

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146342.5 + phase: 0
(507 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
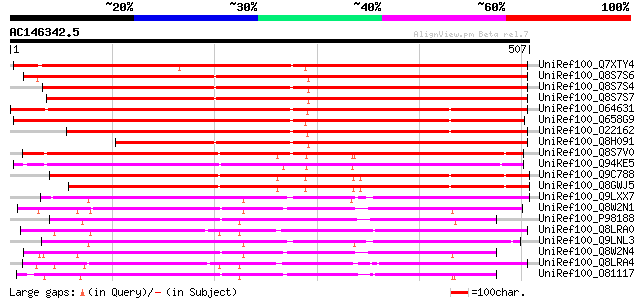
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q7XTY4 OSJNBa0019K04.13 protein [Oryza sativa] 566 e-160
UniRef100_Q8S7S6 Cytochrome P450-like protein [Oryza sativa] 555 e-157
UniRef100_Q8S7S4 Cytochrome P450-like protein [Oryza sativa] 548 e-154
UniRef100_Q8S7S7 Cytochrome P450-like protein [Oryza sativa] 536 e-151
UniRef100_O64631 Putative cytochrome P450 [Arabidopsis thaliana] 515 e-144
UniRef100_Q658G9 Putative cytochrome P450 [Oryza sativa] 500 e-140
UniRef100_O22162 Putative cytochrome P450 [Arabidopsis thaliana] 485 e-135
UniRef100_Q8H091 Putative cytochrome P450-dependent fatty acid h... 476 e-133
UniRef100_Q8S7V0 Putative plant cytochrome P-450 protein [Oryza ... 388 e-106
UniRef100_Q94KE5 Cytochrome P450-like protein [Zea mays] 387 e-106
UniRef100_Q9C788 Cytochrome P450, putative [Arabidopsis thaliana] 386 e-106
UniRef100_Q8GWJ5 Hypothetical protein At1g69500/F10D13_15 [Arabi... 371 e-101
UniRef100_Q9LXX7 Cytochrome P450-like protein [Arabidopsis thali... 321 3e-86
UniRef100_Q8W2N1 Cytochrome P450-dependent fatty acid hydroxylas... 320 8e-86
UniRef100_P98188 Cytochrome P450 94A2 [Vicia sativa] 316 1e-84
UniRef100_Q8LRA0 Putative cytochrome P450-dependent fatty acid h... 310 6e-83
UniRef100_Q9LNL3 F12K21.15 [Arabidopsis thaliana] 309 1e-82
UniRef100_Q8W2N4 Cytochrome P450-dependent fatty acid hydroxylas... 308 2e-82
UniRef100_Q8LRA4 Putative cytochrome P450-dependent fatty acid h... 307 5e-82
UniRef100_O81117 Cytochrome P450 94A1 [Vicia sativa] 301 3e-80
>UniRef100_Q7XTY4 OSJNBa0019K04.13 protein [Oryza sativa]
Length = 506
Score = 566 bits (1458), Expect = e-160
Identities = 282/510 (55%), Positives = 378/510 (73%), Gaps = 11/510 (2%)
Query: 4 LSNPLLFAALSASLTLLLVQLLLRKLNNKSNSMKKKKYHPVAGTVFNQMMNFNRLHHYMT 63
+ +PL A+ A LLLV L L + ++ KK++Y PVAGT+F+Q++NF RL Y T
Sbjct: 1 MESPLSHPAMVALSLLLLVALYLAR---RAVLGKKRRYPPVAGTMFHQLLNFGRLLEYHT 57
Query: 64 DLARKYKTYRLLNPFRSEVYTSEPSNVEYILKTNFENYGKGLYNYQNLKDLLGDGIFTVD 123
+L+RKY+T+R+L P + VYT EP+NVE+ILKTNF NYGKG + L+DLLGDGIF VD
Sbjct: 58 ELSRKYRTFRMLTPTCNYVYTVEPANVEHILKTNFANYGKGPMTHDVLEDLLGDGIFNVD 117
Query: 124 GEKWREQRKISSHEFSTRMLRDFSTSIFRKNAAKVANIVSE--AANSNTKLEIQDIFMKS 181
G WR+QRK++S EFSTR+LRD+S+++FR AA++A I+ AA ++++QD+ M++
Sbjct: 118 GGMWRQQRKVASLEFSTRVLRDYSSAVFRDTAAELAGILERGPAAKGRERVDMQDLLMRA 177
Query: 182 TLDSIFNVVFGTEIDSMCGTSEEGKNFANSFDNASALTLYRYVDVFWKIKKFLNIGSEAA 241
TLDS F V FG + + G+S+EG FA +FD+AS L+R+ D+ WK+K+FLNI SEA
Sbjct: 178 TLDSFFRVGFGVNLGVLSGSSKEGLVFARAFDDASEQVLFRFFDLLWKVKRFLNISSEAT 237
Query: 242 LRNNTEILNEFVIKLINTRIQQMKNSKGDSVRKGGDILSRFLQVKE-----YDTKYLRDI 296
++ + +N+FV +I+ +I+QM + + +K DILSRFL +E +D KY+RDI
Sbjct: 238 MKQSIRTINDFVYSIIDRKIEQMSREQHEFAKKE-DILSRFLLEREKDPGCFDNKYIRDI 296
Query: 297 ILNFVIAGKDTTGGTLSWFMYMLCKYPAVQEKAAQEVREATNTKTVSSCTEFVSSVTDEA 356
ILNFVIAG+DTT GTLSWF+Y +CK VQ+K A+EVR+AT +F S +T++A
Sbjct: 297 ILNFVIAGRDTTAGTLSWFLYAVCKNQRVQDKIAREVRDATTGDRDVGVQDFSSFLTEDA 356
Query: 357 IEKMNYVHAVLTETLRLYPALPFDAKICFADDTLPDGYSVKKGDMVSYQPYAMGRMKFIW 416
I KM Y+HA LTETLRLYP +P D K CF+DDTLPDG++VKKGDMV+YQPY MGRMKF+W
Sbjct: 357 INKMQYLHAALTETLRLYPGVPLDVKYCFSDDTLPDGHAVKKGDMVNYQPYPMGRMKFLW 416
Query: 417 GDDAEEFRPERWLDENGIFQPECPFKFTAFQAGPRICLGKEFAYRQMKIFSAVLLGCFRF 476
GD+AEEF+PERWLD++G+F E PFKFTAFQAGPRICLGKEFAYRQMKI SAVLL FRF
Sbjct: 417 GDNAEEFKPERWLDDSGMFVAESPFKFTAFQAGPRICLGKEFAYRQMKIVSAVLLYFFRF 476
Query: 477 KLNDEKKNVTYKTMITLHIDGGLEIKALYR 506
++ D+ V Y+ M+TL +DG ++AL R
Sbjct: 477 EMWDDDATVGYRPMLTLKMDGPFYLRALAR 506
>UniRef100_Q8S7S6 Cytochrome P450-like protein [Oryza sativa]
Length = 515
Score = 555 bits (1431), Expect = e-157
Identities = 269/502 (53%), Positives = 369/502 (72%), Gaps = 10/502 (1%)
Query: 14 SASLTLLLVQLL---LRKLNNKSNSMKKKKYHPVAGTVFNQMMNFNRLHHYMTDLARKYK 70
+A++ L+LV + L + + ++++ PV GT F+Q+ + R+H Y T L+R++
Sbjct: 15 AAAVGLVLVVAICTYLAVVATRKQKRRRRRRPPVVGTAFHQLYHVRRVHDYHTALSREHM 74
Query: 71 TYRLLNPF-RSEVYTSEPSNVEYILKTNFENYGKGLYNYQNLKDLLGDGIFTVDGEKWRE 129
T+RLL P R ++YT +P+ VE+IL+TNF NYGKG +N+ N+ DL GDGIF VDG+KW++
Sbjct: 75 TFRLLVPAGREQIYTCDPAVVEHILRTNFANYGKGSFNHGNMSDLFGDGIFAVDGDKWKQ 134
Query: 130 QRKISSHEFSTRMLRDFSTSIFRKNAAKVANIVSEAANSNTKLEIQDIFMKSTLDSIFNV 189
QRKI+S++F+TR LRDFS +F++NAAK+A +VS A SN ++ Q M++T+DSIF +
Sbjct: 135 QRKIASYDFTTRALRDFSGDVFKRNAAKLAGVVSSHAASNQSMDFQGFLMRATMDSIFTI 194
Query: 190 VFGTEIDSMCGTSEEGKNFANSFDNASALTLYRYVDVFWKIKKFLNIGSEAALRNNTEIL 249
FG +++++ G S EG+ FA +FD+AS T+ RY++ FWK+ + LN+G+EA L+ +++
Sbjct: 195 AFGQDLNTLDG-SGEGRRFAAAFDDASEFTMLRYLNPFWKLSRLLNVGAEAMLKERIKVV 253
Query: 250 NEFVIKLINTRIQQMKNSKGDSVRKGGDILSRFLQVKEYDT-----KYLRDIILNFVIAG 304
+ FV KLI R ++ N+K DIL+RF+Q D+ KYLRDIILN VIAG
Sbjct: 254 DGFVYKLIRDRSDELSNTKAHDTDSRQDILTRFIQATTSDSGTVDYKYLRDIILNIVIAG 313
Query: 305 KDTTGGTLSWFMYMLCKYPAVQEKAAQEVREATNTKTVSSCTEFVSSVTDEAIEKMNYVH 364
KDTT G+L+WF+YM+CK+P VQEK E EATN +S EF S+TDEA+ KM+Y+H
