + F+ V V+RP + W ++KTPEG + ++ ++ + G
Sbjct: 227 ----SFASLWVMISSAVFIAVCVFVVRPGIAWIIRKTPEGENFSEFHICLILTGVMISGF 282
Query: 178 LGLLTKQSVLGGICIVGLIVPEGPPLGTEMIKQLELFCSWFLFPIFVTSCAMKIDLSVYV 237
+ + G + GL++P GP LG +I++LE F S L P+F +K +++
Sbjct: 283 ITDAIGTHSVFGAFVFGLVIPNGP-LGLTLIEKLEDFVSGLLLPLFFAISGLKTNIAAIQ 341
Query: 238 KSDYIYVWLGIIVAVHLF---KMLVTIGICWYCNMPMADGLCLALMLSCKGVVDFCNNVF 294
WL + + + L K++ T+ + ++ MP+ +G+ L L+L+ KG+V+
Sbjct: 342 GPA---TWLTLFLVIFLACAGKVIGTVIVAFFHGMPVREGITLGLLLNTKGLVEMIVLNV 398
Query: 295 LHDSMLLSSEALSVLSINVLVIGTLARIGVKYLYDPSRKYAGYQKRNILSLKPNSELKIV 354
D +L E + + + LV+ + V LY P +K Y++R I KP+SEL+++
Sbjct: 399 GKDQKVLDDETFATMVLVALVMTGVITPIVTILYKPVKKSVSYKRRTIQQTKPDSELRVL 458
Query: 355 SCILKPSHIIPIKNVLDICSPTSSNPLVIHILHLLELVGRSSPVFISHRLQER---VGSS 411
C+ P ++ I N+L+ PT +P+ I++LHL+EL GR+S + I H ++ +
Sbjct: 459 VCVHTPRNVPTIINLLEASHPTKRSPICIYVLHLVELTGRASAMLIVHNTRKSGRPALNR 518
Query: 412 SHTFSEAVIVTFDLFEHDNAGTASVSTYTAISPVRFMHDDICYLALDKLASIIILPFHLR 471
+ S+ +I F+ +E +A +V TAISP MH+D+C LA DK S II+PFH +
Sbjct: 519 TQAQSDHIINAFENYEQ-HAAFVAVQPLTAISPYSTMHEDVCSLAEDKRVSFIIIPFHKQ 577
Query: 472 WSEDGSVESADETTRSLNTKVLERAPCSVAILVNRG--HSSPFNHNENSKQIAMIFLGGS 529
+ DG +ES + R +N +LE +PCSV ILV+RG ++ N N S Q+A++F GG
Sbjct: 578 QTVDGGMESTNPAYRLVNQNLLENSPCSVGILVDRGLNGATRLNSNTVSLQVAVLFFGGP 637
Query: 530 DDREALCLAKRTIKEDTYHLVVYHLV----------STIKNDEFTSWDVM-------LDD 572
DDREAL A R + L V + + ND M LDD
Sbjct: 638 DDREALAYAWRMAQHPGITLTVLRFIHDEDEADTASTRATNDSDLKIPKMDHRKQRQLDD 697
Query: 573 ELLKGVKGVYGSVDNVTYEKVEVENTSDTTEFISDIAIQHDFIIVGRRNGIKSPQTQALA 632
+ + + +++ Y + V N +T + + HD IVGR G+ SP T L
Sbjct: 698 DYINLFRAENAEYESIVYIEKLVSNGEETVAAVRSMDSSHDLFIVGRGEGMSSPLTAGLT 757
Query: 633 SWTEYPELGVLGDLLASPDTNTKASILVVQQQV 665
W+E PELG +GDLLAS D S+LVVQQ V
Sbjct: 758 DWSECPELGAIGDLLASSDFAATVSVLVVQQYV 790
>UniRef100_Q9ZUV9 Hypothetical protein At2g28170 [Arabidopsis thaliana]
Length = 617
Score = 332 bits (850), Expect = 3e-89
Identities = 205/624 (32%), Positives = 346/624 (54%), Gaps = 30/624 (4%)
Query: 54 IVIGQSGCYFAVIASLLSDLEILNSELGRLALSTAMVMDSFNSIVTGIGTAFISSIKTDS 113
I++ QS FA + LL+DL+I +SE GR+ S A V D I+ GT + K
Sbjct: 9 IILLQSLSSFAGVNGLLTDLKINHSEFGRMVQSCAAVTDLVIFIMVS-GTVLLKGQKGLP 67
Query: 114 HDNGDGKGTLKAFLNVFYYLCFMVVTPLVLRPILKWFVKKTPEGRPMKKVYMYIVFIIAL 173
H + + + F+V ++ P++ W +K+TPEGR +K VY+Y+V A
Sbjct: 68 HG-----------IVIVLVIGFLVY---IVWPVMLWIIKQTPEGRLVKDVYIYLVMATAY 113
Query: 174 AVGMLGLLTKQSVLGGICIVGLIVPEGPPLGTEMIKQLELFCSWFLFPIFVTSCAMKIDL 233
V M L Q G I+GL P GPPLG+ +I++ E F L P+F + ++D+
Sbjct: 114 FVYMFWLNFFQFSTYGWFIIGLATPAGPPLGSALIQRFECFNVGVLLPLFGSLSMEQLDI 173
Query: 234 SVYVKS--------DYIYVWLGIIVAVHLFKMLVTIGICWYCNMPMADGLCLALMLSCKG 285
S ++ + Y + +I+ V + K +VT + +P D + LA++LS +
Sbjct: 174 SWLMREILNLKHMEGFAYEAISVILIVTVVKFVVTAITAFAVRIPYRDSIVLAMVLSNRS 233