Sbjct: 314 KDTTAGSLAWFLYMMCKHPEVQEKICHEAMEATNAGEAASIDEFSQSLTDEALNKMHYLH 373
Query: 365 AVLTETLRLYPALPFDAKICFADDTLPDGYSVKKGDMVSYQPYAMGRMKFIWGDDAEEFR 424
A LTETLRLYPA+P D K CF+DD LP+G++V KGD+V Y PYAMGRM+ +WG DAE FR
Sbjct: 374 AALTETLRLYPAVPLDNKQCFSDDVLPNGFNVSKGDIVFYIPYAMGRMESLWGKDAESFR 433
Query: 425 PERWLDENGIFQPECPFKFTAFQAGPRICLGKEFAYRQMKIFSAVLLGCFRFKLNDEKKN 484
PERWLDENG+FQ E PFKFTAFQAGPRICLGK+FAYRQMKIF+AVLL F KL DEK+
Sbjct: 434 PERWLDENGVFQQESPFKFTAFQAGPRICLGKDFAYRQMKIFAAVLLRFFVLKLRDEKEI 493
Query: 485 VTYKTMITLHIDGGLEIKALYR 506
++Y+TMITL +D GL + A+ R
Sbjct: 494 ISYRTMITLSVDQGLHLTAMAR 515
>UniRef100_Q8S7S4 Cytochrome P450-like protein [Oryza sativa]
Length = 495
Score = 548 bits (1413), Expect = e-154
Identities = 267/479 (55%), Positives = 356/479 (73%), Gaps = 8/479 (1%)
Query: 33 SNSMKKKKYHPVAGTVFNQMMNFNRLHHYMTDLARKYKTYRLLNPFRSE-VYTSEPSNVE 91
SN K+++ PV GTVF+Q+ N R+H Y T L+R++ T+R+L P + +YT +P+ VE
Sbjct: 20 SNKQKRRRRPPVVGTVFHQLYNVRRIHDYHTALSREHTTFRMLVPAGGDQIYTCDPAVVE 79
Query: 92 YILKTNFENYGKGLYNYQNLKDLLGDGIFTVDGEKWREQRKISSHEFSTRMLRDFSTSIF 151
+ILKTNF NYGKG +N+ N KDL GDGIF +DGEKW++QRKI+S++FSTR LRDFS ++F
Sbjct: 80 HILKTNFANYGKGPFNHGNAKDLFGDGIFAIDGEKWKQQRKIASYDFSTRALRDFSCAVF 139
Query: 152 RKNAAKVANIVSEAANSNTKLEIQDIFMKSTLDSIFNVVFGTEIDSMCGTSEEGKNFANS 211
++NAAK+A IVS A SN ++ Q + +++T+DSIF + FGT+++++ G S EG FA +
Sbjct: 140 KRNAAKLAGIVSNHAASNQSMDFQGLMLRATMDSIFTIAFGTDLNTLDG-SGEGSRFAAA 198
Query: 212 FDNASALTLYRYVDVFWKIKKFLNIGSEAALRNNTEILNEFVIKLINTRIQQMKNSKGDS 271
FD+AS T+ RY+ WK+ + LN+G EA L+ ++++EFV +LI R ++ NS
Sbjct: 199 FDDASEFTMLRYISPLWKLARLLNVGVEAMLKERIKVVDEFVYRLIRARSDELSNSHDSG 258
Query: 272 VRKGGDILSRFLQVKEYDT----KYLRDIILNFVIAGKDTTGGTLSWFMYMLCKYPAVQE 327
R+ DILSRFLQ D+ KYLRDIILN VIAGKDTT G L+WF+YM+CK+P VQE
Sbjct: 259 SRQ--DILSRFLQATTSDSVVDYKYLRDIILNIVIAGKDTTAGALAWFLYMVCKHPEVQE 316
Query: 328 KAAQEVREATNTKTVSSCTEFVSSVTDEAIEKMNYVHAVLTETLRLYPALPFDAKICFAD 387
K E AT+ +S EF+ S+TD+A+ M+Y+HA LTETLRLYP++P + K CF+D
Sbjct: 317 KICHEAMVATSAGDTASVDEFLQSLTDQALNNMHYLHAALTETLRLYPSVPMENKQCFSD 376
Query: 388 DTLPDGYSVKKGDMVSYQPYAMGRMKFIWGDDAEEFRPERWLDENGIFQPECPFKFTAFQ 447
D LP+G++V KGD+V + PYAMGRM+ +WG DAE FRPERWLDENG+FQ E PFKFTAFQ
Sbjct: 377 DVLPNGFNVSKGDIVFFIPYAMGRMESLWGKDAEYFRPERWLDENGVFQQESPFKFTAFQ 436
Query: 448 AGPRICLGKEFAYRQMKIFSAVLLGCFRFKLNDEKKNVTYKTMITLHIDGGLEIKALYR 506
AGPRICLGKEFAYRQMKIF+AVLL F KL DEK+ V+Y+T +TL ID GL + A R
Sbjct: 437 AGPRICLGKEFAYRQMKIFAAVLLRFFVLKLRDEKEIVSYRTTLTLAIDQGLHLTATAR 495
>UniRef100_Q8S7S7 Cytochrome P450-like protein [Oryza sativa]
Length = 516
Score = 536 bits (1382), Expect = e-151
Identities = 257/476 (53%), Positives = 352/476 (72%), Gaps = 7/476 (1%)
Query: 37 KKKKYHPVAGTVFNQMMNFNRLHHYMTDLARKYKTYRLL-NPFRSEVYTSEPSNVEYILK 95
+++++ PV GTVF+Q+ + RLH Y T L R++ T+RLL P R +YT +P+ VE+IL+
Sbjct: 42 RRRRHPPVVGTVFHQLYHVRRLHDYYTALCREHTTFRLLATPGRRNIYTCDPAVVEHILR 101
Query: 96 TNFENYGKGLYNYQNLKDLLGDGIFTVDGEKWREQRKISSHEFSTRMLRDFSTSIFRKNA 155
TNF +YGKG N + L DL G+GIF VDGEKW+ QRKI+S++F+TR LRDFS+ +F++NA
Sbjct: 102 TNFPSYGKGPLNSEILNDLFGEGIFAVDGEKWKTQRKIASYDFTTRALRDFSSDVFKRNA 161
Query: 156 AKVANIVSEAANSNTKLEIQDIFMKSTLDSIFNVVFGTEIDSMCGTSEEGKNFANSFDNA 215
AK+A +VS A SN ++ + + ++T+DSIF + FG +++++ G S EG++FA +FD+A
Sbjct: 162 AKLAGVVSNHAASNQSMDFKGLLTRATMDSIFTIAFGQDLNTLDG-SGEGRHFAKAFDDA 220
Query: 216 SALTLYRYVDVFWKIKKFLNIGSEAALRNNTEILNEFVIKLINTRIQQMKNSKGDSVRKG 275
L RY++ FWK+ + LN+G+EA L+ ++++EFV KLI R ++ N+ R
Sbjct: 221 GEYLLLRYLNPFWKLARLLNVGAEATLKERIKVVDEFVYKLIRARSDELSNTMAQDHRSR 280
Query: 276 GDILSRFLQVKEYDT-----KYLRDIILNFVIAGKDTTGGTLSWFMYMLCKYPAVQEKAA 330
D+LSRF+Q D+ KYLRDI+LN VIA KD+T G+L+WF+YM CK P VQEK
Sbjct: 281 DDLLSRFIQATTSDSGTVDYKYLRDIVLNIVIAAKDSTSGSLAWFLYMACKRPEVQEKIF 340
Query: 331 QEVREATNTKTVSSCTEFVSSVTDEAIEKMNYVHAVLTETLRLYPALPFDAKICFADDTL 390
EV E TN +S EF++S+TD+A+ KM+Y+HA LTETLRLYP++P + K CF+DD L
Sbjct: 341 DEVMETTNAGDCASIDEFLTSLTDQALNKMHYLHAALTETLRLYPSVPLENKQCFSDDVL 400
Query: 391 PDGYSVKKGDMVSYQPYAMGRMKFIWGDDAEEFRPERWLDENGIFQPECPFKFTAFQAGP 450
P+G+SV KGD V Y PYAMGRM+F+WG DAE FRPERWLDE+G+FQ E PFKFTAFQAGP
Sbjct: 401 PNGFSVSKGDGVFYMPYAMGRMEFLWGKDAEAFRPERWLDEHGVFQQESPFKFTAFQAGP 460
Query: 451 RICLGKEFAYRQMKIFSAVLLGCFRFKLNDEKKNVTYKTMITLHIDGGLEIKALYR 506
RIC+GK+FAYRQMKIF+AVL+ F FKL D+K NV+Y+T ITL ID L + A R
Sbjct: 461 RICIGKDFAYRQMKIFAAVLIRSFVFKLRDKKDNVSYRTAITLAIDQDLHLTATAR 516
>UniRef100_O64631 Putative cytochrome P450 [Arabidopsis thaliana]
Length = 511
Score = 515 bits (1326), Expect = e-144
Identities = 263/513 (51%), Positives = 355/513 (68%), Gaps = 12/513 (2%)
Query: 1 MDFLSNPLLFAALSASLTLLL-VQLLLRKLNNKSNSMKKKKYHPVAGTVFNQMMNFNRLH 59
M+ L++ + A + + L + L++R KS + K+Y PV TVF+ + + + L+
Sbjct: 1 MEILTSIAITVATTIFIVLCFTIYLMIRIFTGKSRN--DKRYAPVHATVFDLLFHSDELY 58
Query: 60 HYMTDLARKYKTYRLLNPFRSEVYTSEPSNVEYILKTNFENYGKGLYNYQNLKDLLGDGI 119
Y T++AR+ TYR L+P +SE+ T++P NVE+ILKT F+NY KG + +N+ DLLG GI
Sbjct: 59 DYETEIAREKPTYRFLSPGQSEILTADPRNVEHILKTRFDNYSKGHSSRENMADLLGHGI 118
Query: 120 FTVDGEKWREQRKISSHEFSTRMLRDFSTSIFRKNAAKVANIVSEAANSNTKLEIQDIFM 179
F VDGEKWR+QRK+SS EFSTR+LRDFS S+FR+NA+K+ VSE A S + QD+ M