Query: 286 VVDFCNNVFLHDSMLLSSEALSVLSINVLVIGTLARIGVKYLYDPSRKYAGYQKRNILSL 345
+ + ++ + + +++ ++ +++VLV L I ++++Y+P ++ Y+ RN+L+L
Sbjct: 234 IFELGYLGYIVELKMFDNKSFTIAALSVLVSSLLTPIAIEFMYEPQHIFSSYRDRNMLTL 293
Query: 346 KPNSELKIVSCILKPSHIIPIKNVLDICSPTSSNPLVIHILHLLELVGRSSPVFISHRLQ 405
K +S+LK + CI KP HI + N +++ +PT + L ++LHL+EL+G++ P FISH++Q
Sbjct: 294 KHDSKLKTLVCIHKPDHITSMVNFVELFNPTQESKLECNVLHLVELIGQAIPTFISHKMQ 353
Query: 406 E-RVGSSSHTFSEAVIVTF-DLFEHDNAGTASVSTYTAISPVRFMHDDICYLALDKLASI 463
+ +VG+ S S VI F L H S+ +T+ S V MH+D+C+LALDK ++
Sbjct: 354 KPKVGTRS--CSRNVITAFLSLRRHLTKEAISIDIFTSASLVEHMHEDLCWLALDKNVAL 411
Query: 464 IILPFHLRWSEDGS-VESADETTRSLNTKVLERAPCSVAILVNRGHSSPFNHNENSKQIA 522
++LPFH WS D S + S D+ ++LN KVL+RA CSV I V R + + ++
Sbjct: 412 VVLPFHRSWSVDRSTIVSDDKAMQNLNHKVLKRASCSVGIFVYRKPLWESQMHGSCYKVC 471
Query: 523 MIFLGGSDDREALCLAKRTIKEDTYHLVVYHLVSTIKNDEFTSWDVMLDDELLKGVKGVY 582
I +GG DD+EAL R + + + HL+ + +E LD + +K +
Sbjct: 472 AIVVGGKDDKEALAFTNRMRRNKQTSVTILHLIPQLTTEESEDSVQKLDYDDIKEIMKTE 531
Query: 583 GSVDNVTYEKVE--VENTSDTTEFISDIAIQHDFIIVGRRNGIKSPQTQALASWTEYPEL 640
S +N ++ +E V+ ++T+ + IA +D IVGR +G+ S T+ L WTE+ EL
Sbjct: 532 DSNENDSWICIEKSVKEGAETSVILRSIAYDYDLFIVGRSSGMNSAVTKGLNEWTEFEEL 591
Query: 641 GVLGDLLASPDTNTKASILVVQQQ 664
G LGD++AS + ++AS+LV+QQQ
Sbjct: 592 GALGDVIASKEFPSRASVLVLQQQ 615
>UniRef100_Q75J76 Putative cation/hydrogen exchanger [Oryza sativa]
Length = 874
Score = 301 bits (772), Expect = 3e-80
Identities = 207/693 (29%), Positives = 346/693 (49%), Gaps = 48/693 (6%)
Query: 1 MDFTMITRTGRKAWTIAFCSFVIPMFFGLVVCYRFQEFWKLEMGNFEAKNLPVIVIGQSG 60
MD I R+G+KA IA +P G + F+ ++ +A L + + S
Sbjct: 149 MDVNTIRRSGKKALIIAVAGMALPFCIGTATSFIFRH--QVSKNVHQASFLLFLGVALSV 206
Query: 61 CYFAVIASLLSDLEILNSELGRLALSTAMVMDSFNSIVTGIGTAFISSIKTDSHDNGDGK 120
F V+A +L+++++LNS+LGR+A+S A+V D I+ + A S N
Sbjct: 207 TAFPVLARILAEVKLLNSDLGRIAMSAAIVNDMCAWILLALAIAI-------SEVNSSAF 259
Query: 121 GTLKAFLN--VFYYLCFMVVTPLVLRPILKWFVKKTPEGRPMKKVYMYIVFIIALAVGML 178
+L + F CF VV RP++ W V++ PEG + V++ ++ + G+
Sbjct: 260 SSLWVLIAGVAFVLACFYVV-----RPLMWWIVRRVPEGEAIGDVHITLILTGVMVAGVC 314
Query: 179 GLLTKQSVLGGICIVGLIVPEGPPLGTEMIKQLELFCSWFLFPIFVTSCAMKIDLSVYVK 238
+ G + GL++P GP LG +I++LE F + L P+F ++ +++
Sbjct: 315 TDAIGIHSVFGAFVYGLVMPSGP-LGVVLIEKLEDFVTGLLLPLFFAISGLRTNVTKVRD 373
Query: 239 SDYIYVWLGIIVAVHLFKMLVTIGICWYCNMPMADGLCLALMLSCKGVVDFCNNVFLHDS 298
+ + + + V K++ TI I M DG+ L +++ +G+V+ D
Sbjct: 374 PITVGLLVLVFVMASFAKIMGTILIAVSYTMTFRDGVALGFLMNTRGLVEMIVLNIGRDK 433
Query: 299 MLLSSEALSVLSINVLVIGTLARIGVKYLYDPSRKYAGYQKRNILSLKPNSELKIVSCIL 358
+L E+ +V+ + + + L V +Y P+R+ GY++RN+ K ++EL++++C+
Sbjct: 434 EVLDDESFAVMVLVSVAMTALVTPVVTTVYRPARRLVGYKRRNLQRSKHDAELRMLACVH 493
Query: 359 KPSHIIPIKNVLDICSPTSSNPLVIHILHLLELVGRSSPVFISHRLQERVGSSS-HTFSE 417