Sbjct: 119 FAVDGEKWRQQRKLSSFEFSTRVLRDFSCSVFRRNASKLVGFVSEFALSGKAFDAQDLLM 178
Query: 180 KSTLDSIFNVVFGTEIDSMCGTSEEGKNFANSFDNASALTLYRYVDVFWKIKKFLNIGSE 239
+ TLDSIF V FG E+ + G S+EG+ F +FD + T R++D WK+K F NIGS+
Sbjct: 179 RCTLDSIFKVGFGVELKCLDGFSKEGQEFMEAFDEGNVATSSRFIDPLWKLKWFFNIGSQ 238
Query: 240 AALRNNTEILNEFVIKLINTRIQQMKNSKGDSVRKGGDILSRFLQVKEYD-----TKYLR 294
+ L+ + +++FV LI T+ +++ + VR+ DILSRFL E D KYLR
Sbjct: 239 SKLKKSIATIDKFVYSLITTKRKELAKEQNTVVRE--DILSRFLVESEKDPENMNDKYLR 296
Query: 295 DIILNFVIAGKDTTGGTLSWFMYMLCKYPAVQEKAAQEVREAT-NTKTVSSCTEFVSSVT 353
DIILNF+IAGKDTT LSWF+YMLCK P VQEK QE+R+ T + + + FV S+
Sbjct: 297 DIILNFMIAGKDTTAALLSWFLYMLCKNPLVQEKIVQEIRDVTFSHEKTTDVNGFVESIN 356
Query: 354 DEAIEKMNYVHAVLTETLRLYPALPFDAKICFADDTLPDGYSVKKGDMVSYQPYAMGRMK 413
+EA+++M+Y+HA L+ETLRLYP +P D + DD LPDG+ V KGD + Y YAMGRM
Sbjct: 357 EEALDEMHYLHAALSETLRLYPPVPVDMRCAENDDVLPDGHRVSKGDNIYYIAYAMGRMT 416
Query: 414 FIWGDDAEEFRPERWLDENGIFQPECPFKFTAFQAGPRICLGKEFAYRQMKIFSAVLLGC 473
+IWG DAEEF+PERWL ++G+FQPE PFKF +F AGPRICLGK+FAYRQMKI S LL
Sbjct: 417 YIWGQDAEEFKPERWL-KDGLFQPESPFKFISFHAGPRICLGKDFAYRQMKIVSMALLHF 475
Query: 474 FRFKLNDEKKNVTYKTMITLHIDGGLEIKALYR 506
FRFK+ DE V YK M+TLH+DGGL + A+ R
Sbjct: 476 FRFKMADENSKVYYKRMLTLHVDGGLHLCAIPR 508
>UniRef100_Q658G9 Putative cytochrome P450 [Oryza sativa]
Length = 525
Score = 500 bits (1288), Expect = e-140
Identities = 251/506 (49%), Positives = 351/506 (68%), Gaps = 9/506 (1%)
Query: 4 LSNPLLFAALSASLTLLLVQLLLRKLNNKSNSMKKKKYHPVAGTVFNQMMNFNRLHHYMT 63
L++ L A LS +L ++ V L K + + P+ GTVF Q+ NF+R+
Sbjct: 16 LASALAIALLSIALYIIGVVASFAVLCAKEFAERAHDRPPLVGTVFRQLKNFDRMFDEHV 75
Query: 64 DLARKYKTYRLLNPFRSEVYTSEPSNVEYILKTNFENYGKGLYNYQNLKDLLGDGIFTVD 123
+ A ++T R++ P EV+TS+P+ VE++LK +F Y KG + +KDL GDGIF D
Sbjct: 76 NYATAHRTSRIVYPGHCEVFTSDPAVVEHVLKNSFSKYSKGDFLTTAMKDLFGDGIFATD 135
Query: 124 GEKWREQRKISSHEFSTRMLRDFSTSIFRKNAAKVANIVSEAANSNTKLEIQDIFMKSTL 183
G+ WR QRK++S+EFST++LRDFS+ FR+NAAK+A +S AA + + IQD+ M++T+
Sbjct: 136 GDMWRHQRKLASYEFSTKVLRDFSSDTFRRNAAKLAEKISCAAANRISINIQDLLMRATM 195
Query: 184 DSIFNVVFGTEIDSMCGTSEEGKNFANSFDNASALTLYRYVDVFWKIKKFLNIGSEAALR 243
DSIF V FG E++++ G+ E G F+ +FD A++L YR+VD+ WK+K++LNIGSEA L+
Sbjct: 196 DSIFKVGFGFELNTLSGSDESGIQFSKAFDEANSLVYYRFVDIMWKLKRYLNIGSEAKLK 255
Query: 244 NNTEILNEFVIKLINTRIQQMKNSKGDSVRKGGDILSRFLQVKE-----YDTKYLRDIIL 298
N +I++ FV+KLI+ + +QMK + ++ DILSRF+ E D +YLRDI+L
Sbjct: 256 RNIQIIDSFVMKLIHQKREQMKIAADYKTKE--DILSRFVLASEQDPGTMDDRYLRDIVL 313
Query: 299 NFVIAGKDTTGGTLSWFMYMLCKYPAVQEKAAQEVREATN-TKTVSSCTEFVSSVTDEAI 357
NF+IAGKDTTG TL+WF Y+LCK P VQ+K A E+RE +K ++ F + + AI
Sbjct: 314 NFLIAGKDTTGNTLTWFFYLLCKNPIVQDKVALEIREFVEWSKEDNTIESFTKRLDEGAI 373
Query: 358 EKMNYVHAVLTETLRLYPALPFDAKICFADDTLPDGYSVKKGDMVSYQPYAMGRMKFIWG 417
KM+Y+ A ++ETLRLYPA+P DAK+ DD LP+GY V KGD ++Y YAMGRM ++WG
Sbjct: 374 SKMHYLQATISETLRLYPAVPVDAKMADEDDVLPNGYRVVKGDGINYMIYAMGRMTYLWG 433
Query: 418 DDAEEFRPERWLDENGIFQPECPFKFTAFQAGPRICLGKEFAYRQMKIFSAVLLGCFRFK 477
+DA+EFRPERWL NG++Q E PFKF +F AGPRICLGKEFA+RQMKI +A L+ F+F+
Sbjct: 434 EDAQEFRPERWL-VNGVYQQESPFKFVSFNAGPRICLGKEFAHRQMKIMAATLIHFFKFR 492
Query: 478 LNDEKKNVTYKTMITLHIDGGLEIKA 503
L DE K YKTM TLHID GL + A
Sbjct: 493 LEDESKEPIYKTMFTLHIDNGLHLLA 518
>UniRef100_O22162 Putative cytochrome P450 [Arabidopsis thaliana]
Length = 490
Score = 485 bits (1248), Expect = e-135
Identities = 242/457 (52%), Positives = 323/457 (69%), Gaps = 9/457 (1%)
Query: 56 NRLHHYMTDLARKYKTYRLLNPFRSEVYTSEPSNVEYILKTNFENYGKGLYNYQNLKDLL 115
++L+ Y T++AR T+R L+P +SE++T++P NVE+ILKT F NY KG NL DLL
Sbjct: 34 HKLYDYETEIARTKPTFRFLSPGQSEIFTADPRNVEHILKTRFHNYSKGPVGTVNLADLL 93
Query: 116 GDGIFTVDGEKWREQRKISSHEFSTRMLRDFSTSIFRKNAAKVANIVSEAANSNTKLEIQ 175
G GIF VDGEKW++QRK+ S EFSTR+LR+FS S+FR +A+K+ ++E A S + Q
Sbjct: 94 GHGIFAVDGEKWKQQRKLVSFEFSTRVLRNFSYSVFRTSASKLVGFIAEFALSGKSFDFQ 153
Query: 176 DIFMKSTLDSIFNVVFGTEIDSMCGTSEEGKNFANSFDNASALTLYRYVDVFWKIKKFLN 235
D+ MK TLDSIF V FG E+ + G S+EG+ F +FD + T R D FWK+K FLN
Sbjct: 154 DMLMKCTLDSIFKVGFGVELGCLDGFSKEGEEFMKAFDEGNGATSSRVTDPFWKLKCFLN 213
Query: 236 IGSEAALRNNTEILNEFVIKLINTRIQQMKNSKGDSVRKGGDILSRFLQVKEYD-----T 290
IGSE+ L+ + I+++FV LI T+ +++ + SVR+ DILS+FL E D
Sbjct: 214 IGSESRLKKSIAIIDKFVYSLITTKRKELSKEQNTSVRE--DILSKFLLESEKDPENMND 271
Query: 291 KYLRDIILNFVIAGKDTTGGTLSWFMYMLCKYPAVQEKAAQEVREATNT-KTVSSCTEFV 349
KYLRDIILN ++AGKDTT +LSWF+YMLCK P VQEK QE+R+ T++ + + F+
Sbjct: 272 KYLRDIILNVMVAGKDTTAASLSWFLYMLCKNPLVQEKIVQEIRDVTSSHEKTTDVNGFI 331
Query: 350 SSVTDEAIEKMNYVHAVLTETLRLYPALPFDAKICFADDTLPDGYSVKKGDMVSYQPYAM 409
SVT+EA+ +M Y+HA L+ET+RLYP +P + DD LPDG+ V KGD + Y YAM
Sbjct: 332 ESVTEEALAQMQYLHAALSETMRLYPPVPEHMRCAENDDVLPDGHRVSKGDNIYYISYAM 391
Query: 410 GRMKFIWGDDAEEFRPERWLDENGIFQPECPFKFTAFQAGPRICLGKEFAYRQMKIFSAV 469
GRM +IWG DAEEF+PERWL ++G+FQPE FKF +F AGPRIC+GK+FAYRQMKI S
Sbjct: 392 GRMTYIWGQDAEEFKPERWL-KDGVFQPESQFKFISFHAGPRICIGKDFAYRQMKIVSMA 450
Query: 470 LLGCFRFKLNDEKKNVTYKTMITLHIDGGLEIKALYR 506
LL FRFK+ DE V+YK M+TLH+DGGL + A+ R
Sbjct: 451 LLHFFRFKMADENSKVSYKKMLTLHVDGGLHLCAIPR 487