++ I ++L++ +PT +P+ I+ LHL+EL GR+S + +H G +S H F+
Sbjct: 494 TTRNVPSIISLLELSNPTKRSPIFIYALHLVELTGRASNMLAAHHSASNPGGASDHIFN- 552
Query: 418 AVIVTFDLFEHDNAGTASVSTYTAISPVRFMHDDICYLALDKLASIIILPFHLRWSEDGS 477
F+ +E + G SV TA+SP + MH+D+C LA DK S+I+LPFH + + DG
Sbjct: 553 ----AFESYE-EMVGGVSVQALTAVSPYQTMHEDVCVLAEDKHVSLIVLPFHKQQTVDGG 607
Query: 478 VESADETTRSLNTKVLERAPCSVAILVNRGHSSPFNHNENSKQIAMIFLGGSDDREALCL 537
+E + + R N +L APCSV ILV+RG S+ +A++F GG DDRE L
Sbjct: 608 MEPINASLRGFNESILASAPCSVGILVDRGLSAAAARMAAVHHVALLFFGGPDDREGLAY 667
Query: 538 AKRTIKEDTYHLVVYHLV--------------------STIKN--DEFTSWDVMLDDELL 575
A R ++ L + L+ S N E + +D+E L
Sbjct: 668 AWRMVENPGVCLTIVRLIPPGYTAPAISPPQPPMPAAHSRAINVVPEVAKSERQMDEEYL 727
Query: 576 KGVKGVYGSVDNVTYEKVEVENTSDTTEFI-SDIAIQHDFIIVGRRNG-IKSPQTQALAS 633
+ D + Y + V N+ +T I S + H+ IVGR G SP T ALA
Sbjct: 728 NEFRSRNLGNDAILYVEQVVANSEETVAAIRSQLDNAHELYIVGRHPGEASSPLTSALAE 787
Query: 634 WTEYPELGVLGDLLASPDTNTKASILVVQQQVM 666
W E PELG +GDLL S + + AS+LV+QQ V+
Sbjct: 788 WMESPELGPIGDLLVSSEFSKMASVLVMQQYVI 820
>UniRef100_Q6ZJ60 Putative cation/hydrogen exchanger [Oryza sativa]
Length = 825
Score = 301 bits (771), Expect = 4e-80
Identities = 216/693 (31%), Positives = 336/693 (48%), Gaps = 43/693 (6%)
Query: 2 DFTMITRTGRKAWTIAFCSFVIP----MFFGLVVCYRFQEFWKLEMGNFEAKNLPVIVIG 57
D MI ++G KA IA P GL + R W +F NL
Sbjct: 132 DLGMIRKSGNKAIAIAVLGTASPHLAMYITGLALKARVPAAWA---ASFLLTNLNS---W 185
Query: 58 QSGCYFAVIASLLSDLEILNSELGRLALSTAMVMDSFNSIVTGIGTAFISSIKTDSHDNG 117
S F V+ L DL +L+S+LGRLA+S A++ D N+ T+++ +
Sbjct: 186 WSLSAFIVVCCTLHDLNLLSSKLGRLAMSAALIGDFANTFAIAGVTSYLLAASPSEKLQR 245
Query: 118 DGKGTLKAFLNVFYYLCFMVVTPLVLRPILKWFVKKTPEGRPMKKVYMYIVFIIALAVGM 177
G ++ AF ++ FM LV RP + ++ PEG + + + V +I L
Sbjct: 246 IGIASVIAFTT---FIAFMA---LVARPAILRLIRDVPEGALLTEARLIAVLLICLTCSF 299
Query: 178 LGLLTKQSVLGGICIVGLIVPEGPPLGTEMIKQLELFCSWFLFPIFVTSCAMKIDLSVYV 237
G L G ++GL++P G PLG M ++L+ + L P+ M++++
Sbjct: 300 TGELLGLHATYGPFMLGLMLPGGAPLGVTMAERLDRLVAGVLMPLLFAQGGMRLNVKKIT 359
Query: 238 KSDYIYVWLGIIVAVHLFKMLVTIGICWYCNMPMADGLCLALMLSCKGVVDFCNNVFLHD 297
+ + +V + K + +I C Y MP+ D + + LM++ KG+ + D
Sbjct: 360 DASTCALLETFLVVGVVSKFVASIMPCLYFRMPVRDAVVVGLMMNFKGITEVVYASAFED 419
Query: 298 SMLLSSEALSVLSINVLVIGTLARIGVKYLYDPSRKYAGYQKRNILSLKPNSELKIVSCI 357
+ +L + + INVL+IG + VKY+Y P KY Y++R + K EL++V+CI
Sbjct: 420 AQVLDEQVYAAFMINVLLIGAASASAVKYMYHPEEKYVAYRRRTVEHKKLGEELRVVACI 479
Query: 358 LKPSHIIPIKNVLDICSPTSSNPLVIHILHLLELVGRSSPVF--ISHRLQERVGSSSHTF 415
+ P+ +LD SPT +PL +++LHL+ L G +S V H + V S + T
Sbjct: 480 HSQDDVGPMLALLDASSPTPMSPLSVYLLHLMPLAGLTSSVLRHFKHGKRNCVPSGT-TD 538
Query: 416 SEAVIVTFDLF-EHDNAGTASVSTYTAISPVRFMHDDICYLALDKLASIIILPFHLRWSE 474
SE V+ F F + G AS+ Y I+P MHDD+C +AL+K A +I++PFH R +
Sbjct: 539 SERVVNAFQFFVQQRPPGAASLVPYVCIAPYATMHDDVCAVALEKRAMLIVVPFHKRLAI 598