>UniRef100_Q8H091 Putative cytochrome P450-dependent fatty acid hydroxylase,
5'-partial [Oryza sativa]
Length = 404
Score = 476 bits (1224), Expect = e-133
Identities = 231/407 (56%), Positives = 304/407 (73%), Gaps = 7/407 (1%)
Query: 104 GLYNYQNLKDLLGDGIFTVDGEKWREQRKISSHEFSTRMLRDFSTSIFRKNAAKVANIVS 163
G +N+ N KDL GDGIF +DGEKW++QRKI+S++FSTR LRDFS ++F++NAAK+A IVS
Sbjct: 1 GPFNHGNAKDLFGDGIFAIDGEKWKQQRKIASYDFSTRALRDFSCAVFKRNAAKLAGIVS 60
Query: 164 EAANSNTKLEIQDIFMKSTLDSIFNVVFGTEIDSMCGTSEEGKNFANSFDNASALTLYRY 223
A SN ++ Q + +++T+DSIF + FGT+++++ G S EG FA +FD+AS T+ RY
Sbjct: 61 NHAASNQSMDFQGLMLRATMDSIFTIAFGTDLNTLDG-SGEGSRFAAAFDDASEFTMLRY 119
Query: 224 VDVFWKIKKFLNIGSEAALRNNTEILNEFVIKLINTRIQQMKNSKGDSVRKGGDILSRFL 283
+ WK+ + LN+G EA L+ ++++EFV +LI R ++ NS R+ DILSRFL
Sbjct: 120 ISPLWKLARLLNVGVEAMLKERIKVVDEFVYRLIRARSDELSNSHDSGSRQ--DILSRFL 177
Query: 284 QVKEYDT----KYLRDIILNFVIAGKDTTGGTLSWFMYMLCKYPAVQEKAAQEVREATNT 339
Q D+ KYLRDIILN VIAGKDTT G L+WF+YM+CK+P VQEK E AT+
Sbjct: 178 QATTSDSVVDYKYLRDIILNIVIAGKDTTAGALAWFLYMVCKHPEVQEKICHEAMVATSA 237
Query: 340 KTVSSCTEFVSSVTDEAIEKMNYVHAVLTETLRLYPALPFDAKICFADDTLPDGYSVKKG 399
+S EF+ S+TD+A+ M+Y+HA LTETLRLYP++P + K CF+DD LP+G++V KG
Sbjct: 238 GDTASVDEFLQSLTDQALNNMHYLHAALTETLRLYPSVPMENKQCFSDDVLPNGFNVSKG 297
Query: 400 DMVSYQPYAMGRMKFIWGDDAEEFRPERWLDENGIFQPECPFKFTAFQAGPRICLGKEFA 459
D+V + PYAMGRM+ +WG DAE FRPERWLDENG+FQ E PFKFTAFQAGPRICLGKEFA
Sbjct: 298 DIVFFIPYAMGRMESLWGKDAEYFRPERWLDENGVFQQESPFKFTAFQAGPRICLGKEFA 357
Query: 460 YRQMKIFSAVLLGCFRFKLNDEKKNVTYKTMITLHIDGGLEIKALYR 506
YRQMKIF+AVLL F KL DEK+ V+Y+T +TL ID GL + A R
Sbjct: 358 YRQMKIFAAVLLRFFVLKLRDEKEIVSYRTTLTLAIDQGLHLTATAR 404
>UniRef100_Q8S7V0 Putative plant cytochrome P-450 protein [Oryza sativa]
Length = 544
Score = 388 bits (997), Expect = e-106
Identities = 215/521 (41%), Positives = 320/521 (61%), Gaps = 37/521 (7%)
Query: 13 LSASLTLLLVQLLLRKLNNKSNSMKKKKYHPVAGTVFNQMMNFNRLHHYMTDLARKYKTY 72
L A ++L +L+ K + ++ K + P+ G Q+ N++R+H ++ + K +T
Sbjct: 25 LIAIFLVVLSWILVHKWSLRNQ--KGPRSWPIIGATVEQLKNYHRMHDWLVEYLSKDRTV 82
Query: 73 RLLNPFRSEVYTSEPSNVEYILKTNFENYGKGLYNYQNLKDLLGDGIFTVDGEKWREQRK 132
+ PF S Y ++P NVE++LKTNF NY KG + LLGDGIF DGE WR+QRK
Sbjct: 83 TVDMPFTSYTYIADPVNVEHVLKTNFTNYPKGEVYRSYMDVLLGDGIFNADGEMWRKQRK 142
Query: 133 ISSHEFSTRMLRDFSTSIFRKNAAKVANIVSEAANSNTKLEIQDIFMKSTLDSIFNVVFG 192
+S EF+++ LRDFST +FR+ + K+++I+S+A + +++Q++FM+ TLDSI V FG
Sbjct: 143 TASFEFASKNLRDFSTVVFREYSLKLSSILSQACKAGRVVDMQELFMRMTLDSICKVGFG 202
Query: 193 TEIDSMCGTSEEGKNFANSFDNASALTLYRYVDVFWKIKKFLNIGSEAALRNNTEILNEF 252
EI ++ E +FA +FD A+ + R++D W++KKFL++GSEA L + +++++F
Sbjct: 203 VEIGTLSPDLPE-NSFAQAFDAANIIVTLRFIDPLWRLKKFLHVGSEALLEQSMKLVDDF 261
Query: 253 VIKLINTR----IQQMKNSKGDSVRKGGDILSRFLQVKEY----------DTKYLRDIIL 298
+I R +Q + K + ++ DILSRF+++ E D K LRD++L
Sbjct: 262 TYSVIRRRKAEILQARASGKQEKIKH--DILSRFIELGEAGGDEGGGSFGDDKSLRDVVL 319
Query: 299 NFVIAGKDTTGGTLSWFMYMLCKYPAVQEKAAQEV--------RE--------ATNTKTV 342
NFVIAG+DTT TLSWF YM +PAV +K +E+ RE A
Sbjct: 320 NFVIAGRDTTATTLSWFTYMAMTHPAVADKLRRELAAFEDERAREEGVALADAAGEASFA 379
Query: 343 SSCTEFVSSVTDEAIEKMNYVHAVLTETLRLYPALPFDAKICFADDTLPDGYSVKKGDMV 402
+ +F S ++ +A+ K+ Y+HA +TETLRLYPA+P D K DD LPDG V+ G MV
Sbjct: 380 ARVAQFASLLSYDAVGKLVYLHACVTETLRLYPAVPQDPKGIVEDDVLPDGTKVRAGGMV 439
Query: 403 SYQPYAMGRMKFIWGDDAEEFRPERWLD-ENGIFQPECPFKFTAFQAGPRICLGKEFAYR 461
+Y PY+MGRM++ WG DA FRPERWL + G F+ PFKFTAFQAGPRICLGK+ AY
Sbjct: 440 TYVPYSMGRMEYNWGPDAASFRPERWLSGDGGAFRNASPFKFTAFQAGPRICLGKDSAYL 499
Query: 462 QMKIFSAVLLGCFRFKLNDEKKNVTYKTMITLHIDGGLEIK 502
QMK+ A+L + F L ++ V Y+ M L + GL+++
Sbjct: 500 QMKMALAILFRFYTFDLVEDHP-VKYRMMTILSMAHGLKVR 539
>UniRef100_Q94KE5 Cytochrome P450-like protein [Zea mays]
Length = 543
Score = 387 bits (994), Expect = e-106
Identities = 211/529 (39%), Positives = 322/529 (59%), Gaps = 37/529 (6%)
Query: 4 LSNPLLFAALSASLTLLLVQLLLRKLNNKSNSMKKKKYHPVAGTVFNQMMNFNRLHHYMT 63
L+ P + AL L ++L +L+++ + + K + PV G Q+ N++R+H ++
Sbjct: 17 LAGPHKYIAL---LLVVLSWILVQRWSLRKQ--KGPRSWPVIGATVEQLRNYHRMHDWLV 71
Query: 64 DLARKYKTYRLLNPFRSEVYTSEPSNVEYILKTNFENYGKGLYNYQNLKDLLGDGIFTVD 123
+++T + PF S Y ++P NVE++LKTNF NY KG+ + LLGDGIF D
Sbjct: 72 GYLSRHRTVTVDMPFTSYTYIADPVNVEHVLKTNFTNYPKGIVYRSYMDVLLGDGIFNAD 131
Query: 124 GEKWREQRKISSHEFSTRMLRDFSTSIFRKNAAKVANIVSEAANSNTKLEIQDIFMKSTL 183
GE WR+QRK +S EF+++ LRDFS +FR+ + K++ I+S+A+ + +++Q+++M+ TL
Sbjct: 132 GELWRKQRKTASFEFASKNLRDFSAIVFREYSLKLSGILSQASKAGKVVDMQELYMRMTL 191
Query: 184 DSIFNVVFGTEIDSMCGTSEEGKNFANSFDNASALTLYRYVDVFWKIKKFLNIGSEAALR 243
DSI V FG EI ++ E +FA +FD A+ + R++D W+IK+F ++GSEA L
Sbjct: 192 DSICKVGFGVEIGTLSPDLPEN-SFAQAFDAANIIITLRFIDPLWRIKRFFHVGSEALLA 250
Query: 244 NNTEILNEFVIKLINTRIQQMKN--SKGDSVRKGGDILSRFLQVKEY--------DTKYL 293
+ ++++EF +I R ++ + G + DILSRF+++ E D K L
Sbjct: 251 QSIKLVDEFTYSVIRRRKAEIVEVRASGKQEKMKHDILSRFIELGEAGDDGGGFGDDKSL 310
Query: 294 RDIILNFVIAGKDTTGGTLSWFMYMLCKYPAVQEKAAQEV-------------------- 333
RD++LNFVIAG+DTT TLSWF +M +P V EK +E+
Sbjct: 311 RDVVLNFVIAGRDTTATTLSWFTHMAMSHPDVAEKLRRELCAFEAERAREEGVTLVLCGG 370
Query: 334 REATNTKTVSSCTEFVSSVTDEAIEKMNYVHAVLTETLRLYPALPFDAKICFADDTLPDG 393
+A + + +F +T +++ K+ Y+HA +TETLRLYPA+P D K DD LPDG
Sbjct: 371 ADADDKAFAARVAQFAGLLTYDSLGKLVYLHACVTETLRLYPAVPQDPKGILEDDVLPDG 430
Query: 394 YSVKKGDMVSYQPYAMGRMKFIWGDDAEEFRPERWLDENGIFQPECPFKFTAFQAGPRIC 453
V+ G MV+Y PY+MGRM++ WG DA FRPERW++E+G F+ PFKFTAFQAGPRIC
Sbjct: 431 TKVRAGGMVTYVPYSMGRMEYNWGPDAASFRPERWINEDGAFRNASPFKFTAFQAGPRIC 490
Query: 454 LGKEFAYRQMKIFSAVLLGCFRFKLNDEKKNVTYKTMITLHIDGGLEIK 502
LGK+ AY QMK+ A+L + F+L E V Y+ M L + GL+++
Sbjct: 491 LGKDSAYLQMKMALAILFRFYSFRLL-EGHPVQYRMMTILSMAHGLKVR 538
>UniRef100_Q9C788 Cytochrome P450, putative [Arabidopsis thaliana]
Length = 524
Score = 386 bits (992), Expect = e-106
Identities = 212/499 (42%), Positives = 306/499 (60%), Gaps = 34/499 (6%)
Query: 40 KYHPVAGTVFNQMMNFNRLHHYMTDLARKYKTYRLLNPFRSEVYTSEPSNVEYILKTNFE 99
K P+ G Q+ NF+R+H ++ + +T + PF + Y ++P NVEY+LKTNF
Sbjct: 29 KTWPLVGAAIEQLTNFDRMHDWLVEYLYNSRTVVVPMPFTTYTYIADPINVEYVLKTNFS 88
Query: 100 NYGKGLYNYQNLKDLLGDGIFTVDGEKWREQRKISSHEFSTRMLRDFSTSIFRKNAAKVA 159
NY KG + ++ LLGDGIF DGE WR+QRK +S EF+++ LRDFST +F++ + K+
Sbjct: 89 NYPKGETYHSYMEVLLGDGIFNSDGELWRKQRKTASFEFASKNLRDFSTVVFKEYSLKLF 148
Query: 160 NIVSEAANSNTKLEIQDIFMKSTLDSIFNVVFGTEIDSMCGTSEEGKNFANSFDNASALT 219
I+S+A+ ++++Q++ M+ TLDSI V FG EI ++ E +FA +FD A+ +
Sbjct: 149 TILSQASFKEQQVDMQELLMRMTLDSICKVGFGVEIGTLAPELPE-NHFAKAFDTANIIV 207
Query: 220 LYRYVDVFWKIKKFLNIGSEAALRNNTEILNEFVIKLINTR--------IQQMKNSKGDS 271
R++D WK+KKFLNIGSEA L + +++N+F +I R I N+ ++
Sbjct: 208 TLRFIDPLWKMKKFLNIGSEALLGKSIKVVNDFTYSVIRRRKAELLEAQISPTNNNNNNN 267
Query: 272 VRKGGDILSRFLQVKE-----YDTKYLRDIILNFVIAGKDTTGGTLSWFMYMLCKYPAVQ 326
+ DILSRF+++ + K LRDI+LNFVIAG+DTT TL+W +YM+ V
Sbjct: 268 NKVKHDILSRFIEISDDPDSKETEKSLRDIVLNFVIAGRDTTATTLTWAIYMIMMNENVA 327
Query: 327 EKAAQEVR-------EATNTKT-----------VSSCTEFVSSVTDEAIEKMNYVHAVLT 368
EK E++ EATNT TEF + +++ K++Y+HAV+T
Sbjct: 328 EKLYSELQELEKESAEATNTSLHQYDTEDFNSFNEKVTEFAGLLNYDSLGKLHYLHAVIT 387
Query: 369 ETLRLYPALPFDAKICFADDTLPDGYSVKKGDMVSYQPYAMGRMKFIWGDDAEEFRPERW 428
ETLRLYPA+P D K DD LP+G VK G MV+Y PY+MGRM++ WG DA F+PERW
Sbjct: 388 ETLRLYPAVPQDPKGVLEDDMLPNGTKVKAGGMVTYVPYSMGRMEYNWGSDAALFKPERW 447
Query: 429 LDENGIFQPECPFKFTAFQAGPRICLGKEFAYRQMKIFSAVLLGCFRFKLNDEKKNVTYK 488
L ++G+FQ PFKFTAFQAGPRICLGK+ AY QMK+ A+L ++F L V Y+
Sbjct: 448 L-KDGVFQNASPFKFTAFQAGPRICLGKDSAYLQMKMAMAILCRFYKFHLVPNHP-VKYR 505
Query: 489 TMITLHIDGGLEIKALYRN 507
M L + GL++ R+
Sbjct: 506 MMTILSMAHGLKVTVSRRS 524
>UniRef100_Q8GWJ5 Hypothetical protein At1g69500/F10D13_15 [Arabidopsis thaliana]
Length = 478
Score = 371 bits (953), Expect = e-101
Identities = 205/481 (42%), Positives = 296/481 (60%), Gaps = 34/481 (7%)
Query: 58 LHHYMTDLARKYKTYRLLNPFRSEVYTSEPSNVEYILKTNFENYGKGLYNYQNLKDLLGD 117
+H ++ + +T + PF + Y ++P NVEY+LKTNF NY KG + ++ LLGD
Sbjct: 1 MHDWLVEYLYNSRTVVVPMPFTTYTYIADPINVEYVLKTNFSNYPKGETYHSYMEVLLGD 60
Query: 118 GIFTVDGEKWREQRKISSHEFSTRMLRDFSTSIFRKNAAKVANIVSEAANSNTKLEIQDI 177
GIF DGE WR+QRK +S EF+++ LRDFST +F++ + K+ I+S+A+ ++++Q++
Sbjct: 61 GIFNSDGELWRKQRKTASFEFASKNLRDFSTVVFKEYSLKLFTILSQASFKEQQVDMQEL 120
Query: 178 FMKSTLDSIFNVVFGTEIDSMCGTSEEGKNFANSFDNASALTLYRYVDVFWKIKKFLNIG 237
M+ TLDSI V FG EI ++ E +FA +FD A+ + R++D WK+KKFLNIG
Sbjct: 121 LMRMTLDSICKVGFGVEIGTLAPELPE-NHFAKAFDTANIIVTLRFIDPLWKMKKFLNIG 179
Query: 238 SEAALRNNTEILNEFVIKLINTR--------IQQMKNSKGDSVRKGGDILSRFLQVKE-- 287
SEA L + +++N+F +I R I N+ ++ + DILSRF+++ +
Sbjct: 180 SEALLGKSIKVVNDFTYSVIRRRKAELLEAQISPTNNNNNNNNKVKHDILSRFIEISDDP 239
Query: 288 ---YDTKYLRDIILNFVIAGKDTTGGTLSWFMYMLCKYPAVQEKAAQEVR-------EAT 337
K LRDI+LNFVIAG+DTT TL+W +YM+ V EK E++ EAT
Sbjct: 240 DSKETEKSLRDIVLNFVIAGRDTTATTLTWAIYMIMMNENVAEKLYSELQELEKESAEAT 299
Query: 338 NTKT-----------VSSCTEFVSSVTDEAIEKMNYVHAVLTETLRLYPALPFDAKICFA 386
NT TEF + +++ K++Y+HAV+TETLRLYPA+P D K
Sbjct: 300 NTSLHQYDTEDFNSFNEKVTEFAGLLNYDSLGKLHYLHAVITETLRLYPAVPQDPKGVLE 359
Query: 387 DDTLPDGYSVKKGDMVSYQPYAMGRMKFIWGDDAEEFRPERWLDENGIFQPECPFKFTAF 446
DD LP+G VK G MV+Y PY+MGRM++ WG DA F+PERWL ++G+FQ PFKFTAF
Sbjct: 360 DDMLPNGTKVKAGGMVTYVPYSMGRMEYNWGSDAALFKPERWL-KDGVFQNASPFKFTAF 418
Query: 447 QAGPRICLGKEFAYRQMKIFSAVLLGCFRFKLNDEKKNVTYKTMITLHIDGGLEIKALYR 506
QAGPRICLGK+ AY QMK+ A+L ++F L V Y+ M L + GL++ R
Sbjct: 419 QAGPRICLGKDSAYLQMKMAMAILCRFYKFHLVPNHP-VKYRMMTILSMAHGLKVTVSRR 477
Query: 507 N 507
+
Sbjct: 478 S 478
>UniRef100_Q9LXX7 Cytochrome P450-like protein [Arabidopsis thaliana]
Length = 499
Score = 321 bits (823), Expect = 3e-86
Identities = 190/489 (38%), Positives = 284/489 (57%), Gaps = 25/489 (5%)
Query: 31 NKSNSMKKKKYHPVAGTVFNQMMNFNRLHHYMTDLARKYKTYRLL--NPFRSE-VYTSEP 87
N S+ K Y P+ G++ + N +R + + + T + P + + V T+ P
Sbjct: 24 NSSSEFGFKSY-PIVGSLPGLVNNRHRFLDWTVETLSRCPTQTAIFRRPGKLQFVMTANP 82
Query: 88 SNVEYILKTNFENYGKGLYNYQNLKDLLGDGIFTVDGEKWREQRKISSHEFSTRMLRDFS 147
+NVEY+LKT FE++ KG L+D LG GIF DGE W +QRK +S+EFST+ LRDF
Sbjct: 83 ANVEYMLKTKFESFPKGERFISILEDFLGRGIFNSDGEMWWKQRKTASYEFSTKSLRDFV 142
Query: 148 TS-IFRKNAAKVANIVSEAANSNTKLEIQDIFMKSTLDSIFNVVFGTEIDSMCGTSEEGK 206
S + + ++ +++EAA + +++QDI + D+I + F + + G