Query: 475 DGSVESADETT---RSLNTKVLERAPCSVAILVNRGHSSP----------FNHNENSKQI 521
DGSVE ++ NT +L +PCSVAILV+RG S F H ++
Sbjct: 599 DGSVEPTSHNAGAIQAANTNILNYSPCSVAILVDRGSLSTVAAAAAAADGFPH-----RV 653
Query: 522 AMIFLGGSDDREALCLAKRTIKEDTYHLVVYHLV----STIKNDEFTSWDVMLDDELLKG 577
A+ FLGG DDREAL LA ++ T L V+ + + E + D+ L+
Sbjct: 654 ALYFLGGPDDREALALAATMAEDATIGLTVFRFMLPADRQSRGGEGDGEEDRRDEAELQE 713
Query: 578 VKGVYGSVDNVTYEKVEVENTSDTTEFISDIAIQHDFIIVGRRN-GIKSPQTQALASWTE 636
+ V Y + V + + + I + + ++VGRR+ +SP T ++ W+E
Sbjct: 714 FVRRWVDDHRVAYSENMVGGSDEMVDVIRKTSPAFNLLVVGRRSESPESPLTAGISDWSE 773
Query: 637 YPELGVLGDLLASPDTNTKASILVVQQQVMPKA 669
+ ELGVLGDLL S D + S LVVQQQ M A
Sbjct: 774 HLELGVLGDLLTSTDFGCRVSTLVVQQQTMAAA 806
>UniRef100_Q9LMJ1 F10K1.31 protein [Arabidopsis thaliana]
Length = 829
Score = 301 bits (770), Expect = 5e-80
Identities = 185/672 (27%), Positives = 345/672 (50%), Gaps = 26/672 (3%)
Query: 1 MDFTMITRTGRKAWTIAFCSFVIPMFFGLVVCYRFQEFWKLEMGNFEAKNLPVIVIGQSG 60
+D ++I + G KA I S+ +P G + + + L + ++ +
Sbjct: 132 IDASIIRKAGSKAILIGTASYALPFSLGNLTVLFLKNTYNLPPDVVHC--ISTVISLNAM 189
Query: 61 CYFAVIASLLSDLEILNSELGRLALSTAMVMDSFNSIVTGIGTAFISSIKTDSHDNGDGK 120
F V ++L++L ILNS+LGRLA + ++V ++F+ IV + F+
Sbjct: 190 TSFPVTTTVLAELNILNSDLGRLATNCSIVCEAFSWIVALVFRMFLRD------------ 237
Query: 121 GTLKAFLNVFYYLCFMVVTPLVLRPILKWFVKKTPEGRPMK-KVYMYIVFIIALAVGMLG 179
GTL + + + ++V V RP + W ++ ++ + + ++ L + +
Sbjct: 238 GTLASVWSFVWVTALILVIFFVCRPAIIWLTERRSISIDKAGEIPFFPIIMVLLTISLTS 297
Query: 180 LLTKQSVLGGICIVGLIVPEGPPLGTEMIKQLELFCSWFLFPIFVTSCAMKIDLSVYVKS 239
+ G +G+ +P+GPPLGT + +LE+F + + P F++ ++ + + +S
Sbjct: 298 EVLGVHAAFGAFWLGVSLPDGPPLGTGLTTKLEMFATSLMLPCFISISGLQTNFFIIGES 357
Query: 240 DYIYVWLGIIVAVHLFKMLVTIGICWYCNMPMADGLCLALMLSCKGVVDFCNNVFLHDSM 299
++ + +I+ + K L T YCN+ + D LAL++ C+GV++ V D
Sbjct: 358 -HVKIIEAVILITYGCKFLGTAAASAYCNIQIGDAFSLALLMCCQGVIEIYTCVMWKDEK 416
Query: 300 LLSSEALSVLSINVLVIGTLARIGVKYLYDPSRKYAGYQKRNILSLKP-NSELKIVSCIL 358
+L++E ++L I +L++ ++R V LYDPS++Y KR IL + N + +++ C+
Sbjct: 417 VLNTECFNLLIITLLLVTGISRFLVVCLYDPSKRYRSKSKRTILDTRQRNLQFRLLLCVY 476
Query: 359 KPSHIIPIKNVLDICSPTSSNPLVIHILHLLELVGRSSPVFISHRLQERVGSSSHTFSEA 418
++ + N+L+ P+ +P+ + LHL+EL GR+ V + H ++ ++ S
Sbjct: 477 NVENVPSMVNLLEASYPSRFSPISVFTLHLVELKGRAHAVLVPHHQMNKLDPNT-VQSTH 535
Query: 419 VIVTFDLFEHDNAGTASVSTYTAISPVRFMHDDICYLALDKLASIIILPFHLRWSEDGSV 478
++ F FE N GT +TA +P ++DDIC LALDK A++I++PFH +++ DG+V
Sbjct: 536 IVNGFQRFEQQNQGTLMAQHFTAAAPFSSINDDICTLALDKKATLIVIPFHKQYAIDGTV 595
Query: 479 ESADETTRSLNTKVLERAPCSVAILVNRGHSSPFNH---NENSKQIAMIFLGGSDDREAL 535
+ + + R++N VLE+APCSV I ++RG + + + +A+IF+ G DD EAL
Sbjct: 596 DHVNPSIRNINLNVLEKAPCSVGIFIDRGETEGRRSVLMSYTWRNVAVIFIEGRDDAEAL 655
Query: 536 CLAKRTIKEDTYHLVVYHL---VSTIKNDEFTSWDVMLDDELLKGVKGVYGSVDNVTYEK 592
+ R + + + H S +N + + L+ K S ++Y +
Sbjct: 656 AFSMRIAEHPEVSVTMIHFRHKSSLQQNHVVDVESELAESYLINDFKNFAMSKPKISYRE 715
Query: 593 VEVENTSDTTEFISDIAIQHDFIIVGRRNGIKSPQTQALASWTEYPELGVLGDLLASPDT 652
V + +TT+ IS + D ++VGR + ++S L W+E PELGV+GD+ AS D
Sbjct: 716 EIVRDGVETTQVISSLGDSFDLVVVGRDHDLESSVLYGLTDWSECPELGVIGDMFASSDF 775
Query: 653 NTKASILVVQQQ 664
+ S+LV+ QQ
Sbjct: 776 H--FSVLVIHQQ 785
>UniRef100_O22920 Putative Na/H antiporter [Arabidopsis thaliana]
Length = 831
Score = 298 bits (763), Expect = 3e-79
Identities = 191/673 (28%), Positives = 343/673 (50%), Gaps = 29/673 (4%)
Query: 1 MDFTMITRTGRKAWTIAFCSFVIPMFFGLVVCYRFQEFWKLEMGNFEAKNLPVIVIGQSG 60
+D ++I + G KA I S+ P G + + L + + + +
Sbjct: 134 IDGSIIRKAGSKAILIGTASYAFPFSLGNLTIMFISKTMGLPSDVISCTSSAISLSSMTS 193
Query: 61 CYFAVIASLLSDLEILNSELGRLALSTAMVMDSFNSIVTGIGTAFISSIKTDSHDNGDGK 120
F V ++L++L ILNSELGRLA +MV + + V A ++ T
Sbjct: 194 --FPVTTTVLAELNILNSELGRLATHCSMVCEVCSWFV-----ALAFNLYTRDR------ 240
Query: 121 GTLKAFLNVFYYLCFMVVTPLVLRPILKWFVKKTPEGRPMKKVYMYIVFIIALAVGML-G 179
T+ + + + ++V V RPI+ W ++ + K V + ++ L++ L G
Sbjct: 241 -TMTSLYALSMIIGLLLVIYFVFRPIIVWLTQRKTKSMDKKDVVPFFPVLLLLSIASLSG 299
Query: 180 LLTKQSVLGGICIVGLIVPEGPPLGTEMIKQLELFCSWFLFPIFVTSCAMKIDLSVYVKS 239
G +G+ +P+GPPLGTE+ +LE+F S P F+ ++ + +S
Sbjct: 300 EAMGVHAAFGAFWLGVSLPDGPPLGTELAAKLEMFASNLFLPCFIAISGLQTNFFEITES 359
Query: 240 -DYIYVWLGIIVAV-HLFKMLVTIGICWYCNMPMADGLCLALMLSCKGVVDFCNNVFLHD 297
++ V + II+ + + K L T YC + D LCLA ++ C+G+++ + D
Sbjct: 360 HEHHVVMIEIILLITYGCKFLGTAAASAYCQTQIGDALCLAFLMCCQGIIEVYTTIVWKD 419
Query: 298 SMLLSSEALSVLSINVLVIGTLARIGVKYLYDPSRKYAGYQKRNILSLKP-NSELKIVSC 356
+ ++ +E +++ I +L + ++R V YLYDPS++Y KR IL+ + N +L+++
Sbjct: 420 AQVVDTECFNLVIITILFVTGISRFLVVYLYDPSKRYKSKSKRTILNTRQHNLQLRLLLG 479
Query: 357 ILKPSHIIPIKNVLDICSPTSSNPLVIHILHLLELVGRSSPVFISHRLQERVGSSSHTFS 416
+ ++ + N+L+ PT NP+ LHL+EL GR+ + H ++ ++ S
Sbjct: 480 LYNVENVPSMVNLLEATYPTRFNPISFFTLHLVELKGRAHALLTPHHQMNKLDPNTAQ-S 538
Query: 417 EAVIVTFDLFEHDNAGTASVSTYTAISPVRFMHDDICYLALDKLASIIILPFHLRWSEDG 476
++ F FE G +TA +P +++DIC LALDK A++I++PFH +++ DG
Sbjct: 539 THIVNAFQRFEQKYQGALMAQHFTAAAPYSSINNDICTLALDKKATLIVIPFHKQYAIDG 598
Query: 477 SVESADETTRSLNTKVLERAPCSVAILVNRGHSSPFNH---NENSKQIAMIFLGGSDDRE 533
+V + R++N VL+ APCSVAI ++RG + + +AM+F+GG DD E
Sbjct: 599 TVGQVNGPIRTINLNVLDAAPCSVAIFIDRGETEGRRSVLMTNTWQNVAMLFIGGKDDAE 658
Query: 534 ALCLAKRTIKEDTYHLVVYHL--VSTIKNDEFTSWDVMLDDELLKGVKGVYGSVDNVTYE 591
AL L R ++ ++ + H S +++++++ M + L+ K + + Y
Sbjct: 659 ALALCMRMAEKPDLNVTMIHFRHKSALQDEDYSD---MSEYNLISDFKSYAANKGKIHYV 715
Query: 592 KVEVENTSDTTEFISDIAIQHDFIIVGRRNGIKSPQTQALASWTEYPELGVLGDLLASPD 651
+ V + +TT+ IS + +D ++VGR + ++S L W+E PELGV+GD+L SPD
Sbjct: 716 EEIVRDGVETTQVISSLGDAYDMVLVGRDHDLESSVLYGLTDWSECPELGVIGDMLTSPD 775
Query: 652 TNTKASILVVQQQ 664