Sbjct: 143 MSNVTVEINTRLVPVLAEAATNGKLIDLQDILERFAFDNICKLAFNVDSACLGDDGAAGV 202
Query: 207 NFANSFDNASALTLYRYVDVF---WKIKKFLNIGSEAALRNNTEILNEFVIKLINTRIQQ 263
NF +F+ A+ + R+ V WKIKK LNIGSE LR + I+++F +++ RI+Q
Sbjct: 203 NFMQAFETAATIISQRFQSVISYSWKIKKKLNIGSERVLRESIMIVHKFADEIVRNRIEQ 262
Query: 264 MKNSKGDSVRKGGDILSRFLQVKEYDT-KYLRDIILNFVIAGKDTTGGTLSWFMYMLCKY 322
K S D+LSRF+ +E ++ + LRDI+++F++AG+DTT LSWF ++L +
Sbjct: 263 GKVSDHKE-----DLLSRFISKEEMNSPEILRDIVISFILAGRDTTSSALSWFFWLLSMH 317
Query: 323 PAVQEKAAQE---VREATNTKTVSSCTEFVSSVTDEAIEKMNYVHAVLTETLRLYPALPF 379
P V++K QE +RE T K + F E ++ MNY+HA +TE+LRLYP +P
Sbjct: 318 PEVKDKILQELNSIRERTG-KRIGEVYGF------EDLKLMNYLHAAITESLRLYPPVPV 370
Query: 380 DAKICFADDTLPDGYSVKKGDMVSYQPYAMGRMKFIWGDDAEEFRPERWLDE-NGIFQPE 438
D C D+ LPDG + K +SY YAMGRM+ IWG D + F PERW+DE NG F+ E
Sbjct: 371 DTMSCAEDNVLPDGTFIGKDWGISYNAYAMGRMESIWGKDCDRFDPERWIDETNGGFRGE 430
Query: 439 CPFKFTAFQAGPRICLGKEFAYRQMKIFSAVLLGCFRFKLNDEKKNVTYKTMITLHIDGG 498
P+KF AF AGPR+CLGKE AY QMK A +L F ++ +K+ +TL I GG
Sbjct: 431 NPYKFPAFHAGPRMCLGKEMAYIQMKSIVAAVLERFVVEVPGKKERPEILMSVTLRIRGG 490
Query: 499 LEIKALYRN 507
L ++ R+
Sbjct: 491 LNVRVQERS 499
>UniRef100_Q8W2N1 Cytochrome P450-dependent fatty acid hydroxylase [Nicotiana
tabacum]
Length = 511
Score = 320 bits (819), Expect = 8e-86
Identities = 190/519 (36%), Positives = 299/519 (57%), Gaps = 41/519 (7%)
Query: 8 LLFAALSASLTLLLVQLLL-----------RKLNNKSNSMKKKKYHPVAGTVFNQMMNFN 56
LL LS SL LV LL L + +NS K + +P+ G+ F+ + N +
Sbjct: 3 LLDLQLSTSLLFCLVPLLFLFFVKFKKTITNTLLSSNNSSKIPRSYPLIGSYFSILANHD 62
Query: 57 RLHHYMTD--LARKYKTYRLLNP--FRSEVYTSEPSNVEYILKTNFENYGKGLYNYQNLK 112
R +++D L T+ L+ P FR+ ++T+ PSNV+++LKTNF+ Y KG +Y LK
Sbjct: 63 RRIKWISDIILTTPNLTFTLIRPLNFRT-IFTANPSNVQHVLKTNFQVYQKGHGSYSTLK 121
Query: 113 DLLGDGIFTVDGEKWREQRKISSHEFSTRMLRDF-STSIFRKNAAKVANIVSEAANSNTK 171
D L +GIF VDG+ W+ QR+++SHEF+TR LR F T + + + ++ I++ AA + T
Sbjct: 122 DFLSNGIFNVDGDIWKYQRQVASHEFNTRSLRKFVETVVDTELSERLIPILATAAANKTV 181
Query: 172 LEIQDIFMKSTLDSIFNVVFGTEIDSMCGTSEEGKNFANSFDNASALTLYRYVDVF---W 228
L+ QDI + D+I + FG + + + E + FA +F++A L+ R++ F W
Sbjct: 182 LDFQDILQRFAFDNICKIAFGYDPGYLLPSLPEAE-FAVAFEDAVRLSTERFIVPFSLIW 240
Query: 229 KIKKFLNIGSEAALRNNTEILNEFVIKLINTRIQQMKNSKGDSVRKGGDILSRFLQVKEY 288
KIK+ LNIGSE LR E + EF +++ + +++ + S D+LSRFL
Sbjct: 241 KIKRALNIGSEKKLRVAVEQVREFAKEIVREKQKELNDK---SSLDSADLLSRFLSTGHS 297
Query: 289 DTKYLRDIILNFVIAGKDTTGGTLSWFMYMLCKYPAVQEKAAQEVREATNTKTVSSCTEF 348
D ++ DI+++F++AG+DTT L+WF +++ K+P V+ + +EV E +
Sbjct: 298 DEDFVTDIVISFILAGRDTTSAALTWFFWLISKHPEVESQIMKEVGEKSE---------- 347
Query: 349 VSSVTDEAIEKMNYVHAVLTETLRLYPALPFDAKICFADDTLPDGYSVKKGDMVSYQPYA 408
S + + ++ M Y HA L E++R YP +P D+K DD LPDG VKKG V+Y PYA
Sbjct: 348 -SLLLYDEVKNMMYTHASLCESMRFYPPVPMDSKEATKDDILPDGTFVKKGTRVTYHPYA 406
Query: 409 MGRMKFIWGDDAEEFRPERWLDE-----NGIFQPECPFKFTAFQAGPRICLGKEFAYRQM 463
MGR++ +WG+D EF+PERWLD+ N F P+ + + FQAGPRICLGKE A+ QM
Sbjct: 407 MGRVEKVWGEDWAEFKPERWLDKDEVTGNWTFVPKDAYTYPVFQAGPRICLGKEMAFLQM 466
Query: 464 KIFSAVLLGCFR-FKLNDEKKNVTYKTMITLHIDGGLEI 501
K A +L F+ + ++ + + +T + GG +
Sbjct: 467 KRVVAGVLRRFKVVPVVEQGVEPVFISYLTAKMKGGFPV 505
>UniRef100_P98188 Cytochrome P450 94A2 [Vicia sativa]
Length = 513
Score = 316 bits (809), Expect = 1e-84
Identities = 172/447 (38%), Positives = 267/447 (59%), Gaps = 24/447 (5%)
Query: 40 KYHPVAGTVFNQMMNFNRLHHYMTDLARKY--KTYRLLNPFRS-EVYTSEPSNVEYILKT 96
K +P+ G+ F+ + NF+R + +D+ + T+ L PF + +V+T++P+ V++IL+T
Sbjct: 44 KSYPIFGSAFSLLANFHRRIQWTSDILQTIPSSTFVLHRPFGARQVFTAQPAVVQHILRT 103
Query: 97 NFENYGKGLYNYQNLKDLLGDGIFTVDGEKWREQRKISSHEFSTRMLRDF-STSIFRKNA 155
NF YGKGL YQ++ D LGDGIF DGE W+ QR+ISSHEF+TR LR F T + + +
Sbjct: 104 NFTCYGKGLTFYQSINDFLGDGIFNADGESWKFQRQISSHEFNTRSLRKFVETVVDVELS 163
Query: 156 AKVANIVSEAANSNTKLEIQDIFMKSTLDSIFNVVFGTEIDSMCGTSEEGKNFANSFDNA 215
++ ++S+A+NS T L+ QDI + T D+I + FG + + + + E FA +FD +
Sbjct: 164 DRLVPVLSQASNSQTTLDFQDILQRLTFDNICMIAFGYDPEYLLPSLPEIP-FAKAFDES 222
Query: 216 SALTLYRY---VDVFWKIKKFLNIGSEAALRNNTEILNEFVIKLINTRIQQMKNSKGDSV 272
S L++ R + + WK+K+FLNIG E L+ + K++ + +++K S
Sbjct: 223 SQLSIERLNALIPLLWKVKRFLNIGVERQLKEAVAEVRGLATKIVKNKKKELKEKALQSE 282
Query: 273 RKGGDILSRFLQVKEYDTKYLRDIILNFVIAGKDTTGGTLSWFMYMLCKYPAVQEKAAQE 332
+ D+LSRFL D ++ D++++ ++AG+DTT L+WF ++L K+ V+ + +E
Sbjct: 283 SESVDLLSRFLSSGHSDESFVTDMVISIILAGRDTTSAALTWFFWLLSKHSHVENEILKE 342
Query: 333 VREATNTKTVSSCTEFVSSVTDEAIEKMNYVHAVLTETLRLYPALPFDAKICFADDTLPD 392
+ + T V + ++ M Y HA L E++RLYP LP D K+ DD LPD
Sbjct: 343 ITGKSET------------VGYDEVKDMVYTHAALCESMRLYPPLPVDTKVAVHDDVLPD 390
Query: 393 GYSVKKGDMVSYQPYAMGRMKFIWGDDAEEFRPERWLDENGI----FQPECPFKFTAFQA 448
G VKKG V+Y YAMGR + IWG D EFRPERWL + + F + + FQA
Sbjct: 391 GTLVKKGWRVTYHIYAMGRSEKIWGPDWAEFRPERWLSRDEVGKWSFVGIDYYSYPVFQA 450
Query: 449 GPRICLGKEFAYRQMKIFSAVLLGCFR 475
GPR+C+GKE A+ QMK A ++G FR
Sbjct: 451 GPRVCIGKEMAFLQMKRVVAGIMGRFR 477
>UniRef100_Q8LRA0 Putative cytochrome P450-dependent fatty acid hydroxylase [Oryza
sativa]
Length = 511
Score = 310 bits (794), Expect = 6e-83
Identities = 185/514 (35%), Positives = 279/514 (53%), Gaps = 25/514 (4%)
Query: 11 AALSASLTLLLVQLLLRKLNNKSNSMKKKKYH-----PVAGTVFNQMMNFNRLHHYMTDL 65
++ SASL L+L+ L+ L + KK + H PV GT+ + + N +R + T +
Sbjct: 4 SSTSASLLLILLLTLVYFLYLHQDPKKKPRTHGLKSYPVVGTLPHFINNKDRFLEWSTGV 63
Query: 66 ARKYKTYRLLNP---FRSEVYTSEPSNVEYILKTNFENYGKGLYNYQNLKDLLGDGIFTV 122
++ T+ + V T+ P+NVE+ILK NF NY KG L+D LG GIF
Sbjct: 64 MKRSPTHTMSFKELGLTGGVITANPANVEHILKANFGNYPKGELAVSLLEDFLGHGIFNS 123
Query: 123 DGEKWREQRKISSHEFSTRMLRDFSTSIFRKNAA-KVANIVSEAANSNTKLEIQDIFMKS 181
DGE+W QRK +S+EF+ R LR+F R ++ ++ A L++QD+ +
Sbjct: 124 DGEQWLWQRKAASYEFNKRSLRNFVVDTVRFEVVERLLPLLEYAGRHGRTLDVQDVLERF 183
Query: 182 TLDSIFNVVFGTEIDSMCGTSE-----EGKNFANSFDNASALTLYRY---VDVFWKIKKF 233
D+I V F + D C T E + F +F++A L R+ W+IKK
Sbjct: 184 AFDNICRVAF--DEDPACLTEESMAAPQSAEFMRAFNDAQNAILDRFNSPAKSLWRIKKL 241
Query: 234 LNIGSEAALRNNTEILNEFVIKLINTRIQQMKNSKGDSVRKGGDILSRFLQVKEYDTKYL 293
N+ E +R++ ++ + +++ R + + + + D LSRF E+ + L
Sbjct: 242 FNMEPERRMRDSLATIHGYAERIVRER----RERREARLERRDDFLSRFAASGEHSDESL 297
Query: 294 RDIILNFVIAGKDTTGGTLSWFMYMLCKYPAVQEKAAQEVREATNTKTVSSCTEFVSSVT 353
RD++ NF++AG+DTT L+WF ++L P V++K +E+R + S T +
Sbjct: 298 RDVVTNFILAGRDTTSSALTWFFWLLSGRPDVEDKIVREIRAVRQSSAGSEGTRGATFSL 357
Query: 354 DEAIEKMNYVHAVLTETLRLYPALPFDAKICFADDTLPDGYSVKKGDMVSYQPYAMGRMK 413
DE + M Y+HA +TE++RLYP +PFD C ++ LPDG KG +V+Y YAMGR++
Sbjct: 358 DE-LRDMQYLHAAITESMRLYPPVPFDTHSCKEEEFLPDGTFAGKGWLVTYCAYAMGRVE 416
Query: 414 FIWGDDAEEFRPERWLDENGIFQPECPFKFTAFQAGPRICLGKEFAYRQMKIFSAVLLGC 473
IWG D EEFRPERWLDE G F+PE FK+ F AGPR+CLGKE AY QMK A +L
Sbjct: 417 DIWGADCEEFRPERWLDEAGAFRPESTFKYPVFHAGPRMCLGKEMAYIQMKSIVACVLEQ 476
Query: 474 FRFK-LNDEKKNVTYKTMITLHIDGGLEIKALYR 506
F + D K + +TL ++GGL +K R
Sbjct: 477 FSLRYAGDAKGHPGLVVALTLRMEGGLPMKVTIR 510
>UniRef100_Q9LNL3 F12K21.15 [Arabidopsis thaliana]
Length = 498
Score = 309 bits (792), Expect = 1e-82
Identities = 182/479 (37%), Positives = 278/479 (57%), Gaps = 23/479 (4%)
Query: 32 KSNSMKKKKYHPVAGTVFNQMMNFNRLHHYMTDLARKYKTYRLL--NPFRSE-VYTSEPS 88
KS+S K +P+ G+ + N +R + + + T + P + + + T+ PS
Sbjct: 24 KSSSEFGFKSYPIVGSFPGLVNNRHRFLDWTVETLSRCPTQTAIFRRPGKQQLIMTANPS 83
Query: 89 NVEYILKTNFENYGKGLYNYQNLKDLLGDGIFTVDGEKWREQRKISSHEFSTRMLRDFST 148
NVEY+LKT FE++ KG L+D LG GIF DG+ W +QRK +S+EFST+ LRDF
Sbjct: 84 NVEYMLKTKFESFPKGQQFTSVLEDFLGHGIFNSDGDMWWKQRKTASYEFSTKSLRDFVM 143
Query: 149 S-IFRKNAAKVANIVSEAANSNTKLEIQDIFMKSTLDSIFNVVFGTEIDSMCGTSEEGKN 207
S + + ++ ++ EAA + +++QDI + D+I + F + + G N
Sbjct: 144 SNVTVEINTRLVPVLVEAATTGKLIDLQDILERFAFDNICKLAFNVDCACLGHDGAVGVN 203
Query: 208 FANSFDNASALTLYRYVDVF---WKIKKFLNIGSEAALRNNTEILNEFVIKLINTRIQQM 264
F +F+ A+ + R+ V W+IKK LNIGSE LR + +++F +++ RI Q
Sbjct: 204 FMRAFETAATIISQRFRSVASCAWRIKKKLNIGSERVLRESIATVHKFADEIVRNRIDQG 263
Query: 265 KNSKGDSVRKGGDILSRFLQVKEYDT-KYLRDIILNFVIAGKDTTGGTLSWFMYMLCKYP 323
++S D+LSRF+ +E ++ + LRDI+++F++AG+DTT LSWF ++L +P
Sbjct: 264 RSSDHKE-----DLLSRFISKEEMNSPEILRDIVISFILAGRDTTSSALSWFFWLLSMHP 318
Query: 324 AVQEKAAQEVRE--ATNTKTVSSCTEFVSSVTDEAIEKMNYVHAVLTETLRLYPALPFDA 381
V++K QE+ A K + F E ++ MNY+HA +TE+LRLYP +P D
Sbjct: 319 EVEDKILQELNSIRARTGKRIGEVYGF------EHLKMMNYLHAAITESLRLYPPVPVDI 372
Query: 382 KICFADDTLPDGYSVKKGDMVSYQPYAMGRMKFIWGDDAEEFRPERWLDE-NGIFQPECP 440
K C D+ LPDG V KG ++Y +AMGRM+ IWG D + F PERW+DE NG F+ E P
Sbjct: 373 KSCAEDNVLPDGTFVGKGWAITYNIFAMGRMESIWGKDCDRFDPERWIDETNGCFRGEDP 432
Query: 441 FKFTAFQAGPRICLGKEFAYRQMKIFSAVLLGCFRFKLNDEKKNVTYKTMITLHIDGGL 499
KF AF AGPR+C+GK+ AY QMK A +L F ++ +++ +M TL I GGL
Sbjct: 433 SKFPAFHAGPRMCVGKDMAYIQMKSIVAAVLERFVVEVPGKERPEILLSM-TLRIKGGL 490
>UniRef100_Q8W2N4 Cytochrome P450-dependent fatty acid hydroxylase [Nicotiana
tabacum]
Length = 510
Score = 308 bits (789), Expect = 2e-82
Identities = 180/484 (37%), Positives = 281/484 (57%), Gaps = 37/484 (7%)
Query: 14 SASLTLLLVQLLLR---KLNN-------KSNSMKKKKYHPVAGTVFNQMMNFNRLHHYMT 63
S SL LV LL K N SNS K + +P+ G+ F+ + N ++ +++
Sbjct: 9 STSLLFCLVPLLFLFFIKFNKTITNTLLSSNSSKIPRSYPLIGSYFSILANHDQRIKWIS 68
Query: 64 D--LARKYKTYRLLNPFRSE-VYTSEPSNVEYILKTNFENYGKGLYNYQNLKDLLGDGIF 120
D L+ T+ L+ P ++T+ PSNV+++LKTNF+ Y KG + LKD L +GIF
Sbjct: 69 DIILSTPNLTFTLIRPLNFHTIFTANPSNVQHMLKTNFQVYQKGHNSNTTLKDFLSNGIF 128
Query: 121 TVDGEKWREQRKISSHEFSTRMLRDF-STSIFRKNAAKVANIVSEAANSNTKLEIQDIFM 179
VDG+ W+ QR+++SHEF+TR LR F T + + + ++ I++ AA + T L+ QDI
Sbjct: 129 NVDGDIWKYQRQVASHEFNTRSLRKFVETVVDTELSERLIPILATAAANKTVLDFQDILQ 188
Query: 180 KSTLDSIFNVVFGTEIDSMCGTSEEGKNFANSFDNASALTLYRYVDVF---WKIKKFLNI 236
+ D+I + FG + + + E + FA +F++A L+ R++ F WK+K+ LNI
Sbjct: 189 RFAFDNICKIAFGYDPGYLLPSLPEAE-FAVAFEDAVRLSTERFILPFPLIWKMKRALNI 247
Query: 237 GSEAALRNNTEILNEFVIKLINTRIQQMKNSKGDSVRKGGDILSRFLQVKEYDTKYLRDI 296
GSE LR E + EF +++ + +++K+ S D+LSRFL D ++ DI
Sbjct: 248 GSEKKLRFAVEQVREFAKEIVREKQRELKDK---SSLDSADLLSRFLSTGHSDENFVTDI 304
Query: 297 ILNFVIAGKDTTGGTLSWFMYMLCKYPAVQEKAAQEVREATNTKTVSSCTEFVSSVTDEA 356
+++F++AG+DTT L+WF +++ K+P V+ + +E+ E + S + +
Sbjct: 305 VISFILAGRDTTSAALTWFFWLISKHPEVESQILKEIGEKSE-----------SLLLYDE 353
Query: 357 IEKMNYVHAVLTETLRLYPALPFDAKICFADDTLPDGYSVKKGDMVSYQPYAMGRMKFIW 416