+ S+LVV QQ
Sbjct: 776 FH--FSVLVVHQQ 786
>UniRef100_Q94DV8 Na+/H+ antiporter-like protein [Oryza sativa]
Length = 875
Score = 295 bits (755), Expect = 3e-78
Identities = 202/702 (28%), Positives = 349/702 (48%), Gaps = 47/702 (6%)
Query: 1 MDFTMITRTGRKAWTIAFCSFVIPMFFGLVVCYRFQEFWKLEMGNFEAKNLPVIVIGQSG 60
MD +I R+G+KA +A +P G+ + F+ ++ + L + + S
Sbjct: 139 MDLDVIRRSGKKALFVAVAGMALPFCIGIATSFIFRH--QVSRNVHQTSFLLFLGVALSV 196
Query: 61 CYFAVIASLLSDLEILNSELGRLALSTAMVMDSFNSIVTGIGTAFISSIKTDSHDNGDGK 120
F V+A +L+++++LN+ELGR+A+S A+V D I+ + A S N
Sbjct: 197 TAFPVLARILAEIKLLNTELGRIAMSAAIVNDMCAWILLALAIAI-------SEVNSTAL 249
Query: 121 GTLKAFLN--VFYYLCFMVVTPLVLRPILKWFVKKTPEGRPMKKVYMYIVFIIALAVGML 178
+L L +F CF VV RP + W +++ PEG + + + ++ + G+
Sbjct: 250 SSLWVLLAGVLFVLFCFYVV-----RPGMWWLIRRIPEGEVVSDMQVSLILTGVMLAGVC 304
Query: 179 GLLTKQSVLGGICIVGLIVPEGPPLGTEMIKQLELFCSWFLFPIFVTSCAMKIDLSVYVK 238
+ G + GL++P G LG +I++LE F + L P+F ++ ++S
Sbjct: 305 TDAIGIHSVFGAFVYGLVIPGGQ-LGVALIEKLEDFVTGLLLPLFFAISGLRTNISKIRD 363
Query: 239 SDYIYVWLGIIVAVHLFKMLVTIGICWYCNMPMADGLCLALMLSCKGVVDFCNNVFLHDS 298
+ + + + K++ TI I MP +G+ L +++ +G+V+ D
Sbjct: 364 PITVGLLVLVFTMASFAKIMGTIIIAALYTMPFREGIALGFLMNTRGLVEMIVLNIGRDK 423
Query: 299 MLLSSEALSVLSINVLVIGTLARIGVKYLYDPSRKYAGYQKRNILSLKPNSELKIVSCIL 358
+L E+ +V+ + + + TL V +Y PSR+ GY++RN+ ++ +SEL+++ C+
Sbjct: 424 EVLDDESFAVMVLVSVAMTTLVTPVVTGVYRPSRRLVGYKRRNLQRIRHDSELRMLICVH 483
Query: 359 KPSHIIPIKNVLDICSPTSSNPLVIHILHLLELVGRSSPVFISHRL---QERVGSSSHTF 415
++ + ++L++ +PT +P+ I+ LHL+EL GR+S + + ++ SSS T
Sbjct: 484 TTRNVPSVLSLLELSNPTKRSPIFIYALHLVELTGRASNMLAAAAASASKQNRSSSSSTL 543
Query: 416 SEAVIVTFDLFEH--DNAGTASVSTYTAISPVRFMHDDICYLALDKLASIIILPFHLRWS 473
F+ FE+ + G S+ T A+SP + MHDD+ LA DK S+I++PFH + +
Sbjct: 544 PPVTEHIFNAFENYERHTGGISIQTLAAVSPYQTMHDDVSVLAEDKHVSLIVVPFHKQQT 603
Query: 474 EDGSVESADETTRSLNTKVLERAPCSVAILVNRGHSSPFNHNENSKQIAMIFLGGSDDRE 533
DG++E + + R N +L +PCSVAILV+RG S+ ++A+ F GG DDRE
Sbjct: 604 VDGAMEPINPSIRGFNESLLSTSPCSVAILVDRGLSAAAARMAALHRVALFFFGGPDDRE 663
Query: 534 ALCLAKRTIKEDTYHLVVYHLV-----------------------STIKNDEFTSWDVML 570
AL A R ++ L V V +I D ++ +
Sbjct: 664 ALAYAWRMVEHPGVALTVVRFVPPDYRVRSYSNTNYRSVASDADPRSIGIDTEGKTELQM 723
Query: 571 DDELLKGVKGVYGSVDNVTYEKVEVENTSDTTEFISDIAIQ-HDFIIVGRRNG-IKSPQT 628
D+E L + D ++Y V N+ +T I ++ H+ IVGRR G SP T
Sbjct: 724 DEEYLGDFRTRNIGNDAISYSDKVVANSEETVSAIRNMDDSLHELYIVGRRPGEAGSPMT 783
Query: 629 QALASWTEYPELGVLGDLLASPDTNTKASILVVQQQVMPKAS 670
+L W E PELG +GD+L S D + S+LVVQQ V+ A+
Sbjct: 784 ASLEDWMECPELGPIGDMLVSSDFSMSVSVLVVQQYVVAAAA 825
>UniRef100_Q942J4 B1148D12.14 protein [Oryza sativa]
Length = 952
Score = 295 bits (755), Expect = 3e-78
Identities = 202/702 (28%), Positives = 349/702 (48%), Gaps = 47/702 (6%)
Query: 1 MDFTMITRTGRKAWTIAFCSFVIPMFFGLVVCYRFQEFWKLEMGNFEAKNLPVIVIGQSG 60
MD +I R+G+KA +A +P G+ + F+ ++ + L + + S
Sbjct: 216 MDLDVIRRSGKKALFVAVAGMALPFCIGIATSFIFRH--QVSRNVHQTSFLLFLGVALSV 273
Query: 61 CYFAVIASLLSDLEILNSELGRLALSTAMVMDSFNSIVTGIGTAFISSIKTDSHDNGDGK 120
F V+A +L+++++LN+ELGR+A+S A+V D I+ + A S N
Sbjct: 274 TAFPVLARILAEIKLLNTELGRIAMSAAIVNDMCAWILLALAIAI-------SEVNSTAL 326
Query: 121 GTLKAFLN--VFYYLCFMVVTPLVLRPILKWFVKKTPEGRPMKKVYMYIVFIIALAVGML 178
+L L +F CF VV RP + W +++ PEG + + + ++ + G+
Sbjct: 327 SSLWVLLAGVLFVLFCFYVV-----RPGMWWLIRRIPEGEVVSDMQVSLILTGVMLAGVC 381
Query: 179 GLLTKQSVLGGICIVGLIVPEGPPLGTEMIKQLELFCSWFLFPIFVTSCAMKIDLSVYVK 238
+ G + GL++P G LG +I++LE F + L P+F ++ ++S
Sbjct: 382 TDAIGIHSVFGAFVYGLVIPGGQ-LGVALIEKLEDFVTGLLLPLFFAISGLRTNISKIRD 440
Query: 239 SDYIYVWLGIIVAVHLFKMLVTIGICWYCNMPMADGLCLALMLSCKGVVDFCNNVFLHDS 298
+ + + + K++ TI I MP +G+ L +++ +G+V+ D
Sbjct: 441 PITVGLLVLVFTMASFAKIMGTIIIAALYTMPFREGIALGFLMNTRGLVEMIVLNIGRDK 500
Query: 299 MLLSSEALSVLSINVLVIGTLARIGVKYLYDPSRKYAGYQKRNILSLKPNSELKIVSCIL 358
+L E+ +V+ + + + TL V +Y PSR+ GY++RN+ ++ +SEL+++ C+
Sbjct: 501 EVLDDESFAVMVLVSVAMTTLVTPVVTGVYRPSRRLVGYKRRNLQRIRHDSELRMLICVH 560
Query: 359 KPSHIIPIKNVLDICSPTSSNPLVIHILHLLELVGRSSPVFISHRL---QERVGSSSHTF 415
++ + ++L++ +PT +P+ I+ LHL+EL GR+S + + ++ SSS T
Sbjct: 561 TTRNVPSVLSLLELSNPTKRSPIFIYALHLVELTGRASNMLAAAAASASKQNRSSSSSTL 620
Query: 416 SEAVIVTFDLFEH--DNAGTASVSTYTAISPVRFMHDDICYLALDKLASIIILPFHLRWS 473
F+ FE+ + G S+ T A+SP + MHDD+ LA DK S+I++PFH + +
Sbjct: 621 PPVTEHIFNAFENYERHTGGISIQTLAAVSPYQTMHDDVSVLAEDKHVSLIVVPFHKQQT 680
Query: 474 EDGSVESADETTRSLNTKVLERAPCSVAILVNRGHSSPFNHNENSKQIAMIFLGGSDDRE 533
DG++E + + R N +L +PCSVAILV+RG S+ ++A+ F GG DDRE
Sbjct: 681 VDGAMEPINPSIRGFNESLLSTSPCSVAILVDRGLSAAAARMAALHRVALFFFGGPDDRE 740
Query: 534 ALCLAKRTIKEDTYHLVVYHLV-----------------------STIKNDEFTSWDVML 570
AL A R ++ L V V +I D ++ +
Sbjct: 741 ALAYAWRMVEHPGVALTVVRFVPPDYRVRSYSNTNYRSVASDADPRSIGIDTEGKTELQM 800
Query: 571 DDELLKGVKGVYGSVDNVTYEKVEVENTSDTTEFISDIAIQ-HDFIIVGRRNG-IKSPQT 628
D+E L + D ++Y V N+ +T I ++ H+ IVGRR G SP T
Sbjct: 801 DEEYLGDFRTRNIGNDAISYSDKVVANSEETVSAIRNMDDSLHELYIVGRRPGEAGSPMT 860
Query: 629 QALASWTEYPELGVLGDLLASPDTNTKASILVVQQQVMPKAS 670
+L W E PELG +GD+L S D + S+LVVQQ V+ A+
Sbjct: 861 ASLEDWMECPELGPIGDMLVSSDFSMSVSVLVVQQYVVAAAA 902
>UniRef100_Q9LUN4 Na+/H+ exchangeing protein-like [Arabidopsis thaliana]
Length = 800
Score = 259 bits (662), Expect = 2e-67
Identities = 186/683 (27%), Positives = 343/683 (49%), Gaps = 47/683 (6%)
Query: 1 MDFTMITRTGRKAWTIAFCSFVIPMFFGLVVCYRFQEFWKLEMGNFEAKNLPVIV---IG 57
+DF I +TG+K+ IA +P G+ + + G LP IV +
Sbjct: 114 LDFAAIKKTGKKSLLIAIAGISLPFIVGVGTSFVLSA--TISKG---VDQLPFIVFMGVA 168
Query: 58 QSGCYFAVIASLLSDLEILNSELGRLALSTAMVMDSFNSIVTGIGTAFISSIKTDSHDNG 117
S F V+A +L++L++L +++GR+A+S A V D I+ + A +G