++ M Y HA L E++R YP +P D K DD LPDG VKKG+ V+Y PYAMGR++ +W
Sbjct: 354 VKNMIYTHASLCESMRFYPPVPMDTKEATKDDILPDGTFVKKGNRVTYHPYAMGRVEKVW 413
Query: 417 GDDAEEFRPERWLDE-----NGIFQPECPFKFTAFQAGPRICLGKEFAYRQMKIFSAVLL 471
G D EFRPERWLD+ N F + + + FQAGPR+CLGKE A+ QMK A +L
Sbjct: 414 GKDWAEFRPERWLDKDEVTGNWTFVSKDAYTYPVFQAGPRVCLGKEMAFLQMKRVVAGVL 473
Query: 472 GCFR 475
F+
Sbjct: 474 RRFK 477
>UniRef100_Q8LRA4 Putative cytochrome P450-dependent fatty acid hydroxylase [Oryza
sativa]
Length = 511
Score = 307 bits (786), Expect = 5e-82
Identities = 194/518 (37%), Positives = 281/518 (53%), Gaps = 37/518 (7%)
Query: 13 LSASLTLLLVQL----LLRKLNNKSNSMKKKKYH-----PVAGTVFNQMMNFNRLHHYMT 63
+SASL L+L+ L LL L + KK + H PV GT+ + + N +R + T
Sbjct: 6 ISASLLLILILLAFLPLLYFLYMHQDPKKKPRIHGLKSYPVVGTLPHIIKNKHRFLKWST 65
Query: 64 DLARKYKT----YRLLNPFRSEVYTSEPSNVEYILKTNFENYGKGLYNYQNLKDLLGDGI 119
+ + T Y+ L V T+ P+NVE+ILKTNF+NY KG L+D LG GI
Sbjct: 66 SIMKCSPTNTMSYKALG-LTGGVITANPANVEHILKTNFDNYPKGKLTVSMLEDFLGHGI 124
Query: 120 FTVDGEKWREQRKISSHEFSTRMLRDFSTSIFRKNAAK-VANIVSEAANSNTKLEIQDIF 178
F DGE+W QRK +S+EF+ R LR+F R K + ++ +A L++QD+
Sbjct: 125 FNSDGEQWLWQRKAASYEFNKRSLRNFVVDTVRFEIVKRLLPLLEQAGLDGRTLDLQDVL 184
Query: 179 MKSTLDSIFNVVFGTEIDSMCGTSE-----EGKNFANSFDNASALTLYRY---VDVFWKI 230
+ D+I V FG D C T E + F +F++A L R+ W++
Sbjct: 185 ERFAFDNICLVAFGE--DPACLTKERMAAPQSAEFMRAFNDAQNAILARFNSPAKSLWRV 242
Query: 231 KKFLNIGSEAALRNNTEILNEFVIKLINTRIQQMKNSKGDS-VRKGGDILSRFLQVKEYD 289
KK N+ E +R ++ F +++ R +G++ + +G D LSRF E+
Sbjct: 243 KKLFNMEPERRMREALATIHGFAERIVRER-----RERGEAGLARGDDFLSRFAASGEHS 297
Query: 290 TKYLRDIILNFVIAGKDTTGGTLSWFMYMLCKYPAVQEKAAQEVREATNTKTVSSCTEFV 349
+ LRD++ NFV+AG+DTT L+WF +++ P V+++ +E+R + S T+
Sbjct: 298 DESLRDVVTNFVLAGRDTTSSALTWFFWIVSGRPDVEDRVVREIRAV---RASSGSTDAT 354
Query: 350 SSVTDEAIEKMNYVHAVLTETLRLYPALPFDAKICFADDTLPDGYSVKKGDMVSYQPYAM 409
S DE EK +Y+HA +TE++RLYP + D C DD LPDG V KG +V Y YAM
Sbjct: 355 FSF-DELREK-HYLHAAITESMRLYPPVAIDTHSCKEDDFLPDGTFVGKGWLVMYSAYAM 412
Query: 410 GRMKFIWGDDAEEFRPERWLDENGIFQPECPFKFTAFQAGPRICLGKEFAYRQMKIFSAV 469
GRM+ IWG D EE+RPERWLDE G F+PE FK+ F AGPRIC+GKE AY QMK A
Sbjct: 413 GRMEGIWGADCEEYRPERWLDEAGAFRPESTFKYPVFNAGPRICIGKEMAYIQMKSIVAC 472
Query: 470 LLGCFRFK-LNDEKKNVTYKTMITLHIDGGLEIKALYR 506
+L F + +D + +TL + GL +K R
Sbjct: 473 VLEKFSLRYASDANERPRSVLSLTLRMKWGLPMKVTIR 510
>UniRef100_O81117 Cytochrome P450 94A1 [Vicia sativa]
Length = 514
Score = 301 bits (771), Expect = 3e-80
Identities = 174/488 (35%), Positives = 277/488 (56%), Gaps = 41/488 (8%)
Query: 7 PLLFAALSASLTLLLVQLLLRKLNNK-------SNSMKKKKYHPVAGTVFNQMMNFNRLH 59
PLL L ++ L K NNK +N + K +P+ G+ + N +R
Sbjct: 15 PLLLLILPTTI------FFLTKPNNKVSSTSTNNNIITLPKSYPLIGSYLSFRKNLHRRI 68
Query: 60 HYMTDLAR--KYKTYRLLNPF-RSEVYTSEPSNVEYILKTNFENYGKGLYNYQNLKDLLG 116
+++D+ + T++L + ++ T PS V++ILK F NY KG L D LG
Sbjct: 69 QWLSDIVQISPSATFQLDGTLGKRQIITGNPSTVQHILKNQFSNYQKGTTFTNTLSDFLG 128
Query: 117 DGIFTVDGEKWREQRKISSHEFSTRMLRDFSTSIFRKNAA-KVANIVSEAANSNTKLEIQ 175
GIF +G W+ QR+++SHEF+T+ +R+F I ++ I++ + +N L+ Q
Sbjct: 129 TGIFNTNGPNWKFQRQVASHEFNTKSIRNFVEHIVDTELTNRLIPILTSSTQTNNILDFQ 188
Query: 176 DIFMKSTLDSIFNVVFGTEIDSMCGTSEEGKNFANSFDNASALTLYRY---VDVFWKIKK 232
DI + T D+I N+ FG + + + ++ K FA ++++A+ ++ R+ + + WKIKK
Sbjct: 189 DILQRFTFDNICNIAFGYDPEYLTPSTNRSK-FAEAYEDATEISSKRFRLPLPIIWKIKK 247
Query: 233 FLNIGSEAALRNNTEILNEFVIKLINTRIQQMKNSKGDSVRKGGDILSRFLQVKEYDTKY 292
+ NIGSE L+ + F KL+ + ++++ S + D+LSRFL D +
Sbjct: 248 YFNIGSEKRLKEAVTEVRSFAKKLVREKKRELEEK---SSLETEDMLSRFLSSGHSDEDF 304
Query: 293 LRDIILNFVIAGKDTTGGTLSWFMYMLCKYPAVQEKAAQEVREATNTKTVSSCTEFVSSV 352
+ DI+++F++AGKDTT L+WF ++L K P V+E+ E+ + + V
Sbjct: 305 VADIVISFILAGKDTTSAALTWFFWLLWKNPRVEEEIVNELSKKSELM-----------V 353
Query: 353 TDEAIEKMNYVHAVLTETLRLYPALPFDAKICFADDTLPDGYSVKKGDMVSYQPYAMGRM 412
DE +++M Y HA L+E++RLYP +P D+K DD LPDG+ VKKG +V+Y YAMGRM
Sbjct: 354 YDE-VKEMVYTHAALSESMRLYPPVPMDSKEAVNDDVLPDGWVVKKGTIVTYHVYAMGRM 412
Query: 413 KFIWGDDAEEFRPERWLDE---NG--IFQPECPFKFTAFQAGPRICLGKEFAYRQMKIFS 467
K +WGDD EFRPERWL++ NG +F + + FQAGPR+CLGKE A+ QMK
Sbjct: 413 KSLWGDDWAEFRPERWLEKDEVNGKWVFVGRDSYSYPVFQAGPRVCLGKEMAFMQMKRIV 472
Query: 468 AVLLGCFR 475
A ++G F+
Sbjct: 473 AGIVGKFK 480
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.321 0.136 0.395
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 830,224,543
Number of Sequences: 2790947
Number of extensions: 35568999
Number of successful extensions: 98471
Number of sequences better than 10.0: 4160
Number of HSP's better than 10.0 without gapping: 1749
Number of HSP's successfully gapped in prelim test: 2411
Number of HSP's that attempted gapping in prelim test: 88364
Number of HSP's gapped (non-prelim): 4840
length of query: 507
length of database: 848,049,833
effective HSP length: 132
effective length of query: 375
effective length of database: 479,644,829
effective search space: 179866810875
effective search space used: 179866810875
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 77 (34.3 bits)
Medicago: description of AC146342.5