Sbjct: 169 LSITAFPVLARILAELKLLTTDIGRMAMSAAGVNDVAAWILLALAIAL----------SG 218
Query: 118 DGKGTLKAFLNVFYYLC---FMVVTPLVLRPILKWFVKKTPEGRPMKKVYMYIVFIIALA 174
DG L ++V+ LC F++ + ++P+L + ++ PEG P+K++Y+ + + LA
Sbjct: 219 DGTSPL---VSVWVLLCGTGFVIFAVVAIKPLLAYMARRCPEGEPVKELYVCVTLTVVLA 275
Query: 175 VGMLGLLTKQSVLGGICIVGLIVPEGPPLGTEMIKQLELFCSWFLFPIFVTSCAMKIDLS 234
+ L G +VG++ P+ P + +++E S L P++ + +K D++
Sbjct: 276 ASFVTDTIGIHALFGAFVVGIVAPKEGPFCRILTEKIEDLVSGLLLPLYFAASGLKTDVT 335
Query: 235 VYVKSDYIYVWLGIIVAVHLFKMLVTIGICWYCNMPMADGLCLALMLSCKGVVDFCNNVF 294
+ + + +I+ K++ T+G C +P + + L +++ KG+V+
Sbjct: 336 TIRGAQSWGLLVLVILTTCFGKIVGTVGSSMLCKVPFREAVTLGFLMNTKGLVELIVLNI 395
Query: 295 LHDSMLLSSEALSVLSINVLVIGTLARIGVKYLYDPSRKYAGYQKRNILSLKPNSELKIV 354
D +L+ +A ++L + L + V +Y P+RK A Y+ R I +SEL+I+
Sbjct: 396 GKDRKVLNDQAFAILVLMALFTTFITTPIVMLIYKPARKGAPYKHRTIQRKDHDSELRIL 455
Query: 355 SCILKPSHIIPIKNVLDICSPTSSNP-LVIHILHLLELVGRSSPVFISHRLQER---VGS 410
+C +I + N+++ T L ++ +HL+EL RSS + + H+ + + +
Sbjct: 456 ACFHSTRNIPTLINLIESSRGTGKKGRLCVYAMHLMELSERSSAIAMVHKARNNGLPIWN 515
Query: 411 SSHTFSEAVIVTFDLFEHDNAGTASVSTYTAISPVRFMHDDICYLALDKLASIIILPFHL 470
++ +++ F+ ++H A +V TAIS + +H+DIC A K ++I+LPFH
Sbjct: 516 KIERSTDQMVIAFEAYQHLRA--VAVRPMTAISGLSSIHEDICTSAHQKRVAMILLPFHK 573
Query: 471 RWSEDGSVESADETTRSLNTKVLERAPCSVAILVNR--GHSSPFNHNENSKQIAMIFLGG 528
DG++ES +N +VL+RAPCSV ILV+R G +S +E + ++ + F GG
Sbjct: 574 HQRMDGAMESIGHRFHEVNQRVLQRAPCSVGILVDRGLGGTSQVVASEVAYKVVIPFFGG 633
Query: 529 SDDREALCLAKRTIKEDTYHLVVYHLVS---TIK------NDEFTSWDVMLDDELLKGVK 579
DDREAL + ++ L VY V+ T+K +DE + D+E ++ +
Sbjct: 634 LDDREALAYGMKMVEHPGITLTVYKFVAARGTLKRFEKSEHDEKEKKEKETDEEFVRELM 693
Query: 580 GVYGSVDNVTYEKVEVENTSDTTEFISDIAIQHDFIIVGRRNGIKSPQTQALASWTEYPE 639
+++ YE+ VE+ D + ++ + + +VGR + S L T+ PE
Sbjct: 694 NDPRGNESLAYEERVVESKDDIIATLKSMS-KCNLFVVGRNAAVAS-----LVKSTDCPE 747
Query: 640 LGVLGDLLASPDTNTKASILVVQ 662
LG +G LL+S + +T AS+LVVQ
Sbjct: 748 LGPVGRLLSSSEFSTTASVLVVQ 770
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.324 0.139 0.417
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,084,200,948
Number of Sequences: 2790947
Number of extensions: 45294105
Number of successful extensions: 140178
Number of sequences better than 10.0: 131
Number of HSP's better than 10.0 without gapping: 50
Number of HSP's successfully gapped in prelim test: 81
Number of HSP's that attempted gapping in prelim test: 139788
Number of HSP's gapped (non-prelim): 174
length of query: 670
length of database: 848,049,833
effective HSP length: 134
effective length of query: 536
effective length of database: 474,062,935
effective search space: 254097733160
effective search space used: 254097733160
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.5 bits)
S2: 78 (34.7 bits)
Medicago: description of AC146343.3


